A Loop-Mediated Isothermal Amplification Assay for Rapid Detection of Pectobacterium aroidearum that Causes Soft Rot in Konjac
Abstract
1. Introduction
2. Results
2.1. Isolation and Pathogenicity of Konjac Soft Rot Pathogen
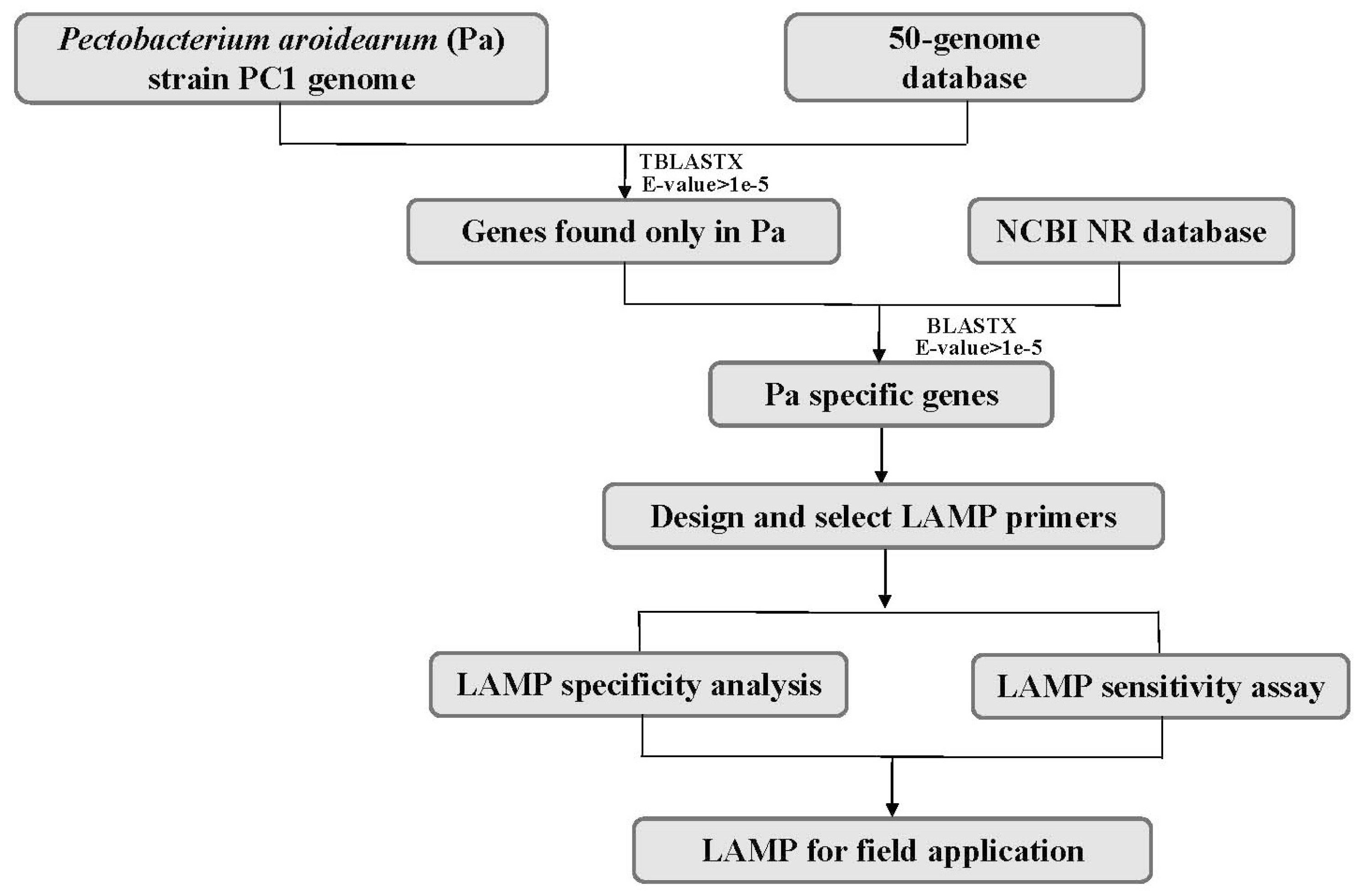
2.2. Mining of Specific Genes in the P. aroidearum Genome for LAMP Primer Design
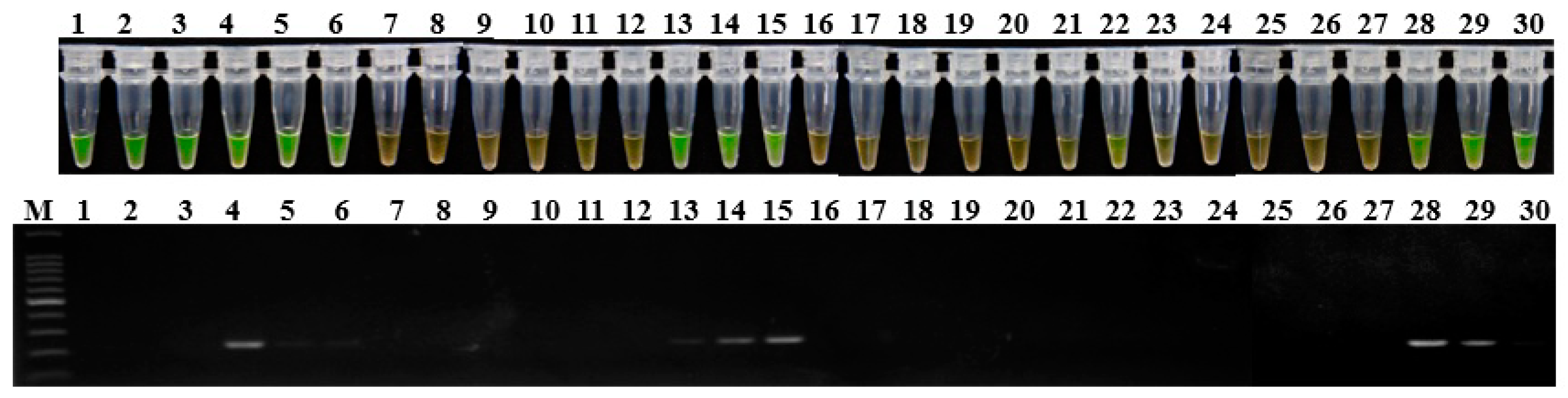
2.3. Specificity of LAMP Assay
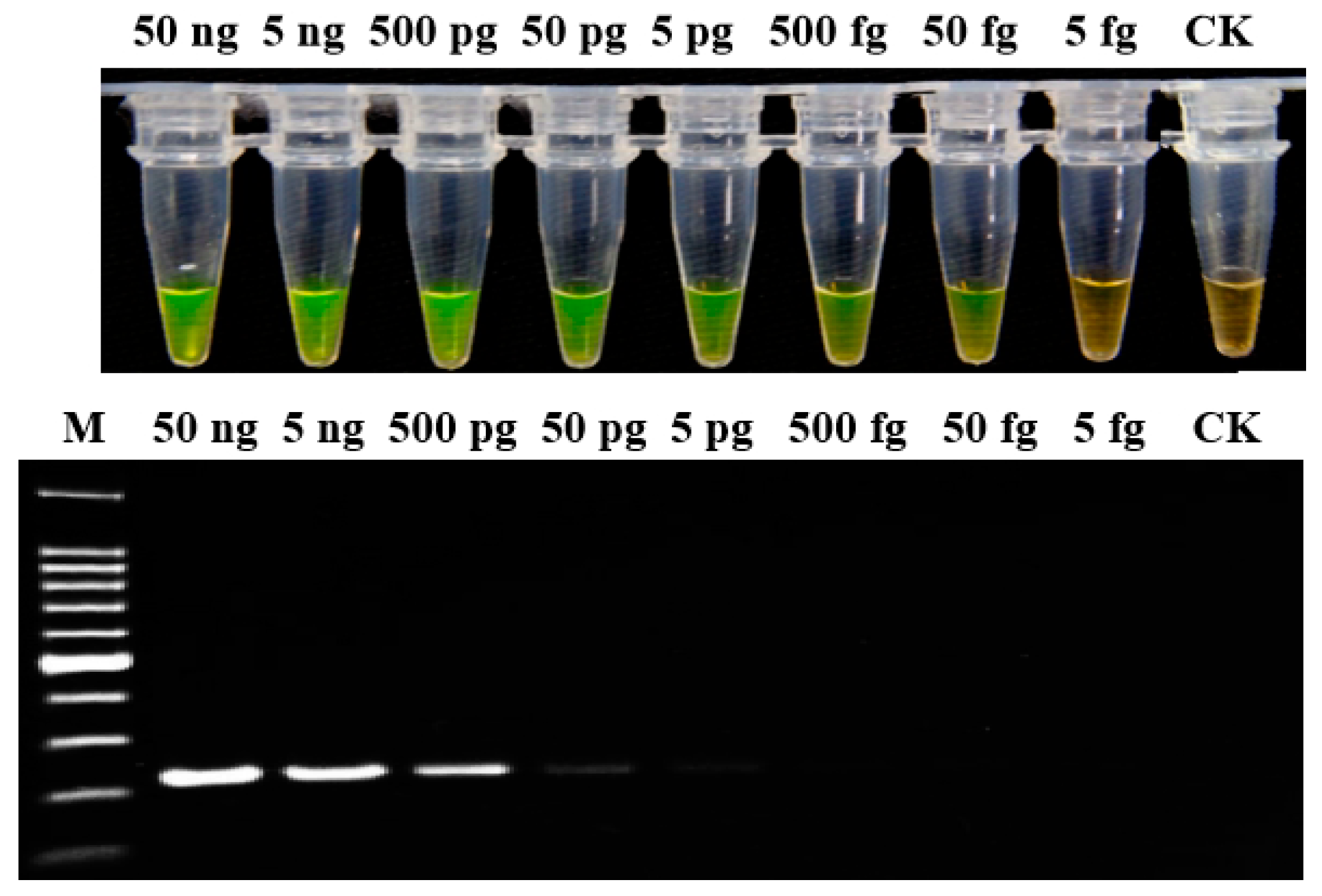
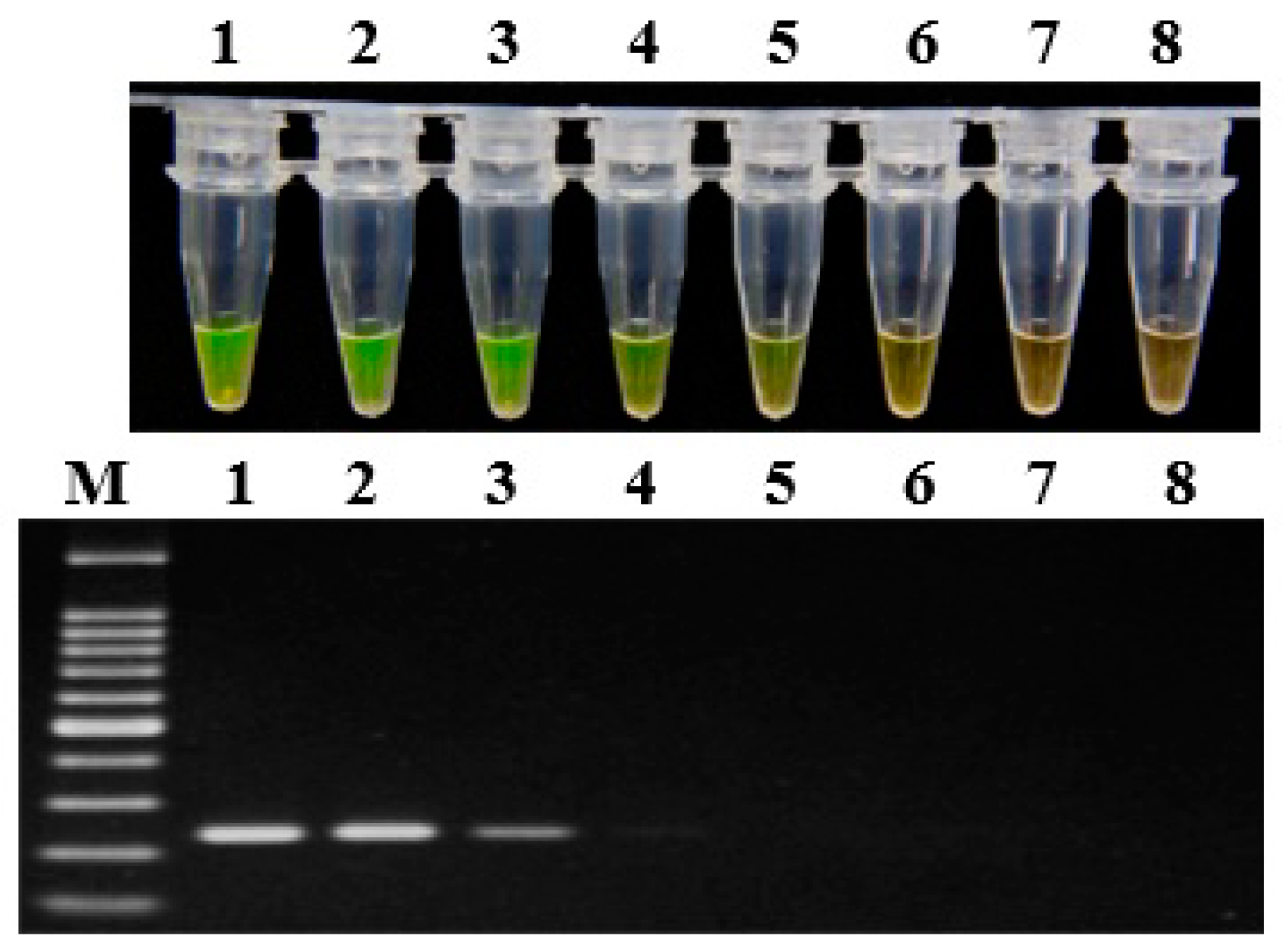
2.4. Sensitivity of LAMP Assay
2.5. LAMP Detection of Artificially Inoculated Soil Samples
2.6. Application of LAMP to Konjac Plants and Rhizosphere Soil in the Field
3. Discussion
4. Materials and Methods
4.1. Isolation and Pathogenicity of Konjac Soft Rot Pathogen
4.2. Bacterial Strains and DNA Preparation
4.3. Soil Samples and DNA Preparation
4.4. Sources of Genome Sequence Data
4.5. Mining of Specific Targets of P. Aroidearum
4.6. Design and Screening of LAMP Primers
4.7. LAMP Reaction and Product Validation
4.8. Conventional PCR Assay
4.9. LAMP Specificity and Sensitivity Assays
4.10. Application of LAMP Assay in Naturally Infected Konjac and Field Rhizosphere Soil
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Vasques, C.A.; Rossetto, S.; Halmenschlager, G.; Linden, R.; Heckler, E.; Fernandez, M.S.; Alonso, J.L. Evaluation of the pharmacotherapeutic efficacy of Garcinia cambogia plus Amorphophallus konjac for the treatment of obesity. Phytother. Res. 2008, 22, 1135–1140. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Wei, W.G.; Shao, W.; Chen, B.L.; Huang, Y.P. Lead and other elemental content of normal and Erwinia carotovora infected Amorphallus konjac corms. Bull. Environ. Contam. Toxicol. 2010, 84, 744–748. [Google Scholar] [CrossRef] [PubMed]
- Chua, M.; Baldwin, T.C.; Hocking, T.J.; Chan, K. Traditional uses and potential health benefits of Amorphophallus konjac K. Koch ex N.E.Br. J. Ethnopharmacol. 2010, 128, 268–278. [Google Scholar] [CrossRef]
