In Vitro Assessment of the Genotoxic Hazard of Novel Hydroxamic Acid- and Benzamide-Type Histone Deacetylase Inhibitors (HDACi)
Abstract
1. Introduction
2. Results and Discussion
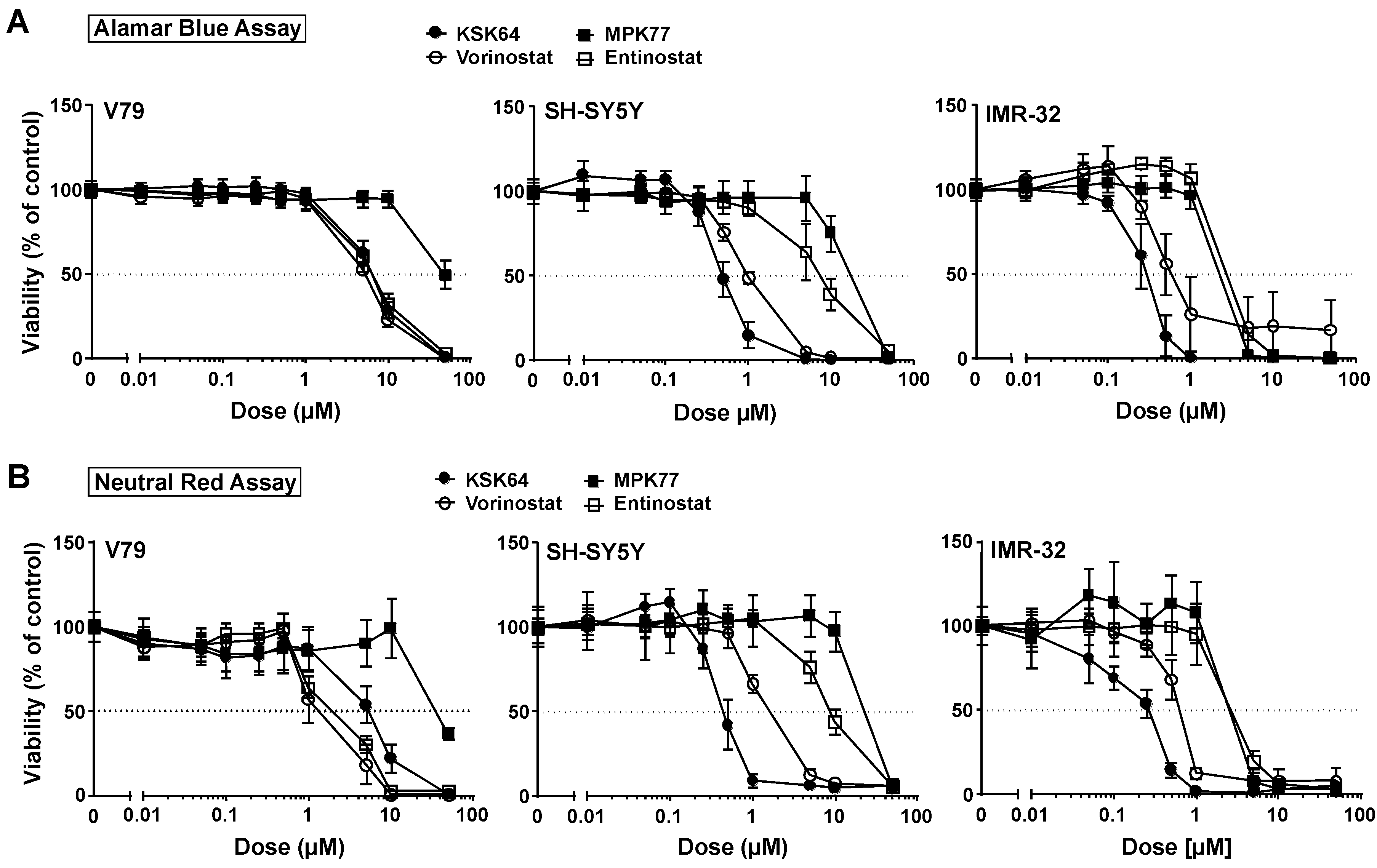
2.1. The Therapeutic Window of Novel Hydroxamic Acid- and Benzamide-Type HDACi
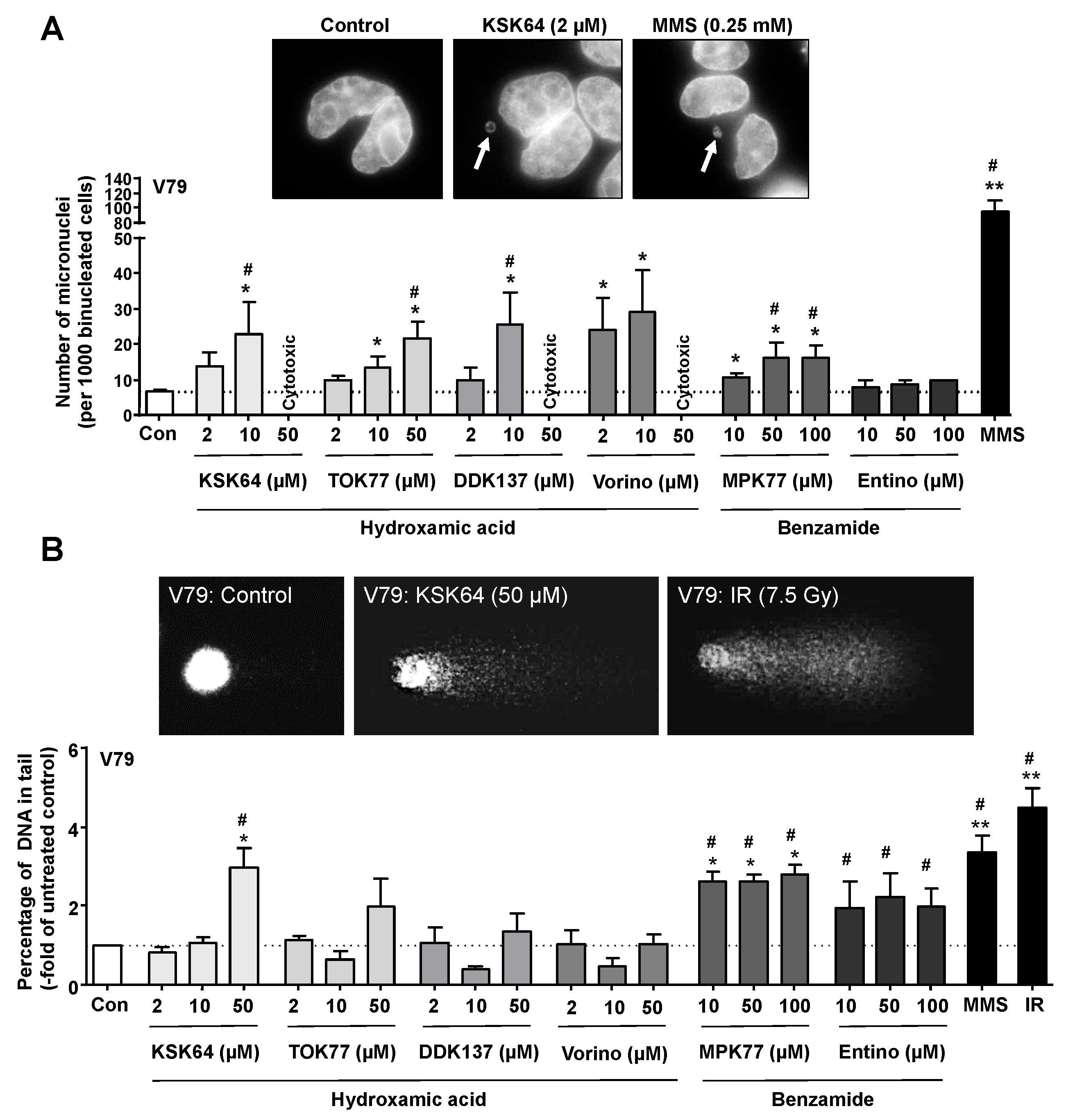
2.2. Formation of Micronuclei and DNA Strand Breaks by HDACi
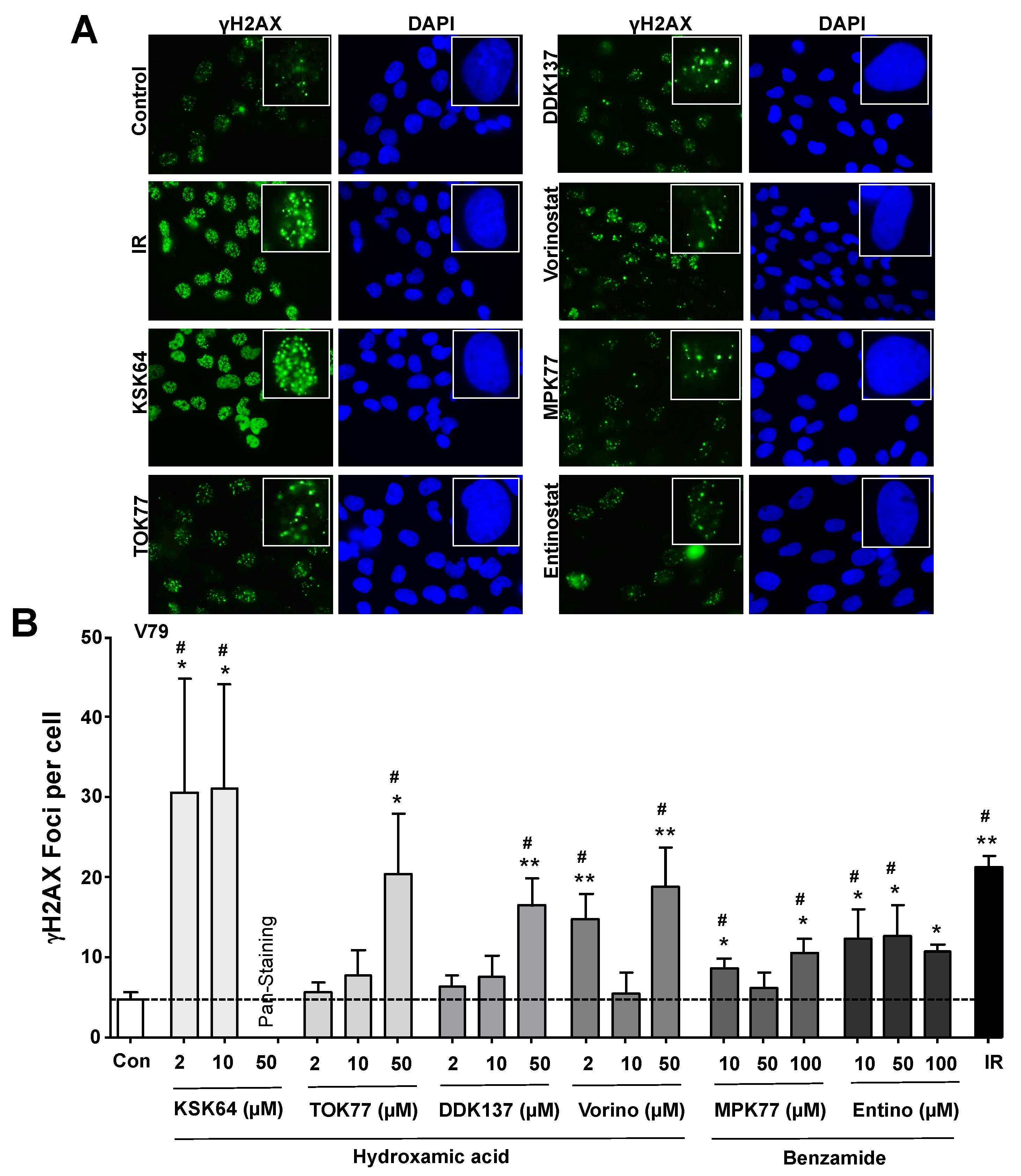
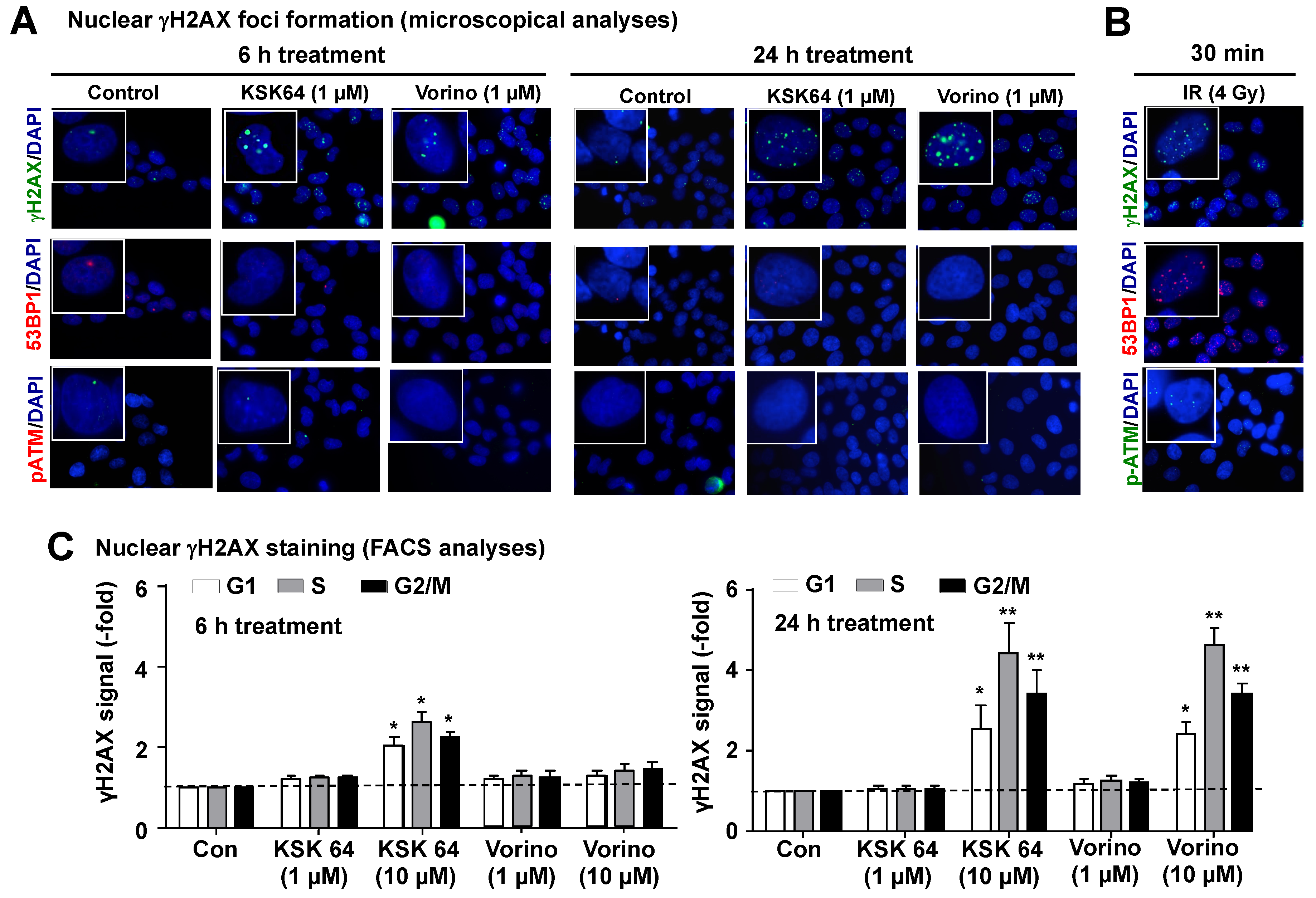
2.3. Influence of HDACi on the Formation of Nuclear γH2AX (Ser139 Phosphorylated Histone H2AX) Foci
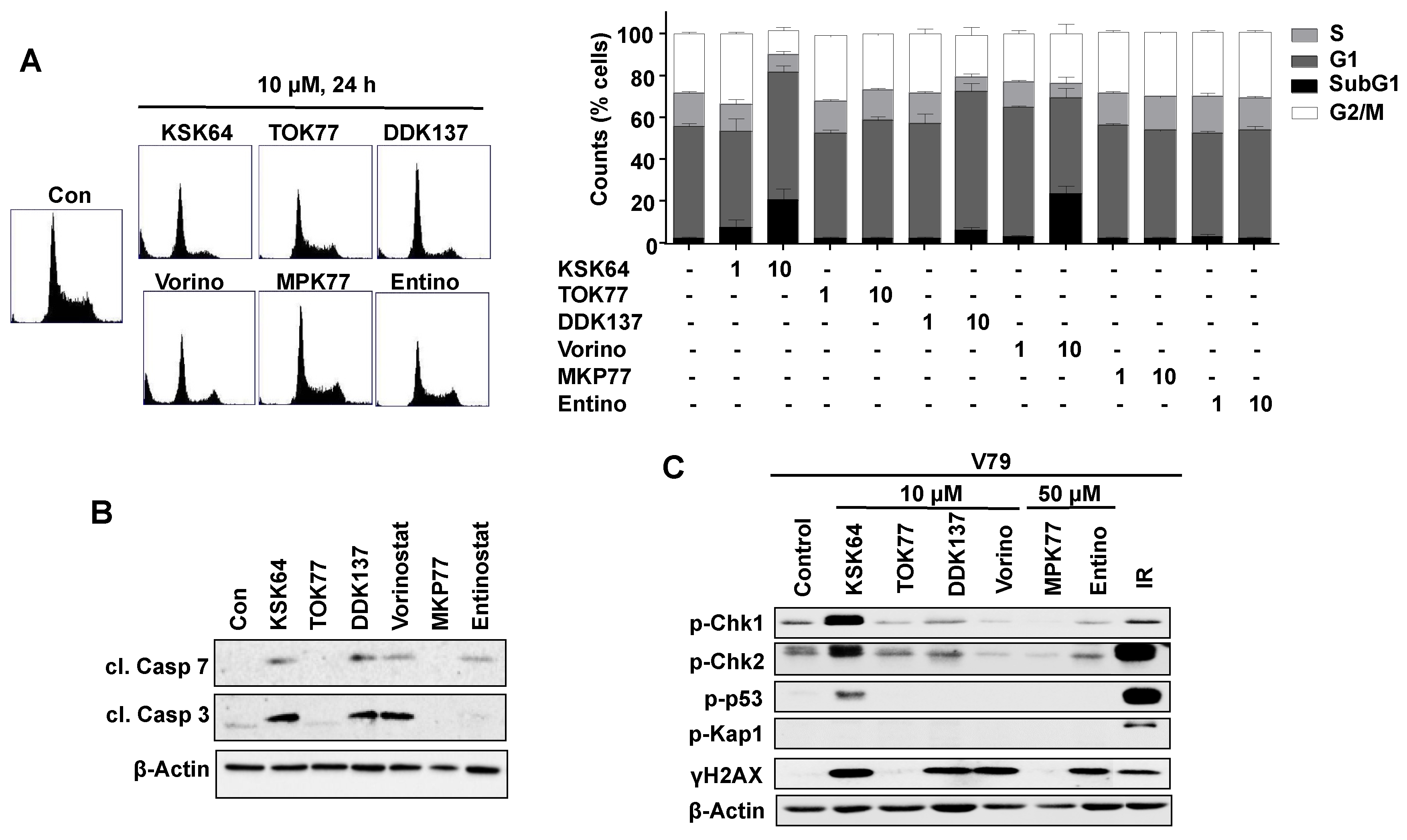
2.4. Influence of HDACi on the Activation of Cell Death Pathways and Mechanisms of the DNA Damage Response (DDR) in Non-Malignant V79 Cells
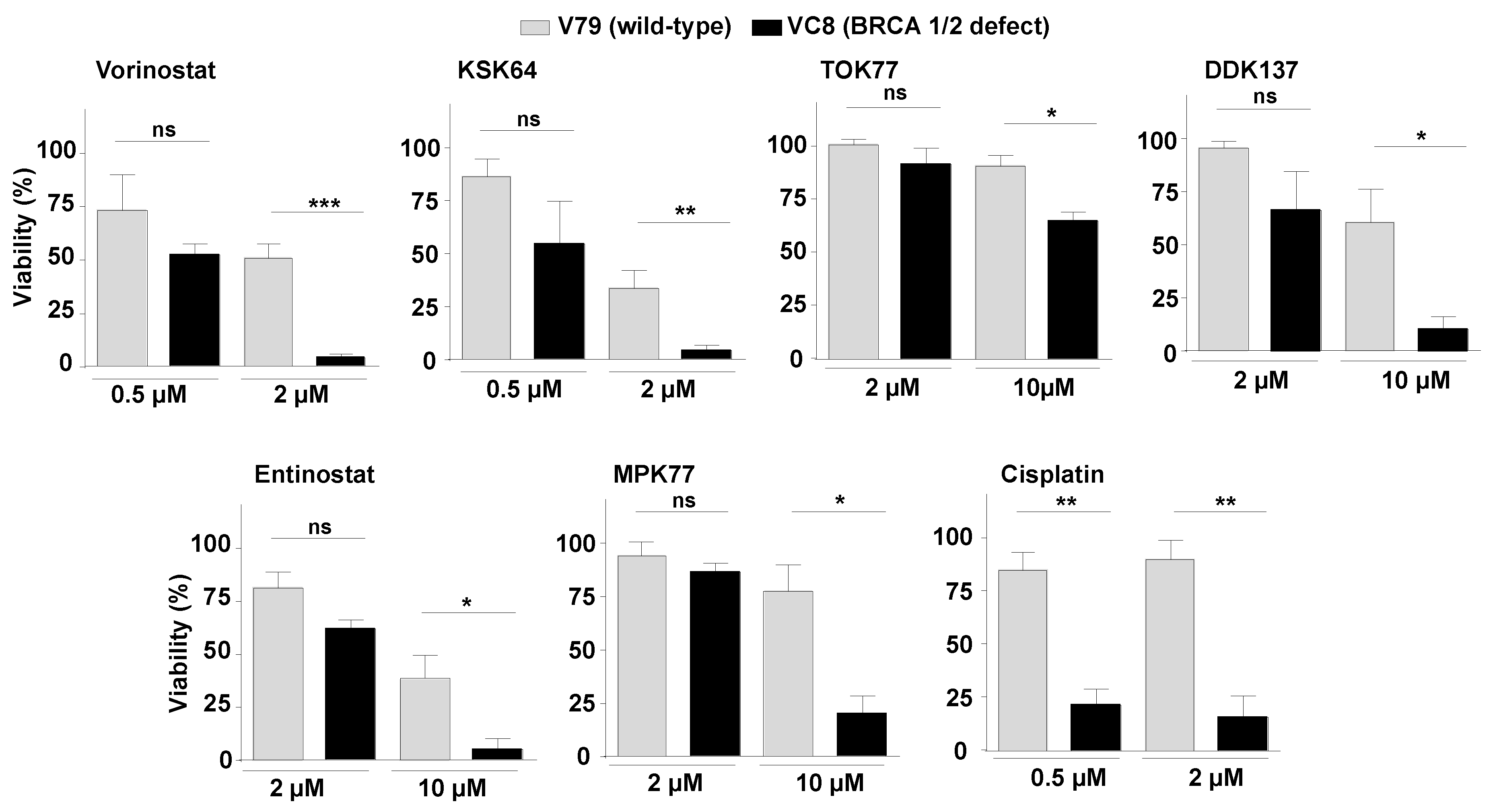
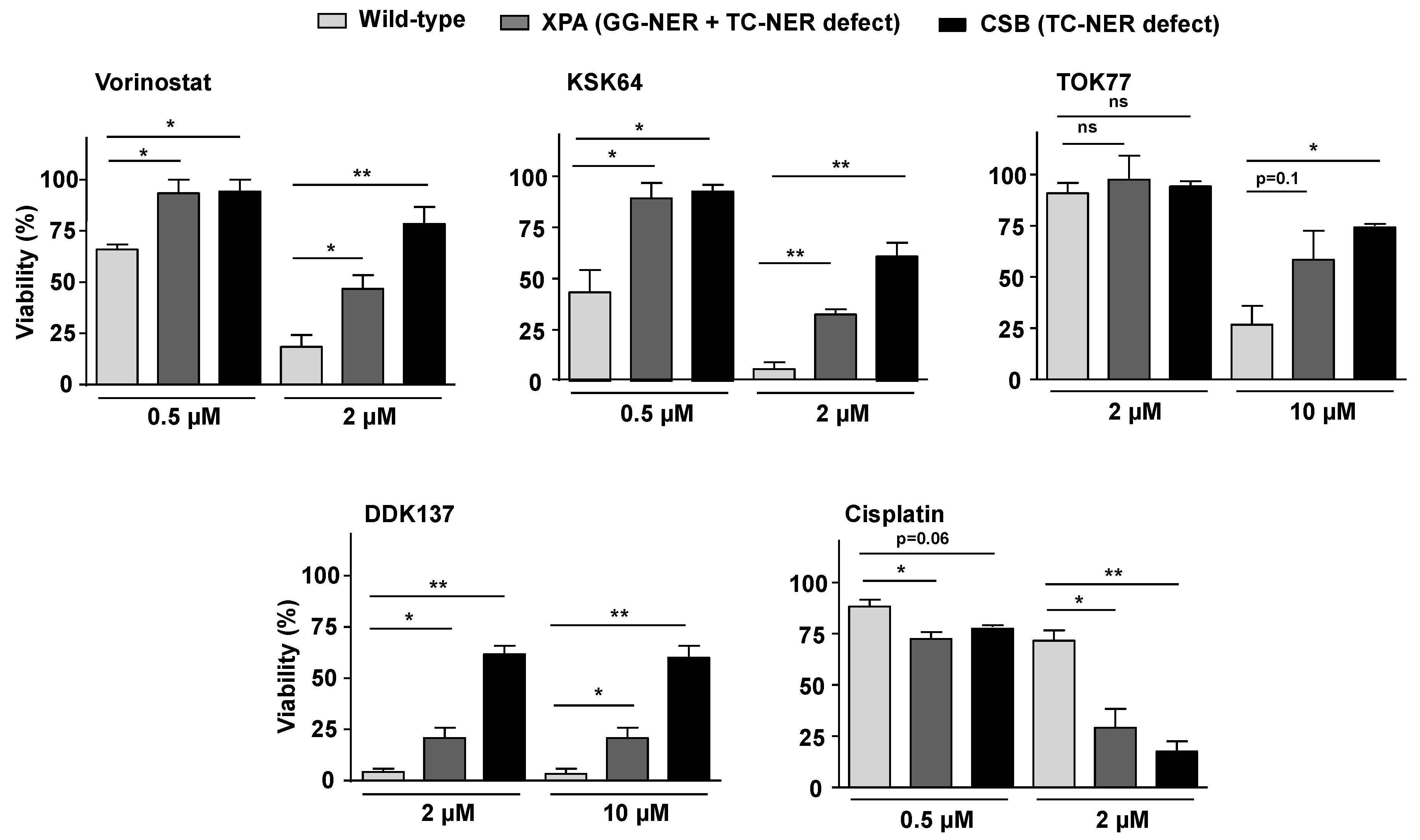
2.5. DSB Repair Defective Cells as a Tool to Predict the DSB-Inducing Potency of HDACi
3. Materials and Methods
3.1. Materials
3.2. Chemical Synthesis of HDACi—Reaction, Monitoring, Purification, and Analytics
3.3. Cell Culture and Treatment of Cells
3.4. Determination of Cell Viability and Cytotoxicity
3.5. Cell Cycle Analysis
3.6. Micronucleus Assay
3.7. Alkaline Comet Assay
3.8. Analysis of DNA Double-Strand Break Formation
3.9. Western Blot Analysis
3.10. Statistical Analyses
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ATM | Ataxia telangiectasia mutated |
| ATR | ATM- and Rad-3 related |
| BRCA1/2 | Breast cancer associated gene 1/2 |
| Chk1/2 | Checkpoint kinase 1/2 (CHEK1/2) |
| CSB | Cockayne syndrome protein B |
| DDR | DNA damage response |
| DSB | DNA double-strand breaks |
| ERK2 | Extracellular regulated kinase 2 |
| γH2AX | Ser139 phosphorylated histone H2AX |
| HDACi | Histone deacetylase inhibitor |
| HR | Homologous recombination |
| Kap1 | KRAB-associated protein 1 (TRIM28) |
| NER | Nucleotide excision repair |
| NHEJ | Non-homologous end-joining |
| XPA | Xeroderma pigmentosum complementation group A |
References
- Roos, W.P.; Krumm, A. The multifaceted influence of histone deacetylases on DNA damage signalling and DNA repair. Nucleic Acids Res. 2016, 44, 10017–10030. [Google Scholar] [CrossRef] [PubMed]
- Haberland, M.; Montgomery, R.L.; Olson, E.N. The many roles of histone deacetylases in development and physiology: Implications for disease and therapy. Nat. Rev. Genet. 2009, 10, 32–42. [Google Scholar] [CrossRef] [PubMed]
- Seto, E.; Yoshida, M. Erasers of histone acetylation: The histone deacetylase enzymes. Cold Spring Harb. Perspect. Biol. 2014, 6, a018713. [Google Scholar] [CrossRef]
- Wang, C.; Fu, M.; Mani, S.; Wadler, S.; Senderowicz Adrian, M.; Pestell Richard, G. Histone acetylation and the cell-cycle in cancer. Front. Biosci. 2001, 6, d610–d629. [Google Scholar] [CrossRef] [PubMed]
- Hadley, M.; Noonepalle, S.; Banik, D.; Villagra, A. Functional Analysis of HDACs in Tumorigenesis. Methods Mol. Biol. 2019, 1983, 279–307. [Google Scholar] [PubMed]
