Abstract
We present a case study report into nutritional competition between Trichoderma spp. isolated from wild raspberries and fungal phytopathogenic isolates (Colletotrichum sp., Botrytis sp., Verticillium sp. and Phytophthora sp.), which infect soft fruit ecological plantations. The competition was evaluated on the basis of nutritional potentiates. Namely, these were consumption and growth, calculated on the basis of substrate utilization located on Biolog® Filamentous Fungi (FF) plates. The niche size, total niche overlap and Trichoderma spp. competitiveness indices along with the occurrence of a stressful metabolic situation towards substrates highlighted the unfolding step-by-step approach. Therefore, the Trichoderma spp. and pathogen niche characteristics were provided. As a result, the substrates in the presence of which Trichoderma spp. nutritionally outcompete pathogens were denoted. These were adonitol, D-arabitol, i-erythritol, glycerol, D-mannitol and D-sorbitol. These substrates may serve as additives in biopreparations of Trichoderma spp. dedicated to plantations contaminated by phytopathogens of the genera Colletotrichum sp., Botrytis sp., Verticillium sp. and Phytophthora sp.
1. Introduction
Fungi are the dominant components of most terrestrial ecosystems [1]. However, there is one great concern that has been widely highlighted, which is the prevalence of plant fungal diseases. Fungi may attack both plants and fruit, thereby contributing to the unpredictable spoilage of agriproducts [2,3,4]. The crucial plant pathogens are those from the fungal genera Colletotrichum, Botrytis as well as Verticillium and fungal-like Phytophthora. These pathogens may wreak havoc, especially in organic production [5].
This is an important matter, since organic soft fruit production has been increasing constantly in recent years and has also enlarged its market share in worldwide food production [6]. Strawberry, raspberry and blueberry fruits are often included as crucial products of central Europe, with a seemingly endless increase in consumer demand to introduce organic methods of fruit cultivation. This constitutes a major reason to seek alternative ways to reduce the losses caused by pathogenic fungi [7].
In particular, Colletotrichum sp. and Botrytis sp., the causal actors of anthracnose and gray mould, respectively, require effective management strategies to combat them [8]. The Verticillium genus representatives, which cause wilt, are also critical from an economic point of view [9,10]. Phytophthora species are responsible for many losses in all of the production areas of the world [11].
Notably, the dynamic of ecological communities is not only rather menacingly shaped by pathogens but also by beneficial microorganisms [12]. Beneficial species share host plants with an array of pathogenic fungi that also inhabit the plant’s internal tissues. Therefore, interactions between these fungi are likely to occur on multiple temporal and spatial scales. Beneficial fungi occurrence in plants may alter disease symptoms to a significant extent [13,14].
In recent times, the process of the interaction between Trichoderma spp., pathogen and plant in securing pre-harvest organic soft fruit production was summarized and highlighted [15]. Trichoderma spp. fungi function as biocontrol agents and manifest a few common mechanisms. In fact, there are five main responses involved in attacking other fungi and promoting plant growth, these have been summarized recently [7]. They are, the production of inhibitory compounds, mycoparasitism, the inactivation of pathogen enzymes, induced resistance and finally, but not the least important mechanism, providing competition for nutrients and therefore for living space by forming mycelium biomass.
Nutritional competition is one of the most common biological control activities. What is more, this property is very useful for plant protection [16]. Fungi belonging to the Trichoderma genus are widely known for very rapid growth and are regarded as aggressive competitors. They quickly colonize substrates and exclude slower growing pathogens. Recent studies have also explored the meaning of the endophytic activities of Trichoderma spp. for the welfare of plants [17].
In terms of ecosystem function, the survival opportunity of fungal pathogens and beneficial strains in a given environment depends on their ability to tolerate nutritional conditions within a stress-like competition [1].
The niche overlapping concept that pertains to competition for substrates, posits that niches are defined by the requirements and impacts of the species that are occupying those niches. This, in turn, determines whether a given set of species can coexist in a given ecological community and therefore provides a prospective tool for biocontrol [18]. Researchers have only recently begun to investigate the relevance of phenotypic heterogeneity for the competitive success of microorganisms in different natural scenarios. [19]. This is in agreement with the principle of competitive exclusion (limiting similarity) [20].
The nutritional niche of fungi with the Phenotype Microarray (PM), among others that use Biolog® Filamentous Fungi (FF) plates, was examined previously [18]. An array of substrates on a microtiter plate was used to assess the exoenzymatic capacity of the tested fungi, since it is impossible to accurately mimic the in chemical environment of the plant inside the plant host. The method of PM that emerged has been proven to have a realistic potential of providing a high throughput of information about the phenotypes of microbial isolates [21,22,23,24].
Thus, the phenotypic consumption response is taken into consideration in the PM approach, namely how the cells respond colorimetrically to the nutritional conditions. This potentiates the results from respiration activity, which accompanies catabolic activity and is monitored at 490 nm [25,26]. On the other hand, the phenotypic turbidity response explaining the increase in microbial cell biomass formation as a reaction to nutritional conditions is also used. This potentiate is recorded as a change in the optical density at 750 nm [27,28,29]. The more intensive the colour formation in the PM method, the better the organism is able to nutritionally use (to consume or to grow on) the provided substrate.
