F-Box Gene D5RF Is Regulated by Agrobacterium Virulence Protein VirD5 and Essential for Agrobacterium-Mediated Plant Transformation
Abstract
1. Introduction
2. Results
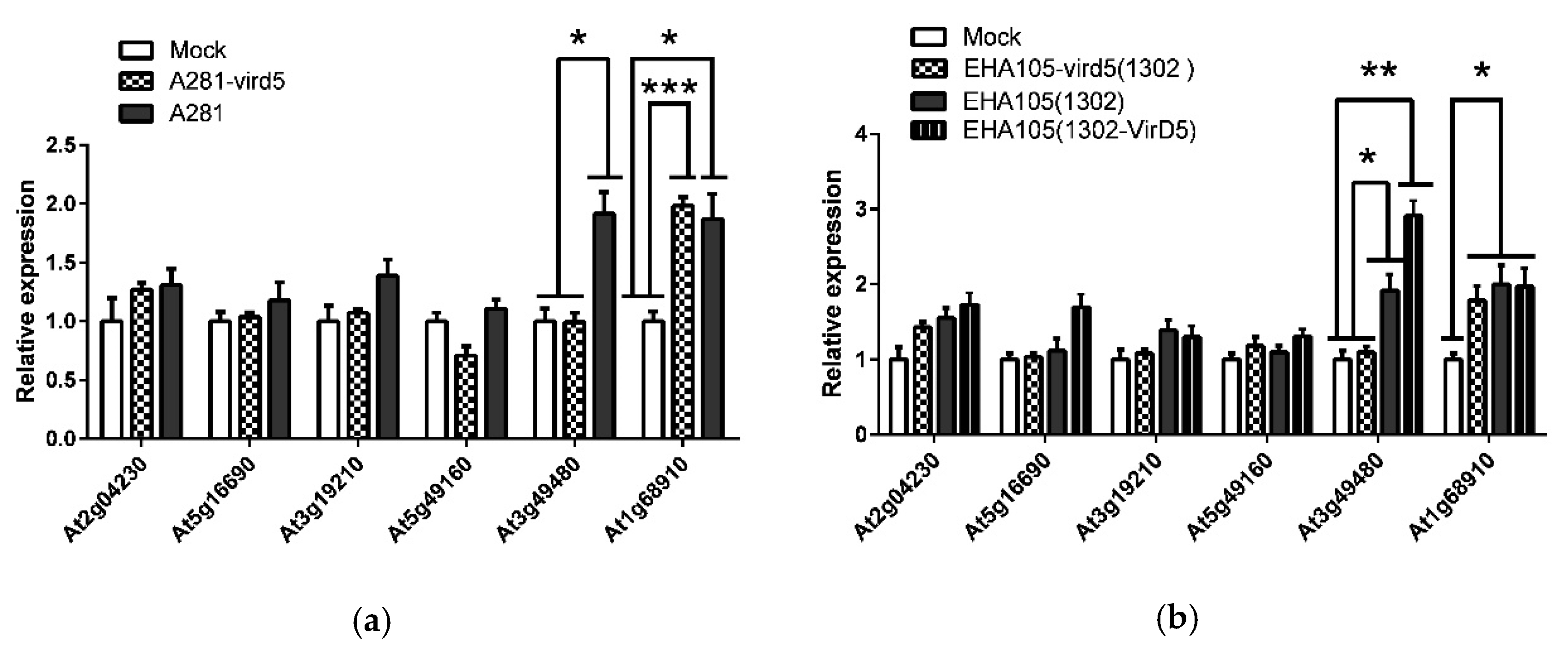
2.1. The Arabidopsis D5RF Gene Is Regulated by VirD5 at the Transcription Level
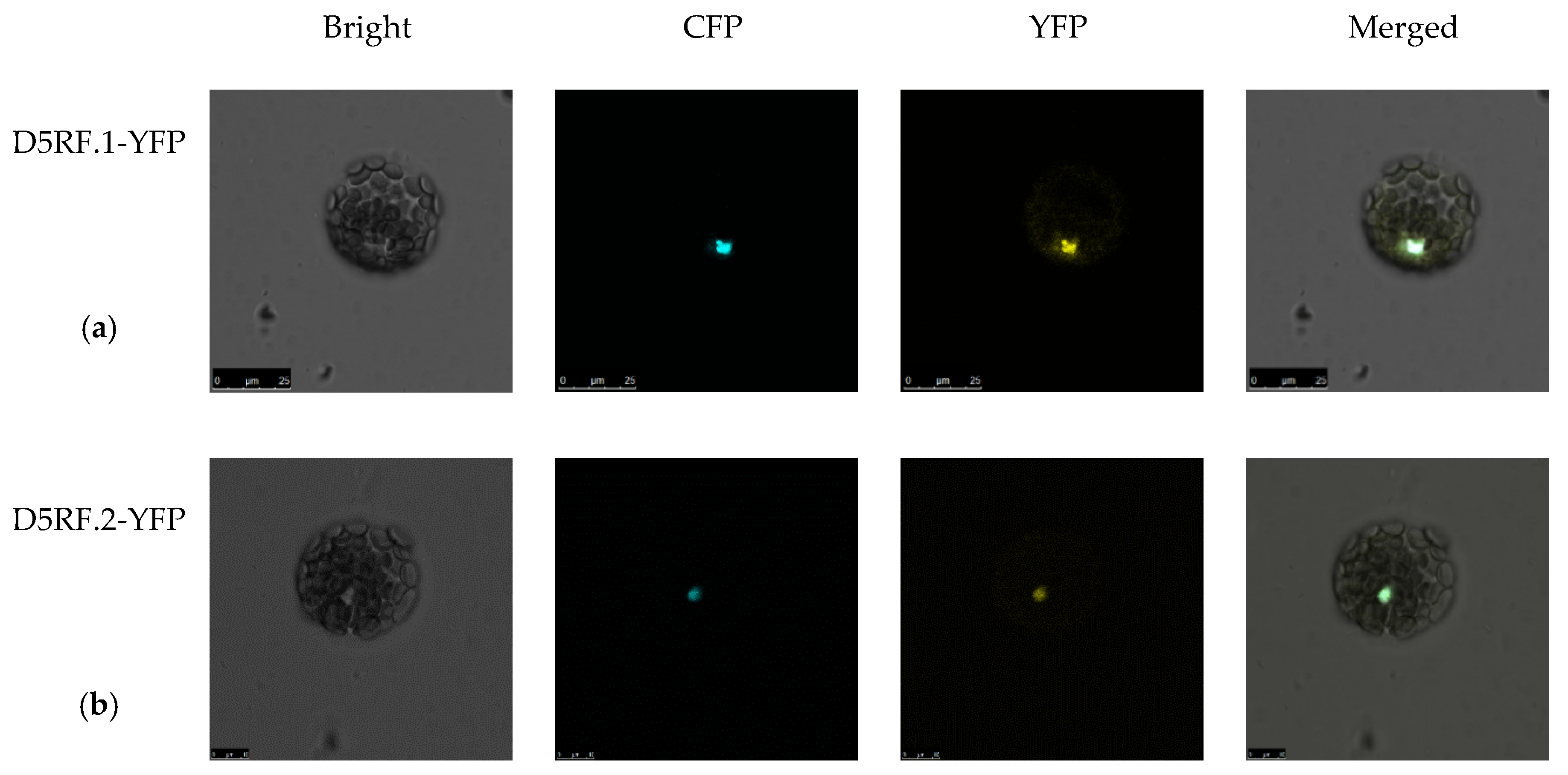
2.2. Bioinformatics Analysis and Subcellular Localization of D5RF
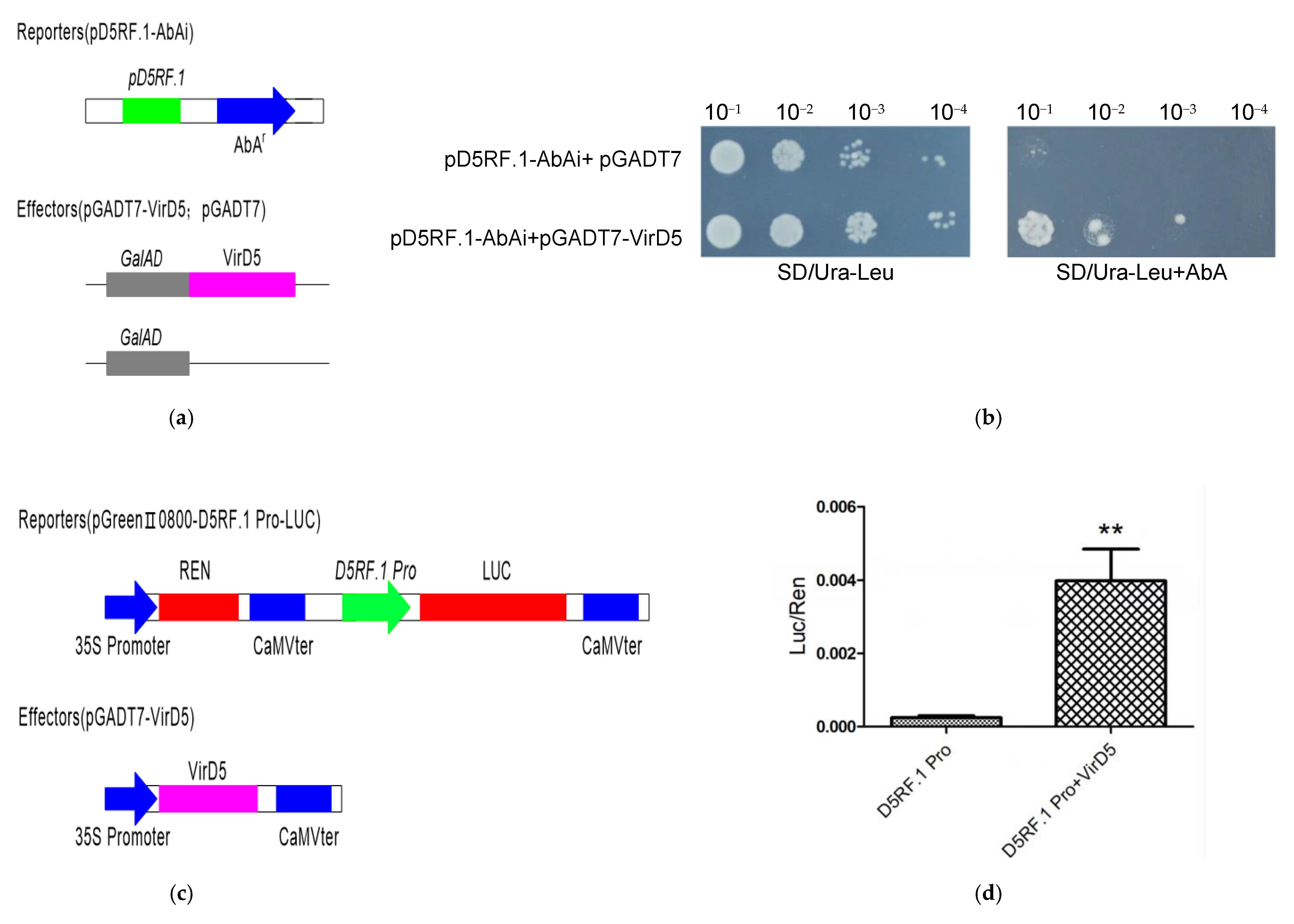
2.3. VirD5 Can Activate the Promotor of D5RF.1
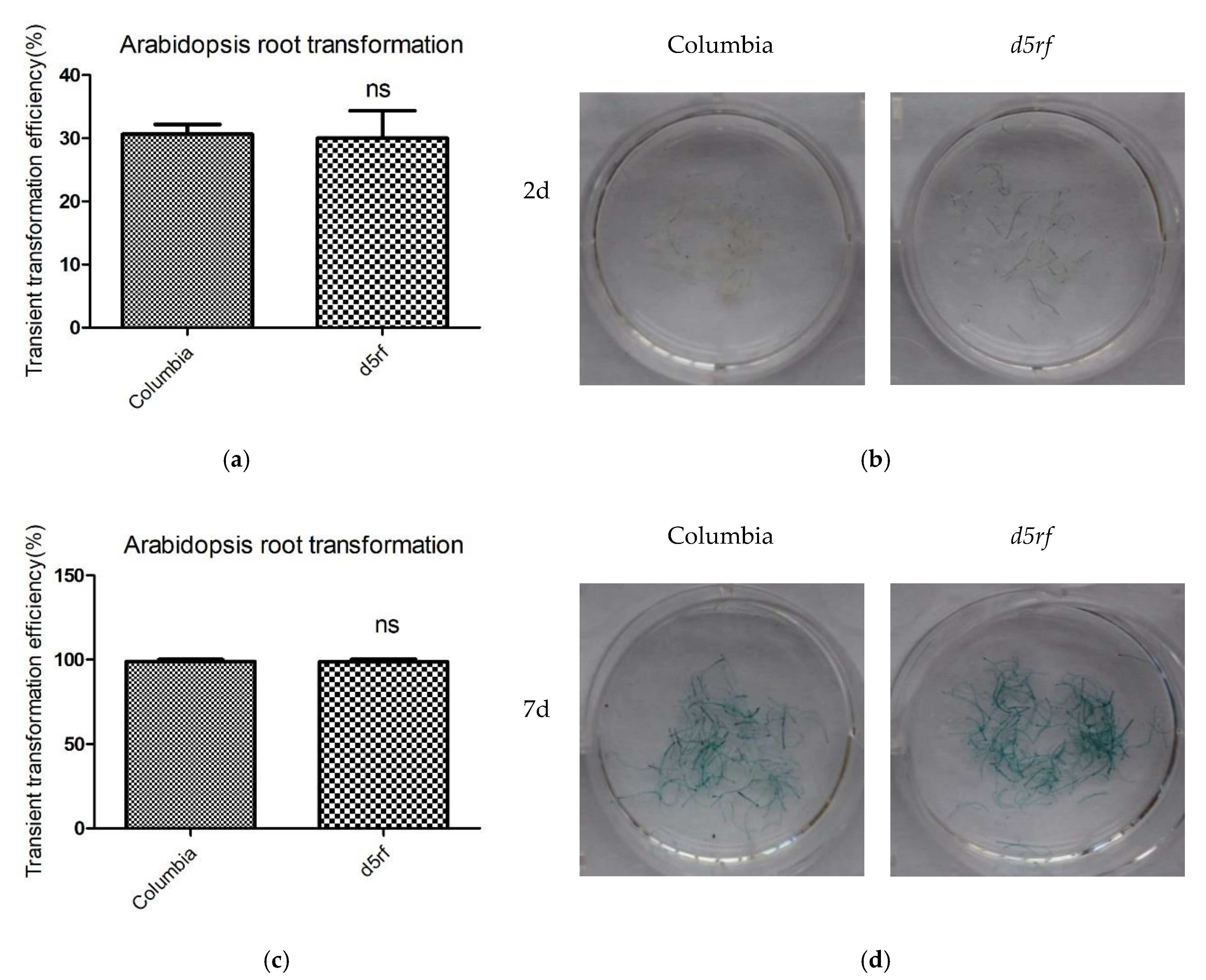
2.4. D5RF Is Required for Plant Stable Transformation, But Not for Transient Transformation
2.5. Pathogen-Responsive Genes Are Possibly Downregulated in d5rf
3. Discussions
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Yeast One-Hybrid Assay
4.3. Dual-Luciferase Reporter Assay System
4.4. Agrobacterium-Mediated Plant Stable and Transient Transformation Assay
4.5. Subcellular Localization Assay in Arabidopsis Protoplasts
4.6. Reverse Transcription PCR and RNA-Sequencing
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ABA | Abscisic Acid |
| DEGs | Differentially Expressed Genes |
| D5RE | VirD5 response element |
| D5RF | VirD5 response F-box protein |
| RT-PCR | Reverse Transcription-Polymerase Chain Reaction |
| RT-QPCR | Real-time quantitative PCR |
| WT | Wild Type |
| YFP | Yellow Fluorescent Protein |
| Y2H | Yeast Two-Hybrid |
References
- Tzfira, T.; Citovsky, V. From host recognition to T-DNA integration: The function of bacterial and plant genes in the Agrobacterium-plant cell interaction. Mol. Plant Pathol. 2000, 1, 201–212. [Google Scholar] [CrossRef]
- Gelvin, S.B. Agrobacterium and plant genes involved in T-DNA transfer and integration. Annu. Rev. Plant Biol. 2000, 51, 223–256. [Google Scholar] [CrossRef] [PubMed]
- Pitzschke, A.; Hirt, H. New insights into an old story: Agrobacterium-induced tumour formation in plants by plant transformation. EMBO J. 2010, 29, 1021–1032. [Google Scholar] [CrossRef] [PubMed]
