Host Immune Response and Novel Diagnostic Approach to NTM Infections
Abstract
1. Introduction
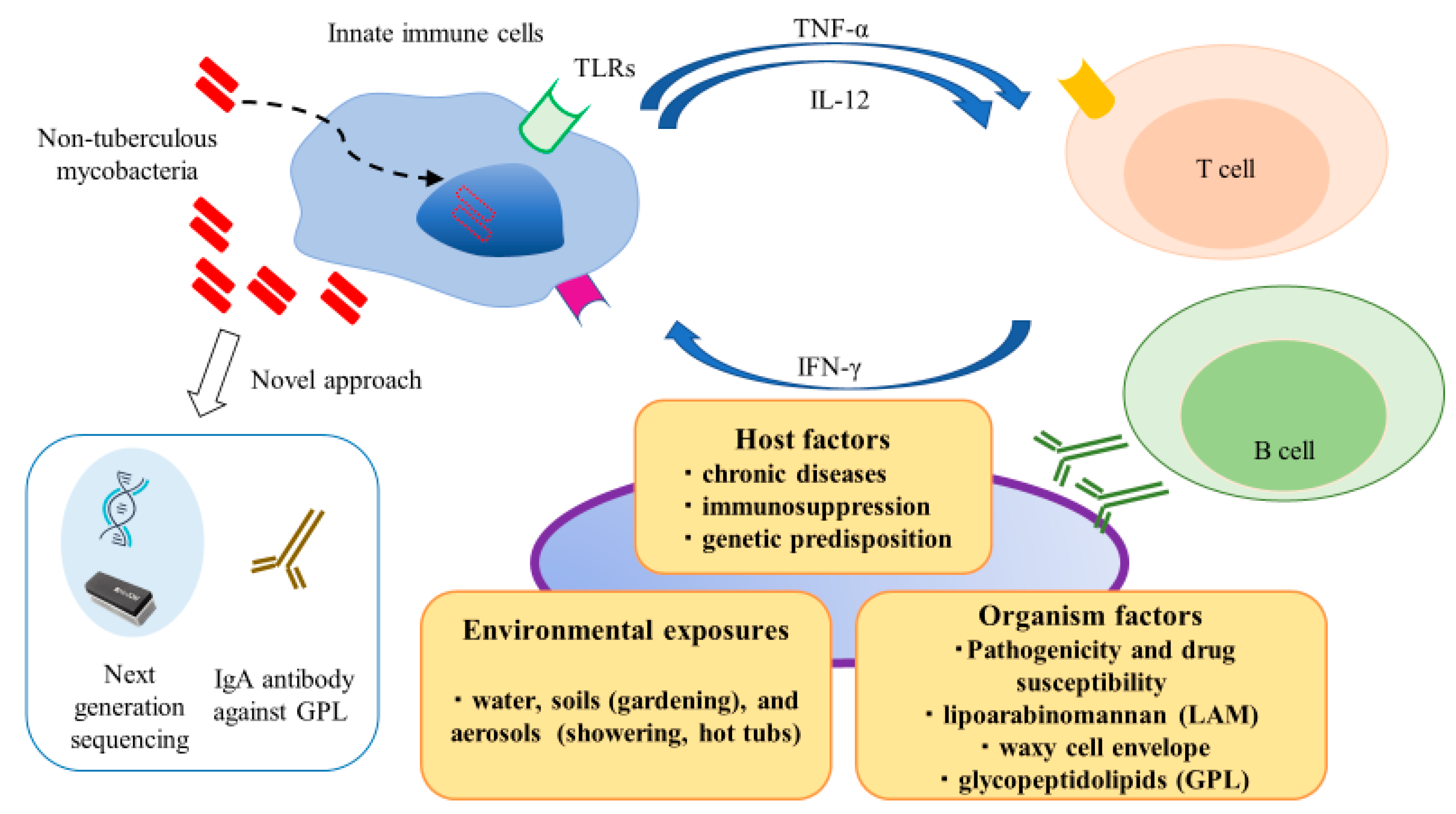
2. Innate Immune Response to NTM Infection
3. Cellular-Mediated Immune Responses to NTM Infection
NTM Infections Affect Immunocompromised Organ Transplanted Patients
4. Humoral Immune Response to NTM Infection
5. Novel Diagnostic Approach to NTM Infections
5.1. Serodiagnosis for Mycobacterium Avium Complex Pulmonary Disease
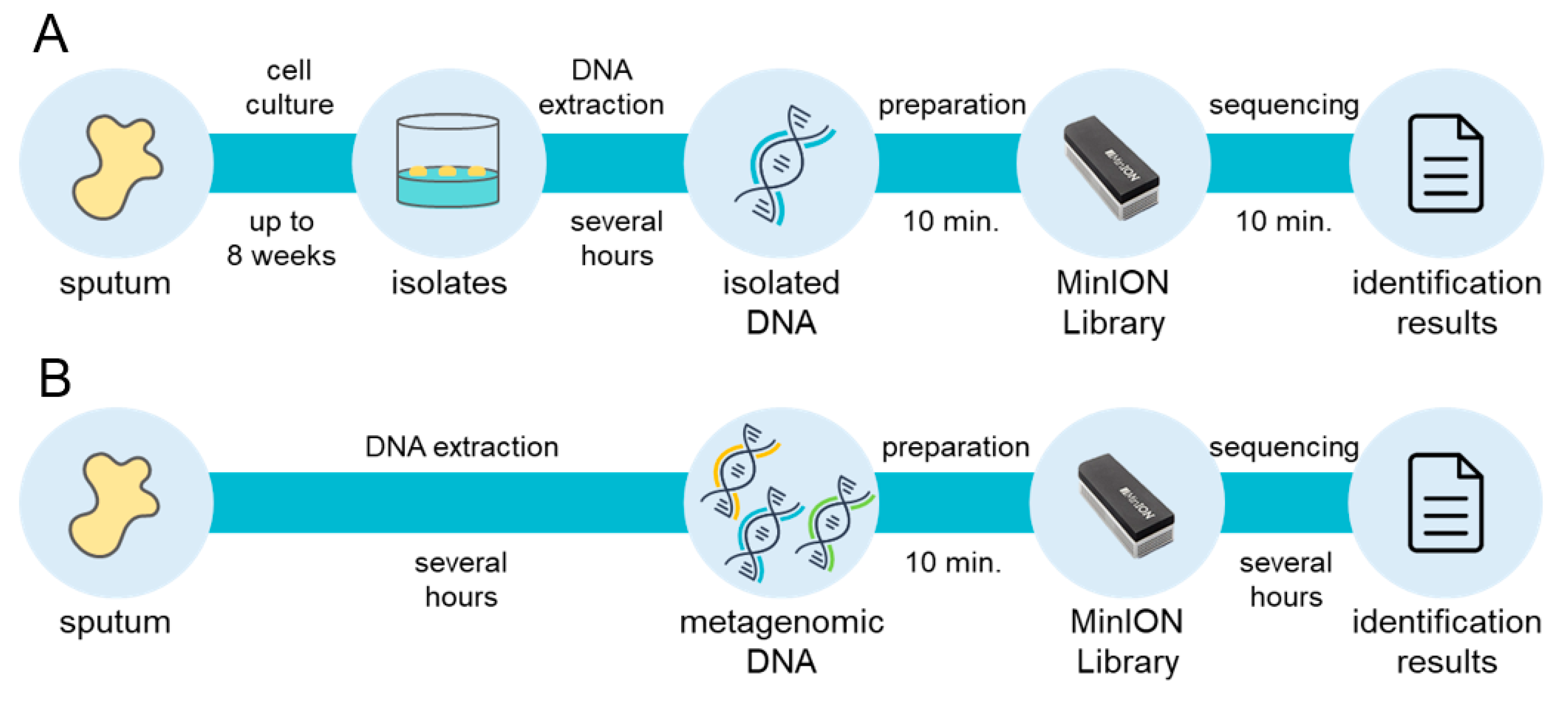
5.2. Rapid and Comprehensive Identification of NTM Species by MLST Database
6. Conclusion and Future Direction
Funding
Acknowledgments
Conflicts of Interest
References
- Falkinham, J.O., III. Ecology of Nontuberculous Mycobacteria—Where Do Human Infections Come from? Semin. Respir. Crit. Care Med. 2013, 34, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Jennifer, A.; Timothy, B.F.; Yihe, G.D.; Jennifer, R.H.; Kenneth, N.O.; Adrian, Z.; Stacey, H.D.; Rebecca, P. Epidemiology of Nontuberculous Mycobacterial Lung Disease and Tuberculosis, Hawaii, USA. Emerg. Infect. Dis. 2017, 23, 439–447. [Google Scholar]
- Honda, J.R.; Alper, S.; Bai, X.; Chan, E.D. Acquired and genetic host susceptibility factors and microbial pathogenic factors that predispose to nontuberculous mycobacterial infections. Curr. Opin. Immunol. 2018, 54, 66–73. [Google Scholar] [CrossRef]
- Gochi, M.; Takayanagi, N.; Kanauchi, T.; Ishiguro, T.; Yanagisawa, T.; Sugita, Y. Retrospective study of the predictors of mortality and radiographic deterioration in 782 patients with nodular/bronchiectatic Mycobacterium avium complex lung disease. BMJ Open. 2015, 5, e008058. [Google Scholar] [CrossRef]
- Fukushima, K.; Kitada, S.; Abe, Y.; Yamamoto, Y.; Matsuki, T.; Kagawa, H.; Oshitani, Y.; Tsujino, K.; Yoshimura, K.; Miki, M.; et al. Long-term treatment outcome of progressive mycobacterium avium complex pulmonary disease. J. Clin. Med. 2020, 9, 1315. [Google Scholar] [CrossRef] [PubMed]
- Syal, K.; Maiti, K.; Naresh, K.; Avaji, P.G.; Chatterji, D.; Jayaraman, N. Synthetic arabinomannan glycolipids impede mycobacterial growth, sliding motility and biofilm structure. Glycoconj. J. 2016, 33, 763–777. [Google Scholar] [CrossRef]
- Dulberger, C.L.; Rubin, E.J.; Boutte, C.C. The mycobacterial cell envelope – A moving target. Nat. Rev. Microbiol. 2020, 18, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Van de Weerd, R.; Boot, M.; Maaskant, J.; Sparrius, M.; Verboom, T.; van Leeuwen, L.M.; Burggraaf, M.J.; Paauw, N.J.; Dainese, E.; Manganelli, R.; et al. Inorganic Phosphate Limitation Modulates Capsular Polysaccharide Composition in Mycobacteria. J. Biol. Chem. 2016, 291, 11787–11799. [Google Scholar] [CrossRef]
- Dufrisne, B.M.; Jorge, C.D.; Timoteo, C.G.; Petrou, V.I.; Ashraf, K.U.; Banerjee, S.; Clarke, O.B.; Santos, H.; Mancia, F. Structural and Functional Characterization of Phosphatidylinositol-Phosphate Biosynthesis in Mycobacteria. J. Mol. Biol. 2020. [Google Scholar]
- Tokunaga, T.; Yamamoto, T.; Yamamoto, S. How bcg led to the discovery of immunostimulatory DNA. Jpn. J. Infect. Dis. 1999, 52, 1–11. [Google Scholar] [PubMed]
- Fitzgerald, K.A.; Kagan, J.C. Toll-like Receptors and the Control of Immunity. Cell. 2020, 180, 1044–1066. [Google Scholar] [CrossRef] [PubMed]