Nevertheless, as it was recently proved that fungal biomass can, relatively speaking, be developed without consuming too much substrate [30]. This was then explored in, e.g., the evaluation of the phytochemicals of apple pomace as prospective bio-fungicide agents against mycotoxigenic fungi [31]. However, this aspect, to the best of our knowledge, has not been previously taken into consideration within competition evaluation. We regard this approach as a future tool that may provide a wider insight into the probability of biocontrol effectiveness upon niche overlapping phenomena.
Therefore, we hypothesized that substrates for nutritional growth as well as nutritional consumption potentiates obtained using the PM method, based on Biolog FF plates®, will be diversified between Trichoderma spp. and soft fruit pathogens: Colletotrichum sp., Botrytis sp., Verticillium sp. and Phytophthora sp. We aimed to characterize nutritional niches in order to assess the probability of niche colonization by these fungi. We intended to give the example of strawberry fruit cell walls and emphasize the substrates in the presence of which Trichoderma spp. is more nutritionally competitive.
2. Results
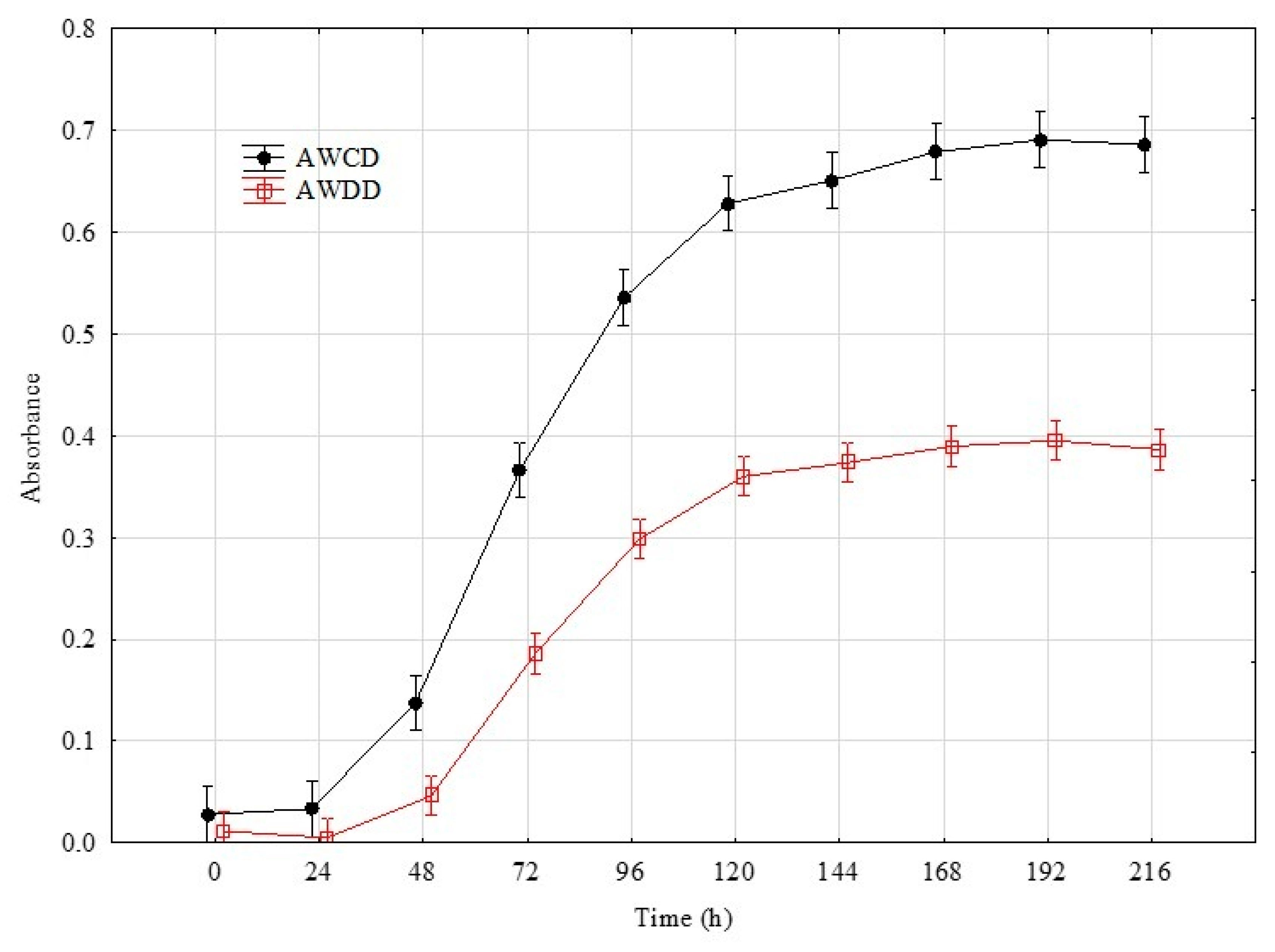
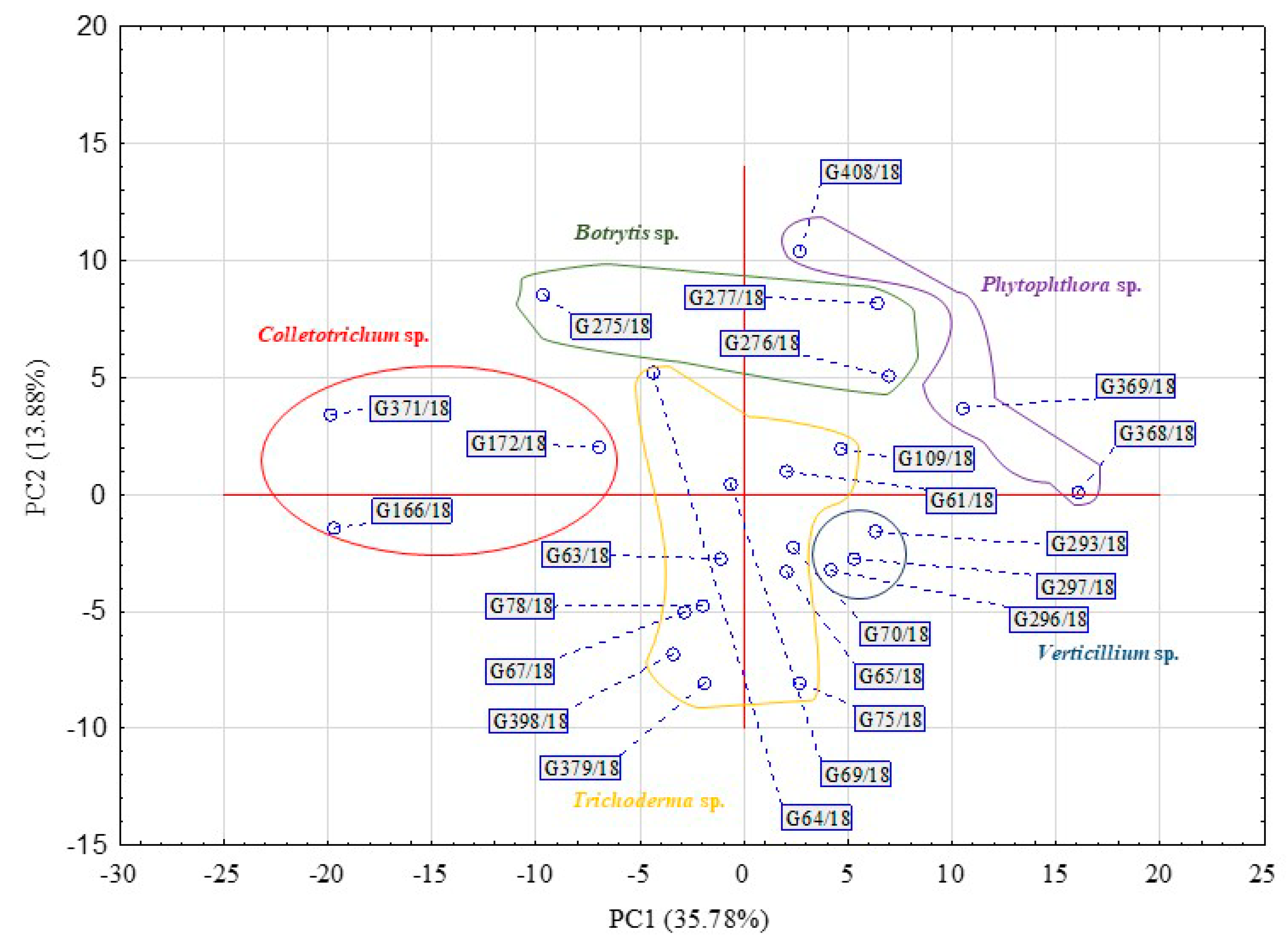
The rates in the total Average Well Colour Development (AWCD) and Average Well Density Development (AWDD) indices values were used to identify the time point that represents the greatest response of the set of tested fungal isolates. This approach was according to their consumption of different substrates and growth response, respectively (Figure 1). For further analyses, the time point of 192 h was taken into consideration with an average AWCD index value 0.7 and 0.4 for AWDD for the whole experiment data set. Principal component analysis (PCA) on the data of 192 h confirmed that the individual isolates belonging to the same genus clustered with respect to the isolate’s ability to consume and/or grow on particular sources (Figure 2).
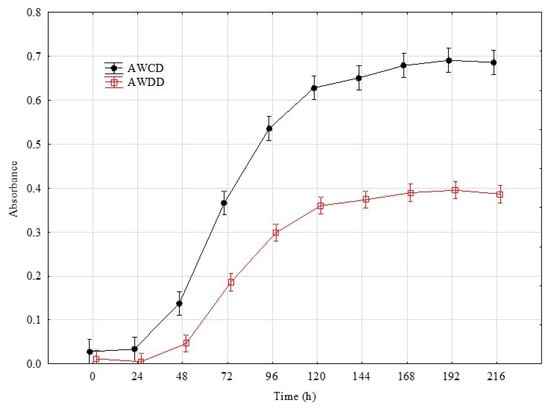
Figure 1.
Changes in total AWCD and AWDD values calculated on the basis of the pathogenic fungi and beneficial Trichoderma isolates’ response to Filamentous Fungi (FF) Biolog® substrates. Definitions: AWCD—Average Well Colour Development (A490nm), AWDD—Average Well Density Development (A750nm) (n = 3).
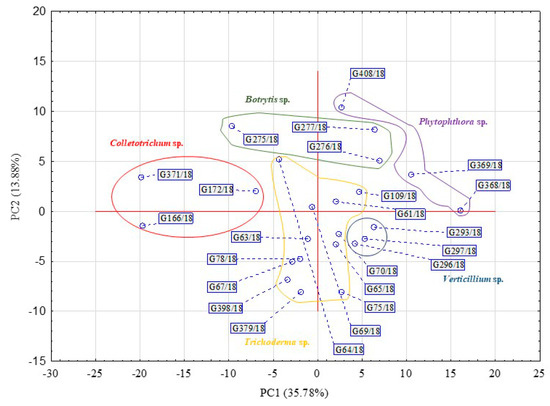
Figure 2.
Principal component analysis (PCA) calculated on the basis of the pathogenic fungi and beneficial Trichoderma isolates’ response to FF Biolog® substrates at 192 h (n = 3).
Table 1 presents the rotated factor loadings and highly influential principal components (PC1: 35.78% and PC2 13.88%). There were 38 substrates, from the 95 available on an FF plate, which strongly and positively influence PC1 or PC2 for a fungal dataset. For the most part, if a substrate influences principal components, this influence originated from both consumption and growth potentiates. Only a minor difference was noted in the factors value obtained between the consumption and growth potentials. For example, it was for 2-amino ethanol that these were 0.807 and 0.779, respectively, influencing PC1 or for adenosine 0.825 and 0.719, respectively, influencing PC2.

Table 1.
Rotated factor loadings with the principal components (PC) distinguished (PC1: 35.78% and PC2 13.88%), calculated on the basis of the pathogenic fungi and beneficial Trichoderma isolates response to FF Biolog® substrates, namely the consumption (490 nm) and growth (750 nm) potentiates (A > 0.2, n = 3, 192 h); “-“ means lack of particular substrate group influence on PC; bold numbers mean its significant influence on PC (PC ≥ 0.7).
For some substrates, there were higher factor values noted for those represented by growth potentiates (putrescine, L-alanine, L-asparagine, L-serine, L-threonine, γ-amino-butyric acid, lactulose, succinic acid mono-methyl ester, D-ribose, maltitol, D-glucuronic acid). Moreover, exclusively, the consumption potentiates of such substrates as D-trehalose from the glucosides group, L-phenylalanine from L-amino acids, D-sorbitol from polyols and the growth potentiates of proline from L-amino acids, p-hydroxyphenyl acetic acid from other groups, fumaric acid belonging to the Tricarboxylic Acid (TCA) cycle-intermediates, have influenced principal components with no influence noted coming opposite from the growth and consumption of these substrates, respectively. This clearly suggests that both consumption and growth potentiates matter to a significant extent when evaluating fungi for nutritional differences that eventually make up the competition features.
The niche size, based on the total number of substrates used for the consumption and/or growth of the fungi of interest, varied among Trichoderma spp. and the genera group of specific pathogens (Botrytis sp., Colletotrichum sp., Phytophthora sp., Verticillium sp.) (Table 2). Nevertheless, the substrates that are most ubiquitously used by all of the tested fungi were found to belong to the oligosaccharides and peptides groups, when niche size evaluating. The most comparable niche size for the Trichoderma spp. among all of the tested pathogenic fungi was revealed to be Colletotrichum sp. Phytophthora sp. was noted to have the lowest niche size, being able to consume and grow on a low number of substrates. Trichoderma spp. was found to be superior to all other pathogens consuming 100% and growing on 88% of available hexoses; consuming 100% and growing on 25% of available aliphatic organic acids; consuming 75% of hexosamines and growing on 80% of pentoses. Botrytis sp. was found to have the greatest niche size among all tested fungi, when it comes to consuming 100% polysaccharides. Colletotrichum sp. and Verticillium sp. were found to most easily consume and grow on biogenic and heterocyclic amines (75% and 50%, respectively).

