OsbZIP81, A Homologue of Arabidopsis VIP1, May Positively Regulate JA Levels by Directly Targetting the Genes in JA Signaling and Metabolism Pathway in Rice
Abstract
1. Introduction
2. Results
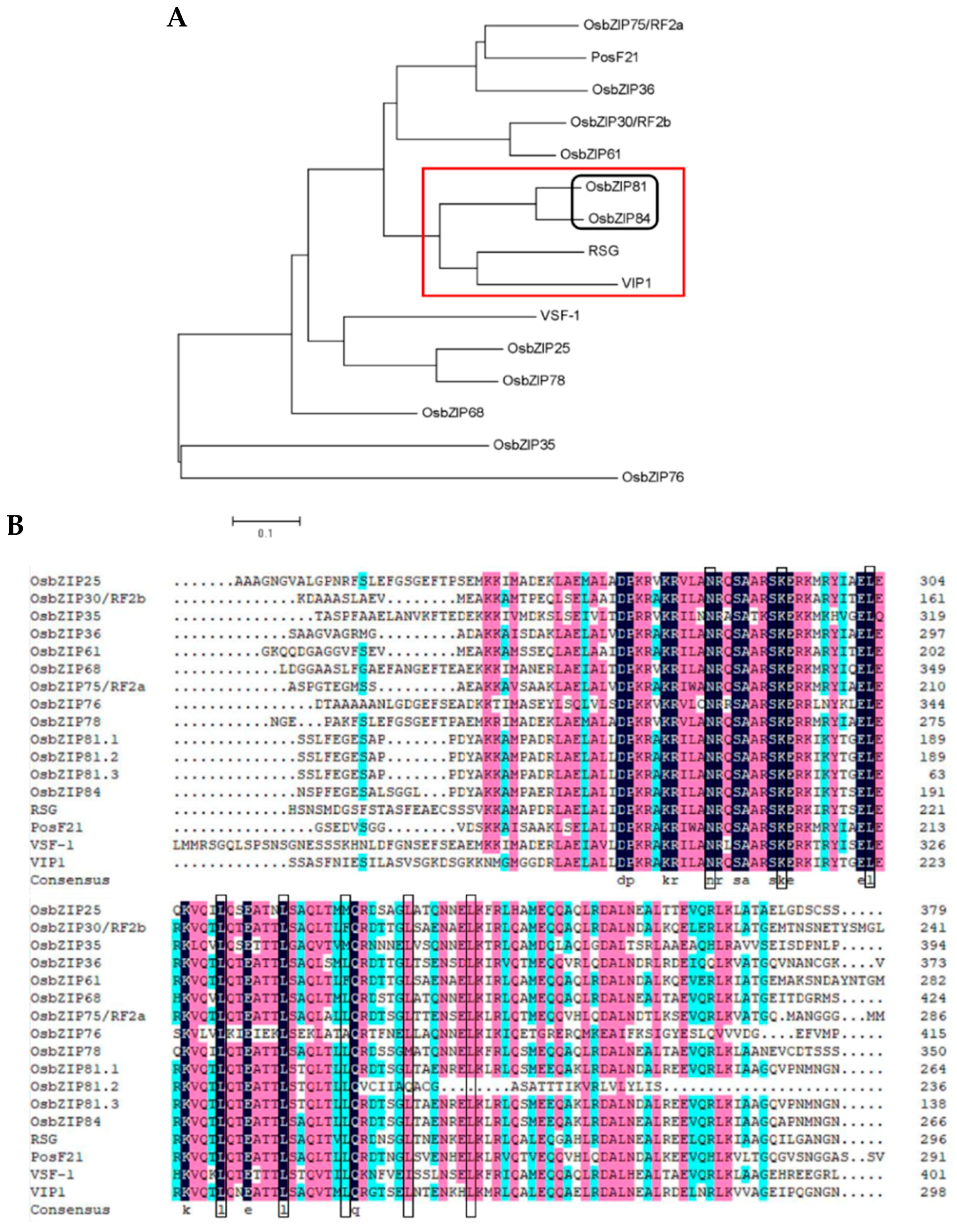
2.1. Identification of Arabidopsis VIP1 Homologue(s) in Rice
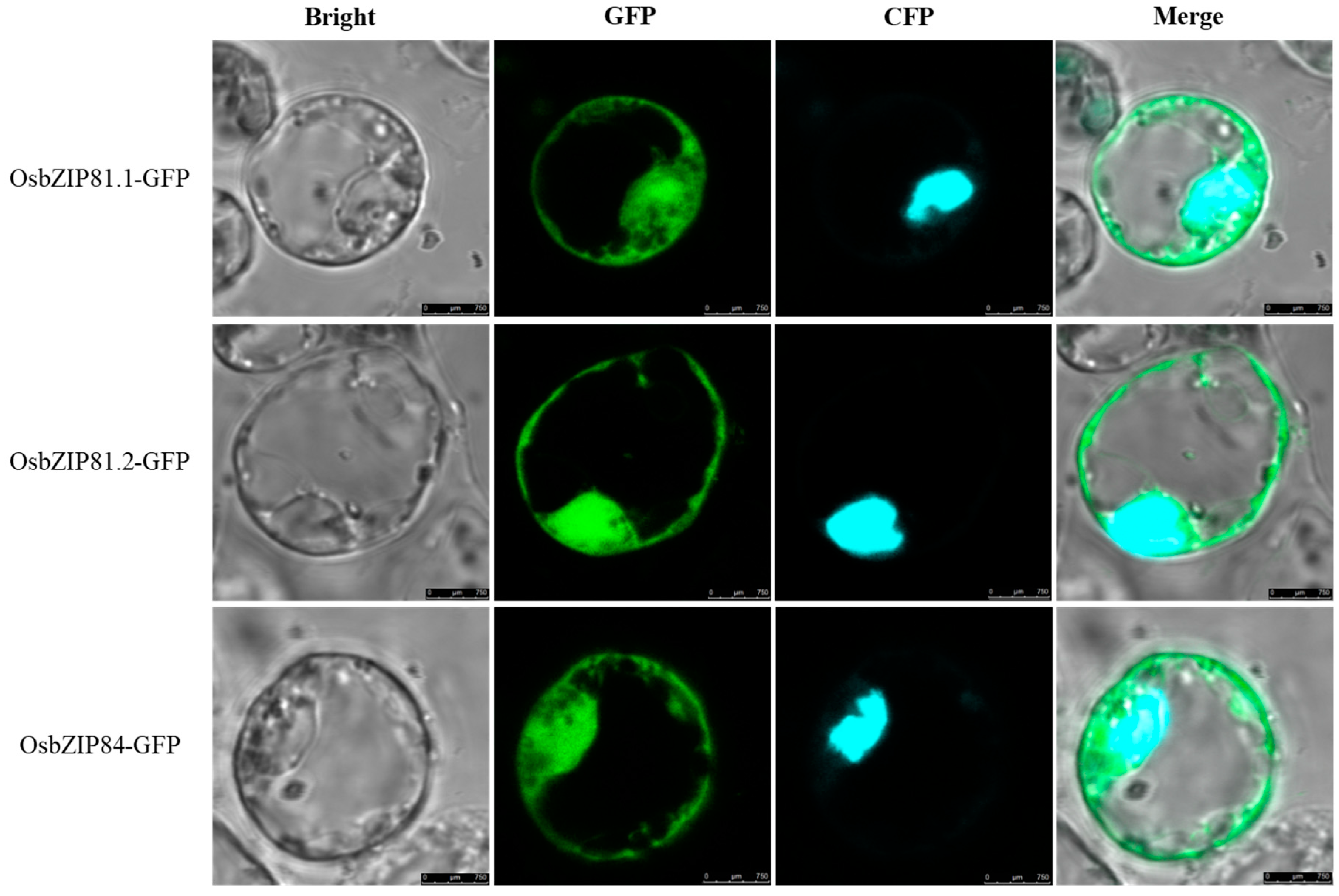
2.2. Subcellular Localization of OsbZIP81 and OsbZIP84
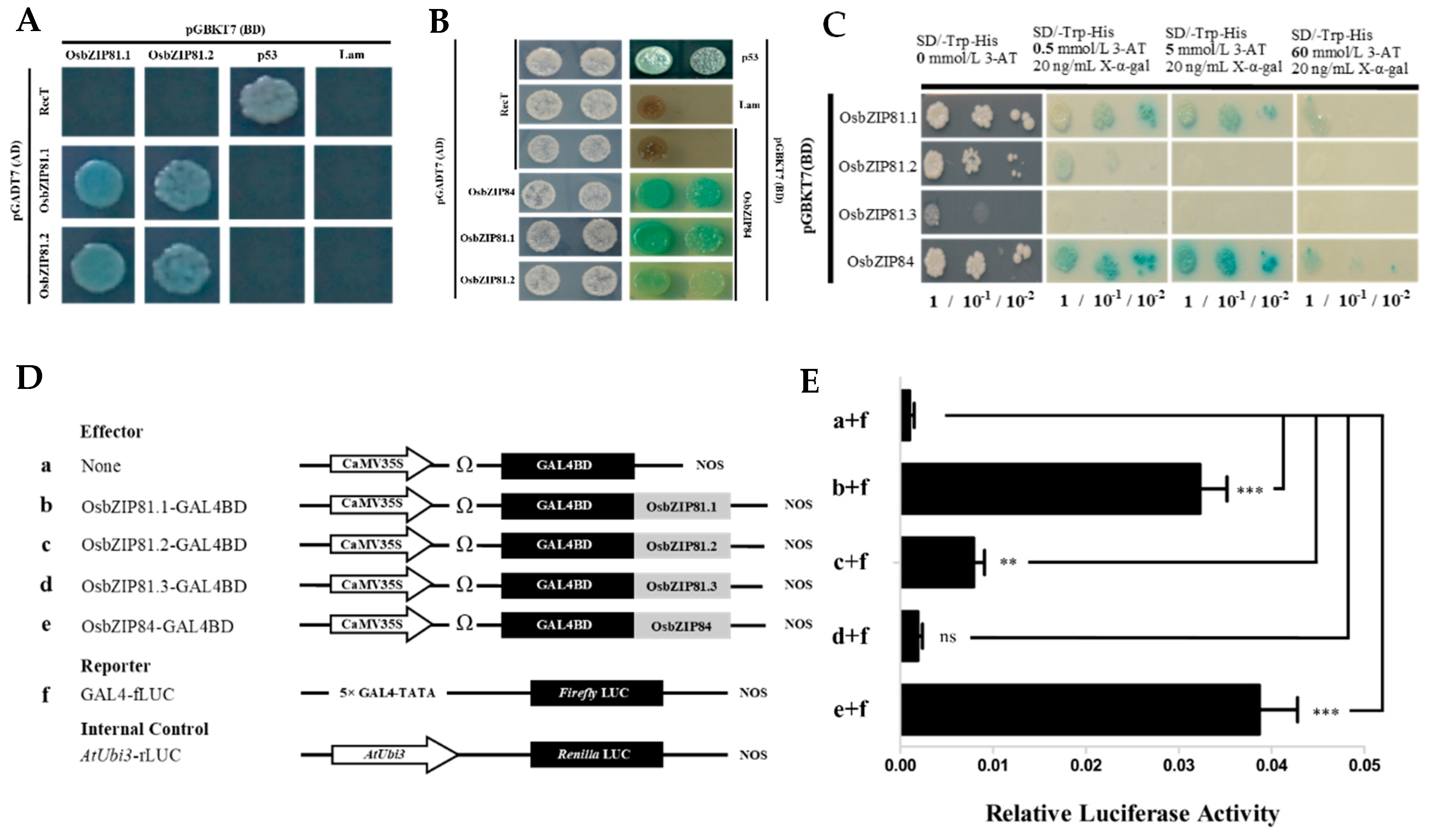
2.3. OsbZIP81.1 and OsbZIP84 Have Strong Self-Activation and Transactivation Activities
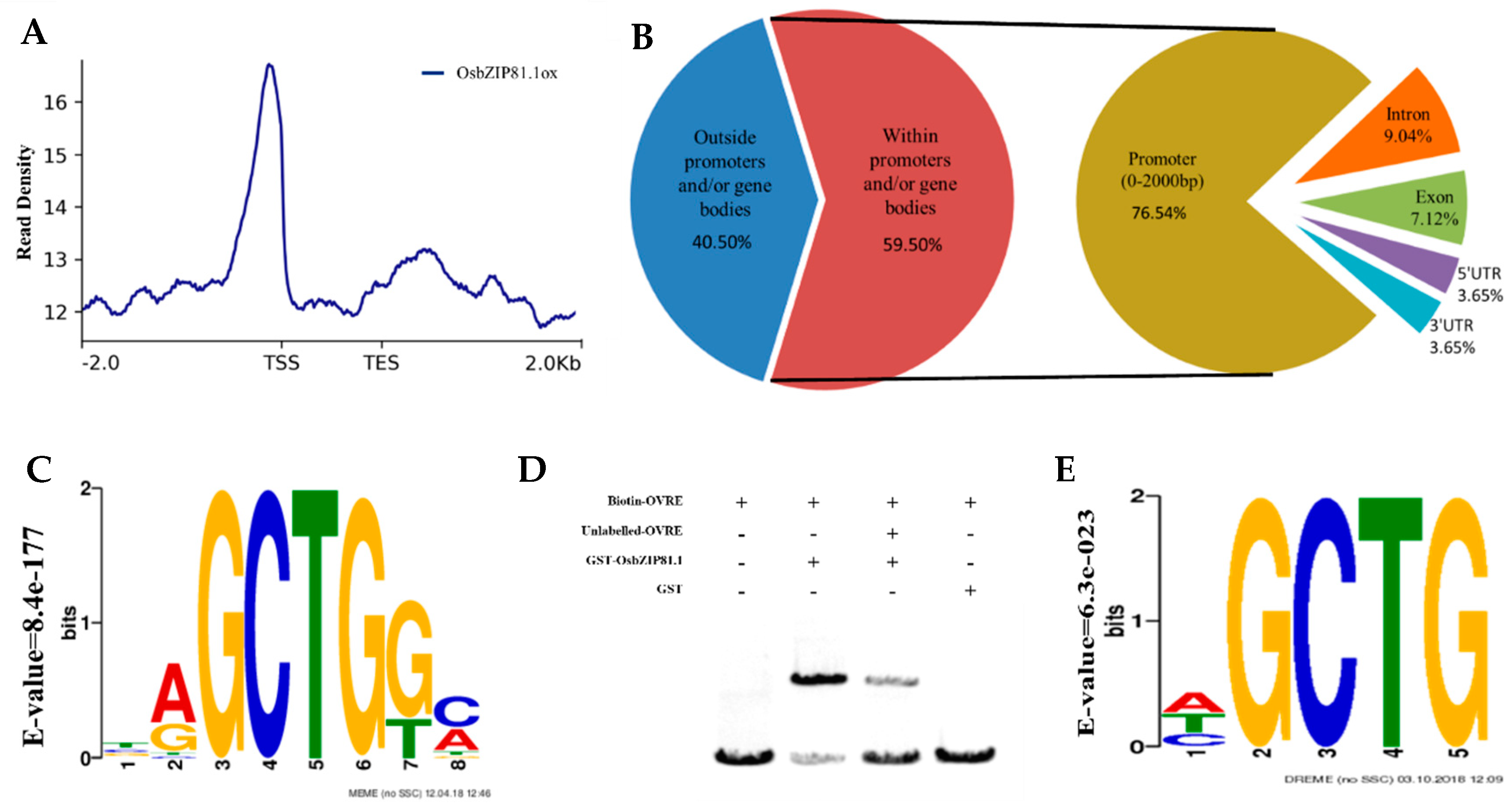
2.4. OsbZIP81.1 and OsbZIP84 Can Bind to the Motif Containing GCTG-Core Sequence
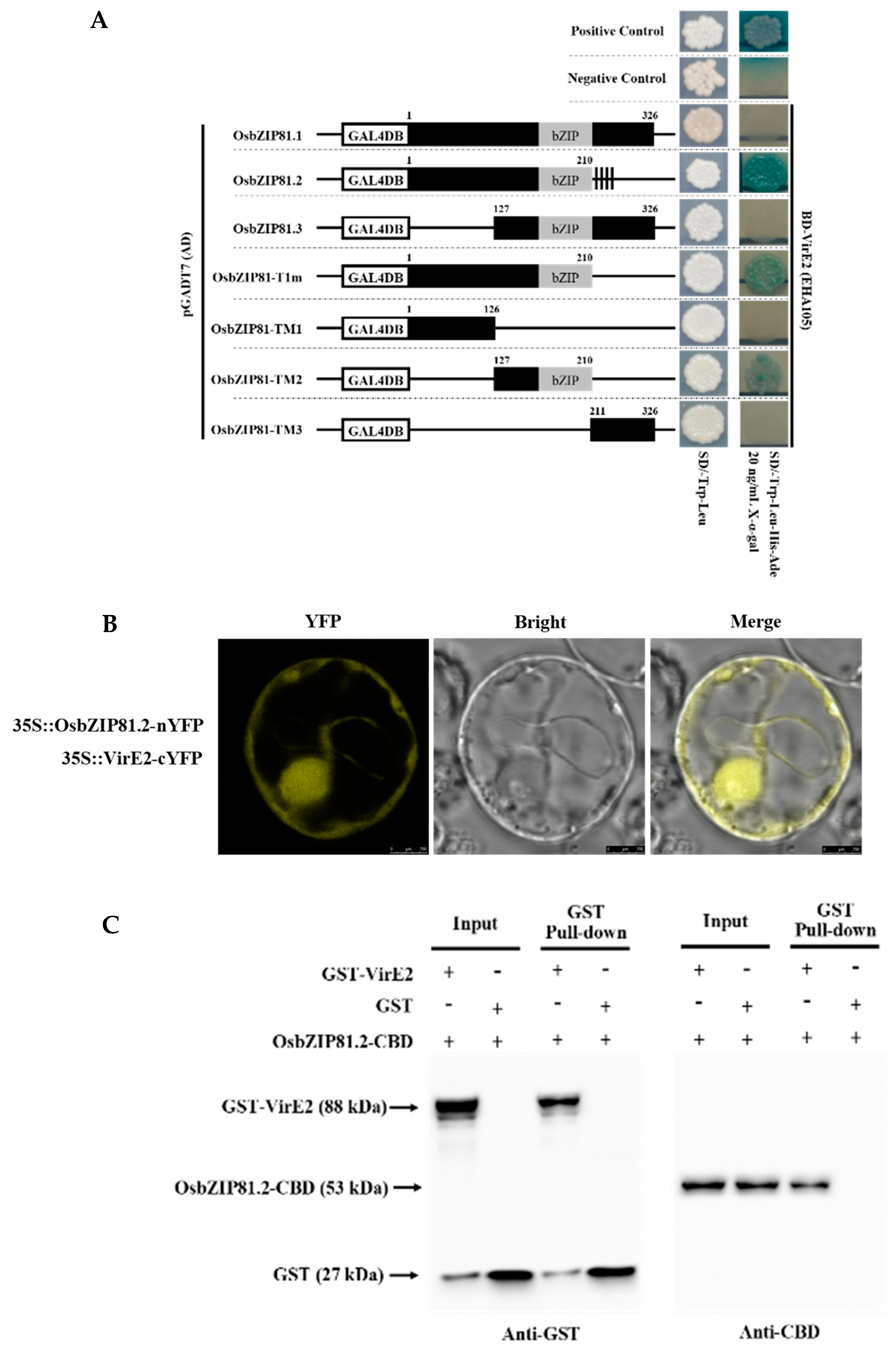
2.5. OsbZIP81.2 Can Interact with VirE2
2.6. OsbZIP81 Can Be Strongly Induced by Agrobacterium, Methyl Jasmonic Acid (MeJA), and PGE6000
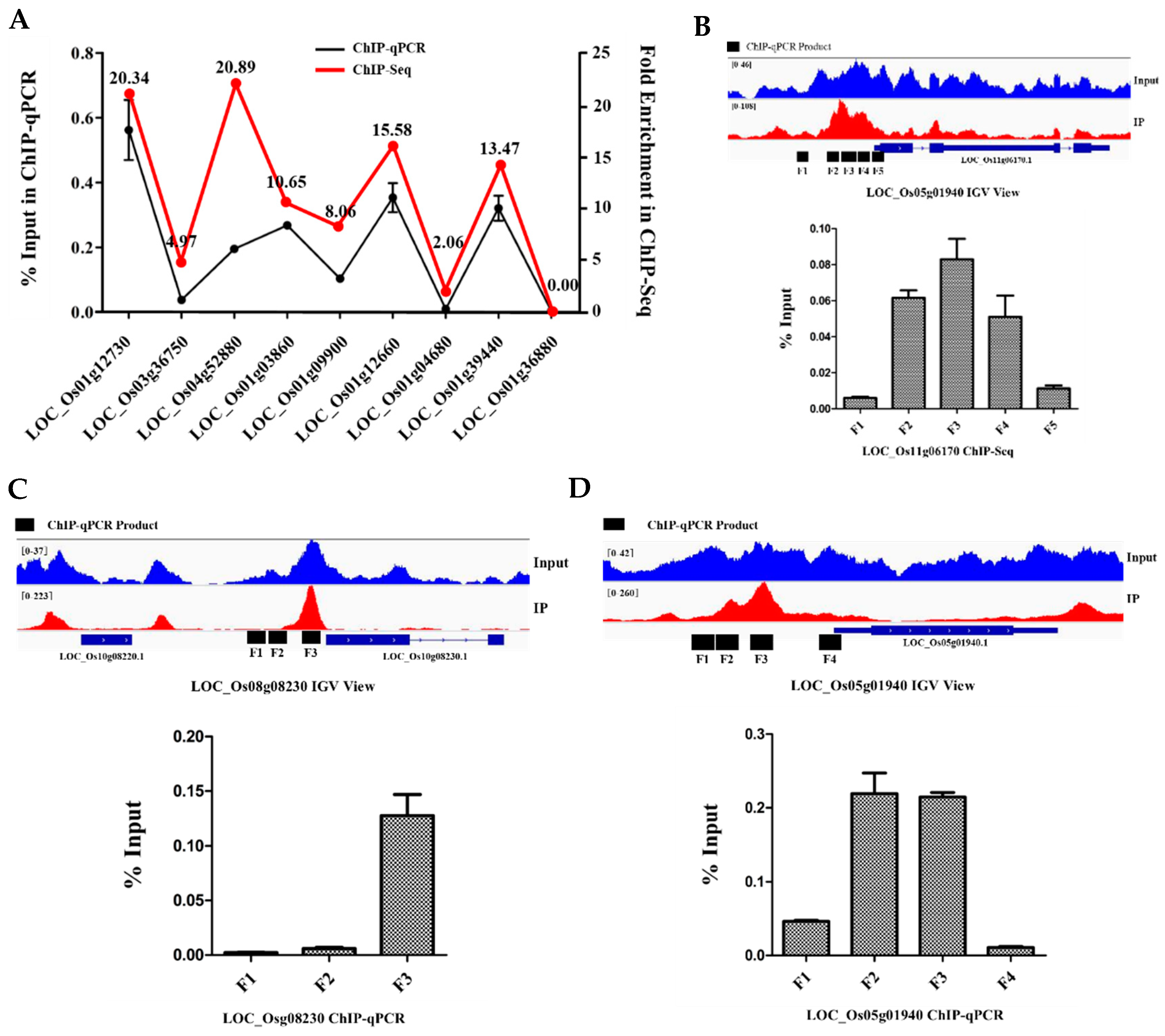
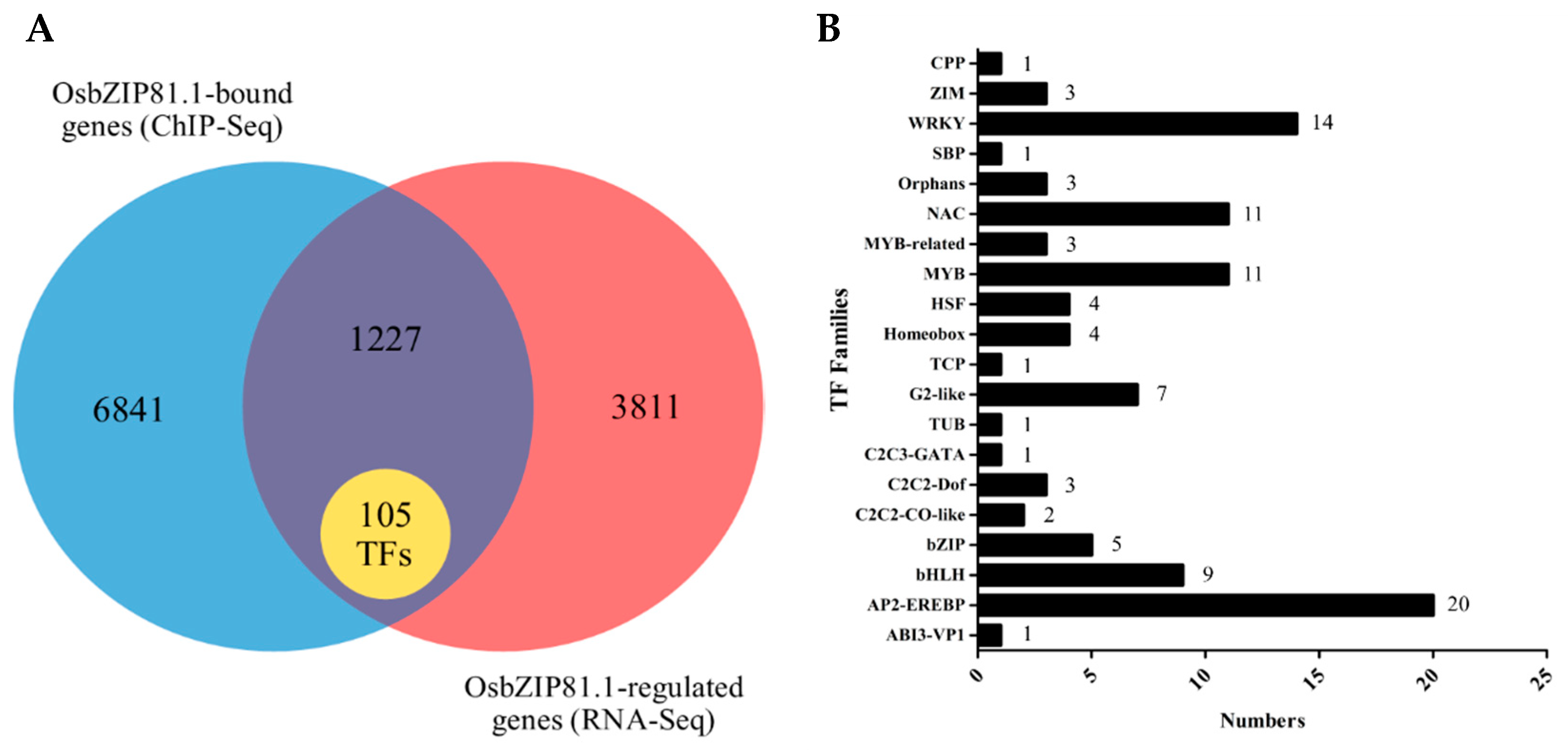
2.7. Genome-Wide Identification of the Bound and Regulated Genes of OsbZIP81
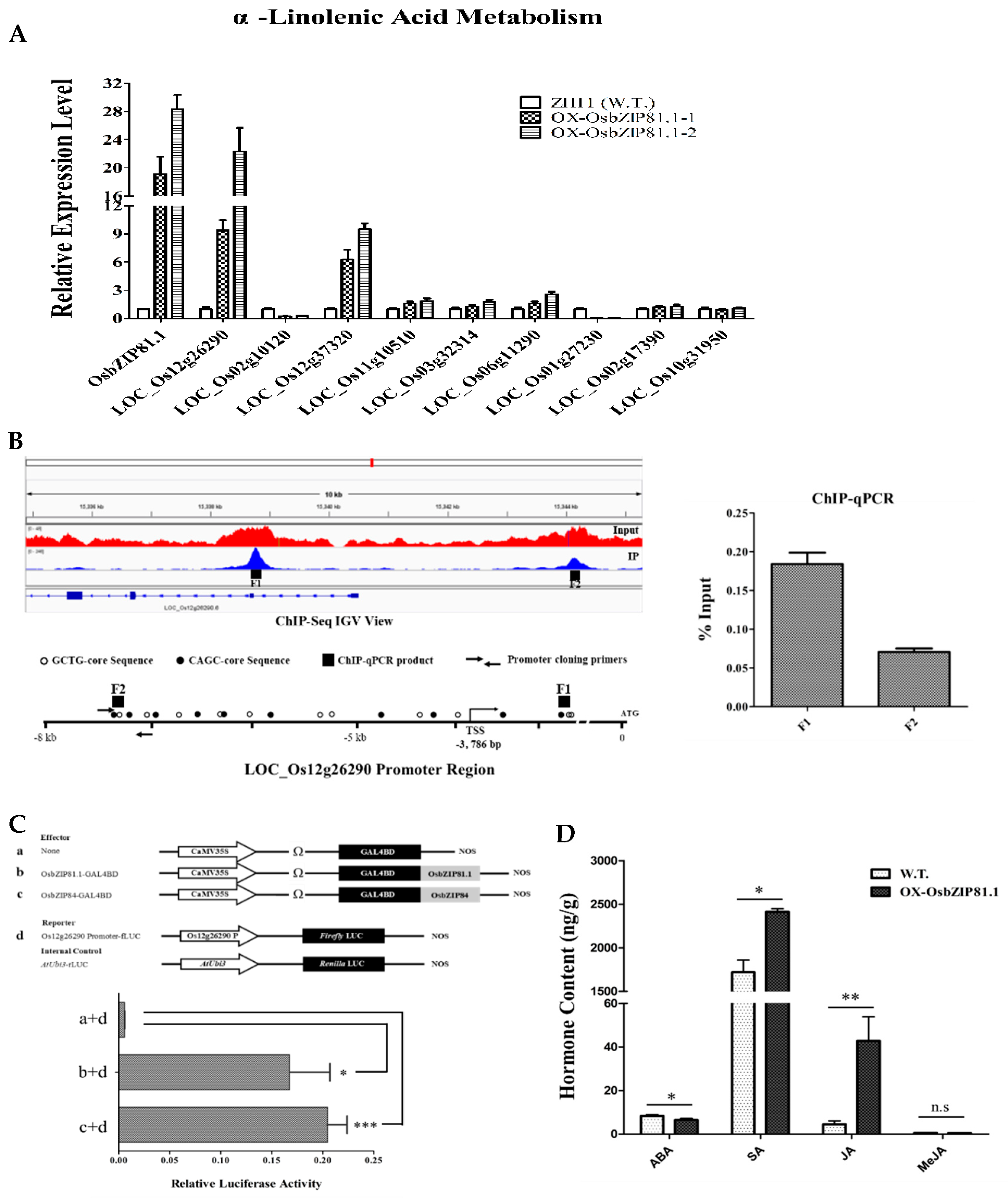
2.8. OsbZIP81.1 May Positively Affects JA levels through Directly Targetting the Genes in JA Signaling and Metabolism Pathway
2.9. OsbZIP81 can Interact with PR Proteins in Yeast
3. Discussion
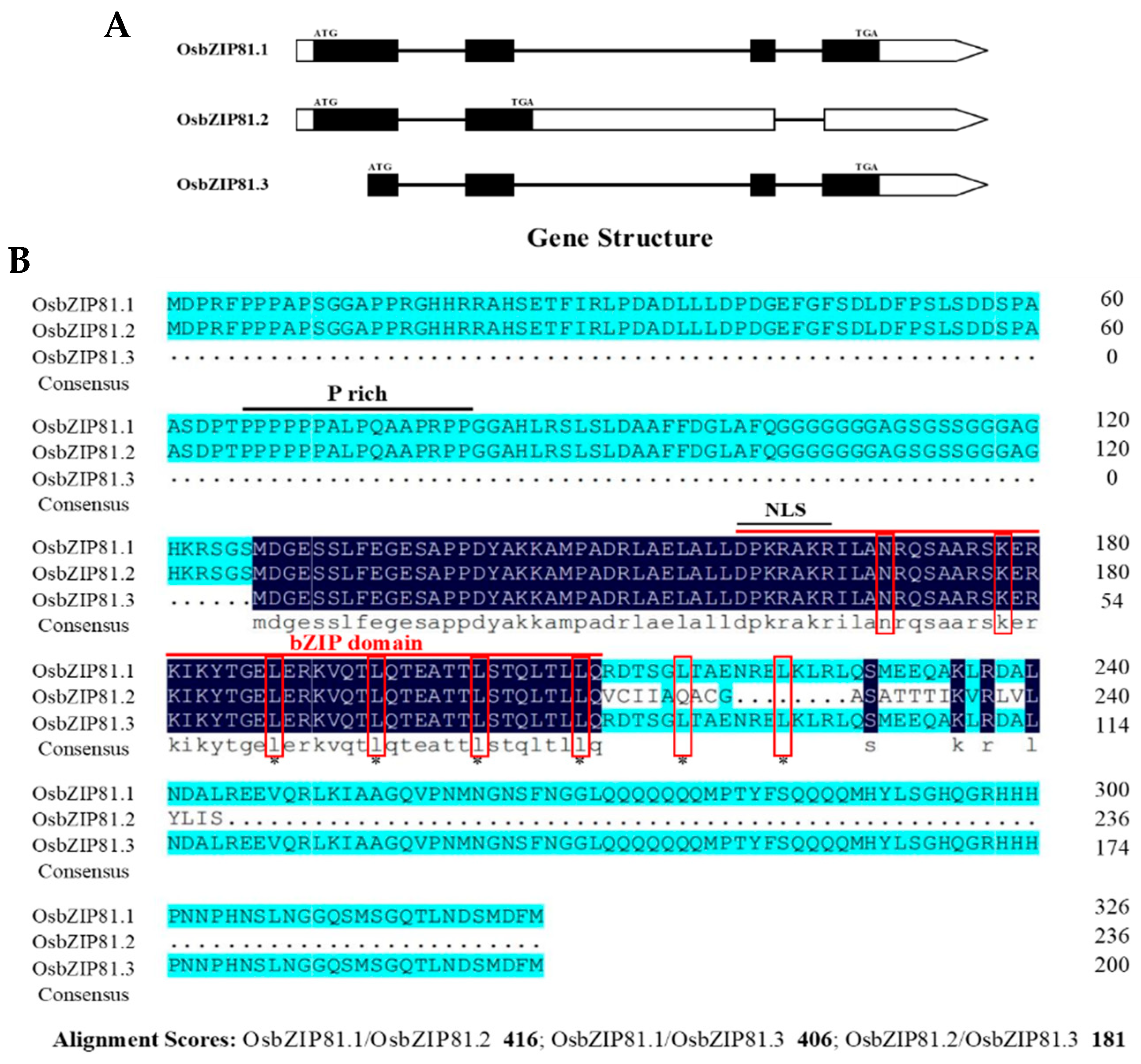
3.1. OsbZIP81 is an AS Gene
3.2. OsbZIP81.1 and OsbZIP84 are Typical bZIP Transcription Factors
3.3. OVRE is a Novel Motif for Group IX bZIPs in Rice
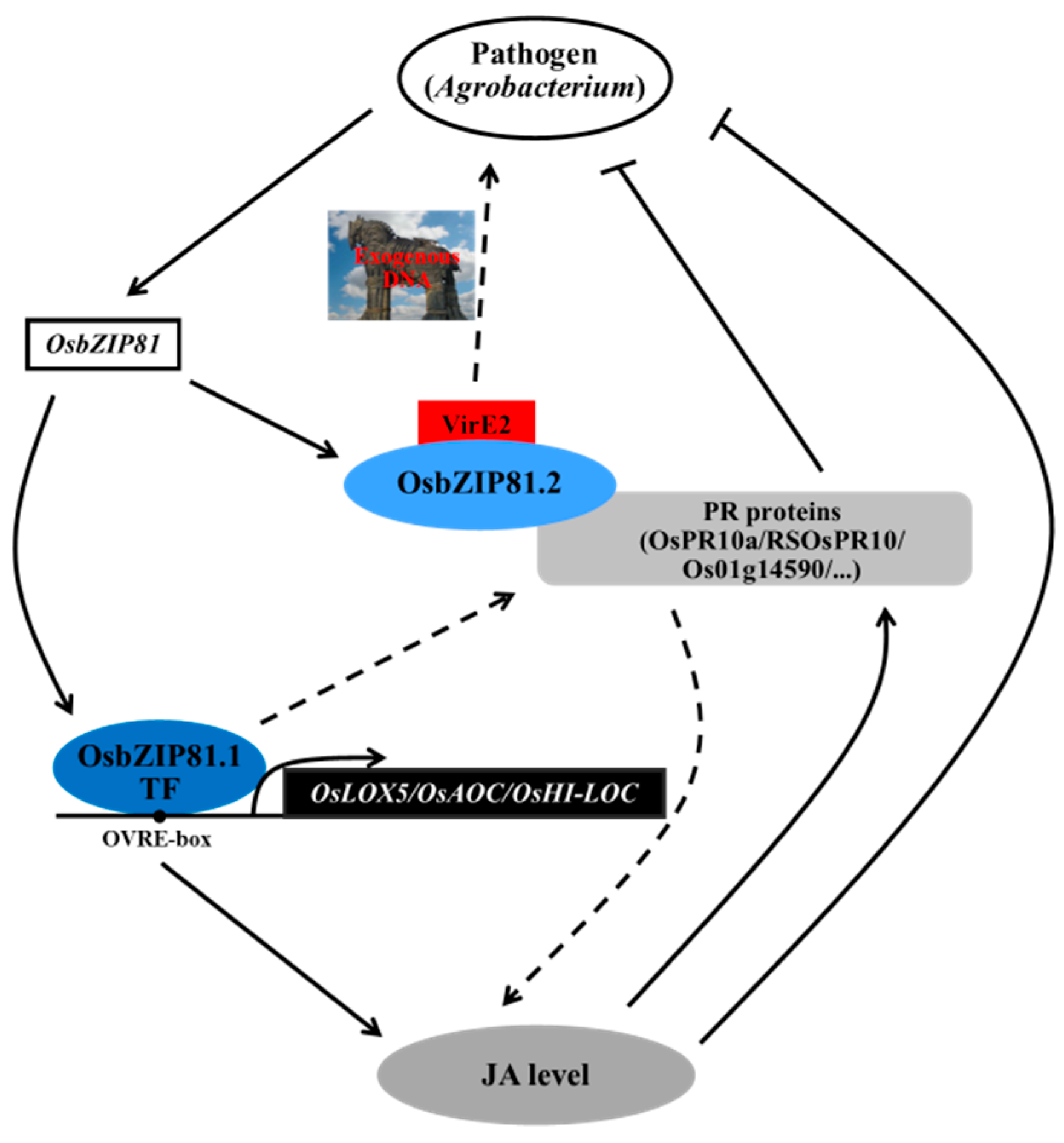
3.4. OsbZIP81 May Positively Affect Endogenous JA Levels through Directly Binding and Regulating Genes in JA Signaling and Metabolism Pathway
3.5. OsbZIP81 MAY Play Roles in Agrobacterium-Mediated Transformation and Pathogen Resistance in Rice
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Subcellular Localization Assay in Rice Protoplasts
4.3. Dual Luciferase Transcriptional Activity Assay in Rice Protoplasts
4.4. ChIP Sequencing
4.5. ChIP-qPCR
4.6. Random DNA Binding Selection Assay (RDSA)
4.7. Electrophoretic Mobility Shift Assay
4.8. Yeast Two-Hybrid and Library Screening Assay
4.9. BiFC Assay
4.10. Glutathione S-Transferase (GST) Pull-down Assay
4.11. Multiple Stress Treatment
4.12. RNA-Seq and Data Analysis
4.13. Real-Time qPCR
4.14. Quantification of Endogenous JA, MeJA, SA and ABA
4.15. Accession Numbers and Data Availability
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| OVRE | Oryza VIP1 Response Element |
| TF(s) | Transcription Factor(s) |
| ChIP-Seq | Chromatin Immunoprecipitation Sequencing |
| RNA-Seq | RNA Sequencing |
| bZIP | Basic region/leucine zipper |
| 3-AT | 3-Amino-1,2,4-triazole |
| Y2H | Yeast Two-Hybrid |
| SA | Salicylic Acid |
| ABA | Abscisic Acid |
| JA | Jasmonic Acid |
| MeJA | Methyl Jasmonic Acid |
| EMSA | Electrophoretic Mobility Shift Assay |
| TSS | Transcription Start Site |
| WT | Wild Type |
| GFP | Green Fluorescent Protein |
| CFP | Yellow Fluorescent Protein |
| ChIP-qPCR | ChIP quantitative PCR |
| DEGs | Differentially Expressed Genes |
| α-LeA | α-Linolenic Acid |
| 12-OPDA | 12-Oxophytodienoic Acid |
| PR | Pathogen-Related |
| RT-PCR | Reverse Transcription-Polymerase Chain Reaction |
| CDS | Coding Sequence |
| RDSA | Random DNA Binding Selection Assay |
| LOX | Lipoxygenase |
| AOC | Allene Oxide Cyclase |
| RT-qPCR | Real-time quantitative PCR |
References
- Liu, C.; Mao, B.; Ou, S.; Wang, W.; Liu, L.; Wu, Y.; Chu, C.; Wang, X. OsbZIP71, a bZIP transcription factor, confers salinity and drought tolerance in rice. Plant Mol. Biol. 2014, 84, 19–36. [Google Scholar] [CrossRef] [PubMed]
- Turner, J.G.; Ellis, C.; Devoto, A. The Jasmonate Signal Pathway. Plant Cell 2002, 14, s153–s164. [Google Scholar] [CrossRef] [PubMed]
