Long Noncoding RNA PVT1 Is Regulated by Bromodomain Protein BRD4 in Multiple Myeloma and Is Associated with Disease Progression
Abstract
1. Introduction
2. Results
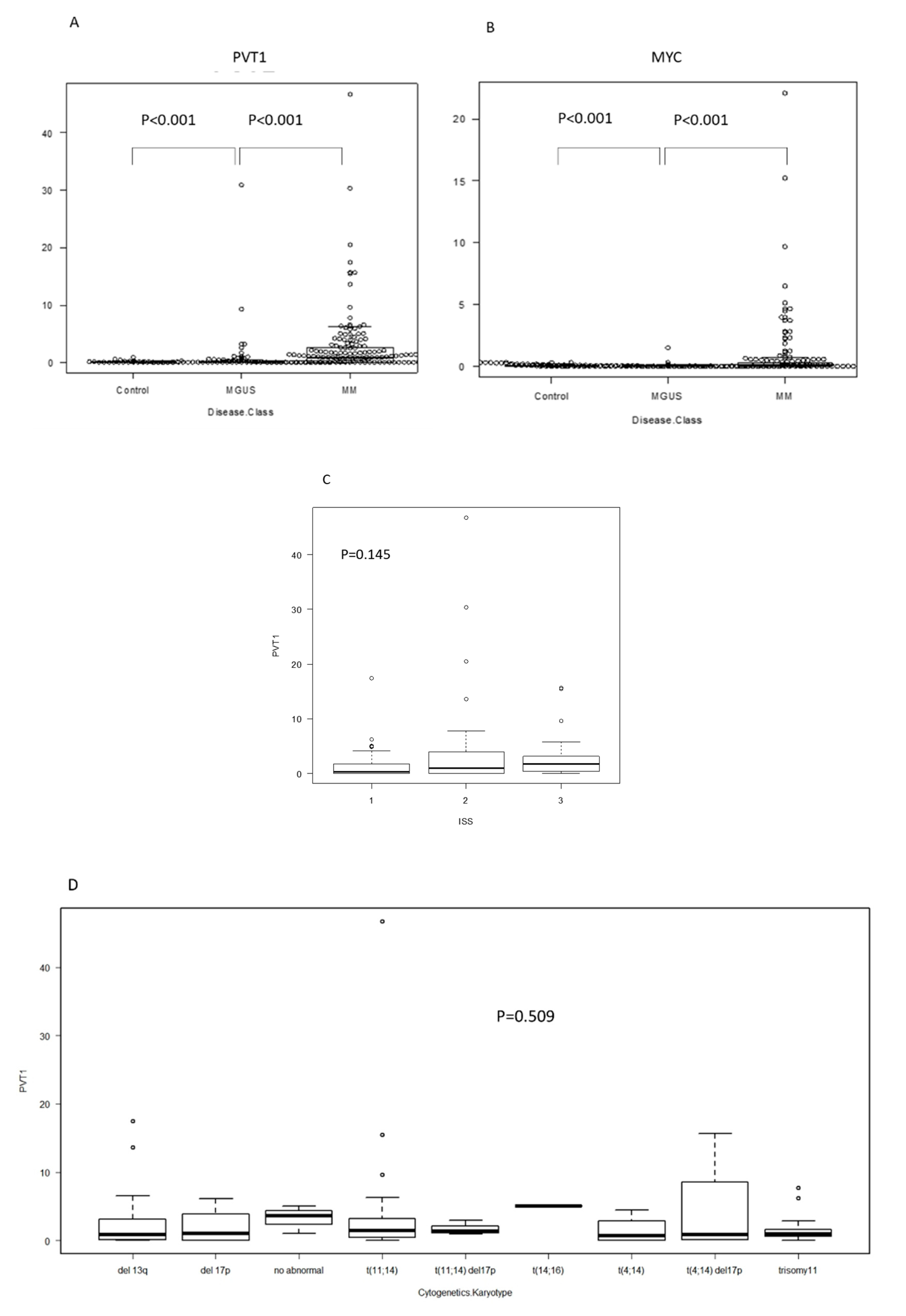
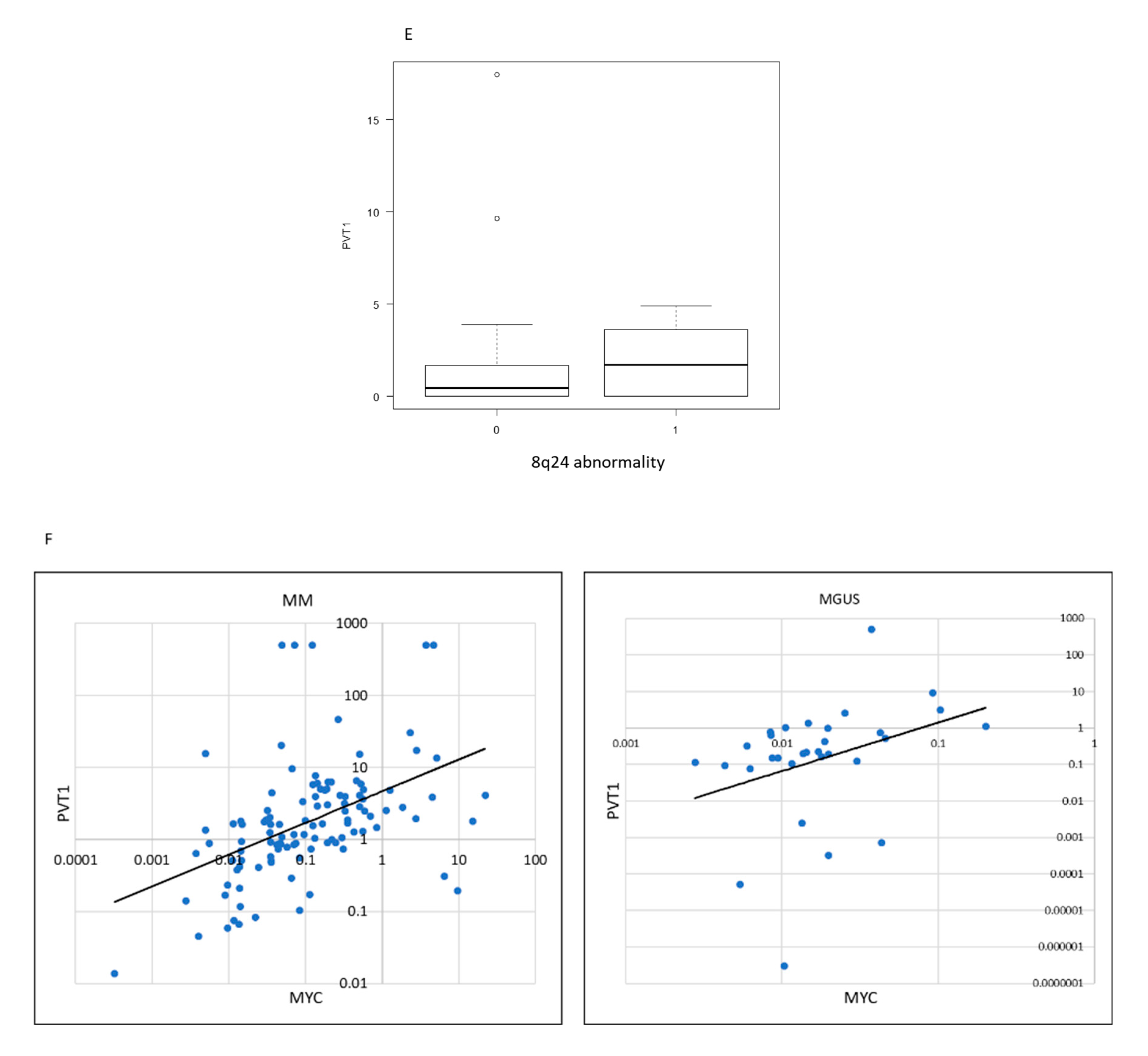
2.1. PVT1 and MYC Expression in Plasma Cells of MM Is Higher than in MGUS or Control
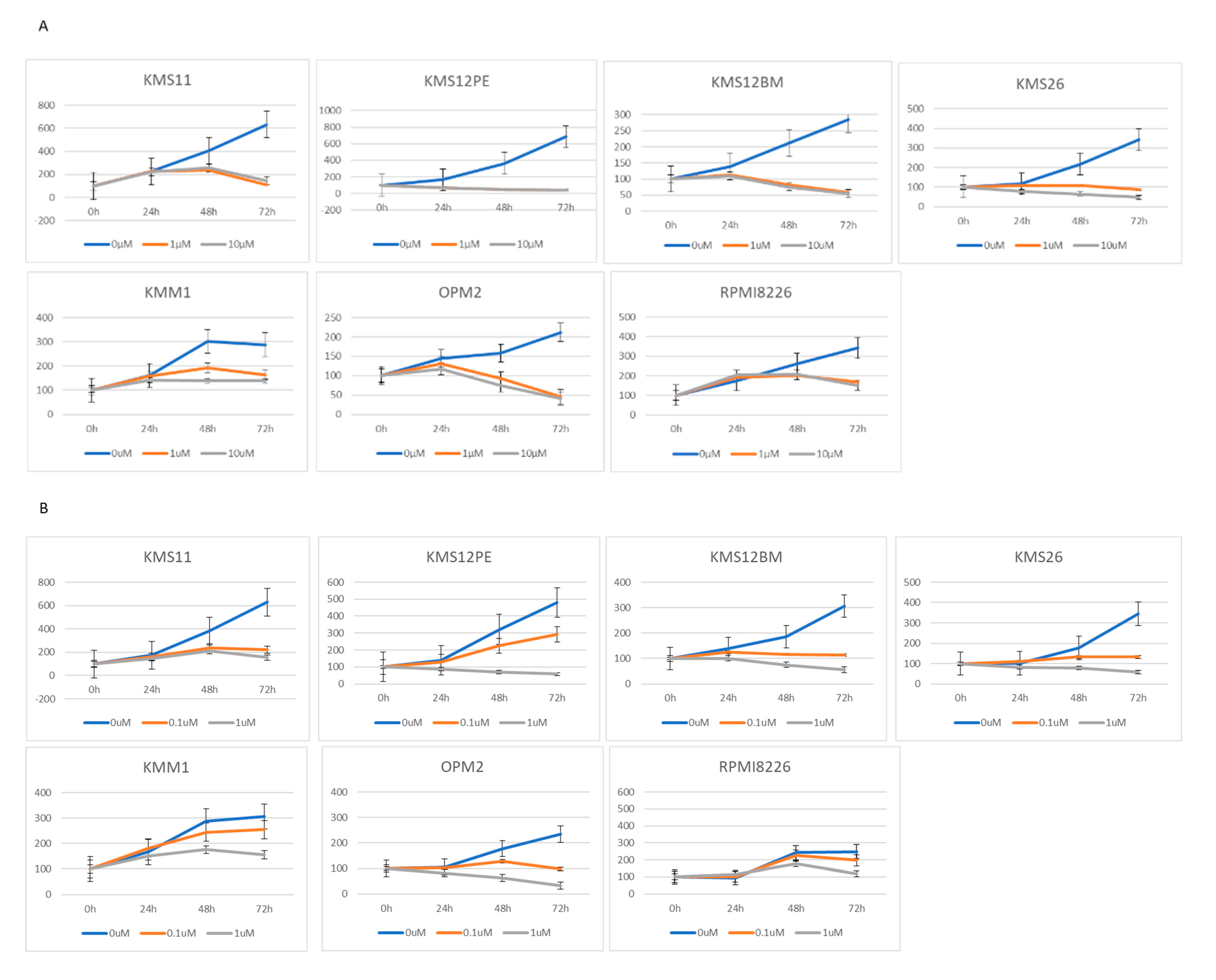
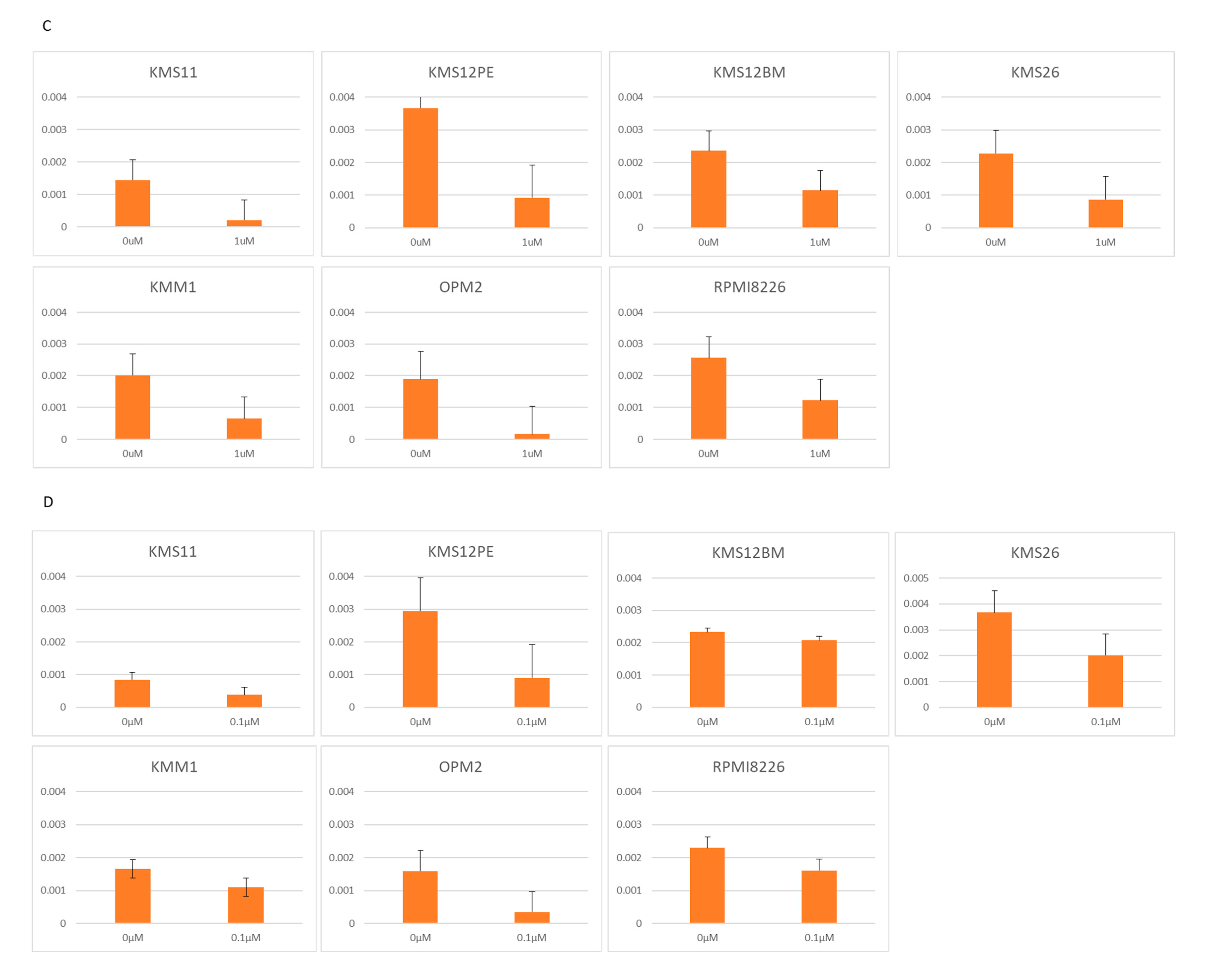
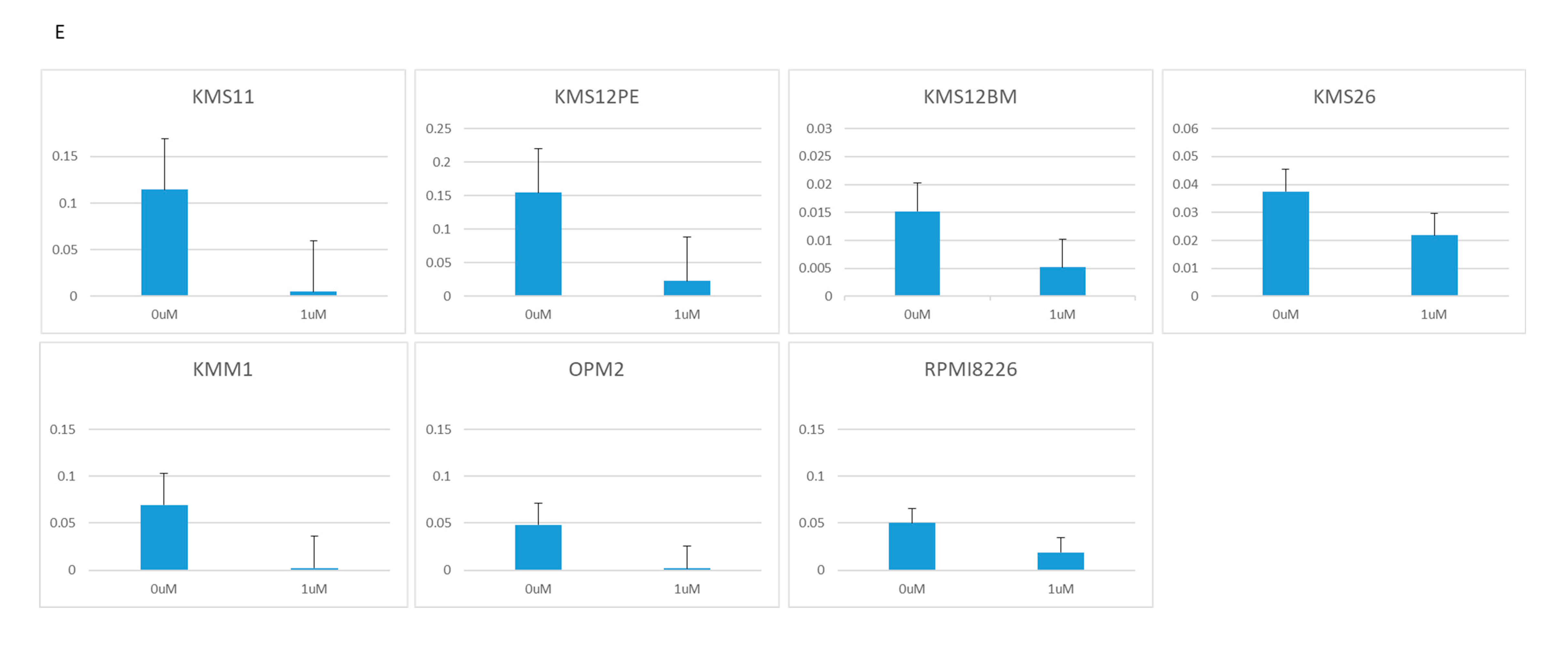
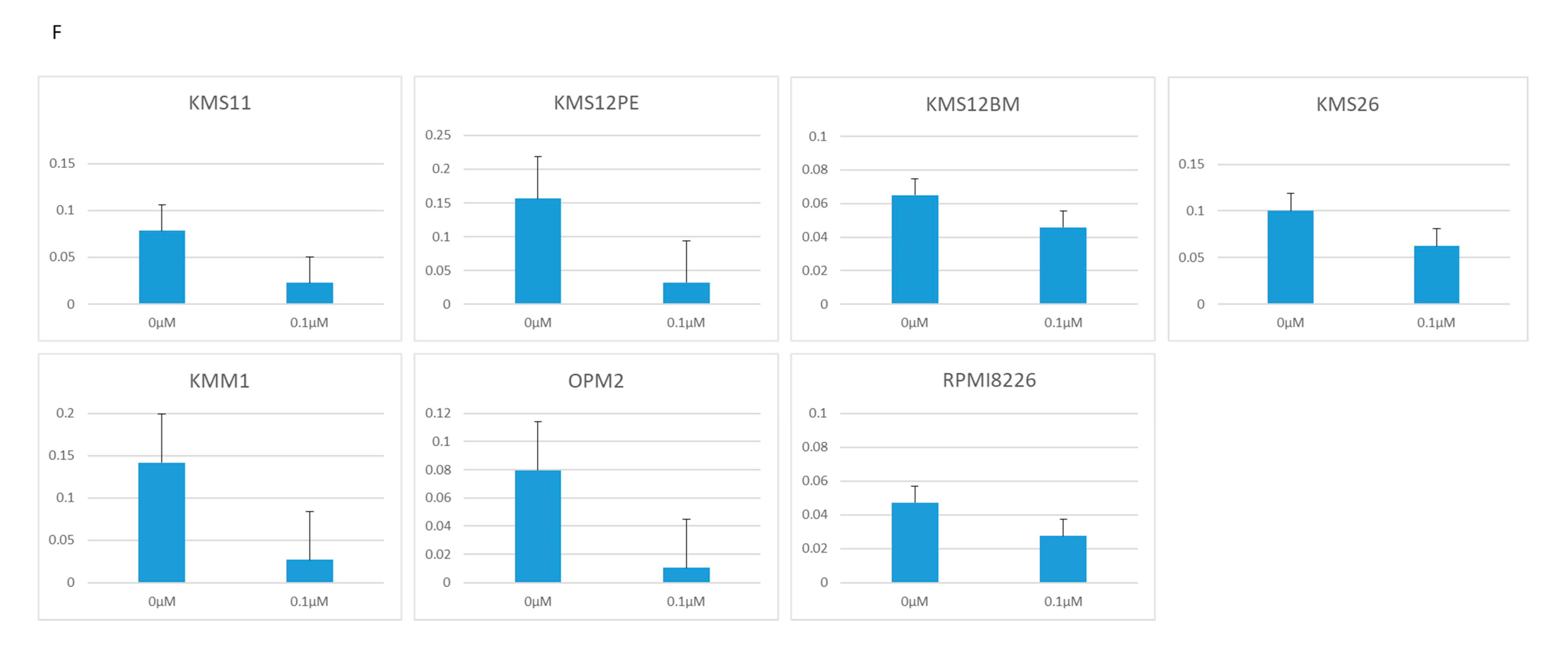
2.2. BRD4 Inhibitors Inhibit MM Cell Proliferation and Downregulate PVT1 and MYC Expression
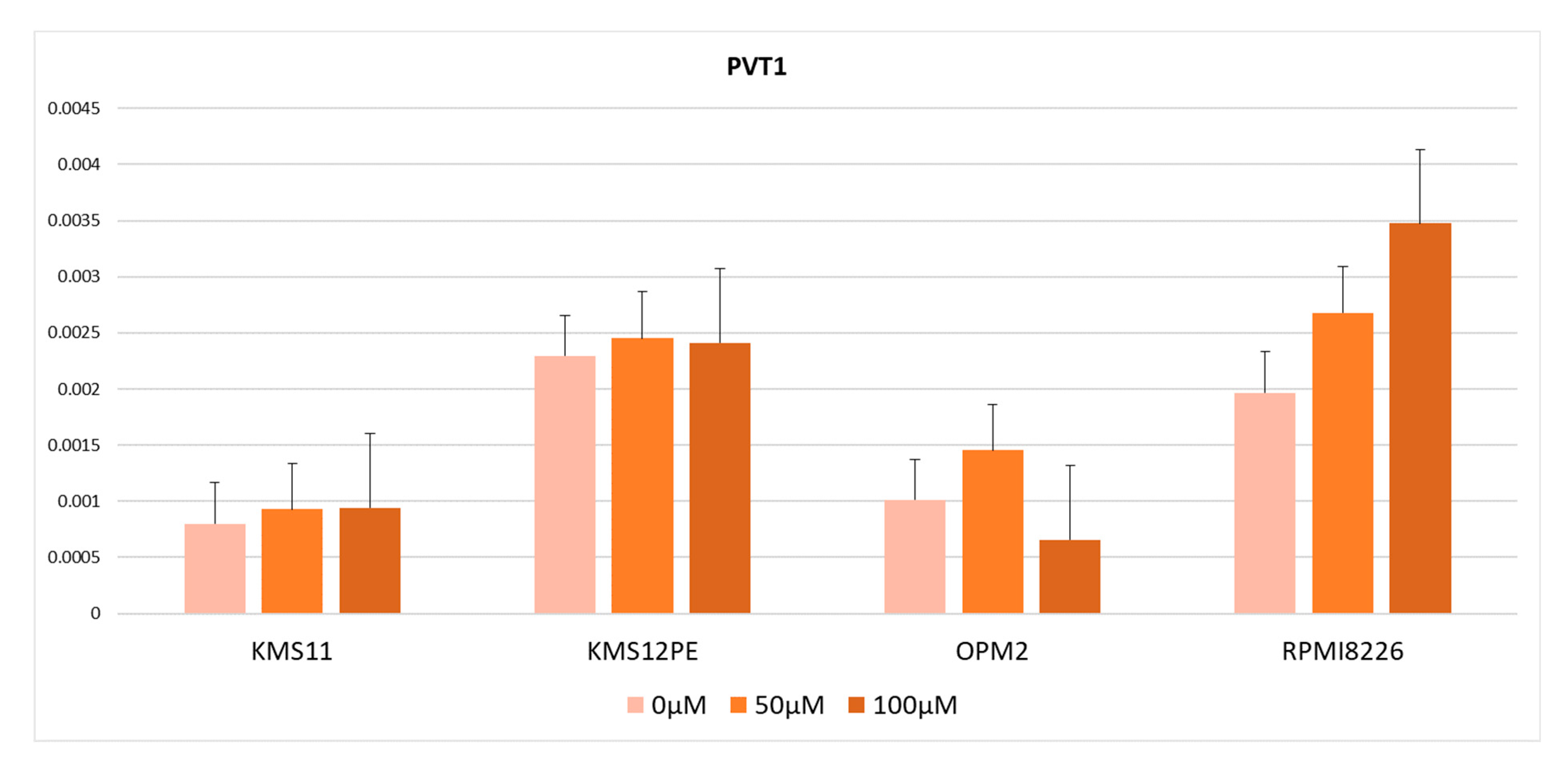
2.3. MYC Inhibitor Did Not Reduce PVT1 Expression
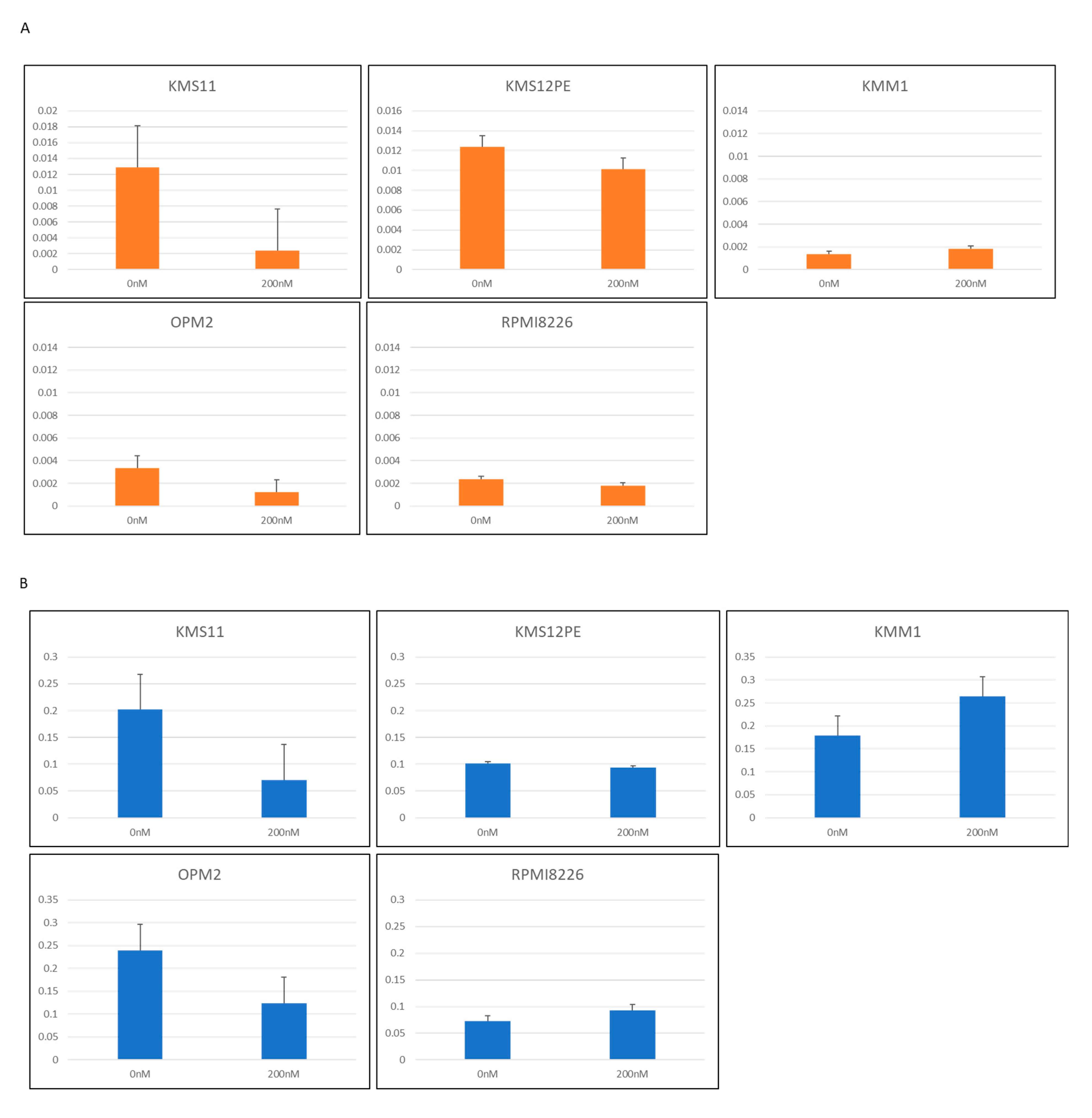
2.4. PVT1 Downregulation by Locked Anti-Sense Nucleotide Reduced MYC Expression
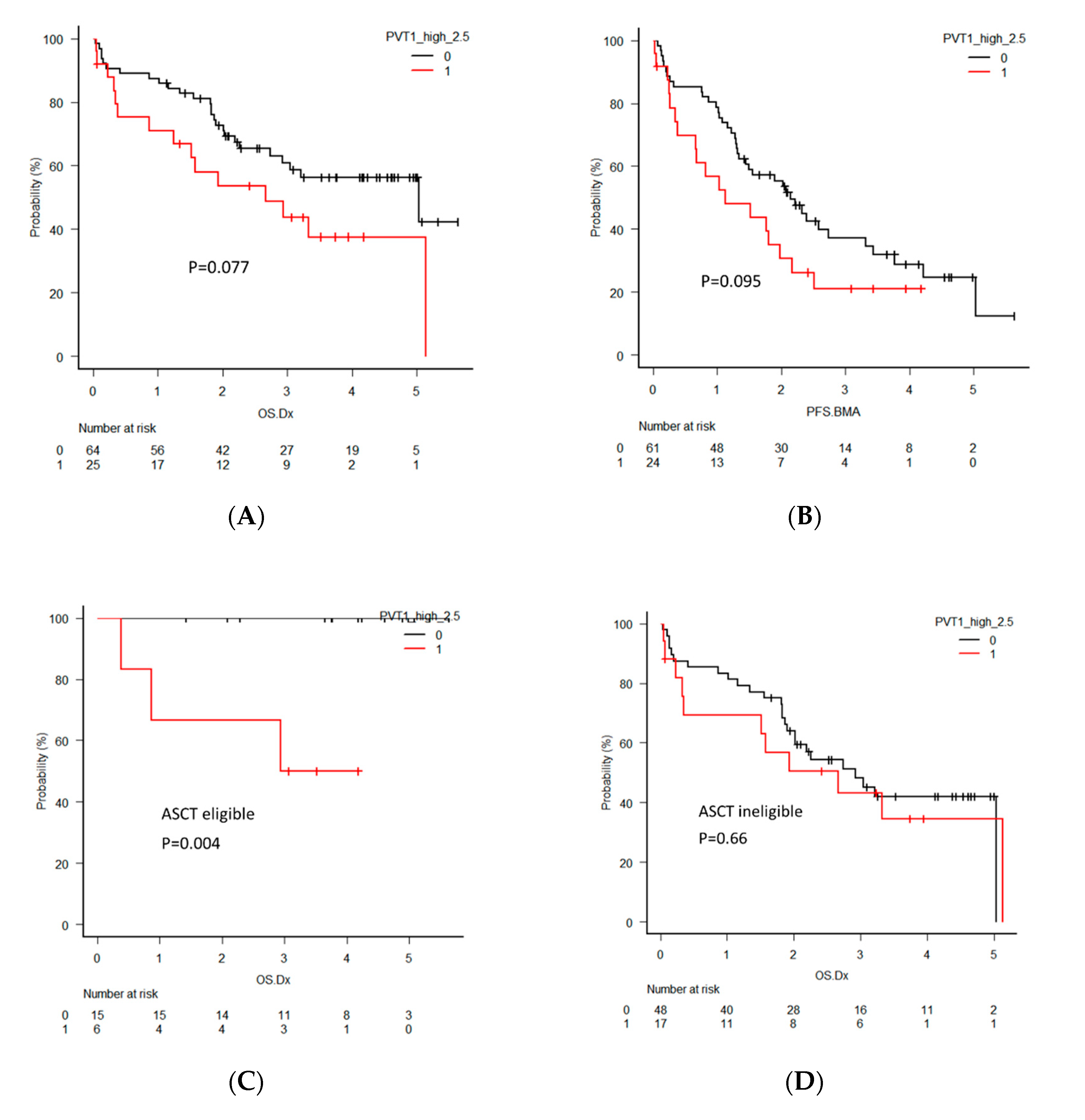
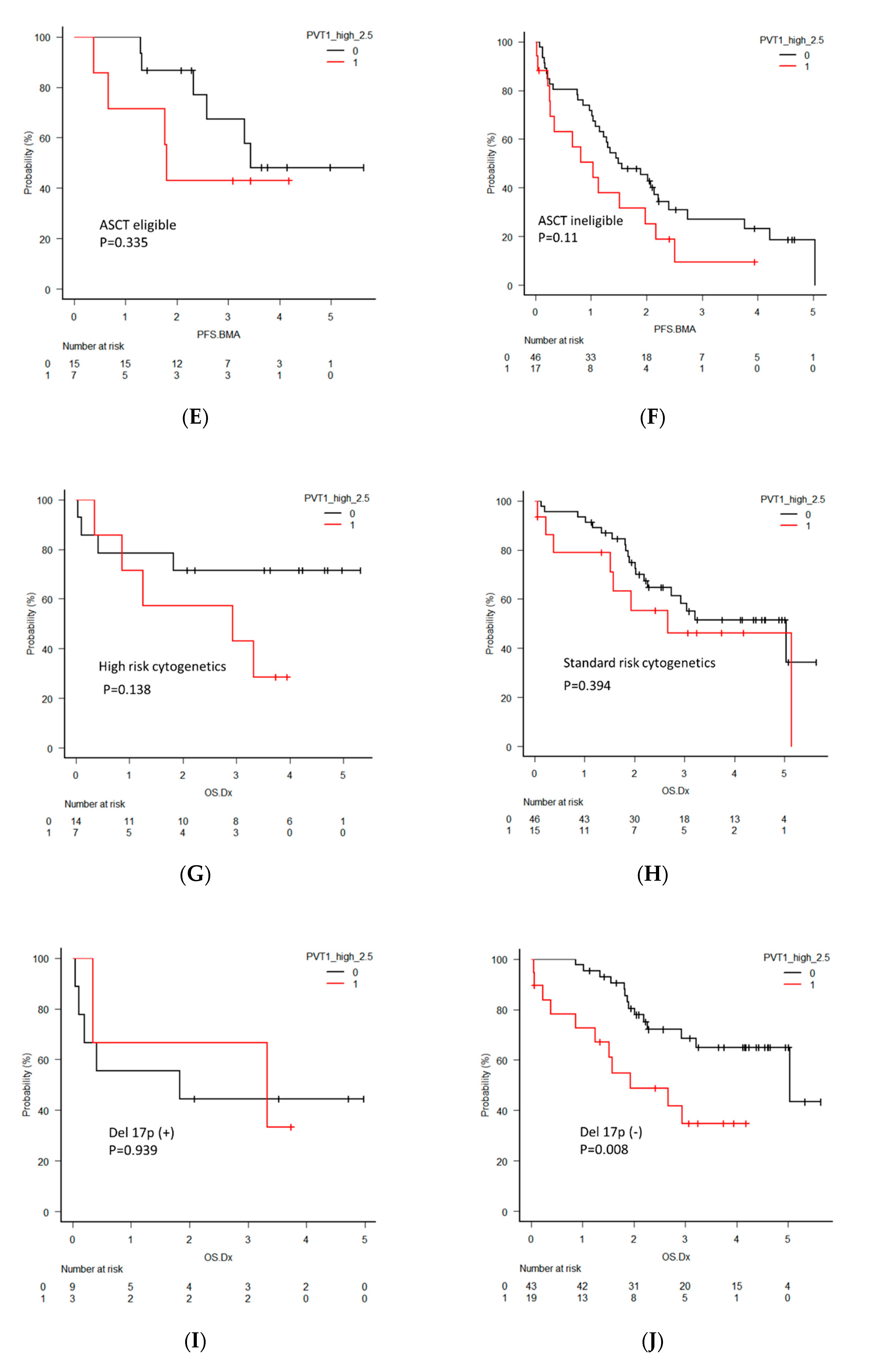
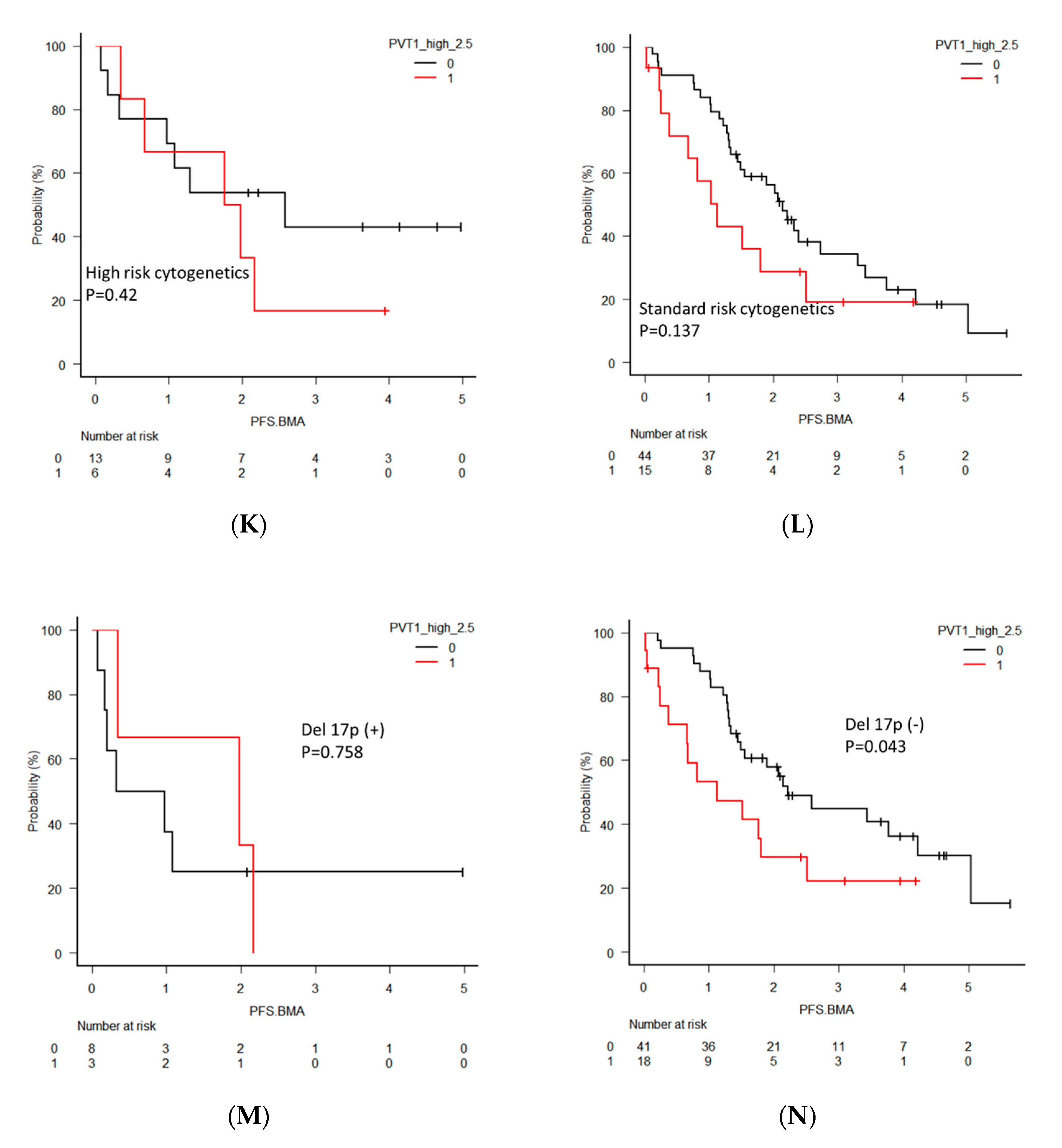
2.5. Clinical Significance of PVT1 Expression in MM
3. Discussion
4. Materials and Methods
4.1. Cell Lines
4.2. Patients
4.3. Treatment with Inhibitors
4.4. PVT1 Silencing in Myeloma Cell Lines
4.5. Isolation of Nucleic Acids
4.6. Real-Time PCR Analysis of PVT1 Expression
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kyle, R.A.; Rajkumar, S.V. Multiple Myeloma. Blood 2008, 111, 2962–2972. [Google Scholar] [CrossRef] [PubMed]