Table 2.
The share of FF Biolog® substrate group positive response (%) of the pathogenic fungi and beneficial Trichoderma strains, calculated on the basis of consumption (Average Well Colour Development—AWCD, 490 nm) and growth (Average Well Density Development—AWDD) potentiates (A > 0.2, n = 3).
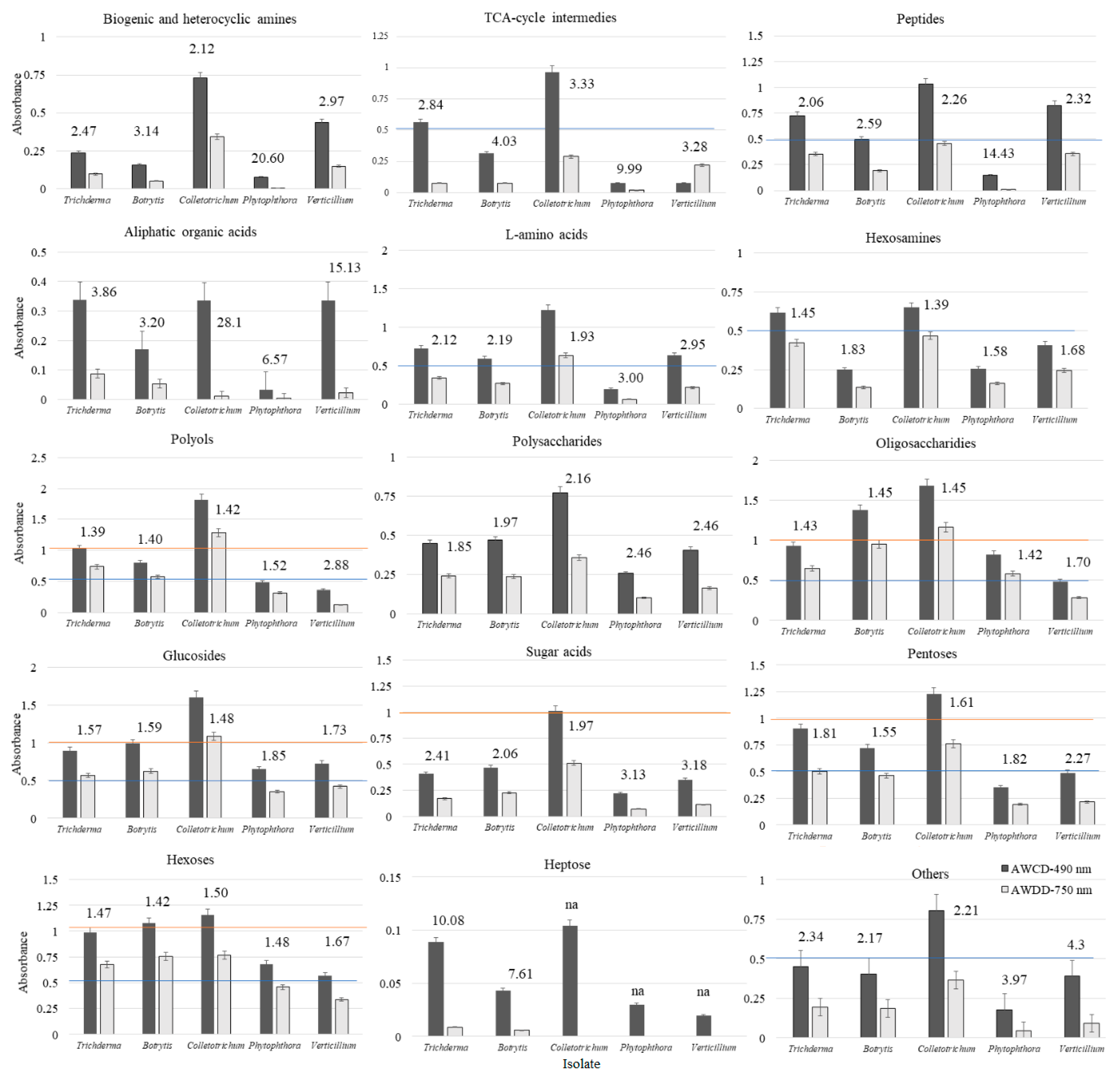
However, if the average nutritional response is analysed, namely the AWCD and AWDD values of the tested groups of substrates (Figure 3), it did not exactly match the findings noted for niche size. The greatest nutritional response (AWCD ≥1.0 and AWDD ≥0.5) among all of the tested fungi was met for such a group of substrates as polyols, oligosaccharides, glucosides, pentoses, hexoses. However, the lowest response (AWCD and AWDD ≤0.5) was for biogenic and heterocyclic amines, aliphatic and heterocyclic amines, polysaccharides, sugar acids, and heptose. The medium response (AWCD 0.5–1.0) was encountered on TCA-cycle intermediates, peptides, L-amino acids, hexosamines, and other groups. Nevertheless, for most substrate groups, at least one pathogen exceeded the response of Trichoderma spp. This was encountered for Colletotrichum sp. on polyols, glucosides and pentoses, and for Colletotrichum sp. and Botrytis sp. on oligosaccharides and hexoses. There was a trend met that Colletotrichum sp. dominates over Trichoderma spp. and other pathogens nutritionally, revealing greater catabolism and/or growth. Trichoderma spp. can match nutritionally Colletotrichum sp. on hexosamines intermediates. The average nutritional response was almost the same.
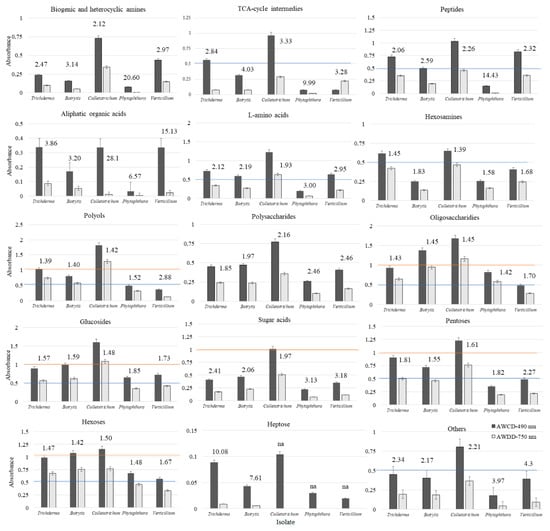
Figure 3.
AWCD and AWDD ratio values of substrate groups calculated on the basis of consumption (Average Well Colour Development—AWCD, 490 nm) and growth (Average Well Density Development—AWDD) potentiates (A > 0.2, n = 3); “na” – indicates not available.
What is more, a stressful metabolic situation, indicated by the ratio of both AWCD to AWDD (Figure 3) was met when using L-amino acids, sugar acids and others for Phytophthora sp. and Verticillium sp. Stressful situation was noted on TCA-cycle intermediates and biogenic and heterocyclic amines for all tested pathogens, but not Trichoderma spp. Peptides caused metabolic stress only for Phytophthora sp., heptose for Trichoderma spp. and Botrytis sp., whereas aliphatic organic acids caused metabolic stress for Trichoderma spp. and all pathogens.
The NOITOT index (Table 3) was found to reach 1 mainly for Colletotrichum sp. due to polyols, L-amino acids, TCA- cycle intermediates consumption as was the case for growth polysaccharides, biogenic and heterocyclic amine, glucosides and polyols. As for Botrytis sp. and Verticillium sp. and the growth for these genera, the greatest NOITOT for polysaccharides and biogenic and heterocyclic amines, respectively was revealed. It was also confirmed that the most versatile substrates for all tested fungi belonged to oligosaccharides and peptides.

Table 3.
Total Niche Overlap Index (NOITOT) and Trichoderma Competitiveness Index (COMTRICH). The presented indices were calculated on the basis of consumption (Average Well Colour Development—AWCD, 490 nm) and growth (Average Well Density Development—AWDD) potentiates (A > 0.2, n = 3).
The COMTRICH index (Table 3) reached >2.0, giving competitive consumption and growth dominance over Botrytis sp., on hexosamines. For Phytophthora sp., this phenomenon was observed on hexosamines and L-amino-acids. Consumption dominance over Botrytis sp. was met on aliphatic and organic acids. Growth dominance over Botrytis sp. was met on peptides and other groups, over Phytophthora sp. on pentoses and over Verticillium sp. on polysaccharides, glucosides, polyols and pentoses.
It should be mentioned that high COMTRICH index values were also noted, thereby explaining the growth dominance over Colletotrichum sp. and Verticillium sp. on biogenic and heterocyclic amines. However, simultaneously for this substrate group, a very low COMTRICH index value was encountered, indicating the consumption superiority of those two pathogens over Trichoderma spp.
Trichoderma spp. was able to produce a higher response than Phytophthora sp. on L-aminoamides, polysaccharides, sugar acids and others. Phytophthora sp. and Botrytis sp. reacted greatly on biogenic and heterocyclic amines and peptides. Phytophthora sp. and Verticillium sp. responded well on oligosaccharides, glucosides and hexoses, whereas Phytophthora sp., Verticillium sp. and Botrytis sp. on TCA-cycle intermediates, hexosamines, polyols and pentoses.