- Cheong, J.-J.; Choi, Y.D. Methyl jasmonate as a vital substance in plants. Trends Genet. 2003, 19, 409–413. [Google Scholar] [CrossRef]
- Carvalhais, L.C.; Schenk, P.M.; Dennis, P.G. Jasmonic acid signalling and the plant holobiont. Curr. Opin. Microbiol. 2017, 37, 42–47. [Google Scholar] [CrossRef]
- Han, G.Z. Evolution of jasmonate biosynthesis and signaling mechanisms. J. Exp. Bot. 2017, 68, 1323–1331. [Google Scholar] [CrossRef]
- Xu, C.; Cao, H.; Zhang, Q.; Wang, H.; Xin, W.; Xu, E.; Zhang, S.; Yu, R.; Yu, D.; Hu, Y. Control of auxin-induced callus formation by bZIP59–LBD complex in Arabidopsis regeneration. Nat. Plants 2018, 4, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Alves, M.S.; Dadalto, S.P.; Gonçalves, A.B.; De Souza, G.B.; Barros, V.A.; Fietto, L.G. Plant bZIP Transcription Factors Responsive to Pathogens: A Review. Int. J. Mol. Sci. 2013, 14, 7815–7828. [Google Scholar] [CrossRef]
- Fürstenberg-Hägg, J.; Zagrobelny, M.; Bak, S. Plant Defense against Insect Herbivores. Int. J. Mol. Sci. 2013, 14, 10242–10297. [Google Scholar] [CrossRef]
- Wasternack, C.; Strnad, M. Jasmonates: News on Occurrence, Biosynthesis, Metabolism and Action of an Ancient Group of Signaling Compounds. Int. J. Mol. Sci. 2018, 19, 2539. [Google Scholar] [CrossRef]
- Wasternack, C. Action of jasmonates in plant stress responses and development—Applied aspects. Biotechnol. Adv. 2014, 32, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Chen, Y.; Shi, Z.; Jin, Y.; Sun, H.; Xie, F.; Zhang, L. Biosynthesis and signal transduction of ABA, JA, and BRs in response to drought stress of Kentucky bluegrass. Int. J. Mol. Sci. 2019, 20, 1289. [Google Scholar] [CrossRef]
- Jakoby, M.; Weisshaar, B.; Dröge-Laser, W.; Vicente-Carbajosa, J.; Tiedemann, J.; Kroj, T.; Parcy, F. bZIP transcription factors in Arabidopsis. Trends. Sci. 2002, 7, 106–111. [Google Scholar] [CrossRef]
- Dröge-Laser, W.; Snoek, B.L.; Snel, B.; Weiste, C. The Arabidopsis bZIP transcription factor family—An update. Curr. Opp. Plant Biol. 2018, 45, 36–49. [Google Scholar] [CrossRef]
- Nijhawan, A.; Jain, M.; Tyagi, A.K.; Khurana, J.P. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol. 2008, 146, 333–350. [Google Scholar] [CrossRef]
- Corrêa, L.G.G.; Riaño-Pachón, D.M.; Schrago, C.G.; Dos Santos, R.V.; Mueller-Roeber, B.; Vincentz, M. The Role of bZIP Transcription Factors in Green Plant Evolution: Adaptive Features Emerging from Four Founder Genes. PLoS ONE 2008, 3, e2944. [Google Scholar] [CrossRef]
- Wei, K.; Chen, J.; Wang, Y.; Chen, Y.; Chen, S.; Lin, Y.; Pan, S.; Zhong, X.; Xie, D. Genome-wide analysis of bZIP-encoding genes in maize. DNA Res. 2012, 19, 463–476. [Google Scholar] [CrossRef]
- Zhou, Y.; Xu, D.; Jia, L.; Huang, X.; Ma, G.; Wang, S.; Zhu, M.; Zhang, A.; Guan, M.; Lu, K.; et al. Genome-wide identification and structural analysis of bZIP transcription factor genes in brassica napus. Genes 2017, 8, 288. [Google Scholar] [CrossRef]
- Tzfira, T.; Vaidya, M.; Citovsky, V. VIP1, an Arabidopsis protein that interacts with Agrobacterium VirE2, is involved in VirE2 nuclear import and Agrobacterium infectivity. EMBO J. 2001, 20, 3596–3607. [Google Scholar] [CrossRef]
- Citovsky, V.; Kapelnikov, A.; Oliel, S.; Zakai, N.; Rojas, M.R.; Gilbertson, R.L.; Tzfira, T.; Loyter, A. Protein interactions involved in nuclear import of the agrobacterium VirE2 protein in vivo and in vitro. J. Boil. Chem. 2004, 279, 29528–29533. [Google Scholar] [CrossRef]
- Djamei, A.; Pitzschke, A.; Nakagami, H.; Rajh, I.; Hirt, H. Trojan Horse Strategy in agrobacterium transformation: Abusing MAPK defense signaling. Science 2007, 318, 453–456. [Google Scholar] [CrossRef]
