Bone Marrow Endothelial Cells Influence Function and Phenotype of Hematopoietic Stem and Progenitor Cells after Mixed Neutron/Gamma Radiation
Abstract
1. Background
2. Statistical Analysis
3. Results
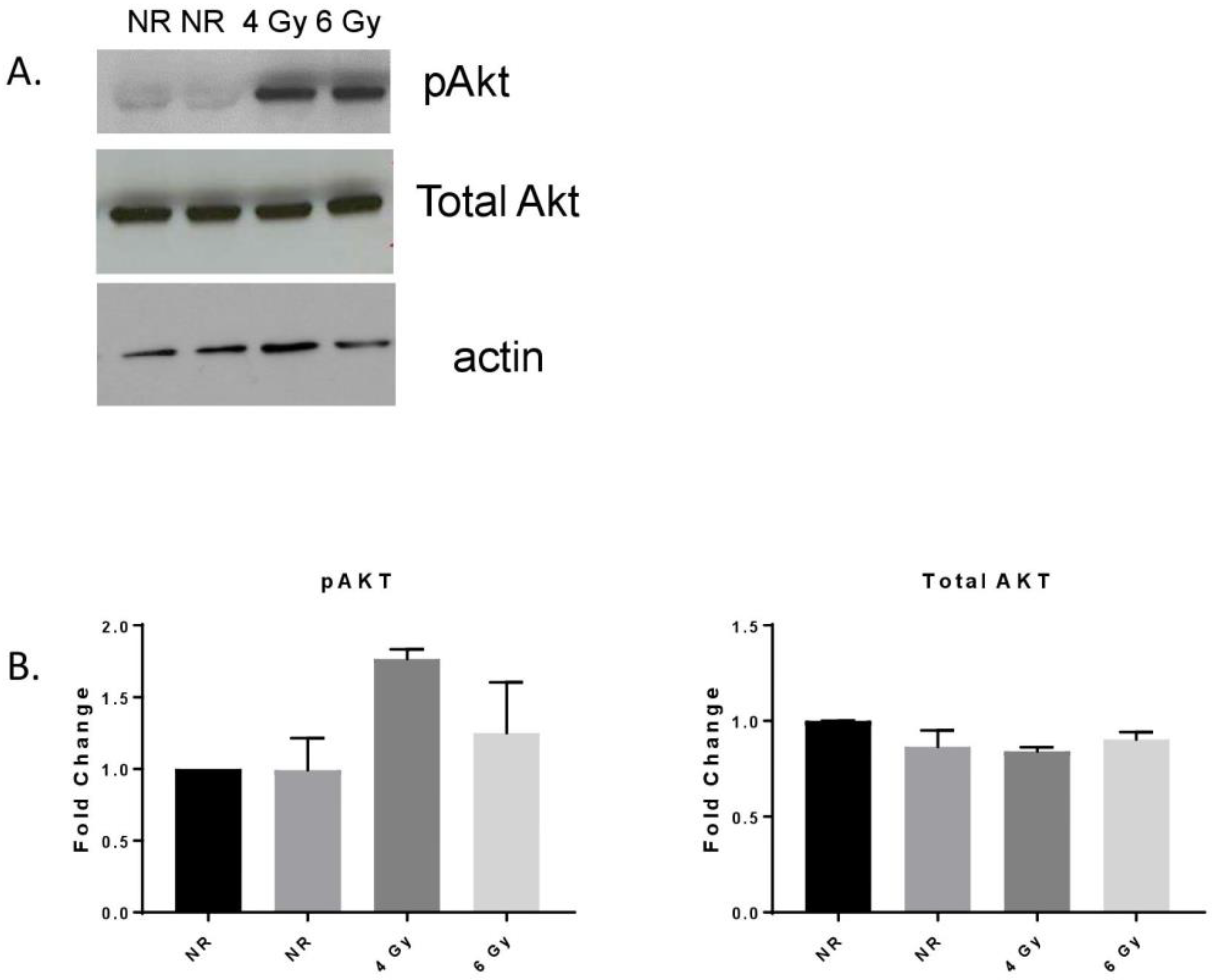
3.1. MF Radiation-Induced Akt Signaling in hBMEC.
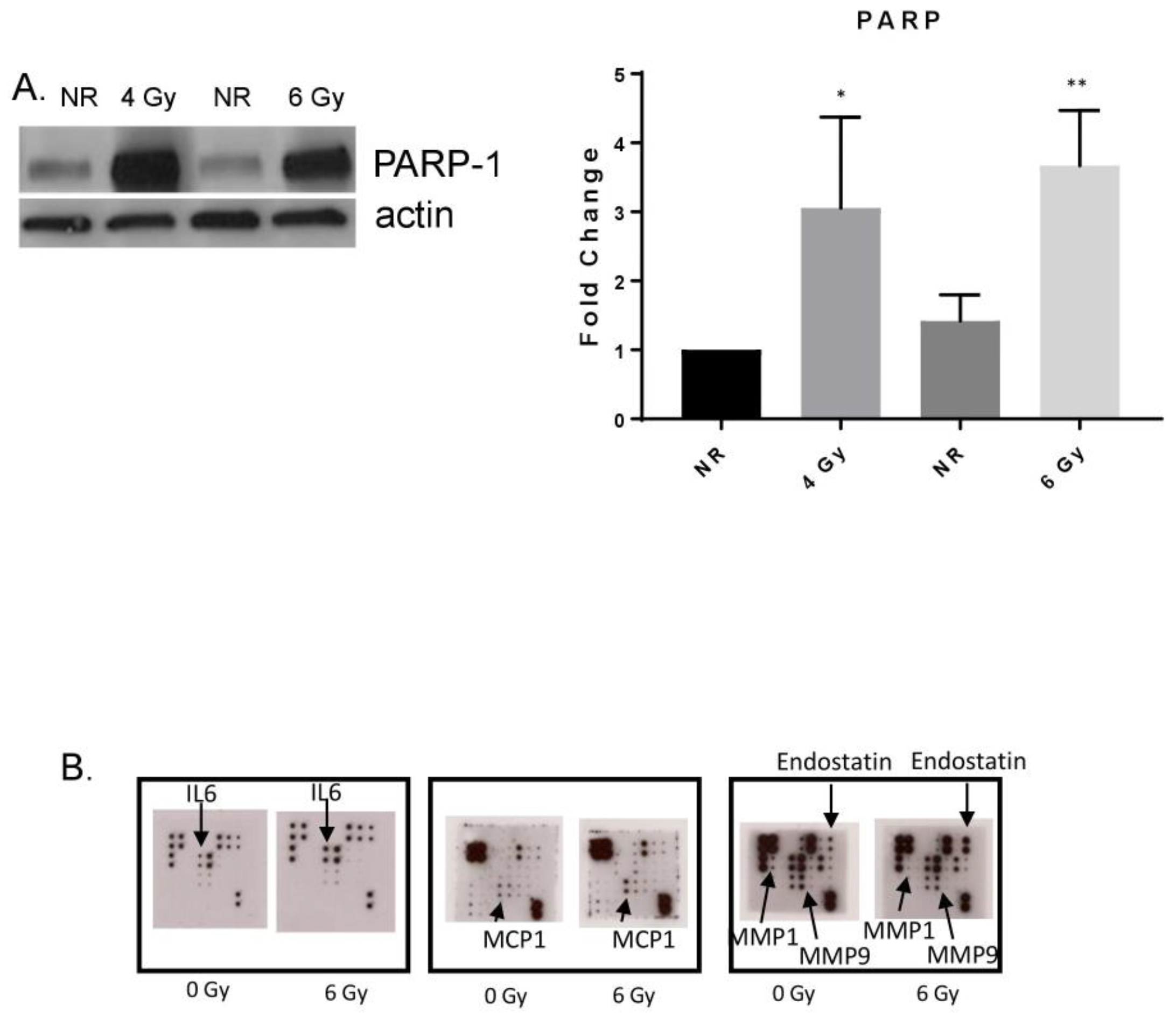
3.2. MF Radiation Altered Protein Expression in hBMEC.
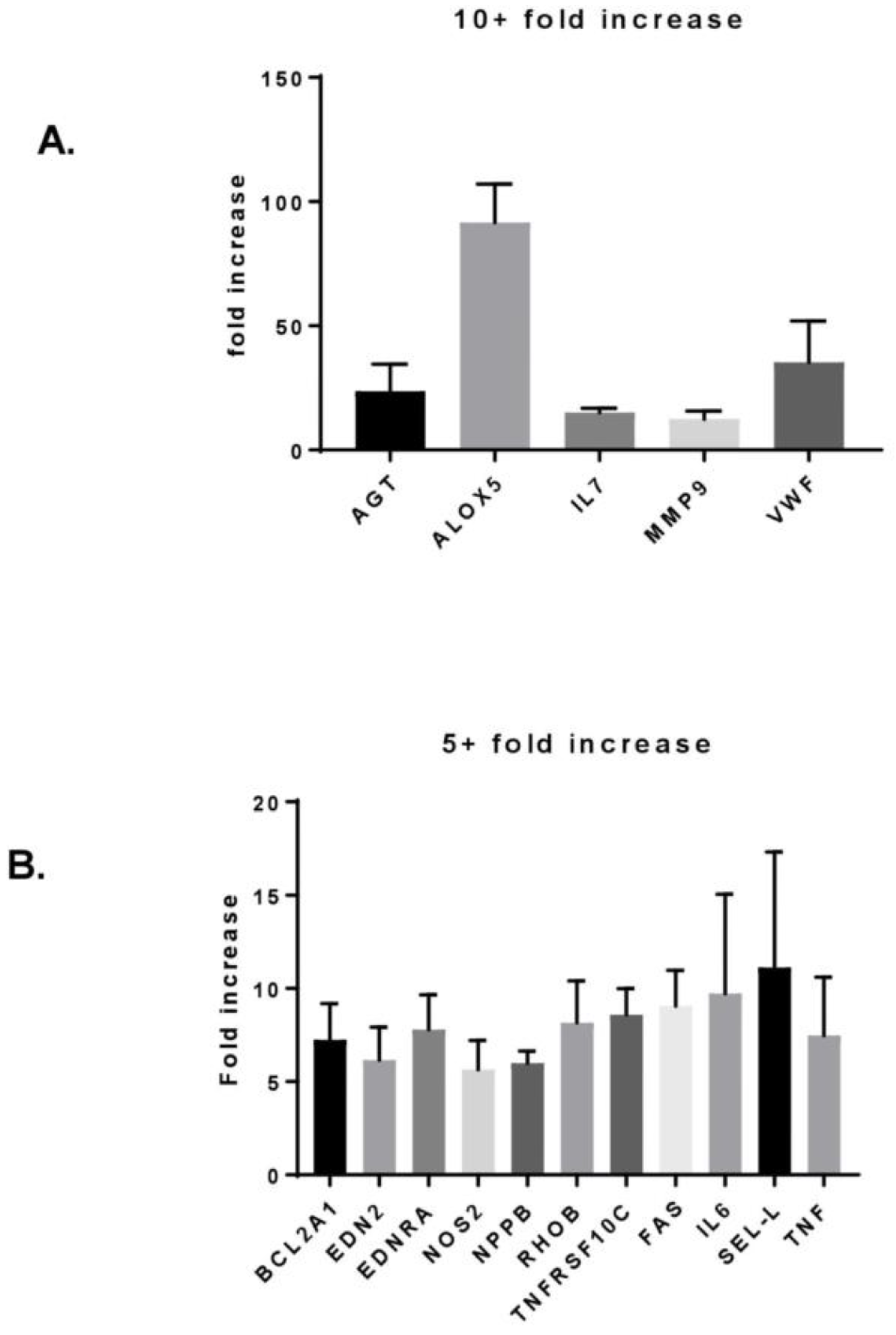
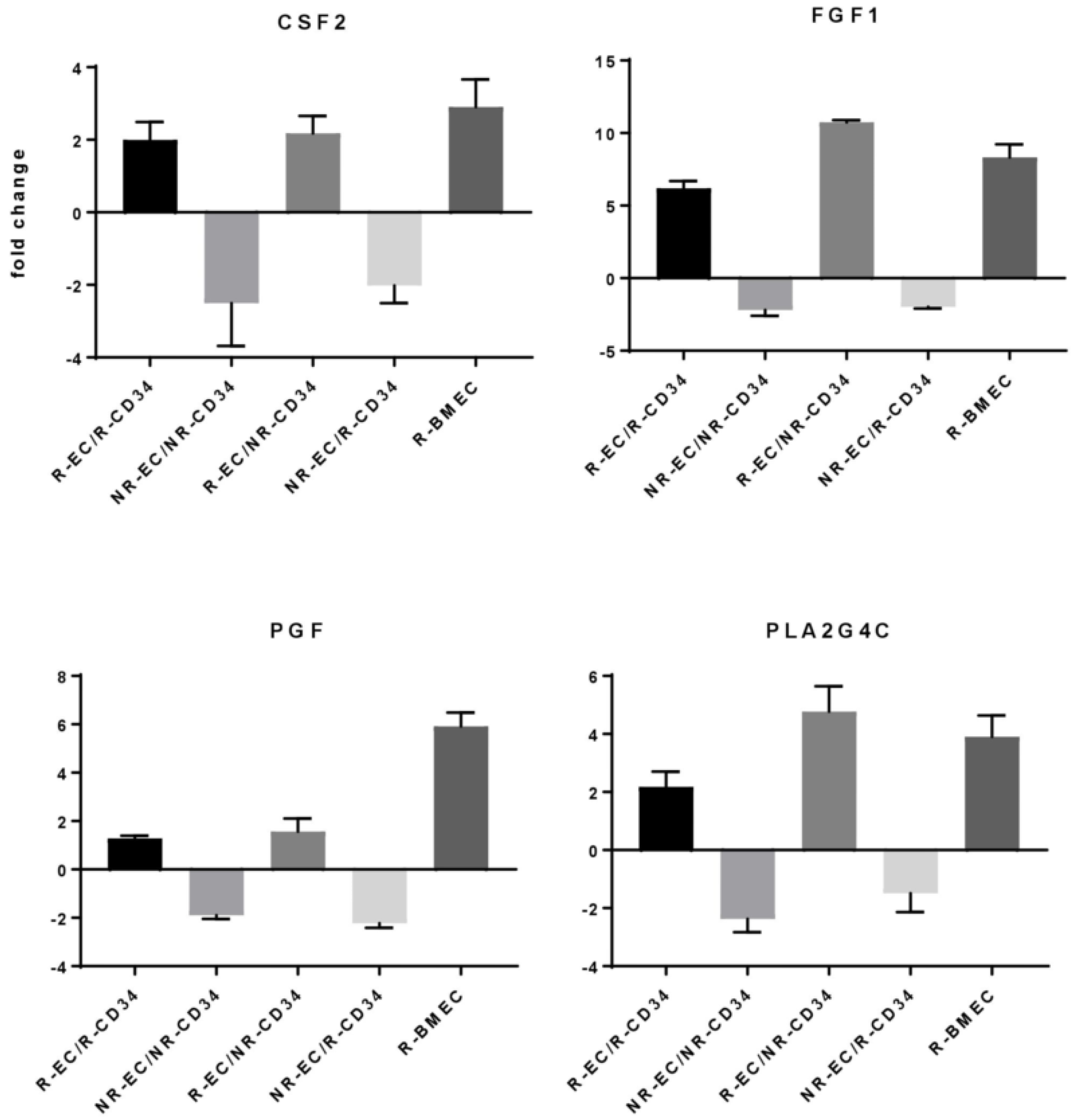
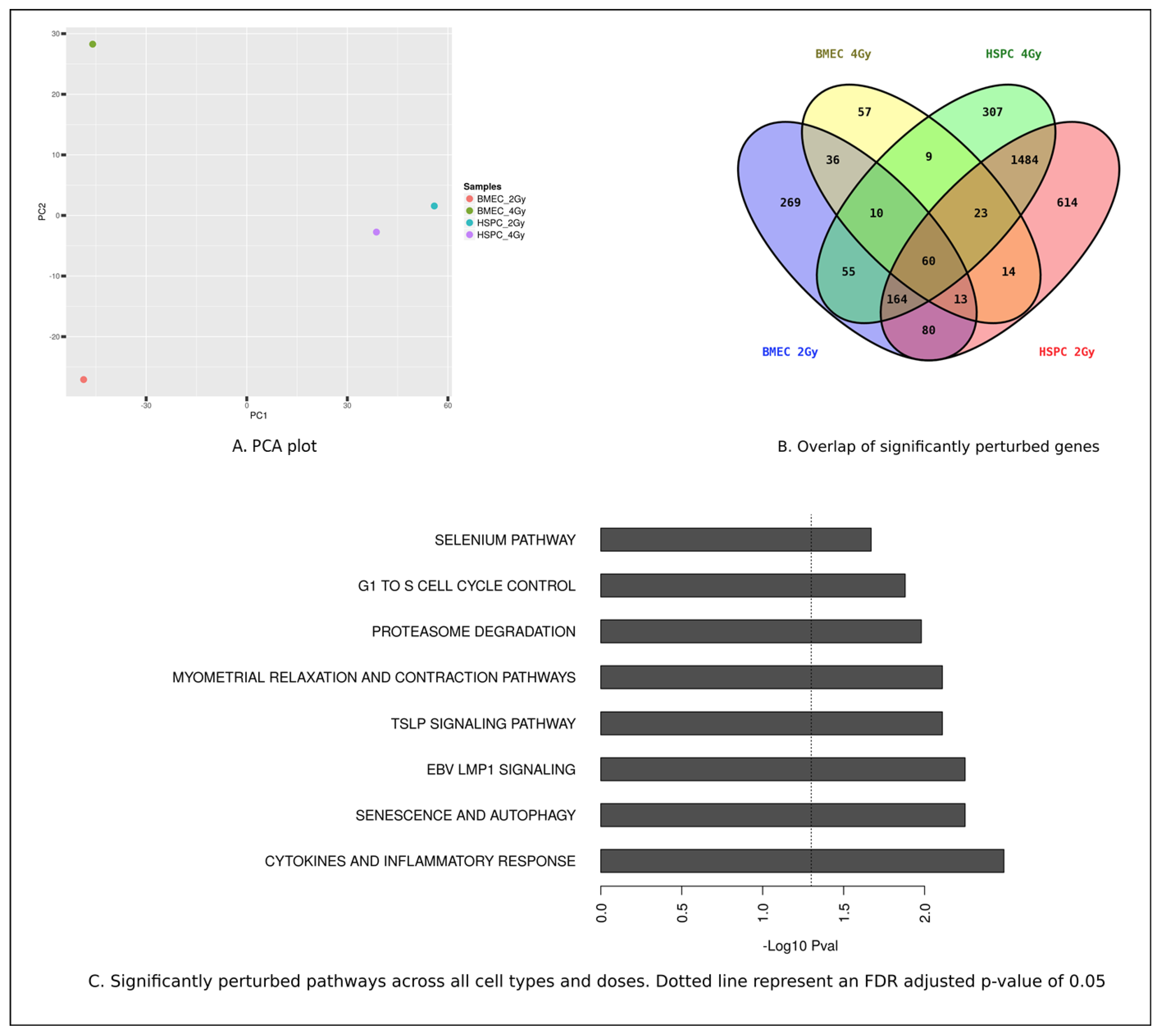
3.3. MF Radiation and CD34+ Coculture Altered mRNA Expression in hBMEC
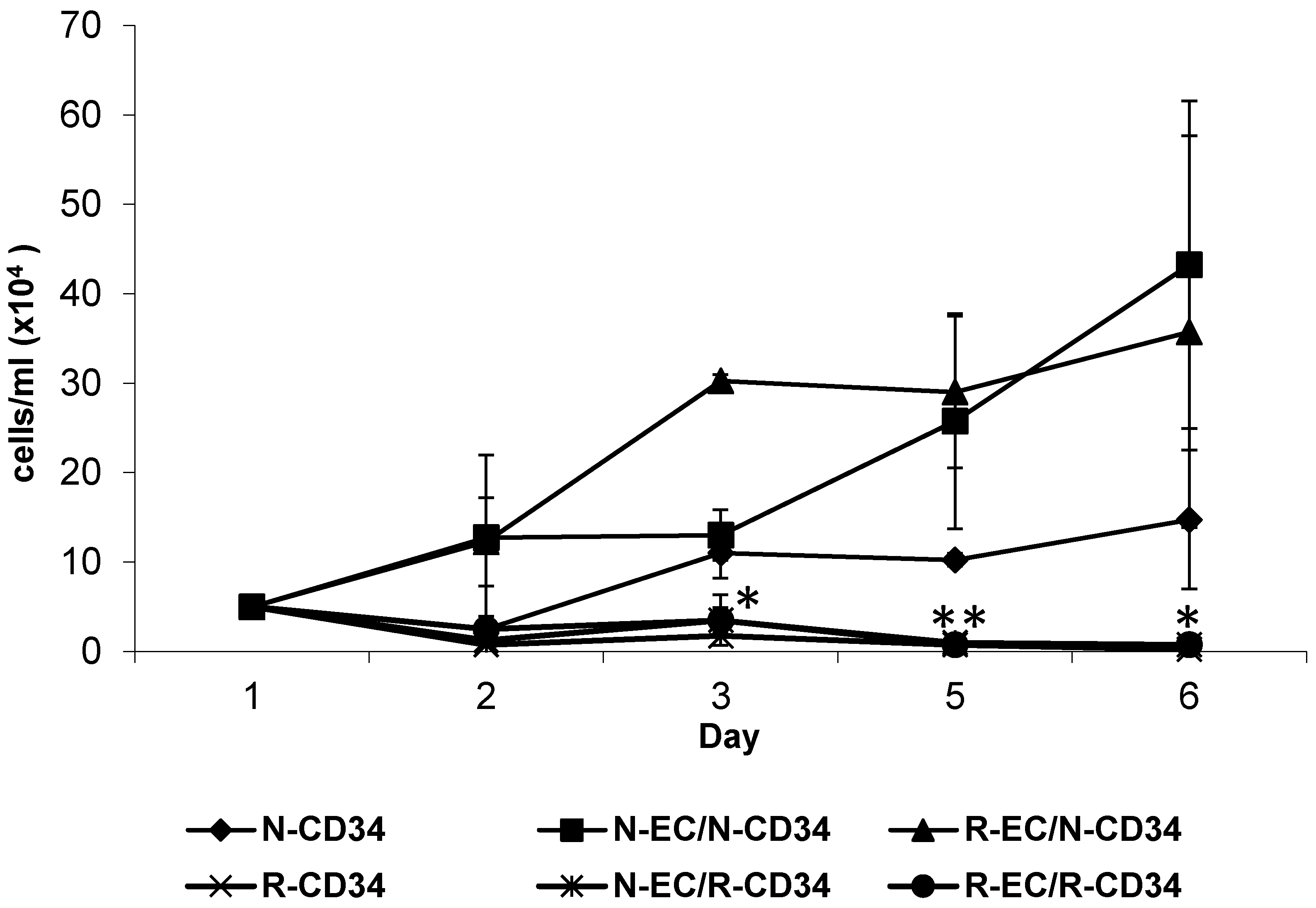
3.4. hBMEC Altered Stem Cell Proliferation
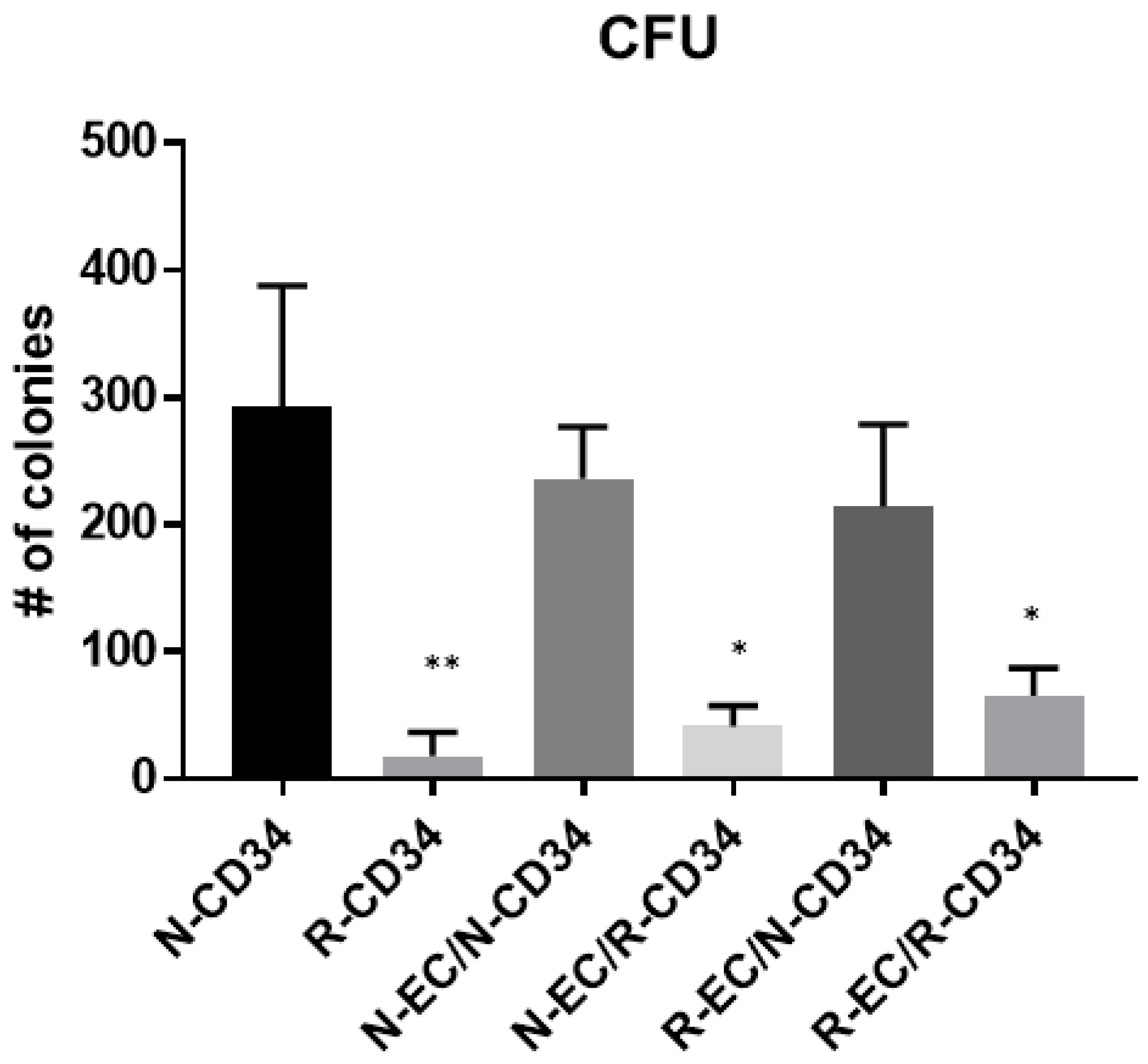
3.5. hBMEC Improved HSPC Differentiation after MF Radiation
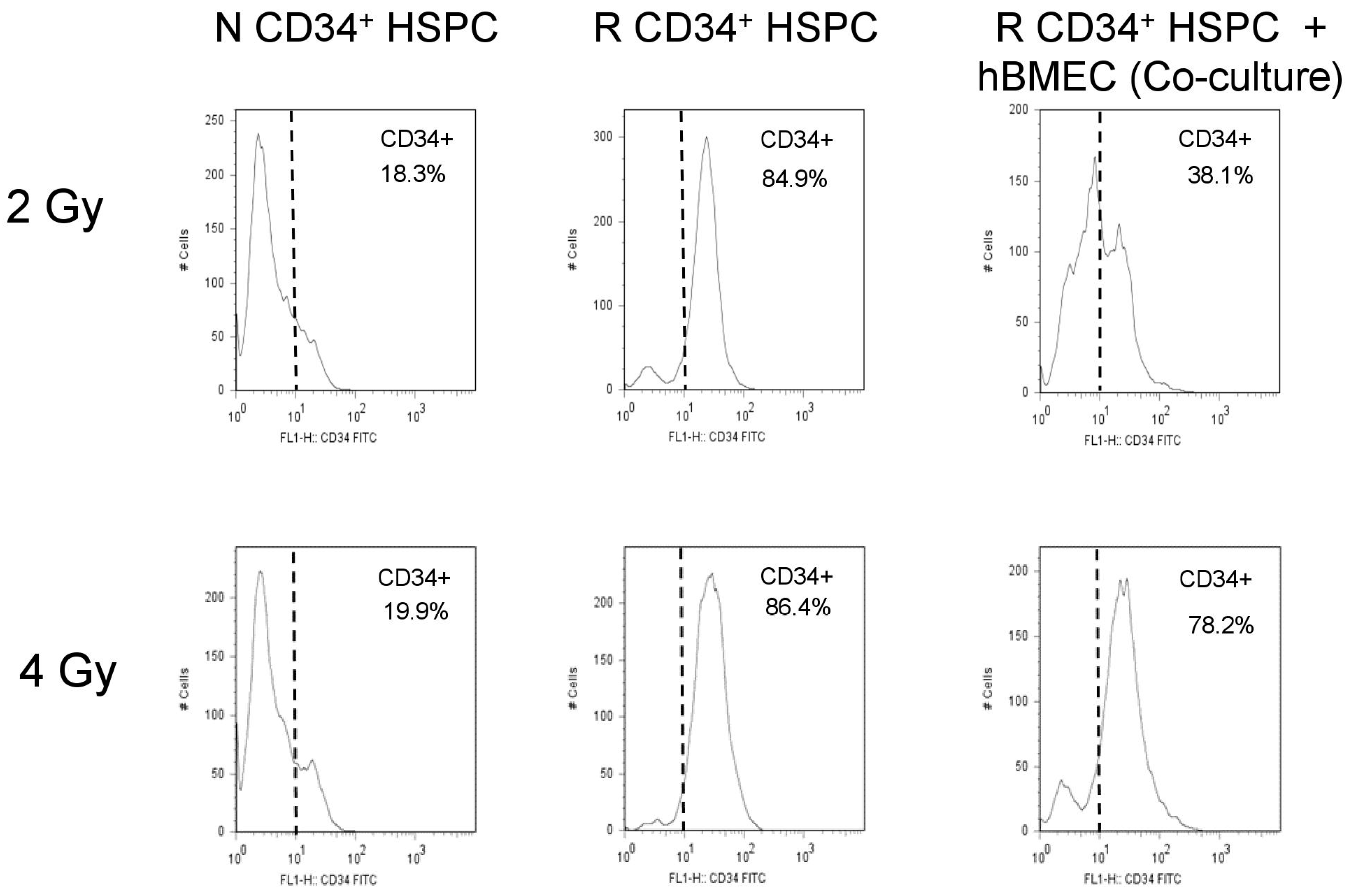
3.6. hBMEC Protected HSPC Phenotype after Radiation
4. Discussion
5. Materials and Methods
5.1. Cell Culture
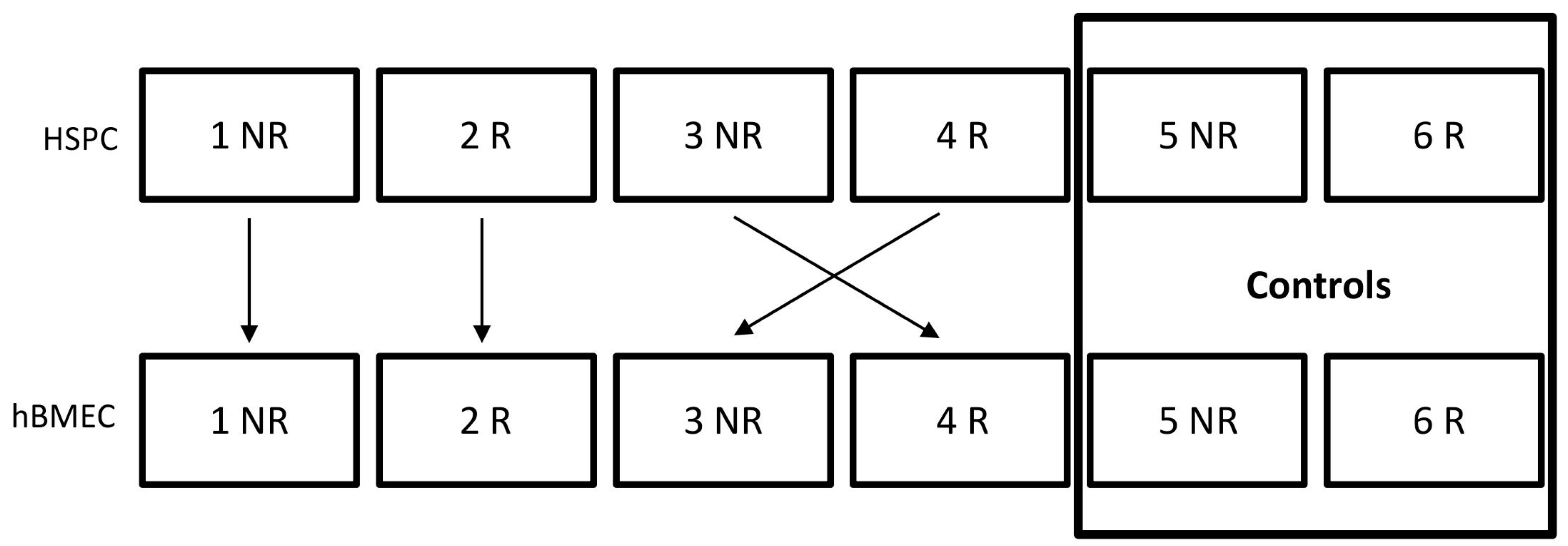
5.2. Coculture
5.3. MF Irradiation
5.4. Western Blot
5.5. Protein Array
5.6. PCR Array Analysis
5.7. Microarray Analysis
5.8. Survival and Proliferation
5.9. Hematopoietic Progenitor Clonogenic Assay
5.10. Flow Cytometry
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ARS | acute radiation syndrome |
| BM | bone marrow |
| CFU | colony forming unit |
| EC | endothelial cell |
| ECM | extracellular matrix |
| FBS | fetal bovine serum |
| HSC/HSPC | hematopoietic stem/hematopoietic stem and progenitor cells |
| ICAM | intracellular adhesion molecule |
| IND | improvised nuclear device |
| IR | ionizing radiation |
| LET | linear energy transfer |
| MF | mixed field |
| MM | multiple myeloma |
| NR | nonirradiated |
| R | irradiated |
| RIN | RNA Integration Number |
| SASP | senescence-associated secretory phenotypes |
| SC | stem cell |
| SCF | stem cell factor |
| TBI | total body irradiation |
| ARS | acute radiation syndrome |
| BM | bone marrow |
References
- Jacobson, L.O.; Simmons, E.L.; Marks, E.K.; Robson, M.J.; Bethard, W.F.; Gaston, E.O. The role of the spleen in radiation injury and recovery. J. Lab. Clin. Med. 1950, 35, 746–770. [Google Scholar]
