Plasticity and Potency of Mammary Stem Cell Subsets During Mammary Gland Development
Abstract
1. Introduction
2. Lessons from Normal Mammary Gland Stem Cells
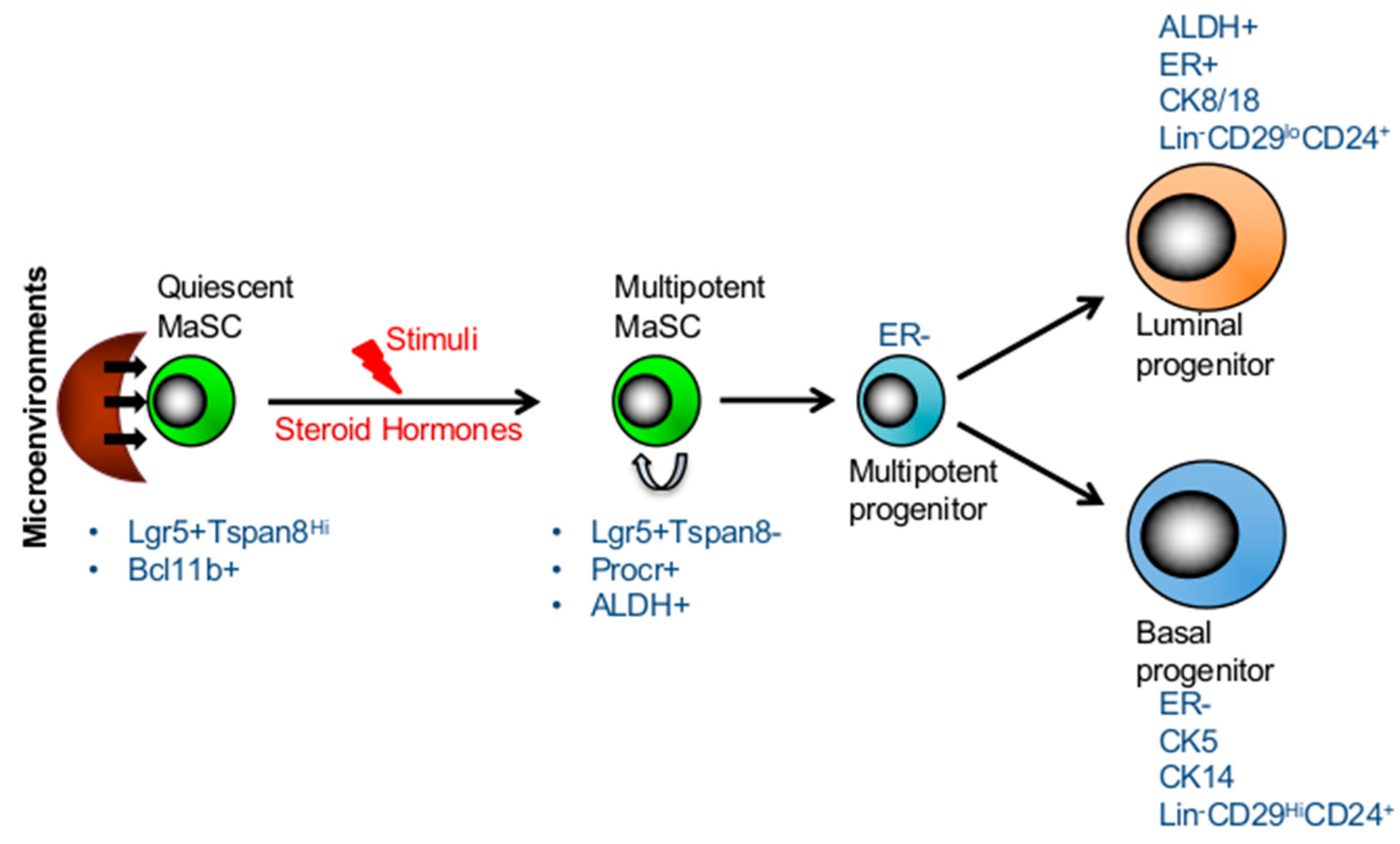
3. Long-Lived Quiescent MaSCs Which Are Activated in Response to Stimuli
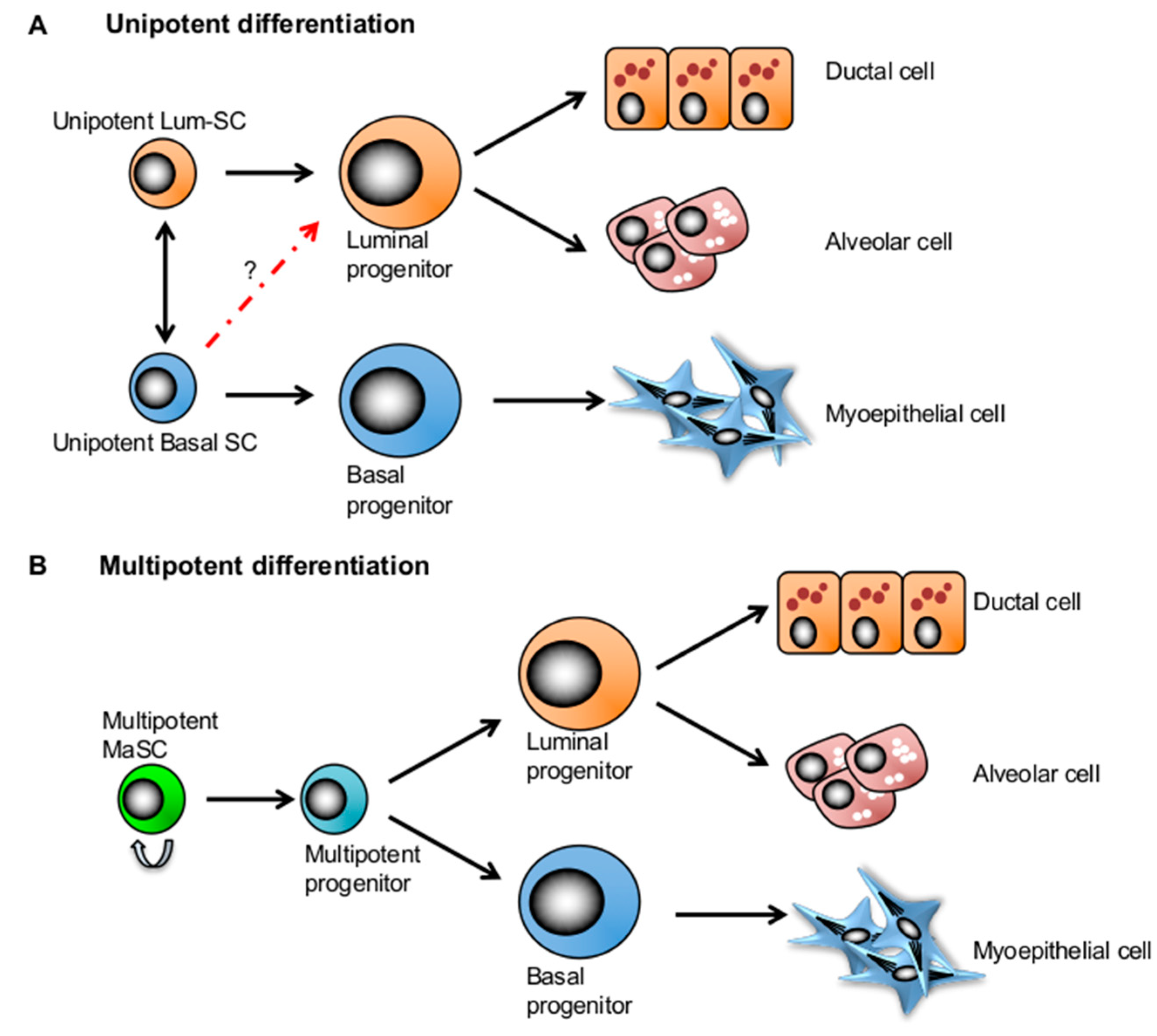
4. Competing Models: Multipotency and Lineage-Restricted Unipotent MaSCs
5. Mammary Stem Cell Plasticity Regulated by the Microenvironment during Mammary Development
6. Markers of Mammary Stem Cells and Progenitors
7. Concluding Remarks
Funding
Conflicts of Interest
References
- Merrell, A.J.; Stanger, B.Z. Adult cell plasticity in vivo: De-differentiation and transdifferentiation are back in style. Nat. Rev. Mol. Cell Biol. 2016, 17, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Morrison, S.J. Stem cell potential: Can anything make anything? Curr. Biol. 2001, 11, R7–R9. [Google Scholar] [CrossRef][Green Version]
- Okita, K.; Nakagawa, M.; Hyenjong, H.; Ichisaka, T.; Yamanaka, S. Generation of mouse induced pluripotent stem cells without viral vectors. Science 2008, 322, 949–953. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef]
- Chen, Y.; Yang, Z.; Zhao, Z.A.; Shen, Z. Direct reprogramming of fibroblasts into cardiomyocytes. Stem Cell Res. Ther. 2017, 8, 118. [Google Scholar] [CrossRef]
- Hou, S.; Lu, P. Direct reprogramming of somatic cells into neural stem cells or neurons for neurological disorders. Neural. Regen. Res. 2016, 11, 28–31. [Google Scholar] [PubMed]
- Kelaini, S.; Cochrane, A.; Margariti, A. Direct reprogramming of adult cells: Avoiding the pluripotent state. Stem Cells Cloning 2014, 7, 19–29. [Google Scholar]
- Ambasudhan, R.; Talantova, M.; Coleman, R.; Yuan, X.; Zhu, S.; Lipton, S.A.; Ding, S. Direct reprogramming of adult human fibroblasts to functional neurons under defined conditions. Cell Stem Cell 2011, 9, 113–118. [Google Scholar] [CrossRef]
- Zhou, Q.; Brown, J.; Kanarek, A.; Rajagopal, J.; Melton, D.A. In vivo reprogramming of adult pancreatic exocrine cells to beta-cells. Nature 2008, 455, 627–632. [Google Scholar] [CrossRef] [PubMed]