- Takeo, K.; Ito, T. Subcellular localization of VIP1 is regulated by phosphorylation and 14-3-3 proteins. FEBS Lett. 2017, 591, 1972–1981. [Google Scholar] [CrossRef]
- Wu, Y.; Zhao, Q.; Gao, L.; Yu, X.-M.; Fang, P.; Oliver, D.J.; Xiang, C.-B. Isolation and characterization of low-sulphur-tolerant mutants of Arabidopsis. J. Exp. Bot. 2010, 61, 3407–3422. [Google Scholar] [CrossRef]
- Tsugama, D.; Liu, S.; Takano, T. A bZIP Protein, VIP1, is a regulator of Osmosensory signaling in Arabidopsis. Plant Physiol. 2012, 159, 144–155. [Google Scholar] [CrossRef]
- Lacroix, B.; Citovsky, V. Characterization of VIP1 activity as a transcriptional regulator in vitro and in planta. Sci. Rep. 2013, 3, 2440. [Google Scholar] [CrossRef]
- Tsugama, D.; Liu, S.; Takano, T. Metal-Binding Ability of VIP1: A bZIP protein in Arabidopsis thaliana. Protein J. 2013, 32, 526–532. [Google Scholar] [CrossRef]
- Tsugama, D.; Liu, S.; Takano, T. Analysis of functions of VIP1 and its close homologs in Osmosensory responses of Arabidopsis thaliana. PLoS ONE 2014, 9, e103930. [Google Scholar] [CrossRef]
- Tsugama, D.; Liu, S.; Takano, T. VIP1 is very important/interesting protein 1 regulating touch responses of Arabidopsis. Plant Signal. Behav. 2016, 11, e1187358. [Google Scholar] [CrossRef][Green Version]
- Tsugama, D.; Liu, S.; Takano, T. The bZIP Protein VIP1 is involved in touch responses in arabidopsis Roots. Plant Physiol. 2016, 171, 1355–1365. [Google Scholar]
- Lapham, R.; Lee, L.Y.; Tsugama, D.; Lee, S.; Mengiste, T.; Gelvin, S.B. VIP1 and Its homologs are not required for Agrobacterium-mediated transformation, but play a role in botrytis and salt stress responses. Front Plant. Sci. 2018, 9, 749. [Google Scholar] [CrossRef]
- Tsugama, D.; Liu, S.; Fujino, K.; Takano, T. Calcium signalling regulates the functions of the bZIP protein VIP1 in touch responses in Arabidopsis thaliana. Ann. Bot. 2018, 122, 1219–1229. [Google Scholar] [CrossRef]
- Pitzschke, A.; Djamei, A.; Teige, M.; Hirt, H. VIP1 response elements mediate mitogen-activated protein kinase 3-induced stress gene expression. Proc. Natl. Acad. Sci. USA 2009, 106, 18414–18419. [Google Scholar] [CrossRef]
- O’Malley, R.C.; Huang, S.-S.C.; Song, L.; Lewsey, M.G.; Bartlett, A.; Nery, J.R.; Galli, M.; Gallavotti, A.; Ecker, J.R. Cistrome and Epicistrome Features Shape the Regulatory DNA Landscape. Cell 2016, 166, 1598. [Google Scholar] [CrossRef]
- Pyo, H.; Demura, T.; Fukuda, H. Vascular cell expression patterns of Arabidopsis bZIP group I genes. Plant Biotechnol. 2006, 23, 497–501. [Google Scholar] [CrossRef]
- Wang, L.Y.; Lacroix, B.; Guo, J.H.; Citovsky, V. The Agrobacterium VirE2 effector interacts with multiple members of the Arabidopsis VIP1 protein family. Mol. Plant Pathol. 2018, 19, 1172–1183. [Google Scholar] [CrossRef]
- Van Leene, J.; Blomme, J.; Kulkarni, S.R.; Cannoot, B.; De Winne, N.; Eeckhout, D.; Persiau, G.; Van De Slijke, E.; Vercruysse, L.; Bossche, R.V.; et al. Functional characterization of the Arabidopsis transcription factor bZIP29 reveals its role in leaf and root development. J. Exp. Bot. 2016, 67, 5825–5840. [Google Scholar] [CrossRef]
- Gibalová, A.; Steinbachová, L.; Hafidh, S.; Bláhová, V.; Gadiou, Z.; Michailidis, C.; Műller, K.; Pleskot, R.; Dupľáková, N.; Honys, D. Characterization of pollen-expressed bZIP protein interactions and the role of ATbZIP18 in the male gametophyte. Plant Reprod. 2017, 30, 1–17. [Google Scholar] [CrossRef]
- Yin, Y.; Zhu, Q.; Dai, S.; Lamb, C.; Beachy, R.N. RF2a, a bZIP transcriptional activator of the phloem-specific rice tungro bacilliform virus promoter, functions in vascular development. EMBO J. 1997, 16, 5247–5259. [Google Scholar] [CrossRef]
