Effect of trans-Cinnamaldehyde on Methicillin-Resistant Staphylococcus aureus Biofilm Formation: Metabolic Activity Assessment and Analysis of the Biofilm-Associated Genes Expression
Abstract
1. Introduction
2. Results
2.1. Antibacterial Activity of TC against MRSA Strains
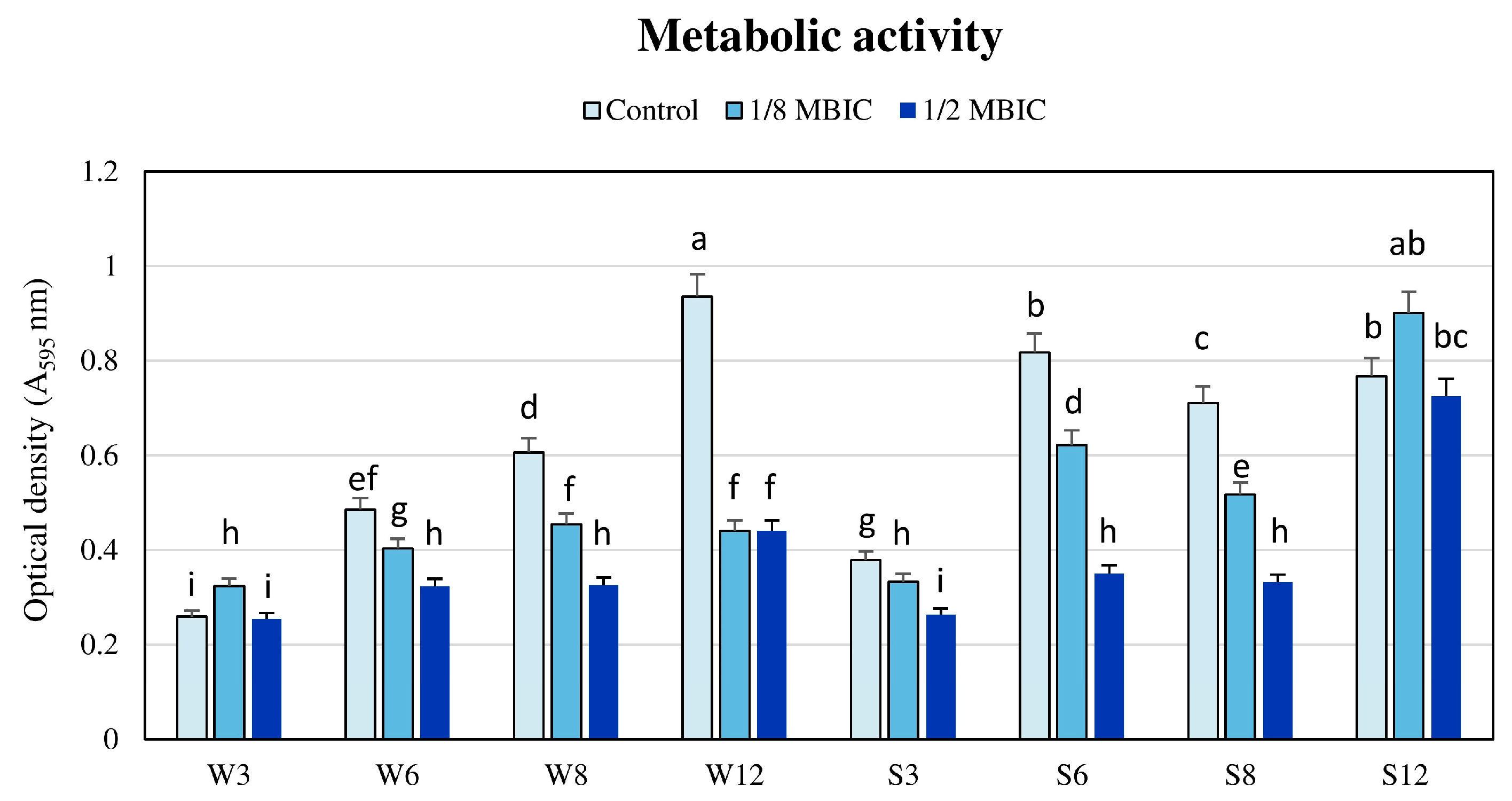
2.2. The Influence of TC on Metabolic Activity of Biofilms in Different Time Intervals
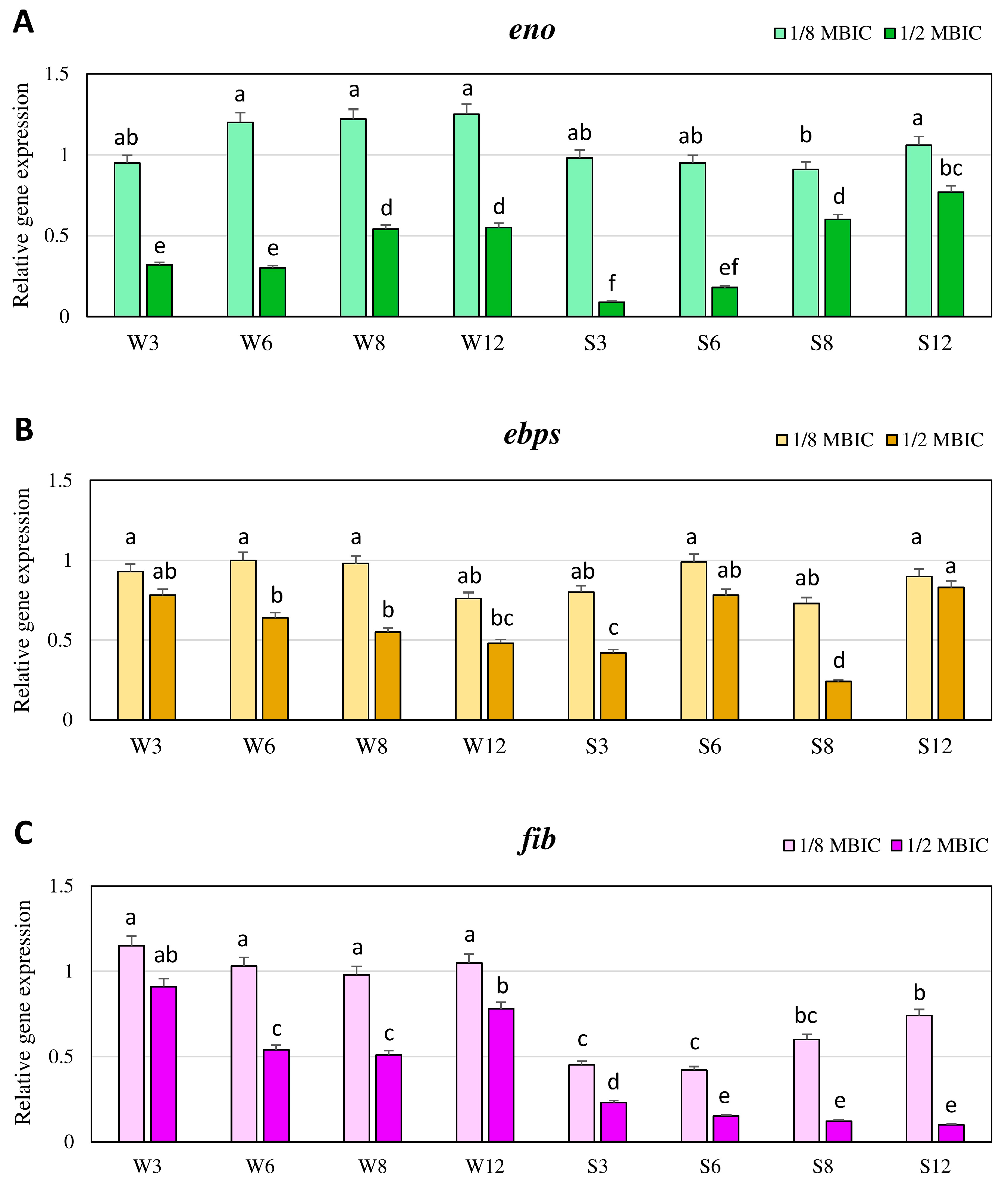
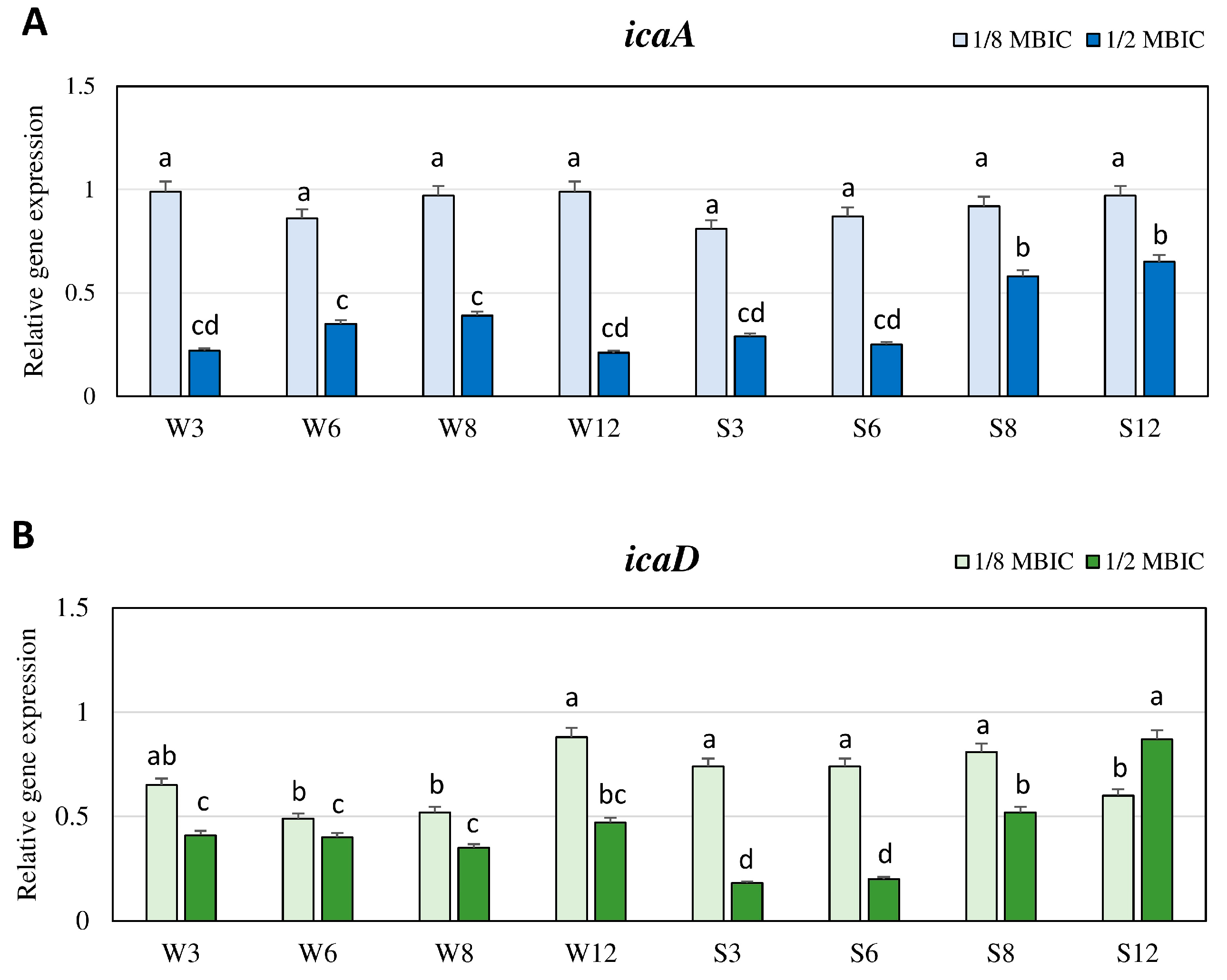
2.3. Expression Levels of Genes Associated with Biofilm Formation in the Presence of TC Quantified by Real-Time qRT-PCR
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains
4.2. Determination of Minimum Inhibitory Concentrations (MIC) and Minimum Bactericidal Concentrations (MBC) of TC
4.3. Determination of Minimum Biofilm Inhibition Concentrations (MBIC) and Minimum Biofilm Eradication Concentrations (MBEC) of TC
4.4. TC Inhibiting Biofilm Formation Assay in Different Time Intervals
4.5. Growth Conditions of MRSA Strains
4.6. Preparation of the Lysate from Bacterial Cells, RNA Extraction and cDNA Synthesis
4.7. Gene Quantification
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kot, B.; Wierzchowska, K.; Piechota, M.; Grużewska, A. Antimicrobial resistance patterns in methicillin-resistant Staphylococcus aureus from patients hospitalized during 2015–2017 in hospitals in Poland. Med. Princ. Pract. 2019. [Google Scholar] [CrossRef] [PubMed]
- Morgenstern, M.; Erichsen, C.; Hackl, S.; Mily, J.; Militz, M.; Friederichs, J.; Hungerer, S.; Bühren, V.; Moriarty, T.F.; Post, V.; et al. Antibiotic resistance of commensal Staphylococcus aureus and coagulase-negative staphylococci in an international cohort of surgeons: A prospective point-prevalence study. PLoS ONE 2016, 11, e0148437. [Google Scholar] [CrossRef] [PubMed]
- Macià, M.D.; Perez, J.L.; Molin, S.; Oliver, A. Dynamics of mutator and antibiotic-resistant populations in a pharmacokinetic/pharmacodynamics model of Pseudomonas aeruginosa biofilm treatment. Antimicrob. Agents Chemother. 2011, 55, 5230–5237. [Google Scholar] [CrossRef] [PubMed]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef]
