TIE1 as a Candidate Gene for Lymphatic Malformations with or without Lymphedema
Abstract
1. Introduction
2. Results
2.1. Clinical Results
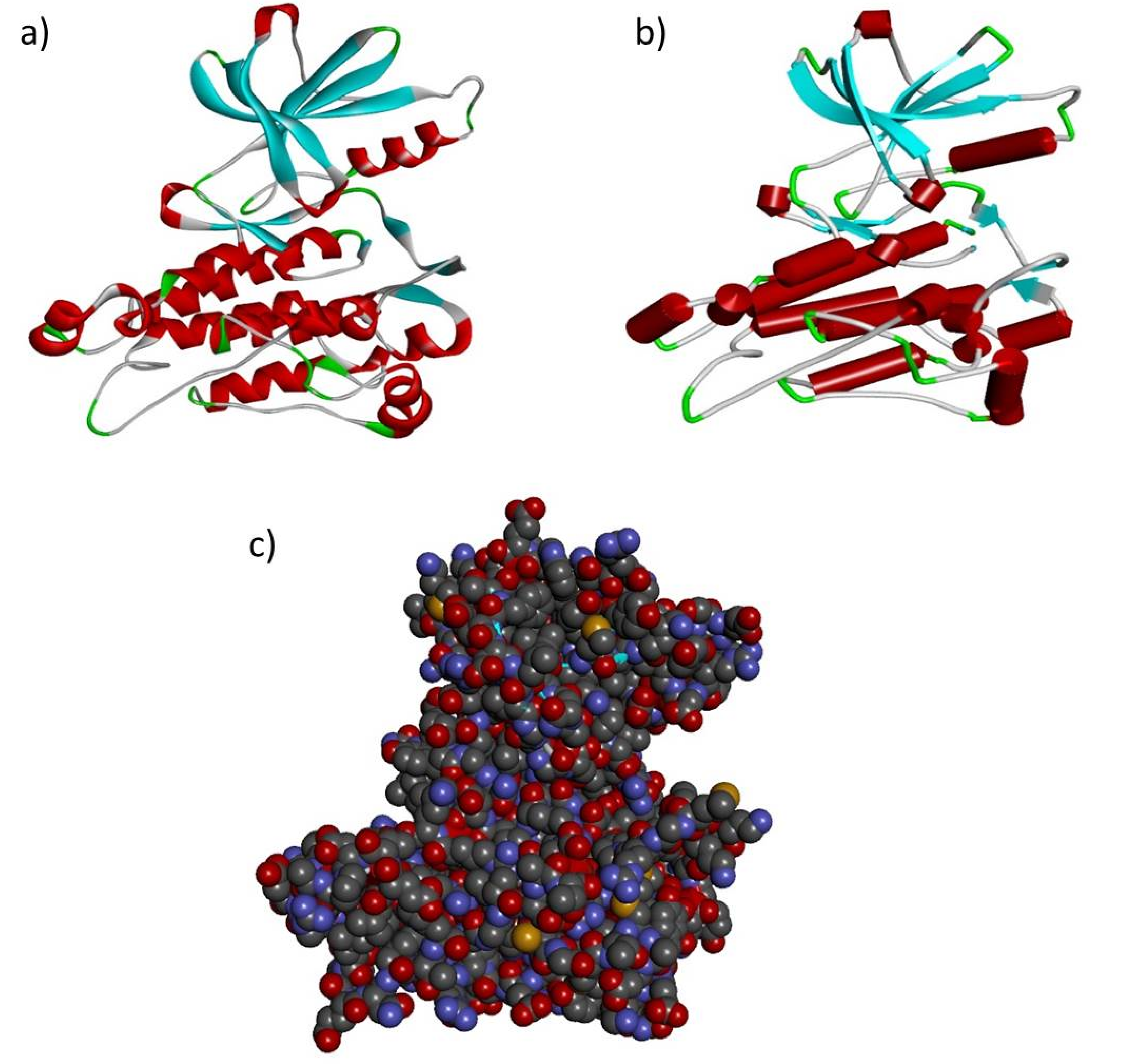
2.2. In-Silico Analysis, Template Selection and Model Building
3. Discussion
4. Materials and Methods
4.1. Clinical Evaluation
4.2. Genetic Analysis
4.3. In-Silico Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| NGS | Next Generation Sequencing |
| TIE1 | Tyrosine Kinase with Immunoglobulin-Like and EGF-Like Domains 1 |
References
- Partanen, J.; Armstrong, E.; Mäkelä, T.P.; Korhonen, J.; Sandberg, M.; Renkonen, R.; Knuutila, S.; Huebner, K.; Alitalo, K. A novel endothelial cell surface receptor tyrosine kinase with extracellular epidermal growth factor homology domains. Mol. Cell. Boil. 1992, 12, 1698–1707. [Google Scholar] [CrossRef]
- Eklund, L.; Kangas, J.; Saharinen, P. Angiopoietin–Tie signalling in the cardiovascular and lymphatic systems. Clin. Sci. 2016, 131, 87–103. [Google Scholar] [CrossRef]
- Saharinen, P.; Eklund, L.; Miettinen, J.; Wirkkala, R.; Anisimov, A.; Winderlich, M.; Nottebaum, A.; Vestweber, D.; Deutsch, U.; Koh, G.Y.; et al. Angiopoietins assemble distinct Tie2 signalling complexes in endothelial cell–cell and cell–matrix contacts. Nature 2008, 10, 527–537. [Google Scholar] [CrossRef]
- Kim, I.; Kim, H.G.; So, J.N.; Kim, J.H.; Kwak, H.J.; Koh, G.Y. Angiopoietin-1 regulates endothelial cell survival through the phosphatidylinositol 3’-kinase/Akt signal transduction pathway. Circ. Res. 2000, 86, 24–29. [Google Scholar] [CrossRef]
- Sato, T.N.; Tozawa, Y.; Deutsch, U.; Wolburg-Buchholz, K.; Fujiwara, Y.; Gendron-Maguire, M.; Gridley, T.; Wolburg, H.; Risau, W.; Qin, Y. Distinct roles of the receptor tyrosine kinases Tie-1 and Tie-2 in blood vessel formation. Nature 1995, 376, 70–74. [Google Scholar] [CrossRef]
- Puri, M.C.; Rossant, J.; Alitalo, K.; Bernstein, A.; Partanen, J. The receptor tyrosine kinase TIE is required for integrity and survival of vascular endothelial cells. EMBO J. 1995, 14, 5884–5891. [Google Scholar] [CrossRef]