- Rackham, O.J.; Firas, J.; Fang, H.; Oates, M.E.; Holmes, M.L.; Knaupp, A.S.; Consortium, F.; Suzuki, H.; Nefzger, C.M.; Daub, C.O.; et al. A predictive computational framework for direct reprogramming between human cell types. Nat. Genet. 2016, 48, 331–335. [Google Scholar] [CrossRef] [PubMed]
- Rajapakse, I.; Groudine, M.; Mesbahi, M. Dynamics and control of state-dependent networks for probing genomic organization. Proc. Natl. Acad. Sci. USA 2011, 108, 17257–17262. [Google Scholar] [CrossRef]
- Fu, N.Y.; Rios, A.C.; Pal, B.; Law, C.W.; Jamieson, P.; Liu, R.; Vaillant, F.; Jackling, F.; Liu, K.H.; Smyth, G.K.; et al. Identification of quiescent and spatially restricted mammary stem cells that are hormone responsive. Nat. Cell Biol. 2017, 19, 164–176. [Google Scholar] [CrossRef]
- Cai, S.; Kalisky, T.; Sahoo, D.; Dalerba, P.; Feng, W.; Lin, Y.; Qian, D.; Kong, A.; Yu, J.; Wang, F.; et al. A quiescent bcl11b high stem cell population is required for maintenance of the mammary gland. Cell Stem Cell 2017, 20, 247–260. [Google Scholar] [CrossRef]
- Wang, D.; Cai, C.; Dong, X.; Yu, Q.C.; Zhang, X.O.; Yang, L.; Zeng, Y.A. Identification of multipotent mammary stem cells by protein c receptor expression. Nature 2015, 517, 81–84. [Google Scholar] [CrossRef] [PubMed]
- Rios, A.C.; Fu, N.Y.; Lindeman, G.J.; Visvader, J.E. In situ identification of bipotent stem cells in the mammary gland. Nature 2014, 506, 322–327. [Google Scholar] [CrossRef] [PubMed]
- Shackleton, M.; Vaillant, F.; Simpson, K.J.; Stingl, J.; Smyth, G.K.; Asselin-Labat, M.L.; Wu, L.; Lindeman, G.J.; Visvader, J.E. Generation of a functional mammary gland from a single stem cell. Nature 2006, 439, 84–88. [Google Scholar] [CrossRef] [PubMed]
- Kordon, E.C.; Smith, G.H. An entire functional mammary gland may comprise the progeny from a single cell. Development 1998, 125, 1921–1930. [Google Scholar] [PubMed]
- Stingl, J.; Eirew, P.; Ricketson, I.; Shackleton, M.; Vaillant, F.; Choi, D.; Li, H.I.; Eaves, C.J. Purification and unique properties of mammary epithelial stem cells. Nature 2006, 439, 993–997. [Google Scholar] [CrossRef]
- Makarem, M.; Kannan, N.; Nguyen, L.V.; Knapp, D.J.; Balani, S.; Prater, M.D.; Stingl, J.; Raouf, A.; Nemirovsky, O.; Eirew, P.; et al. Developmental changes in the in vitro activated regenerative activity of primitive mammary epithelial cells. PLoS Biol. 2013, 11, e1001630. [Google Scholar] [CrossRef]