- Uematsu, S.; Akira, S. Toll-like receptors and Type I interferons. J. Biol. Chem. 2007, 282, 15319–15323. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.J.; Im, E.; Pothoulakis, C.; Rhee, S.H. TRIF modulates TLR5-dependent responses by inducing proteolytic degradation of TLR5. J. Biol. Chem. 2010, 285, 21382–21390. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, O.; Akira, S. Pattern recognition receptors and inflammation. Cell 2010, 140, 805–820. [Google Scholar] [CrossRef]
- Vu, A.; Calzadilla, A.; Gidfar, S.; Calderon-Candelario, R.; Mirsaeidi, M. Toll-like receptors in mycobacterial infection. Eur. J. Pharmacol. 2017, 808, 1–7. [Google Scholar] [CrossRef]
- Mortaz, E.; Adcock, I.M.; Tabarsi, P.; Masjedi, M.R.; Mansouri, D.; Velayati, A.A.; Casanova, J.L.; Barnes, P.J. Interaction of Pattern Recognition Receptors with Mycobacterium Tuberculosis. J. Clin. Immunol. 2015, 35, 1–10. [Google Scholar] [CrossRef]
- Kurt-Jones, E.A.; Popova, L.; Kwinn, L.; Haynes, L.M.; Jones, L.P.; Tripp, R.A.; Walsh, E.E.; Freeman, M.W.; Golenbock, D.T.; Anderson, L.J.; et al. Pattern recognition receptors tlr4 and cd14 mediate response to respiratory syncytial virus. Nat. Immunol. 2000, 1, 398–401. [Google Scholar] [CrossRef]
- Saraav, I.; Singh, S.; Sharma, S. Outcome of Mycobacterium tuberculosis and Toll-like receptor interaction: Immune response or immune evasion? Immunol. Cell Biol. 2014, 92, 741–746. [Google Scholar] [CrossRef]
- Shukla, S.; Richardson, E.T.; Drage, M.G.; Boom, W.H.; Harding, C.V. Mycobacterium tuberculosis Lipoprotein and Lipoglycan Binding to Toll-Like Receptor 2 Correlates with Agonist Activity and Functional Outcomes. Infect. Immun. 2018, 86, e00450-18. [Google Scholar] [CrossRef]
- Lien, E.; Sellati, T.J.; Yoshimura, A.; Flo, T.H.; Rawadi, G.; Finberg, R.W.; Carroll, J.D.; Espevik, T.; Ingalls, R.R.; Radolf, J.D.; et al. Toll-like receptor 2 functions as a pattern recognition receptor for diverse bacterial products. J. Biol. Chem. 1999, 274, 33419–33425. [Google Scholar] [CrossRef]
- Sweet, L.; Schorey, J.S. Glycopeptidolipids from mycobacterium avium promote macrophage activation in a tlr2- and myd88-dependent manner. J. Leukoc. Biol. 2006, 80, 415–423. [Google Scholar] [CrossRef]
- Feng, C.G.; Scanga, C.A.; Collazo-Custodio, C.M.; Cheever, A.W.; Hieny, S.; Caspar, P.; Sher, A. Mice lacking myeloid differentiation factor 88 display profound defects in host resistance and immune responses to mycobacterium avium infection not exhibited by toll-like receptor 2 (tlr2)- and tlr4-deficient animals. J. Immunol. 2003, 171, 4758–4764. [Google Scholar] [CrossRef] [PubMed]
- Marinho, F.A.; de Paula, R.R.; Mendes, A.C.; de Almeida, L.A.; Gomes, M.T.; Carvalho, N.B.; Oliveira, F.S.; Caliari, M.V.; Oliveira, S.C. Toll-like receptor 6 senses mycobacterium avium and is required for efficient control of mycobacterial infection. Eur. J. Immunol. 2013, 43, 2373–2385. [Google Scholar] [CrossRef]
- Carvalho, N.B.; Oliveira, F.S.; Duraes, F.V.; de Almeida, L.A.; Florido, M.; Prata, L.O.; Caliari, M.V.; Appelberg, R.; Oliveira, S.C. Toll-like receptor 9 is required for full host resistance to mycobacterium avium infection but plays no role in induction of th1 responses. Infect. Immun. 2011, 79, 1638–1646. [Google Scholar] [CrossRef]
- Ryu, Y.J.; Kim, E.J.; Lee, S.H.; Kim, S.Y.; Suh, G.Y.; Chung, M.P.; Kim, H.; Kwon, O.J.; Koh, W.J. Impaired expression of toll-like receptor 2 in nontuberculous mycobacterial lung disease. Eur. Respir. J. 2007, 30, 736–742. [Google Scholar] [CrossRef] [PubMed]
- Meylan, E.; Tschopp, J.; Karin, M. Intracellular pattern recognition receptors in the host response. Nature 2006, 442, 39–44. [Google Scholar] [CrossRef] [PubMed]