- Richard, T. konjac flour properties and applications. Food Technol. 1991, 45, 82–92. [Google Scholar]
- Wu, J.P.; Jiao, Z.B.; Chen, L.F.; Jiao, Z.J.; Qiu, Z.M. The factors affecting adsorption and ingression of bacterial soft rot to konjac root. Hubei Agric. Sci. 2014, 53, 5734–5737. [Google Scholar]
- Wu, J.P.; Yang, C.Z.; Jiao, Z.B.; Hu, Z.L.; Ding, Z.L.; Qiu, Z.M. Genetic relationships of soft rot bacteria isolated from konjac in China by amplified fragment length polymorphism (AFLP) and 16S rDNA gene sequences. Agric. Sci. 2015, 6, 717–723. [Google Scholar] [CrossRef]
- Wu, J.P.; Diao, Y.; Gu, Y.C.; Hu, Z.L. Molecular detection of Pectobacterium species causing soft rot of Amorphophallus konjac. World J. Microbiol. Biotechnol. 2011, 27, 613–618. [Google Scholar] [CrossRef]
- Nabhan, S. Taxonomic relationships among species of Pectobacterium and description of two novel taxa within the heterogeneous species P. carotovorum, Namely P. carotovorum subsp. brasiliense subsp. nov. and P. aroideae sp. nov. Ph.D. Thesis, Gottfried Wilhem Leibniz Universität Hannover, Hannover, Germany, December 2011. [Google Scholar]
- Nabhan, S.; De Boer, S.H.; Maiss, E.; Wydra, K. Pectobacterium aroidearum sp. nov., a soft rot pathogen with preference for monocotyledonous plants. Int. J. Syst. Evol. Microbiol. 2013, 63, 2520–2525. [Google Scholar] [CrossRef]
- Xu, W.; Wang, T.T.; Liu, L.P.; Wang, X.B.; Wei, G.H. Isolation and identification of Amorphophallus bacterial soft rot in Langao county, Shaanxi Province. J. Northwest A&F Univ. (Nat. Sci. Ed.) 2011, 39, 97–102. [Google Scholar]
- Ma, B.; Hibbing, M.E.; Kim, H.S.; Reedy, R.M.; Yedidia, I.; Breuer, J.; Breuer, J.; Glasner, J.M.; Perna, N.T.; Kelman, A.; et al. Host range and molecular phylogenies of the soft rot enterobacterial genera Pectobacterium and Dickeya. Phytopathology 2007, 97, 1150–1163. [Google Scholar] [CrossRef]
- Cuppels, D.; Kelman, A. Evaluation of selective media for isolation of soft-rot bacteria from soil and plant tissue. Phytopathology 1974, 64, 468–475. [Google Scholar] [CrossRef]