- Kot, B.; Sytykiewicz, H.; Sprawka, I. Expression of the biofilm-associated genes in methicillin-resistant Staphylococcus aureus in biofilm and planktonic conditions. Int. J. Mol. Sci. 2018, 19, 3487. [Google Scholar] [CrossRef]
- Carneiro, C.R.; Postol, E.; Nomizo, R.; Reis, L.F.; Brentani, R.R. Identification of enolase as a laminin-binding protein on the surface of Staphylococcus aureus. Microbes. Infect. 2004, 6, 604–608. [Google Scholar] [CrossRef]
- Downer, R.; Roche, F.; Park, P.W.; Mecham, R.P.; Foster, T.J. The elastin-binding protein of Staphylococcus aureus (EbpS) is expressed at the cell surface as an integral membrane protein and not as a cell wall-associated protein. J. Biol. Chem. 2002, 277, 243–250. [Google Scholar] [CrossRef]
- Hartford, O.M.; Wann, E.R.; Höök, M.; Foster, T.J. Identification of residues in the Staphylococcus aureus fibrinogen-binding MSCRAMM clumping factor A (ClfA) that are important for ligand binding. J. Biol. Chem. 2001, 276, 2466–2473. [Google Scholar] [CrossRef]
- Cramton, S.E.; Ulrich, M.; Gotz, F.; Döring, G. Anaerobic conditions induce expression of polysaccharide intercellular adhesin in Staphylococcus aureus and Staphylococcus epidermidis. Infect. Immun. 2001, 69, 4079–4085. [Google Scholar] [CrossRef]
- Cue, D.; Lei, M.G.; Lee, C.Y. Genetic regulation of the intercellular adhesion locus in staphylococci. Front. Cell Infect. Microbiol. 2012, 2, 38. [Google Scholar] [CrossRef]
- Cerca, N.; Jefferson, K.K.; Maira-Litrán, T.; Pier, D.B.; Kelly-Quintos, C.; Goldmann, D.A.; Azeredo, J.; Pier, G.B. Molecular basis for preferential protective efficacy of antibodies directed to the poorly acetylated form of staphylococcal poly-N-acetyl-β-(1-6)-glucosamine. Infect. Immun. 2007, 75, 3406–3413. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, A.; Jain, A. Glucose & sodium chloride induced biofilm production & ica operon in clinical isolates of staphylococci. Indian J. Med. Res. 2013, 138, 262–266. [Google Scholar] [PubMed]