- Palumbo, A.; Anderson, K. Multiple Myeloma. N. Engl. J. Med. 2011, 364, 1046–1060. [Google Scholar] [CrossRef] [PubMed]
- Kyle, R.A.; Larson, D.R.; Therneau, T.M.; Dispenzieri, A.; Kumar, S.; Cerhan, J.R.; Rajkumar, S.V. Long-Term Follow-Up of Monoclonal Gammopathy of Undetermined Significance. N. Engl. J. Med. 2018, 378, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Chesi, M.; Bergsagel, P.L. Molecular Pathogenesis of Multiple Myeloma: Basic and Clinical Updates. Int. J. Hematol. 2013, 97, 313–323. [Google Scholar] [CrossRef]
- Bakkus, M.H.; Brakel-van Peer, K.M.; Michiels, J.J.; van’t Veer, M.B.; Benner, R. Amplification of the c-myc and the pvt-Like Region in Human Multiple Myeloma. Oncogene 1990, 5, 1359–1364. [Google Scholar]
- Iyer, M.K.; Niknafs, Y.S.; Malik, R.; Singhal, U.; Sahu, A.; Hosono, Y.; Barrette, T.R.; Prensner, J.R.; Evans, J.R.; Zhao, S.; et al. The Landscape of Long Noncoding RNAs in the Human Transcriptome. Nat. Genet. 2015, 47, 199–208. [Google Scholar] [CrossRef] [PubMed]
- Djebali, S.; Davis, C.A.; Merkel, A.; Dobin, A.; Lassmann, T.; Mortazavi, A.; Tanzer, A.; Lagarde, J.; Lin, W.; Schlesinger, F.; et al. Landscape of Transcription in Human Cells. Nature 2012, 489, 101–108. [Google Scholar] [CrossRef]
