Greatest Hits—Innovative Technologies for High Throughput Identification of Bispecific Antibodies
Abstract
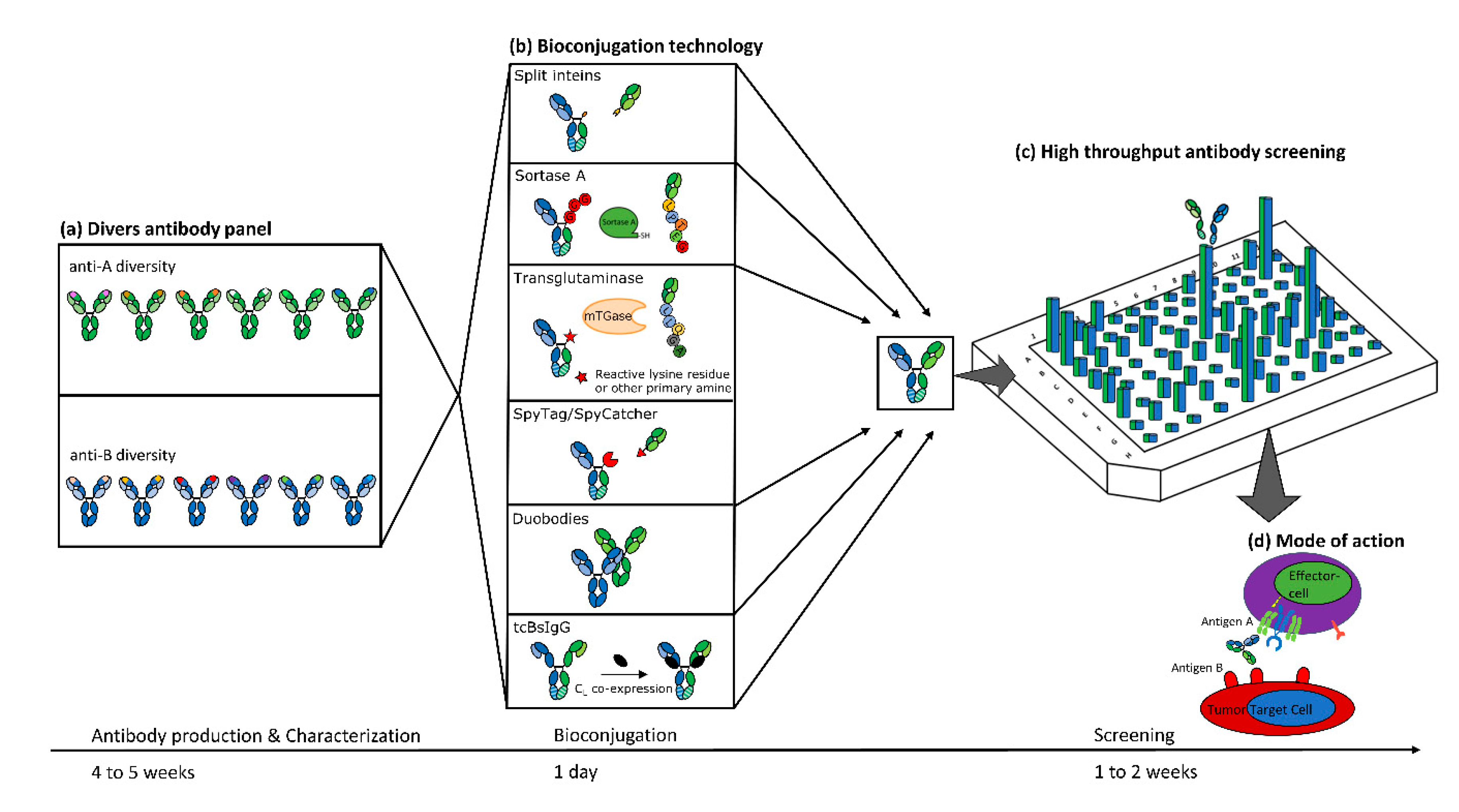
1. Introduction
2. Controlled Fab-Arm Exchange (“DuoBodies”)
3. Paired Light Chain Single Cell Production Approaches
4. Microbial Transglutaminase and Sortase A
5. The SpyTag/SpyCatcher System
6. Split Inteins
7. Conclusions and Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| aa | Amino acid |
| ADC | Antibody-Drug Conjugate |
| BLI | Biolayer interferometry |
| CL | Constant domain of a Light chain (of an antibody) |
| DARPin | Designed Ankyrin Repeat Proteins |
| Fab | Fragment antigen binding (of an antibody) |
| FAE | Fab-arm exchange |
| LC | (Antibody) light chain |
| LCMS | Liquid Chromatography-Mass Spectrometry |
| mTGase | Microbial Transglutaminase |
| MoA | Mode of Action |
| NSCLC | Non-Small Cell Lung Cancer |
| PTS | Protein Trans-Splicing |
| RPLC | Reverse Phase Liquid Chromatography |
| sdAb | Single domain Antibody |
| SrtA | Sortase A |
| TNF | Tumor Necrosis Factor |
| VHH | Variable Heavy chain domain of Heavy chain antibodies |
References
- Akira, S.; Uematsu, S.; Takeuchi, O. Pathogen Recognition and Innate Immunity. Cell 2006, 124, 783–801. [Google Scholar] [CrossRef] [PubMed]
- Janssens, S.; Beyaert, R. Role of Toll-Like Receptors in Pathogen Recognition. Clin. Microbiol. Rev. 2003, 16, 637–646. [Google Scholar] [CrossRef] [PubMed]
- Kang, T.H.; Jung, S.T. Boosting therapeutic potency of antibodies by taming Fc domain functions. Exp. Mol. Med. 2019, 51, 1–9. [Google Scholar] [CrossRef] [PubMed]
- De Taeye, S.W.; Rispens, T.; Vidarsson, G. The Ligands for Human IgG and Their Effector Functions. Antibodies 2019, 8, 30. [Google Scholar] [CrossRef]
- Kaplon, H.; Muralidharan, M.; Schneider, Z.; Reichert, J.M. Antibodies to watch in 2020. mAbs 2020, 12, 1703531. [Google Scholar] [CrossRef]
- Lu, R.M.; Hwang, Y.-C.; Liu, I.-J.; Lee, C.-C.; Tsai, H.-Z.; Li, H.-J.; Wu, H. Development of therapeutic antibodies for the treatment of diseases. J. Biomed. Sci. 2020, 27, 1. [Google Scholar] [CrossRef]
- Krah, S.; Kolmar, H.; Becker, S.; Zielonka, S. Engineering IgG-Like Bispecific Antibodies—An Overview. Antibodies 2018, 7, 28. [Google Scholar] [CrossRef]
- Wu, C.; Ying, H.; Grinnell, C.; Bryant, S.; Miller, R.; Clabbers, A.; Bose, S.; McCarthy, D.; Zhu, R.-R.; Santora, L.; et al. Simultaneous targeting of multiple disease mediators by a dual-variable-domain immunoglobulin. Nat. Biotechnol. 2007, 25, 1290–1297. [Google Scholar] [CrossRef]
- Labrijn, A.F.; Janmaat, M.L.; Reichert, J.M.; Parren, P.W.H.I. Bispecific antibodies: A mechanistic review of the pipeline. Nat. Rev. Drug Discov. 2019, 18, 585–608. [Google Scholar] [CrossRef]
- Sedykh, S.; Prinz, V.; Buneva, V.; Nevinsky, G. Bispecific antibodies: Design, therapy, perspectives. Drug Des. Dev. Ther. 2018, 12, 195–208. [Google Scholar] [CrossRef]
- Suurs, F.V.; Lub-de Hooge, M.N.; de Vries, E.G.E.; de Groot, D.J.A. A review of bispecific antibodies and antibody constructs in oncology and clinical challenges. Pharmacol. Ther. 2019, 201, 103–119. [Google Scholar] [CrossRef] [PubMed]
- Brinkmann, U.; Kontermann, R.E. The making of bispecific antibodies. mAbs 2017, 9, 182–212. [Google Scholar] [CrossRef] [PubMed]
- Doerner, A.; Rhiel, L.; Zielonka, S.; Kolmar, H. Therapeutic antibody engineering by high efficiency cell screening. FEBS Lett. 2014, 588, 278–287. [Google Scholar] [CrossRef] [PubMed]
- Frenzel, A.; Schirrmann, T.; Hust, M. Phage display-derived human antibodies in clinical development and therapy. mAbs 2016, 8, 1177–1194. [Google Scholar] [CrossRef]
- Frenzel, A.; Kügler, J.; Helmsing, S.; Meier, D.; Schirrmann, T.; Hust, M.; Dübel, S. Designing Human Antibodies by Phage Display. Transfus. Med. Hemother. 2017, 44, 312–318. [Google Scholar] [CrossRef]
- Boder, E.T.; Wittrup, K.D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 1997, 15, 553–557. [Google Scholar] [CrossRef]
- Rashidian, J.; Lloyd, J. Single B Cell Cloning and Production of Rabbit Monoclonal Antibodies. In Genotype Phenotype Coupling; Zielonka, S., Krah, S., Eds.; Springer: New York, NY, USA, 2020; Volume 2070, pp. 423–441. [Google Scholar]