- Arciola, C.R.; Campoccia, D.; Baldassari, L.; Donati, M.E.; Pirini, V. Detection of biofilm formation in Staphylococcus epidermidis from implant infections. Comparison of a PCR-method that recognizes the presence of ica genes with two classic phenotypic methods. J. Biomed. Mater. Res. 2006, 76, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Ceylan, O.; Ugur, A. Chemical composition and anti-biofilm activity of Thymus sipyleus BOISS. subsp. sipyleus BOISS. var. davisianus RONNIGER essential oil. Arch. Pharm. Res. 2015, 38, 957–965. [Google Scholar] [CrossRef] [PubMed]
- Macià, M.D.; Rojo-Molinero, E.; Oliver, A. Antimicrobial susceptibility testing in biofilm-growing bacteria. Clin. Microbiol. Infect. 2014, 20, 981–990. [Google Scholar] [CrossRef] [PubMed]
- Beenken, K.E.; Dunman, P.M.; McAleese, F.; Macapagal, D.; Murphy, E.; Projan, S.J.; Blevins, J.S.; Smeltzer, M.S. Global gene expression in Staphylococcus aureus biofilms. J. Bacteriol. 2004, 186, 4665–4684. [Google Scholar] [CrossRef]
- Seo, Y.S.; Lee, D.Y.; Rayamahji, N.; Kang, M.L.; Yoo, H.S. Biofilm-forming associated genotypic and phenotypic characteristics of Staphylococcus spp. isolated from animals and air. Res. Vet. Sci. 2008, 85, 433–438. [Google Scholar] [CrossRef]
- Ghasemian, A.; Najar Peerayeh, S.; Bakhshi, B.; Mirzaee, M. The microbial surface components recognizing adhesive matrix molecules (MSCRAMMs) genes among clinical isolates of Staphylococcus aureus from hospitalized children. Iran. J. Pathol. 2015, 10, 258–264. [Google Scholar]
- Yuan, J.H.; Dieter, M.P.; Bucher, J.R.; Jameson, C.W. Toxicokinetics of cinnamaldehyde in F344 rats. Food Chem. Toxicol. 1992, 30, 997–1004. [Google Scholar] [CrossRef]