- Potrykus, M.; Sledz, W.; Golanowska, M.; Slawiak, M.; Binek, A.; Motyka, A.; Zoledowska, S.; Czajkowski, R.; Lojkowska, E. Simultaneous detection of major blackleg and soft rot bacterial pathogens in potato by multiplex polymerase chain reaction. Ann. Appl. Biol. 2014, 165, 474–487. [Google Scholar] [CrossRef]
- Czajkowski, R.; Pérombelon, M.C.M.; Jafra, S.; Lojkowska, E.; Potrykus, M.; van der Wolf, J.M.; Sledz, W. Detection, identification and differentiation of Pectobacterium and Dickeya species causing potato blackleg and tuber soft rot: A review. Ann. Appl. Biol. 2015, 166, 18–38. [Google Scholar] [CrossRef]
- Darrasse, A.; Priou, S.; Kotoujansky, A.; Bertheau, Y. PCR and restriction fragment length polymorphism of a pel gene as a tool to identify Erwinia carotovora in relation to potato diseases. Appl. Environ. Microbiol. 1994, 60, 1437–1443. [Google Scholar]
- Li, X.; Nie, J.; Ward, L.J.; Nickerson, J.; and De Boer, S.H. Development and evaluation of a loop-mediated isothermal amplification assay for rapid detection and identification of Pectobacterium atrosepticum. Can. J. Plant Pathol. 2011, 33, 447–457. [Google Scholar] [CrossRef]
- Duarte, V.; De Boer, S.H.; Ward, L.J.; De Oliveira, A.M.R. Characterization of atypical Erwinia carotovora strains causing blackleg of potato in Brazil. J. Appl. Microbiol. 2004, 9, 535–545. [Google Scholar] [CrossRef]
- Park, D.S.; Shim, J.K.; Kim, J.S.; Kim, B.Y.; Kang, M.J.; Seol, Y.J.; Hahn, J.H.; Sherstha, R.; Lim, C.K.; Go, S.J.; et al. PCR-based sensitive and specific detection of Pectobacterium atrosepticum using primers based on Rhs family gene sequences. Plant Pathol. 2006, 55, 625–629. [Google Scholar] [CrossRef]
- Ash, G.J.; Lang, J.M.; Triplett, L.R.; Stodart, B.J.; Verdier, V.; Vera Cruz, C.; Rott, P.; Leach, J.E. Development of a genomics-based LAMP (loop-mediated isothermal amplification) assay for detection of Pseudomonas fuscovaginae from rice. Plant Dis. 2014, 98, 909–915. [Google Scholar] [CrossRef]
