Transcriptomic Analysis Reveals a Comprehensive Calcium- and Phytohormone-Dominated Signaling Response in Leymus chinensis Self-Incompatibility
Abstract
1. Introduction
2. Results
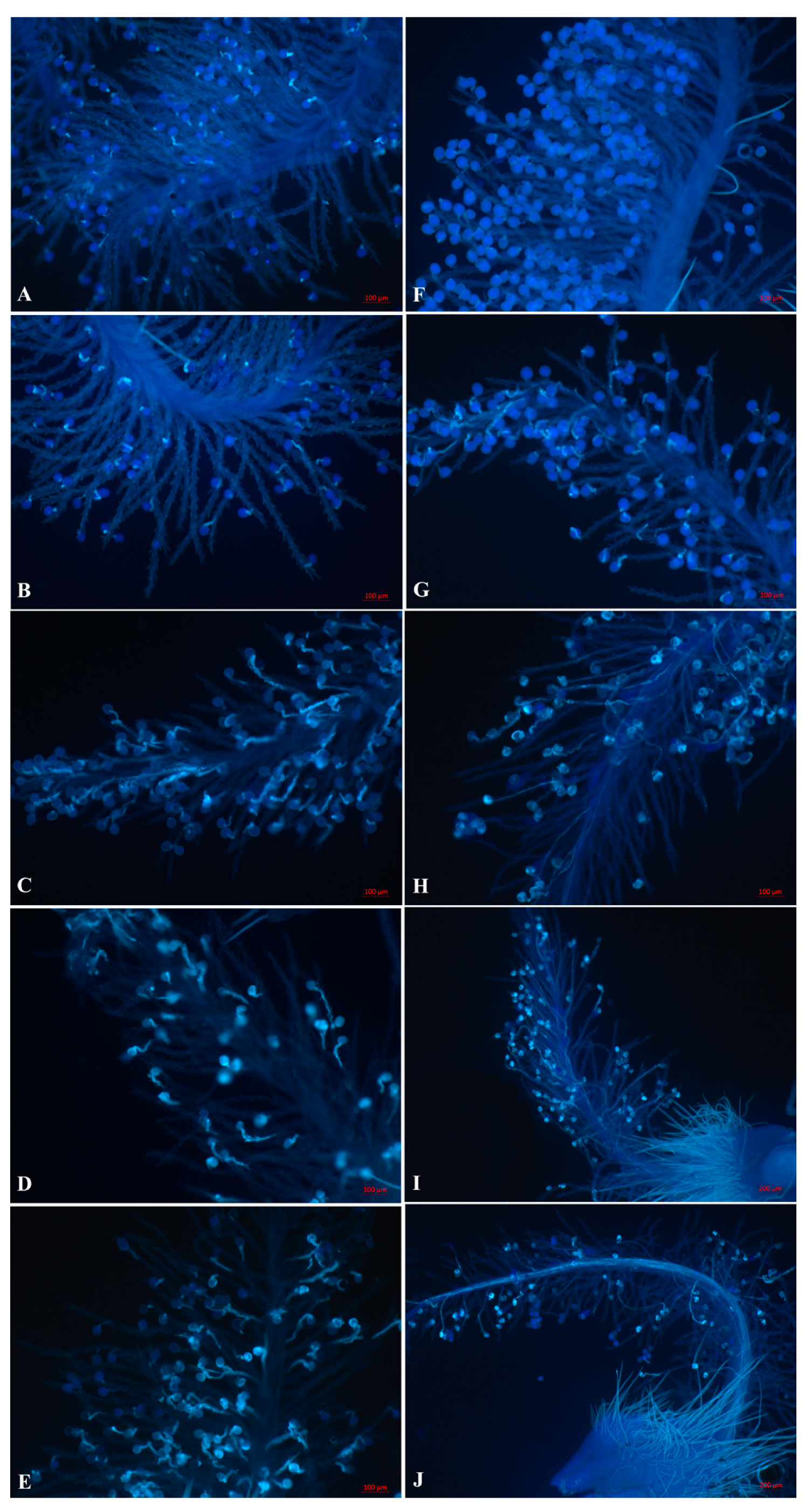
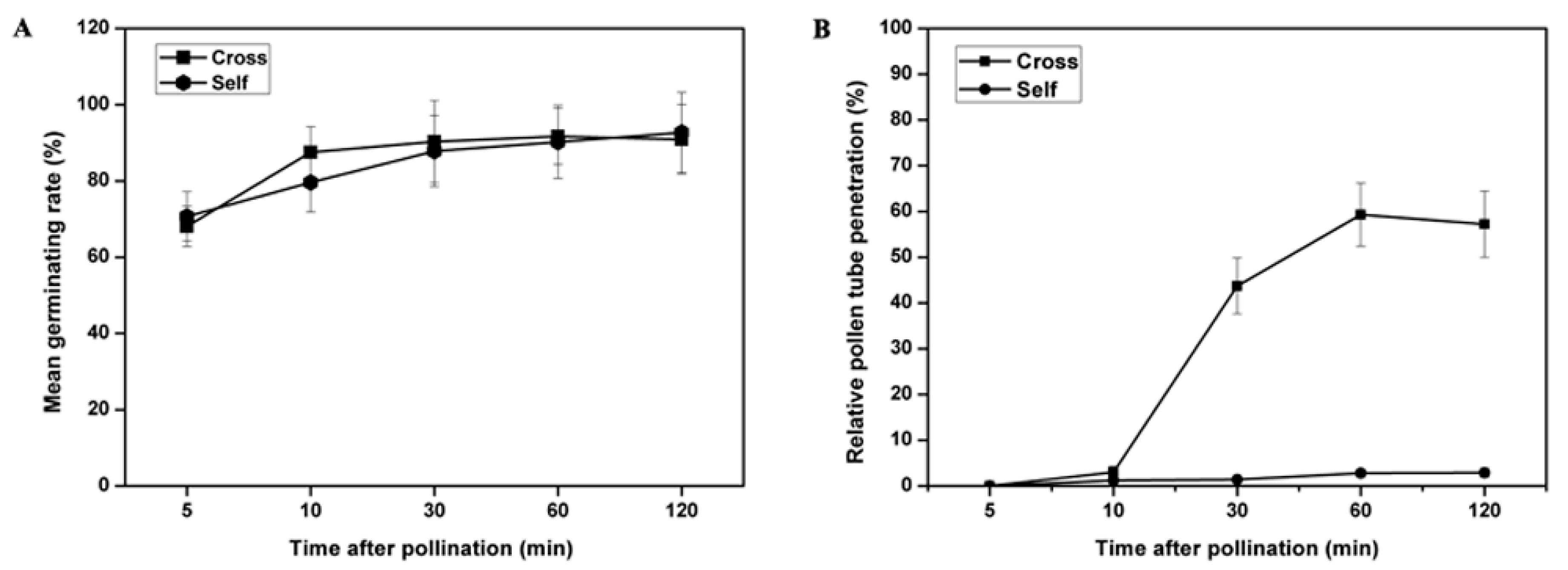
2.1. Pollen Tube Growth Patterns are Different in Self- and Cross-Pollination of Sheepgrass
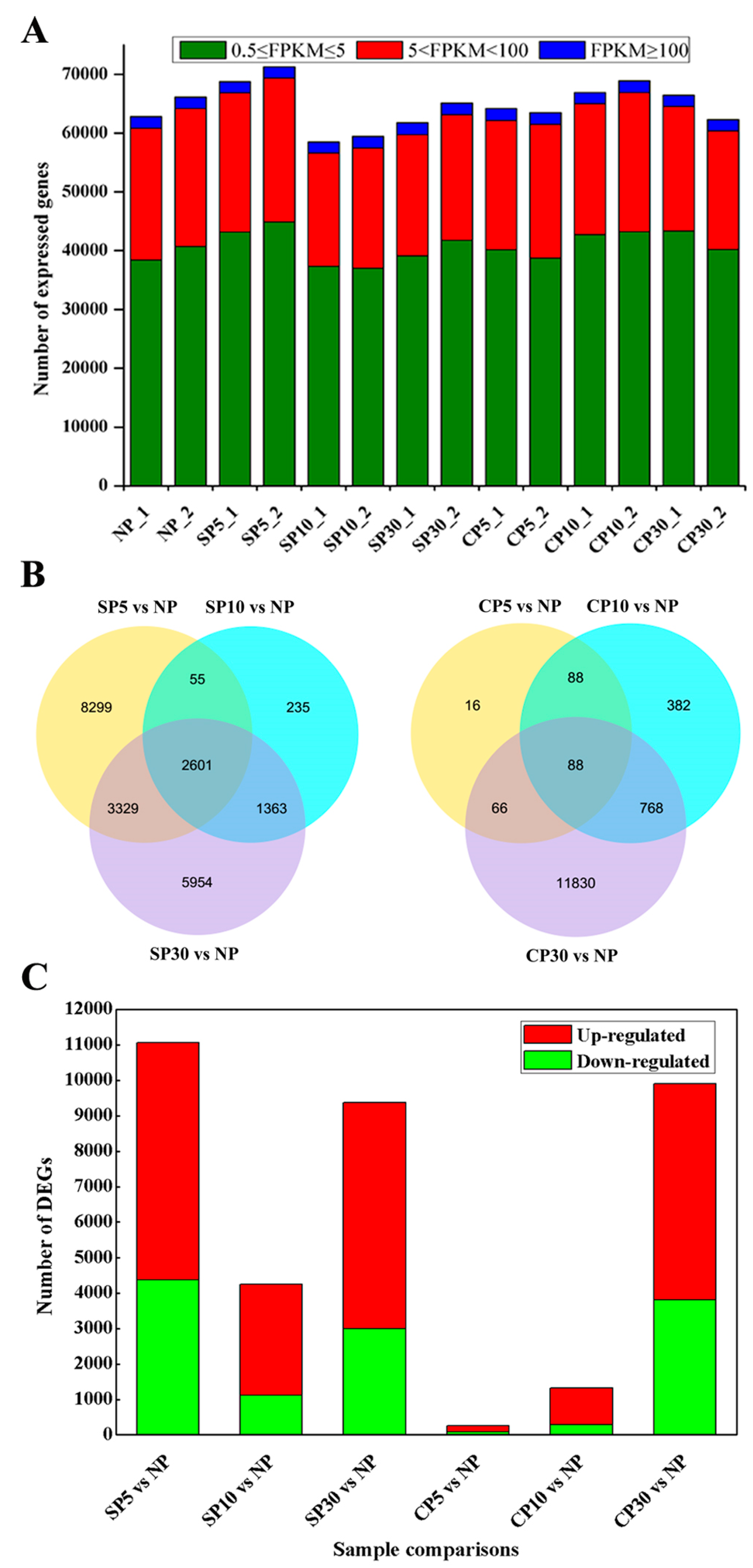
2.2. Global Comparison of Pistil Transcriptomes during Self- and Cross-Pollination in Sheepgrass
2.3. Differentially Expressed Genes are Distinct in Self- and Cross-Pollinated Pistils
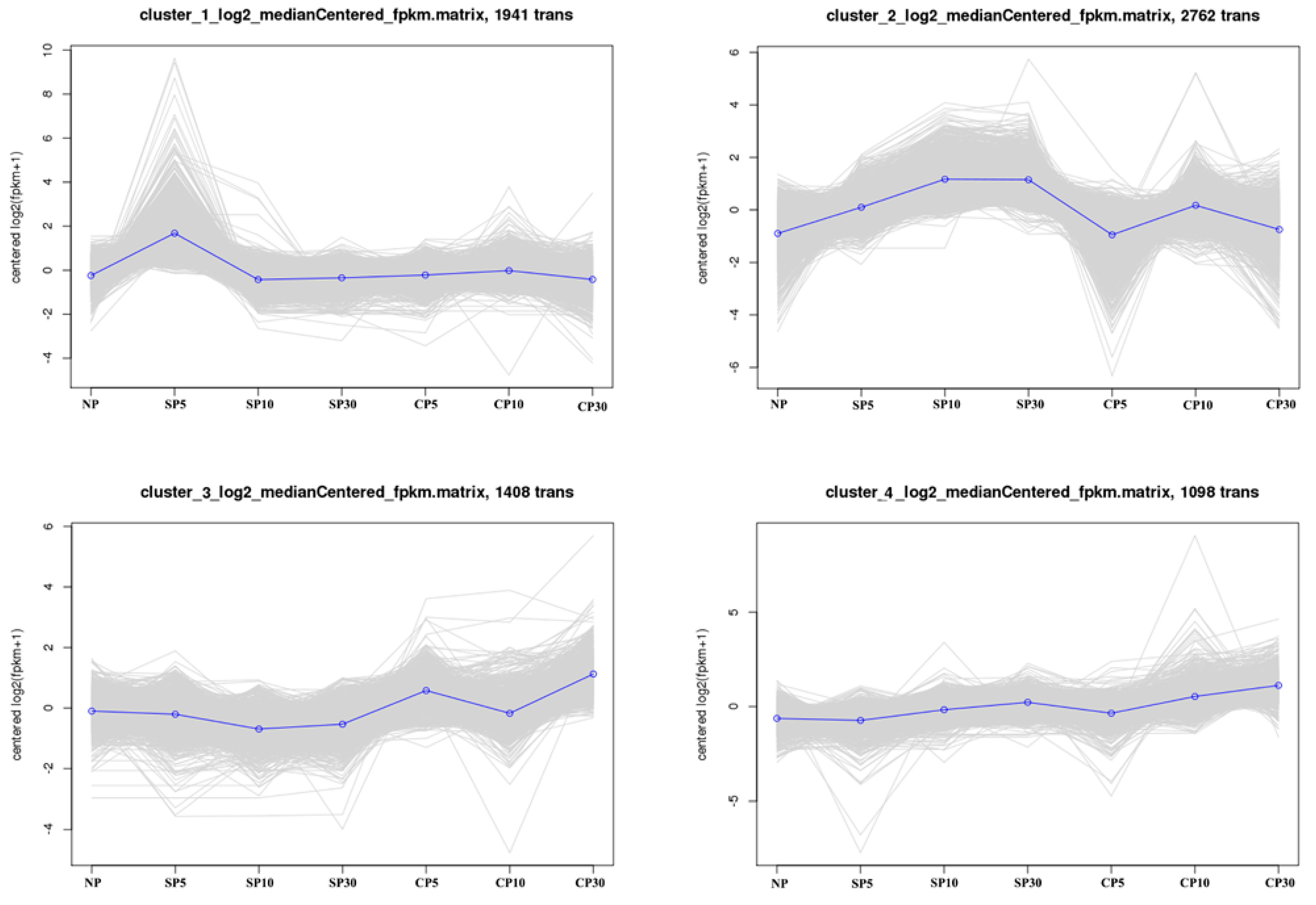
2.4. K-Means Clustering Analysis of DEGs
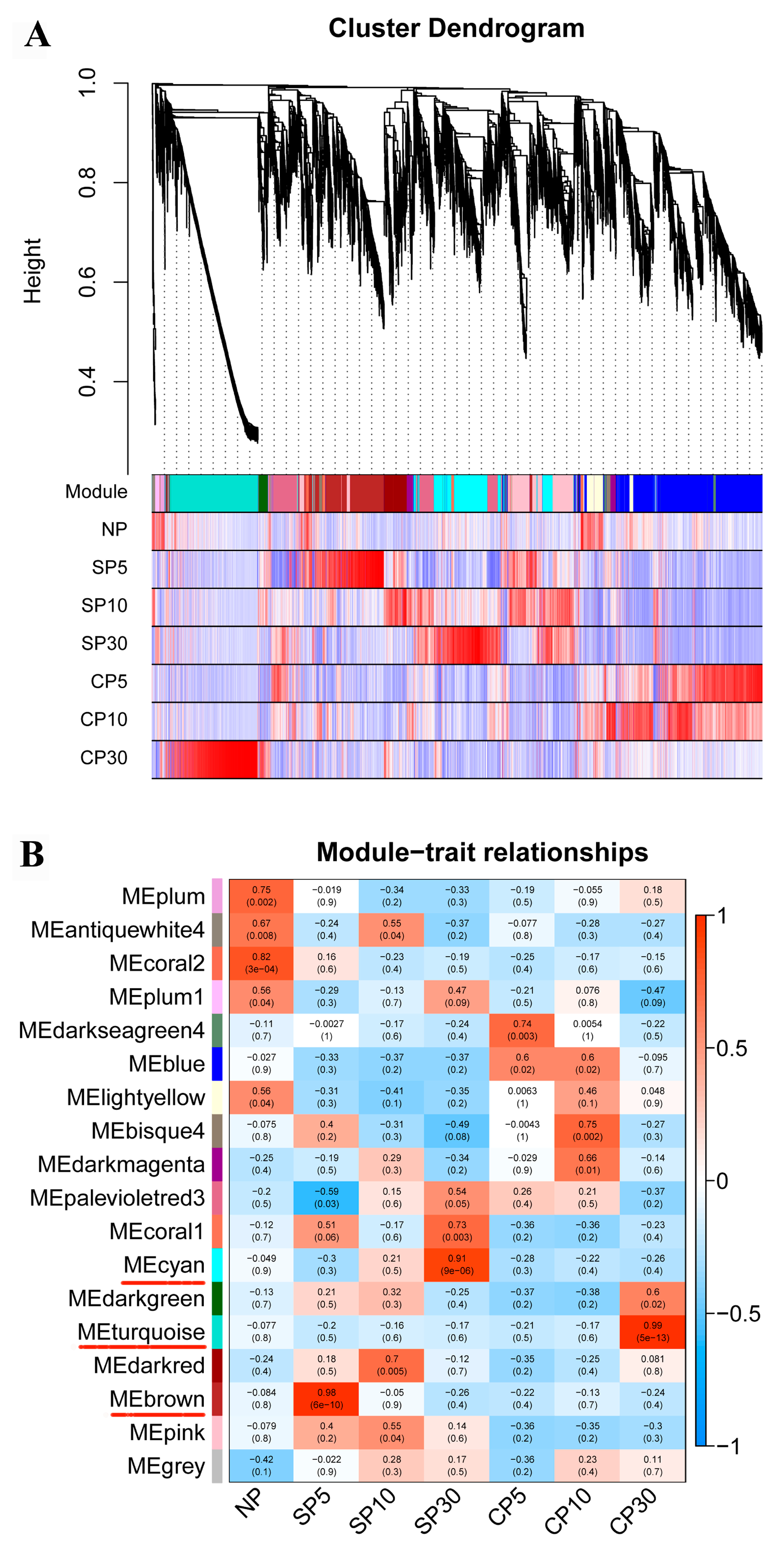
2.5. Weighted Gene Coexpression Network Analysis of DEGs
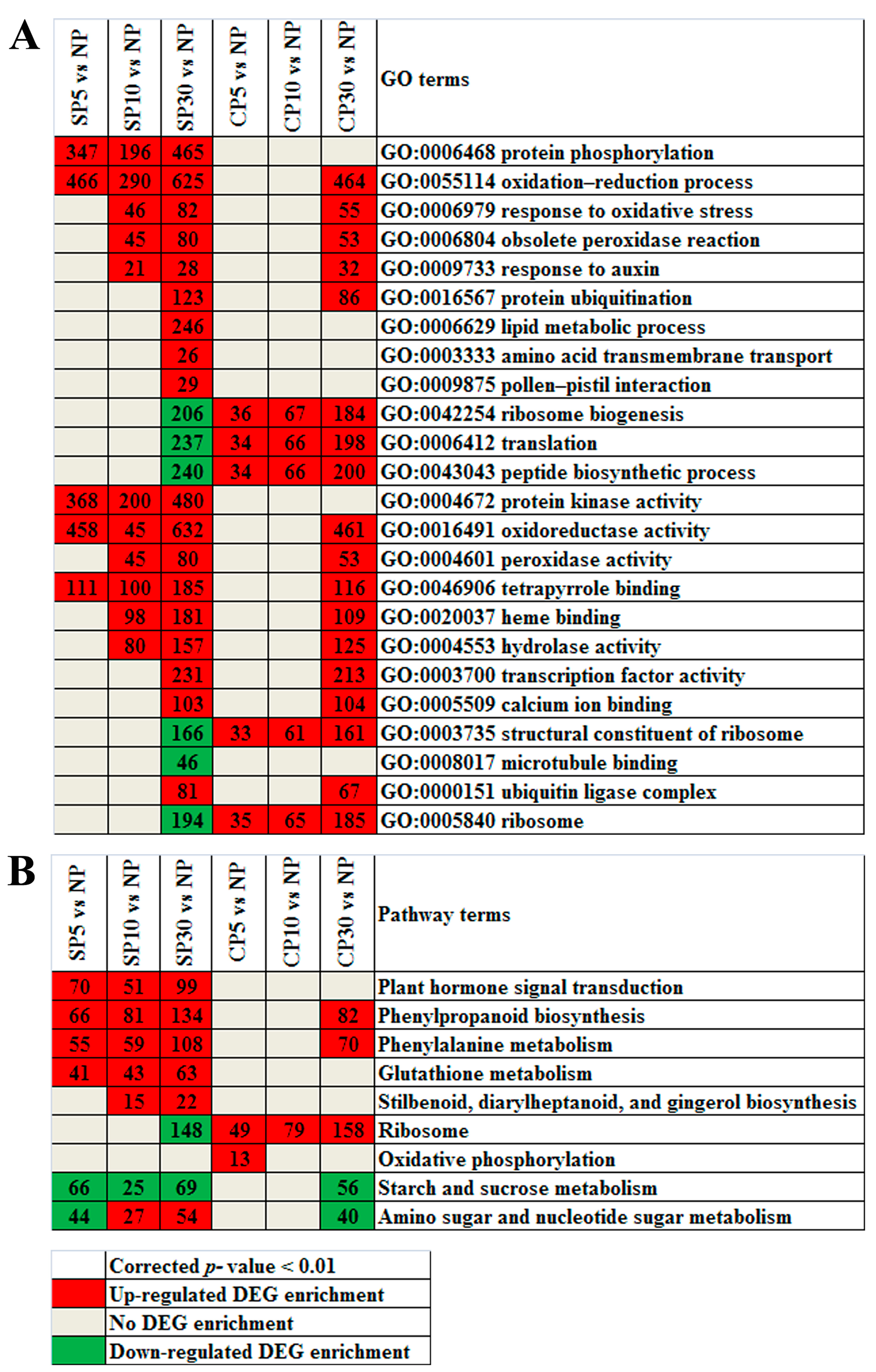
2.6. Enriched Gene Ontology Terms and Kyoto Encyclopedia of Genes and Genomes Pathways of DEGs
2.7. A Pool of SI-Related Gene Categories
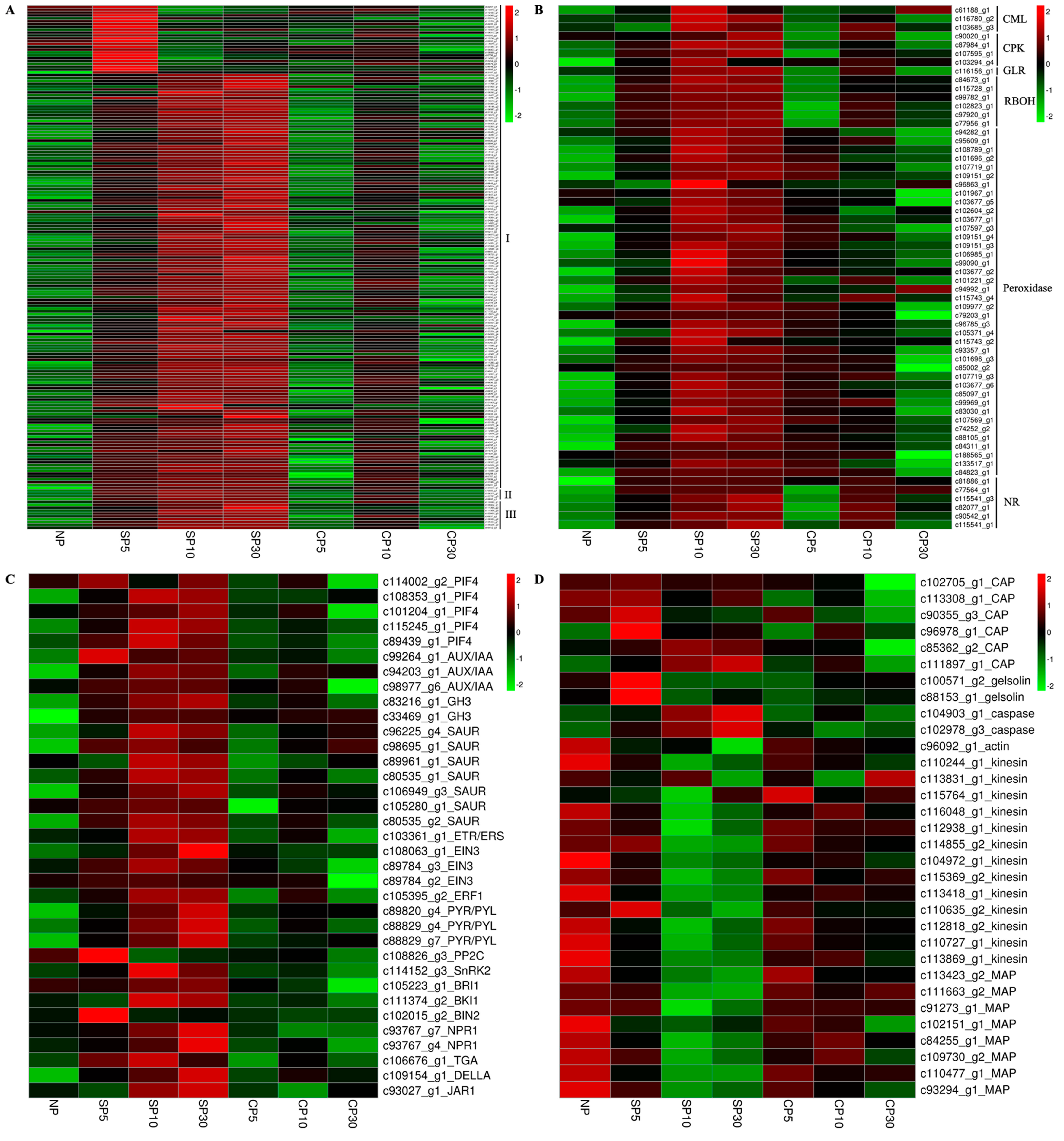
2.7.1. SI Triggers Protein Phosphorylation-Related Gene Expression in Sheepgrass
2.7.2. SI Triggers Calcium, ROS, and NO Signaling-Related Gene Expression in Sheepgrass
2.7.3. SI Triggers Plant Hormone Signaling-Related Gene Expression in Sheepgrass
2.7.4. SI Triggers Cytoskeleton-Related Gene Expression in Sheepgrass
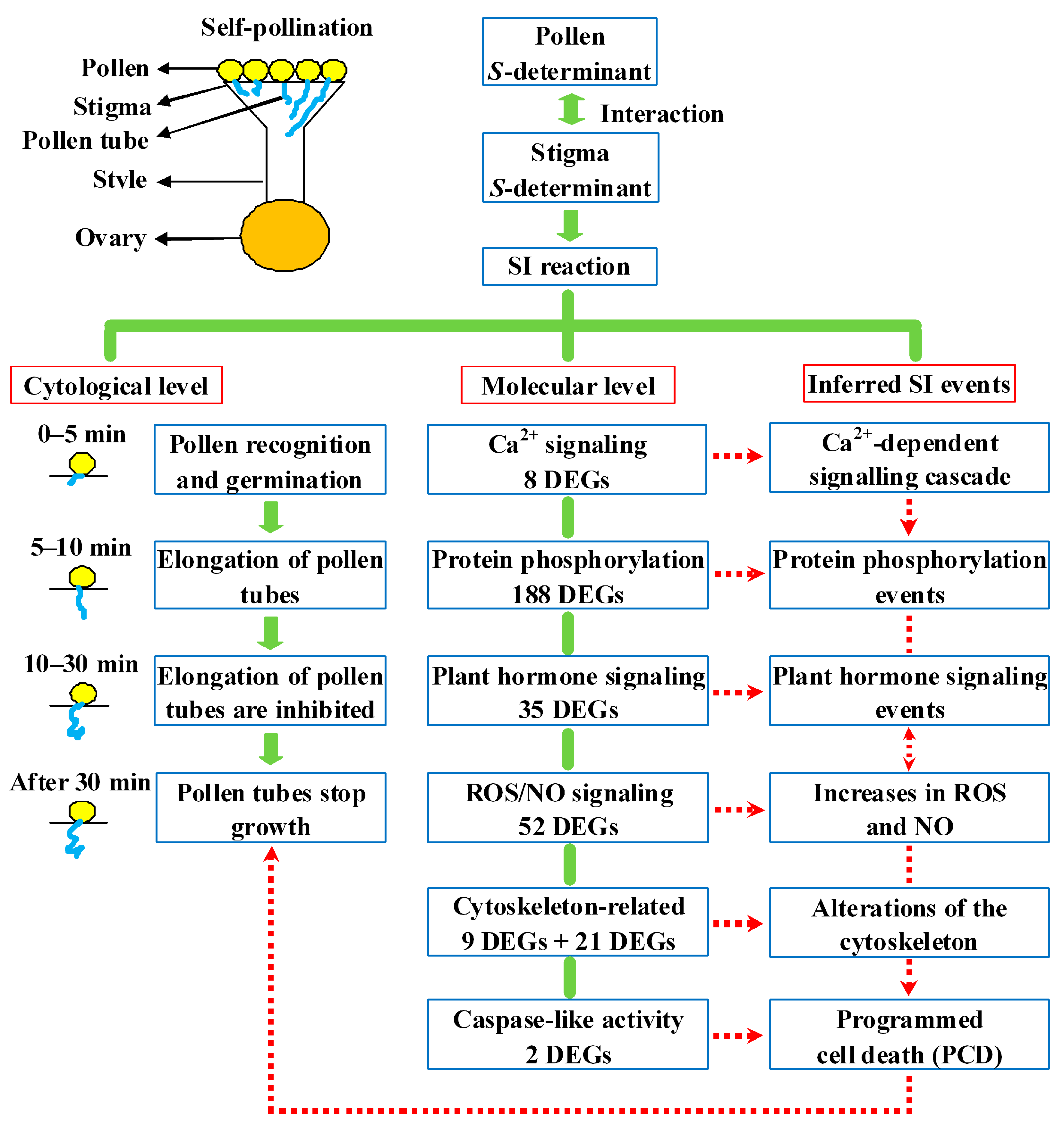
2.8. Putative SI Response Molecular Model in Sheepgrass
3. Discussion
3.1. Inhibition of Incompatible Pollen Tube Growth Is Rapid in Sheepgrass
3.2. Protein Phosphorylation Events Involved in the Sheepgrass SI Response
3.3. Sheepgrass SI is Involved in Calcium, ROS, and NO Signaling
3.4. Roles of Plant Hormones in the Sheepgrass SI Response
3.5. Sheepgrass SI May Trigger Alterations to the Cytoskeleton and PCD
4. Materials and Methods
4.1. Plant Materials
4.2. Observation of Pollen Tube Growth
4.3. RNA Extraction, Library Construction, and Sequencing
4.4. De Novo Assembly and Functional Annotation
4.5. Differential Expression, Cluster Analysis, and GO Enrichment Analysis
4.6. Quantitative Real-Time PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| SI | Self-incompatibility |
| SSI | Sporophytic self-incompatibility |
| GSI | Gametophytic self-incompatibility |
| PM | Plasma membrane |
| sPPase | Soluble inorganic pyrophosphatase |
| ARC1 | Arm repeat containing 1 |
| MLPK | M-locus protein kinase |
| THL1 | Thioredoxin h-like1 |
| ROS | Reactive oxygen species |
| NO | Nitric oxide |
| PCD | Programmed cell death |
| SLF | S locus F-box |
| FPKM | Fragments per kilobase of transcript per million mapped reads |
| DEGs | Differentially expressed genes |
| WGCNA | Weighted gene coexpression network analysis |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| CRPs | Cysteine-rich peptides |
| SCR | S locus cysteine-rich |
| SRK | S-receptor kinase |
| GLR | Glutamate receptor-like channel |
| CML | Calcium-binding protein |
| CPK | Calcium-dependent protein kinase |
| RBOH | Respiratory burst oxidase homolog |
| NR | Nitrate reductase |
| Aux/IAA | Auxin/indole-3-acetic acid |
| GH3 | Gretchen Hagen 3 |
| SAUR | Small auxin-up RNA |
| PIFs | Phytochrome-interacting factors |
| ETR, ERS | Ethylene receptor |