- Upadhyaya, I.; Upadhyay, A.; Kollanoor-Johny, A.; Mooyottu, S.; Baskaran, S.A.; Yin, H.B.; Schreiber, D.T.; Khan, M.I.; Darre, M.J.; Curtis, P.A.; et al. In-feed supplementation of trans-cinnamaldehyde reduces layer-chicken eggborne transmission of Salmonella enterica serovar enteritidis. Appl. Environ. Microbiol. 2015, 81, 2985–2994. [Google Scholar] [CrossRef]
- Ribeiro, M.; Malheiro, J.; Grenho, L.; Fernandes, M.H.; Simões, M. Cytotoxicity and antimicrobial action of selected phytochemicals against planktonic and sessile Streptococcus mutans. PeerJ 2018, 6, e4872. [Google Scholar] [CrossRef] [PubMed]
- Ferro, T.A.; Araújo, J.M.; Dos Santos Pinto, B.L.; Dos Santos, J.S.; Souza, E.B.; da Silva, B.L.; Colares, V.L.; Novais, T.M.; Filho, C.M.; Struve, C.; et al. Cinnamaldehyde inhibits Staphylococcus aureus virulence factors and protects against infection in a Galleria mellonella model. Front. Microbiol. 2016, 7, 2052. [Google Scholar] [CrossRef] [PubMed]
- Kot, B.; Wicha, J.; Piechota, M.; Wolska, K.; Grużewska, A. Antibiofilm activity of trans-cinnamaldehyde, p-coumaric, and ferulic acids on uropathogenic Escherichia coli. Turk. J. Med. Sci. 2015, 45, 919–924. [Google Scholar] [CrossRef] [PubMed]
- Nabavi, S.F.; Di Lorenzo, A.; Izadi, M.; Sobarzo-Sanchez, E.; Daglia, M.; Nabavi, S.M. Antibacterial effects of cinnamon: From farm to food, cosmetic and pharmaceutical industries. Nutrients 2015, 7, 7729–7748. [Google Scholar] [CrossRef] [PubMed]
- Kot, B.; Wierzchowska, K.; Grużewska, A.; Lohinau, D. The effects of selected phytochemicals on biofilm formed by five methicillin-resistant Staphylococcus aureus. Nat. Prod. Res. 2018, 32, 1299–1302. [Google Scholar] [CrossRef]
- Vasconcelos, N.G.; Croda, J.; Simionatto, S. Antibacterial mechanisms of cinnamon and its constituents: A review. Microb. Pathog. 2018, 120, 198–203. [Google Scholar] [CrossRef]
- Firmino, D.F.; Cavalcante, T.T.A.; Gomes, G.A.; Firmino, N.C.S.; Rosa, L.D.; de Carvalho, M.G.; Catunda, F.E.A., Jr. Antibacterial and antibiofilm activities of cinnamomum sp. essential oil and cinnamaldehyde: Antimicrobial activities. TSWJ 2018, 2018, 7405736. [Google Scholar] [CrossRef]