- Kachhap, S.K.; Rosmus, N.; Collis, S.J.; Kortenhorst, M.S.; Wissing, M.D.; Hedayati, M.; Shabbeer, S.; Mendonca, J.; Deangelis, J.; Marchionni, L.; et al. Downregulation of homologous recombination DNA repair genes by HDAC inhibition in prostate cancer is mediated through the E2F1 transcription factor. PLoS ONE 2010, 5, e11208. [Google Scholar] [CrossRef]
- Bolden, J.E.; Peart, M.J.; Johnstone, R.W. Anticancer activities of histone deacetylase inhibitors. Nat. Rev. Drug Discov. 2006, 5, 769–784. [Google Scholar] [CrossRef]
- Van Dyke, M.W. Lysine deacetylase (KDAC) regulatory pathways: An alternative approach to selective modulation. ChemMedChem 2014, 9, 511–522. [Google Scholar] [CrossRef]
- Nakagawa, M.; Oda, Y.; Eguchi, T.; Aishima, S.; Yao, T.; Hosoi, F.; Basaki, Y.; Ono, M.; Kuwano, M.; Tanaka, M.; et al. Expression profile of class I histone deacetylases in human cancer tissues. Oncol. Rep. 2007, 18, 769–774. [Google Scholar] [CrossRef]
- Lernoux, M.; Schnekenburger, M.; Dicato, M.; Diederich, M. Epigenetic mechanisms underlying the therapeutic effects of HDAC inhibitors in chronic myeloid leukemia. Biochem. Pharmacol. 2020, 173, 113698. [Google Scholar] [CrossRef]
- Bian, J.; Zhang, L.; Han, Y.; Wang, C.; Zhang, L. Histone deacetylase inhibitors: Potent anti-leukemic agents. Curr. Med. Chem. 2015, 22, 2065–2074. [Google Scholar] [CrossRef]
- Valente, S.; Mai, A. Small-molecule inhibitors of histone deacetylase for the treatment of cancer and non-cancer diseases: A patent review (2011–2013). Expert Opin. Ther. Pat. 2014, 24, 401–415. [Google Scholar] [CrossRef] [PubMed]
- Marks, P.A. The clinical development of histone deacetylase inhibitors as targeted anticancer drugs. Expert Opin. Investig. Drugs 2010, 19, 1049–1066. [Google Scholar] [CrossRef] [PubMed]
- Newbold, A.; Matthews, G.M.; Bots, M.; Cluse, L.A.; Clarke, C.J.; Banks, K.M.; Cullinane, C.; Bolden, J.E.; Christiansen, A.J.; Dickins, R.A.; et al. Molecular and biologic analysis of histone deacetylase inhibitors with diverse specificities. Mol. Cancer Ther. 2013, 12, 2709–2721. [Google Scholar] [CrossRef] [PubMed]
- Bishton, M.J.; Johnstone, R.W.; Dickinson, M.; Harrison, S.; Prince, H.M. Overview of Histone Deacetylase Inhibitors in Haematological Malignancies. Pharmaceuticals 2010, 3, 2674–2688. [Google Scholar] [CrossRef] [PubMed]
- De Souza, C.; Chatterji, B.P. HDAC Inhibitors as Novel Anti-Cancer Therapeutics. Recent Pat. Anticancer Drug Discov. 2015, 10, 145–162. [Google Scholar] [CrossRef]
- Vancurova, I.; Uddin, M.M.; Zou, Y.; Vancura, A. Combination Therapies Targeting HDAC and IKK in Solid Tumors. Trends Pharmacol. Sci. 2018, 39, 295–306. [Google Scholar] [CrossRef]
- Chun, P. Histone deacetylase inhibitors in hematological malignancies and solid tumors. Arch. Pharm. Res. 2015, 38, 933–949. [Google Scholar] [CrossRef]
- Banik, D.; Moufarrij, S.; Villagra, A. Immunoepigenetics Combination Therapies: An Overview of the Role of HDACs in Cancer Immunotherapy. Int. J. Mol. Sci. 2019, 20, 2241. [Google Scholar] [CrossRef]
- Asfaha, Y.; Schrenk, C.; Alves Avelar, L.A.; Hamacher, A.; Pflieger, M.; Kassack, M.U.; Kurz, T. Recent advances in class IIa histone deacetylases research. Bioorg. Med. Chem. 2019, 27, 115087. [Google Scholar] [CrossRef]