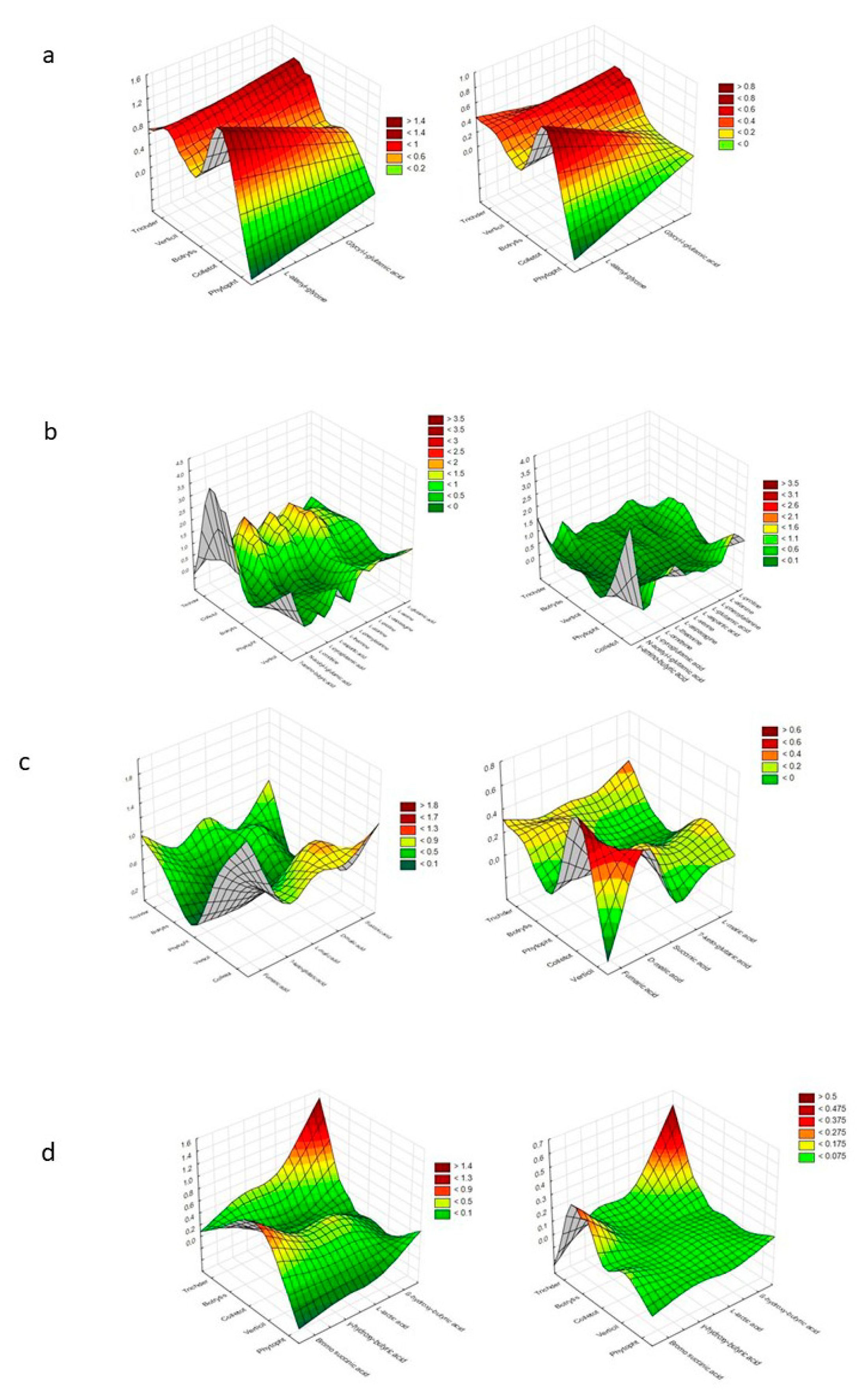
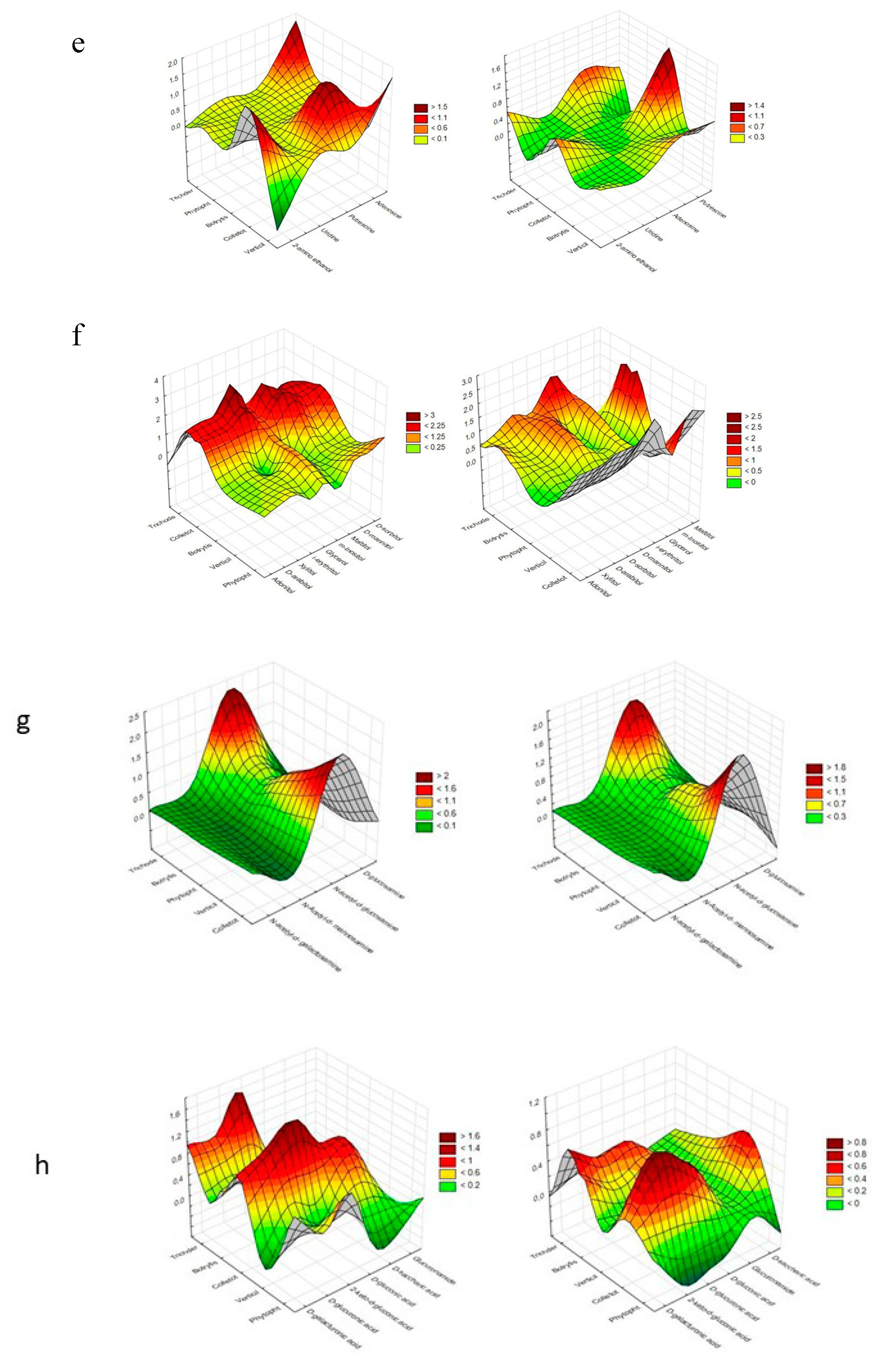
Figure 4 presents a denotation of preferred and non-preferred particular substrates among each group (those groups that provoke a stressful metabolic situation for Trichoderma spp. were excluded).
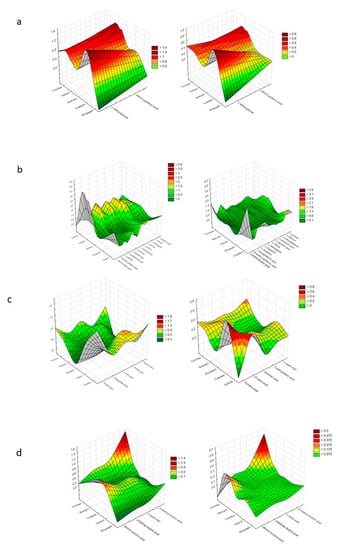
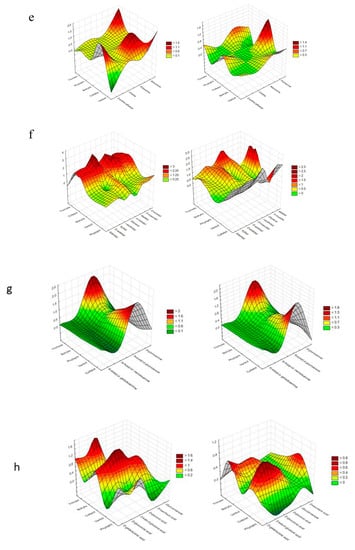
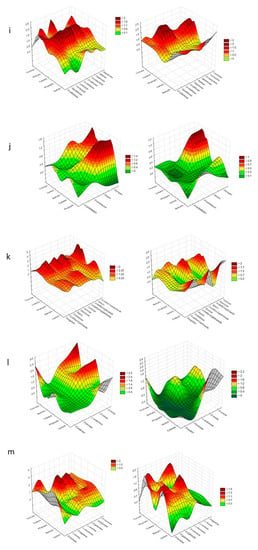
Figure 4.