- Dai, S.; Zhang, Z.; Chen, S.; Beachy, R.N. RF2b, a rice bZIP transcription activator, interacts with RF2a and is involved in symptom development of rice tungro disease. Proc. Natl. Acad. Sci. USA 2004, 101, 687–692. [Google Scholar] [CrossRef]
- Dai, S.; Petruccelli, S.; Ordiz, M.I.; Zhang, Z.; Chen, S.; Beachy, R.N. Functional analysis of RF2a, a rice transcription factor. J. Boil. Chem. 2003, 278, 36396–36402. [Google Scholar] [CrossRef]
- Dai, S.; Wei, X.; Alfonso, A.A.; Pei, L.; Duque, U.G.; Zhang, Z.; Babb, G.M.; Beachy, R.N. Transgenic rice plants that overexpress transcription factors RF2a and RF2b are tolerant to rice tungro virus replication and disease. Proc. Natl. Acad. Sci. USA 2008, 105, 21012–21016. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, S.; Huang, F.; Zhou, X.; Chen, Z.; Peng, W.; Luo, M. VirD5 is required for efficient Agrobacterium infection and interacts with Arabidopsis VIP2. New Phytol. 2018, 217, 726–738. [Google Scholar] [CrossRef]
- Wang, Y.; Peng, W.; Zhou, X.; Huang, F.; Shao, L.; Luo, M. The putativeAgrobacteriumtranscriptional activator-like virulence protein VirD5 may target T-complex to prevent the degradation of coat proteins in the plant cell nucleus. New Phytol. 2014, 203, 1266–1281. [Google Scholar] [CrossRef]
- Wang, Y.; Zeng, H.; Zhou, X.; Huang, F.; Peng, W.; Liu, L.; Xiong, W.; Shi, X.; Luo, M. Transformation of rice with large maize genomic DNA fragments containing high content repetitive sequences. Plant Cell Rep. 2015, 34, 1049–1061. [Google Scholar] [CrossRef]
- Aeschbacher, R.; Schrott, M.; Potrykus, I.; Saul, M. Isolation and molecular characterization of PosF21, an Arabidopsis thaliana gene which shows characteristics of a b-Zip class transcription factor. Plant J. 1991, 1, 303–316. [Google Scholar] [CrossRef]
- Fukazawa, J. Repression of Shoot Growth, a bZIP Transcriptional Activator, Regulates Cell Elongation by Controlling the Level of Gibberellins. Plant Cell 2000, 12, 901–915. [Google Scholar] [CrossRef]
- Torres-Schumann, S.; Ringli, C.; Heierli, D.; Amrhein, N.; Keller, B.; Torres-Schumann, S. In vitro binding of the tomato bZIP transcriptional activator VSF-1 to a regulatory element that controls xylem-specific gene expression. Plant J. 1996, 9, 283–296. [Google Scholar] [CrossRef]
- Chou, K.-C.; Shen, H.-B. Cell-PLoc: A package of Web servers for predicting subcellular localization of proteins in various organisms. Nat. Protoc. 2008, 3, 153–162. [Google Scholar] [CrossRef]
- Xue, W.; Xing, Y.; Weng, X.; Zhao, Y.; Tang, W.; Wang, L.; Zhou, H.; Yu, S.; Xu, C.; Li, X.; et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 2008, 40, 761–767. [Google Scholar] [CrossRef]
- Machanick, P.; Bailey, T.L. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 2011, 27, 1696–1697. [Google Scholar] [CrossRef]
- Kawahara, Y.; de la Bastide, M.; Hamilton, J.P.; Kanamori, H.; McCombie, W.R.; Ouyang, S.; Schwartz, D.C.; Tanaka, T.; Wu, J.; Zhou, S.; et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 2013, 6, 4. [Google Scholar] [CrossRef]
- Yan, H.; Tian, T.; Liu, Y.; You, Q.; Yi, X.; Du, Z.; Xu, W.; Su, Z. agriGO v2.0: A GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 2017, 45, W122–W129. [Google Scholar]
- Zou, M.; Guan, Y.; Ren, H.; Zhang, F.; Chen, F. Characterization of alternative splicing products of bZIP transcription factors OsABI5. Biochem. Biophys. Commun. 2007, 360, 307–313. [Google Scholar] [CrossRef]
- Kelemen, O.; Convertini, P.; Zhang, Z.; Wen, Y.; Shen, M.; Falaleeva, M.; Stamm, S. Function of alternative splicing. Gene 2013, 514, 1–30. [Google Scholar] [CrossRef]