- Spike, B.T.; Kelber, J.A.; Booker, E.; Kalathur, M.; Rodewald, R.; Lipianskaya, J.; La, J.; He, M.; Wright, T.; Klemke, R.; et al. Cripto/grp78 signaling maintains fetal and adult mammary stem cells ex vivo. Stem Cell Rep. 2014, 2, 427–439. [Google Scholar] [CrossRef][Green Version]
- Nguyen, L.V.; Makarem, M.; Carles, A.; Moksa, M.; Kannan, N.; Pandoh, P.; Eirew, P.; Osako, T.; Kardel, M.; Cheung, A.M.; et al. Clonal analysis via barcoding reveals diverse growth and differentiation of transplanted mouse and human mammary stem cells. Cell Stem Cell 2014, 14, 253–263. [Google Scholar] [CrossRef] [PubMed]
- Van Keymeulen, A.; Rocha, A.S.; Ousset, M.; Beck, B.; Bouvencourt, G.; Rock, J.; Sharma, N.; Dekoninck, S.; Blanpain, C. Distinct stem cells contribute to mammary gland development and maintenance. Nature 2011, 479, 189–193. [Google Scholar] [CrossRef] [PubMed]
- Wuidart, A.; Sifrim, A.; Fioramonti, M.; Matsumura, S.; Brisebarre, A.; Brown, D.; Centonze, A.; Dannau, A.; Dubois, C.; Van Keymeulen, A.; et al. Early lineage segregation of multipotent embryonic mammary gland progenitors. Nat. Cell Biol. 2018, 20, 666–676. [Google Scholar] [CrossRef]
- Lilja, A.M.; Rodilla, V.; Huyghe, M.; Hannezo, E.; Landragin, C.; Renaud, O.; Leroy, O.; Rulands, S.; Simons, B.D.; Fre, S. Clonal analysis of notch1-expressing cells reveals the existence of unipotent stem cells that retain long-term plasticity in the embryonic mammary gland. Nat. Cell Biol. 2018, 20, 677–687. [Google Scholar] [CrossRef]
- Donati, G.; Watt, F.M. Stem cell heterogeneity and plasticity in epithelia. Cell Stem Cell 2015, 16, 465–476. [Google Scholar] [CrossRef] [PubMed]
- Parmar, H.; Cunha, G.R. Epithelial-stromal interactions in the mouse and human mammary gland in vivo. Endocr. Relat. Cancer 2004, 11, 437–458. [Google Scholar] [CrossRef] [PubMed]
- Ingthorsson, S.; Briem, E.; Bergthorsson, J.T.; Gudjonsson, T. Epithelial plasticity during human breast morphogenesis and cancer progression. J. Mammary Gland. Biol. Neoplasia 2016, 21, 139–148. [Google Scholar] [CrossRef]