- De Cleene, M.; De Ley, J. The host range of crown gall. Bot. Rev. 1976, 42, 389–466. [Google Scholar] [CrossRef]
- Michielse, C.B.; Hooykaas, P.; Van Den Hondel, C.A.; Ram, A.F. Agrobacterium-mediated transformation as a tool for functional genomics in fungi. Curr. Genet. 2005, 48, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Gelvin, S.B. Plant proteins involved in agrobacterium-mediated genetic transformation. Annu. Rev. Phytopathol. 2010, 48, 45–68. [Google Scholar] [CrossRef] [PubMed]
- Gelvin, S.B. Integration of agrobacteriumT-DNA into the plant genome. Annu. Rev. Genet. 2017, 51, 195–217. [Google Scholar] [CrossRef] [PubMed]
- Zupan, J.; Muth, T.R.; Draper, O.; Zambryski, P. The transfer of DNA from Agrobacterium tumefaciens into plants: A feast of fundamental insights. Plant J. 2000, 23, 11–28. [Google Scholar] [CrossRef] [PubMed]
- Fronzes, R.; Christie, P.J.; Waksman, G. The structural biology of type IV secretion systems. Nat. Rev. Genet. 2009, 7, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Citovsky, V.; Wong, M.L.; Zambryski, P. Cooperative interaction of agrobacterium VirE2 protein with single-stranded DNA: Implications for the T-DNA transfer process. Proc. Natl. Acad. Sci. USA 1989, 86, 1193–1197. [Google Scholar] [CrossRef]
- Tzfira, T.; Citovsky, V. Agrobacterium-mediated genetic transformation of plants: Biology and biotechnology. Curr. Opin. Biotechnol. 2006, 17, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Vergunst, A.C.; Schrammeijer, B.; Dulk-Ras, A.D.; De Vlaam, C.M.T.; Regensburg-Tuı̈nk, T.J.G.; Hooykaas, P. VirB/D4-dependent protein translocation from agrobacterium into plant cells. Science 2000, 290, 979–982. [Google Scholar] [CrossRef] [PubMed]
- Tzfira, T.; Vaidya, M.; Citovsky, V. VIP1, an Arabidopsis protein that interacts with Agrobacterium VirE2, is involved in VirE2 nuclear import and Agrobacterium infectivity. EMBO J. 2001, 20, 3596–3607. [Google Scholar] [CrossRef]
