ChimeraMiner: An Improved Chimeric Read Detection Pipeline and Its Application in Single Cell Sequencing
Abstract
1. Introduction
2. Results
2.1. Data Downloaded and Initial Processing
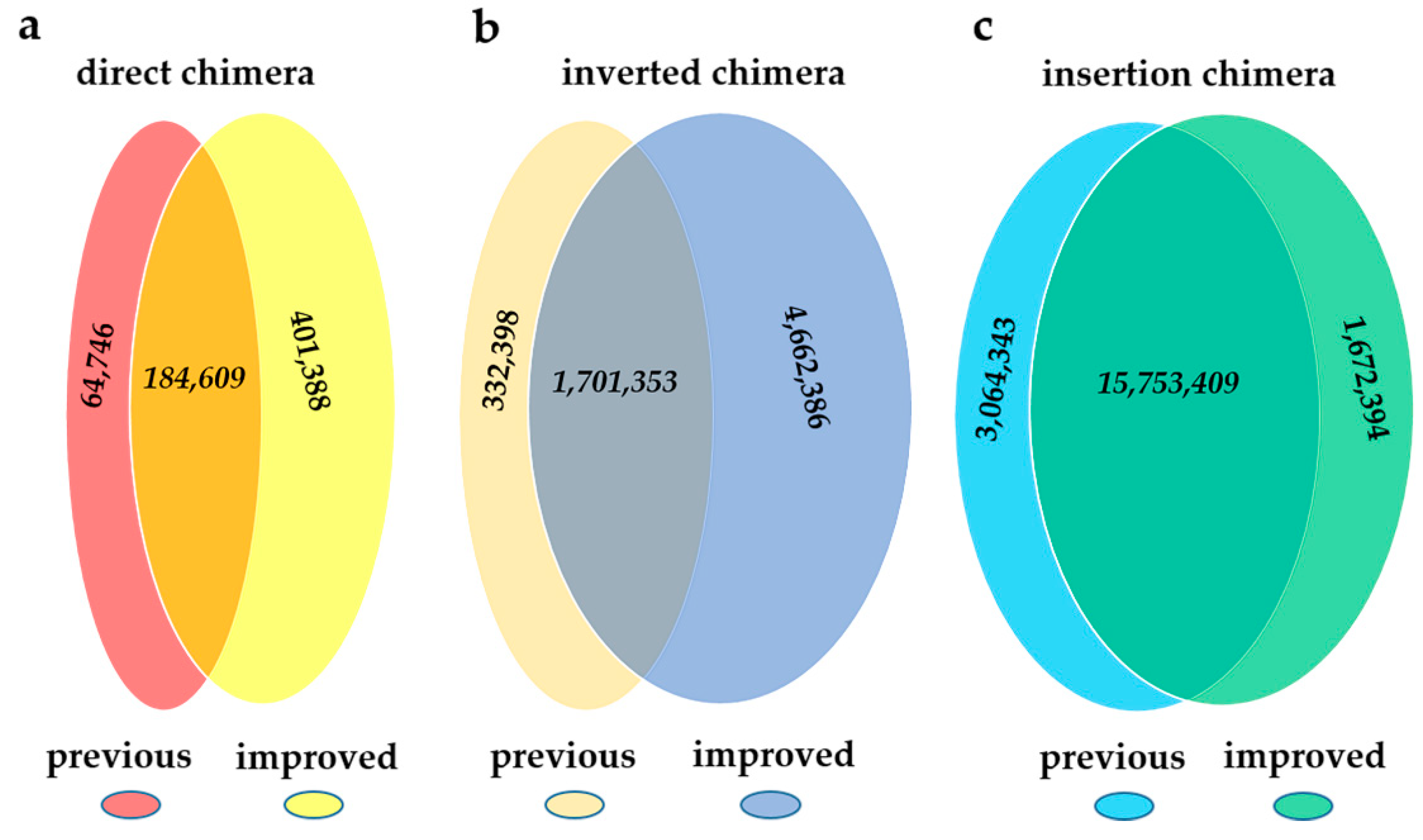
2.2. Comparison between the Previous Pipeline and ChimeraMiner
2.3. The Performance in Single-Cell MDA Sequencing Data
2.4. The Impact of Chimeras on Structural Variation
3. Discussion
4. Materials and Methods
4.1. Data Sources
4.2. Bioinformatics Analysis for Chimera Detecting
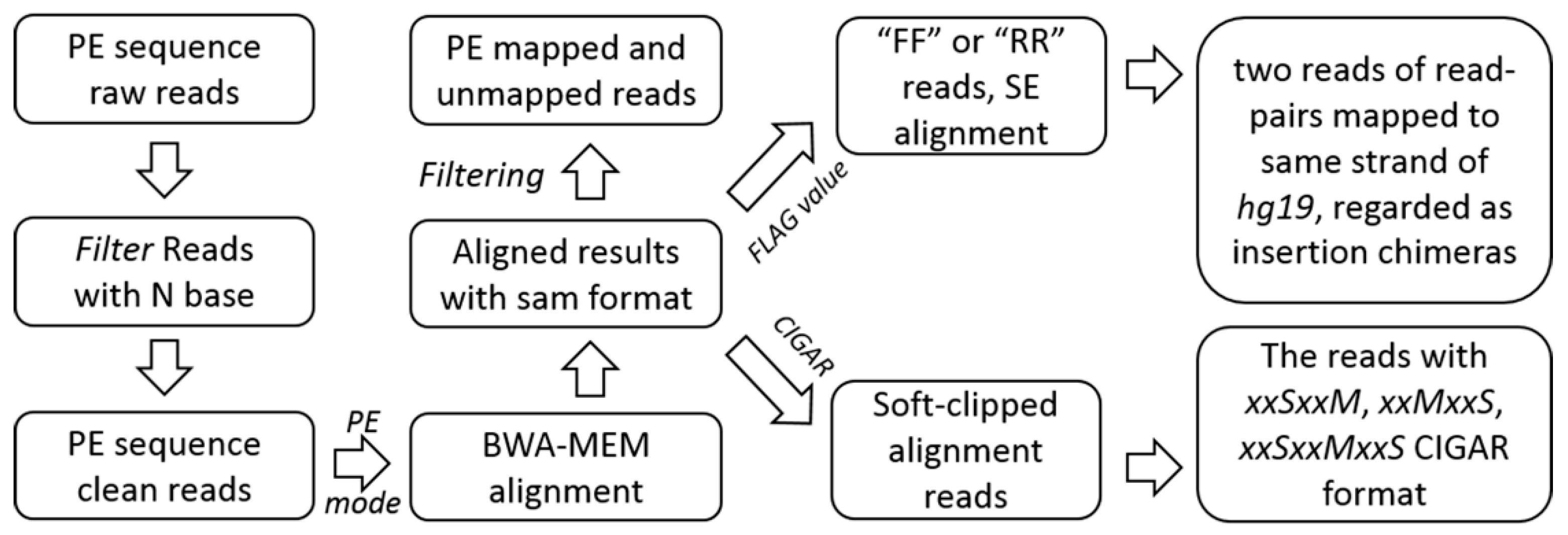
4.2.1. Pre-Processing, Alignment, and Filtrating
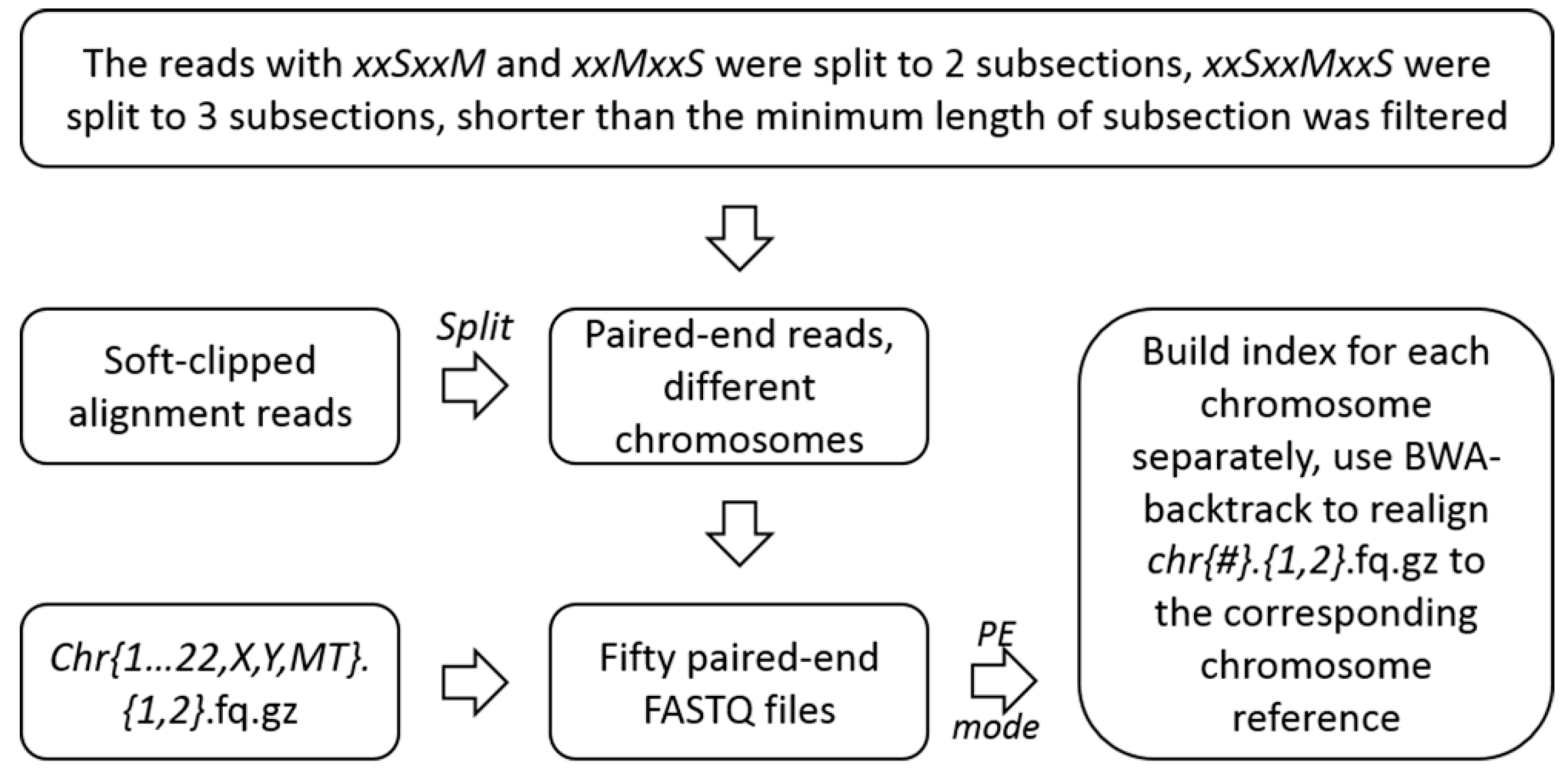
4.2.2. Unscrambling of the Soft-Clipped Alignment Reads and Realignment
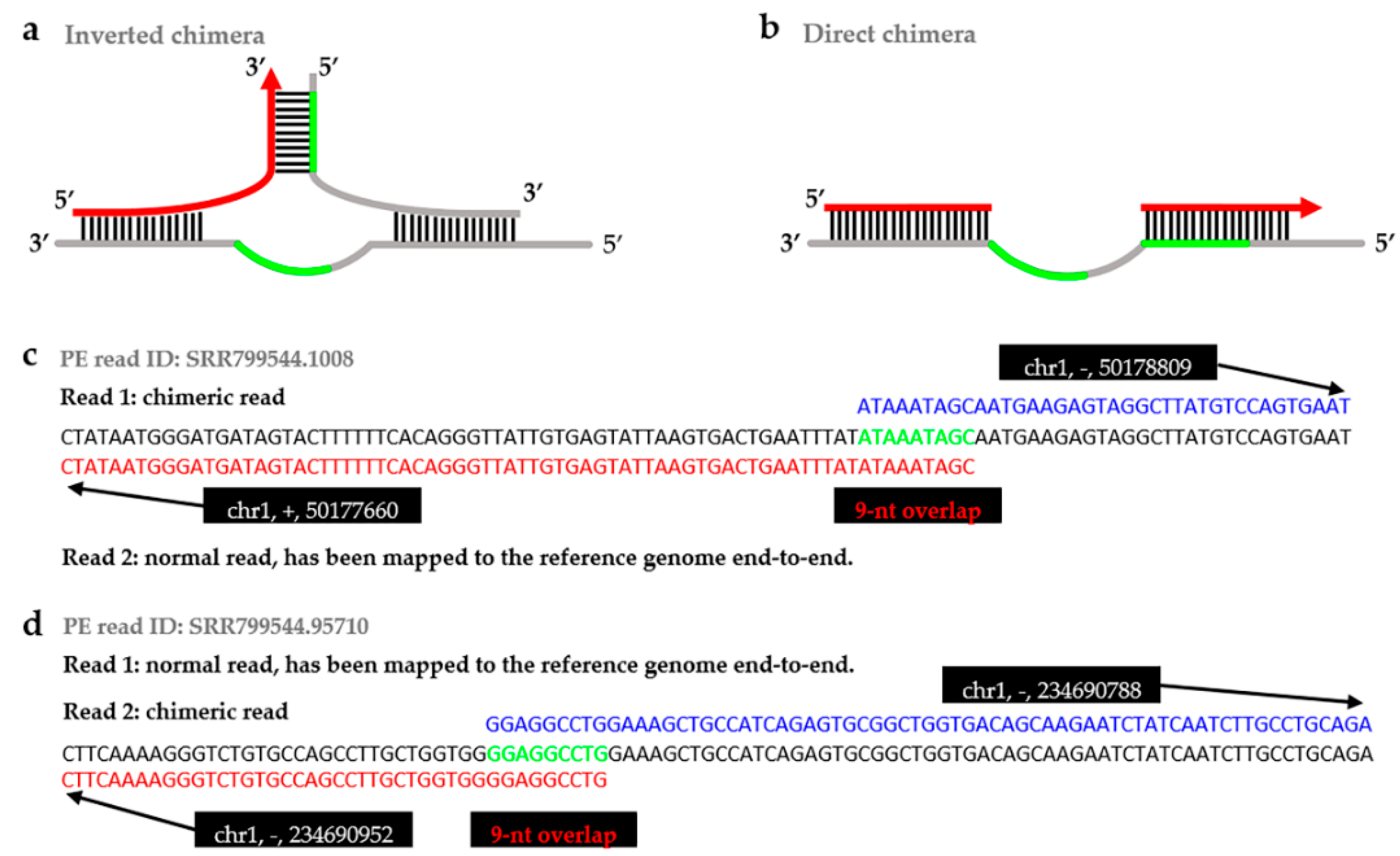
4.2.3. Searching Overlap and Achieving Valid Chimeras
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| MDA | Multiple displacement amplification |
| WGA | Whole genome amplification |
| Phi29DNApol | Phi29 DNA polymerase |
| Chimeras | Chimeric sequence reads |
| SOAP2 | Short Oligonucleotide Analysis Package 2 |
| BWA | Burrows-Wheeler Aligner |
| SCMDA | Single cell multiple displacement amplification |
| Hg19 | Human reference genome Feb. 2009 assembly |
| SRA | Sequence Read Archive |
| UCSC | University of California, Santa Cruz |
| NCBI | National Center for Biotechnology Information |
| B-Lymphocyte | Peripheral vein B-Lymphocyte |
| BJ, ATCC | BJ primary human foreskin fibroblast |
| HUVEC | Human umbilical vein endothelial cell |
| YH | YH lymphoblastoid cell line |
| BWA-MEM | Burrows-Wheeler Aligner-Maximal Exact Matches |
| SVs | Structural Variations |
References