- Landskroner-Eiger, S.; Park, J.; Israel, D.; Pollard, J.W.; Scherer, P.E. Morphogenesis of the developing mammary gland: Stage-dependent impact of adipocytes. Dev. Biol. 2010, 344, 968–978. [Google Scholar] [CrossRef] [PubMed]
- Hendrix, M.J.; Seftor, E.A.; Hess, A.R.; Seftor, R.E. Vasculogenic mimicry and tumour-cell plasticity: Lessons from melanoma. Nat. Rev. Cancer 2003, 3, 411–421. [Google Scholar] [CrossRef]
- Maniotis, A.J.; Folberg, R.; Hess, A.; Seftor, E.A.; Gardner, L.M.; Pe’er, J.; Trent, J.M.; Meltzer, P.S.; Hendrix, M.J. Vascular channel formation by human melanoma cells in vivo and in vitro: Vasculogenic mimicry. Am. J. Pathol. 1999, 155, 739–752. [Google Scholar] [CrossRef]
- Bissell, M.J. Tumor plasticity allows vasculogenic mimicry, a novel form of angiogenic switch. A rose by any other name? Am. J. Pathol. 1999, 155, 675–679. [Google Scholar] [CrossRef]
- Bissell, M. Mammary gland development. Adv. Dent. Res. 1995, 9, 9. [Google Scholar] [CrossRef] [PubMed]
- Daniel, C.W.; Smith, G.H. The mammary gland: A model for development. J. Mammary Gland. Biol. Neoplasia 1999, 4, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Kondo, M.; Wagers, A.J.; Manz, M.G.; Prohaska, S.S.; Scherer, D.C.; Beilhack, G.F.; Shizuru, J.A.; Weissman, I.L. Biology of hematopoietic stem cells and progenitors: Implications for clinical application. Annu. Rev. Immunol. 2003, 21, 759–806. [Google Scholar] [CrossRef] [PubMed]
- Potten, C.S.; Booth, C.; Pritchard, D.M. The intestinal epithelial stem cell: The mucosal governor. Int. J. Exp. Pathol. 1997, 78, 219–243. [Google Scholar] [CrossRef] [PubMed]
- Keller, P.J.; Arendt, L.M.; Kuperwasser, C. Stem cell maintenance of the mammary gland: It takes two. Cell Stem Cell 2011, 9, 496–497. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Liao, M.J.; Zhang, C.C.; Zhou, B.; Zimonjic, D.B.; Mani, S.A.; Kaba, M.; Gifford, A.; Reinhardt, F.; Popescu, N.C.; Guo, W.; et al. Enrichment of a population of mammary gland cells that form mammospheres and have in vivo repopulating activity. Cancer Res. 2007, 67, 8131–8138. [Google Scholar] [CrossRef]
- Ginestier, C.; Wicha, M.S. Mammary stem cell number as a determinate of breast cancer risk. Breast Cancer Res. 2007, 9, 109. [Google Scholar] [CrossRef]