- Shi, Y.; Lee, L.Y.; Gelvin, S.B. Is VIP1 important for Agrobacterium-mediated transformation? Plant J. 2014, 79, 848–860. [Google Scholar] [CrossRef] [PubMed]
- Meza, T.J. Analyses of single-copy Arabidopsis T-DNA-transformed lines show that the presence of vector backbone sequences, short inverted repeats and DNA methylation is not sufficient or necessary for the induction of transgene silencing. Nucleic Acids Res. 2002, 30, 4556–4566. [Google Scholar] [CrossRef] [PubMed]
- Pelczar, P.; Kalck, V.; Gomez, D.; Hohn, B. Agrobacterium proteins VirD2 and VirE2 mediate precise integration of synthetic T-DNA complexes in mammalian cells. EMBO Rep. 2004, 5, 632–637. [Google Scholar] [CrossRef]
- Paszkowski, J.; Baur, M.; Bogucki, A.; Potrykus, I. Gene targeting in plants. EMBO J. 1988, 7, 4021–4026. [Google Scholar] [CrossRef]
- Gheysen, G.; Villarroel, R.; Van Montagu, M. Illegitimate recombination in plants: A model for T-DNA integration. Genes Dev. 1991, 5, 287–297. [Google Scholar] [CrossRef]
- Van Kregten, M.; De Pater, S.; Romeijn, R.; Van Schendel, R.; Hooykaas, P.; Tijsterman, M. T-DNA integration in plants results from polymerase-θ-mediated DNA repair. Nat. Plants 2016, 2, 16164. [Google Scholar] [CrossRef]
- Mysore, K.S.; Nam, J.; Gelvin, S.B. An Arabidopsis histone H2A mutant is deficient in Agrobacterium T-DNA integration. Proc. Natl. Acad. Sci. USA 2000, 97, 948–953. [Google Scholar] [CrossRef]
- Zhu, Y.; Nam, J.; Humara, J.M.; Mysore, K.S.; Lee, L.Y.; Cao, H.; Valentine, L.; Li, J.; Kaiser, A.D.; Kopecky, A.L.; et al. Identification of Arabidopsis rat mutants. Plant Physiol. 2003, 132, 494–505. [Google Scholar] [CrossRef] [PubMed]
- Crane, Y.M.; Gelvin, S.B. RNAi-mediated gene silencing reveals involvement of Arabidopsis chromatin-related genes in Agrobacterium-mediated root transformation. Proc. Natl. Acad. Sci. USA 2007, 104, 15156–15161. [Google Scholar] [CrossRef] [PubMed]