- Kitazawa, T.; Igawa, T.; Sampei, Z.; Muto, A.; Kojima, T.; Soeda, T.; Yoshihashi, K.; Okuyama-Nishida, Y.; Saito, H.; Tsunoda, H.; et al. A bispecific antibody to factors IXa and X restores factor VIII hemostatic activity in a hemophilia a model. Nat. Med. 2012, 18, 1570–1574. [Google Scholar] [CrossRef]
- DiGiammarino, E.L.; Harlan, J.E.; Walter, K.A.; Ladror, U.S.; Edalji, R.P.; Hutchins, C.W.; Lake, M.R.; Greischar, A.J.; Liu, J.; Ghayur, T.; et al. Ligand association rates to the inner-variable-domain of a dual-variable-domain immunoglobulin are significantly impacted by linker design. mAbs 2011, 3, 487–494. [Google Scholar] [CrossRef]
- Mayer, K.; Baumann, A.-L.; Grote, M.; Seeber, S.; Kettenberger, H.; Breuer, S.; Killian, T.; Schäfer, W.; Brinkmann, U. TriFabs—Trivalent IgG-Shaped Bispecific Antibody Derivatives: Design, Generation, Characterization and Application for Targeted Payload Delivery. Int. J. Mol. Sci. 2015, 16, 27497–27507. [Google Scholar] [CrossRef]
- Lu, D.; Jimenez, X.; Witte, L.; Zhu, Z. The effect of variable domain orientation and arrangement on the antigen-binding activity of a recombinant human bispecific diabody. Biochem. Biophys. Res. Commun. 2004, 318, 507–513. [Google Scholar] [CrossRef]
- Labrijn, A.F.; Meesters, J.I.; de Goeij, B.E.C.G.; van den Bremer, E.T.J.; Neijssen, J.; van Kampen, M.D.; Strumane, K.; Verploegen, S.; Kundu, A.; Gramer, M.J.; et al. Efficient generation of stable bispecific IgG1 by controlled Fab-arm exchange. Proc. Natl. Acad. Sci. USA 2013, 110, 5145–5150. [Google Scholar] [CrossRef] [PubMed]
- Geuijen, C.A.W.; De Nardis, C.; Maussang, D.; Rovers, E.; Gallenne, T.; Hendriks, L.J.; Visser, T.; Nijhuis, R.; Logtenberg, T.; De Kruif, J.; et al. Unbiased Combinatorial Screening Identifies a Bispecific IgG1 that Potently Inhibits HER3 Signaling via HER2-Guided Ligand Blockade. Cancer Cell 2018, 33, 922–936.e10. [Google Scholar] [CrossRef] [PubMed]
- Plagmann, I.; Chalaris, A.; Kruglov, A.A.; Nedospasov, S.; Rosenstiel, P.; Rose-John, S.; Scheller, J. Transglutaminasecatalyzed covalent multimerization of Camelidae anti-human TNF single domain antibodies improves neutralizing activity. J. Biotechnol. 2009, 142, 170–178. [Google Scholar] [CrossRef] [PubMed]
- Andres, F.; Schwill, M.; Boersma, Y.L.; Plückthun, A. High-Throughput Generation of Bispecific Binding Proteins by Sortase A-mediated Coupling for direct functional screening in cell culture. Mol. Cancer Ther. 2020, 19, 1980–1988. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, A.R.; Alam, M.K.; Geyer, C.R. Post-translational Assembly of Protein Parts into Complex Devices by Using SpyTag/SpyCatcher Protein Ligase. Chembiochem 2019, 20, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Hatlem, D.; Trunk, T.; Linke, D.; Leo, J.C. Catching a SPY: Using the SpyCatcher-SpyTag and Related Systems for Labeling and Localizing Bacterial Proteins. Int. J. Mol. Sci. 2019, 20, 2129. [Google Scholar] [CrossRef]
- Han, L.; Chen, J.; Ding, K.; Zong, H.; Xie, Y.; Jiang, H.; Zhang, B.; Lu, H.; Yin, W.; Gilly, J.; et al. Efficient generation of bispecific IgG antibodies by split intein mediated protein trans-splicing system. Sci. Rep. 2017, 7, 8360. [Google Scholar] [CrossRef]
- van der Neut Kolfschoten, M.; Schuurman, J.; Losen, M.; Bleeker, W.K.; Martínez-Martínez, P.; Vermeulen, E.; den Bleker, T.H.; Wiegman, L.; Vink, T.; Aarden, L.A.; et al. Anti-inflammatory activity of human IgG4 antibodies by dynamic Fab arm exchange. Science 2007, 317, 1554–1557. [Google Scholar] [CrossRef]
- Ridgway, J.B.; Presta, L.G.; Carter, P. ‘Knobs-into-holes’ engineering of antibody CH3 domains for heavy chain heterodimerization. Protein Eng. 1996, 9, 617–621. [Google Scholar] [CrossRef]
- Schaefer, W.; Regula, J.T.; Bähner, M.; Schanzer, J.; Croasdale, R.; Dürr, H.; Gassner, C.; Georges, G.; Kettenberger, H.; Imhof-Jung, S.; et al. Immunoglobulin domain crossover as a generic approach for the production of bispecific IgG antibodies. Proc. Natl. Acad. Sci. USA 2011, 108, 11187–11192. [Google Scholar] [CrossRef]