| ERF1 | Ethylene-responsive transcription factor 1 |
| EIN3 | Ethylene-insensitive protein 3 |
| PYR/PYL | Abscisic acid receptors |
| PP2C | Protein phosphatase 2C |
| SnRK2 | Serine/threonine protein kinase SRK2 |
| ABPs | Actin-binding proteins |
| MAPs | Microtubule-associated proteins |
| CAP | Cyclase-associated protein |
| Ca2+ | Calcium |
| NAA | 1-Naphthaleneacetic acid |
| ARF3 | Auxin response factor 3 |
References
- Fujii, S.; Kubo, K.; Takayama, S. Non-self- and self-recognition models in plant self-incompatibility. Nat. Plants 2016, 2, 16130. [Google Scholar] [CrossRef] [PubMed]
- Hayman, D.L.; Richter, J. Mutations affecting self-Incompatibility in Phalaris-coerulescens Desf (Poaceae). Heredity 1992, 68, 495–503. [Google Scholar] [CrossRef][Green Version]
- Thorogood, D.; Hayward, M.D. Self-compatibility in Lolium-temulentum L.—its genetic-control and transfer into Lolium-perenne L. and L-multiflorum Lam. Heredity 1992, 68, 71–78. [Google Scholar] [CrossRef][Green Version]
- Klaas, M.; Yang, B.C.; Bosch, M.; Thorogood, D.; Manzanares, C.; Armstead, I.P.; Franklin, F.C.H.; Barth, S. Progress towards elucidating the mechanisms of self-incompatibility in the grasses: Further insights from studies in Lolium. Ann. Bot. 2011, 108, 677–685. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.S.; Huang, S.S.; Kao, T.H. S-proteins control rejection of incompatible pollen in Petunia-inflata. Nature 1994, 367, 560–563. [Google Scholar] [CrossRef]
- Murfett, J.; Atherton, T.L.; Mou, B.; Gasser, C.S.; Mcclure, B.A. S-Rnase expressed in transgenic Nicotiana causes S-allele-specific pollen rejection. Nature 1994, 367, 563–566. [Google Scholar] [CrossRef]
- Mcclure, B.A.; Haring, V.; Ebert, P.R.; Anderson, M.A.; Simpson, R.J.; Sakiyama, F.; Clarke, A.E. Style self-incompatibility gene-products of Nicotiana-alata are ribonucleases. Nature 1989, 342, 955–957. [Google Scholar] [CrossRef] [PubMed]
- Sijacic, P.; Wang, X.; Skirpan, A.L.; Wang, Y.; Dowd, P.E.; McCubbin, A.G.; Huang, S.; Kao, T.H. Identification of the pollen determinant of S-RNase-mediated self-incompatibility. Nature 2004, 429, 302–305. [Google Scholar] [CrossRef]
- Entani, T.; Kubo, K.; Isogai, S.; Fukao, Y.; Shirakawa, M.; Isogai, A.; Takayama, S. Ubiquitin-proteasome-mediated degradation of S-RNase in a solanaceous cross-compatibility reaction. Plant J. 2014, 78, 1014–1021. [Google Scholar] [CrossRef] [PubMed]
- Williams, J.S.; Wu, L.H.; Li, S.; Sun, P.L.; Kao, T.H. Insight into S-RNase-based self-incompatibility in Petunia: Recent findings and future directions. Front. Plant Sci. 2015, 6, 41. [Google Scholar] [CrossRef]
- Foote, H.C.C.; Ride, J.P.; Franklintong, V.E.; Walker, E.A.; Lawrence, M.J.; Franklin, F.C.H. Cloning and expression of a distinctive class of self-incompatibility (S) gene from Papaver-rhoeas L. Proc. Natl. Acad. Sci. USA 1994, 91, 2265–2269. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, M.J.; de Graaf, B.H.J.; Hadjiosif, N.; Perry, R.M.; Poulter, N.S.; Osman, K.; Vatovec, S.; Harper, A.; Franklin, F.C.H.; Franklin-Tong, V.E. Identification of the pollen self-incompatibility determinant in Papaver rhoeas. Nature 2009, 459, 992–995. [Google Scholar] [CrossRef] [PubMed]