- Mottamal, M.; Zheng, S.; Huang, T.L.; Wang, G. Histone deacetylase inhibitors in clinical studies as templates for new anticancer agents. Molecules 2015, 20, 3898–3941. [Google Scholar] [CrossRef] [PubMed]
- Eckschlager, T.; Plch, J.; Stiborova, M.; Hrabeta, J. Histone Deacetylase Inhibitors as Anticancer Drugs. Int. J. Mol. Sci. 2017, 18, 1414. [Google Scholar] [CrossRef] [PubMed]
- Dickinson, M.; Johnstone, R.W.; Prince, H.M. Histone deacetylase inhibitors: Potential targets responsible for their anti-cancer effect. Invest. New Drugs 2010, 28 (Suppl. 1), 3–20. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, T.; Wu, W.; Okada, M.; Maeda, I.; Kojima, Y.; Hayami, R.; Miyoshi, Y.; Tsugawa, K.; Ohta, T. Class I histone deacetylase inhibitors inhibit the retention of BRCA1 and 53BP1 at the site of DNA damage. Cancer Sci. 2015, 106, 1050–1056. [Google Scholar] [CrossRef]
- Miller, K.M.; Tjeertes, J.V.; Coates, J.; Legube, G.; Polo, S.E.; Britton, S.; Jackson, S.P. Human HDAC1 and HDAC2 function in the DNA-damage response to promote DNA nonhomologous end-joining. Nat. Struct. Mol. Biol. 2010, 17, 1144–1151. [Google Scholar] [CrossRef]
- Nikolova, T.; Kiweler, N.; Kramer, O.H. Interstrand Crosslink Repair as a Target for HDAC Inhibition. Trends Pharmacol. Sci. 2017, 38, 822–836. [Google Scholar] [CrossRef]
- Zhang, M.; Xiang, S.; Joo, H.Y.; Wang, L.; Williams, K.A.; Liu, W.; Hu, C.; Tong, D.; Haakenson, J.; Wang, C.; et al. HDAC6 deacetylates and ubiquitinates MSH2 to maintain proper levels of MutSalpha. Mol. Cell 2014, 55, 31–46. [Google Scholar] [CrossRef]
- Goder, A.; Emmerich, C.; Nikolova, T.; Kiweler, N.; Schreiber, M.; Kuhl, T.; Imhof, D.; Christmann, M.; Heinzel, T.; Schneider, G.; et al. HDAC1 and HDAC2 integrate checkpoint kinase phosphorylation and cell fate through the phosphatase-2A subunit PR130. Nat. Commun. 2018, 9, 764. [Google Scholar] [CrossRef]
- Gomez-Gonzalez, B.; Ortega, P.; Aguilera, A. Histone deacetylases facilitate the accurate repair of broken forks. Mol. Cell. Oncol. 2020, 7, 1705731. [Google Scholar] [CrossRef]
- Stenzel, K.; Hamacher, A.; Hansen, F.K.; Gertzen, C.G.W.; Senger, J.; Marquardt, V.; Marek, L.; Marek, M.; Romier, C.; Remke, M.; et al. Alkoxyurea-Based Histone Deacetylase Inhibitors Increase Cisplatin Potency in Chemoresistant Cancer Cell Lines. J. Med. Chem. 2017, 60, 5334–5348. [Google Scholar] [CrossRef]
- Xie, C.; Drenberg, C.; Edwards, H.; Caldwell, J.T.; Chen, W.; Inaba, H.; Xu, X.; Buck, S.A.; Taub, J.W.; Baker, S.D.; et al. Panobinostat enhances cytarabine and daunorubicin sensitivities in AML cells through suppressing the expression of BRCA1, CHK1, and Rad51. PLoS ONE 2013, 8, e79106. [Google Scholar] [CrossRef] [PubMed]
- Gerelchuluun, A.; Maeda, J.; Manabe, E.; Brents, C.A.; Sakae, T.; Fujimori, A.; Chen, D.J.; Tsuboi, K.; Kato, T.A. Histone Deacetylase Inhibitor Induced Radiation Sensitization Effects on Human Cancer Cells after Photon and Hadron Radiation Exposure. Int. J. Mol. Sci. 2018, 19, 496. [Google Scholar] [CrossRef] [PubMed]