- Anand, A.; Krichevsky, A.; Schornack, S.; Lahaye, T.; Tzfira, T.; Tang, Y.; Citovsky, V.; Mysore, K.S. Arabidopsis Vire2 interacting Protein2 is required for agrobacterium t-DNA integration in plants. Plant Cell 2007, 19, 1695–1708. [Google Scholar] [CrossRef] [PubMed]
- Tzfira, T.; Vaidya, M.; Citovsky, V. Involvement of targeted proteolysis in plant genetic transformation by Agrobacterium. Nature 2004, 431, 87–92. [Google Scholar] [CrossRef]
- Zaltsman, A.; Krichevsky, A.; Loyter, A.; Citovsky, V. Agrobacterium induces expression of a host F-box protein required for tumorigenicity. Cell Host Microbe 2010, 7, 197–209. [Google Scholar] [CrossRef]
- Melchers, L.S.; Maroney, M.J.; Dulk-Ras, A.D.; Thompson, D.V.; Van Vuuren, H.A.J.; Schilperoort, R.A.; Hooykaas, P. Octopine and nopaline strains of Agrobacterium tumefaciens differ in virulence; molecular characterization of the virF locus. Plant Mol. Biol. 1990, 14, 249–259. [Google Scholar] [CrossRef]
- Regensburg-Tuïnk, A.J.G.; Hooykaas, P. Transgenic N. glauca plants expressing bacterial virulence gene virF are converted into hosts for nopaline strains of A. tumefaciens. Nature 1993, 363, 69–71. [Google Scholar] [CrossRef]
- Schrammeijer, B.; Risseeuw, E.; Pansegrau, W.; Regensburg-Tuink, T.J.; Crosby, W.L.; Hooykaas, P.J. Interaction of the virulence protein VirF of Agrobacterium tumefaciens with plant homologs of the yeast Skp1 protein. Curr. Biol. 2001, 11, 258–262. [Google Scholar] [CrossRef]
- Lacroix, B.; Loyter, A.; Citovsky, V. Association of the Agrobacterium T-DNA–protein complex with plant nucleosomes. Proc. Natl. Acad. Sci. USA 2008, 105, 15429–15434. [Google Scholar] [CrossRef]
- García-Rodríguez, F.M.; Schrammeijer, B.; Hooykaas, P. The Agrobacterium VirE3 effector protein: A potential plant transcriptional activator. Nucleic Acids Res. 2006, 34, 6496–6504. [Google Scholar] [CrossRef]
