Sub-Nanometer Cryo-EM Density Map of the Human Heterodimeric Amino Acid Transporter 4F2hc-LAT2
Abstract
1. Introduction
2. Results and Discussion
3. Conclusions
4. Materials and Methods
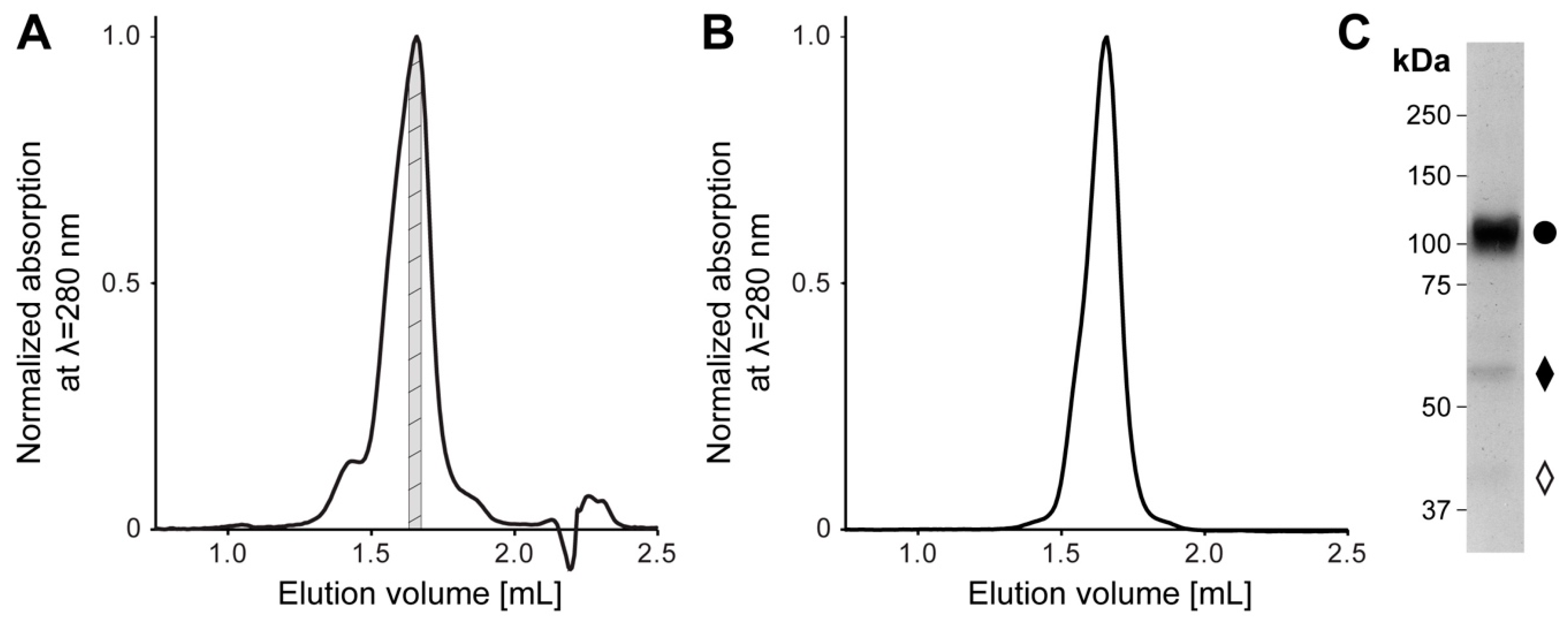
4.1. Cloning, Overexpression, and Purification of Human 4F2hc-LAT2
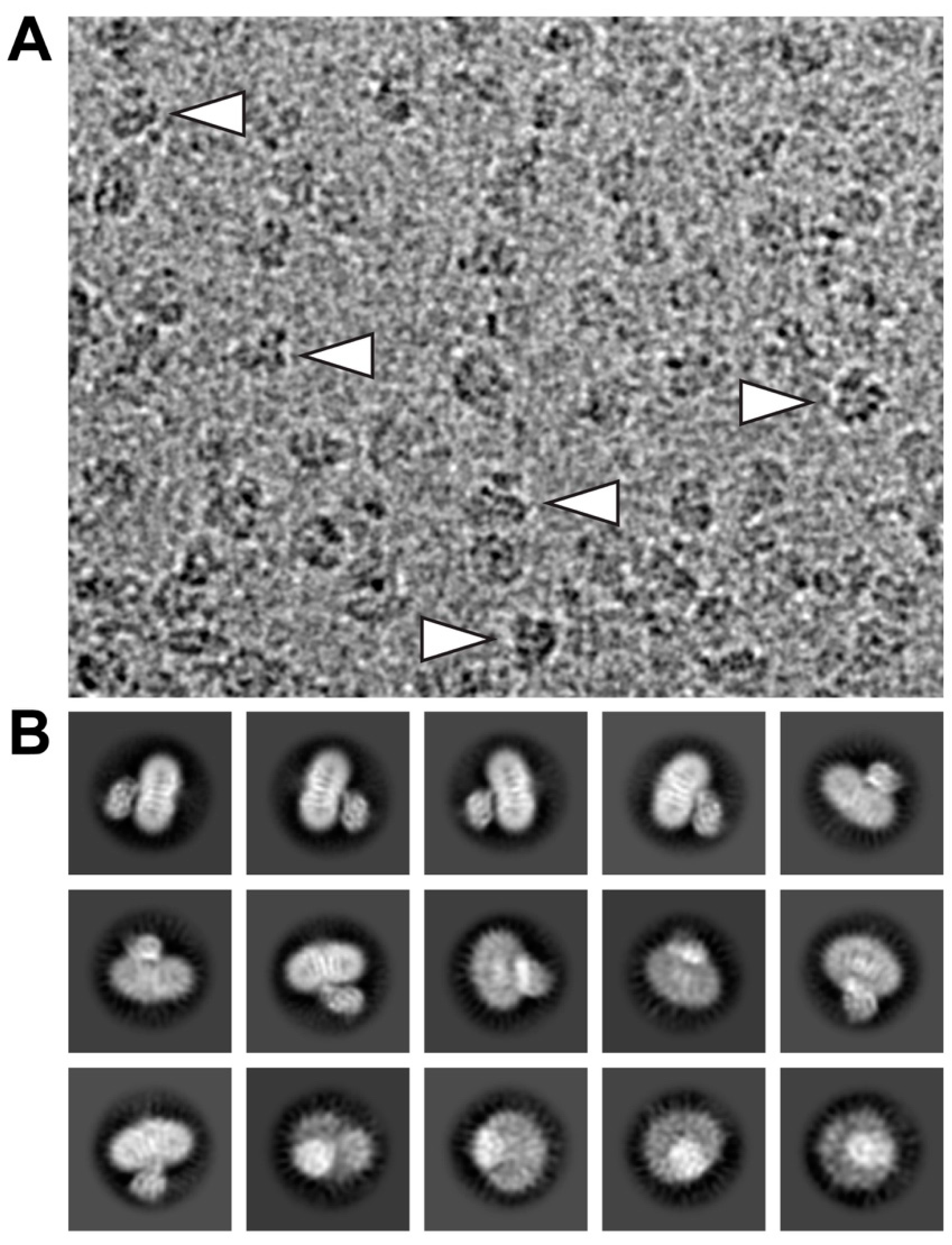
4.2. Grid Preparation and Cryo-EM Data Collection
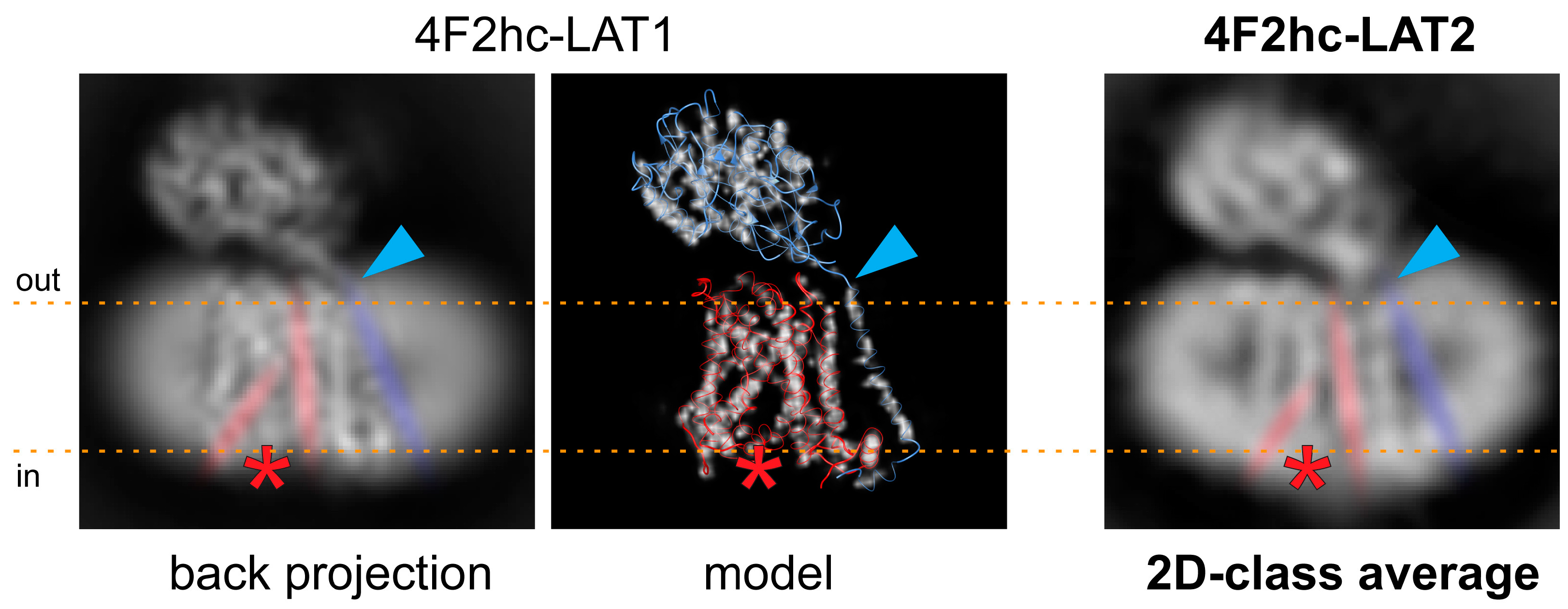
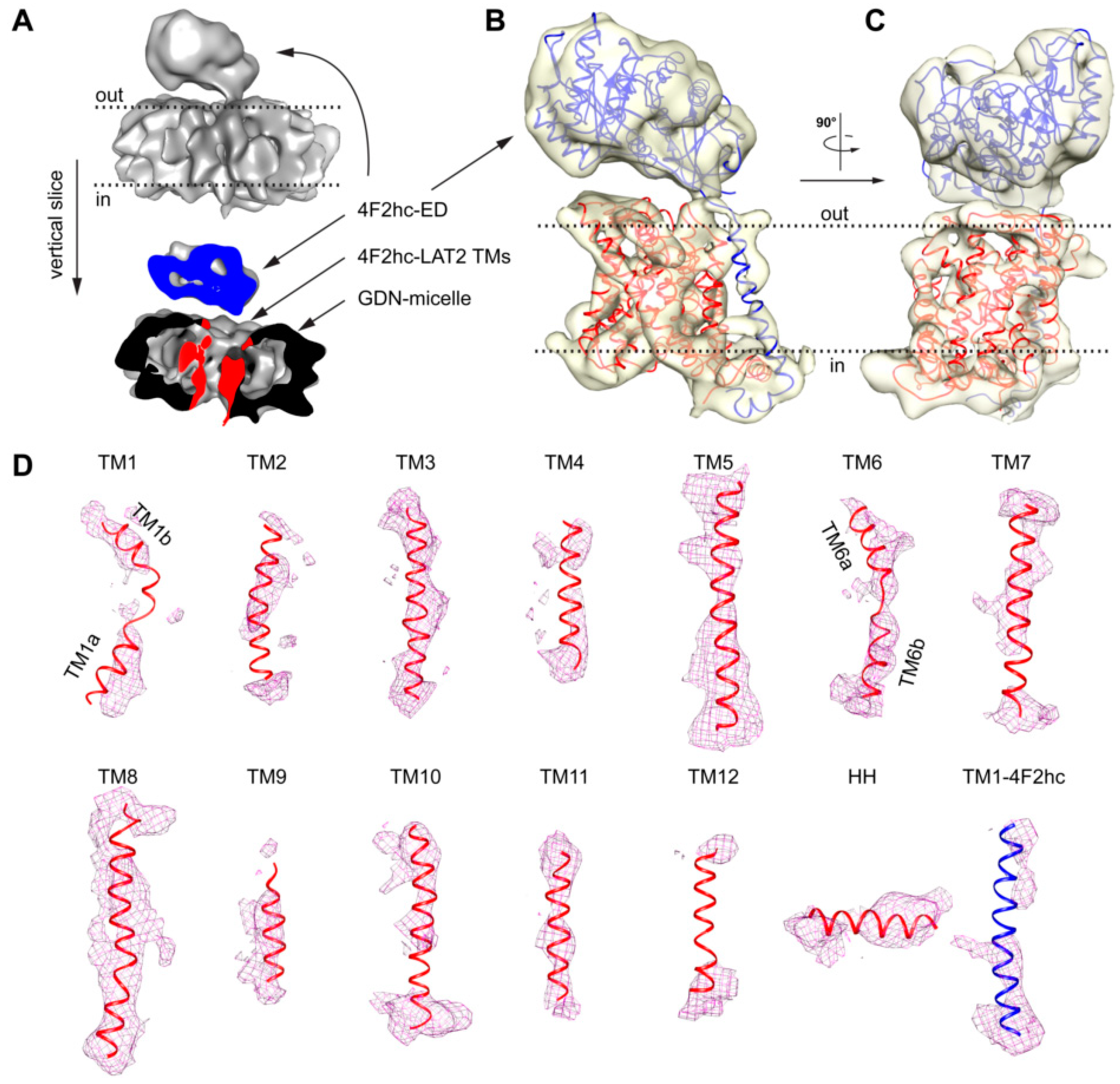
4.3. Calculation of 2D-Class Averages and Cryo-EM Density Map
4.4. Sequence Alignment Calculation of LATs and Homology Model Generation of 4F2hc-LAT2hom
4.5. Figure Preparation
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| 3D | three-dimensional |
| 4F2hc-LAT2Thom | homology model of 4F2hc-LAT2 |
| CHS | cholesteryl hemisuccinate |
| CL | cytoplasmic loop |
| CTF | contrast transfer function |
| ED | ectodomain |
| EL | extracellular loop |
| EM | electron microscopy |
| EMD | electron microscopy databank |
| GDN | glyco-diosgenin |
| HH | horizontal α-helix |
| HAT | heterodimeric amino acid transporter |
| LAT | L-type amino acid transporter |
| LMNG | lauryl maltose neopentyl glycol |
| MW | molecular weight |
| PDB | protein databank |
| PPM | positioning of proteins in membrane |
| RMSD | root-mean-square deviation |
| SDS-PAGE | SDS-polyacrylamid gel electrophoresis |
| SEC | size exclusion chromatography |
| TM | transmembrane helix |
| UniProt-ID | protein knowledgebase-identifier |
References