- Dong, C.; He, F.; Berkowitz, O.; Liu, J.; Cao, P.; Tang, M.; Shi, S.; Wang, W.; Li, Q.; Whelan, J.; et al. Alternative splicing plays a critical role in maintaining mineral nutrient homeostasis in rice (Oryza sativa). Plant Cell. 2018, 30, 2267–2285. [Google Scholar] [CrossRef]
- Szakonyi, D.; Duque, P. Alternative splicing as a regulator of early plant development. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Zhang, G.; Sun, M.; Wang, J.; Lei, M.; Li, C.; Zhao, D.; Huang, J.; Li, W.; Li, S.; Li, J.; et al. PacBio full-length cDNA sequencing integrated with RNA-seq reads drastically improves the discovery of splicing transcripts in rice. Plant J. 2018, 97, 296–305. [Google Scholar] [CrossRef]
- Pruneda-Paz, J.L.; Breton, G.; Nagel, D.H.; Kang, S.E.; Bonaldi, K.; Doherty, C.J.; Ravelo, S.; Galli, M.; Ecker, J.R.; Kay, S.A. A genome-scale resource for the functional characterization of Arabidopsis transcription factors. Cell Rep. 2014, 8, 622–632. [Google Scholar] [CrossRef]
- Ringli, C.; Keller, B. Specific interaction of the tomato bZIP transcription factor VSF-1 with a non-palindromic DNA sequence that controls vascular gene expression. Plant. Mol. Biol. 1998, 37, 977–988. [Google Scholar] [CrossRef]
- Pieterse, C.M.J.; Leon-Reyes, A.; Van Der Ent, S.; Van Wees, S.C.M. Networking by small-molecule hormones in plant immunity. Nat. Methods 2009, 5, 308–316. [Google Scholar] [CrossRef]
- Tsuda, K.; Sato, M.; Stoddard, T.; Glazebrook, J.; Katagiri, F. Network Properties of Robust Immunity in Plants. PLoS Genet. 2009, 5, e1000772. [Google Scholar] [CrossRef]
- Pieterse, C.M.; Van Der Does, D.; Zamioudis, C.; Leon-Reyes, A.; Van Wees, S.C. Hormonal modulation of plant immunity. Annu. Cell Dev. Boil. 2012, 28, 489–521. [Google Scholar] [CrossRef] [PubMed]
- Ballaré, C.L. Jasmonate-induced defenses: a tale of intelligence, collaborators and rascals. Trends Plant Sci. 2011, 16, 249–257. [Google Scholar] [CrossRef]
- Broekgaarden, C.; Caarls, L.; Vos, I.A.; Pieterse, C.M.; Van Wees, S.C. Ethylene: Traffic controller on hormonal crossroads to Defense1. Plant Physiol. 2015, 169, 2371–2379. [Google Scholar] [CrossRef] [PubMed]
- Chini, A.; Cimmino, A.; Masi, M.; Reveglia, P.; Nocera, P.; Solano, R.; Evidente, A. The fungal phytotoxin lasiojasmonate A activates the plant jasmonic acid pathway. J. Exp. Bot. 2018, 69, 3095–3102. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Ding, C.; Baerson, S.R.; Lian, F.; Lin, X.; Zhang, L.; Wu, C.; Hwang, S.-Y.; Zeng, R.; Song, Y. The roles of jasmonate signalling in nitrogen uptake and allocation in rice (Oryza sativa L.). Plant Cell 2018, 42, 659–672. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.W.; Lim, S.; Baek, W.; Han, S.-W.; Lee, S.C. Expression and functional roles of the pepper pathogen-induced bZIP transcription factor, CabZIP2, in Enhanced disease resistance to bacterial pathogen infection. Mol. Plant Microbe Interact. 2015, 28, 825–833. [Google Scholar] [CrossRef] [PubMed]
- Moon, S.-J.; Park, H.J.; Kim, T.-H.; Kang, J.-W.; Lee, J.-Y.; Cho, J.-H.; Lee, J.-H.; Park, D.-S.; Byun, M.-O.; Kim, B.-G.; et al. OsTGA2 confers disease resistance to rice against leaf blight by regulating expression levels of disease related genes via interaction with NH1. PLoS ONE 2018, 13, e0206910. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.L.; Kohchi, T.; Yamato, K.T.; Jenkins, J.; Shu, S.; Ishizaki, K.; Yamaoka, S.; Nishihama, R.; Nakamura, Y.; Berger, F.; et al. Insights into land plant evolution garnered from the marchantia polymorpha genome. Cell 2017, 171, 287.e15–304.e15. [Google Scholar] [CrossRef]