- Ginestier, C.; Hur, M.H.; Charafe-Jauffret, E.; Monville, F.; Dutcher, J.; Brown, M.; Jacquemier, J.; Viens, P.; Kleer, C.G.; Liu, S.; et al. Aldh1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 2007, 1, 555–567. [Google Scholar] [CrossRef]
- Fridriksdottir, A.J.; Petersen, O.W.; Ronnov-Jessen, L. Mammary gland stem cells: Current status and future challenges. Int. J. Dev. Biol. 2011, 55, 719–729. [Google Scholar] [CrossRef]
- Boras-Granic, K.; Dann, P.; Wysolmerski, J.J. Embryonic cells contribute directly to the quiescent stem cell population in the adult mouse mammary gland. Breast Cancer Res. 2014, 16, 487. [Google Scholar] [CrossRef]
- Baldridge, M.T.; King, K.Y.; Boles, N.C.; Weksberg, D.C.; Goodell, M.A. Quiescent haematopoietic stem cells are activated by ifn-gamma in response to chronic infection. Nature 2010, 465, 793–797. [Google Scholar] [CrossRef]
- Cicalese, A.; Bonizzi, G.; Pasi, C.E.; Faretta, M.; Ronzoni, S.; Giulini, B.; Brisken, C.; Minucci, S.; Di Fiore, P.P.; Pelicci, P.G. The tumor suppressor p53 regulates polarity of self-renewing divisions in mammary stem cells. Cell 2009, 138, 1083–1095. [Google Scholar] [CrossRef]
- Pellacani, D.; Tan, S.; Lefort, S.; Eaves, C.J. Transcriptional regulation of normal human mammary cell heterogeneity and its perturbation in breast cancer. EMBO J. 2019. [Google Scholar] [CrossRef]
- Visvader, J.E.; Stingl, J. Mammary stem cells and the differentiation hierarchy: Current status and perspectives. Genes Dev. 2014, 28, 1143–1158. [Google Scholar] [CrossRef]
- Oakes, S.R.; Gallego-Ortega, D.; Ormandy, C.J. The mammary cellular hierarchy and breast cancer. Cell Mol. Life Sci. 2014, 71, 4301–4324. [Google Scholar] [CrossRef]
- Domenici, G.; Aurrekoetxea-Rodriguez, I.; Simoes, B.M.; Rabano, M.; Lee, S.Y.; Millan, J.S.; Comaills, V.; Oliemuller, E.; Lopez-Ruiz, J.A.; Zabalza, I.; et al. A sox2-sox9 signalling axis maintains human breast luminal progenitor and breast cancer stem cells. Oncogene 2019, 38, 3151–3169. [Google Scholar] [CrossRef]
- Shehata, M.; Teschendorff, A.; Sharp, G.; Novcic, N.; Russell, I.A.; Avril, S.; Prater, M.; Eirew, P.; Caldas, C.; Watson, C.J.; et al. Phenotypic and functional characterisation of the luminal cell hierarchy of the mammary gland. Breast Cancer Res. 2012, 14, R134. [Google Scholar] [CrossRef]