- Johnsson, P.; Morris, K.V. Expanding the Functional Role of Long Noncoding RNAs. Cell Res. 2014, 24, 1284–1285. [Google Scholar] [CrossRef][Green Version]
- Zhang, R.; Xia, L.Q.; Lu, W.W.; Zhang, J.; Zhu, J.S. LncRNAs and Cancer. Oncol. Lett. 2016, 12, 1233–1239. [Google Scholar] [CrossRef]
- Yarmishyn, A.A.; Kurochkin, I.V. Long Noncoding RNAs: A Potential Novel Class of Cancer Biomarkers. Front. Genet. 2015, 6, 145. [Google Scholar] [CrossRef]
- Schmitt, A.M.; Chang, H.Y.L. Long Noncoding RNAs in Cancer Pathways. Cancer Cell 2016, 29, 452–463. [Google Scholar] [CrossRef] [PubMed]
- Handa, H.; Kuroda, Y.; Kimura, K.; Masuda, Y.; Hattori, H.; Alkebsi, L.; Matsumoto, M.; Kasamatsu, T.; Kobayashi, N.; Tahara, K.I.; et al. Long Non-Coding RNA MALAT1 Is an Inducible Stress Response Gene Associated with Extramedullary Spread and Poor Prognosis of Multiple Myeloma. Br. J. Haematol. 2017, 179, 449–460. [Google Scholar] [CrossRef]
- Cui, Y.S.; Song, Y.P.; Fang, B.J. The Role of Long Non-Coding RNAs in Multiple Myeloma. Eur. J. Haematol. 2019, 103, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Butova, R.; Vychytilova-Faltejskova, P.; Souckova, A.; Sevcikova, S.; Hajek, R.L. Long Non-Coding RNAs in Multiple Myeloma. Noncoding RNA 2019, 5, 13. [Google Scholar] [CrossRef]