- Bühlmann, A.; Pothier, J.F.; Tomlinson, J.A.; Frey, J.E.; Smits, T.H.M.; Duffy, B. Genomics-informed design of loop-mediated isothermal amplification for detection of phytopathogenic Xanthomonas arboricola pv. pruni at the intraspecific level. Plant Pathol. 2013, 62, 475–484. [Google Scholar] [CrossRef]
- Malapi-Wight, M.; Demers, J.E.; Veltri, D.; Marra, R.E.; Crouch, J.A. LAMP detection assays for boxwood blight pathogens: A comparative genomics approach. Sci. Rep. 2016, 6, 26140. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.T.; Zheng, L.; Jia, Q.; Liu, H.; Hsiang, T.; Huang, J.B. PCR markers derived from comparative genomics for detection and identification of the rice pathogen Ustilaginoidea virens in plant tissues. Plant Dis. 2017, 101, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, E63. [Google Scholar] [CrossRef] [PubMed]
- Nagamine, K.; Watanabe, K.; Ohtsuka, K.; Hase, T.; Notomi, T. Loop-mediated isothermal amplification reaction using a nondenatured template. Clin. Chem. 2001, 47, 1742–1743. [Google Scholar]
- Nagamine, K.; Hase, T.; Notomi, T. Accelerated reaction by loop-mediated isothermal amplification using loop primers. Mol. Cell. Probes 2002, 16, 223–229. [Google Scholar] [CrossRef] [PubMed]
- Tomotada, I.; Toshiaki, S.; Kozaburo, H. Loop-mediated isothermal amplification for direct detection of Mycobacterium tuberculosis complex, M. avium, and M. intracellulare in sputum samples. J. Clin. Microbiol. 2003, 41, 2616–2622. [Google Scholar]
- Mori, Y.; Nagamine, K.; Tomita, N.; Notomi, T. Detection of loop-mediated isothermal amplification reaction by turbidity derived from magnesium pyrophosphate formation. Biochem. Biophys. Res. Commun. 2001, 289, 150–154. [Google Scholar] [CrossRef]
- Yasuhara-Bell, J.; Marrero, G.; De Silva, A.; Alvarez, A.M. Specific detection of Pectobacterium carotovorum by loop-mediated isothermal amplification. Mol. Plant Pathol. 2016, 17, 1499–1505. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.J.; Chen, W.Y.; Wang, D.P.; He, X.H.; Zhu, X.N.; Shing, X.M. Species-specific PCR detection of the food-borne pathogen Vibrio parahaemolyticus using the irgB gene identified by comparative genomic analysis. FEMS Microbiol. Lett. 2010, 307, 65–71. [Google Scholar] [CrossRef]