- Eaves, D.J.; Flores-Ortiz, C.; Haque, T.; Lin, Z.C.; Teng, N.J.; Franklin-Tong, V.E. Self-incompatibility in Papaver: Advances in integrating the signalling network. Biochem. Soc. Trans. 2014, 42, 370–376. [Google Scholar] [CrossRef]
- Wilkins, K.A.; Poulter, N.S.; Franklin-Tong, V.E. Taking one for the team: Self-recognition and cell suicide in pollen. J. Exp. Bot. 2014, 65, 1331–1342. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, K.A.; Bosch, M.; Haque, T.; Teng, N.; Poulter, N.S.; Franklin-Tong, V.E. Self-incompatibility-induced programmed cell death in field Poppy pollen involves dramatic acidification of the incompatible pollen tube cytosol. Plant Physiol. 2015, 167, 766–779. [Google Scholar] [CrossRef]
- Bedinger, P.A.; Broz, A.K.; Tovar-Mendez, A.; McClure, B. Pollen-pistil interactions and their role in mate selection. Plant Physiol. 2017, 173, 79–90. [Google Scholar] [CrossRef] [PubMed]
- Schopfer, C.R.; Nasrallah, M.E.; Nasrallah, J.B. The male determinant of self-incompatibility in Brassica. Science 1999, 286, 1697–1700. [Google Scholar] [CrossRef]
- Kachroo, A.; Schopfer, C.R.; Nasrallah, M.E.; Nasrallah, J.B. Allele-specific receptor-ligand interactions in Brassica self-incompatibility. Science 2001, 293, 1824–1826. [Google Scholar] [CrossRef] [PubMed]
- Shiba, H.; Takayama, S.; Iwano, M.; Shimosato, H.; Funato, M.; Nakagawa, T.; Che, F.S.; Suzuki, G.; Watanabe, M.; Hinata, K.; et al. A pollen coat protein, SP11/SCR, determines the pollen S-specificity in the self-incompatibility of Brassica species. Plant Physiol. 2001, 125, 2095–2103. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, R.; Fobis-Loisy, I.; Gaude, T. When no means no: Guide to Brassicaceae self-incompatibility. Trends Plant Sci. 2010, 15, 387–394. [Google Scholar] [CrossRef]
- Takayama, S.; Isogai, A. Self-incompatibility in plants. Annu. Rev. Plant Biol. 2005, 56, 467–489. [Google Scholar] [CrossRef]
- Nasrallah, J.B.; Nasrallah, M.E. Robust self-incompatibility in the absence of a functional ARC1 gene in Arabidopsis thaliana. Plant Cell 2014, 26, 3838–3841. [Google Scholar] [CrossRef][Green Version]
- Iwano, M.; Ito, K.; Fujii, S.; Kakita, M.; Asano-Shimosato, H.; Igarashi, M.; Kaothien-Nakayama, P.; Entani, T.; Kanatani, A.; Takehisa, M.; et al. Calcium signalling mediates self-incompatibility response in the Brassicaceae. Nat. Plants 2015, 1, 15128. [Google Scholar] [CrossRef]
- Gu, T.S.; Mazzurco, M.; Sulaman, W.; Matias, D.D.; Goring, D.R. Binding of an arm repeat protein to the kinase domain of the S-locus receptor kinase. Proc. Natl. Acad. Sci. USA 1998, 95, 382–387. [Google Scholar] [CrossRef] [PubMed]
- Murase, K.; Shiba, H.; Iwano, M.; Che, F.S.; Watanabe, M.; Isogai, A.; Takayama, S. A membrane-anchored protein kinase involved in Brassica self-incompatibility signaling. Science 2004, 303, 1516–1519. [Google Scholar] [CrossRef]
- Cabrillac, D.; Cock, J.M.; Dumas, C.; Gaude, T. The S-locus receptor kinase is inhibited by thioredoxins and activated by pollen coat proteins. Nature 2001, 410, 220–223. [Google Scholar] [CrossRef] [PubMed]