- Li, T.; Wang, D.; Liu, N.; Ma, Y.; Ding, T.; Mei, Y.; Li, J. Inhibition of quorum sensing-controlled virulence factors and biofilm formation in Pseudomonas fluorescens by cinnamaldehyde. Int. J. Food Microbiol. 2018, 269, 98–106. [Google Scholar] [CrossRef]
- Albano, M.; Crulhas, B.P.; Alves, F.C.B.; Pereira, A.F.M.; Andrade, B.F.; Barbosa, L.N.; Furlanetto, A.; da Silveira Lyra, L.P.; Rall, V.L.M.; Fernandes Júnior, A. Antibacterial and anti-biofilm activities of cinnamaldehyde against S. epidermidis. Microb. Pathog. 2019, 126, 231–238. [Google Scholar] [CrossRef]
- Kim, Y.G.; Lee, J.H.; Kim, S.I.; Baek, K.H.; Lee, J. Cinnamon bark oil and its components inhibit biofilm formation and toxin production. Int. J. Food Microbiol. 2015, 195, 30–39. [Google Scholar] [CrossRef]
- Nowotarska, S.W.; Nowotarski, K.; Grant, I.R.; Elliott, C.T.; Friedman, M.; Situ, C. Mechanisms of antimicrobial action of cinnamon and oregano oils, cinnamaldehyde, carvacrol, 2,5-dihydroxybenzaldehyde, and 2-hydroxy-5-methoxybenzaldehyde against Mycobacterium avium subsp. paratuberculosis (Map). Foods 2017, 6, 72. [Google Scholar] [CrossRef] [PubMed]
- Shen, S.; Zhang, T.; Yuan, Y.; Lin, S.; Xu, J.; Ye, H. Effects of cinnamaldehyde on Escherichia coli and Staphylococcus aureus membrane. Food Control 2015, 47, 196–202. [Google Scholar] [CrossRef]
- Abdelhamed, H.; Ozdemir, O.; Ibrahim, I.; Lawrence, M.; Karsi, A. Antibacterial activities of trans-cinnamaldehyde, caprylic acid, and β-resorcylic acid against catfish pathogens. Aquaculture 2019, 504, 334–344. [Google Scholar] [CrossRef]
- Mousavi, F.; Bojko, B.; Bessonneau, V.; Pawliszyn, J. Cinnamaldehyde characterization as an antibacterial agent toward E. coli metabolic profile using 96-blade solid-phase microextraction coupled to liquid chromatography-mass spectrometry. J. Proteome Res. 2016, 15, 963–975. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Niu, H.; Zhang, W.; Mu, H.; Sun, C.; Duan, J. Synergy among thymol, eugenol, berberine, cinnamaldehyde and streptomycin against planktonic and biofilm-associated food-borne pathogens. Lett. Appl. Microbiol. 2015, 60, 421–430. [Google Scholar] [CrossRef] [PubMed]