- Krah, S.; Sellmann, C.; Rhiel, L.; Schröter, C.; Dickgiesser, S.; Beck, J.; Becker, S.; Toleikis, L.; Hock, B.; Kolmar, H.; et al. Engineering bispecific antibodies with defined chain pairing. N. Biotechnol. 2017, 39 Pt B, 167–173. [Google Scholar] [CrossRef]
- Chiu, M. How the Bispecific Antibody Targeting EGFR and cMET has superior preclinical activity and potentially better safety profile. In Proceedings of the Presentation at the 2nd Annual Antibody Summit: Discovery, Engineering & Therapeutics, Barcelona, Spain, 18–19 May 2017. [Google Scholar]
- Steinhardt, J.; Wu, Y.; Fleming, R.; Ruddle, B.T.; Patel, P.; Wu, H.; Gao, C.; DiMasi, N. Fab-Arm Exchange Combined with Selective Protein A Purification Results in a Platform for Rapid Preparation of Monovalent Bispecific Antibodies Directly from Culture Media. Pharmaceutics 2019, 12, 3. [Google Scholar] [CrossRef] [PubMed]
- Oostindie, S.C.; van der Horst, H.J.; Kil, L.P.; Strumane, K.; Overdijk, M.B.; Brink, E.N.V.D.; Brakel, J.H.N.V.D.; Rademaker, H.J.; Van Kessel, B.; Noort, J.V.D.; et al. DuoHexaBody-CD37®, a novel biparatopic CD37 antibody with enhanced Fc-mediated hexamerization as a potential therapy for B-cell malignancies. Blood Cancer J. 2020, 10, 30. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.S.; Dunshee, D.R.; Yee, A.; Tong, R.K.; Kim, I.; Farahi, F.; Hongo, J.-A.; Ernst, J.A.; Sonoda, J.; Spiess, C. Tethered-variable CL bispecific IgG: An antibody platform for rapid bispecific antibody screening. Protein Eng. Des. Sel. 2017, 30, 627–637. [Google Scholar] [CrossRef]
- Clarke, D.D.; Mycek, M.J.; Neidle, A.; Waelsch, H. The incorporation of amines into proteins. Arch. Biochem. Biophys. 1957, 79, 338–354. [Google Scholar] [CrossRef]
- Ando, H.; Adachi, M.; Umeda, K.; Matsuura, A.; Nonaka, M.; Uchio, R.; Tanaka, H.; Motoki, M. Purification and characteristics of a novel transglutaminase derived from microorganisms. Agric. Biol. Chem. 1989, 53, 2613–2617. [Google Scholar]
- Griffin, M.; Casadio, R.; Bergamini, C.M. Transglutaminases: Nature’s biological glues. Biochem. J. 2002, 368, 377–396. [Google Scholar] [CrossRef]
- Ohtsuka, T.; Ota, M.; Nio, N.; Motoki, M. Comparison of substrate specificities of transglutaminases using synthetic peptides as acyl donors. Biosci. Biotechnol. Biochem. 2000, 64, 2608–2613. [Google Scholar] [CrossRef]
- Jeger, S.; Zimmermann, K.; Blanc, A.; Grunberg, J.; Honer, M.; Hunziker, P.; Struthers, H.; Schibli, R. Site-specific and stoichiometric modification of antibodies by bacterial transglutaminase. Angew. Chem. Int. Ed. 2010, 49, 9995–9997. [Google Scholar] [CrossRef]
- Deweid, L.; Avrutina, O.; Kolmar, H. Microbial transglutaminase for biotechnological and biomedical engineering. Biol. Chem. 2019, 400, 257–274. [Google Scholar] [CrossRef]
- Strop, P.; Liu, S.H.; Dorywalska, M.; Delaria, K.; Dushin, R.G.; Tran, T.T.; Ho, W.H.; Farias, S.; Casas, M.G.; Abdiche, Y.; et al. Location matters: Site of conjugation modulates stability and pharmacokinetics of antibody drug conjugates. Chem. Biol. 2013, 20, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Siegmund, V.; Schmelz, S.; Dickgiesser, S.; Beck, J.; Ebenig, A.; Fittler, H.; Frauendorf, H.; Piater, B.; Betz, U.A.; Avrutina, O.; et al. Locked by design: A conformationally constrained transglutaminase tag enables efficient site-specific conjugation. Angew. Chem. Int. Ed. 2015, 54, 13420–13424. [Google Scholar] [CrossRef] [PubMed]
- Takazawa, T.; Kamiya, N.; Ueda, H.; Nagamune, T. Enzymatic labeling of a single chain variable fragment of an antibody with alkaline phosphatase by microbial transglutaminase. Biotechnol. Bioeng. 2004, 86, 399–404. [Google Scholar] [CrossRef] [PubMed]
- Kamiya, N.; Takazawa, T.; Tanaka, T.; Ueda, H.; Nagamune, T. Site-specific cross-linking of functional proteins by transglutamination. Enzyme Microb. Technol. 2003, 33, 492–496. [Google Scholar] [CrossRef]
- Mazmanian, S.K.; Liu, G.; Ton-That, H.; Schneewind, O. Staphylococcus aureus sortase, an enzyme that anchors surface proteins to the cell wall. Science 1999, 285, 760–763. [Google Scholar] [CrossRef]
- Huang, X.; Aulabaugh, A.; Ding, W.; Kapoor, B.; Alksne, L.; Tabie, K.; Ellestad, G. Kinetic mechanism of Staphylococcus aureaus Sortase Srt A. Biochemistry 2003, 42, 11307–11315. [Google Scholar] [CrossRef]