- D’Amico, G.; Korhonen, E.A.; Waltari, M.; Saharinen, P.; Laakkonen, P.; Alitalo, K. Loss of endothelial tie1 receptor impairs lymphatic vessel development-brief report. Arter. Thromb. Vasc. Boil. 2010, 30, 207–209. [Google Scholar] [CrossRef]
- Qu, X.; Tompkins, K.; Batts, L.E.; Puri, M.; Baldwin, H.S.; Baldwin, H.S. Abnormal embryonic lymphatic vessel development in Tie1 hypomorphic mice. Development 2010, 137, 1285–1295. [Google Scholar] [CrossRef]
- Qu, X.; Zhou, B.; Baldwin, H.S. Tie1 is required for lymphatic valve and collecting vessel development. Dev. Boil. 2015, 399, 117–128. [Google Scholar] [CrossRef]
- Shen, B.; Shang, Z.; Wang, B.; Zhang, L.; Zhou, F.; Li, T.; Chu, M.; Jiang, H.; Wang, Y.; Qiao, T.; et al. Genetic dissection of tie pathway in mouse lymphatic maturation and valve development. Arter. Thromb. Vasc. Boil. 2014, 34, 1221–1230. [Google Scholar] [CrossRef]
- Oliver, G.; Detmar, M. The rediscovery of the lymphatic system: Old and new insights into the development and biological function of the lymphatic vasculature. Genes Dev. 2002, 16, 773–783. [Google Scholar] [CrossRef]
- Schulte-Merker, S.; Sabine, A.; Petrova, T.V. Lymphatic vascular morphogenesis in development, physiology, and disease. J. Cell Biol. 2011, 193, 607–618. [Google Scholar] [CrossRef]
- Petrova, T.V.; Karpanen, T.; Norrmén, C.; Mellor, R.; Tamakoshi, T.; Finegold, D.; Ferrell, R.; Kerjaschki, D.; Mortimer, P.; Ylä-Herttuala, S.; et al. Defective valves and abnormal mural cell recruitment underlie lymphatic vascular failure in lymphedema distichiasis. Nat. Med. 2004, 10, 974–981. [Google Scholar] [CrossRef]
- Liu, N.; Kim, Y.-W.; Lee, B.B.; Yakes, W.F.; Do, Y.S. Truncular Lymphatic Malformation (LM): Primary lymphedema. In Congenital Vascular Malformations; Springer: Berlin/Heidelberg, Germany, 2017; pp. 121–128. [Google Scholar]