- Cory, S.; Graham, M.; Webb, E.; Corcoran, L.; Adams, J.M. Variant (6;15) Translocations in Murine Plasmacytomas Involve a chromosome 15 Locus at Least 72 Kb From the c-myc Oncogene. EMBO J. 1985, 4, 675–681. [Google Scholar] [CrossRef] [PubMed]
- Lu, D.; Luo, P.; Wang, Q.; Ye, Y.; Wang, B. lncRNA PVT1 in Cancer: A Review and Meta-Analysis. Clin. Chim. Acta 2017, 474, 1–7. [Google Scholar] [CrossRef]
- Colombo, T.; Farina, L.; Macino, G.; Paci, P. PVT1: A Rising Star Among Oncogenic Long Noncoding RNAs. BioMed Res. Int. 2015, 2015, 304208. [Google Scholar] [CrossRef]
- Shtivelman, E.; Bishop, J.M. The PVT Gene Frequently Amplifies with MYC in Tumor Cells. Mol. Cell. Biol. 1989, 9, 1148–1154. [Google Scholar] [CrossRef]
- Felsher, D.W.; Bishop, J.M. Reversible Tumorigenesis by MYC in Hematopoietic Lineages. Mol. Cell 1999, 4, 199–207. [Google Scholar] [CrossRef]
- Lin, C.Y.; Lovén, J.; Rahl, P.B.; Paranal, R.M.; Burge, C.B.; Bradner, J.E.; Lee, T.I.; Young, R.A. Transcriptional Amplification in Tumor Cells With Elevated c-Myc. Cell 2012, 151, 56–67. [Google Scholar] [CrossRef]
- Yang, M.; Zhang, L.; Wang, X.; Zhou, Y.; Wu, S. Down-Regulation of miR-203a by lncRNA PVT1 in Multiple Myeloma Promotes Cell Proliferation. Arch. Med. Sci. 2018, 14, 1333–1339. [Google Scholar] [CrossRef]