- Lorenz, E.; Uphoff, D.; Reid, T.R.; Shelton, E. Modification of irradiation injury in mice and guinea pigs by bone marrow injections. J. Natl. Cancer Inst. 1951, 12, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Inoue, T.; Hirabayashi, Y.; Mitsui, H.; Sasaki, H.; Cronkite, E.P.; Bullis, J.E., Jr.; Bond, V.P.; Yoshida, K. Survival of spleen colony-forming units (CFU-S) of irradiated bone marrow cells in mice: Evidence for the existence of a radioresistant subfraction. Exp. Hematol. 1995, 23, 1296–1300. [Google Scholar] [PubMed]
- Van Vekkum, D.W. Radiation sensitivity of the hemopoietic stem cell. Radiat. Res. 1991, 128, S4–S8. [Google Scholar] [CrossRef]
- Rodgerson, D.O.; Reidenberg, B.E.; Harris, A.G.; Pecora, A.L. Potential for a pluripotent adult stem cell treatment for acute radiation sickness. World J. Exp. Med. 2012, 2, 37–44. [Google Scholar] [CrossRef] [PubMed]
- Pinzur, L.; Akyuez, L.; Levdansky, L.; Blumenfeld, M.; Volinsky, E.; Aberman, Z.; Reinke, P.; Ofir, R.; Volk, H.D.; Gorodetsky, R. Rescue from lethal acute radiation syndrome (ARS) with severe weight loss by secretome of intramuscularly injected human placental stromal cells. J. Cachexia Sarcopenia Muscle 2018, 9, 1079–1092. [Google Scholar] [CrossRef] [PubMed]
- Giralt, S.; Costa, L.; Schriber, J.; Dipersio, J.; Maziarz, R.; McCarty, J.; Shaughnessy, P.; Snyder, E.; Bensinger, W.; Copelan, E.; et al. Optimizing autologous stem cell mobilization strategies to improve patient outcomes: Consensus guidelines and recommendations. Biol. Blood Marrow Transplant. 2014, 20, 295–308. [Google Scholar] [CrossRef] [PubMed]
- Arora, S.; Majhail, N.S.; Liu, H. Hematopoietic progenitor cell mobilization for autologous stem cell transplantation in multiple myeloma in contemporary era. Clin. Lymphoma Myeloma Leuk. 2018, in press. [Google Scholar] [CrossRef]
- Juric, M.; Ghimire, S.; Ogonek, J.; Weissinger, E.; Holler, E.; Van Rood, J.; Oudshoorn, M.; Dickinson, A.; Greinix, H. Milestones of Hematopoietic Stem Cell Transplantation—From First Human Studies to Current Developments. Front. Immunol. 2016, 7, 470. [Google Scholar] [CrossRef]
- Singh, V.K.; Fatanmi, O.O.; Singh, P.K.; Whitnall, M.H. Role of radiation-induced granulocyte colony-stimulating factor in recovery from whole body gamma-irradiation. Cytokine 2012, 58, 406–414. [Google Scholar] [CrossRef] [PubMed]
- Bonig, H.; Papayannopoulou, T. Mobilization of hematopoietic stem/progenitor cells: General principles and molecular mechanisms. Methods Mol. Biol. 2012, 904, 1–14. [Google Scholar]
- Shao, L.; Luo, Y.; Zhou, D. Hematopoietic stem cell injury induced by ionizing radiation. Antioxid. Redox Signal. 2014, 20, 1447–1462. [Google Scholar] [CrossRef] [PubMed]