- Eirew, P.; Kannan, N.; Knapp, D.J.; Vaillant, F.; Emerman, J.T.; Lindeman, G.J.; Visvader, J.E.; Eaves, C.J. Aldehyde dehydrogenase activity is a biomarker of primitive normal human mammary luminal cells. Stem Cells 2012, 30, 344–348. [Google Scholar] [CrossRef]
- Lim, E.; Vaillant, F.; Wu, D.; Forrest, N.C.; Pal, B.; Hart, A.H.; Asselin-Labat, M.L.; Gyorki, D.E.; Ward, T.; Partanen, A.; et al. Aberrant luminal progenitors as the candidate target population for basal tumor development in brca1 mutation carriers. Nat. Med. 2009, 15, 907–913. [Google Scholar] [CrossRef]
- Rodilla, V.; Dasti, A.; Huyghe, M.; Lafkas, D.; Laurent, C.; Reyal, F.; Fre, S. Luminal progenitors restrict their lineage potential during mammary gland development. PLoS Biol. 2015, 13, e1002069. [Google Scholar] [CrossRef]
- Wahl, G.M.; Spike, B.T. Cell state plasticity, stem cells, emt, and the generation of intra-tumoral heterogeneity. NPJ Breast Cancer 2017, 3, 14. [Google Scholar] [CrossRef]
- Wuidart, A.; Ousset, M.; Rulands, S.; Simons, B.D.; Van Keymeulen, A.; Blanpain, C. Quantitative lineage tracing strategies to resolve multipotency in tissue-specific stem cells. Genes Dev 2016, 30, 1261–1277. [Google Scholar] [CrossRef]
- Barker, N.; van Es, J.H.; Kuipers, J.; Kujala, P.; van den Born, M.; Cozijnsen, M.; Haegebarth, A.; Korving, J.; Begthel, H.; Peters, P.J.; et al. Identification of stem cells in small intestine and colon by marker gene lgr5. Nature 2007, 449, 1003–1007. [Google Scholar] [CrossRef]
- Trejo, C.L.; Luna, G.; Dravis, C.; Spike, B.T.; Wahl, G.M. Lgr5 is a marker for fetal mammary stem cells, but is not essential for stem cell activity or tumorigenesis. NPJ Breast Cancer 2017, 3, 16. [Google Scholar] [CrossRef]
- Jaks, V.; Barker, N.; Kasper, M.; van Es, J.H.; Snippert, H.J.; Clevers, H.; Toftgard, R. Lgr5 marks cycling, yet long-lived, hair follicle stem cells. Nat. Genet. 2008, 40, 1291–1299. [Google Scholar] [CrossRef]
- Hoeck, J.D.; Biehs, B.; Kurtova, A.V.; Kljavin, N.M.; de Sousa, E.M.F.; Alicke, B.; Koeppen, H.; Modrusan, Z.; Piskol, R.; de Sauvage, F.J. Stem cell plasticity enables hair regeneration following lgr5+ cell loss. Nat. Cell Biol. 2017, 19, 666–676. [Google Scholar] [CrossRef]