- Pritchard, L.; Humphris, S.; Saddler, G.S.; Parkinson, N.M.; Bertrand, V.; Elphinstone, J.G.; Toth, I.K. Detection of phytopathogens of the genus Dickeya using a PCR primer prediction pipeline for draft bacterial genome sequences. Plant Pathol. 2012, 62, 587–596. [Google Scholar] [CrossRef]
- Van Dam, P.; de Sain, M.; ter Horst, A.; van der Gragt, M.; Rep, M. Use of comparative genomics-based markers for discrimination of host specificity in Fusarium oxysporum. Appl. Environ. Microbiol. 2018, 84, e01868-17. [Google Scholar] [CrossRef]
- Kong, X.J.; Qin, W.T.; Huang, X.Q.; Kong, F.F.; Schoen, C.D.; Feng, J.; Wang, Z.Y.; Zhang, H. Development and application of loop-mediated isothermal amplification (LAMP) for detection of Plasmopara viticola. Sci. Rep. 2016, 6, 28935. [Google Scholar] [CrossRef] [PubMed]
- Gloster, T.M. Advances in understanding glycosyltransferases from a structural perspective. Curr Opin Struct. Biol. 2014, 28, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Sun, Z.; Yan, J.; Luo, Y.; Wang, H.; Ma, Z. Rapid and precise detection of latent infections of wheat stripe rust in wheat leaves using loop-mediated isothermal amplification. J. Phytopathol. 2011, 159, 582–584. [Google Scholar] [CrossRef]
- Hsiang, T.; Baillie, D.L. Comparison of the yeast proteome to other fungal genomes to find core fungal genes. J. Mol. Evol. 2005, 60, 475–483. [Google Scholar] [CrossRef]


| Gene No. | Length (bp) | BLASTX Prediction | Predicted Domain a | Signal Peptide b |
|---|---|---|---|---|
| PC1_0970 | 303 | Hypothetical protein | YCII | No |
| PC1_1421 | 1320 | Hypothetical protein | No | Yes |
| PC1_1622 | 452 | Hypothetical protein | No | No |
| PC1_1675 | 1809 | Glycosyl transferase | Glyco_transf_8 | No |
| PC1_2056 | 891 | Hypothetical protein | No | Yes |
| PC1_2248 | 792 | Hypothetical protein | No | No |