- Ray, S.; Kulkarni, S.; Chakraborty, K.; Pessu, R.; Hauer-Jensen, M.; Kumar, K.; Ghosh, S. Mobilization of progenitor cells into peripheral blood by gamma-tocotrienol: A promising radiation countermeasure. Int. Immunopharmacol. 2013, 15, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.; Brown, D.; Kao, T. Alpha-tocopherol succinate protects mice from gamma-radiation by induction of granulocyte-colony stimulating factor. Int. J. Radiat. Biol. 2010, 86, 12–21. [Google Scholar] [CrossRef]
- Available online: https://www.fda.gov/emergencypreparedness/counterterrorism/medicalcountermeasures/aboutmcmi/ucm443245.htm (accessed on 30 March 2019).
- Zhang, J.; Niu, C.; Ye, L.; Huang, H.; He, X.; Tong, W.; Ross, J.; Haug, J.; Johnson, T.; Fend, J.; et al. Identification of the haematopoietic stem cell niche and control of the niche size. Nature 2003, 425, 836–841. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Yin, T.; Wiegraebe, W.; He, X.; Miller, D.; Stark, D.; Perko, K.; Alexander, R.; Schwartz, J.; Grindley, J.; et al. Detection of functional haematopoietic stem cell niche using real-time imaging. Nature 2009, 457, 97–101. [Google Scholar] [CrossRef] [PubMed]
- Mikkola, H.K.; Orkin, S.H. The journey of developing hematopoietic stem cells. Development 2006, 133, 3733–3744. [Google Scholar] [CrossRef] [PubMed]
- Chotinantakul, K.; Leeanansaksiri, W. Hematopoietic Stem Cell Development, Niches, and Signaling Pathways. Bone Marrow Res. 2012, 2012, 270425. [Google Scholar] [CrossRef]
- Richter, R.; Forssmann, W.; Henschler, R. Current Developments in Mobilization of Hematopoietic Stem and Progenitor Cells and Their Interaction with Niches in Bone Marrow. Transfus. Med. Hemother. 2017, 44, 151–164. [Google Scholar] [CrossRef]
- Kiel, M.; Radice, G.; Morrison, J. Lack of evidence that hematopoietic stem cells depend on N-cadherin-mediated adhesion to osteoblasts for their maintenance. Cell Stem Cell 2007, 1, 204–217. [Google Scholar] [CrossRef]
- Kiel, M.; Yilmaz, O.; Iwashita, T.; Terhorst, C.; Morrison, S. SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell 2005, 121, 1109–1121. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Andres, M.; Timiryasova, T.; Fodor, I.; Slater, J.; Gridley, D. Radiation-enhanced endostatin gene expression and effects of combination treatment. Technol. Cancer Res. Treat. 2005, 4, 193–202. [Google Scholar] [CrossRef]