- Dean, F.B.; Hosono, S.; Fang, L.H.; Wu, X.H.; Faruqi, A.F.; Bray-Ward, P.; Sun, Z.Y.; Zong, Q.L.; et al. Comprehensive human genome amplification using multiple displacement amplification. Proc. Natl. Acad. Sci. USA 2002, 99, 5261–5266. [Google Scholar] [CrossRef] [PubMed]
- Dean, F.B.; Nelson, J.R.; Giesler, T.L.; Lasken, R.S. Rapid amplification of plasmid and phage DNA using phi29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 2001, 11, 1095–1099. [Google Scholar] [CrossRef]
- Zhang, D.Y.; Brandwein, M.; Hsuih, T.; Li, H.B. Ramification amplification: A novel isothermal DNA amplification method. Mol. Diagn. 2001, 6, 141–150. [Google Scholar] [CrossRef] [PubMed]
- Garmendia, C.; Bernad, A.; Esteban, J.A.; Blanco, L.; Salas, M. The Bacteriophage-Phi-29 DNA-Polymerase, a Proofreading Enzyme. J. Biol. Chem. 1992, 267, 2594–2599. [Google Scholar] [PubMed]
- Zhang, K.; Martiny, A.C.; Reppas, N.B.; Barry, K.W.; Malek, J.; Chisholm, S.W.; Church, G.M. Sequencing genomes from single cells by polymerase cloning. Nat. Biotechnol. 2006, 24, 680–686. [Google Scholar] [CrossRef]
- Lasken, R.S.; Stockwell, T.B. Mechanism of chimera formation during the Multiple Displacement Amplification reaction. BMC Biotechnol. 2007, 7, 19. [Google Scholar] [CrossRef]
- Marcy, Y.; Ishoey, T.; Lasken, R.S.; Stockwell, T.B.; Walenz, B.P.; Halpern, A.L.; Beeson, K.Y.; Goldberg, S.M.; Quake, S.R. Nanoliter reactors improve multiple displacement amplification of genomes from single cells. PLoS Genet. 2007, 3, 1702–1708. [Google Scholar] [CrossRef]
- Tu, J.; Guo, J.; Li, J.J.; Gao, S.; Yao, B.; Lu, Z.H. Systematic Characteristic Exploration of the Chimeras Generated in Multiple Displacement Amplification through Next Generation Sequencing Data Reanalysis. PLoS ONE 2015, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Tu, J.; Lu, N.; Duan, M.Q.; Huang, M.T.; Chen, L.; Li, J.J.; Guo, J.; Lu, Z.H. Hotspot Selective Preference of the Chimeric Sequences Formed in Multiple Displacement Amplification. Int. J. Mol. Sci. 2017, 18, 492. [Google Scholar] [CrossRef]