- Michelini, S.; Paolacci, S.; Manara, E.; Eretta, C.; Mattassi, R.; Lee, B.B.; Bertelli, M. Genetic tests in lymphatic vascular malformations and lymphedema. J. Med. Genet. 2018, 55, 222–232. [Google Scholar] [CrossRef]
- Brouillard, P.; Boon, L.; Vikkula, M. Genetics of lymphatic anomalies. J. Clin. Investig. 2014, 124, 898–904. [Google Scholar] [CrossRef]
- Maltese, P.E.; Michelini, S.; Ricci, M.; Maitz, S.; Fiorentino, A.; Serrani, R.; Lazzerotti, A.; Bruson, A.; Paolacci, S.; Benedetti, S.; et al. Increasing evidence of hereditary lymphedema caused byCELSR1loss-of-function variants. Am. J. Med Genet. Part A 2019, 179, 1718–1724. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–423. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The protein data bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.S.; Bealer, K.; Madden, T.L. BLAST: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Remmert, M.; Biegert, A.; Hauser, A.; Söding, J. HHblits: Lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat. Methods 2011, 9, 173–175. [Google Scholar] [CrossRef]
- McLaren, W.; Gil, L.; Hunt, S.E.; Riat, H.S.; Ritchie, G.R.S.; Thormann, A.; Flicek, P.; Cunningham, F. The Ensembl Variant Effect Predictor. Genome Biol. 2016, 17, 122. [Google Scholar] [CrossRef]
- Schwarz, J.M.; Cooper, D.N.; Schuelke, M.; Seelow, D. MutationTaster2: Mutation prediction for the deep-sequencing age. Nat. Methods 2014, 11, 4361–4362. [Google Scholar] [CrossRef]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alföldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; Beer, T.A.P.D.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef]
- Mackerell, A.D.; Feig, M.; Brooks, C.L. Extending the treatment of backbone energetics in protein force fields: Limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 2004, 25, 1400–1415. [Google Scholar] [CrossRef]
- Guex, N.; Peitsch, M.C.; Schwede, T. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: A historical perspective. Electrophoresis 2009, 30, S162–S173. [Google Scholar] [CrossRef]