- Niu, X.; Zhou, M.; Henkel, C.V.; Van Heusden, G.P.H.; Hooykaas, P. The Agrobacterium tumefaciens virulence protein VirE3 is a transcriptional activator of the F-box gene VBF. Plant J. 2015, 84, 914–924. [Google Scholar] [CrossRef]
- Li, X.; Tu, H.; Pan, S.Q. Agrobacterium Delivers Anchorage Protein VirE3 for Companion VirE2 to Aggregate at Host Entry Sites for T-DNA Protection. Cell Rep. 2018, 25, 302–311.e6. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Peng, W.; Zhou, X.; Huang, F.; Shao, L.; Luo, M. The putative Agrobacterium transcriptional activator-like virulence protein VirD5 may target T-complex to prevent the degradation of coat proteins in the plant cell nucleus. New Phytol. 2014, 203, 1266–1281. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Van Heusden, G.P.H.; Hooykaas, P.J.J. Virulence protein VirD5 of Agrobacterium tumefaciens binds to kinetochores in host cells via an interaction with Spt4. Proc. Natl. Acad. Sci. USA 2017, 114, 10238–10243. [Google Scholar] [CrossRef]
- Zhang, X.; Hooykaas, P. The Agrobacterium VirD5 protein hyperactivates the mitotic Aurora kinase in host cells. New Phytol. 2019, 222, 1551–1560. [Google Scholar] [CrossRef] [PubMed]
- Magori, S.; Citovsky, V. Agrobacterium counteracts host-induced degradation of its effector F-box protein. Sci. Signal. 2011, 4, ra69. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, S.; Huang, F.; Zhou, X.; Chen, Z.; Peng, W.; Luo, M. VirD5 is required for efficient Agrobacterium infection and interacts with Arabidopsis VIP2. New Phytol. 2018, 217, 726–738. [Google Scholar] [CrossRef]
- Liu, D.; Shi, S.; Hao, Z.; Xiong, W.; Luo, M. OsbZIP81, a homologue of Arabidopsis VIP1, may positively regulate JA levels by directly targetting the genes in JA signaling and metabolism pathway in rice. Int. J. Mol. Sci. 2019, 20, 2360. [Google Scholar] [CrossRef]
- Pierleoni, A.; Martelli, P.L.; Fariselli, P.; Casadio, R. BaCelLo: A balanced subcellular localization predictor. Bioinformatics. 2006, 22, e408–e416. [Google Scholar] [CrossRef]