- Chitsaz, H.; Yee-Greenbaum, J.L.; Tesler, G.; Lombardo, M.J.; Dupont, C.L.; Badger, J.H.; Novotny, M.; Rusch, D.B.; Fraser, L.J.; Gormley, N.A.; et al. Efficient de novo assembly of single-cell bacterial genomes from short-read data sets. Nat. Biotechnol. 2011, 29, 915. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Navin, N.E. Advances and Applications of Single-Cell Sequencing Technologies. Mol/ Cell 2015, 58, 598–609. [Google Scholar] [CrossRef] [PubMed]
- Abel, H.J.; Duncavage, E.J.; Becker, N.; Armstrong, J.R.; Magrini, V.J.; Pfeifer, J.D. SLOPE: A quick and accurate method for locating non-SNP structural variation from targeted next-generation sequence data. Bioinformatics 2010, 26, 2684–2688. [Google Scholar] [CrossRef] [PubMed]
- Voet, T.; Kumar, P.; Van Loo, P.; Cooke, S.L.; Marshall, J.; Lin, M.L.; Zamani Esteki, M.; Van der Aa, N.; Mateiu, L.; McBride, D.J.; et al. Single-cell paired-end genome sequencing reveals structural variation per cell cycle. Nucleic Acids Res/ 2013, 41, 6119–6138. [Google Scholar] [CrossRef] [PubMed]
- Van Loo, P.; Voet, T. Single cell analysis of cancer genomes. Curr. Opin. Genet. Dev. 2014, 24, 82–91. [Google Scholar] [CrossRef]
- Hou, Y.; Wu, K.; Shi, X.L.; Li, F.Q.; Song, L.T.; Wu, H.J.; Dean, M.; Li, G.B.; Tsang, S.; Jiang, R.Z.; et al. Comparison of variations detection between whole-genome amplification methods used in single-cell resequencing. Gigascience 2015, 4, 37. [Google Scholar] [CrossRef]