- Benkert, P.; Biasini, M.; Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 2010, 27, 343–350. [Google Scholar] [CrossRef]
- Biovia Discovery Studio. Discovery Studio Modeling Environment; Dassault Systèmes BIOVIA: San Diego, CA, USA, 2016. [Google Scholar]
- Sleigh, B.C.; Manna, B. Lymphedema; StatPearls Publishing: Treasure Island, FL, USA, 2020. [Google Scholar]
- Fiedler, U.; Krissl, T.; Koidl, S.; Weiss, C.; Koblizek, T.; Deutsch, U.; Martiny-Baron, G.; Marme, D.; Augustin, H.G. Angiopoietin-1 and Angiopoietin-2 share the same binding domains in the tie-2 receptor involving the first ig-like loop and the epidermal growth factor-like repeats. J. Boil. Chem. 2002, 278, 1721–1727. [Google Scholar] [CrossRef]
- Bernier, J.; Petrova, T.V. Intestinal lymphatic vasculature: Structure, mechanisms and functions. Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 510–526. [Google Scholar] [CrossRef]
- Tammela, T.; Alitalo, K. Review lymphangiogenesis: Molecular mechanisms and future promise. Cell 2010, 140, 460–476. [Google Scholar] [CrossRef]
| Organism | Gene | Variant | Embryonic Lethality | Lymphatic Phenotype |
|---|---|---|---|---|
| Mouse | TIE1 | TIE1−/− | E13.5 due to hemorrhage and cardiovascular malformations [5,6] | Edema at E12.5, abnormal patterning of lymphatic sacs and impaired integrity of lymphatic capillaries [7] |
| Mouse | TIE1 | TIE1neo/neo hypomorphic allele | After E18.5, although some mice survived to adulthood | Edema, lymphatic vasculature abnormalities, overgrown jugular lymphatic vessels, dysfunctional fluid drainage in skin [8] |
| Mouse | TIE1 | Deletion of intracellular domain of TIE1 | E18.5; some mice were born alive, but none survived | Subcutaneous edema at E13.5, failed primary lymphatic system remodeling, lymphatic system malformation in newborn mice with induced mutation [10] |