- Fotiadis, D.; Kanai, Y.; Palacín, M. The SLC3 and SLC7 families of amino acid transporters. Mol. Asp. Med. 2013, 34, 139–158. [Google Scholar] [CrossRef] [PubMed]
- Saier, M.H.; Reddy, V.S.; Tsu, B.V.; Ahmed, M.S.; Li, C.; Moreno-Hagelsieb, G. The Transporter Classification Database (TCDB): Recent advances. Nucleic Acids Res. 2016, 44, D372–D379. [Google Scholar] [CrossRef] [PubMed]
- Chillarón, J.; Roca, R.; Valencia, A.; Zorzano, A.; Palacín, M. Heteromeric amino acid transporters: Biochemistry, genetics, and physiology. Am. J. Physiol. Physiol. 2001, 281, F995–F1018. [Google Scholar] [CrossRef] [PubMed]
- Verrey, F.; Closs, E.I.; Wagner, C.A.; Palacin, M.; Endou, H.; Kanai, Y. CATs and HATs: The SLC7 family of amino acid transporters. Pflug. Arch. Eur. J. Physiol. 2004, 447, 532–542. [Google Scholar] [CrossRef] [PubMed]
- Rosell, A.; Meury, M.; Alvarez-Marimon, E.; Costa, M.; Perez-Cano, L.; Zorzano, A.; Fernandez-Recio, J.; Palacin, M.; Fotiadis, D. Structural bases for the interaction and stabilization of the human amino acid transporter LAT2 with its ancillary protein 4F2hc. Proc. Natl. Acad. Sci. USA 2014, 111, 2966–2971. [Google Scholar] [CrossRef]
- Mastroberardino, L.; Spindler, B.; Pfeiffer, R.; Skelly, P.J.; Loffing, J.; Shoemaker, C.B.; Verrey, F. Amino-acid transport by heterodimers of 4F2hc/CD98 and members of a permease family. Nature 1998, 395, 288–291. [Google Scholar] [CrossRef]
- Meier, C.; Ristic, Z.; Klauser, S.; Verrey, F. Activation of system L heterodimeric amino acid exchangers by intracellular substrates. EMBO J. 2002, 21, 580–589. [Google Scholar] [CrossRef]