- van de Veerdonk, F.L.; Netea, M.G.; Dinarello, C.A.; Joosten, L.A. Inflammasome activation and IL-1beta and IL-18 processing during infection. Trends Immunol. 2011, 32, 110–116. [Google Scholar] [CrossRef]
- Wu, M.F.; Shu, C.C.; Wang, J.Y.; Yan, B.S.; Lai, H.C.; Chiang, B.L.; Wu, L.S.; Yu, C.J. NLRP3 inflammasome is attenuated in patients with Mycobacterium avium complex lung disease and correlated with decreased interleukin-1beta response and host susceptibility. Sci. Rep. 2019, 9, 12534. [Google Scholar] [CrossRef]
- Lee, H.M.; Yuk, J.M.; Kim, K.H.; Jang, J.; Kang, G.; Park, J.B.; Son, J.W.; Jo, E.K. Mycobacterium abscessus activates the NLRP3 inflammasome via Dectin-1-Syk and p62/SQSTM1. Immunol. Cell Biol. 2012, 90, 601–610. [Google Scholar] [CrossRef]
- Miyake, Y.; Toyonaga, K.; Mori, D.; Kakuta, S.; Hoshino, Y.; Oyamada, A.; Yamada, H.; Ono, K.; Suyama, M.; Iwakura, Y.; et al. C-type lectin mcl is an fcrgamma-coupled receptor that mediates the adjuvanticity of mycobacterial cord factor. Immunity 2013, 38, 1050–1062. [Google Scholar] [CrossRef]
- Toyonaga, K.; Torigoe, S.; Motomura, Y.; Kamichi, T.; Hayashi, J.M.; Morita, Y.S.; Noguchi, N.; Chuma, Y.; Kiyohara, H.; Matsuo, K.; et al. C-type lectin receptor dcar recognizes mycobacterial phosphatidyl-inositol mannosides to promote a th1 response during infection. Immunity 2016, 45, 1245–1257. [Google Scholar] [CrossRef] [PubMed]
- Yonekawa, A.; Saijo, S.; Hoshino, Y.; Miyake, Y.; Ishikawa, E.; Suzukawa, M.; Inoue, H.; Tanaka, M.; Yoneyama, M.; Oh-Hora, M.; et al. Dectin-2 is a direct receptor for mannose-capped lipoarabinomannan of mycobacteria. Immunity 2014, 41, 402–413. [Google Scholar] [CrossRef]
- Kang, P.B.; Azad, A.K.; Torrelles, J.B.; Kaufman, T.M.; Beharka, A.; Tibesar, E.; DesJardin, L.E.; Schlesinger, L.S. The human macrophage mannose receptor directs mycobacterium tuberculosis lipoarabinomannan-mediated phagosome biogenesis. J. Exp. Med. 2005, 202, 987–999. [Google Scholar] [CrossRef] [PubMed]
- Geijtenbeek, T.B.; Van Vliet, S.J.; Koppel, E.A.; Sanchez-Hernandez, M.; Vandenbroucke-Grauls, C.M.; Appelmelk, B.; Van Kooyk, Y. Mycobacteria target dc-sign to suppress dendritic cell function. J. Exp. Med. 2003, 197, 7–17. [Google Scholar] [CrossRef]
- Akaki, T.; Sato, K.; Shimizu, T.; Sano, C.; Kajitani, H.; Dekio, S.; Tomioka, H. Effector molecules in expression of the antimicrobial activity of macrophages against mycobacterium avium complex: Roles of reactive nitrogen intermediates, reactive oxygen intermediates, and free fatty acids. J. Leukoc. Biol. 1997, 62, 795–804. [Google Scholar] [CrossRef] [PubMed]
- Gomes, M.S.; Florido, M.; Pais, T.F.; Appelberg, R. Improved clearance of mycobacterium avium upon disruption of the inducible nitric oxide synthase gene. J. Immunol. 1999, 162, 6734–6739. [Google Scholar]
- Gutierrez, M.G.; Mishra, B.B.; Jordao, L.; Elliott, E.; Anes, E.; Griffiths, G. Nf-kappa b activation controls phagolysosome fusion-mediated killing of mycobacteria by macrophages. J. Immunol. 2008, 181, 2651–2663. [Google Scholar] [CrossRef]
- Sexton, P.; Harrison, A.C. Susceptibility to nontuberculous mycobacterial lung disease. Eur. Respir. J. 2008, 31, 1322–1333. [Google Scholar] [CrossRef]
- Doffinger, R.; Smahi, A.; Bessia, C.; Geissmann, F.; Feinberg, J.; Durandy, A.; Bodemer, C.; Kenwrick, S.; Dupuis-Girod, S.; Blanche, S.; et al. X-linked anhidrotic ectodermal dysplasia with immunodeficiency is caused by impaired nf-kappab signaling. Nat. Genet. 2001, 27, 277–285. [Google Scholar] [CrossRef]