- Chen, I.; Dorr, B.M.; Liu, D.R. A general strategy for the evolution of bond-forming enzymes using yeast display. Proc. Natl. Acad. Sci. USA 2011, 108, 11399–11404. [Google Scholar] [CrossRef]
- Hirakawa, H.; Ishikawa, S.; Nagamune, T. Ca2+-independent sortase—A exhibits high selective protein ligation activity in the cytoplasm of Escherichia coli. Biotechnol. J. 2015, 10, 1487–1492. [Google Scholar] [CrossRef]
- Antos, J.M.; Truttmann, M.C.; Ploegh, H.L. Recent advances in sortase-catalyzed ligation methodology. Curr. Opin. Struct. Biol. 2016, 38, 111–118. [Google Scholar] [CrossRef]
- Schmohl, L.; Schwarzer, D. Sortase-mediated ligations for the site-specific modification of proteins. Curr. Opin. Chem. Biol. 2014, 22, 122–128. [Google Scholar] [CrossRef]
- Mao, H.; Hart, S.A.; Schink, A.; Pollok, B.A. Sortase-mediated protein ligation: A new method for protein engineering. J. Am. Chem Soc. 2004, 126, 2670–2671. [Google Scholar] [CrossRef] [PubMed]
- Popp, M.W.; Antos, J.M.; Grotenbreg, G.M.; Spooner, E.; Ploegh, H.L. Sortagging: A versatile method for protein labeling. Nat. Chem. Biol. 2007, 3, 707–708. [Google Scholar] [CrossRef] [PubMed]
- Levary, D.A.; Parthasarathy, R.; Boder, E.T.; Ackerman, M.E. Protein-Protein Fusion Catalyzed by Sortase A. PLoS ONE 2011, 6, e18342. [Google Scholar] [CrossRef] [PubMed]
- Popp, M.W.; Dougan, S.K.; Chuang, T.Y.; Spooner, E.; Ploegh, H.L. Sortase-catalyzed transformations that improve the properties of cytokines. Proc. Natl. Acad. Sci. USA 2011, 108, 3169–3174. [Google Scholar] [CrossRef]
- Madej, M.P.; Coia, G.; Williams, C.C.; Caine, J.M.; Pearce, L.A.; Attwood, R.; Bartone, N.A.; Dolezal, O.; Nisbet, R.M.; Nuttall, S.D.; et al. Engineering of an anti-epidermal growth factor receptor antibody to single chain format and labeling by sortase A-mediated protein ligation. Biotechnol. Bioeng. 2012, 109, 1461–1470. [Google Scholar] [CrossRef]
- Bartels, L.; Ploegh, H.L.; Spits, H.; Wagner, K. Preparation of bispecific antibody-protein adducts by site-specific chemoenzymatic conjugation. Methods 2019, 154, 93–101. [Google Scholar] [CrossRef]
- Wagner, K.; Kwakkenbos, M.J.; Claassen, Y.B.; Maijoor, K.; Bohne, M.; Van Der Sluijs, K.F.; Witte, M.D.; Van Zoelen, D.J.; Cornelissen, L.A.; Beaumont, T.; et al. Bispecific antibody generated with sortase and click chemistry has broad antiinfluenza virus activity. Proc. Natl. Acad. Sci. USA 2014, 111, 16820–16825. [Google Scholar] [CrossRef]
- Telford, J.L.; Barocchi, M.A.; Margarit, I.; Rappuoli, R.; Grandi, G. Pili in gram-positive pathogens. Nat. Rev. Microbiol. 2006, 4, 509–519. [Google Scholar] [CrossRef]
- Zakeri, B.; Howarth, M. Spontaneous intermolecular amide bond formation between side chains for irreversible peptide targeting. J. Am. Chem Soc. 2010, 132, 4526–4527. [Google Scholar] [CrossRef]
- Zakeri, B.; Fierer, J.O.; Celik, E.; Chittock, E.C.; Schwarz-Linek, U.; Moy, V.T.; Howarth, M. Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc. Natl. Acad. Sci. USA 2012, 109, E690–E697. [Google Scholar] [CrossRef]
- Veggiani, G.; Zakeri, B.; Howarth, M. Superglue from bacteria: Unbreakable bridges for protein nanotechnology. Trends Biotechnol. 2014, 32, 506–512. [Google Scholar] [CrossRef] [PubMed]
- Zakeri, B. Synthetic Biology: A New Tool for the Trade. Chembiochem 2015, 16, 2277–2282. [Google Scholar] [CrossRef] [PubMed]
- Keeble, A.H.; Banerjee, A.; Ferla, M.P.; Reddington, S.C.; Anuar, I.N.A.K.; Howarth, M. Evolving Accelerated Amidation by SpyTag/SpyCatcher to Analyze Membrane Dynamics. Angew. Chem. Int. Ed. 2017, 56, 16521–16525. [Google Scholar] [CrossRef] [PubMed]
- Si, M.; Xu, Q.; Jiang, L.; Huang, H. SpyTag/SpyCatcher Cyclization Enhances the Thermostability of Firefly Luciferase. PLoS ONE 2016, 11, e0162318. [Google Scholar] [CrossRef]
- Zhou, Y.B.; Cao, J.W.; Sun, X.B.; Wang, H.; Gao, D.-Y.; Li, Y.-N.; Wu, K.-Y.; Wang, J.-K.; Qian, G.-Y.; Wang, Q. Enhanced stability of a rumen-derived xylanase using SpyTag/SpyCatcher cyclization. World J. Microbiol. Biotechnol. 2020, 36, 33. [Google Scholar] [CrossRef]