- Pineda, M.; Fernández, E.; Torrents, D.; Estévez, R.; López, C.; Camps, M.; Lloberas, J.; Zorzano, A.; Palacín, M. Identification of a membrane protein, LAT-2, that co-expresses with 4F2 heavy chain, an L-type amino acid transport activity with broad specificity for small and large zwitterionic amino acids. J. Biol. Chem. 1999, 274, 19738–19744. [Google Scholar] [CrossRef]
- Rossier, G.; Meier, C.; Bauch, C.; Summa, V.; Sordat, B.; Verrey, F.; Kühn, L.C. LAT2, a new basolateral 4F2hc/CD98-associated amino acid transporter of kidney and intestine. J. Biol. Chem. 1999, 274, 34948–34954. [Google Scholar] [CrossRef]
- Zevenbergen, C.; Meima, M.E.; De Souza, E.C.L.; Peeters, R.P.; Kinne, A.; Krause, G.; Edward Visser, W.; Visser, T.J. Transport of iodothyronines by human l-type amino acid transporters. Endocrinology 2015, 156, 4345–4355. [Google Scholar] [CrossRef][Green Version]
- Teixeira, S.; Di Grandi, S.; Kuhn, L.C. Primary structure of the human 4F2 antigen heavy chain predicts a transmembrane protein with a cytoplasmic NH2-terminus. J. Biol. Chem. 1987, 262, 9574–9580. [Google Scholar] [PubMed]
- Fort, J.; De La Ballina, L.R.; Burghardt, H.E.; Ferrer-Costa, C.; Turnay, J.; Ferrer-Orta, C.; Usón, I.; Zorzano, A.; Fernández-Recio, J.; Orozco, M.; et al. The structure of human 4F2hc ectodomain provides a model for homodimerization and electrostatic interaction with plasma membrane. J. Biol. Chem. 2007, 282, 31444–31452. [Google Scholar] [CrossRef] [PubMed]