- Davis, C.F.; Ritter, D.I.; Wheeler, D.A.; Wang, H.; Ding, Y.; Dugan, S.P.; Bainbridge, M.N.; Muzny, D.M.; Rao, P.H.; Man, T.K.; et al. SV-STAT accurately detects structural variation via alignment to reference-based assemblies. Source Code Biol. Med. 2016, 11, 8. [Google Scholar] [CrossRef]
- Guan, P.; Sung, W.K. Structural variation detection using next-generation sequencing data A comparative technical review. Methods 2016, 102, 36–49. [Google Scholar] [CrossRef]
- Rodrigue, S.; Malmstrom, R.R.; Berlin, A.M.; Birren, B.W.; Henn, M.R.; Chisholm, S.W. Whole Genome Amplification and De novo Assembly of Single Bacterial Cells. PLoS ONE 2009, 4, 9. [Google Scholar] [CrossRef] [PubMed]
- Nurk, S.; Bankevich, A.; Antipov, D.; Gurevich, A.A.; Korobeynikov, A.; Lapidus, A.; Prjibelski, A.D.; Pyshkin, A.; Sirotkin, A.; Sirotkin, Y.; et al. Assembling Single-Cell Genomes and Mini-Metagenomes From Chimeric MDA Products. J. Comput. Biol. 2013, 20, 714–737. [Google Scholar] [CrossRef] [PubMed]
- Hosokawa, M.; Nishikawa, Y.; Kogawa, M.; Takeyama, H. Massively parallel whole genome amplification for single-cell sequencing using droplet microfluidics. Sci. Rep. 2017, 7, 5199. [Google Scholar] [CrossRef] [PubMed]
- Kogawa, M.; Hosokawa, M.; Nishikawa, Y.; Mori, K.; Takeyama, H. Obtaining high-quality draft genomes from uncultured microbes by cleaning and co-assembly of single-cell amplified genomes. Sci. Rep. 2018, 8, 2059. [Google Scholar] [CrossRef] [PubMed]
- Peccoud, J.; Lequime, S.; Moltini-Conclois, I.; Giraud, I.; Lambrechts, L.; Gilbert, C. A Survey of Virus Recombination Uncovers Canonical Features of Artificial Chimeras Generated During Deep Sequencing Library Preparation. G3 Genes Genom Genet. 2018, 8, 1129–1138. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Ma, F.; Chapman, A.; Lu, S.J.; Xie, X.S. Single-Cell Whole-Genome Amplification and Sequencing: Methodology and Applications. Annu. Rev. Genom. Hum. G 2015, 16, 79–102. [Google Scholar] [CrossRef]