- Asfaha, Y.; Schrenk, C.; Alves Avelar, L.A.; Lange, F.; Wang, C.; Bandolik, J.J.; Hamacher, A.; Kassack, M.U.; Kurz, T. Novel alkoxyamide-based histone deacetylase inhibitors reverse cisplatin resistance in chemoresistant cancer cells. Bioorg. Med. Chem. 2020, 28, 115108. [Google Scholar] [CrossRef] [PubMed]
- Groselj, B.; Sharma, N.L.; Hamdy, F.C.; Kerr, M.; Kiltie, A.E. Histone deacetylase inhibitors as radiosensitisers: Effects on DNA damage signalling and repair. Br. J. Cancer 2013, 108, 748–754. [Google Scholar] [CrossRef]
- Feingold, P.L.; Surman, D.R.; Brown, K.; Xu, Y.; McDuffie, L.A.; Shukla, V.; Reardon, E.S.; Crooks, D.R.; Trepel, J.B.; Lee, S.; et al. Induction of Thioredoxin-Interacting Protein by a Histone Deacetylase Inhibitor, Entinostat, Is Associated with DNA Damage and Apoptosis in Esophageal Adenocarcinoma. Mol. Cancer Ther. 2018, 17, 2013–2023. [Google Scholar] [CrossRef]
- Stiborova, M.; Eckschlager, T.; Poljakova, J.; Hrabeta, J.; Adam, V.; Kizek, R.; Frei, E. The synergistic effects of DNA-targeted chemotherapeutics and histone deacetylase inhibitors as therapeutic strategies for cancer treatment. Curr. Med. Chem. 2012, 19, 4218–4238. [Google Scholar] [CrossRef] [PubMed]
- Cerna, T.; Hrabeta, J.; Eckschlager, T.; Frei, E.; Schmeiser, H.H.; Arlt, V.M.; Stiborova, M. The Histone Deacetylase Inhibitor Valproic Acid Exerts a Synergistic Cytotoxicity with the DNA-Damaging Drug Ellipticine in Neuroblastoma Cells. Int. J. Mol. Sci. 2018, 19, 164. [Google Scholar] [CrossRef]
- Suraweera, A.; O’Byrne, K.J.; Richard, D.J. Combination Therapy With Histone Deacetylase Inhibitors (HDACi) for the Treatment of Cancer: Achieving the Full Therapeutic Potential of HDACi. Front Oncol 2018, 8, 92. [Google Scholar] [CrossRef]
- Wawruszak, A.; Luszczki, J.J.; Kalafut, J.; Okla, K.; Halasa, M.; Rivero-Muller, A.; Stepulak, A. Additive Pharmacological Interaction between Cisplatin (CDDP) and Histone Deacetylase Inhibitors (HDIs) in MDA-MB-231 Triple Negative Breast Cancer (TNBC) Cells with Altered Notch1 Activity-An Isobolographic Analysis. Int. J. Mol. Sci. 2019, 20, 3663. [Google Scholar] [CrossRef]
- Shen, S.; Kozikowski, A.P. Why Hydroxamates May Not Be the Best Histone Deacetylase Inhibitors--What Some May Have Forgotten or Would Rather Forget? ChemMedChem 2016, 11, 15–21. [Google Scholar] [CrossRef]
- Habibian, J.; Ferguson, B.S. The Crosstalk between Acetylation and Phosphorylation: Emerging New Roles for HDAC Inhibitors in the Heart. Int. J. Mol. Sci. 2018, 20, 102. [Google Scholar] [CrossRef] [PubMed]
- Thomas, E.A.; D’Mello, S.R. Complex neuroprotective and neurotoxic effects of histone deacetylases. J. Neurochem. 2018, 145, 96–110. [Google Scholar] [CrossRef] [PubMed]
- Dobbin, M.M.; Madabhushi, R.; Pan, L.; Chen, Y.; Kim, D.; Gao, J.; Ahanonu, B.; Pao, P.C.; Qiu, Y.; Zhao, Y.; et al. SIRT1 collaborates with ATM and HDAC1 to maintain genomic stability in neurons. Nat. Neurosci. 2013, 16, 1008–1015. [Google Scholar] [CrossRef] [PubMed]
- De Simone, A.; Milelli, A. Histone Deacetylase Inhibitors as Multitarget Ligands: New Players in Alzheimer’s Disease Drug Discovery? ChemMedChem 2019, 14, 1067–1073. [Google Scholar] [CrossRef]