- Vilches, C.; Boiadjieva-Knöpfel, E.; Bodoy, S.; Camargo, S.; López de Heredia, M.; Prat, E.; Ormazabal, A.; Artuch, R.; Zorzano, A.; Verrey, F.; et al. Cooperation of antiporter LAT2/CD98hc with uniporter TAT1 for renal reabsorption of neutral amino acids. J. Am. Soc. Nephrol. 2018, 29, 1624–1635. [Google Scholar] [CrossRef] [PubMed]
- Guarch, M.E.; Font-Llitjós, M.; Murillo-Cuesta, S.; Errasti-Murugarren, E.; Celaya, A.M.; Girotto, G.; Vuckovic, D.; Mezzavilla, M.; Vilches, C.; Bodoy, S.; et al. Mutations in L-type amino acid transporter-2 support SLC7A8 as a novel gene involved in age-related hearing loss. eLife 2018, 7, 1–27. [Google Scholar]
- Knöpfel, E.B.; Vilches, C.; Camargo, S.M.R.R.; Errasti-Murugarren, E.; Stäubli, A.; Mayayo, C.; Munier, F.L.; Miroshnikova, N.; Poncet, N.; Junza, A.; et al. Dysfunctional LAT2 amino acid transporter is associated with cataract in mouse and humans. Front. Physiol. 2019, 10, 688. [Google Scholar] [CrossRef]
- Tina, E.; Prosén, S.; Lennholm, S.; Gasparyan, G.; Lindberg, M.; Eremo, A.G. Expression profile of the amino acid transporters SLC7A5, SLC7A7, SLC7A8 and the enzyme TDO2 in basal cell carcinoma. Br. J. Dermatol. 2019, 180, 130–140. [Google Scholar] [CrossRef]
- Looser, V.; Bruhlmann, B.; Bumbak, F.; Stenger, C.; Costa, M.; Camattari, A.; Fotiadis, D.; Kovar, K. Cultivation strategies to enhance productivity of Pichia pastoris: A review. Biotechnol. Adv. 2014, 33, 1177–1193. [Google Scholar] [CrossRef]