| PC1_2623 | 831 | Hypothetical protein | DUF4393 | No |
| PC1_2625 | 345 | Hypothetical protein | No | No |
| PC1_2831 | 303 | Hypothetical protein | No | Yes |
| Primer | Sequence (5′-3′) | Description |
|---|---|---|
| F3 | GCACAAGCTTGACTGCATAC | Outer primer |
| B3 | TGGCGAGTTGTCCCCATAG | Outer primer |
| FIP | CGCCGTATCGGCACAGAAGAAAGCTTGGCGTTTCTCTCTCA | Inner primer |
| BIP | GCCCTCATCTCGCTGGCAATCATGTGCTTCCGGCAACAC | Inner primer |
| LF | AGAAACGGGATGGGGTGG | Loop primer |
| LB | GCCATCGAGCGTAGCGAA | Loop primer |
| Species | Strain | Host | Location | LAMP a (1675-1) | PCR (1675F3/B3) |
|---|---|---|---|---|---|
| P. aroidearum | YCC1 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC2 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC3 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC4 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC5 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC6 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC7 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC8 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC9 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC10 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC11 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC12 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC13 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | YCC14 | Konjac | Enshi, Hubei | + | + |
| P. aroidearum | NS-1 | Konjac | Enshi, Hubei | + | + |
| P. aroidearum | NS-2 | Konjac | Enshi, Hubei | + | + |
| P. aroidearum | NS-3 | Konjac | Enshi, Hubei | + | + |
| P. aroidearum | NS-4 | Konjac | Enshi, Hubei | + | + |
| P. aroidearum | NS-5 | Konjac | Enshi, Hubei | + | + |
| P. aroidearum | WF-1 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | WF-2 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | WF-3 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | WF-4 | Konjac | Yichang, Hubei | + | + |
| P. aroidearum | WF-5 | Konjac | Yichang, Hubei | + | + |