- Gawad, C.; Koh, W.; Quake, S.R. Single-cell genome sequencing: Current state of the science. Nat. Rev. Genet. 2016, 17, 175–188. [Google Scholar] [CrossRef] [PubMed]
- Jiao, X.; Rosenlund, M.; Hooper, S.D.; Tellgren-Roth, C.; He, L.Q.; Fu, Y.T.; Mangion, J.; Sjoblom, T. Structural Alterations from Multiple Displacement Amplification of a Human Genome Revealed by Mate-Pair Sequencing. PLoS ONE 2011, 6, 7. [Google Scholar] [CrossRef] [PubMed]
- Kaper, F.; Swamy, S.; Klotzle, B.; Munchel, S.; Cottrell, J.; Bibikova, M.; Chuang, H.Y.; Kruglyak, S.; Ronaghi, M.; Eberle, M.A.; et al. Whole-genome haplotyping by dilution, amplification, and sequencing. Proc. Natl. Acad. Sci. USA 2013, 110, 5552–5557. [Google Scholar] [CrossRef] [PubMed]
- Li, R.Q.; Li, Y.R.; Kristiansen, K.; Wang, J. SOAP: Short oligonucleotide alignment program. Bioinformatics 2008, 24, 713–714. [Google Scholar] [CrossRef]
- Li, R.Q.; Yu, C.; Li, Y.R.; Lam, T.W.; Yiu, S.M.; Kristiansen, K.; Wang, J. SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 2009, 25, 1966–1967. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv, 2013; arXiv:1303.3997. [Google Scholar]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Layer, R.M.; Chiang, C.; Quinlan, A.R.; Hall, I.M. LUMPY: A probabilistic framework for structural variant discovery. Genome Biol. 2014, 15, R84. [Google Scholar] [CrossRef] [PubMed]