- Ogus, A.C.; Yoldas, B.; Ozdemir, T.; Uguz, A.; Olcen, S.; Keser, I.; Coskun, M.; Cilli, A.; Yegin, O. The arg753gln polymorphism of the human toll-like receptor 2 gene in tuberculosis disease. Eur. Respir. J. 2004, 23, 219–223. [Google Scholar] [CrossRef]
- Lindestam Arlehamn, C.S.; Lewinsohn, D.; Sette, A.; Lewinsohn, D. Antigens for CD4 and CD8 T cells in tuberculosis. Cold Spring Harb. Perspect. Med. 2014, 4, a018465. [Google Scholar] [CrossRef] [PubMed]
- Ramirez-Alejo, N.; Santos-Argumedo, L. Innate defects of the il-12/ifn-gamma axis in susceptibility to infections by mycobacteria and salmonella. J. Interferon Cytokine Res. 2014, 34, 307–317. [Google Scholar] [CrossRef] [PubMed]
- Kaufmann, S.H. Protection against tuberculosis: Cytokines, T cells, and macrophages. Ann. Rheum. Dis. 2002, 61. [Google Scholar] [CrossRef] [PubMed]
- Serbina, N.V.; Flynn, J.L. CD8(+) T cells participate in the memory immune response to Mycobacterium tuberculosis. Infect. Immun. 2001, 69, 4320–4328. [Google Scholar] [CrossRef]
- Matsuyama, M.; Ishii, Y.; Yageta, Y.; Ohtsuka, S.; Ano, S.; Matsuno, Y.; Morishima, Y.; Yoh, K.; Takahashi, S.; Ogawa, K.; et al. Role of Th1/Th17 balance regulated by T-bet in a mouse model of Mycobacterium avium complex disease. J. Immunol. 2014, 192, 1707–1717. [Google Scholar] [CrossRef]
- Al-Herz, W.; Bousfiha, A.; Casanova, J.L.; Chatila, T.; Conley, M.E.; Cunningham-Rundles, C.; Etzioni, A.; Franco, J.L.; Gaspar, H.B.; Holland, S.M.; et al. Primary immunodeficiency diseases: An update on the classification from the international union of immunological societies expert committee for primary immunodeficiency. Front. Immunol. 2014, 5, 162. [Google Scholar]
- Otome, O.; O’Reilly, M.; Lim, L. Disseminated mycobacterium haemophilum skeletal disease in a patient with interferon-gamma deficiency. Intern. Med. J. 2015, 45, 1073–1076. [Google Scholar] [CrossRef]
- Valour, F.; Perpoint, T.; Senechal, A.; Kong, X.F.; Bustamante, J.; Ferry, T.; Chidiac, C.; Ader, F.; Lyon, T. Interferon-gamma Autoantibodies as Predisposing Factor for Nontuberculous Mycobacterial Infection. Emerg. Infect. Dis. 2016, 22, 1124–1126. [Google Scholar] [CrossRef]
- Marciano, B.E.; Huang, C.Y.; Joshi, G.; Rezaei, N.; Carvalho, B.C.; Allwood, Z.; Ikinciogullari, A.; Reda, S.M.; Gennery, A.; Thon, V.; et al. Bcg vaccination in patients with severe combined immunodeficiency: Complications, risks, and vaccination policies. J. Allergy Clin. Immunol. 2014, 133, 1134–1141. [Google Scholar] [CrossRef]
- Yarmohammadi, H.; Cunningham-Rundles, C. Idiopathic CD4 lymphocytopenia: Pathogenesis, etiologies, clinical presentations and treatment strategies. Ann. Allergy Asthma Immunol. 2017, 119, 374–378. [Google Scholar] [CrossRef]
- Hariadi, N.I.; Blackwood, R.A. Disseminated Mycobacterium Avium Complex in an Adolescent with Perinatally-Acquired HIV Infection. Infect. Dis. Rep. 2017, 9, 6884. [Google Scholar] [CrossRef] [PubMed]
- Parodi, M.; Favoreel, H.; Candiano, G.; Gaggero, S.; Sivori, S.; Mingari, M.C.; Moretta, L.; Vitale, M.; Cantoni, C. NKp44-NKp44 Ligand Interactions in the Regulation of Natural Killer Cells and Other Innate Lymphoid Cells in Humans. Front. Immunol. 2019, 10, 719. [Google Scholar] [CrossRef] [PubMed]
- Rocco, J.M.; Irani, V.R. Mycobacterium avium and modulation of the host macrophage immune mechanisms. Int. J. Tuberc. Lung Dis. 2011, 15, 447–452. [Google Scholar] [CrossRef] [PubMed]