- Byrne, B. Pichia pastoris as an expression host for membrane protein structural biology. Curr. Opin. Struct. Biol. 2015, 32, 9–17. [Google Scholar] [CrossRef]
- Karlsson, M.; Fotiadis, D.; Sjövall, S.; Johansson, I.; Hedfalk, K.; Engel, A.; Kjellbom, P. Reconstitution of water channel function of an aquaporin overexpressed and purified from Pichia pastoris. FEBS Lett. 2003, 537, 68–72. [Google Scholar] [CrossRef]
- Costa, M.; Rosell, A.; Álvarez-Marimon, E.; Zorzano, A.; Fotiadis, D.; Palacín, M. Expression of human heteromeric amino acid transporters in the yeast Pichia pastoris. Protein Expr. Purif. 2013, 87, 35–40. [Google Scholar] [CrossRef]
- Meury, M.; Costa, M.; Harder, D.; Stauffer, M.; Jeckelmann, J.-M.; Brühlmann, B.; Rosell, A.; Ilgü, H.; Kovar, K.; Palacín, M.; et al. Detergent-induced stabilization and improved 3D map of the human heteromeric amino acid transporter 4F2hc-LAT2. PLoS ONE 2014, 9, e109882. [Google Scholar] [CrossRef] [PubMed]
- Jeckelmann, J.-M.; Fotiadis, D. Volta phase plate cryo-EM structure of the human heterodimeric amino acid transporter 4F2hc-LAT2. Int. J. Mol. Sci. 2019, 20, 931. [Google Scholar] [CrossRef] [PubMed]
- Yan, R.; Zhao, X.; Lei, J.; Zhou, Q. Structure of the human LAT1–4F2hc heteromeric amino acid transporter complex. Nature 2019, 568, 127–130. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Wiriyasermkul, P.; Jin, C.; Quan, L.; Ohgaki, R.; Okuda, S.; Kusakizako, T.; Nishizawa, T.; Oda, K.; Ishitani, R.; et al. Cryo-EM structure of the human L-type amino acid transporter 1 in complex with glycoprotein CD98hc. Nat. Struct. Mol. Biol. 2019, 26, 510–517. [Google Scholar] [CrossRef] [PubMed]