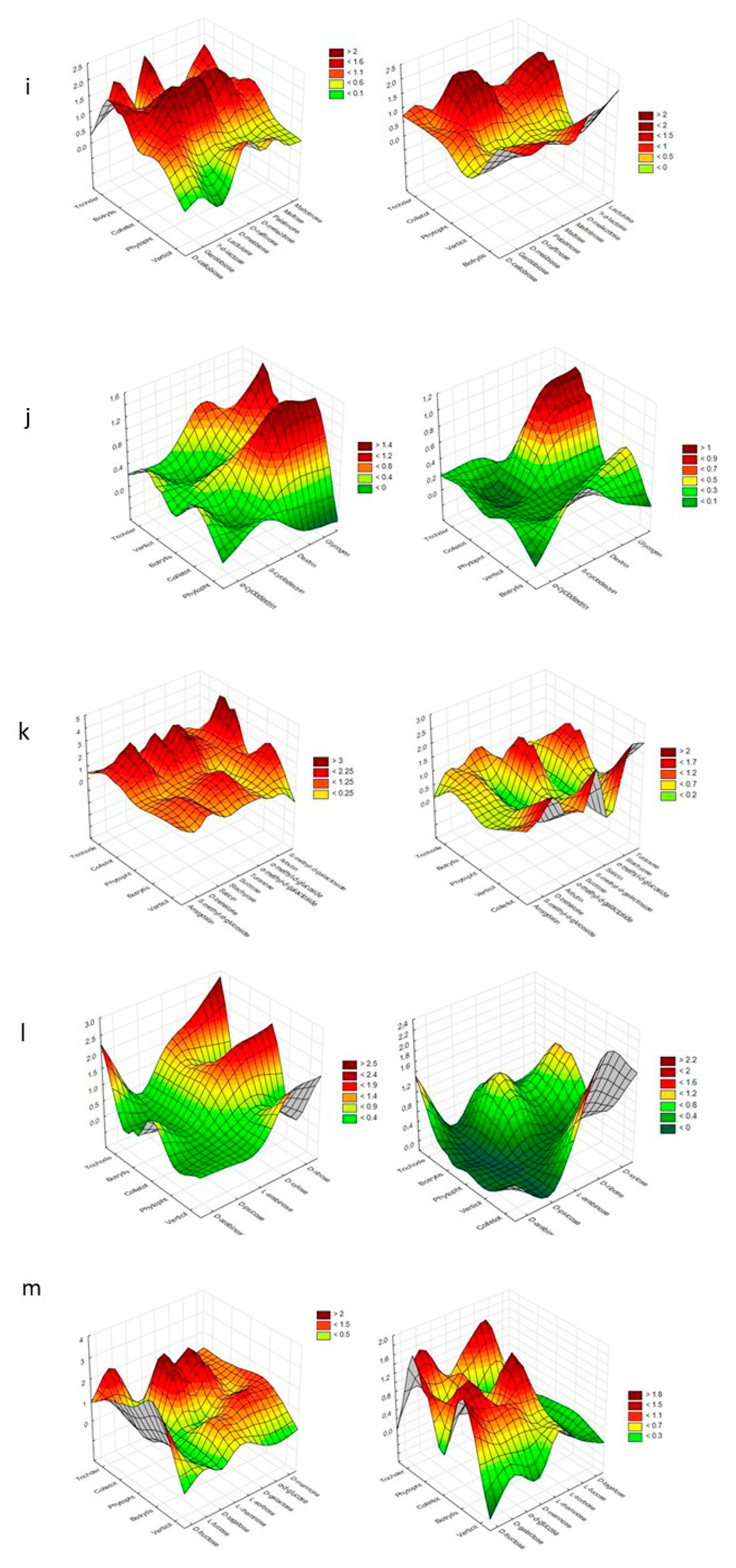
Cluster Analysis depicting the consumption response (A 490 nm, shown at left) and growth response (A 750 nm, shown at right) of microorganisms to substrates located on Biolog® FF plates, shown as the following substrate groups: (a) peptides, (b) L-amino acids, (c) Tricarboxylic Acid (TCA) cycle-intermediates, (d) aliphatic organic acids, (e) biogenic and heterocyclic amines, (f) polyols, (g) hexosamines, (h) sugar acids, (i) oligosaccharides, (j) polysaccharides, (k) glucosides, (l) pentoses, and (m) hexoses. The analysis was performed on the basis of consumption (490 nm) and growth potentiates (A > 0.2, n = 3).
It was assumed that mainly Trichoderma spp. and Colletotrichum sp. preferred the same particular substrates. These were as follows, from the peptides—L-alanyl-glycine and glycyl-l-glutamic acid; from L-amino acids—γ-amino-butyric acid; from TCA-cycle intermediates— fumaric acid; from hexosamines—N-acetyl-D-glucosamine.
Polysaccharides such as dextrin and glycogen, hexoses such as D-fructose, D-galactose, α-D-glucose, D-mannose, L-rhamnose, D-tagatose, and L-sorbose, sugar acids such as D-galacturonic acid, D-gluconic acid, D-glucuronic acid, 2-keto-D-gluconic acid, and D-saccharic acid, polyols such as adonitol, D-arabitol, i-erythritol, glycerol, D-mannitol, D-sorbitol, xylitol, inositol, and maltitol, and biogenic and heterocyclic amines such as 2-amino ethanol and putrescine were even more preferred by Colletotrichum sp. than Trichoderma spp.
Trichoderma spp. preferred the polyols adonitol, D-arabitol, i-erythritol, glycerol, D-mannitol, and D-sorbitol, and, from biogenic and heterocyclic amines, adenosine.
Pentoses such as D-arabinose, L-arabinose, D-ribose, and D-xylose were preferred to a high degree by Trichoderma spp., Colletotrichum sp. and Verticillium sp.
Glucosides—amygdalin, arbutin, α-methyl-D-galactoside, β-methyl-D-glucoside, salicin, stachyose, sucrose, D-trehalose, and turanose—were preferred to an equal extent by Trichoderma spp. and Botritis sp.
Table 4 presents the saccharide composition of cell wall material extracted from strawberries cv. Dipret from organic farming. Sugar acids (galacturonic acid), pentoses (arabinose, xylose) and hexoses (rhamnose, galactose, glucose, mannose) were measured. Galacturonic acid (47.9 mol%), glucose (24.2 mol%), arabinose (12.1 mol%), and galactose (9.1 mol%) were revealed to be the most abundant. A low content of xylose (1.8 mol%), rhamnose (3.0 mol%) and mannose (1.9 mol%) was determined.

Table 4.
Saccharide composition of strawberry fruit (mol%).
3. Discussion
Nutritional potentiates were previously reported to be useful in niche overlap evaluation following the phenotype Microarray approach and based on substrate consumption. This was applied to Dutch elm fungal endophytes and pathogens [18].
As for the microorganisms of interest in soft fruit plantation, global substrate assimilation within mycelial growth was previously assessed and described as the metabolic profiles characteristic of Trichoderma spp. strains [32], Phytophthora sp. and Botrytis sp. [23]. Moreover, to the best of our knowledge, Verticillium sp. and Colletotrichum sp. have not been tested in this way. An array of substrates on a microtiter plate was used to assess the exoenzymatic capacity of the tested fungi.
Following our hypothesis of nutritional growth as well as nutritional consumption, potentiates obtained using the PM method, based on Biolog FF plates®, were noted to be diversified between Trichoderma spp. and Colletotrichum sp., Botrytis sp., Verticillium sp. and Phytophthora sp., and thus it has an important meaning in niche evaluation, which included the competition for substrate groups (niche size, niche overlap index, competitiveness), the stressful metabolic situation, and substrates usage selectivity.
We regard this approach as a future tool for providing a wider insight into the probability of biocontrol effectiveness. Substrates, such as those preferred by Trichoderma spp., but not by pathogens, may be considered as additives to Trichoderma spp. biopreparations and are expected to increase their competitiveness in the destined microbial community, e.g., community of soil or plant tissue beset by pathogens.
It was revealed that Trichoderma spp. has the most similar niche to Colletotrichum sp. and follows the limiting similarity principle (competitive exclusion), these two species cannot occupy the same ecological niche [33]. Therefore, additives’ conception with adenosine, revealed in this study, may bring about a positive effect, especially against Colletotrichum sp.
Apart from adenosine, adonitol, D-arabitol, i-erythritol, glycerol, D-mannitol, and D-sorbitol can also be added to Trichoderma spp. biopreparations dedicated to plantations, where Colletotrichum sp., Botrytis sp., Verticillium sp. and Phytophthora sp. appear. Nevertheless, there are few substrates denoted that are preferred not only by Trichoderma spp. but also by particular pathogens, and, therefore, these may be considered as additives but, interestingly, it depends on which plates the fungal infection occurs. For example, L-alanyl-glycine and glycyl-l-glutamic acid, γ-amino-butyric acid, fumaric acid, N-acetyl-d-glucosamine could be less effective as additives applied, if Colletotrichum sp. occurred, but not for Verticillium sp., Phytophthora sp. and Botrytis sp.
Our studies showed that, in particular, hexosamines groups are expected to increase Trichoderma spp. competitiveness against Botrytis sp. and Phytophthora sp. Oligosaccharides and peptides groups were the most ubiquitously used by all of the fungi tested and therefore would probably not give Trichoderma spp. much predominance in the community.