- James, C.A.; Seshadri, C. T Cell Responses to Mycobacterial Glycolipids: On the Spectrum of "Innateness". Front. Immunol. 2020, 11, 170. [Google Scholar] [CrossRef] [PubMed]
- Layre, E.; Collmann, A.; Bastian, M.; Mariotti, S.; Czaplicki, J.; Prandi, J.; Mori, L.; Stenger, S.; De Libero, G.; Puzo, G.; et al. Mycolic acids constitute a scaffold for mycobacterial lipid antigens stimulating cd1-restricted t cells. Chem. Biol. 2009, 16, 82–92. [Google Scholar] [CrossRef] [PubMed]
- Kinjo, Y.; Takatsuka, S.; Kitano, N.; Kawakubo, S.; Abe, M.; Ueno, K.; Miyazaki, Y. Functions of CD1d-Restricted Invariant Natural Killer T Cells in Antimicrobial Immunity and Potential Applications for Infection Control. Front. Immunol. 2018, 9, 1266. [Google Scholar] [CrossRef]
- Kwon, B.E.; Ahn, J.H.; Park, E.K.; Jeong, H.; Lee, H.J.; Jung, Y.J.; Shin, S.J.; Jeong, H.S.; Yoo, J.S.; Shin, E.; et al. B Cell-Based Vaccine Transduced With ESAT6-Expressing Vaccinia Virus and Presenting alpha-Galactosylceramide Is a Novel Vaccine Candidate Against ESAT6-Expressing Mycobacterial Diseases. Front. Immunol. 2019, 10, 2542. [Google Scholar] [CrossRef]
- Hagiya, H.; Koyama, T.; Zamami, Y.; Minato, Y.; Tatebe, Y.; Mikami, N.; Teratani, Y.; Ohshima, A.; Shinomiya, K.; Kitamura, Y.; et al. Trends in incidence and mortality of tuberculosis in japan: A population-based study, 1997–2016. Epidemiol. Infect. 2018, 147, e38. [Google Scholar] [CrossRef]
- Daley, C.L. Nontuberculous mycobacterial disease in transplant recipients: Early diagnosis and treatment. Curr. Opin. Organ. Transplant. 2009, 14, 619–624. [Google Scholar] [CrossRef]
- Longworth, S.A.; Daly, J.S. Management of infections due to nontuberculous mycobacteria in solid organ transplant recipients-guidelines from the American society of transplantation infectious diseases community of practice. Clin. Transplant. 2019, 33, e13588. [Google Scholar] [CrossRef]
- Malouf, M.A.; Glanville, A.R. The spectrum of mycobacterial infection after lung transplantation. Am. J. Respir. Crit. Care Med. 1999, 160, 1611–1616. [Google Scholar] [CrossRef] [PubMed]
- Ose, N.; Minami, M.; Funaki, S.; Kanou, T.; Kanzaki, R.; Shintani, Y. Nontuberculous mycobacterial infection after lung transplantation: A report of four cases. Surg. Case Rep. 2019, 5, 11. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.K.; McAnally, K.J.; Seoane, L.; Lombard, G.A.; LaPlace, S.G.; Lick, S.; Dhillon, G.S.; Valentine, V.G. Analysis of pulmonary non-tuberculous mycobacterial infections after lung transplantation. Transpl. Infect. Dis. 2016, 18, 585–591. [Google Scholar] [CrossRef] [PubMed]
- Yamamura, Y.; Maeda, H.; Ogawa, Y.; Hashimoto, T. Experimental pulmonary cavity formation by mycobacterial components and synthetic adjuvants. Microbiol. Immunol. 1986, 30, 1175–1187. [Google Scholar] [CrossRef]
- Tsai, M.C.; Chakravarty, S.; Zhu, G.; Xu, J.; Tanaka, K.; Koch, C.; Tufariello, J.; Flynn, J.; Chan, J. Characterization of the tuberculous granuloma in murine and human lungs: Cellular composition and relative tissue oxygen tension. Cell Microbiol. 2006, 8, 218–232. [Google Scholar] [CrossRef]
- Torrado, E.; Fountain, J.J.; Robinson, R.T.; Martino, C.A.; Pearl, J.E.; Rangel-Moreno, J.; Tighe, M.; Dunn, R.; Cooper, A.M. Differential and site specific impact of b cells in the protective immune response to mycobacterium tuberculosis in the mouse. PLoS ONE 2013, 8, e61681. [Google Scholar] [CrossRef]