- Scheres, S.H.W. RELION: Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 2012, 180, 519–530. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Lu, F.; Zhou, L.; Dang, S.; Sun, L.; Li, X.; Wang, J.; Shi, Y. Structure and mechanism of an amino acid antiporter. Science 2009, 324, 1565–1568. [Google Scholar] [CrossRef] [PubMed]
- Ilgü, H.; Jeckelmann, J.-M.; Gapsys, V.; Ucurum, Z.; de Groot, B.L.D.; Fotiadis, D. Insights into the molecular basis for substrate binding and specificity of the wild-type L-arginine/agmatine antiporter AdiC. Proc. Natl. Acad. Sci. USA 2016, 113, 10358–10363. [Google Scholar] [CrossRef]
- Drew, D.; Boudker, O. Shared molecular mechanisms of membrane transporters. Annu. Rev. Biochem. 2016, 85, 543–572. [Google Scholar] [CrossRef]
- Bosshart, P.D.; Fotiadis, D. Secondary active transporters. Subcell. Biochem. 2019, 92, 275–299. [Google Scholar]
- Fotiadis, D.; Jeckelmann, J.-M. The structure of the human 4F2hc-LAT1 heteromeric amino acid transporter. Sci. China Life Sci. 2019, 794, 5–8. [Google Scholar] [CrossRef]
- Punjani, A.; Rubinstein, J.L.; Fleet, D.J.; Brubaker, M.A. CryoSPARC: Algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 2017, 14, 290–296. [Google Scholar] [CrossRef] [PubMed]
- Thonghin, N.; Kargas, V.; Clews, J.; Ford, R.C. Cryo-electron microscopy of membrane proteins. Methods 2018, 147, 176–186. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; Beer, T.A.P.D.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed]