- Chiang, C.; Layer, R.M.; Faust, G.G.; Lindberg, M.R.; Rose, D.B.; Garrison, E.P.; Marth, G.T.; Quinlan, A.R.; Hall, I.M. SpeedSeq: Ultra-fast personal genome analysis and interpretation. Nat. Methods 2015, 12, 966–968. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Zhu, F.; Vongsangnak, W.; Tang, Y.F.; Zhang, W.Y.; Shen, B.R. Evaluation and Comparison of Multiple Aligners for Next-Generation Sequencing Data Analysis. BioMed Res. Int. 2014, 2014, 309650. [Google Scholar] [CrossRef]
- Choi, J.; Park, K.; Cho, S.B.; Chung, M. HIA: A genome mapper using hybrid index-based sequence alignment. Algorithm Mol. Biol. 2015, 10. [Google Scholar] [CrossRef]
- Ziemann, M. Accuracy, speed and error tolerance of short DNA sequence aligners. bioRxiv 2016. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Genome Project Data Processing, S. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar]
- Kent, W.J.; Sugnet, C.W.; Furey, T.S.; Roskin, K.M.; Pringle, T.H.; Zahler, A.M.; Haussler, D. The human genome browser at UCSC. Genome Res. 2002, 12, 996–1006. [Google Scholar] [CrossRef]
- Cock, P.J.A.; Fields, C.J.; Goto, N.; Heuer, M.L.; Rice, P.M. The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic Acids Res. 2010, 38, 1767–1771. [Google Scholar] [CrossRef]
- Fu, Y.S.; Li, C.M.; Lu, S.J.; Zhou, W.X.; Tang, F.C.; Xie, X.S.; Huang, Y.Y. Uniform and accurate single-cell sequencing based on emulsion whole-genome amplification. Proc. Natl. Acad. Sci. USA 2015, 112, 11923–11928. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.; Song, L.T.; Zhu, P.; Zhang, B.; Tao, Y.; Xu, X.; Li, F.Q.; Wu, K.; Liang, J.; Shao, D.; et al. Single-Cell Exome Sequencing and Monoclonal Evolution of a JAK2-Negative Myeloproliferative Neoplasm. Cell 2012, 148, 873–885. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.Y.; Xing, D.; Tan, L.Z.; Li, H.; Zhou, G.Y.; Huang, L.; Xie, X.S. Single-cell whole-genome analyses by Linear Amplification via Transposon Insertion (LIANTI). Science 2017, 356, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, R.L.; Foy, B.D.; Phoenix, T. Learning Perl, 6th ed.; O’Reilly: Beijing, China; Sebastopol, CA, USA, 2011; 363p. [Google Scholar]
| Sample | Pipelines | Times/h | Direct 1 | Inverted 2 | Insertion 3 | Chimeric Rate 4 (%) |
|---|---|---|---|---|---|---|
| MDA1 | previous | 137 | 249,355 | 2,033,751 | 18,817,752 | 5.46 |
| ChimeraMiner | 60 | 585,997 | 6,363,739 | 17,425,803 | 6.31 | |
| MDA2 | previous | 114 | 203,343 | 1,534,870 | 11,194,470 | 4.90 |
| ChimeraMiner | 49 | 521,820 | 5,376,608 | 11,582,554 | 6.62 |
| Protocol | SampleID | Direct | Inverted | Insertion | Chimeric Rate (%) |
|---|---|---|---|---|---|
| MDA 1 | MDA1 | 585,997 | 6,363,739 | 17,425,803 | 6.31 |
| MDA | MDA2 | 521,820 | 5,376,608 | 11,582,554 | 6.62 |
| scMDA 2 | BGIYH1 | 240,456 | 1,763,085 | 3,334,791 | 2.37 |
| scMDA | BGIYH2 | 64,154 | 598,660 | 1,600,853 | 1.42 |
| scMDA | HUMDA | 2,708,326 | 4,559,919 | 4,397,551 | 3.24 |
| scMDA | Qiagen1 | 167,175 | 2,206,328 | 157,692 | 0.93 |
| scMDA | Qiagen5 | 227,852 | 2,867,365 | 163,770 | 1.17 |
| scMDA | Qiagen9 | 224,273 | 2,993,157 | 234,983 | 1.16 |
| scMDA | GE2 | 3,336,137 | 10,315,253 | 310,998 | 4.38 |
| scMDA | GE4 | 5,112,524 | 12,681,862 | 323,529 | 4.24 |
| scMDA | GE10 | 4,844,590 | 8,899,910 | 387,391 | 4.68 |
| SVTypes | Before Filter 2 | After Filter 3 | Bulk |
|---|---|---|---|
| BND 1 | 172,238 | 16,416 | 3,462 |
| DEL | 1,611 | 1,433 | 2,924 |
| DUP | 28,921 | 20,300 | 848 |
| INV | 593,858 | 90,726 | 55 |
| SRA Number | Sample ID | Cell Line | Protocol | Reads’ Type |
|---|---|---|---|---|