- Esmail, H.; Lai, R.P.; Lesosky, M.; Wilkinson, K.A.; Graham, C.M.; Horswell, S.; Coussens, A.K.; Barry, C.E., 3rd; O’Garra, A.; Wilkinson, R.J. Complement pathway gene activation and rising circulating immune complexes characterize early disease in hiv-associated tuberculosis. Proc. Natl. Acad. Sci. USA 2018, 115, E964–E973. [Google Scholar] [CrossRef]
- Achkar, J.M.; Casadevall, A. Antibody-mediated immunity against tuberculosis: Implications for vaccine development. Cell Host Microbe 2013, 13, 250–262. [Google Scholar] [CrossRef]
- Maekura, R.; Kitada, S.; Osada-Oka, M.; Tateishi, Y.; Ozeki, Y.; Fujicawa, T.; Miki, M.; Jyunnko, O.; Mori, M.; Matsumoto, S. Serum antibody profiles in individuals with latent mycobacterium tuberculosis infection. Microbiol. Immunol. 2019, 63, 130–138. [Google Scholar] [CrossRef]
- Hamasur, B.; Haile, M.; Pawlowski, A.; Schroder, U.; Kallenius, G.; Svenson, S.B. A mycobacterial lipoarabinomannan specific monoclonal antibody and its f(ab’) fragment prolong survival of mice infected with mycobacterium tuberculosis. Clin. Exp. Immunol. 2004, 138, 30–38. [Google Scholar] [CrossRef]
- Balu, S.; Reljic, R.; Lewis, M.J.; Pleass, R.J.; McIntosh, R.; van Kooten, C.; van Egmond, M.; Challacombe, S.; Woof, J.M.; Ivanyi, J. A novel human iga monoclonal antibody protects against tuberculosis. J. Immunol. 2011, 186, 3113–3119. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, N.; Thormann, V.; Hu, B.; Kohler, A.B.; Imai-Matsushima, A.; Locht, C.; Arnett, E.; Schlesinger, L.S.; Zoller, T.; Schurmann, M.; et al. Human isotype-dependent inhibitory antibody responses against mycobacterium tuberculosis. EMBO Mol. Med. 2016, 8, 1325–1339. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.L.; Chung, A.W.; Rosebrock, T.R.; Ghebremichael, M.; Yu, W.H.; Grace, P.S.; Schoen, M.K.; Tafesse, F.; Martin, C.; Leung, V.; et al. A functional role for antibodies in tuberculosis. Cell 2016, 167, 433–443. [Google Scholar] [CrossRef] [PubMed]
- Jain, D.; Ghosh, S.; Teixeira, L.; Mukhopadhyay, S. Pathology of pulmonary tuberculosis and non-tuberculous mycobacterial lung disease: Facts, misconceptions, and practical tips for pathologists. Semin. Diagn. Pathol. 2017, 34, 518–529. [Google Scholar] [CrossRef] [PubMed]
- Fujita, J.; Ohtsuki, Y.; Suemitsu, I.; Shigeto, E.; Yamadori, I.; Obayashi, Y.; Miyawaki, H.; Dobashi, N.; Matsushima, T.; Takahara, J. Pathological and radiological changes in resected lung specimens in Mycobacterium avium intracellulare complex disease. Eur. Respir. J. 1999, 13, 535–540. [Google Scholar] [CrossRef]
- Kunnath-Velayudhan, S.; Gennaro, M.L. Immunodiagnosis of tuberculosis: A dynamic view of biomarker discovery. Clin. Microbiol. Rev. 2011, 24, 792–805. [Google Scholar] [CrossRef]
- Schorey, J.S.; Sweet, L. The mycobacterial glycopeptidolipids: Structure, function, and their role in pathogenesis. Glycobiology 2008, 18, 832–841. [Google Scholar] [CrossRef]
- Kitada, S.; Kobayashi, K.; Ichiyama, S.; Takakura, S.; Sakatani, M.; Suzuki, K.; Takashima, T.; Nagai, T.; Sakurabayashi, I.; Ito, M.; et al. Serodiagnosis of mycobacterium avium-complex pulmonary disease using an enzyme immunoassay kit. Am. J. Respir. Crit. Care Med. 2008, 177, 793–797. [Google Scholar] [CrossRef]
- Shibata, Y.; Horita, N.; Yamamoto, M.; Tsukahara, T.; Nagakura, H.; Tashiro, K.; Watanabe, H.; Nagai, K.; Nakashima, K.; Ushio, R.; et al. Diagnostic test accuracy of anti-glycopeptidolipid-core iga antibodies for mycobacterium avium complex pulmonary disease: Systematic review and meta-analysis. Sci. Rep. 2016, 6, 29325. [Google Scholar] [CrossRef]