- Xue, W.; Xing, Y.; Weng, X.; Zhao, Y.; Tang, W.; Wang, L.; Zhou, H.; Yu, S.; Xu, C.; Li, X.; et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 2008, 40, 761–767. [Google Scholar] [CrossRef]
- Yan, W.-H.; Liu, H.; Zhou, X.; Li, Q.; Zhang, J.; Lu, L.; Liu, T.; Liu, H.; Zhang, C.; Zhang, Z.; et al. Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res. 2013, 23, 969–971. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Nam, J.; Carpita, N.C.; Matthysse, A.G.; Gelvin, S.B. Agrobacterium-mediated root transformation is inhibited by mutation of an arabidopsis cellulose synthase-like gene. Plant Physiol. 2003, 133, 1000–1010. [Google Scholar] [CrossRef] [PubMed]
- Vergunst, A.C.; Van Lier, M.C.M.; Dulk-Ras, A.D.; Stüve, T.A.G.; Ouwehand, A.; Hooykaas, P. Positive charge is an important feature of the C-terminal transport signal of the VirB/D4-translocated proteins of Agrobacterium. Proc. Natl. Acad. Sci. USA 2005, 102, 832–837. [Google Scholar] [CrossRef]
- Koukolíková-Nicola, Z.; Raineri, D.; Stephens, K.; Ramos, C.; Tinland, B.; Nester, E.W.; Hohn, B. Genetic analysis of the virD operon of Agrobacterium tumefaciens: A search for functions involved in transport of T-DNA into the plant cell nucleus and in T-DNA integration. J. Bacteriol. 1993, 175, 723–731. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kalogeraki, V.S.; Zhu, J.; Stryker, J.L.; Winans, S.C. The right end of the vir region of an octopine-type ti plasmid contains four new members of the vir regulon that are not essential for pathogenesis. J. Bacteriol. 2000, 182, 1774–1778. [Google Scholar] [CrossRef][Green Version]
- Zhao, Q.; Brkljacic, J.; Meier, I. Two distinct interacting classes of nuclear envelope-associated coiled-coil proteins are required for the tissue-specific nuclear envelope targeting of Arabidopsis RanGAP. Plant Cell 2008, 20, 1639–1651. [Google Scholar] [CrossRef]