- Yumura, K.; Akiba, H.; Nagatoishi, S.; Kusano-Arai, O.; Iwanari, H.; Hamakubo, T.; Tsumoto, K. Use of SpyTag/SpyCatcher to construct bispecific antibodies that target two epitopes of a single antigen. J. Biochem. 2017, 162, 203–210. [Google Scholar] [CrossRef]
- Alam, M.K.; Brabant, M.; Viswas, R.S.; Barreto, K.; Fonge, H.; Ronald Geyer, C. A novel synthetic trivalent single chain variable fragment (tri-scFv) construction platform based on the SpyTag/SpyCatcher protein ligase system. BMC Biotechnol. 2018, 18, 55. [Google Scholar] [CrossRef]
- Falck, G.; Müller, K.M. Enzyme-Based Labeling Strategies for Antibody-Drug Conjugates and Antibody Mimetics. Antibodies 2018, 7, 4. [Google Scholar] [CrossRef]
- Akiba, H.; Takayanagi, K.; Kusano-Arai, O.; Iwanari, H.; Hamakubo, T.; Tsumoto, K. Generation of biparatopic antibody through two-step targeting of fragment antibodies on antigen using SpyTag and SpyCatcher. Biotechnol. Rep. (Amst.) 2020, 25, e00418. [Google Scholar] [CrossRef]
- Singh, S.K.; Thrane, S.; Janitzek, C.M.; Nielsen, M.A.; Theander, T.G.; Theisen, M.; Salanti, A.; Sander, A.F. Improving the malaria transmission-blocking activity of a Plasmodium falciparum 48/45 based vaccine antigen by SpyTag/SpyCatcher mediated virus-like display. Vaccine 2017, 35, 3726–3732. [Google Scholar] [CrossRef]
- Li, D.; Zhang, H.; Yang, L.; Chen, J.; Zhang, Y.; Yu, X.; Zheng, Q.; Hou, J. Surface display of classical swine fever virus E2 glycoprotein on gram-positive enhancer matrix (GEM) particles via the SpyTag/SpyCatcher system. Protein Expr Purif. 2020, 167, 105526. [Google Scholar] [CrossRef] [PubMed]
- Okba, N.M.A.; Widjaja, I.; van Dieren, B.; Aebischer, A.; Van Amerongen, G.; De Waal, L.; Stittelaar, K.J.; Schipper, D.; Martina, B.; Brand, J.M.A.V.D.; et al. Particulate multivalent presentation of the receptor binding domain induces protective immune responses against MERS-CoV. Emerg. Microbes Infect. 2020, 9, 1080–1091. [Google Scholar] [CrossRef] [PubMed]
- Ji, M.; Zhu, J.; Xie, X.X.; Liu, D.-Q.; Wang, B.; Yu, Z.; Liu, R.-T. A novel rapid modularized hepatitis B core virus-like particle-based platform for personalized cancer vaccine preparation via fixed-point coupling. Nanomedicine 2020, 28, 102223. [Google Scholar] [CrossRef] [PubMed]
- Sharma, J.; Shepardson, K.; Johns, L.L.; Wellham, J.; Avera, J.; Schwarz, B.; Apple, A.R.; Douglas, T. A Self-Adjuvanted, Modular, Antigenic VLP for Rapid Response to Influenza Virus Variability. ACS Appl. Mater. Interfaces 2020, 12, 18211–18224. [Google Scholar] [CrossRef]
- Lin, Z.; Lin, Q.; Li, J.; Pistolozzi, M.; Zhao, L.; Yang, X.; Ye, Y. Spy chemistry-enabled protein directional immobilization and protein purification. Biotechnol. Bioeng. 2020. [Google Scholar] [CrossRef]
- Anuar, I.N.A.K.; Banerjee, A.; Keeble, A.H.; Carella, A.; Nikov, G.I.; Howarth, M. Spy & Go purification of SpyTag-proteins using pseudo-SpyCatcher to access an oligomerization toolbox. Nat. Commun. 2019, 10, 1734. [Google Scholar]
- Fierle, J.K.; Abram-Saliba, J.; Brioschi, M.; deTiani, M.; Coukos, G.; Dunn, S.M. Integrating SpyCatcher/SpyTag covalent fusion technology into phage display workflows for rapid antibody discovery. Sci. Rep. 2019, 9, 12815. [Google Scholar] [CrossRef]
- Alam, M.K.; El-Sayed, A.; Barreto, K.; Bernhard, W.; Fonge, H.; Geyer, C.R. Site-Specific Fluorescent Labeling of Antibodies and Diabodies Using SpyTag/SpyCatcher System for In Vivo Optical Imaging. Mol. Imaging Biol. 2019, 21, 54–66. [Google Scholar] [CrossRef]
- Bedbrook, C.N.; Kato, M.; Ravindra Kumar, S.; Lakshmanan, A.; Nath, R.D.; Sun, F.; Sternberg, P.W.; Arnold, F.H.; Gradinaru, V. Genetically Encoded Spy Peptide Fusion System to Detect Plasma Membrane-Localized Proteins In Vivo. Chem. Biol. 2015, 22, 1108–1121. [Google Scholar] [CrossRef]
- Lim, S.; Kim, J.; Kim, Y.; Xu, D.; Clark, D.S. CRISPR/Cas-directed programmable assembly of multi-enzyme complexes. Chem. Commun. (Camb.) 2020, 56, 4950–4953. [Google Scholar] [CrossRef]
- Veggiani, G.; Nakamura, T.; Brenner, M.D.; Gayet, R.; Yan, J.; Robinson, C.V.; Howarth, M. Programmable polyproteams built using twin peptide superglues. Proc. Natl. Acad. Sci. USA 2016, 113, 1202–1207. [Google Scholar] [CrossRef] [PubMed]