- Teijido, O.; Cacabelos, R. Pharmacoepigenomic Interventions as Novel Potential Treatments for Alzheimer’s and Parkinson’s Diseases. Int. J. Mol. Sci. 2018, 19, 3199. [Google Scholar] [CrossRef]
- von Knethen, A.; Heinicke, U.; Weigert, A.; Zacharowski, K.; Brune, B. Histone Deacetylation Inhibitors as Modulators of Regulatory T Cells. Int. J. Mol. Sci. 2020, 21, 2356. [Google Scholar] [CrossRef]
- Yoon, S.; Eom, G.H. HDAC and HDAC Inhibitor: From Cancer to Cardiovascular Diseases. Chonnam Med. J. 2016, 52, 1–11. [Google Scholar] [CrossRef]
- Rotili, D.; Simonetti, G.; Savarino, A.; Palamara, A.T.; Migliaccio, A.R.; Mai, A. Non-cancer uses of histone deacetylase inhibitors: Effects on infectious diseases and beta-hemoglobinopathies. Curr. Top. Med. Chem. 2009, 9, 272–291. [Google Scholar] [CrossRef]
- Xu, K.; Dai, X.L.; Huang, H.C.; Jiang, Z.F. Targeting HDACs: A promising therapy for Alzheimer’s disease. Oxid. Med. Cell. Longev. 2011, 2011, 143269. [Google Scholar] [CrossRef]
- Ecker, J.; Oehme, I.; Mazitschek, R.; Korshunov, A.; Kool, M.; Hielscher, T.; Kiss, J.; Selt, F.; Konrad, C.; Lodrini, M.; et al. Targeting class I histone deacetylase 2 in MYC amplified group 3 medulloblastoma. Acta Neuropathol. Commun. 2015, 3, 22. [Google Scholar] [CrossRef]
- Attia, S.M.; Al-Hamamah, M.A.; Alotaibi, M.R.; Harisa, G.I.; Attia, M.M.; Ahmad, S.F.; Ansari, M.A.; Nadeem, A.; Bakheet, S.A. Investigation of belinostat-induced genomic instability by molecular cytogenetic analysis and pathway-focused gene expression profiling. Toxicol. Appl. Pharmacol. 2018, 350, 43–51. [Google Scholar] [CrossRef]
- Lee, M.S.; Isobe, M. Metabolic activation of the potent mutagen, 2-naphthohydroxamic acid, in Salmonella typhimurium TA98. Cancer Res. 1990, 50, 4300–4307. [Google Scholar] [PubMed]
- Roos, W.P.; Thomas, A.D.; Kaina, B. DNA damage and the balance between survival and death in cancer biology. Nat. Rev. Cancer 2016, 16, 20–33. [Google Scholar] [CrossRef] [PubMed]
- Harper, J.W.; Elledge, S.J. The DNA damage response: Ten years after. Mol. Cell 2007, 28, 739–745. [Google Scholar] [CrossRef]
- Shiloh, Y.; Ziv, Y. The ATM protein kinase: Regulating the cellular response to genotoxic stress, and more. Nat. Rev. Mol. Cell Biol. 2013, 14, 197–210. [Google Scholar] [CrossRef] [PubMed]
- Olive, P.L. Detection of DNA damage in individual cells by analysis of histone H2AX phosphorylation. Methods Cell Biol. 2004, 75, 355–373. [Google Scholar] [PubMed]
- Marti, T.M.; Hefner, E.; Feeney, L.; Natale, V.; Cleaver, J.E. H2AX phosphorylation within the G1 phase after UV irradiation depends on nucleotide excision repair and not DNA double-strand breaks. Proc. Natl. Acad. Sci. USA 2006, 103, 9891–9896. [Google Scholar] [CrossRef]
- Kelly, W.K.; O’Connor, O.A.; Krug, L.M.; Chiao, J.H.; Heaney, M.; Curley, T.; MacGregore-Cortelli, B.; Tong, W.; Secrist, J.P.; Schwartz, L.; et al. Phase I study of an oral histone deacetylase inhibitor, suberoylanilide hydroxamic acid, in patients with advanced cancer. J. Clin. Oncol. 2005, 23, 3923–3931. [Google Scholar] [CrossRef]
- Zimmermann, M.; de Lange, T. 53BP1: Pro choice in DNA repair. Trends Cell Biol. 2014, 24, 108–117. [Google Scholar] [CrossRef]