- Nishiyama, T.; Sakayama, H.; De Vries, J.; Buschmann, H.; Saint-Marcoux, D.; Ullrich, K.K.; Haas, F.B.; Vanderstraeten, L.; Becker, D.; Lang, D.; et al. The Chara genome: Secondary complexity and implications for plant terrestrialization. Cell 2018, 174, 448.e24–464.e24. [Google Scholar] [CrossRef]
- Kombrink, E. Chemical and genetic exploration of jasmonate biosynthesis and signaling paths. Planta 2012, 236, 1351–1366. [Google Scholar] [CrossRef]
- Wasternack, C.; Hause, B. Jasmonates: Biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann. Bot. 2013, 111, 1021–1058. [Google Scholar] [CrossRef]
- Zhang, X.; Bao, Y.; Shan, D.; Wang, Z.; Song, X.; Wang, Z.; Wang, J.; He, L.; Wu, L.; Zhang, Z.; et al. Magnaporthe oryzae induces the expression of a MicroRNA to suppress the immune response in Rice. Plant Physiol. 2018, 177, 352–368. [Google Scholar] [PubMed]
- Zhou, G.; Qi, J.; Ren, N.; Cheng, J.; Erb, M.; Mao, B.; Lou, Y. SilencingOsHI-LOXmakes rice more susceptible to chewing herbivores, but enhances resistance to a phloem feeder. Plant J. 2009, 60, 638–648. [Google Scholar] [CrossRef]
- Qi, J.; Zhou, G.; Yang, L.; Erb, M.; Lu, Y.; Sun, X.; Cheng, J.; Lou, Y. The Chloroplast-localized phospholipases D α4 and α5 regulate herbivore-induced direct and indirect defenses in Rice. Plant Physiol. 2011, 157, 1987–1999. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Afsheen, S.; Xin, Z.J.; Han, X.; Lou, Y.G. OsNPR1 negatively regulates herbivore-induced JA and ethylene signaling and plant resistance to a chewing herbivore in rice. Physiol Plant. 2013, 147, 340–351. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, J.; Hu, L.; Zhang, T.; Zhang, G.; Lou, Y. OsMPK3 positively regulates the JA signaling pathway and plant resistance to a chewing herbivore in rice. Plant Cell Rep. 2013, 32, 1075–1084. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, G. Cloning of novel rice allene oxide cyclase (OsAOC): mRNA expression and comparative analysis with allene oxide synthase (OsAOS) gene provides insight into the transcriptional regulation of octadecanoid pathway biosynthetic genes in rice. Plant Sci. 2003, 164, 979–992. [Google Scholar] [CrossRef]
- Riemann, M.; Haga, K.; Shimizu, T.; Okada, K.; Ando, S.; Mochizuki, S.; Nishizawa, Y.; Yamanouchi, U.; Nick, P.; Yano, M.; et al. Identification of rice Allene Oxide Cyclase mutants and the function of jasmonate for defence against Magnaporthe oryzae. Plant. J. 2013, 74, 226–238. [Google Scholar] [CrossRef]
- Liu, L.; Zou, Z.; Qian, K.; Xia, C.; He, Y.; Zeng, H.; Zhou, X.; Riemann, M.; Yin, C. Jasmonic acid deficiency leads to scattered floret opening time in cytoplasmic male sterile rice Zhenshan 97A. J. Exp. Bot. 2017, 68, 4613–4625. [Google Scholar] [CrossRef]
- Koeduka, T.; Matsui, K.; Akakabe, Y.; Kajiwara, T. Catalytic properties of rice alpha-oxygenase - A comparison with mammalian prostaglandin H synthases. J. Biol. Chem. 2002, 277, 22648–22655. [Google Scholar] [CrossRef] [PubMed]
- Hamberg, M.; Sanz, A.; Castresana, C. alpha-oxidation of fatty acids in higher plants. Identification of a pathogen-inducible oxygenase (piox) as an alpha-dioxygenase and biosynthesis of 2-hydroperoxylinolenic acid. J. Boil. Chem. 1999, 274, 24503–24513. [Google Scholar] [CrossRef] [PubMed]
- Pitzschke, A.; Hirt, H. New insights into an old story: Agrobacterium-induced tumour formation in plants by plant transformation. EMBO J. 2010, 29, 1021–1032. [Google Scholar] [CrossRef] [PubMed]
- Winans, S.C.; Christie, P.J.; E Ward, J.; Nester, E.W. The Agrobacterium tumefaciens virE2 gene product is a single-stranded-DNA-binding protein that associates with T-DNA. J. Bacteriol. 1988, 170, 2659–2667. [Google Scholar]