- Andersson, A.C.; Buldun, C.M.; Pattinson, D.J.; Draper, S.J.; Howarth, M. SnoopLigase peptide-peptide conjugation enables modular vaccine assembly. Sci. Rep. 2019, 9, 4625. [Google Scholar] [CrossRef] [PubMed]
- Bluemel, C.; Hausmann, S.; Fluhr, P.; Sriskandarajah, M.; Stallcup, W.B.; Baeuerle, P.A.; Kufer, P. Epitope distance to the target cell membrane and antigen size determine the potency of T cell-mediated lysis by BiTE antibodies specific for a large melanoma surface antigen. Cancer Immunol. Immunother. 2010, 59, 1197–1209. [Google Scholar] [CrossRef] [PubMed]
- Dickopf, S.; Georges, G.J.; Brinkmann, U. Format and geometries matter: Structure-based design defines the functionality of bispecific antibodies. Comput. Struct. Biotechnol. J. 2020, 18, 1221–1227. [Google Scholar] [CrossRef] [PubMed]
- Scott, M.J.; Lee, J.A.; Wake, M.S.; Batt, K.V.; Wattam, T.A.; Hiles, I.D.; Batuwangala, T.D.; Ashman, C.I.; Steward, M. ‘In-Format’ screening of a novel bispecific antibody format reveals significant potency improvements relative to unformatted molecules. mAbs 2017, 9, 85–93. [Google Scholar] [CrossRef]
- Li, L.; Fierer, J.O.; Rapoport, T.A.; Howarth, M. Structural analysis and optimization of the covalent association between SpyCatcher and a peptide Tag. J. Mol. Biol. 2014, 426, 309–317. [Google Scholar] [CrossRef]
- Gogarten, J.P.; Senejani, A.G.; Zhaxybayeva, O.; Olendzenski, L.; Hilario, E. Inteins: Structure, Function, and Evolution. Annu. Rev. Microbiol. 2002, 56, 263–287. [Google Scholar] [CrossRef]
- Aranko, A.S.; Volkmanna, G. Protein trans-splicing as a protein ligation tool to study protein structure and function. Biomol. Concepts 2011, 2, 183–198. [Google Scholar] [CrossRef]
- Wu, H.; Hu, Z.; Liu, X.Q. Protein trans-splicing by a split intein encoded in a split DnaE gene of Synechocystis sp. Proc. Natl. Acad. Sci. USA 1998, 95, 9226–9231. [Google Scholar] [CrossRef]
- Carvajal-Vallejos, P.; Palliss, R.; Mootz, H.D.; Schmidt, S.R. Unprecedented rates and efficiencies revealed for new natural split inteins from metagenomic sources. J. Biol. Chem. 2012, 287, 28686–28696. [Google Scholar] [CrossRef]
- Iwai, H.; Züger, S.; Jin, J.; Tam, P.H. Highly efficient protein trans-splicing by a naturally split DnaE intein from Nostoc punctiforme. FEBS Lett. 2006, 580, 1853–1858. [Google Scholar] [CrossRef] [PubMed]
- Zettler, J.; Schütz, V.; Mootz, H.D. The naturally split Npu DnaE intein exhibits an extraordinarily high rate in the protein trans-splicing reaction. FEBS Lett. 2009, 583, 909–914. [Google Scholar] [CrossRef] [PubMed]
- Miraula, M.; Enculescu, C.; Schenk, G.; Mitić, N. Inteins—A Focus on the Biotechnological Applications of Splicing-Promoting Proteins. Am. J. Mol. Biol. 2015, 5, 42–56. [Google Scholar] [CrossRef]
- Perler, F.B. Protein splicing mechanisms and applications. IUBMB Life 2005, 57, 469–476. [Google Scholar] [CrossRef]
- Friedel, K.; Popp, M.A.; Matern, J.C.J.; Gazdag, E.M.; Thiel, I.V.; Volkmann, G.; Blankenfeldt, W.; Mootz, H.D. Chemical Science A functional interplay between intein and extein sequences in protein splicing compensates for the essential block B histidine. Chem. Sci. 2019, 10, 239–251. [Google Scholar] [CrossRef]
- Vangelis, A.; Ouafâa, E.l.M.; Vincent, D.; Marine, C.; Jean-Christophe, M.M.; Oleg, M. Native Chemical Ligation and Extended Methods: Mechanisms, Catalysis, Scope, and Limitations. Chem. Rev. 2019, 119, 7328–7443. [Google Scholar]
- Liu, H.; May, K. Disulfide Bond Structures of IgG Molecules: Structural Variations, Chemical Modifications and Possible Impacts to Stability and Biological Function. mAbs 2012, 4, 17–23. [Google Scholar] [CrossRef]
- Ciragan, A.; Aranko, A.S.; Tascon, I.; Iwaï, H. Salt-inducible Protein Splicing in cis and trans by Inteins from Extremely Halophilic Archaea as a Novel Protein-Engineering Tool. J. Mol. Biol. 2016, 428, 4573–4588. [Google Scholar] [CrossRef]
- Bhagawati, M.; Terhorst, T.M.E.; Füsser, F.; Hoffmann, S.; Pasch, T.A. Mesophilic cysteine-less split intein for protein trans—Splicing applications under oxidizing conditions. Proc. Natl. Acad. Sci. USA 2019, 116, 22164–22172. [Google Scholar] [CrossRef]