- Vasa, M.; Fichtlscherer, S.; Adler, K.; Aicher, A.; Martin, H.; Zeiher, A.; Dimmeler, S. Increase in circulating endothelial progenitor cells by statin therapy in patients with stable coronary artery disease. Circulation 2001, 103, 2885–2890. [Google Scholar] [CrossRef]
- Méndez-Ferrer, S.; Michurina, T.V.; Ferraro, F.; Mazloom, A.R.; Macarthur, B.D.; Lira, S.A.; Scadden, D.T.; Ma’ayan, A.; Enikolopov, G.N.; Frenette, P.S. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature 2010, 466, 829–834. [Google Scholar] [CrossRef] [PubMed]
- Muramoto, G.G.; Chen, B.; Cui, X.; Chao, N.J.; Chute, J.P. Vascular endothelial cells produce soluble factors that mediate the recovery of human hematopoietic stem cells after radiation injury. Biol. Blood Marrow Transplant. 2006, 12, 530–540. [Google Scholar] [CrossRef] [PubMed]
- Himburg, H.A.; Muramoto, G.G.; Daher, P.; Meadows, S.K.; Russell, J.L.; Doan, P.; Chi, J.T.; Salter, A.B.; Lento, W.E.; Reya, T.; et al. Pleiotrophin regulates the expansion and regeneration of hematopoietic stem cells. Nat. Med. 2010, 16, 475–482. [Google Scholar] [CrossRef] [PubMed]
- Doan, P.L.; Himburg, H.A.; Helms, K.; Russell, J.L.; Fixsen, E.; Quarmyne, M.; Harris, J.R.; Deoliviera, D.; Sullivan, J.M.; Chao, N.J.; et al. Epidermal growthfactor regulates hematopoietic regeneration after radiation injury. Nat. Med. 2013, 19, 295–304. [Google Scholar] [CrossRef] [PubMed]
- Yazlovitskaya, E.; Linkous, A.; Thotala, D.; Cuneo, K.; Hallahan, D. Cytosolic phospholipase A2 regulates viability of irradiated vascular endothelium. Cell Death Differ. 2008, 15, 1641–1653. [Google Scholar] [CrossRef]
- Edwards, E.; Geng, L.; Tan, J.; Onishko, H.; Donnelly, E.; Hallahan, D. Phosphatidylinositol 3-Kinase/Akt Signaling in the response of vascular endothelium to ionizing radiation. Cancer Res. 2002, 62, 4671–4677. [Google Scholar]
- Chou, C.H.; Chen, S.U.; Cheng, J.C. Radiation-induced interleukin-6 expression through MAPK/p38/NF-kappaB signaling pathway and the resultant antiapoptotic effect on endothelial cells through Mcl-1 expression with sIL6-Ralpha. Int. J. Radiat. Oncol. Biol. Phys. 2009, 75, 1553–1561. [Google Scholar] [CrossRef]
- Nalla, A.; Gogineni, V.; Gupta, R.; Dinh, D.; Rao, J. Suppression of uPA and uPAR blocks radiation-induced MCP-1 mediated recruitment of endothelial cells in meningioma. Cell. Signal. 2011, 23, 1299–1310. [Google Scholar] [CrossRef]