- Fradet-Turcotte, A.; Canny, M.D.; Escribano-Diaz, C.; Orthwein, A.; Leung, C.C.; Huang, H.; Landry, M.C.; Kitevski-LeBlanc, J.; Noordermeer, S.M.; Sicheri, F.; et al. 53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark. Nature 2013, 499, 50–54. [Google Scholar] [CrossRef]
- Bunting, S.F.; Callen, E.; Wong, N.; Chen, H.T.; Polato, F.; Gunn, A.; Bothmer, A.; Feldhahn, N.; Fernandez-Capetillo, O.; Cao, L.; et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 2010, 141, 243–254. [Google Scholar] [CrossRef] [PubMed]
- Chapman, J.R.; Sossick, A.J.; Boulton, S.J.; Jackson, S.P. BRCA1-associated exclusion of 53BP1 from DNA damage sites underlies temporal control of DNA repair. J. Cell Sci. 2012, 125 Pt 15, 3529–3534. [Google Scholar] [CrossRef]
- Lopez-Martinez, D.; Liang, C.C.; Cohn, M.A. Cellular response to DNA interstrand crosslinks: The Fanconi anemia pathway. Cell. Mol. Life Sci. 2016, 73, 3097–3114. [Google Scholar] [CrossRef] [PubMed]
- Paulsen, R.D.; Cimprich, K.A. The ATR pathway: Fine-tuning the fork. DNA Repair 2007, 6, 953–966. [Google Scholar] [CrossRef] [PubMed]
- Redon, C.E.; Nakamura, A.J.; Martin, O.A.; Parekh, P.R.; Weyemi, U.S.; Bonner, W.M. Recent developments in the use of gamma-H2AX as a quantitative DNA double-strand break biomarker. Aging (Albany NY) 2011, 3, 168–174. [Google Scholar] [CrossRef] [PubMed]
- Sedelnikova, O.A.; Pilch, D.R.; Redon, C.; Bonner, W.M. Histone H2AX in DNA damage and repair. Cancer Biol. Ther. 2003, 2, 233–235. [Google Scholar]
- de Feraudy, S.; Revet, I.; Bezrookove, V.; Feeney, L.; Cleaver, J.E. A minority of foci or pan-nuclear apoptotic staining of gammaH2AX in the S phase after UV damage contain DNA double-strand breaks. Proc. Natl. Acad. Sci. USA 2010, 107, 6870–6875. [Google Scholar] [CrossRef]
- Roos, W.P.; Kaina, B. DNA damage-induced cell death: From specific DNA lesions to the DNA damage response and apoptosis. Cancer Lett. 2013, 332, 237–248. [Google Scholar] [CrossRef]
- Nikolova, T.; Ensminger, M.; Lobrich, M.; Kaina, B. Homologous recombination protects mammalian cells from replication-associated DNA double-strand breaks arising in response to methyl methanesulfonate. DNA Repair 2010, 9, 1050–1063. [Google Scholar] [CrossRef]
- Nagasawa, H.; Wilson, P.F.; Chen, D.J.; Thompson, L.H.; Bedford, J.S.; Little, J.B. Low doses of alpha particles do not induce sister chromatid exchanges in bystander Chinese hamster cells defective in homologous recombination. DNA Repair 2008, 7, 515–522. [Google Scholar] [CrossRef][Green Version]
- Christmann, M.; Tomicic, M.T.; Roos, W.P.; Kaina, B. Mechanisms of human DNA repair: An update. Toxicology 2003, 193, 3–34. [Google Scholar] [CrossRef]
- Friedberg, E.C.; Wallner, G.C.; Siede, W. DNA Repair and Mutagenesis; ASM Press: Washington, DC, USA, 1995. [Google Scholar]
- Sugitani, N.; Voehler, M.W.; Roh, M.S.; Topolska-Wos, A.M.; Chazin, W.J. Analysis of DNA binding by human factor xeroderma pigmentosum complementation group A (XPA) provides insight into its interactions with nucleotide excision repair substrates. J. Biol. Chem. 2017, 292, 16847–16857. [Google Scholar] [CrossRef] [PubMed]