| D. fangzhongdai | YCH1 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | YCH2 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | YCH3 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | YCH4 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | YCH5 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | YCH6 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | YCH7 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | YCH8 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | YCH9 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | WFH-1 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | WFH-2 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | WFH-3 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | WFH-4 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | WFH-5 | Konjac | Yichang, Hubei | − | − |
| D. fangzhongdai | WFH-6 | Konjac | Yichang, Hubei | − | − |
| Pcc | CGMCC1.141 | Cabbage | Beijing | − | − |
| P. carotovorum subsp. actinidiae | HN-1 | Chili | Hunan | − | − |
| P. carotovorum subsp. brasiliense | YCB1 | Konjac | Yichang, Hubei | − | − |
| D. chrysanthemi | CGMCC1.7280 | Unknown | Beijing | − | − |
| D. zeae | CGMCC1.3614 | Unknown | Beijing | − | − |
| Erwinia amylovora | CGMCC1.7276 | Unknown | Beijing | − | − |
| E. pyrifoliae | CGMCC1.7277 | Unknown | Beijing | − | − |
| Ralstonia solanacearum | GMI1000 | Potato | Wuhan, Hubei | − | − |
| Acidovorax citrulli | HN-2 | Watermelon | Hunan | − | − |
| Xanthomonas oryzae pv. oryzae | HW-67 | Rice | Wuhan, Hubei | − | − |
| Kosakonia pseudosacchari | M1 | Konjac | Yichang, Hubei | − | − |
| Acinetobacter calcoaceticus | M2 | Konjac | Yichang, Hubei | − | − |
| Bacillus aryabhattai | M3 | Konjac | Yichang, Hubei | − | − |
| B. firmus | M4 | Konjac | Yichang, Hubei | − | − |
| B. simplex | M5 | Konjac | Yichang, Hubei | − | − |
| Citrobacter farmeri | M6 | Konjac | Yichang, Hubei | − | − |
| C. freundii | M7 | Konjac | Yichang, Hubei | − | − |
| Enterobacter aerogenes | M8 | Konjac | Yichang, Hubei | − | − |
| E. asburiae | M9 | Konjac | Yichang, Hubei | − | − |