- El Atki, Y.; Aouam, I.; El Kamari, F.; Taroq, A.; Nayme, K.; Timinouni, M.; Lyoussi, B.; Abdellaoui, A. Antibacterial activity of cinnamon essential oils and their synergistic potential with antibiotics. J. Adv. Pharm. Technol. Res. 2019, 10, 63–67. [Google Scholar] [CrossRef]
- Domadia, P.; Swarup, S.; Bhunia, A.; Sivaraman, J.; Dasgupta, D. Inhibition of bacterial cell division protein FtsZ by cinnamaldehyde. Biochem. Pharmacol. 2007, 74, 831–840. [Google Scholar] [CrossRef]
- Rachid, S.; Ohlsen, K.; Witte, W.; Hacker, J.; Ziebuhr, W. Effect of subinhibitory antibiotic concentration on polysaccharide intercellular adhesion expression in biofilm-forming Staphylococcus epidermidis. Antimicrob. Agents Chemother. 2000, 44, 3357–3363. [Google Scholar] [CrossRef]
- Knobloch, J.K.M.; Horstkotte, M.A.; Rohde, H.; Kaulfers, P.; Mack, D. Alcoholic ingredients in skin disinfectants increase biofilm expression of Staphylococcus epidermidis. J. Antimicrob. Chemother. 2002, 49, 683–687. [Google Scholar] [CrossRef]
- Kaplan, J.B.; Izano, E.A.; Gopal, P.; Karwacki, M.T.; Kim, S.; Bose, J.L.; Bayles, K.W.; Horswill, A.R. Lowlevelsof β-lactam antibiotics induce extracellular DNA release and biofilm formation in Staphylococcus aureus. MBio 2012, 3, e00198-12. [Google Scholar] [CrossRef]
- Schilcher, K.; Andreoni, F.; Haunreiter, V.D.; Seidl, K.; Hasse, B.; Zinkernagel, A.S. Modulation of Staphylococcus aureus biofilm matrix by subinhibitory concentrations of clindamycin. Antimicrob. Agents Chemother. 2016, 60, 5957–5967. [Google Scholar] [CrossRef] [PubMed]
- Nuryastuti, T.; van der Mei, H.C.; Busscher, H.J.; Iravati, S.; Aman, A.T.; Krom, B.P. Effect of cinnamon oil on icaA expression and biofilm formation by Staphylococcus epidermidis. Appl. Environ. Microbiol. 2009, 75, 6850–6855. [Google Scholar] [CrossRef] [PubMed]