- Nirmala, C.; Jasti, S.; Sawaya, R.; Kyritsis, A.; Konduri, S.; Ali-Osman, F.; Rao, J.; Mohanam, S. Effects of radiation on the levels of MMP-2, MMP-9 and TIMP-1 during morphogenic glial-endothelial cell interactions. Int. J. Cancer 2000, 88, 766–771. [Google Scholar] [CrossRef]
- Clevers, H. Stem cells, asymmetric division and cancer. Nat. Genet. 2005, 37, 1027–1028. [Google Scholar] [CrossRef] [PubMed]
- Liles, W.; Broxmeyer, H.; Rodger, E.; Wood, B.; Hubel, K.; Cooper, S.; Hangoc, G.; Bridger, G.; Henson, G.; Calandra, G.; et al. Mobilization of hematopoietic progenitor cells in healthy volunteers by AMD3100, a CXCR4 antagonist. Blood 2003, 102, 2728–2730. [Google Scholar] [CrossRef] [PubMed]
- Dimmeler, S.; Aicher, A.; Vasa, M.; Mildner-Rihm, C.; Adler, K.; Tiemann, M.; Rütten, H.; Fichtlscherer, S.; Martin, H.; Zeiher, A.M. HMG-CoA reductase inhibitors (statins) increase endothelial progenitor cells via the PI 3-kinase/Akt pathway. J. Clin. Investig. 2001, 108, 391–397. [Google Scholar] [CrossRef] [PubMed]
- Llevadot, J.; Murasawa, S.; Kureishi, Y.; Uchida, S.; Masuda, H.; Kawamoto, A.; Walsh, K.; Isner, J.; Asahara, T. HMG-CoA reductase inhibitor mobilizes bone marrow—derived endothelial progenitor cells. J. Clin. Investig. 2001, 108, 399–405. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.; Newman, V.; Seed, T. Colony-stimulating factors for the treatment of the hematopoietic component of the acute radiation syndrome (H-ARS): A review. Cytokine 2015, 71, 22–37. [Google Scholar] [CrossRef] [PubMed]
- Farese, A.; MacVittie, T. Filgrastim for the treatment of hematopoietic acute radiation syndrome. Drugs Tooday 2015, 51, 537–548. [Google Scholar]
- Durante, M.; Loeffler, J. Charged particles in radiation oncology. Nat. Rev. Clin. Oncol. 2010, 7, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Rall, M.; Kraft, D.; Volcic, M.; Cucu, A.; Nasonova, E.; Taucher-Scholz, G.; Bönig, H.; Wiesmüller, L.; Fournier, C. Impact of charged particle exposure on homologous DNA double-strand break repair in human blood-derived cells. Front. Oncol. 2015, 5, 250. [Google Scholar] [CrossRef]
- Zhang, B.; Davidson, M.; Hei, T. Mitochondria regulate DNA damage and genomic instability induced by high LET radiation. Life Sci. Space Res. 2014, 1, 80–88. [Google Scholar] [CrossRef] [PubMed]