- Topilina, N.I.; Mills, K.V. Recent advances in in vivo applications of intein-mediated protein splicing. Mob. DNA 2014, 5, 5. [Google Scholar] [CrossRef]
- Xu, Y.; Zhang, L.; Ma, B.; Hu, L.; Lu, H.; Dou, T.; Chen, J.; Zhu, J. Intermolecular disulfide bonds between unpaired cysteines retard the C-terminal trans-cleavage of Npu DnaE. Enzyme Microb. Technol. 2018, 118, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Zong, H.; Han, L.; Xie, Y.; Jiang, H.; Gilly, J.; Zhang, B.; Lu, H.; Chen, J.; Sun, R.; et al. A novel bispecific antibody targeting CD3 and prolactin receptor (PRLR) against PRLR-expression breast cancer. J. Exp. Clin. Cancer Res. 2020, 39, 87. [Google Scholar] [CrossRef] [PubMed]
- Hemmi, S.; Asano, R.; Kimura, K.; Umetsu, M.; Nakanishi, T.; Kumagai, I.; Makabe, K. Construction of a Circularly Connected VHH Bispecific Antibody (Cyclobody) for the Desirable Positioning of Antigen-Binding Sites. Biochem. Biophys. Res. Commun. 2020, 523, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Möhlmann, S.; Bringmann, P.; Greven, S.; Harrenga, A. Site-specific modification of ED-B-targeting antibody using intein-fusion technology. BMC Biotechnol. 2011, 11, 76. [Google Scholar] [CrossRef]
- Pirzer, T.; Becher, K.S.; Rieker, M.; Meckel, T.; Mootz, H.D.; Kolmar, H. Generation of Potent Anti-HER1/2 Immunotoxins by Protein Ligation Using Split Inteins. ACS Chem. Biol. 2018, 13, 2058–2066. [Google Scholar] [CrossRef]
- Hofmann, T.; Schmidt, J.; Ciesielski, E.; Becker, S.; Rysiok, T.; Schütte, M.; Toleikis, L.; Kolmar, H.; Doerner, A. Intein mediated high throughput screening for bispecific antibodies. mAbs 2020, 12, 1. [Google Scholar] [CrossRef]
- Busche, A.E.; Aranko, A.S.; Talebzadeh-Farooji, M.; Bernhard, F.; Dötsch, V.; Iwaï, H. Segmental isotopic labeling of a central domain in a multidomain protein by protein trans-splicing using only one robust DnaE intein. Angew. Chem. Int. Ed. 2009, 48, 6128–6131. [Google Scholar] [CrossRef]
- Pinto, F.; Thornton, E.L.; Wang, B. An expanded library of orthogonal split inteins enables modular multi-peptide assemblies. Nat. Commun. 2020, 11, 1529. [Google Scholar] [CrossRef]
- Suresh, M.R.; Cuello, A.C.; Milstein, C. Bispecific monoclonal antibodies from hybrid hybridomas. Methods Enzymol. 1986, 121, 210–228. [Google Scholar]
- Mezzanzanica, D.; Canevari, S.; Ménard, S.; Pupa, S.M.; Tagliabue, E.; Lanzavecchia, A.; Colnaghi, M.I. Human ovarian carcinoma lysis by cytotoxic T cells targeted by bispecific monoclonal antibodies: Analysis of the antibody components. Int. J. Cancer 1988, 41, 609–615. [Google Scholar] [CrossRef]
- Gilliland, L.K.; Clark, M.R.; Waldmann, H. Universal bispecific antibody for targeting tumor cells for destruction by cytotoxic T cells. Proc. Natl. Acad. Sci. USA 1988, 85, 7719–7723. [Google Scholar] [CrossRef] [PubMed]
- Krishnamurthy, A.; Jimeno, A. Bispecific antibodies for cancer therapy: A review. Pharm. Ther. 2018, 185, 122–134. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Wang, J. T cell-redirecting bispecific antibodies in cancer immunotherapy: Recent advances. J. Cancer Res. Clin. Oncol. 2019, 145, 941–956. [Google Scholar] [CrossRef]
- Spiess, C.; Zhai, Q.; Carter, P.J. Alternative molecular formats and therapeutic applications for bispecific antibodies. Mol. Immunol. 2015, 67, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Stieglmaier, J.; Benjamin, J.; Nagorsen, D. Utilizing the BiTE (bispecific T-cell engager) platform for immunotherapy of cancer. Expert Opin. Biol. Ther. 2015, 15, 1093–1099. [Google Scholar] [CrossRef] [PubMed]
- Cruz, E.; Kayser, V. Monoclonal antibody therapy of solid tumors: Clinical limitations and novel strategies to enhance treatment efficacy. Biologics 2019, 13, 33–51. [Google Scholar] [CrossRef] [PubMed]
- Duell, J.; Lammers, P.E.; Djuretic, I.; Chunyk, A.G.; Alekar, S.; Jacobs, I.; Gill, S. Bispecific Antibodies in the Treatment of Hematologic Malignancies. Clin. Pharmacol. Ther. 2019, 106, 781–791. [Google Scholar] [CrossRef]
- Lim, S.I. Site-specific bioconjugation and self-assembly technologies for multi-functional biologics: On the road to the clinic. Drug Discov. Today 2020, 25, 168–176. [Google Scholar] [CrossRef]