- Celia-Terrassa, T.; Liu, D.D.; Choudhury, A.; Hang, X.; Wei, Y.; Zamalloa, J.; Alfaro-Aco, R.; Chakrabarti, R.; Jiang, Y.Z.; Koh, B.I.; et al. Normal and cancerous mammary stem cells evade interferon-induced constraint through the mir-199a-lcor axis. Nat. Cell Biol. 2017, 19, 711–723. [Google Scholar] [CrossRef]
- Pietras, E.M.; Lakshminarasimhan, R.; Techner, J.M.; Fong, S.; Flach, J.; Binnewies, M.; Passegue, E. Re-entry into quiescence protects hematopoietic stem cells from the killing effect of chronic exposure to type i interferons. J. Exp. Med. 2014, 211, 245–262. [Google Scholar] [CrossRef]
- Scheele, C.L.; Hannezo, E.; Muraro, M.J.; Zomer, A.; Langedijk, N.S.; van Oudenaarden, A.; Simons, B.D.; van Rheenen, J. Identity and dynamics of mammary stem cells during branching morphogenesis. Nature 2017, 542, 313–317. [Google Scholar] [CrossRef]
- Deome, K.B.; Faulkin, L.J., Jr.; Bern, H.A.; Blair, P.B. Development of mammary tumors from hyperplastic alveolar nodules transplanted into gland-free mammary fat pads of female c3h mice. Cancer Res. 1959, 19, 515–520. [Google Scholar]
- Jones, P.H.; Harper, S.; Watt, F.M. Stem cell patterning and fate in human epidermis. Cell 1995, 80, 83–93. [Google Scholar] [CrossRef]
- Lim, S.C.; Oh, S.H. The role of cd24 in various human epithelial neoplasias. Pathol. Res. Pract. 2005, 201, 479–486. [Google Scholar] [CrossRef]
- Rietze, R.L.; Valcanis, H.; Brooker, G.F.; Thomas, T.; Voss, A.K.; Bartlett, P.F. Purification of a pluripotent neural stem cell from the adult mouse brain. Nature 2001, 412, 736–739. [Google Scholar] [CrossRef]
- Eirew, P.; Stingl, J.; Raouf, A.; Turashvili, G.; Aparicio, S.; Emerman, J.T.; Eaves, C.J. A method for quantifying normal human mammary epithelial stem cells with in vivo regenerative ability. Nat. Med. 2008, 14, 1384–1389. [Google Scholar] [CrossRef]
- Welm, B.E.; Tepera, S.B.; Venezia, T.; Graubert, T.A.; Rosen, J.M.; Goodell, M.A. Sca-1(pos) cells in the mouse mammary gland represent an enriched progenitor cell population. Dev. Biol. 2002, 245, 42–56. [Google Scholar] [CrossRef]