| Mouse | TIE1 | Conditional deletion in lymphatic endothelium | E18.5 (conditional mutation) | Lymphatic vessel remodeling failure, abnormal lymphatic valve morphogenesis, impaired maturation of the lymphatic system and failed development of lymphatic collecting vessels [9] |
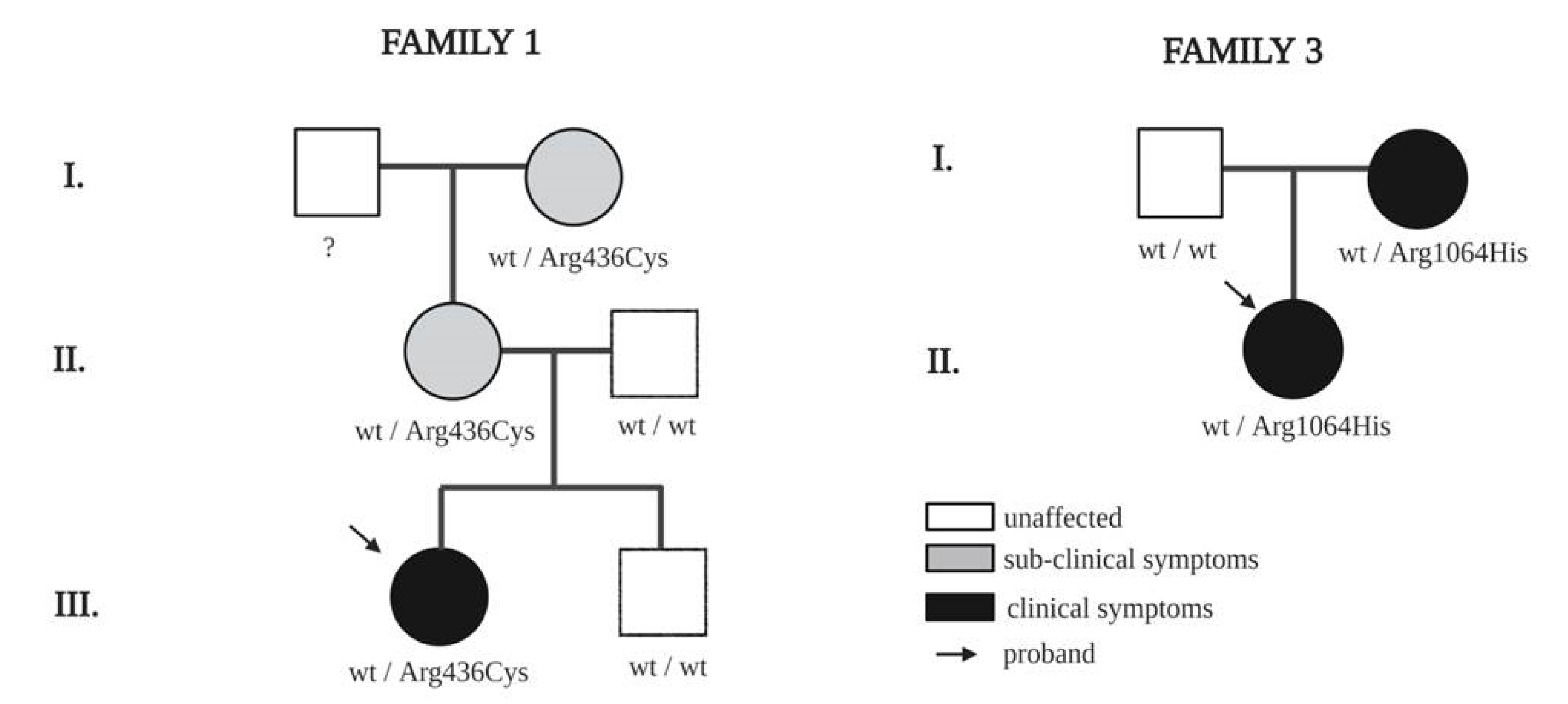
| Family | Pedigree | Sex | Age | Clinical Features | Age of Onset | Familial | Variant Nomenclature |
|---|---|---|---|---|---|---|---|
| 1 | Proband | F | 23 | Edema of lower limbs | 13 | NO | NM_001253357.1:c.1306C>T/wt; NP_001240286.1:p.Arg436Cys |
| 1 | Father | M | 52 | Healthy | / | NO | wt/wt |
| 1 | Mother | F | 49 | Healthy | / | NO | NM_001253357.1:c.1306C>T/wt; NP_001240286.1:p.Arg436Cys |
| 1 | Brother | M | 19 | Healthy | / | NO | wt/wt |
| 1 | Grandmother | F | 72 | Healthy | / | NO | NM_001253357.1:c.1306C>T/wt; NP_001240286.1:p.Arg436Cys |
| 2 | Proband | F | 52 | Edema of right lower limb | 25 | NO | NM_001253357.1:c.3046G>A/wt NP_001240286.1:p.Glu1016Lys |
| 3 | Proband | F | 47 | Lymphedema of lower limbs, from the knee down | 15 | YES | NM_001253357.1:c.3191G>A/wt; NP_001240286.1:p.Arg1064His |
| 3 | Father | M | 74 | Healthy | / | YES | wt /wt |
| 3 | Mother | F | 72 | Lymphedema | / | YES | NM_001253357.1:c.3191G>A/wt; NP_001240286.1:p.Arg1064His |
| Variant | dbSNP ID | SIFT | PolyPhen | Frequency |
|---|---|---|---|---|
| TIE1:NM_001253357.1:c.1306C>T: NP_001240286.1:p.Arg436Cys | rs139244400 | Deleterious | Possibly damaging | 0.00002847 |
| TIE1:NM_001253357.1:c.3046G>A:NP_001240286.1:p.Glu1016Lys | rs760492428 | Deleterious | Probably damaging | 0.0000325 |
| TIE1:NM_001253357.1:c.3191G>A:NP_001240286.1:p.Arg1064His | rs34993202 | Deleterious | Probably damaging | 0.0007575 |
| MVWRVPPFLLPILFLASHVGAAVDLTLLANLRLTDPQRFFLTCVSGEAGAGRGSDAWGPPLLLEKDDRIVRTPPGP PLRLARNGSHQVTLRGFSKPSDLVGVFSCVGGAGARRTRVIYVHNSPGAHLLPDKVTHTVNKGDTAVLSARVHK EKQTDVIWKSNGSYFYTLDWHEAQDGRFLLQLPNVQPPSSGIYSATYLEASPLGSAFFRLIVRGCGAGRWGPGCTK ECPGCLHGGVCHDHDGECVCPPGFTGTRCEQACREGRFGQSCQEQCPGISGCRGLTFCLPDPYGCSCGSGWRGS