- de Boer, J.; Hoeijmakers, J.H. Nucleotide excision repair and human syndromes. Carcinogenesis 2000, 21, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Le Page, F.; Kwoh, E.E.; Avrutskaya, A.; Gentil, A.; Leadon, S.A.; Sarasin, A.; Cooper, P.K. Transcription-coupled repair of 8-oxoguanine: Requirement for XPG, TFIIH, and CSB and implications for Cockayne syndrome. Cell 2000, 101, 159–171. [Google Scholar] [CrossRef]
- Zhao, J.; Xie, C.; Edwards, H.; Wang, G.; Taub, J.W.; Ge, Y. Histone deacetylases 1 and 2 cooperate in regulating BRCA1, CHK1, and RAD51 expression in acute myeloid leukemia cells. Oncotarget 2017, 8, 6319–6329. [Google Scholar] [CrossRef]
- Fahrer, J.; Kaina, B. Impact of DNA repair on the dose-response of colorectal cancer formation induced by dietary carcinogens. Food Chem. Toxicol. 2017, 106 Pt B, 583–594. [Google Scholar] [CrossRef]
- Fahrer, J.; Frisch, J.; Nagel, G.; Kraus, A.; Dorsam, B.; Thomas, A.D.; Reissig, S.; Waisman, A.; Kaina, B. DNA repair by MGMT, but not AAG, causes a threshold in alkylation-induced colorectal carcinogenesis. Carcinogenesis 2015, 36, 1235–1244. [Google Scholar] [CrossRef]
- Porter, N.J.; Osko, J.D.; Diedrich, D.; Kurz, T.; Hooker, J.M.; Hansen, F.K.; Christianson, D.W. Histone Deacetylase 6-Selective Inhibitors and the Influence of Capping Groups on Hydroxamate-Zinc Denticity. J. Med. Chem. 2018, 61, 8054–8060. [Google Scholar] [CrossRef]
- Pflieger, M.; Hamacher, A.; Oz, T.; Horstick-Muche, N.; Boesen, B.; Schrenk, C.; Kassack, M.U.; Kurz, T. Novel alpha,beta-unsaturated hydroxamic acid derivatives overcome cisplatin resistance. Bioorg. Med. Chem. 2019, 27, 115036. [Google Scholar] [CrossRef]
- Mackwitz, M.K.W.; Hesping, E.; Antonova-Koch, Y.; Diedrich, D.; Woldearegai, T.G.; Skinner-Adams, T.; Clarke, M.; Scholer, A.; Limbach, L.; Kurz, T.; et al. Structure-Activity and Structure-Toxicity Relationships of Peptoid-Based Histone Deacetylase Inhibitors with Dual-Stage Antiplasmodial Activity. ChemMedChem 2019, 14, 912–926. [Google Scholar] [CrossRef]
- Krieger, V.; Hamacher, A.; Cao, F.; Stenzel, K.; Gertzen, C.G.W.; Schaker-Hubner, L.; Kurz, T.; Gohlke, H.; Dekker, F.J.; Kassack, M.U.; et al. Synthesis of peptoid-based class I selective histone deacetylase inhibitors with chemosensitizing properties. J. Med. Chem. 2019, 62, 11260–11279. [Google Scholar] [CrossRef] [PubMed]
- Damrot, J.; Helbig, L.; Roos, W.P.; Barrantes, S.Q.; Kaina, B.; Fritz, G. DNA replication arrest in response to genotoxic stress provokes early activation of stress-activated protein kinases (SAPK/JNK). J. Mol. Biol. 2009, 385, 1409–1421. [Google Scholar] [CrossRef] [PubMed]
- Fritz, G.; Kaina, B. Late activation of stress kinases (SAPK/JNK) by genotoxins requires the DNA repair proteins DNA-PKcs and CSB. Mol. Biol. Cell 2006, 17, 851–861. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.; Wilson, I.; Orton, T.; Pognan, F. Investigation of the Alamar Blue (resazurin) fluorescent dye for the assessment of mammalian cell cytotoxicity. Eur. J. Biochem. 2000, 267, 5421–5426. [Google Scholar] [CrossRef] [PubMed]