- Nie, S.; Wang, Z.; Moscoso-Castro, M.; D’Sousa, P.; Lei, C.; Xu, J.; Gu, J. Biology drives the discovery of bispecific antibodies as innovative therapeutics. Antibody Ther. 2020, 3, 18–62. [Google Scholar] [CrossRef]
- Züger, S.; Iwai, H. Intein-based biosynthetic incorporation of unlabeled protein tags into isotopically labeled proteins for NMR studies. Nat. Biotechnol. 2005, 23, 736–740. [Google Scholar] [CrossRef]
- Jaakkonen, A.; Volkmann, G.; Iwaï, H. An off-the-Shelf Approach for the Production of Fc Fusion Proteins by Protein Trans-Splicing towards Generating a Lectibody In Vitro. Int. J. Mol. Sci. 2020, 21, 4011. [Google Scholar] [CrossRef] [PubMed]
- Ling, W.L.; Lua, W.H.; Gan, S.K.E. Sagacity in antibody humanization for therapeutics, diagnostics and research purposes: Considerations of antibody elements and their roles. Antibody Ther. 2020, 3, 71–79. [Google Scholar] [CrossRef][Green Version]
| Technology | Linkage | Component Number | Handle of Motif | Residual Amino Acid Imprint | Activation Conditions | Reaction | Cofactors | Downstream Purification | HTS Compatibility | In Vivo Ligation | Covered Formats |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Duobodies | None | 2 | F405 and Y407 substitution in CH3 | Seamless | Mixed mAb precursor fragments under mild reducing conditions | Controlled Fab-arm exchange | Reducing agent, 2-MEA | Protein A, SEC | Yes | No | Full length-IgG format, DuoHexaBodies |
| Transglutaminase | C-terminal, N-terminal, site specific | 2 | LLQGA | LLQGA | Addition of Transglutaminase in high concentrations | Acyl-transfer | None | Protein A, SEC | No | No | Antibody drug conjugates |
| Sortase A | C-terminal, N-terminal, site specific | 2 | LPXTG | LPXTGGG | Excess of one reconstitution partner required based on a reversible reaction, high concentrations of Sortase A | Transpeptidation | None | IMAC | yes | Yes | Fc fusions, scFv, VHH |
| SpyTag/SpyCatcher | C-terminal, N-terminal | 3 | Lys in SpyCatcher, Asp in SpyTag | SpyTag/SpyCatcher | Isopeptide bond formation after bringing precursor proteins in close proximity | Amidation | None | Protein A, SEC | Yes | Yes | Full length-IgG format, Fc fusions, scFv, VHH |
| Split inteins | C-terminal, N-terminal | 3 or more | Int N and Int C | Extein sequences | Isopeptide bond formation after bringing precursor proteins in close proximity under mild reducing conditions | Protein trans splicing | Reducing agent TCEP, DTT if using cys containing split inteins | IMAC | Yes | Yes | Full length-IgG format, Fc fusions, scFv, VHH |
| Tethered variable CL or common LC bsAbs | None | 2 | VL-HC fusion (VLfH) or none | Short (Gly4Ser)4-linker or none | Intact BsAb generation in a single cell line by fusing the VL domain residues (1-R108) genetically to the antibody HC via (G4S)4 linker and CL co expression | co expression | None | ProteinA | yes if supernatant compatible | Not needed | Full-length-IgG format |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hofmann, T.; Krah, S.; Sellmann, C.; Zielonka, S.; Doerner, A. Greatest Hits—Innovative Technologies for High Throughput Identification of Bispecific Antibodies. Int. J. Mol. Sci. 2020, 21, 6551. https://doi.org/10.3390/ijms21186551
Hofmann T, Krah S, Sellmann C, Zielonka S, Doerner A. Greatest Hits—Innovative Technologies for High Throughput Identification of Bispecific Antibodies. International Journal of Molecular Sciences. 2020; 21(18):6551. https://doi.org/10.3390/ijms21186551
Chicago/Turabian StyleHofmann, Tim, Simon Krah, Carolin Sellmann, Stefan Zielonka, and Achim Doerner. 2020. "Greatest Hits—Innovative Technologies for High Throughput Identification of Bispecific Antibodies" International Journal of Molecular Sciences 21, no. 18: 6551. https://doi.org/10.3390/ijms21186551
APA StyleHofmann, T., Krah, S., Sellmann, C., Zielonka, S., & Doerner, A. (2020). Greatest Hits—Innovative Technologies for High Throughput Identification of Bispecific Antibodies. International Journal of Molecular Sciences, 21(18), 6551. https://doi.org/10.3390/ijms21186551