- Kitada, S.; Maekura, R.; Yoshimura, K.; Miki, K.; Miki, M.; Oshitani, Y.; Nishida, K.; Sawa, N.; Mori, M.; Kobayashi, K. Levels of antibody against glycopeptidolipid core as a marker for monitoring treatment response in mycobacterium avium complex pulmonary disease: A prospective cohort study. J. Clin. Microbiol. 2017, 55, 884–892. [Google Scholar] [CrossRef][Green Version]
- Yamada, K.; Sugiyama, T.; Yasuda, A.; Seki, Y.; Hasegawa, M.; Hayashi, Y.; Tarumi, O.; Nakagawa, T.; Yamada, N.; Ogawa, K. A study of relapse/recurrence cases after surgical treatment for patients with pulmonary nontuberculous mycobacteriosis. Kekkaku 2013, 88, 469–475. [Google Scholar] [PubMed]
- Swenson, C.; Zerbe, C.S.; Fennelly, K. Host variability in ntm disease: Implications for research needs. Front Microbiol. 2018, 9, 2901. [Google Scholar] [CrossRef]
- Maekura, R.; Okuda, Y.; Hirotani, A.; Kitada, S.; Hiraga, T.; Yoshimura, K.; Yano, I.; Kobayashi, K.; Ito, M. Clinical and prognostic importance of serotyping mycobacterium avium-mycobacterium intracellulare complex isolates in human immunodeficiency virus-negative patients. J. Clin. Microbiol. 2005, 43, 3150–3158. [Google Scholar] [CrossRef] [PubMed]
- Neonakis, I.K.; Gitti, Z.; Krambovitis, E.; Spandidos, D.A. Molecular diagnostic tools in mycobacteriology. J. Microbiol. Methods 2008, 75, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Maiden, M.C.; Bygraves, J.A.; Feil, E.; Morelli, G.; Russell, J.E.; Urwin, R.; Zhang, Q.; Zhou, J.; Zurth, K.; Caugant, D.A.; et al. Multilocus sequence typing: A portable approach to the identification of clones within populations of pathogenic microorganisms. Proc. Natl. Acad. Sci. USA 1998, 95, 3140–3145. [Google Scholar] [CrossRef] [PubMed]
- Brown-Elliott, B.A.; Fritsche, T.R.; Olson, B.J.; Vasireddy, S.; Vasireddy, R.; Iakhiaeva, E.; Alame, D.; Wallace, R.J.; Branda, J.A. Comparison of two commercial matrix-assisted laser desorption/ionization-time of flight mass spectrometry (maldi-tof ms) systems for identification of nontuberculous mycobacteria. Am. J. Clin. Pathol. 2019, 152, 527–536. [Google Scholar] [CrossRef]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef]
- Wood, D.E.; Salzberg, S.L. Kraken: Ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 2014, 15, R46. [Google Scholar] [CrossRef]
- Truong, D.T.; Tett, A.; Pasolli, E.; Huttenhower, C.; Segata, N. Microbial strain-level population structure and genetic diversity from metagenomes. Genome Res. 2017, 27, 626–638. [Google Scholar] [CrossRef]
- Chun, J.; Rainey, F.A. Integrating genomics into the taxonomy and systematics of the bacteria and archaea. Int. J. Syst. Evol. Microbiol. 2014, 64, 316–324. [Google Scholar] [CrossRef]
- Konstantinidis, K.T.; Tiedje, J.M. Genomic insights that advance the species definition for prokaryotes. Proc. Natl. Acad. Sci. USA 2005, 102, 2567–2572. [Google Scholar] [CrossRef] [PubMed]
- Parks, D.H.; Chuvochina, M.; Chaumeil, P.A.; Rinke, C.; Mussig, A.J.; Hugenholtz, P. A complete domain-to-species taxonomy for bacteria and archaea. Nat. Biotechnol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Auch, A.F.; von Jan, M.; Klenk, H.P.; Goker, M. Digital DNA-DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand. Genom. Sci. 2010, 2, 117–134. [Google Scholar] [CrossRef] [PubMed]