QCQEACAPGHFGADCRLQCQCQNGGTCDRFSGCVCPSGWHGVHCEKSDRIPQILNMASELEFNLETMPRINCAA AGNPFPVRGSIELRKPDGTVLLSTKAIVEPEKTTAEFEVPRLVLADSGFWECRVSTSGGQDSRRFKVNVKVPPVPLA APRLLTKQSRQLVVSPLVSFSGDGPISTVRLHYRPQDSTMDWSTIVVDPSENVTLMNLRPKTGYSVRVQLSRPGEG GEGAWGPPTLMTTDCPEPLLQPWLEGWHVEGTDRLRVSWSLPLVPGPLVGDGFLLRLWDGTRGQERRENVSSPQ ARTALLTGLTPGTHYQLDVQLYHCTLLGPASPPAHVLLPPSGPPAPRHLHAQALSDSEIQLTWKHPEALPGPISKYV VEVQVAGGAGDPLWIDVDRPEETSTIIRGLNASTRYLFRMRASIQGLGDWSNTVEESTLGNGLQAEGPVQESRAA EEGLDQQLILAVVGSVSATCLTILAALLTLVCIRRSCLHRRRTFTYQSGSGEETILQFSSGTLTLTRRPKLQPEPLSYPV LEWEDITFEDLIGEGNFGQVIRAMIKKDGLKMNAAIKMLKEYASENDHRDFAGELEVLCKLGHHPNIINLLGACK NRGYLYIAIEYAPYGNLLDFLRKSRVLETDPAFAREHGTASTLSSRQLLRFASDAANGMQYLSEKQFIHRDLAARNV LVGENLASKIADFGLSRGEEVYVKKTMGRLPVRWMAIESLNYSVYTTKSDVWSFGVLLWEIVSLGGTPYCGMTCA ELYEKLPQGYRMEQPRNCDDEVYELMRQCWRDRPYERPPFAQIALQLGRMLEARKAYVNMSLFENFTYAGIDATA EEA |
| Template | Seq Identity | Oligo-State | QSQE | Found by | Method | Resolution | Seq Similarity | Coverage | Description |
|---|---|---|---|---|---|---|---|---|---|
| 4k0v.1.A | 38.87 | monomer | − | HHblits | X-ray | 4.51 Å | 0.40 | 0.45 | TEK tyrosine kinase variant |
| 4k0v.1.A | 39.18 | monomer | − | BLAST | X-ray | 4.51 Å | 0.40 | 0.43 | TEK tyrosine kinase variant |
| 2gy5.1.A | 38.70 | monomer | − | HHblits | X-ray | 2.90 Å | 0.40 | 0.37 | Angiopoietin-1 receptor |
| 2gy7.1.A | 38.70 | monomer | − | HHblits | X-ray | 3.70 Å | 0.40 | 0.37 | Angiopoietin-1 receptor |
| 2gy5.1.A | 39.31 | monomer | − | BLAST | X-ray | 2.90 Å | 0.41 | 0.36 | Angiopoietin-1 receptor |
| 2gy7.1.A | 39.31 | monomer | − | BLAST | X-ray | 3.70 Å | 0.41 | 0.36 | Angiopoietin-1 receptor |
| 1fvr.1.A | 81.01 | monomer | − | BLAST | X-ray | 2.20 Å | 0.56 | 0.28 | Tyrosine-Protein Kinase TIE-2 |
| 6mwe.2.A | 81.01 | monomer | − | BLAST | X-ray | 2.05 Å | 0.56 | 0.28 | Angiopoietin-1 receptor |
| 2wqb.1.A | 79.81 | monomer | − | BLAST | X-ray | 2.95 Å | 0.55 | 0.28 | Angiopoietin-1 Receptor |
| 3I8p.1.A | 81.01 | monomer | − | BLAST | X-ray | 2.40 Å | 0.56 | 0.28 | Angiopoietin-1 receptor |