- Kot, B.; Wierzchowska, K.; Piechota, M.; Czerniewicz, P.; Chrzanowski, G. Antimicrobial activity of five essential oils from lamiaceae against multidrug-resistant Staphylococcus aureus. Nat. Prod. Res. 2019, 33, 3587–3591. [Google Scholar] [CrossRef] [PubMed]
- Welch, K.; Cai, Y.; Strømme, M. A method for quantitative determination of biofilm viability. J. Funct. Biomater. 2012, 3, 418–431. [Google Scholar] [CrossRef]
- Pettit, R.K.; Weber, C.A.; Pettit, G.R. Application of a high throughput Alamar blue biofilm susceptibility assay to Staphylococcus aureus biofilms. Ann. Clin. Microbiol. Antimicrob. 2009, 8, 28. [Google Scholar] [CrossRef]
- Borges, A.; . Simões, L.C.; Saavedra, M.J.; Simões, M. The action of selected isothiocyanates on bacterial biofilm prevention and control. Int. Biodeterior. Biodegrad. 2014, 86, 25–33. [Google Scholar] [CrossRef]
- Pettit, R.K.; Weber, C.A.; Kean, M.J.; Hoffmann, H.; Pettit, G.R.; Tan, R.; Franks, K.S.; Horton, M.L. Microplate alamar blue assay for Staphylococcus epidermidis biofilm susceptibility testing. Antimicrob. Agents Chemother. 2005, 49, 2612–2617. [Google Scholar] [CrossRef]
- Mariscal, A.; Lopez-Gigosos, R.M.; Carnero-Varo, M.; Fernandez-Crehuet, J. Fluorescent assay based on resazurin for detection of activity of disinfectants against bacterial biofilm. Appl. Microbiol. Biotechnol. 2009, 82, 773–783. [Google Scholar] [CrossRef]
- Borges, A.; Saavedra, M.J.; Simões, M. The activity of ferulic and gallic acids in biofilm prevention and control of pathogenic bacteria. Biofouling 2012, 28, 755–767. [Google Scholar] [CrossRef]
| Strain | Source | Biofilm | trans-Cinnamaldehyde (mg/mL) | |||
|---|---|---|---|---|---|---|
| Planktonic Cells | Biofilm | |||||
| MIC | MBC | MBIC | MBEC | |||
| 27,887 | Wound | Strong | 0.24 | 0.48 | 0.48 | 0.96 |
| 1037 | Anus | Weak | 0.06 | 0.06 | 0.12 | 1.92 |
| Genes | Accession No. (GenBank) | Sequences of Primers and Probes |
|---|---|---|
| icaA | SAB2541 (K11936) | F: CAATACTATTTCGGGTGTCTTCACTCT R: CAAGAAACTGCAATATCTTCGGTAATCAT P: 5′-FAM-CCCAGTAGCCAACATC-NFQ-3′ |