The nutritional advantage of Trichoderma spp. also results from the fact that this group of fungi demonstrated a relatively low metabolic stress situation in the presence of only a few substrate groups compared to the pathogens. The above fact leads to the premise that Trichoderma spp. has a vast ability to develop in the environment. This beneficial aspect is the most desired activity in a fungal community and indicates the ability of Trichoderma spp. to more effectively colonize numerous and various niches [34]. Colonization is the very first step in the use of a wide array of other biological control mechanisms, such as antibiosis, antagonism, mycoparasitism, and the induction of plant defence responses [35].
Moreover, our findings related to strawberry fruit saccharides composition and the determination of preferred and non-preferred particular substrates among saccharides (glucose, mannose, rhamnose, galacturonic acid, arabinose) indicate that saccharide composition may be one of many conditions that results in Colletotrichum sp. and Verticillium sp. colonization and consequently in anthracnose and Verticillium wilt diseases development. It seems that in Phytophthora sp. and Botrytis sp. colonization, the other substrates play a crucial role. However, it is worth noting that, according to the results obtained, Botrytis sp. intensively utilizes galacturonic acid, which is one of the main components of strawberry, which may explain why these pathogens develop so easily on strawberry fruit, causing them to spoil. Nevertheless, in the community, a suite of traits is selected, which also maximizes the ability of the organism to acquire limiting resources given local environmental conditions in competition with co-occurring species [36].
In summary, to respond to the question of what makes Trichoderma spp. win the competition battle over the pathogens of soft fruits, the metabolic studies of beneficial Trichoderma spp. strains mainly included the determination of the food competition between these fungi, isolated from the rhizosphere and rhizoplane of wild raspberries, and phytopathogens (Colletotrichum sp., Botrytis sp., Verticillium sp., Phytophthora sp.) attacking the organic plantations of soft fruit. Based on the research conducted, it may be concluded that the substrates preferred by Trichoderma spp., but not by pathogens, can be used as additives for biopreparations containing these beneficial fungi.
The results indicate that adenosine enhanced the growth of Trichoderma spp., but it was a source not utilized by Colletotrichum sp. fungi. These findings suggest that the addition of adenosine to biopreparations containing Trichoderma spp. may simultaneously stimulate beneficial fungi growth and negatively affect the phytopathogens of Colletotrichum sp. It has also been shown that adonitol, D-arabitol, i-erythritol, glycerol, D-mannitol and D-sorbitol can be added to the biopreparations of Trichoderma spp., and dedicated to plantations contaminated by phytopathogens of the genera Colletotrichum sp., Botrytis sp., Verticillium sp. and Phytophthora sp.
4. Materials and Methods
4.1. Fungal Strains
The following fungal pathogens were used in the study and were isolated in the Laboratory of Molecular and Environmental Microbiology, Institute of Agrophysics, Polish Academy of Sciences: two strains of Colletotrichum sp. G166/18 (GenBank: MT126798.1), G172/18 (GenBank: MT126803.1)) were isolated from infected strawberry fruit. Three strains of Botrytis sp. G277/18 (GenBank: MT154304.1), G275/18 (GenBank: MT154302.1), G276/18 (GenBank: MT154303.1) and three strains of Verticillium sp. G293/18 (GenBank: MT133324.1), G296/18 (GenBank: MT133320.1), G297/18 (GenBank: MT133316.1) and one strain of Phytophthora sp. G408/18 (GenBank: MT126670.1) were isolated from infected strawberry roots. The two environmental strains of Phytophthora sp. (G368/18 (GenBank: MT558571), G369/18 (GenBank: MT558729)) and one strain of Colletotrichum sp. (G371/18 (GenBank: MT558572)) came from the collection of the Research Institute of Horticulture in Skierniewice (Poland).
Twelve isolates of Trichoderma spp. were used: G109/18, G61/18, G65/18, G67/18, G69/18, G70/18. They were isolated in the Laboratory of Molecular and Environmental Microbiology, Institute of Agrophysics, Polish Academy of Sciences from wild raspberry rhizosphere soil using a serial dilution method; G379/18 and G398/18 were isolated from the external surface roots together with closely adhering soil particles and debris (rhizoplane); and G75/18 (GenBank: MT558563), G63/18 (GenBank: MT558561), G64/18 (GenBank: MT558562), G78/18 were isolated from wild raspberry roots.
All environmental samples used to isolate the fungi were obtained from Poland. Strawberry/raspberry roots were washed in tap water (for a few minutes in a bowl), then thoroughly rinsed with distilled water, then a surface disinfection in 70% ethanol was performed. Then the top layer of the root was removed and the interior part was cut into small fragments (several mm) and laid on a prepared media in Petri dishes (Potato Dextrose Agar, PDA, Biocorp, Warsaw, Poland). The serial dilutions method was used to isolate fungi from the rhizosphere and rhizoplane. In order to obtain the growth of the microbes, incubation was conducted at 22 °C. The passage of the fresh PDA medium was conducted several times to obtain a pure culture of fungi.
Fruit with visible traces of infestation (black lesions) were cut without the removal of changed fruit fragments and placed in a sterile medium (V8 or PDA, Biocorp, Warsaw, Poland). Incubation at 22 °C for several days (until the mycelium appears) allowed for the isolation of the fungi. The passage of the fresh PDA medium was conducted several times to obtain a pure culture of all the strains.
Genetic genus identification was confirmed using Internal Transcribed Spacer (ITS) region [37] or/and D2 Region of the Large Subunit Ribosomal RNA Gene (D2 LSU) [38] gene fragments as described by Frąc et al. [39].
All of the analyses were performed using mean values for the data obtained from the isolates mentioned above (with three independent replicates) divided according to the distinction of being a member of the following groups: Trichoderma spp., Botrytis sp., Colletotrichum sp., Phytophthora sp., Verticillium sp.
4.2. FF Plates® Preparation
The inoculation procedure was performed according to the manufacturer’s protocol with modifications as described in detail by [21] in three replicates (three separate plates for each isolate). In brief, after the homogenization of the mycelium suspension in inoculating fluid (FF-IF, Biolog®, Hayward, CA, USA), the transmittance was adjusted to 75% using a turbidimeter (Biolog®). A volume of 100 μL of the mycelium suspension was added to each well. The inoculated microplates were incubated in darkness at 25 °C within 10 days.
4.3. Group of Substrate Use—Specific Phenotypic Profiles Based on Consumption and Growth Potentiates
The optical density at 490 nm (substrate consumption, catabolism, respiration) and 750 nm (turbidity, growth, biomass formation) was determined using a microplate reader (Biolog®, Hayward, CA, USA) on a daily basis to calculate Average Well Colour Development (AWCD) and Average Well Density Development (AWDD) indices, respectively, as suggested by Jeszka-Skowron et al. [29].
Fifteen groups of these substrates were evaluated in accordance with [40] based on their chemical properties. These were as follows heptoses, hexoses, pentoses, sugar acids, hexosamines, polyols, polysaccharides, oligosaccharides, glucosides, peptides, L-amino acids, biogenic and heterocyclic amines, TCA-cycle intermediates, aliphatic organic acids, and others. For interpretation, the substrates were also divided into more general groups, namely into: monosaccharides (heptose, hexoses, pentoses), monosaccharides-related substrates (sugar acids, hexosamines, polyols), sugar-related substrates (polysaccharides, oligosaccharides), N-containing substrates (peptides, L-amino acids, biogene and heterocyclic amines, TCA-cycle intermediates, aliphatic organic acids) and others (glucosides) [30,41].
4.4. Time Point Selection
The rates of change in the total AWCD and AWDD indices values dynamics (mean for all tested strains) and the principal component analysis (PCA) were used to identify and confirm the time point (the reading hour) that best represents the greatest response of the fungal isolates according to their consumption of different substrates and growth responses, respectively. The varimax-rotated factor loadings substantially influence the principal components (>0.7) which were distinguished. An optical density higher than 0.20 for each substrate was considered to be a positive response.
4.5. Competition for Substrates Groups
In order to compare the substrate group use patterns of the pathogens to those of the beneficial Trichoderma spp., AWCD and AWDD indices, niche size, a total niche overlap index (NOITOT) and Trichoderma spp. competitiveness index (COMTRICH) were calculated according to [18] with their own modification. In brief, the niche size is the share of the positive response (%) to the substrate groups.
The NOITOT index was calculated as the number of substrates shared by both pathogens and Trichoderma spp. divided by the total number of substrates given in a particular group. The NOITOT value of 0.9 or above was assumed to indicate a high degree of niche overlap and a competitive advantage for the target fungus [18,42]. This function was used to quantify the ability of Trichoderma spp. to overcome pathogens. It was assumed that, if the value of this function was greater than 2.0, Trichoderma spp. exhibits a competitive superiority in relation to the pathogen.
The COMTRICH value indicates the relative rate of substrate usage by the pathogenic isolates compared to Trichoderma spp. (calculated as Trichoderma spp. effectiveness at using substrates included in a particular group, in comparison to pathogens). A value of 2.0 or higher indicates that the Trichoderma spp. strain is more effective at utilizing the substrates included in a particular group. A value below 1.0 means that the pathogen is more successful at substrate usage.
4.6. Stressful Metabolic Situation
The ratio was calculated for AWCD and AWDD of the substrate group for each group of fungi to indicate the specific respiration rate for the mean values of each substrate group and shows the catabolic efforts, compared with biomass development. A ratio much higher than 2.0 was regarded as indicating a stressful metabolic situation, when a small biomass yields high respiration rates [30,41].