| Enterobacter sp. | M10 | Konjac | Yichang, Hubei | − | − |
| Klebsiella oxytoca | M11 | Konjac | Yichang, Hubei | − | − |
| K. pneumoniae | M12 | Konjac | Yichang, Hubei | − | − |
| Morganella morganii | M13 | Konjac | Yichang, Hubei | − | − |
| Paenibacillus polymyxa | M14 | Konjac | Yichang, Hubei | − | − |
| Pantoea agglomerans | M15 | Konjac | Yichang, Hubei | − | − |
| Providencia rettgeri | M16 | Konjac | Yichang, Hubei | − | − |
| Pseudomonas plecoglossicida | M17 | Konjac | Yichang, Hubei | − | − |
| P. putida | M18 | Konjac | Yichang, Hubei | − | − |
| Raoultella ornithinolytica | M19 | Konjac | Yichang, Hubei | − | − |
| R. planticola | M20 | Konjac | Yichang, Hubei | − | − |
| R. terrigena | M21 | Konjac | Yichang, Hubei | − | − |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, M.; Liu, H.; Huang, J.; Peng, J.; Fei, F.; Zhang, Y.; Hsiang, T.; Zheng, L. A Loop-Mediated Isothermal Amplification Assay for Rapid Detection of Pectobacterium aroidearum that Causes Soft Rot in Konjac. Int. J. Mol. Sci. 2019, 20, 1937. https://doi.org/10.3390/ijms20081937
Sun M, Liu H, Huang J, Peng J, Fei F, Zhang Y, Hsiang T, Zheng L. A Loop-Mediated Isothermal Amplification Assay for Rapid Detection of Pectobacterium aroidearum that Causes Soft Rot in Konjac. International Journal of Molecular Sciences. 2019; 20(8):1937. https://doi.org/10.3390/ijms20081937
Chicago/Turabian StyleSun, Miaomiao, Hao Liu, Junbin Huang, Jinbo Peng, Fuhua Fei, Ya Zhang, Tom Hsiang, and Lu Zheng. 2019. "A Loop-Mediated Isothermal Amplification Assay for Rapid Detection of Pectobacterium aroidearum that Causes Soft Rot in Konjac" International Journal of Molecular Sciences 20, no. 8: 1937. https://doi.org/10.3390/ijms20081937
APA StyleSun, M., Liu, H., Huang, J., Peng, J., Fei, F., Zhang, Y., Hsiang, T., & Zheng, L. (2019). A Loop-Mediated Isothermal Amplification Assay for Rapid Detection of Pectobacterium aroidearum that Causes Soft Rot in Konjac. International Journal of Molecular Sciences, 20(8), 1937. https://doi.org/10.3390/ijms20081937