- Xiao, M.; Feng, Y.; Liu, C.; Zhang, Z. Prognostic Values of Long Noncoding RNA PVT1 in Various Carcinomas: An Updated Systematic Review and Meta-Analysis. Cell Prolif. 2018, 51, e12519. [Google Scholar] [CrossRef]
- Onagoruwa, O.T.; Pal, G.; Ochu, C.; Ogunwobi, O.O. Oncogenic Role of PVT1 and Therapeutic Implications. Front. Oncol. 2020, 10, 17. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Jin, J.; Liang, D.; Gao, Z.; Zhang, Y.; Guo, T.; He, Y.L. Long Noncoding RNA PVT1 as a Novel Predictor of Metastasis, Clinicopathological Characteristics and Prognosis in Human Cancers: A Meta-Analysis. Pathol. Oncol. Res. 2019, 25, 837–847. [Google Scholar] [CrossRef] [PubMed]
- Guan, Y.; Kuo, W.L.; Stilwell, J.L.; Takano, H.; Lapuk, A.V.; Fridlyand, J.; Mao, J.H.; Yu, M.; Miller, M.A.; Santos, J.L.; et al. Amplification of PVT1 Contributes to the Pathophysiology of Ovarian and Breast Cancer. Clin. Cancer Res. 2007, 13, 5745–5755. [Google Scholar] [CrossRef] [PubMed]
- Hnisz, D.; Abraham, B.J.; Lee, T.I.; Lau, A.; Saint-André, V.; Sigova, A.A.; Hoke, H.A.; Young, R.A. Super-Enhancers in the Control of Cell Identity and Disease. Cell 2013, 155, 934–947. [Google Scholar] [CrossRef]