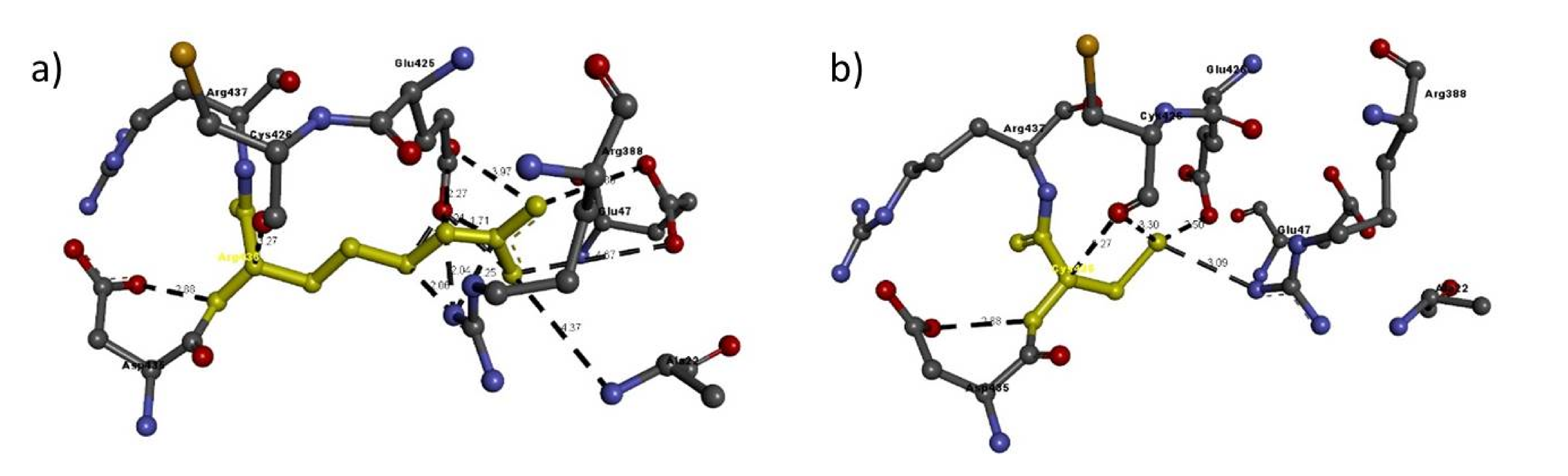
| Mutation | Amino Acid | Molecular Interactions | Bond Length in Å | Bond Type |
|---|---|---|---|---|
| Arg436Cys | Arg436 | Arg388:N-Arg436:C | 2.06 | H-bond |
| Arg388:N-Arg436:N | 2.04 | H-bond | ||
| Arg388:N-Arg436:C | 2.25 | H-bond | ||
| Glu425:C-Arg436:N | 2.27 | H-bond | ||
| Glu425:O-Arg436:C | 1.68 | H-bond | ||
| Glu425:O-Arg436:N | 1.21 | H-bond | ||
| Glu425:O-Arg436:C | 1.71 | H-bond | ||
| Arg436:N-Glu47:O | 4.67 | H-bond | ||
| Arg436:N-Glu425:O | 2.43 | H-bond | ||
| Arg436:N-Glu47:O | 4.86 | H-bond | ||
| Arg436:N-Glu425:O | 3.97 | H-bond | ||
| Arg436:N-Asp435:O | 2.88 | H-bond | ||
| Arg436:C-Cys426:O | 3.27 | H-bond | ||
| Ala22:N-Arg436:N | 4.37 | H-bond | ||
| Cys436 | Arg388:N-Cys436:S | 3.09 | H-bond | |
| Cys436:N-Asp435:O | 2.88 | H-bond | ||
| Cys436:S-Glu425:O | 2.50 | H-bond | ||
| Cys436:S-Cys426:O | 3.30 | H-bond | ||
| Cys436:C-Cys426:O | 3.27 | H-bond |
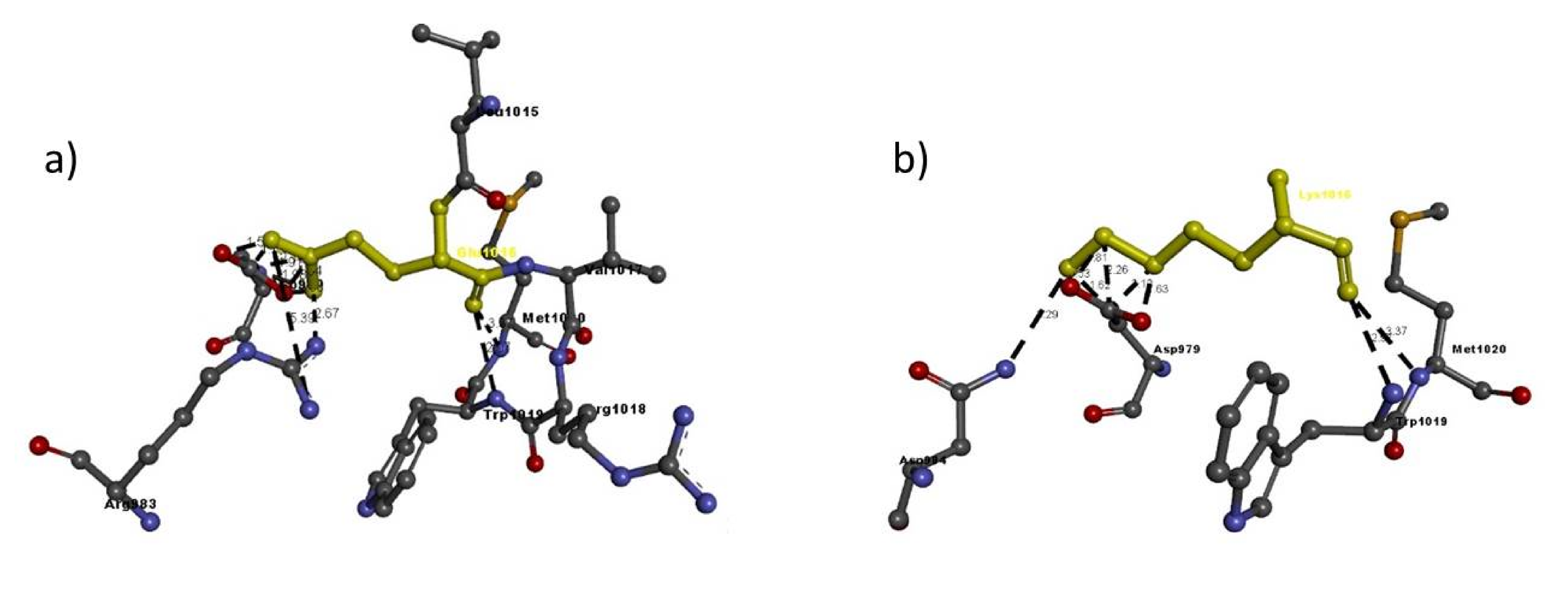
| Mutation | Amino Acid | Molecular Interactions | Bond Length in Å | Bond Type |
|---|---|---|---|---|
| Glu1016Lys | Glu1016 | Asp979:C-Glu1016:C | 2.10 | H-bond |
| Asp979:C-Glu1016:O | 1.73 | H-bond | ||
| Asp979:C-Glu1016:O | 1.98 | H-bond | ||
| Asp979:O-Glu1016:C | 1.54 | H-bond | ||
| Asp979:O-Glu1016:O | 1.91 | H-bond | ||
| Asp979:O-Glu1016:O | 0.88 | H-bond | ||
| Asp979:O-Glu1016:O | 1.56 | H-bond | ||
| Arg983:N-Glu1016:O | 5.39 | H-bond | ||
| Arg983:N-Glu1016:O | 2.67 | H-bond | ||
| Trp1019:N-Glu1016:O | 2.97 | H-bond | ||
| Met1020:N-Glu1016:O | 3.37 | H-bond | ||
| Lys1016 | Asp979:C-Lys1016:C | 2.12 | H-bond | |
| Asp979:C-Lys1016:C | 2.26 | H-bond | ||
| Asp979:C-Lys1016:N | 1.62 | H-bond | ||
| Asp979:O-Lys1016:C | 1.63 | H-bond | ||
| Asp979:O-Lys1016:C | 1.81 | H-bond | ||
| Lys1016:N-Asp979:O | 0.53 | H-bond | ||
| Trp1019:N-Lys1016:O | 2.97 | H-bond | ||
| Met1020:N-Lys1016:O | 3.37 | H-bond | ||