- Cornish, M.A.; Hayward, M.D.; Lawrence, M.J. Self-incompatibility in ryegrass 3. The joint segregation of S and Pgi-2 in Lolium-perenne L. Heredity 1980, 44, 55–62. [Google Scholar] [CrossRef]
- Gertz, A.; Wricke, G. Linkage between the incompatibility locus-Z and a beta-glucosidase locus in rye. Plant Breed. 1989, 102, 255–259. [Google Scholar] [CrossRef]
- Fuong, F.T.; Voylokov, A.V.; Smirnov, V.G. Genetic-studies of self-fertility in rye (Secale-cereale L.) 2. The search for isozyme marker genes linked to self-incompatibility loci. Appl. Genet. 1993, 87, 619–623. [Google Scholar] [CrossRef] [PubMed]
- Hackauf, B.; Wehling, P. Approaching the self-incompatibility locus Z in rye (Secale cereale L.) via comparative genetics. Appl. Genet. 2005, 110, 832–845. [Google Scholar] [CrossRef]
- Bian, X.Y.; Friedrich, A.; Bai, J.R.; Baumann, U.; Hayman, D.L.; Barker, S.J.; Langridge, P. High-resolution mapping of the S and Z loci of Phalaris coerulescens. Genome 2004, 47, 918–930. [Google Scholar] [CrossRef] [PubMed]
- Thorogood, D.; Kaiser, W.J.; Jones, J.G.; Armstead, I. Self-incompatibility in ryegrass 12. Genotyping and mapping the S and Z loci of Lolium perenne L. Heredity 2002, 88, 385–390. [Google Scholar] [CrossRef] [PubMed]
- Li, X.M.; Nield, J.; Hayman, D.; Langridge, P. Cloning a putative self-incompatibility gene from the pollen of the grass Phalaris-coerulescens. Plant Cell 1994, 6, 1923–1932. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Li, X.M.; Nield, J.; Hayman, D.; Langridge, P. Thioredoxin activity in the c-terminus of Phalaris S-protein. Plant J. 1995, 8, 133–138. [Google Scholar] [CrossRef]
- Baumann, U.; Juttner, J.; Bian, X.Y.; Langridge, P. Self-incompatibility in the grasses. Ann. Bot. 2000, 85, 203–209. [Google Scholar] [CrossRef]
- Kakeda, K.; Ibuki, T.; Suzuki, J.; Tadano, H.; Kurita, Y.; Hanai, Y.; Kowyama, Y. Molecular and genetic characterization of the S locus in Hordeum bulbosum L., a wild self-incompatible species related to cultivated barley. Mol. Genet. Genom. 2008, 280, 509–519. [Google Scholar] [CrossRef]
- Kakeda, K. S locus-linked F-box genes expressed in anthers of Hordeum bulbosum. Plant Cell Rep. 2009, 28, 1453–1460. [Google Scholar] [CrossRef]
- Yang, B.C.; Thorogood, D.; Armstead, I.P.; Franklin, F.C.H.; Barth, S. Identification of genes expressed during the self-incompatibility response in perennial ryegrass (Lolium perenne L.). Plant Mol. Biol. 2009, 70, 709–723. [Google Scholar] [CrossRef]
- Shinozuka, H.; Cogan, N.O.I.; Smith, K.F.; Spangenberg, G.C.; Forster, J.W. Fine-scale comparative genetic and physical mapping supports map-based cloning strategies for the self-incompatibility loci of perennial ryegrass (Lolium perenne L.). Plant Mol. Biol. 2010, 72, 343–355. [Google Scholar] [CrossRef]
- Manzanares, C.; Barth, S.; Thorogood, D.; Byrne, S.L.; Yates, S.; Czaban, A.; Asp, T.; Yang, B.C.; Studer, B. A gene encoding a DUF247 domain protein cosegregates with the S self-incompatibility locus in perennial ryegrass. Mol. Biol. Evol. 2016, 33, 870–884. [Google Scholar] [CrossRef]
- Thomas, S.M.; Murray, B.G. A new site for self-incompatibility in the Gramineae. Incompat. Newsl. 1975, 6, 22–23. [Google Scholar]