- Queiroz, K.D. The general lineage concept of species, species chteria, and the process of speciation and terminological recommendations. In Endless Forms Species Speciation; Oxford University Press: Oxford, UK, 1998; pp. 57–75. [Google Scholar]
- Jolley, K.A.; Maiden, M.C. Bigsdb: Scalable analysis of bacterial genome variation at the population level. BMC Bioinform. 2010, 11, 595. [Google Scholar] [CrossRef]
- Matsumoto, Y.; Kinjo, T.; Motooka, D.; Nabeya, D.; Jung, N.; Uechi, K.; Horii, T.; Iida, T.; Fujita, J.; Nakamura, S. Comprehensive subspecies identification of 175 nontuberculous mycobacteria species based on 7547 genomic profiles. Emerg. Microbes Infect. 2019, 8, 1043–1053. [Google Scholar] [CrossRef]
- Votintseva, A.A.; Bradley, P.; Pankhurst, L.; del Ojo Elias, C.; Loose, M.; Nilgiriwala, K.; Chatterjee, A.; Smith, E.G.; Sanderson, N.; Walker, T.M.; et al. Same-day diagnostic and surveillance data for tuberculosis via whole-genome sequencing of direct respiratory samples. J. Clin. Microbiol. 2017, 55, 1285–1298. [Google Scholar] [CrossRef]
- Ratnatunga, C.N.; Lutzky, V.P.; Kupz, A.; Doolan, D.L.; Reid, D.W.; Field, M.; Bell, S.C.; Thomson, R.M.; Miles, J.J. The Rise of Non-Tuberculosis Mycobacterial Lung Disease. Front. Immunol. 2020, 11, 303. [Google Scholar] [CrossRef]
- Seo, H.; Jeon, I.; Kim, B.S.; Park, M.; Bae, E.A.; Song, B.; Koh, C.H.; Shin, K.S.; Kim, I.K.; Choi, K.; et al. IL-21-mediated reversal of NK cell exhaustion facilitates anti-tumour immunity in MHC class I-deficient tumours. Nat. Commun. 2017, 8, 15776. [Google Scholar] [CrossRef]
| No. | Age | Sex | Primary Disease | Procedure of Transplant | Immuno-Suppressive Agents | Species | Radiologic Features | Site of Infection | Time to Isolate (months) | Treatment | Outcome |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 10 | M | PAH | DLT | TAC, MMF, PSL | M. gordonae | Non-cavitary NB | Transplanted | 76 | None | Alive |
| 2 | 35 | F | BE | DLT | CyA, MMF, PSL | M. abscessus | Non-cavitary NB | Transplanted | 82 | IMP, CAM, AMK | Dead |
| 3 | 41 | F | LAM | SLT | CyA, MMF, PSL | M. abscessus | Non-cavitary NB | Transplanted | 58 | IPM, AMK, AZM | Alive |
| 4 | 39 | M | IIP | SLT | CyA, MMF, PSL | M. intracellulare | Non-cavitary NB | Native | 12 | RFP, EB, CAM | Dead |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abe, Y.; Fukushima, K.; Hosono, Y.; Matsumoto, Y.; Motooka, D.; Ose, N.; Nakamura, S.; Kitada, S.; Kida, H.; Kumanogoh, A. Host Immune Response and Novel Diagnostic Approach to NTM Infections. Int. J. Mol. Sci. 2020, 21, 4351. https://doi.org/10.3390/ijms21124351
Abe Y, Fukushima K, Hosono Y, Matsumoto Y, Motooka D, Ose N, Nakamura S, Kitada S, Kida H, Kumanogoh A. Host Immune Response and Novel Diagnostic Approach to NTM Infections. International Journal of Molecular Sciences. 2020; 21(12):4351. https://doi.org/10.3390/ijms21124351
Chicago/Turabian StyleAbe, Yuko, Kiyoharu Fukushima, Yuki Hosono, Yuki Matsumoto, Daisuke Motooka, Naoko Ose, Shota Nakamura, Seigo Kitada, Hiroshi Kida, and Atsushi Kumanogoh. 2020. "Host Immune Response and Novel Diagnostic Approach to NTM Infections" International Journal of Molecular Sciences 21, no. 12: 4351. https://doi.org/10.3390/ijms21124351
APA StyleAbe, Y., Fukushima, K., Hosono, Y., Matsumoto, Y., Motooka, D., Ose, N., Nakamura, S., Kitada, S., Kida, H., & Kumanogoh, A. (2020). Host Immune Response and Novel Diagnostic Approach to NTM Infections. International Journal of Molecular Sciences, 21(12), 4351. https://doi.org/10.3390/ijms21124351