| Not-released | Bulk | B-Lymphocyte 1 | unamplified | 2 × 150 bp |
| SRX247249 | MDA1 | phi29 DNA polymerase | 2 × 101 bp | |
| SRX252522 | MDA2 | |||
| SRR1777307&8 | HUMDA | HUVEC 2 | phi29 DNA polymerase | 2 × 101 bp |
| SRS294760 | BGIYH1 | YH 3 | REPLI-g Mini Kit | 2 × 100 bp |
| SRS294759 | BGIYH2 | |||
| SRR5365373 | Qiagen1 | BJ, ATCC 4 | REPLI-g Single Cell Kit | 2 × 150 bp |
| SRR5365372 | Qiagen5 | |||
| SRR5365371 | Qiagen9 | |||
| SRR5365364 | GE2 | BJ, ATCC 4 | illustra Single Cell GenomiPhiDNA Amplification Kit | 2 × 150 bp |
| SRR5365363 | GE4 | |||
| SRR5365362 | GE10 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, N.; Li, J.; Bi, C.; Guo, J.; Tao, Y.; Luan, K.; Tu, J.; Lu, Z. ChimeraMiner: An Improved Chimeric Read Detection Pipeline and Its Application in Single Cell Sequencing. Int. J. Mol. Sci. 2019, 20, 1953. https://doi.org/10.3390/ijms20081953
Lu N, Li J, Bi C, Guo J, Tao Y, Luan K, Tu J, Lu Z. ChimeraMiner: An Improved Chimeric Read Detection Pipeline and Its Application in Single Cell Sequencing. International Journal of Molecular Sciences. 2019; 20(8):1953. https://doi.org/10.3390/ijms20081953
Chicago/Turabian StyleLu, Na, Junji Li, Changwei Bi, Jing Guo, Yuhan Tao, Kaihao Luan, Jing Tu, and Zuhong Lu. 2019. "ChimeraMiner: An Improved Chimeric Read Detection Pipeline and Its Application in Single Cell Sequencing" International Journal of Molecular Sciences 20, no. 8: 1953. https://doi.org/10.3390/ijms20081953
APA StyleLu, N., Li, J., Bi, C., Guo, J., Tao, Y., Luan, K., Tu, J., & Lu, Z. (2019). ChimeraMiner: An Improved Chimeric Read Detection Pipeline and Its Application in Single Cell Sequencing. International Journal of Molecular Sciences, 20(8), 1953. https://doi.org/10.3390/ijms20081953