- Zhukovsky, M.A.; Filograna, A.; Luini, A.; Corda, D.; Valente, C. Protein amphipathic helix insertion: A mechanism to induce membrane fission. Front. Cell Dev. Biol. 2019, 7, 291. [Google Scholar] [CrossRef]
- Giménez-Andrés, M.; Čopič, A.; Antonny, B. The many faces of amphipathic helices. Biomolecules 2018, 8, 45. [Google Scholar] [CrossRef]
- Lomize, M.A.; Pogozheva, I.D.; Joo, H.; Mosberg, H.I.; Lomize, A.L. OPM database and PPM web server: Resources for positioning of proteins in membranes. Nucleic Acids Res. 2012, 40, D370–D376. [Google Scholar] [CrossRef]
- Rohou, A.; Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 2015, 192, 216–221. [Google Scholar] [CrossRef]
- Tegunov, D.; Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 2019, 16, 1146–1152. [Google Scholar] [CrossRef]
- Henderson, R.; Sali, A.; Baker, M.L.; Carragher, B.; Devkota, B.; Downing, K.H.; Egelman, E.H.; Feng, Z.; Frank, J.; Grigorieff, N.; et al. Outcome of the first electron microscopy validation task force meeting. Structure 2012, 20, 205–214. [Google Scholar] [CrossRef]
- Scheres, S.H.W. A bayesian view on cryo-EM structure determination. J. Mol. Biol. 2012, 415, 406–418. [Google Scholar] [CrossRef]
- Needleman, S.B.; Wunsch, C.D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J. Mol. Biol. 1970, 48, 443–453. [Google Scholar] [CrossRef]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 2015, 10, 845–858. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
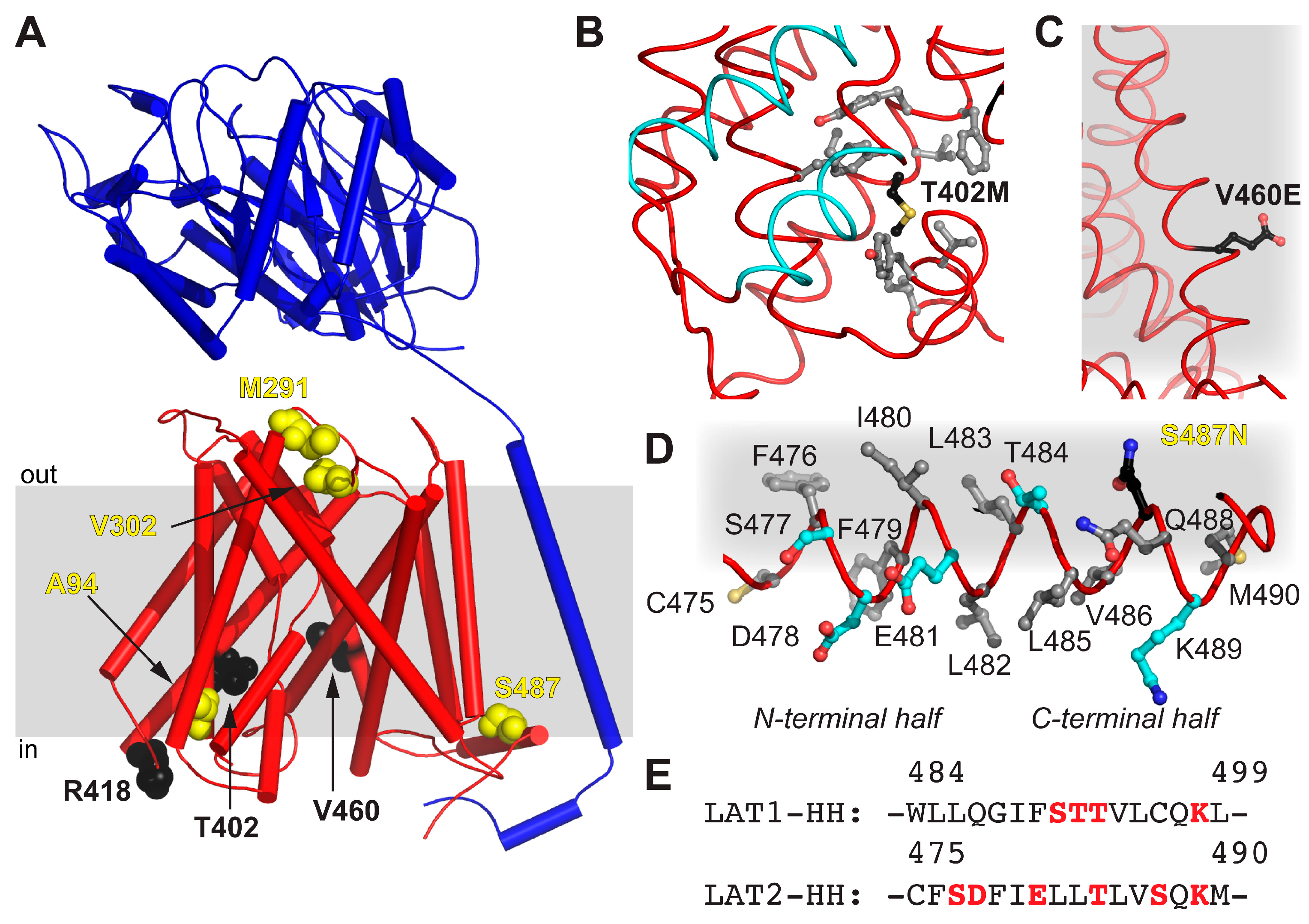