- Lovén, J.; Hoke, H.A.; Lin, C.Y.; Lau, A.; Orlando, D.A.; Vakoc, C.R.; Bradner, J.E.; Lee, T.I.; Young, R.A. Selective Inhibition of Tumor Oncogenes by Disruption of Super-Enhancers. Cell 2013, 153, 320–334. [Google Scholar] [CrossRef]
- Delmore, J.E.; Issa, G.C.; Lemieux, M.E.; Rahl, P.B.; Shi, J.; Jacobs, H.M.; Kastritis, E.; Gilpatrick, T.; Paranal, R.M.; Qi, J.; et al. BET Bromodomain Inhibition as a Therapeutic Strategy to Target c-Myc. Cell 2011, 146, 904–917. [Google Scholar] [CrossRef]
- Tseng, Y.Y.; Moriarity, B.S.; Gong, W.; Akiyama, R.; Tiwari, A.; Kawakami, H.; Ronning, P.; Reuland, B.; Guenther, K.; Beadnell, T.C.; et al. PVT1 Dependence in Cancer With MYC Copy-Number Increase. Nature 2014, 512, 82–86. [Google Scholar] [CrossRef]
- Nagoshi, H.; Taki, T.; Hanamura, I.; Nitta, M.; Otsuki, T.; Nishida, K.; Okuda, K.; Sakamoto, N.; Kobayashi, S.; Yamamoto-Sugitani, M.; et al. Frequent PVT1 Rearrangement and Novel Chimeric Genes PVT1-NBEA and PVT1-WWOX Occur in Multiple Myeloma With 8q24 Abnormality. Cancer Res. 2012, 72, 4954–4962. [Google Scholar] [CrossRef]
- Yin, X.; Giap, C.; Lazo, J.S.; Prochownik, E.V. Low Molecular Weight Inhibitors of Myc-Max Interaction and Function. Oncogene 2003, 22, 6151–6159. [Google Scholar] [CrossRef]
- Carramusa, L.; Contino, F.; Ferro, A.; Minafra, L.; Perconti, G.; Giallongo, A.; Feo, S. The PVT-1 Oncogene Is a Myc Protein Target That Is Overexpressed in Transformed Cells. J. Cell. Physiol. 2007, 213, 511–518. [Google Scholar] [CrossRef] [PubMed]