| icaD | SAB2542 (K21461) | F: TCAAGCCCAGACAGAGGGAATA R: ACACGATATAGCGATAAGTGCTGTTT P: 5′-FAM-CCCAACGCTAAAATC-NFQ-3′ |
| eno | AF065394.1 | F: AAACTGCCGTAGGTGACGAA R: TGTTTCAACAGCATCTTCAGTACCTT P: 5′-FAM- TTCGCTCCTAAATTTG-NFQ-3′ |
| ebps | SAB1343c | F: ACATTCAAATGACGCTCAAAACAAAAGT R: CTTATCTTGAGACGCTTTATCCTCAGT P: 5′-FAM- CAAGGCGAATAACTCG-NFQ-3′ |
| fib | SAB1021 (K14200) | F: GAATATGGTGCACGTCCACAATT R: AAGATTTTGAGCTTGAATCAATTTTTGTTCTTTTT P: 5′-FAM-TCGCTGCTGGTTTATT-NFQ-3′ |
| rpoB | KY086792.1 | F: CAGCTGACGAAGAAGATAGCTATGT R: ACTTCATCATCCATGAAACGACCAT P: 5′-TAGCACAAGCAAACTC-NFQ-3′ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kot, B.; Sytykiewicz, H.; Sprawka, I.; Witeska, M. Effect of trans-Cinnamaldehyde on Methicillin-Resistant Staphylococcus aureus Biofilm Formation: Metabolic Activity Assessment and Analysis of the Biofilm-Associated Genes Expression. Int. J. Mol. Sci. 2020, 21, 102. https://doi.org/10.3390/ijms21010102
Kot B, Sytykiewicz H, Sprawka I, Witeska M. Effect of trans-Cinnamaldehyde on Methicillin-Resistant Staphylococcus aureus Biofilm Formation: Metabolic Activity Assessment and Analysis of the Biofilm-Associated Genes Expression. International Journal of Molecular Sciences. 2020; 21(1):102. https://doi.org/10.3390/ijms21010102
Chicago/Turabian StyleKot, Barbara, Hubert Sytykiewicz, Iwona Sprawka, and Małgorzata Witeska. 2020. "Effect of trans-Cinnamaldehyde on Methicillin-Resistant Staphylococcus aureus Biofilm Formation: Metabolic Activity Assessment and Analysis of the Biofilm-Associated Genes Expression" International Journal of Molecular Sciences 21, no. 1: 102. https://doi.org/10.3390/ijms21010102
APA StyleKot, B., Sytykiewicz, H., Sprawka, I., & Witeska, M. (2020). Effect of trans-Cinnamaldehyde on Methicillin-Resistant Staphylococcus aureus Biofilm Formation: Metabolic Activity Assessment and Analysis of the Biofilm-Associated Genes Expression. International Journal of Molecular Sciences, 21(1), 102. https://doi.org/10.3390/ijms21010102