- Szakonyi, D.; Duque, P. Alternative splicing as a regulator of early plant development. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Klepikova, A.V.; Kasianov, A.S.; Gerasimov, E.S.; Logacheva, M.D.; Penin, A.A. A high resolution map of the Arabidopsis thaliana developmental transcriptome based on RNA-seq profiling. Plant J. 2016, 88, 1058–1070. [Google Scholar] [CrossRef]
- Magori, S.; Citovsky, V. Hijacking of the host SCF ubiquitin ligase machinery by plant pathogens. Front. Plant Sci. 2011, 2, 87. [Google Scholar] [CrossRef]
- Gagne, J.M.; Downes, B.P.; Shiu, S.-H.; Durski, A.M.; Vierstra, R.D. The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc. Natl. Acad. Sci. USA 2002, 99, 11519–11524. [Google Scholar] [CrossRef] [PubMed]
- Hua, Z.; Vierstra, R.D. The cullin-RING ubiquitin-protein ligases. Annu. Rev. Plant Biol. 2011, 62, 299–334. [Google Scholar] [CrossRef]
- Anand, A.; Rojas, C.M.; Tang, Y.; Mysore, K.S. Several components of SKP1/Cullin/F-box E3 ubiquitin ligase complex and associated factors play a role in Agrobacterium-mediated plant transformation. New Phytol. 2012, 195, 203–216. [Google Scholar] [CrossRef] [PubMed]
- Zhao, F.; Chen, L.; Perl, A.; Chen, S.; Ma, H. Proteomic changes in grape embryogenic callus in response to Agrobacterium tumefaciens-mediated transformation. Plant Sci. 2011, 181, 485–495. [Google Scholar] [CrossRef] [PubMed]
- Hellens, R.P.; Allan, A.C.; Friel, E.N.; Bolitho, K.; Grafton, K.; Templeton, M.D.; Karunairetnam, S.; Gleave, A.P.; Laing, W.A. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 2005, 1, 13. [Google Scholar] [CrossRef]
- Gelvin, S.B. Agrobacterium Transformation of Arabidopsis thaliana Roots: A Quantitative Assay. Agrobacterium Protoc. 2006, 343, 105–114. [Google Scholar] [CrossRef]