- Grange, W.; Duckely, M.; Husale, S.; Jacob, S.; Engel, A.; Hegner, M. VirE2: A Unique ssDNA-Compacting Molecular Machine. PLoS Boil. 2008, 6, e44. [Google Scholar] [CrossRef] [PubMed]
- Loyter, A.; Rosenbluh, J.; Zakai, N.; Li, J.; Kozlovsky, S.V.; Tzfira, T.; Citovsky, V. The Plant VirE2 Interacting Protein 1. A Molecular link between the agrobacterium T-Complex and the host cell chromatin?1. Plant Physiol. 2005, 138, 1318–1321. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, A.; Aichi, I.; Matsuoka, M. A protocol for Agrobacterium-mediated transformation in rice. Nat. Protoc. 2006, 1, 2796–2802. [Google Scholar] [CrossRef]
- Ozawa, K. Establishment of a high efficiency Agrobacterium-mediated transformation system of rice (Oryza sativa L.). Plant Sci. 2009, 176, 522–527. [Google Scholar] [CrossRef]
- Ozawa, K. A High-efficiency agrobacterium-mediated transformation system of Rice (Oryza sativa L.). Methods Mol. Biol. 2012, 847, 51–57. [Google Scholar]
- Wu, C.; Sui, Y. Efficient and fast production of transgenic rice plants by agrobacterium-mediated transformation. Methods Mol. Biol. 2019, 1864, 95–103. [Google Scholar] [CrossRef]
- Takeuchi, K.; Gyohda, A.; Tominaga, M.; Kawakatsu, M.; Hatakeyama, A.; Ishii, N.; Shimaya, K.; Nishimura, T.; Riemann, M.; Nick, P.; et al. RSOsPR10 expression in response to environmental stresses is regulated antagonistically by Jasmonate/Ethylene and salicylic acid signaling pathways in rice roots. Plant Cell Physiol. 2011, 52, 1686–1696. [Google Scholar] [CrossRef]
- Van Loon, L.; Rep, M.; Pieterse, C. Significance of inducible defense-related proteins in infected plants. Annu. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef]
- Kumashiro, T.; Hiei, Y.; Ohta, S.; Komari, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 1994, 6, 271–282. [Google Scholar]
- Xie, K.; Yang, Y. RNA-Guided Genome Editing in Plants Using a CRISPR-Cas System. Mol. Plant 2013, 6, 1975–1983. [Google Scholar] [CrossRef]
- Zong, W.; Tang, N.; Yang, J.; Peng, L.; Ma, S.; Xu, Y.; Li, G.; Xiong, L. Feedback regulation of ABA signaling and biosynthesis by a bZIP transcription factor targets drought-resistance-related Genes. Plant Physiol. 2016, 171, 2810–2825. [Google Scholar]
- Zong, W.; Zhong, X.; You, J.; Xiong, L. Genome-wide profiling of histone H3K4-tri-methylation and gene expression in rice under drought stress. Plant Mol. Biol. 2013, 81, 175–188. [Google Scholar] [CrossRef]
- Li, R.; Yu, C.; Li, Y.; Lam, T.-W.; Yiu, S.-M.; Kristiansen, K.; Wang, J. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 2009, 25, 1966–1967. [Google Scholar] [CrossRef]
- Liu, T. Use Model-Based Analysis of ChIP-Seq (MACS) to Analyze Short Reads Generated by Sequencing Protein–DNA Interactions in Embryonic Stem Cells. Methods Mol. Biol. 2014, 1150, 81–95. [Google Scholar]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative genomics viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef]
- Ma, W.; Noble, W.S.; Bailey, T.L. Motif-based analysis of large nucleotide data sets using MEME-ChIP. Nat. Protoc. 2014, 9, 1428–1450. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S.; Sato, Y.; Furumichi, M.; Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 2012, 40, D109–D114. [Google Scholar] [CrossRef]
- Thimm, O.; Nagel, A.; Krüger, P.; Selbig, J.; Müller, L.A.; Bläsing, O.; Gibon, Y.; Meyer, S.; Rhee, S.Y.; Stitt, M. mapman: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004, 37, 914–939. [Google Scholar] [CrossRef] [PubMed]
- Waadt, R.; Schmidt, L.K.; Lohse, M.; Hashimoto, K.; Bock, R.; Kudla, J.; Kudla, J. Multicolor bimolecular fluorescence complementation reveals simultaneous formation of alternative CBL/CIPK complexesin planta. Plant J. 2008, 56, 505–516. [Google Scholar] [CrossRef] [PubMed]