- Houshmand, M.; Yazdi, N.; Kazemi, A.; Atashi, A.; Hamidieh, A.A.; Anjam Najemdini, A.; Mohammadi Pour, M.; Nikougoftar Zarif, M.L. Long Non-Coding RNA PVT1 as a Novel Candidate for Targeted Therapy in Hematologic Malignancies. Int. J. Biochem. Cell Biol. 2018, 98, 54–64. [Google Scholar] [CrossRef] [PubMed]
- Li, M.Y.; Tang, X.H.; Fu, Y.; Wang, T.J.; Zhu, J.M. Regulatory Mechanisms and Clinical Applications of the Long Non-Coding RNA PVT1 in Cancer Treatment. Front. Oncol. 2019, 9, 787. [Google Scholar] [CrossRef]
- Ogunwobi, O.O.; Kumar, A. Chemoresistance Mediated by ceRNA Networks Associated with the PVT1 Locus. Front. Oncol. 2019, 9, 834. [Google Scholar] [CrossRef] [PubMed]
- Conte, F.; Fiscon, G.; Chiara, M.; Colombo, T.; Farina, L.; Paci, P. Role of the Long Non-Coding RNA PVT1 in the Dysregulation of the ceRNA-ceRNA Network in Human Breast Cancer. PLoS ONE 2017, 12, e0171661. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Handa, H.; Honma, K.; Oda, T.; Kobayashi, N.; Kuroda, Y.; Kimura-Masuda, K.; Watanabe, S.; Ishihara, R.; Murakami, Y.; Masuda, Y.; et al. Long Noncoding RNA PVT1 Is Regulated by Bromodomain Protein BRD4 in Multiple Myeloma and Is Associated with Disease Progression. Int. J. Mol. Sci. 2020, 21, 7121. https://doi.org/10.3390/ijms21197121