- Li, J.; Vaidya, M.; White, C.I.; Vainstein, A.; Citovsky, V.; Tzfira, T. Involvement of KU80 in T-DNA integration in plant cells. Proc. Natl. Acad. Sci. USA 2005, 102, 19231–19236. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.D.; Cho, Y.H.; Sheen, J. Arabidopsis mesophyll protoplasts: A versatile cell system for transient gene expression analysis. Nat. Protoc. 2007, 2, 1565–1572. [Google Scholar] [CrossRef]
- Jones, R.W. Application of succulent plant leaves for Agrobacterium infiltration-mediated protein production. J. Microbiol. Methods 2016, 120, 65–67. [Google Scholar] [CrossRef]
- Wu, H.Y.; Liu, K.H.; Wang, Y.C.; Wu, J.F.; Chiu, W.L.; Chen, C.Y.; Wu, S.H.; Sheen, J.; Lai, E.M. AGROBEST: An efficient Agrobacterium-mediated transient expression method for versatile gene function analyses in Arabidopsis seedlings. Plant Methods 2014, 10, 19. [Google Scholar] [CrossRef]



© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, S.; Chen, Z.; Huang, F.; Wang, Y.; Luo, M. F-Box Gene D5RF Is Regulated by Agrobacterium Virulence Protein VirD5 and Essential for Agrobacterium-Mediated Plant Transformation. Int. J. Mol. Sci. 2020, 21, 6731. https://doi.org/10.3390/ijms21186731
Zhang S, Chen Z, Huang F, Wang Y, Luo M. F-Box Gene D5RF Is Regulated by Agrobacterium Virulence Protein VirD5 and Essential for Agrobacterium-Mediated Plant Transformation. International Journal of Molecular Sciences. 2020; 21(18):6731. https://doi.org/10.3390/ijms21186731
Chicago/Turabian StyleZhang, Shaojuan, Zhuo Chen, Fei Huang, Yafei Wang, and Meizhong Luo. 2020. "F-Box Gene D5RF Is Regulated by Agrobacterium Virulence Protein VirD5 and Essential for Agrobacterium-Mediated Plant Transformation" International Journal of Molecular Sciences 21, no. 18: 6731. https://doi.org/10.3390/ijms21186731
APA StyleZhang, S., Chen, Z., Huang, F., Wang, Y., & Luo, M. (2020). F-Box Gene D5RF Is Regulated by Agrobacterium Virulence Protein VirD5 and Essential for Agrobacterium-Mediated Plant Transformation. International Journal of Molecular Sciences, 21(18), 6731. https://doi.org/10.3390/ijms21186731