- Castejón, O. Increased vesicular and vacuolar transendothelial transport in traumatic human brain oedema. A review. Folia Neuropathol. 2013, 51, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Gaugler, M.; Squiban, C.; Claraz, M.; Schweitzer, K.; Weksler, B.; Gourmelon, P.; Van der Meeren, A. Characterization of the response of human bone marrow endothelial cells to in vitro irradiation. Br. J. Haematol. 1998, 103, 980–989. [Google Scholar] [CrossRef]
- Kalash, R.; Berhane, H.; Au, J.; Rhieu, B.; Epperly, M.; Goff, J.; Dixon, T.; Wang, H.; Zhang, X.; Franicola, D.; et al. Differences in irradiated lung gene transcription between fibrosis-prone C57BL/6NHsd and fibrosis-resistant C3H/HeNHsd mice. In Vivo 2014, 28, 147–171. [Google Scholar] [PubMed]
- Gabriels, K.; Hoving, S.; Seemann, I.; Visser, N.; Gijbels, M.; Pol, J.; Daemen, M.; Stewart, F.; Heeneman, S. Local heart irradiation of ApoE(−/−) mice induces microvascular and endocardial damage and accelerates coronary atherosclerosis. Radiother. Oncol. 2012, 105, 358–364. [Google Scholar] [CrossRef] [PubMed]
- Rhieu, B.; Epperly, M.; Cao, S.; Franicola, D.; Shields, D.; Goff, J.; Wang, H.; Greenberger, J. Increased hematopoiesis in long-term bone marrow cultures and reduced irradiation-induced pulmonary fibrosis in Von Willebrand factor homologous deletion recombinant mice. In Vivo 2014, 28, 449–456. [Google Scholar] [PubMed]
- Yabluchanskiy, A.; Ma, Y.; Iyer, R.; Hall, M.; Lindsey, M. Matrix metalloproteinase-9: many shades of function in cardiovascular disease. Physiology 2013, 28, 391–403. [Google Scholar] [CrossRef] [PubMed]
- Iolyeva, M.; Aebischer, D.; Proulx, S.; Willrodt, A.; Ecoiffier, T.; Häner, S.; Bouchaud, G.; Krieg, C.; Onder, L.; Ludewig, B.; et al. Interleukin-7 is produced by afferent lymphatic vessels and supports lymphatic drainage. Blood 2013, 122, 2271–2281. [Google Scholar] [CrossRef] [PubMed]
- Mazzucchelli, R.; Warming, S.; Lawrence, S.; Ishii, M.; Abshari, M.; Washington, A.; Feigenbaum, L.; Warner, A.; Sims, D.; Li, W.; et al. Visualization and identification of IL-7 producing cells in reporter mice. PLoS ONE 2009, 4, e7637. [Google Scholar] [CrossRef]