4.7. Substrate Usage Selectivity—Preferred and Non-Preferred Substrates
The cluster analysis, particularly the grouping of objects and features with superficies visualization, was performed to denote preferred and non-preferred particular substrates among a group.
4.8. Saccharide Composition of Cell Wall Material from Strawberries
The cell wall material was extracted from strawberries cv. Dipret purchased from a local organic farmer according to the method by Renard [43] with slight modifications. The strawberries were first homogenized and then mixed and stirred with 70% ethanol for 1 h. Next, the mixture was filtered and mixed repeatedly with ethanol until a negative result was obtained from the assay concerning the presence of sugars [44].
Saccharide composition was determined according to a modified method described by Lv et al. [45]. In brief, cell wall material from strawberries (CWM) was decomposed by hydrolysis in trifluoroacetic acid (TFA) by the addition of 2 mL of 3M TFA into a glass tube with 20 mg of CWM and incubation in boiling water for 8 h. After cooling, the suspension was centrifuged and the supernatant was freeze-dried and then 1 mL of water was added to the hydrolysate.
Hydrolysed saccharides were subjected to derivatization with 1-Phenyl-3-methyl-5-pyrazolone (PMP). First, 50 μL of 0.3M NaOH and a 0.5 M methanol solution of PMP were added to hydrolysate. The mixture was incubated for 60 min at 70 °C and then cooled, neutralized with 0.3M HCl and extracted three times with 1 mL of chloroform. The aqueous layer was then filtered through a 0.45 μm membrane.
The concentration of PMP-labelled saccharides was then determined using a High Performance Liquid Chromatography (HPLC) system equipped with a S 1130 HPLC quaternary pump, S 5300 sample injector, S4120 column oven and S 3350 PAD detector (Sykam GmbH, Gewerbering, Germany). The HPLC column was a Bionacom Velocity LPH C18 (ID 4.6 × 250 mm, 5 μm), preceded by a 0.5 µm Bionacom ultra filter column protector. The injection volume was 20 μL, the flow rate was 0.8 mL min−1 and the temperature was 35 °C. The chromatograms were recorded at 248 nm. Two mobile phases, A (acetonitrile) and B (0.045% KH2PO4–0.05% triethylamine buffer, pH 7.0), were applied with a gradient elution of 90–89–86% B with a linear decrease from 0–15–40 min. Saccharide concentration was then calculated on the basis of the calibration curves that were composed of five concentrations of PMP-labelled standards: galacturonic acid, arabinose, rhamnose, galactose, glucose, rhamnose, xylose, mannose and fucose.
5. Conclusions
In summary, to respond to the question of what makes Trichoderma spp. gain supremacy in a competition battle with soft fruit pathogens, the metabolic studies of beneficial Trichoderma spp. strains mainly included the determination of food competition between these fungi, isolated from the rhizosphere and rhizoplane of wild raspberries, and phytopathogens (Colletotrichum sp., Botrytis sp., Verticillium sp., Phytophthora sp.) attacking organic plantations of soft fruit. Based on the research conducted, it may be concluded that the substrates, those preferred by Trichoderma spp., but not by pathogens, can be used as additives in biopreparations containing these beneficial fungi. The results indicate that adenosine enhanced the growth of Trichoderma spp., but it was a source that was not utilized by Colletotrichum sp. fungi.
This finding suggests that the addition of adenosine to biopreparations containing Trichoderma spp. can simultaneously stimulate beneficial fungi growth and can also negatively affect the phytopathogens of Colletotrichum sp. It has also been shown that adonitol, D-arabitol, i-erythritol, glycerol, D-mannitol and D-sorbitol can be added into the biopreparations of Trichoderma spp., and dedicated to plantations contaminated by phytopathogens of the genera Colletotrichum sp., Botrytis sp., Verticillium sp. and Phytophthora sp.
Author Contributions
Conceptualization, K.O. and M.F.; methodology, K.O. and M.F.; software, K.O.; investigation, K.O. and J.C.; writing—original draft preparation, K.O.; writing—review and editing, M.F., J.C.; visualization, K.O.; supervision, M.F.; funding acquisition, M.F. All authors have read and agreed to the published version of the manuscript.
Funding
The study was financed by The National Centre for Research and Development within the framework of the project BIOSTRATEG, contract number BIOSTRATEG3/344433/16/NCBR/2018.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| FF | Filamentous Fungi |
| COMTRICH | Trichoderma Competitiveness Index |
| NOITOT | Total Niche Overlap index |
| AWCD | Average Well Colour Development |
| AWDD | Average Well Density Development |
| HPLC | High Performance Liquid Chromatography |
| PCA | Principal Component Analysis |
| PDA | Potato Dextrose Agar |
| TCA | Tricarboxylic Acid Cycle / Citric Acid Cycle |
| CWM | Cell Wall Material |
| TFA | Trifluoroacetic Acid |
| PMP | 1-Phenyl-3-methyl-5-pyrazolone |
| FF-IF | Biolog® Filamentous Fungi Inoculating Fluid |
| ITS | Internal Transcribed Spacer region |
| D2 LSU | D2 Region of the Large Subunit Ribosomal RNA Gene |
References
- Ceci, A.; Pinzari, F.; Russo, F.; Persiani, A.M.; Gadd, G.M. Roles of saprotrophic fungi in biodegradation or transformation of organic and inorganic pollutants in co-contaminated sites. Appl. Microb. Biotech. 2019, 103, 53–68. [Google Scholar] [CrossRef] [PubMed]
- Vega, D.; Gally, M.E.; Romero, A.M.; Poggio, S.L. Functional groups of plant pathogens in agroecosystems: A review. Eur. J. Plant Pathol. 2019, 153, 695–713. [Google Scholar] [CrossRef]
- Aladdin, A.; Dib, J.R.; Malek, R.A.; El Enshasy, H.A. Killer Yeast, a novel biological control of soilborne diseases for good agriculture practice. In Sustainable Technologies for the Management of Agricultural Wastes; Zakaria, Z.A., Ed.; Springer Singapore: Singapore, 2018; pp. 71–86. [Google Scholar] [CrossRef]
- Frąc, M.; Hannula, S.E.; Bełka, M.; Jędryczka, M. Fungal biodiversity and their role in soil health. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Malarczyk, D.; Panek, J.; Frac, M. Alternative molecular-based diagnostic methods of plant pathogenic fungi affecting berry crops—A Review. Molecules 2019, 24. [Google Scholar] [CrossRef]
- Szczałba, M.; Kopta, T.; Gąstoł, M.; Sękara, A. Comprehensive insight into arbuscular mycorrhizal fungi, Trichoderma spp. and plant multilevel interactions with emphasis on biostimulation of horticultural crops. J. Appl. Microb. 2019, 127, 630–647. [Google Scholar] [CrossRef]
- Pylak, M.; Oszust, K.; Frąc, M. Review report on the role of bioproducts, biopreparations, biostimulants and microbial inoculants in organic production of fruit. Rev. Environ. Sci. Bio Technol. 2019, 18, 597–616. [Google Scholar] [CrossRef]
- Weber, R.W.S.; Hahn, M. Grey mould disease of strawberry in northern Germany: Causal agents, fungicide resistance and management strategies. Appl. Microb. Biotech. 2019, 103, 1589–1597. [Google Scholar] [CrossRef]
- Fan, R.; Cockerton, H.M.; Armitage, A.D.; Bates, H.; Cascant-Lopez, E.; Antanaviciute, L.; Xu, X.; Hu, X.; Harrison, R.J. Vegetative compatibility groups partition variation in the virulence of Verticillium dahliae on strawberry. PLoS ONE 2018, 13, e0191824. [Google Scholar] [CrossRef]
- Nellist, C.F.; Vickerstaff, R.J.; Sobczyk, M.K.; Marina-Montes, C.; Wilson, F.M.; Simpson, D.W.; Whitehouse, A.B.; Harrison, R.J. Quantitative trait loci controlling Phytophthora cactorum resistance in the cultivated octoploid strawberry (Fragaria x ananassa). Hortic. Res. 2019, 6, 60. [Google Scholar] [CrossRef]
- Meszka, B.; Michalecka, M. Identification of Phytophthora spp. isolated from plants and soil samples on strawberry plantations in Poland. J. Plant Dis. Protect. 2016, 123, 29–36. [Google Scholar] [CrossRef]
- Ponisio, L.C.; Valdovinos, F.S.; Allhoff, K.T.; Gaiarsa, M.P.; Barner, A.; Guimarães, P.R.; Hembry, D.H.; Morrison, B.; Gillespie, R. A network perspective for community assembly. Front. Ecol. Evol. 2019, 7. [Google Scholar] [CrossRef]
- Kumar, A.; Verma, J.P. The role of microbes to improve crop productivity and soil health. In Ecological Wisdom Inspired Restoration Engineering; Achal, V., Mukherjee, A., Eds.; Springer Singapore: Singapore, 2019; pp. 249–265. [Google Scholar] [CrossRef]