- Wehling, P.; Hackauf, B.; Wricke, G. Phosphorylation of pollen proteins in relation to self-incompatibility in rye (Secale-cereale L.). Sex Plant Reprod. 1994, 7, 67–75. [Google Scholar] [CrossRef]
- Jones, E.S.; Mahoney, N.L.; Hayward, M.D.; Armstead, I.P.; Jones, J.G.; Humphreys, M.O.; King, I.P.; Kishida, T.; Yamada, T.; Balfourier, F.; et al. An enhanced molecular marker based genetic map of perennial ryegrass (Lolium perenne) reveals comparative relationships with other Poaceae genomes. Genome 2002, 45, 282–295. [Google Scholar] [CrossRef] [PubMed]
- Miljus-Dukic, J.; Ninkovic, S.; Neskovic, M. Effects of protein phosphatase inhibitors and calcium antagonists on self-incompatible reaction in buckwheat. Biol. Plant 2003, 46, 475–478. [Google Scholar] [CrossRef]
- Matton, D.P.; Nass, N.; Clarke, A.E.; Newbigin, E. Self-incompatibility—how plants avoid illegitimate offspring. Proc. Natl. Acad. Sci. USA 1994, 91, 1992–1997. [Google Scholar] [CrossRef]
- Igic, B.; Kohn, J.R. Evolutionary relationships among self-incompatibility RNases. Proc. Natl. Acad. Sci. USA 2001, 98, 13167–13171. [Google Scholar] [CrossRef]
- Chen, S.Y.; Huang, X.; Yan, X.Q.; Liang, Y.; Wang, Y.Z.; Li, X.F.; Peng, X.J.; Ma, X.Y.; Zhang, L.X.; Cai, Y.Y.; et al. Transcriptome analysis in sheepgrass (Leymus chinensis): A dominant perennial grass of the Eurasian Steppe. PLoS ONE 2013, 8, e67974. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.D.; Chen, S.Y.; Liu, G.S.; Jan, C.C. Seed-set and pollen-stigma compatibility in Leymus chinensis. Grass Forage Sci. 2004, 59, 180–185. [Google Scholar] [CrossRef]
- Zhou, Q.Y.; Jia, J.T.; Huang, X.; Yan, X.Q.; Cheng, L.Q.; Chen, S.Y.; Li, X.X.; Peng, X.J.; Liu, G.S. The large-scale investigation of gene expression in Leymus chinensis stigmas provides a valuable resource for understanding the mechanisms of poaceae self-incompatibility. BMC Genom. 2014, 15, 399. [Google Scholar] [CrossRef][Green Version]
- Kang, C.Y.; Darwish, O.; Geretz, A.; Shahan, R.; Alkharouf, N.; Liu, Z.C. Genome-scale transcriptomic insights into early-stage fruit development in woodland strawberry Fragaria vesca. Plant Cell 2013, 25, 1960–1978. [Google Scholar] [CrossRef]
- Knox, R.B.; Heslopha, J. Pollen-wall proteins—Fate of intined-held antigens on stigma in compatible and incompatible pollinations of Phalaris-tuberosa L. J. Cell Sci. 1971, 9, 239–251. [Google Scholar]
- Wheeler, M.J.; Franklin-Tong, V.E.; Franklin, F.C.H. The molecular and genetic basis of pollen-pistil interactions. New Phytol. 2001, 151, 565–584. [Google Scholar] [CrossRef]
- Rudd, J.J.; Franklin, F.C.H.; Lord, J.M.; FranklinTong, V.E. Increased phosphorylation of a 26-kD pollen protein is induced by the self-incompatibility response in Papaver rhoeas. Plant Cell 1996, 8, 713–724. [Google Scholar] [CrossRef]
- Rudd, J.J.; Osman, K.; Franklin, F.C.H.; Franklin-Tong, V.E. Activation of a putative MAP kinase in pollen is stimulated by the self-incompatibility (SI) response. FEBS Lett. 2003, 547, 223–227. [Google Scholar] [CrossRef]
- Takayama, S.; Shimosato, H.; Shiba, H.; Funato, M.; Che, F.S.; Watanabe, M.; Iwano, M.; Isogai, A. Direct ligand-receptor complex interaction controls Brassica self-incompatibility. Nature 2001, 413, 534–538. [Google Scholar] [CrossRef]