- Miller, C.; Hartigan-O’Connor, D.; Lee, M.; Laidlaw, G.; Cornelissen, I.; Matloubian, M.; Coughlin, S.; McDonald, D.; McCune, J. IL-7 production in murine lymphatic endothelial cells and induction in the setting of peripheral lymphopenia. Int. Immunol. 2013, 25, 471–483. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Zhao, L.; Wan, Y.; Zhu, B. Mechanism of action of IL-7 and its potential applications and limitations in cancer immunotherapy. Int. J. Mol. Sci. 2015, 16, 10267–10280. [Google Scholar] [CrossRef] [PubMed]
- Tsuboi, I.; Hirabayashi, Y.; Harada, T.; Hiramoto, M.; Kanno, J.; Inoue, T.; Aizawa, S. Predominant regeneration of B-cell lineage, instead of myeloid lineage, of the bone marrow after 1 Gy whole-body irradiation in mice: Role of differential cytokine expression between B-cell stimulation by IL10, Flt3 ligand and IL7 and myeloid suppression by GM-CSF and SCF. Radiat. Res. 2008, 170, 15–22. [Google Scholar]
- Shin, S.; Lee, K.; Kang, Y.; Kim, K.; Lim, S.; Yang, K.; Kim, J.; Nam, S.; Kim, H. Differential expression of immune-associated cancer regulatory genes in low-versus high-dose-rate irradiated AKR/J mice. Genomics 2011, 97, 358–363. [Google Scholar] [CrossRef]
- Werner, A.; de Vries, E.; Tait, S.; Bontjer, I.; Borst, J. Bcl-2 family member Bfl-1/A1 sequesters truncated bid to inhibit is collaboration with pro-apoptotic Bak or Bax. J. Biol. Chem. 2002, 277, 22781–22788. [Google Scholar] [CrossRef] [PubMed]
- Collins, T.; Read, M.; Neish, A.; Whitley, M.; Thanos, D.; Maniatis, T. Transcriptional regulation of endothelial cell adhesion molecules: NF-kappa B and cytokine-inducible enhancers. FASEB J. 1995, 9, 899–909. [Google Scholar] [CrossRef]
- Lanza, V.; Fadda, P.; Iannone, C.; Negri, R. Low-dose ionizing radiation stimulates transcription and production of endothelin by human vein endothelial cells. Radiat. Res. 2007, 68, 193–198. [Google Scholar] [CrossRef] [PubMed]
- Gorbunov, N.; Pogue-Geile, K.; Epperly, M.; Bigbee, W.; Draviam, R.; Day, B.; Wald, N.; Watkins, S.; Greenberger, J. Activation of the nitric oxide synthase 2 pathway in the response of bone marrow stromal cells to high doses of ionizing radiation. Radiat. Res. 2000, 154, 73–86. [Google Scholar] [CrossRef]
- Hong, C.; Kim, Y.; Pyo, H.; Lee, J.; Kim, S.; Lee, S.; Noh, J. Involvement of inducible nitric oxide synthase in radiation-induced vascular endothelial damage. J. Radiat. Res. 2013, 54, 1036–1042. [Google Scholar] [CrossRef] [PubMed]
- Coppé, J.; Desprez, P.; Krtolica, A.; Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 2010, 5, 99–118. [Google Scholar] [CrossRef] [PubMed]