- Latha, P.; Karthikeyan, M.; Rajeswari, E. Endophytic bacteria: Prospects and applications for the plant disease management. In Plant Health Under Biotic Stress: Volume 2: Microbial Interactions; Ansari, R.A., Mahmood, I., Eds.; Springer Singapore: Singapore, 2019; pp. 1–50. [Google Scholar] [CrossRef]
- Silva, R.N.; Monteiro, V.N.; Steindorff, A.S.; Gomes, E.V.; Noronha, E.F.; Ulhoa, C.J. Trichoderma/pathogen/plant interaction in pre-harvest food security. Fungal Biol. 2019, 123, 565–583. [Google Scholar] [CrossRef] [PubMed]
- Köhl, J.; Kolnaar, R.; Ravensberg, W.J. Mode of action of microbial biological control agents against plant diseases: Relevance beyond efficacy. Front. Plant. Sci. 2019, 10, 845. [Google Scholar] [CrossRef] [PubMed]
- Patel, J.; Teli, B.; Bajpai, R.; Meher, J.; Rashid, M.; Mukherjee, A.; Yadav, S.K. 13—Trichoderma-mediated biocontrol and growth promotion in plants: An endophytic approach. In Role of Plant Growth Promoting Microorganisms in Sustainable Agriculture and Nanotechnology; Kumar, A., Singh, A.K., Choudhary, K.K., Eds.; Woodhead Publishing: Cambridge, UK, 2019; pp. 219–239. [Google Scholar] [CrossRef]
- Blumenstein, K.; Albrectsen, B.R.; Martín, J.A.; Hultberg, M.; Sieber, T.N.; Helander, M.; Witzell, J. Nutritional niche overlap potentiates the use of endophytes in biocontrol of a tree disease. BioControl 2015, 60, 655–667. [Google Scholar] [CrossRef]
- Hewitt, S.K.; Foster, D.S.; Dyer, P.S.; Avery, S.V. Phenotypic heterogeneity in fungi: Importance and methodology. Fungal Biol. Rev. 2016, 30, 176–184. [Google Scholar] [CrossRef]
- Mikkelson, G.M. Niche-based vs. neutral models of ecological communities. Biol. Phil. 2005, 20, 557–566. [Google Scholar] [CrossRef]
- Oszust, K.; Panek, J.; Pertile, G.; Siczek, A.; Oleszek, M.; Frąc, M. Metabolic and genetic properties of petriella setifera precultured on waste. Front. Microb. 2018, 9. [Google Scholar] [CrossRef]
- Pawlik, A.; Ruminowicz-Stefaniuk, M.; Frąc, M.; Mazur, A.; Wielbo, J.; Janusz, G. The wood decay fungus Cerrena unicolor adjusts its metabolism to grow on various types of wood and light conditions. PLoS ONE 2019, 14, e0211744. [Google Scholar] [CrossRef]
- Wang, H.-C.; Li, L.-C.; Cai, B.; Cai, L.-T.; Chen, X.-J.; Yu, Z.-H.; Zhang, C.-Q. Metabolic phenotype characterization of Botrytis cinerea, the causal agent of gray mold. Front. Microb. 2018, 9. [Google Scholar] [CrossRef]
- Pertile, G.; Frąc, M.; Fornal, E.; Oszust, K.; Gryta, A.; Yaguchi, T. Molecular and metabolic strategies for postharvest detection of heat-resistant fungus Neosartorya fischeri and its discrimination from Aspergillus fumigatus. Postharv. Biol. Technol. 2020, 161, 111082. [Google Scholar] [CrossRef]
- Blumenstein, K.; Macaya-Sanz, D.; Martín, J.A.; Albrectsen, B.R.; Witzell, J. Phenotype MicroArrays as a complementary tool to next generation sequencing for characterization of tree endophytes. Front. Microb. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Ceci, A.; Pinzari, F.; Riccardi, C.; Maggi, O.; Pierro, L.; Petrangeli Papini, M.; Gadd, G.M.; Persiani, A.M. Metabolic synergies in the biotransformation of organic and metallic toxic compounds by a saprotrophic soil fungus. Appl. Microb. Biotech. 2018, 102, 1019–1033. [Google Scholar] [CrossRef] [PubMed]
- Pertile, G.; Panek, J.; Oszust, K.; Siczek, A.; Frac, M. Intraspecific functional and genetic diversity of Petriella setifera. PeerJ 2018, 6, e4420. [Google Scholar] [CrossRef] [PubMed]
- Frąc, M.; Gryta, A.; Oszust, K.; Kotowicz, N. Fast and accurate microplate method (Biolog MT2) for detection of Fusarium fungicides resistance/sensitivity. Front. Microb. 2016, 7, 489. [Google Scholar] [CrossRef] [PubMed]
- Jeszka-Skowron, M.; Oszust, K.; Zgoła-Grześkowiak, A.; Frąc, M. Quality assessment of goji fruits, cranberries, and raisins using selected markers. Eur. Food Res. Technol. 2018, 244, 2159–2168. [Google Scholar] [CrossRef]
- Pinzari, F.; Maggi, O.; Lunghini, D.; Di Lonardo, D.P.; Persiani, A.M. A simple method for measuring fungal metabolic quotient and comparing carbon use efficiency of different isolates: Application to Mediterranean leaf litter fungi. Plant Biol. 2017, 151, 371–376. [Google Scholar] [CrossRef][Green Version]
- Oleszek, M.; Pecio, L.; Kozachok, S.; Lachowska-Filipiuk, Z.; Oszust, K.; Frac, M. Phytochemicals of apple pomace as prospect bio-fungicide agents against mycotoxigenic fungal species-in vitro experiments. Toxins 2019, 11. [Google Scholar] [CrossRef]
- Lopes, F.A.; Steindorff, A.S.; Geraldine, A.M.; Brandao, R.S.; Monteiro, V.N.; Lobo, M., Jr.; Coelho, A.S.; Ulhoa, C.J.; Silva, R.N. Biochemical and metabolic profiles of Trichoderma strains isolated from common bean crops in the Brazilian Cerrado, and potential antagonism against Sclerotinia sclerotiorum. Fungal Biol. 2012, 116, 815–824. [Google Scholar] [CrossRef]
- Mason, N.W.H.; de Bello, F.; Doležal, J.; Lepš, J. Niche overlap reveals the effects of competition, disturbance and contrasting assembly processes in experimental grassland communities. J. Ecol. 2011, 99, 788–796. [Google Scholar] [CrossRef]
- Dou, K.; Gao, J.; Zhang, C.; Yang, H.; Jiang, X.; Li, J.; Li, Y.; Wang, W.; Xian, H.; Li, S.; et al. Trichoderma biodiversity in major ecological systems of China. J. Microb. 2019, 57, 668–675. [Google Scholar] [CrossRef]
- Jaroszuk-Scisel, J.; Tyskiewicz, R.; Nowak, A.; Ozimek, E.; Majewska, M.; Hanaka, A.; Tyskiewicz, K.; Pawlik, A.; Janusz, G. Phytohormones (auxin, gibberellin) and ACC deaminase in vitro synthesized by the mycoparasitic Trichoderma DEMTkZ3A0 strain and changes in the level of auxin and plant resistance markers in wheat seedlings inoculated with this strain conidia. Int. J. Mol. Sci. 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Lozada-Gobilard, S.; Stang, S.; Pirhofer-Walzl, K.; Kalettka, T.; Heinken, T.; Schroder, B.; Eccard, J.; Joshi, J. Environmental filtering predicts plant-community trait distribution and diversity: Kettle holes as models of meta-community systems. Ecol. Evol. 2019, 9, 1898–1910. [Google Scholar] [CrossRef] [PubMed]
- Pryce, T.M.; Palladino, S.; Kay, I.D.; Coombs, G.W. Rapid identification of fungi by sequencing the ITS1 and ITS2 regions using an automated capillary electrophoresis system. Med. Mycol. 2003, 41, 369–381. [Google Scholar] [CrossRef] [PubMed]
- Rozynek, P.; Gilges, S.; Brüning, T.; Wilhelm, M. Quality test of the MicroSeq D2 LSU Fungal Sequencing Kit for the identification of fungi. Int. J. Hygiene Environ. Health 2004, 207, 297–299. [Google Scholar] [CrossRef]
- Frac, M.; Oszust, K.; Lipiec, J.; Jezierska-Tys, S.; Nwaichi, E.O. Soil microbial functional and fungal diversity as influenced by municipal sewage sludge accumulation. Int. J. Environ. Res. Public Health 2014, 11, 8891–8908. [Google Scholar] [CrossRef]
- Atanasova, L.; Druzhinina, I.S. Review: Global nutrient profiling by Phenotype MicroArrays: A tool complementing genomic and proteomic studies in conidial fungi. J. Zhejiang Univ. Sci. B 2010, 11, 151–168. [Google Scholar] [CrossRef]
- Pinzari, F.; Ceci, A.; Abu-Samra, N.; Canfora, L.; Maggi, O.; Persiani, A. Phenotype MicroArray system in the study of fungal functional diversity and catabolic versatility. Res. Microb. 2016, 167, 710–722. [Google Scholar] [CrossRef]
- Lee, H.B.; Magan, N. Environmental factors and nutritional utilization patterns affect niche overlap indices between Aspergillus ochraceus and other spoilage fungi. Lett. Appl. Microb. 1999, 28, 300–304. [Google Scholar] [CrossRef]
- Renard, C.M.G.C. Variability in cell wall preparations: Quantification and comparison of common methods. Carbohydr. Polym. 2005, 60, 515–522. [Google Scholar] [CrossRef]
- DuBois, M.; Gilles, K.A.; Hamilton, J.K.; Rebers, P.A.; Smith, F. Colorimetric method for determination of sugars and related substances. Anal. Chem. 1956, 28, 350–356. [Google Scholar] [CrossRef]
- Lv, Y.; Yang, X.; Zhao, Y.; Ruan, Y.; Yang, Y.; Wang, Z. Separation and quantification of component monosaccharides of the tea polysaccharides from Gynostemma pentaphyllum by HPLC with indirect UV detection. Food Chem. 2009, 112, 742–746. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).