| Mutation | Location | Possible Implication |
|---|---|---|
| A94T | TM2 | ne 1 |
| M291I | EL 2 TM7–8 | Alteration of EL flexibility due to steric reasons |
| V302I | EL 2 TM7–8 | Alteration of EL flexibility due to steric reasons |
| T402M | TM10 | Impaired structural flexibility due to local hydrophobicity alteration |
| R418C | CL 3-TM10–11 | ne 1 |
| V460E | TM12 | Disturbance of LAT2 hydrophobic interactions with lipidic environment |
| S487N | HH | Contribution to the disruption of the amphipathic nature of HH |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jeckelmann, J.-M.; Fotiadis, D. Sub-Nanometer Cryo-EM Density Map of the Human Heterodimeric Amino Acid Transporter 4F2hc-LAT2. Int. J. Mol. Sci. 2020, 21, 7094. https://doi.org/10.3390/ijms21197094
Jeckelmann J-M, Fotiadis D. Sub-Nanometer Cryo-EM Density Map of the Human Heterodimeric Amino Acid Transporter 4F2hc-LAT2. International Journal of Molecular Sciences. 2020; 21(19):7094. https://doi.org/10.3390/ijms21197094
Chicago/Turabian StyleJeckelmann, Jean-Marc, and Dimitrios Fotiadis. 2020. "Sub-Nanometer Cryo-EM Density Map of the Human Heterodimeric Amino Acid Transporter 4F2hc-LAT2" International Journal of Molecular Sciences 21, no. 19: 7094. https://doi.org/10.3390/ijms21197094
APA StyleJeckelmann, J.-M., & Fotiadis, D. (2020). Sub-Nanometer Cryo-EM Density Map of the Human Heterodimeric Amino Acid Transporter 4F2hc-LAT2. International Journal of Molecular Sciences, 21(19), 7094. https://doi.org/10.3390/ijms21197094