- Byrne, S.L.; Nagy, I.; Pfeifer, M.; Armstead, I.; Swain, S.; Studer, B.; Mayer, K.; Campbell, J.D.; Czaban, A.; Hentrup, S.; et al. A synteny-based draft genome sequence of the forage grass Lolium perenne. Plant J. 2015, 84, 816–826. [Google Scholar] [CrossRef]
- Qu, L.J.; Li, L.; Lan, Z.J.; Dresselhaus, T. Peptide signalling during the pollen tube journey and double fertilization. J. Exp. Bot. 2015, 66, 5139–5150. [Google Scholar] [CrossRef] [PubMed]
- Bircheneder, S.; Dresselhaus, T. Why cellular communication during plant reproduction is particularly mediated by CRP signalling. J. Exp. Bot. 2016, 67, 4849–4861. [Google Scholar] [CrossRef]
- Takayama, S.; Shiba, H.; Iwano, M.; Shimosato, H.; Che, F.S.; Kai, N.; Watanabe, M.; Suzuki, G.; Hinata, K.; Isogai, A. The pollen determinant of self-incompatibility in Brassica campestris. Proc. Natl. Acad. Sci. USA 2000, 97, 1920–1925. [Google Scholar] [CrossRef]
- Wu, J.Y.; Wang, S.; Gu, Y.C.; Zhang, S.L.; Publicover, S.J.; Franklin-Tong, V.E. Self-incompatibility in Papaver rhoeas activates nonspecific cation conductance permeable to Ca2+ and K+. Plant Physiol. 2011, 155, 963–973. [Google Scholar] [CrossRef]
- Yoon, G.M.; Dowd, P.E.; Gilroy, S.; McCubbin, A.G. Calcium-dependent protein kinase isoforms in Petunia have distinct functions in pollen tube growth, including regulating polarity. Plant Cell 2006, 18, 867–878. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.M.; Lan, W.Z.; Jiang, Y.Q.; Fang, W.; Luan, S. A calcium-dependent protein kinase interacts with and activates a calcium channel to regulate pollen tube growth. Mol. Plant 2014, 7, 369–376. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.T.; Gao, Y.B.; Zhou, H.S.; Chen, J.Q.; Wu, J.Y.; Zhang, S.L. Apoplastic calmodulin promotes self-incompatibility pollen tube growth by enhancing calcium influx and reactive oxygen species concentration in Pyrus pyrifolia. Plant Cell Rep. 2014, 33, 255–263. [Google Scholar] [CrossRef]
- Wilkins, K.A.; Bancroft, J.; Bosch, M.; Ings, J.; Smirnoff, N.; Franklin-Tong, V.E. Reactive oxygen species and nitric oxide mediate actin reorganization and programmed cell death in the self-incompatibility response of Papaver. Plant Physiol. 2011, 156, 404–416. [Google Scholar] [CrossRef] [PubMed]
- Serrano, I.; Romero-Puertas, M.C.; Rodriguez-Serrano, M.; Sandalio, L.M.; Olmedilla, A. Peroxynitrite mediates programmed cell death both in papillar cells and in self-incompatible pollen in the olive (Olea europaea L.). J. Exp. Bot. 2012, 63, 1479–1493. [Google Scholar] [CrossRef]
- Serrano, I.; Romero-Puertas, M.C.; Sandalio, L.M.; Olmedilla, A. The role of reactive oxygen species and nitric oxide in programmed cell death associated with self-incompatibility. J. Exp. Bot. 2015, 66, 2869–2876. [Google Scholar] [CrossRef] [PubMed]
- Chamizo-Ampudia, A.; Sanz-Luque, E.; Llamas, A.; Galvan, A.; Fernandez, E. Nitrate reductase regulates plant nitric oxide homeostasis. Trends Plant Sci. 2017, 22, 163–174. [Google Scholar] [CrossRef]