- Alvi, A.J.; Clayton, H.; Joshi, C.; Enver, T.; Ashworth, A.; Vivanco, M.M.; Dale, T.C.; Smalley, M.J. Functional and molecular characterisation of mammary side population cells. Breast Cancer Res. 2003, 5, R1–R8. [Google Scholar] [CrossRef]
- Smalley, M.J.; Clarke, R.B. The mammary gland "side population": A putative stem/progenitor cell marker? J. Mammary Gland. Biol. Neoplasia 2005, 10, 37–47. [Google Scholar] [CrossRef]
- Allen, J.E.; Hart, L.S.; Dicker, D.T.; Wang, W.; El-Deiry, W.S. Visualization and enrichment of live putative cancer stem cell populations following p53 inactivation or bax deletion using non-toxic fluorescent dyes. Cancer Biol. Ther. 2009, 8, 2194–2205. [Google Scholar] [CrossRef]
- Hess, D.A.; Meyerrose, T.E.; Wirthlin, L.; Craft, T.P.; Herrbrich, P.E.; Creer, M.H.; Nolta, J.A. Functional characterization of highly purified human hematopoietic repopulating cells isolated according to aldehyde dehydrogenase activity. Blood 2004, 104, 1648–1655. [Google Scholar] [CrossRef]
- Kastan, M.B.; Schlaffer, E.; Russo, J.E.; Colvin, O.M.; Civin, C.I.; Hilton, J. Direct demonstration of elevated aldehyde dehydrogenase in human hematopoietic progenitor cells. Blood 1990, 75, 1947–1950. [Google Scholar] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, E.; Piranlioglu, R.; Wicha, M.S.; Korkaya, H. Plasticity and Potency of Mammary Stem Cell Subsets During Mammary Gland Development. Int. J. Mol. Sci. 2019, 20, 2357. https://doi.org/10.3390/ijms20092357
Lee E, Piranlioglu R, Wicha MS, Korkaya H. Plasticity and Potency of Mammary Stem Cell Subsets During Mammary Gland Development. International Journal of Molecular Sciences. 2019; 20(9):2357. https://doi.org/10.3390/ijms20092357
Chicago/Turabian StyleLee, Eunmi, Raziye Piranlioglu, Max S. Wicha, and Hasan Korkaya. 2019. "Plasticity and Potency of Mammary Stem Cell Subsets During Mammary Gland Development" International Journal of Molecular Sciences 20, no. 9: 2357. https://doi.org/10.3390/ijms20092357
APA StyleLee, E., Piranlioglu, R., Wicha, M. S., & Korkaya, H. (2019). Plasticity and Potency of Mammary Stem Cell Subsets During Mammary Gland Development. International Journal of Molecular Sciences, 20(9), 2357. https://doi.org/10.3390/ijms20092357