| Asn984:N-Lys1016:N | 3.29 | H-bond |
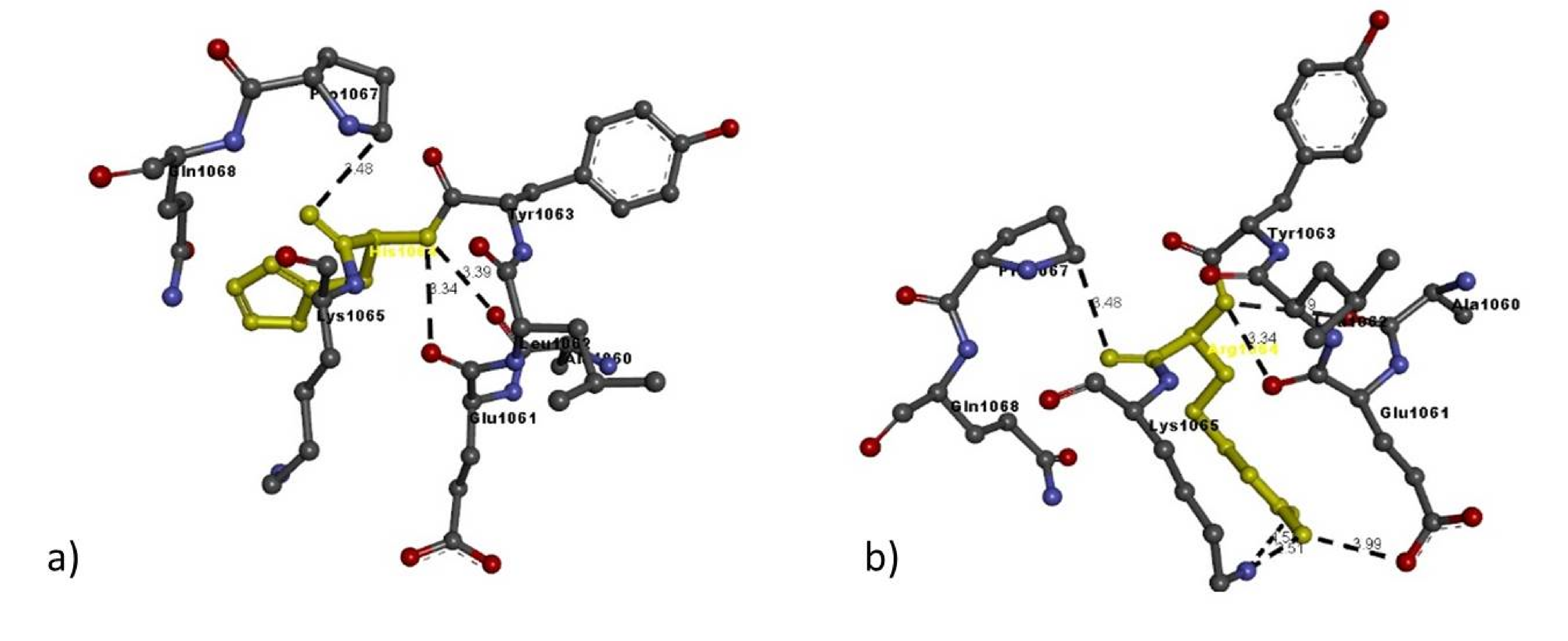
| Mutation | Amino Acid | Molecular Interactions | Bond Length in Å | Bond Type |
|---|---|---|---|---|
| Arg1064His | Arg1064 | Arg1064:N-Glu1061:O | 3.99 | H-bond |
| Arg1064:N-Ala1060:O | 3.39 | H-bond | ||
| Arg1064:N-Glu1061:O | 3.34 | H-bond | ||
| Arg1064:N-Lys1065:N | 4.57 | H-bond | ||
| Arg1064:N-Lys1065:N | 2.51 | H-bond | ||
| Lys1016 | His1064:N-Ala1060:O | 3.39 | H-bond | |
| His1064:N-Glu1061:O | 3.34 | H-bond | ||
| Pro1067:C-His1064:O | 3.48 | H-bond |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Michelini, S.; Ricci, M.; Veselenyiova, D.; Kenanoglu, S.; Kurti, D.; Baglivo, M.; Fiorentino, A.; Basha, S.H.; Priya, S.; Serrani, R.; et al. TIE1 as a Candidate Gene for Lymphatic Malformations with or without Lymphedema. Int. J. Mol. Sci. 2020, 21, 6780. https://doi.org/10.3390/ijms21186780
Michelini S, Ricci M, Veselenyiova D, Kenanoglu S, Kurti D, Baglivo M, Fiorentino A, Basha SH, Priya S, Serrani R, et al. TIE1 as a Candidate Gene for Lymphatic Malformations with or without Lymphedema. International Journal of Molecular Sciences. 2020; 21(18):6780. https://doi.org/10.3390/ijms21186780
Chicago/Turabian StyleMichelini, Sandro, Maurizio Ricci, Dominika Veselenyiova, Sercan Kenanoglu, Danjela Kurti, Mirko Baglivo, Alessandro Fiorentino, Syed Hussain Basha, Sasi Priya, Roberta Serrani, and et al. 2020. "TIE1 as a Candidate Gene for Lymphatic Malformations with or without Lymphedema" International Journal of Molecular Sciences 21, no. 18: 6780. https://doi.org/10.3390/ijms21186780
APA StyleMichelini, S., Ricci, M., Veselenyiova, D., Kenanoglu, S., Kurti, D., Baglivo, M., Fiorentino, A., Basha, S. H., Priya, S., Serrani, R., Krajcovic, J., Dundar, M., Dautaj, A., & Bertelli, M. (2020). TIE1 as a Candidate Gene for Lymphatic Malformations with or without Lymphedema. International Journal of Molecular Sciences, 21(18), 6780. https://doi.org/10.3390/ijms21186780