- Baker, R.P.; Hasenstein, K.H.; Zavada, M.S. Hormonal changes after compatible and incompatible pollination in Theobroma cacao L. Hortscience 1997, 32, 1231–1234. [Google Scholar] [CrossRef]
- Hasenstein, K.H.; Zavada, M.S. Auxin modification of the incompatibility response in Theobroma cacao. Physiol. Plant. 2001, 112, 113–118. [Google Scholar] [CrossRef]
- Tantikanjana, T.; Nasrallah, J.B. Non-cell-autonomous regulation of crucifer self-incompatibility by Auxin Response Factor ARF3. Proc. Natl. Acad. Sci. USA 2012, 109, 19468–19473. [Google Scholar] [CrossRef]
- Zhao, P.F.; Zhang, L.D.; Zhao, L.X. Dissection of the style’s response to pollination using transcriptome profiling in self-compatible (Solanum pimpinellifolium) and self-incompatible (Solanum chilense) tomato species. Bmc Plant Biol. 2015, 15, 119. [Google Scholar] [CrossRef]
- Rodriguez-Serrano, M.; Romero-Puertas, M.C.; Sparkes, I.; Hawes, C.; del Rio, L.A.; Sandalio, L.M. Peroxisome dynamics in Arabidopsis plants under oxidative stress induced by cadmium. Free Radic. Biol. Med. 2009, 47, 1632–1639. [Google Scholar] [CrossRef]
- Simontacchi, M.; Garcia-Mata, C.; Bartoli, C.G.; Santa-Maria, G.E.; Lamattina, L. Nitric oxide as a key component in hormone-regulated processes. Plant Cell Rep. 2013, 32, 853–866. [Google Scholar] [CrossRef]
- Hussey, P.J.; Ketelaar, T.; Deeks, M.J. Control of the actin cytoskeleton in plant cell growth. Annu. Rev. Plant Biol. 2006, 57, 109–125. [Google Scholar] [CrossRef]
- Huang, S.J.; Blanchoin, L.; Chaudhry, F.; Franklin-Tong, V.E.; Staiger, C.J. A gelsolin-like protein from Papaver rhoeas pollen (PrABP80) stimulates calcium-regulated severing and depolymerization of actin filaments. J. Biol. Chem. 2004, 279, 23364–23375. [Google Scholar] [CrossRef]
- Poulter, N.S.; Staiger, C.J.; Rappoport, J.Z.; Franklin-Tong, V.E. Actin-binding proteins implicated in the formation of the punctate actin foci stimulated by the self-incompatibility response in Papaver. Plant Physiol. 2010, 152, 1274–1283. [Google Scholar] [CrossRef]
- Eng, R.C.; Halat, L.S.; Livingston, S.J.; Sakai, T.; Motose, H.; Wasteneys, G.O. The ARM domain of ARMADILLO-REPEAT KINESIN 1 is not required for microtubule catastrophe but can negatively regulate NIMA-RELATED KINASE 6 in Arabidopsis thaliana. Plant Cell Physiol. 2017, 58, 1350–1363. [Google Scholar] [CrossRef]
- Feng, S.J.; Song, Y.L.; Shen, M.H.; Xie, S.S.; Li, W.J.; Lu, Y.; Yang, Y.H.; Ou, G.S.; Zhou, J.; Wang, F.D.; et al. Microtubule-binding protein FOR20 promotes microtubule depolymerization and cell migration. Cell Discov. 2017, 3, 17032. [Google Scholar] [CrossRef]
- Thomas, S.G.; Huang, S.J.; Li, S.T.; Staiger, C.J.; Franklin-Tong, V.E. Actin depolymerization is sufficient to induce programmed cell death in self-incompatible pollen. J. Cell Biol. 2006, 174, 221–229. [Google Scholar] [CrossRef]
- Bosch, M.; Poulter, N.S.; Vatovec, S.; Franklin-Tong, V.E. Initiation of programmed cell death in self-incompatibility: Role for cytoskeleton modifications and several caspase-Like activities. Mol. Plant 2008, 1, 879–887. [Google Scholar] [CrossRef]
- Poulter, N.S.; Vatovec, S.; Franklin-Tong, V.E. Microtubules are a target for self-incompatibility signaling in Papaver pollen. Plant Physiol. 2008, 146, 1358–1367. [Google Scholar] [CrossRef]