Handa H, Honma K, Oda T, Kobayashi N, Kuroda Y, Kimura-Masuda K, Watanabe S, Ishihara R, Murakami Y, Masuda Y, et al. Long Noncoding RNA PVT1 Is Regulated by Bromodomain Protein BRD4 in Multiple Myeloma and Is Associated with Disease Progression. International Journal of Molecular Sciences. 2020; 21(19):7121. https://doi.org/10.3390/ijms21197121
Chicago/Turabian StyleHanda, Hiroshi, Kazuki Honma, Tsukasa Oda, Nobuhiko Kobayashi, Yuko Kuroda, Kei Kimura-Masuda, Saki Watanabe, Rei Ishihara, Yuki Murakami, Yuta Masuda, and et al. 2020. "Long Noncoding RNA PVT1 Is Regulated by Bromodomain Protein BRD4 in Multiple Myeloma and Is Associated with Disease Progression" International Journal of Molecular Sciences 21, no. 19: 7121. https://doi.org/10.3390/ijms21197121
APA StyleHanda, H., Honma, K., Oda, T., Kobayashi, N., Kuroda, Y., Kimura-Masuda, K., Watanabe, S., Ishihara, R., Murakami, Y., Masuda, Y., Tahara, K.-i., Takei, H., Kasamatsu, T., Saitoh, T., & Murakami, H. (2020). Long Noncoding RNA PVT1 Is Regulated by Bromodomain Protein BRD4 in Multiple Myeloma and Is Associated with Disease Progression. International Journal of Molecular Sciences, 21(19), 7121. https://doi.org/10.3390/ijms21197121