| Sample | Total Reads | Mapped Reads | Paired | Single | SelfAND Mate | MapDiff CHR | Quality |
|---|---|---|---|---|---|---|---|
| OsbZIP81.1ox_IP | 44705515 | 43192879 (96.62%) | 21604458 | 373400 | 35769480 | 12784780 | 8179875 |
| OsbZIP81.1ox_Input | 35488065 | 34501154 (97.22%) | 33180942 | 106591 | 34091530 | 775770 | 486020 |
| OsbZIP81.2ox_IP | 48091675 | 45489002 (94.59%) | 43727660 | 146629 | 44858010 | 989478 | 622885 |
| OsbZIP81.2ox_Input | 39256499 | 38410324 (97.84%) | 36286088 | 155149 | 37874736 | 1401852 | 936240 |
| Gene ID | Nearest TSS | Putative Function (Reported Gene Name) | Fold Enrichment |
|---|---|---|---|
| LOC_Os05g08370 | Chr5:4565177-4571461:-:-71 | CESA1 - cellulose synthase | 23.0974 |
| LOC_Os04g56530 | Chr4:33707427-33719236:-:-96 | STE kinases include homologs to sterile 7, sterile 11 and sterile 20 from yeast | 17.6621 |
| LOC_Os08g32170 | Chr8:19954661-19956231:-:-83 | oxidoreductase, 2OG-FeII oxygenase domain containing protein | 16.4529 |
| LOC_Os04g54240 | Chr4:32301622-32302456:-:-204 | wound induced protein | 16.4453 |
| LOC_Os02g26160 | Chr2:15363265-15367217:+:179 | receptor lectin kinase like protein | 16.1431 |
| LOC_Os03g62700 | Chr3:35480584-35486347:-:-161 | protein kinase | 15.8304 |
| LOC_Os06g12660 | Chr6:6915783-6916582:+:460 | NHL repeat-containing protein | 15.5848 |
| LOC_Os05g50910 | Chr5:29213556-29220234:+:154 | extra-large G-protein-related | 14.9281 |
| LOC_Os08g35110 | Chr8:22136791-22137879:+:221 | OsSAUR33 - Auxin-responsive SAUR gene family member | 14.2462 |
| LOC_Os09g35010 | Chr9:20395224-20396205:-:-147 | dehydration-responsive element-binding protein (OsDREB1B) | 13.8964 |
| LOC_Os07g02200 | Chr7:699844-700831:-:-70 | plastocyanin-like domain containing protein | 13.7476 |
| LOC_Os10g39440 | Chr10:21048271-21052794:+:167 | transporter family protein (OsTMT1) | 13.4741 |
| LOC_Os09g07920 | Chr9:4042693-4045478:+:249 | NOI protein, nitrate-induced | 13.3778 |
| LOC_Os03g56820 | Chr3:32375903-32378469:+:99 | fatty acid hydroxylase (OsFAH2) | 13.2101 |
| LOC_Os03g22700 | Chr3:13110912-13115736:-:-161 | cyclin-dependent kinase C-2 | 13.2033 |
| LOC_Os04g58250 | Chr4:34680601-34685203:-:-421 | protein kinase, putative | 13.1943 |
| LOC_Os10g33800 | Chr10:17913818-17917850:+:215 | lactate/malate dehydrogenase | 13.0107 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, D.; Shi, S.; Hao, Z.; Xiong, W.; Luo, M. OsbZIP81, A Homologue of Arabidopsis VIP1, May Positively Regulate JA Levels by Directly Targetting the Genes in JA Signaling and Metabolism Pathway in Rice. Int. J. Mol. Sci. 2019, 20, 2360. https://doi.org/10.3390/ijms20092360
Liu D, Shi S, Hao Z, Xiong W, Luo M. OsbZIP81, A Homologue of Arabidopsis VIP1, May Positively Regulate JA Levels by Directly Targetting the Genes in JA Signaling and Metabolism Pathway in Rice. International Journal of Molecular Sciences. 2019; 20(9):2360. https://doi.org/10.3390/ijms20092360
Chicago/Turabian StyleLiu, Defang, Shaopeng Shi, Zhijun Hao, Wentao Xiong, and Meizhong Luo. 2019. "OsbZIP81, A Homologue of Arabidopsis VIP1, May Positively Regulate JA Levels by Directly Targetting the Genes in JA Signaling and Metabolism Pathway in Rice" International Journal of Molecular Sciences 20, no. 9: 2360. https://doi.org/10.3390/ijms20092360
APA StyleLiu, D., Shi, S., Hao, Z., Xiong, W., & Luo, M. (2019). OsbZIP81, A Homologue of Arabidopsis VIP1, May Positively Regulate JA Levels by Directly Targetting the Genes in JA Signaling and Metabolism Pathway in Rice. International Journal of Molecular Sciences, 20(9), 2360. https://doi.org/10.3390/ijms20092360