- Thomas, S.G.; Franklin-Tong, V.E. Self-incompatibility triggers programmed cell death in Papaver pollen. Nature 2004, 429, 305–309. [Google Scholar] [CrossRef] [PubMed]
- Bosch, M.; Franklin-Tong, V.E. Temporal and spatial activation of caspase-like enzymes induced by self-incompatibility in Papaver pollen. Proc. Natl. Acad. Sci. USA 2007, 104, 18327–18332. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.D.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
- Benjamini, Y.; Yekutieli, D. The control of the false discovery rate in multiple testing under dependency. Ann. Stat. 2001, 29, 1165–1188. [Google Scholar]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef]
- Xie, C.; Mao, X.Z.; Huang, J.J.; Ding, Y.; Wu, J.M.; Dong, S.; Kong, L.; Gao, G.; Li, C.Y.; Wei, L.P. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, W316–W322. [Google Scholar] [CrossRef]
- Wu, F.X.; Zhang, W.J.; Kusalik, A.J. A genetic K-means clustering algorithm applied to gene expression data. Lect. Notes Artif. Int. 2003, 2671, 520–526. [Google Scholar]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef] [PubMed]
| Metric | Value |
|---|---|
| General assembly statistics | |
| Total transcripts | 399,224 |
| Smallest transcript length | 201 |
| Largest transcript length | 13,765 |
| Transcript N50 | 1307 |
| Transcript N90 | 331 |
| Median transcript length | 507 |
| Average transcript length | 820 |
| Total transcript assembled bases | 327,421,926 |
| Statistics based on the longest transcript per gene | |
| Total unigenes | 220,800 |
| Unigene N50 | 901 |
| Unigene N90 | 255 |
| Median unigene length | 352 |
| Average unigene length | 614 |
| Total unigene assembled bases | 135,549,030 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, S.; Jia, J.; Cheng, L.; Zhao, P.; Qi, D.; Yang, W.; Liu, H.; Dong, X.; Li, X.; Liu, G. Transcriptomic Analysis Reveals a Comprehensive Calcium- and Phytohormone-Dominated Signaling Response in Leymus chinensis Self-Incompatibility. Int. J. Mol. Sci. 2019, 20, 2356. https://doi.org/10.3390/ijms20092356
Chen S, Jia J, Cheng L, Zhao P, Qi D, Yang W, Liu H, Dong X, Li X, Liu G. Transcriptomic Analysis Reveals a Comprehensive Calcium- and Phytohormone-Dominated Signaling Response in Leymus chinensis Self-Incompatibility. International Journal of Molecular Sciences. 2019; 20(9):2356. https://doi.org/10.3390/ijms20092356
Chicago/Turabian StyleChen, Shuangyan, Junting Jia, Liqin Cheng, Pincang Zhao, Dongmei Qi, Weiguang Yang, Hui Liu, Xiaobing Dong, Xiaoxia Li, and Gongshe Liu. 2019. "Transcriptomic Analysis Reveals a Comprehensive Calcium- and Phytohormone-Dominated Signaling Response in Leymus chinensis Self-Incompatibility" International Journal of Molecular Sciences 20, no. 9: 2356. https://doi.org/10.3390/ijms20092356
APA StyleChen, S., Jia, J., Cheng, L., Zhao, P., Qi, D., Yang, W., Liu, H., Dong, X., Li, X., & Liu, G. (2019). Transcriptomic Analysis Reveals a Comprehensive Calcium- and Phytohormone-Dominated Signaling Response in Leymus chinensis Self-Incompatibility. International Journal of Molecular Sciences, 20(9), 2356. https://doi.org/10.3390/ijms20092356