- Doherty, A.T. The in vitro micronucleus assay. Methods Mol. Biol. 2012, 817, 121–141. [Google Scholar]
- Olive, P.L.; Banath, J.P. The comet assay: A method to measure DNA damage in individual cells. Nat. Protoc. 2006, 1, 23–29. [Google Scholar] [CrossRef]
- Rogakou, E.P.; Pilch, D.R.; Orr, A.H.; Ivanova, V.S.; Bonner, W.M. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 1998, 273, 5858–5868. [Google Scholar] [CrossRef]
- Nikolova, T.; Dvorak, M.; Jung, F.; Adam, I.; Kramer, E.; Gerhold-Ay, A.; Kaina, B. The gammaH2AX assay for genotoxic and nongenotoxic agents: Comparison of H2AX phosphorylation with cell death response. Toxicol. Sci. 2014, 140, 103–117. [Google Scholar] [CrossRef]
- Nikolova, T.; Marini, F.; Kaina, B. Genotoxicity testing: Comparison of the gammaH2AX focus assay with the alkaline and neutral comet assays. Mutat. Res. 2017, 822, 10–18. [Google Scholar] [CrossRef]
- Panier, S.; Boulton, S.J. Double-strand break repair: 53BP1 comes into focus. Nat. Rev. Mol. Cell Biol. 2014, 15, 7–18. [Google Scholar] [CrossRef]
| HDACi | Comet Assay | Micronucl. Assay | gH2AX Foci | gH2AX Pan-Stain | DDR Activation | DSB Repair def. Cells | Apoptosis | Viability (IC50) 24 h - 72 h |
|---|---|---|---|---|---|---|---|---|
| Vorinostat | − | + | + | ++ | + | ++ | ++ | 10 - 5 |
| KSK64 | ++ | + | ++ | +++ | ++ | ++ | ++ | 18 - 7 |
| TOK77 | − | + | + | 0 | − | + | − | >50 - >50 |
| DDK137 | − | + | + | + | + | + | + | 33 - 11 |
| Entinostat | − | - | + | + | + | + | 0 | >50 - 7 |
| MPK77 | + | + | + | 0 | − | + | − | >50 - 50 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Friedrich, A.; Assmann, A.-S.; Schumacher, L.; Stuijvenberg, J.v.; Kassack, M.U.; Schulz, W.A.; Roos, W.P.; Hansen, F.K.; Pflieger, M.; Kurz, T.; et al. In Vitro Assessment of the Genotoxic Hazard of Novel Hydroxamic Acid- and Benzamide-Type Histone Deacetylase Inhibitors (HDACi). Int. J. Mol. Sci. 2020, 21, 4747. https://doi.org/10.3390/ijms21134747
Friedrich A, Assmann A-S, Schumacher L, Stuijvenberg Jv, Kassack MU, Schulz WA, Roos WP, Hansen FK, Pflieger M, Kurz T, et al. In Vitro Assessment of the Genotoxic Hazard of Novel Hydroxamic Acid- and Benzamide-Type Histone Deacetylase Inhibitors (HDACi). International Journal of Molecular Sciences. 2020; 21(13):4747. https://doi.org/10.3390/ijms21134747
Chicago/Turabian StyleFriedrich, Annabelle, Ann-Sophie Assmann, Lena Schumacher, Jana v. Stuijvenberg, Matthias U. Kassack, Wolfgang A. Schulz, Wynand P. Roos, Finn K. Hansen, Marc Pflieger, Thomas Kurz, and et al. 2020. "In Vitro Assessment of the Genotoxic Hazard of Novel Hydroxamic Acid- and Benzamide-Type Histone Deacetylase Inhibitors (HDACi)" International Journal of Molecular Sciences 21, no. 13: 4747. https://doi.org/10.3390/ijms21134747
APA StyleFriedrich, A., Assmann, A.-S., Schumacher, L., Stuijvenberg, J. v., Kassack, M. U., Schulz, W. A., Roos, W. P., Hansen, F. K., Pflieger, M., Kurz, T., & Fritz, G. (2020). In Vitro Assessment of the Genotoxic Hazard of Novel Hydroxamic Acid- and Benzamide-Type Histone Deacetylase Inhibitors (HDACi). International Journal of Molecular Sciences, 21(13), 4747. https://doi.org/10.3390/ijms21134747