- Lavorgn, A.; Harhaj, E. EBV LMP1: New and shared pathways to NF-κB activation. PNAS 2012, 109, 2188–2189. [Google Scholar] [CrossRef] [PubMed]
- Gewurz, B.E.; Towfic, F.; Mar, J.C.; Shinners, N.P.; Takasaki, K.; Zhao, B.; Cahir-McFarland, E.D.; Quackenbush, J.; Xavier, R.J.; Kieff, E. Genome-wide siRNA screen for mediators of NF-κB activation. PNAS 2012, 109, 2467–2472. [Google Scholar] [CrossRef] [PubMed]
- Jang, Y.; Jeoing, S.; Park, Y.; Bae, H.; Lee, H.; Ryu, W.; Park, G.; Son, S. UVB Induces HIF-1-dependent TSLP expression via the JNK and ERK pathways. J. Investig. Dermatol. 2013, 133, 2601–2608. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Yan, Q. Soluble factors from bone marrow endothelial cells regulate differentiation and proliferation of hematopoietic and endothelial lineages and embryonic stem cells. Acta Physiol. 2013, 65, 433–444. [Google Scholar]
- Da Silva, C.; Gonçalves, R.; Dos Santos, F.; Andrade, P.; Almeida-Porada, G.; Cabral, J. Dynamic cell-cell interactions between cord blood haematopoietic progenitors and the cellular niche are essential for the expansion of CD34(+), CD34(+)CD38(−) and early lymphoid CD7(+) cells. J. Tissue Eng. Regen. Med. 2010, 4, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Almeida-Porada, G.; Ascensão, J. Isolation, characterization, and biologic features of bone marrow endothelial cells. J. Lab. Clin. Med. 1996, 128, 399–407. [Google Scholar] [CrossRef]
- Durdik, M.; Kosik, P.; Kruzliakova, J.; Jakl, L.; Markova, E.; Belyaev, I. Hematopoietic stem/progenitor cells are less prone to undergo apoptosis than lymphocytes despite similar DNA damage response. Oncotarget 2017, 8, 48846–48853. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Neutron Dosimetry for Biology and Medicine; ICRU Report No. 26; International Commission on Radiological Units and Measurements: Bethesda, MD, USA, 1977.
- Protocol for Neutron Beam Dosimetry; AAPM REPORT No. 7; American Institute of Physics: New York, NY, USA, 1980.
- A Practical Guide to Ionization Chamber Dosimetry at the AFRRI Reactor; AFRRI Contract Report CR85-1; Armed Forces Radiobiology Research Institute: Bethesda, MD, USA, 1985; Available online: http://www.dtic.mil/docs/citations/ADA155185 (accessed on 30 March 2019).
- Smyth, G. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.; Shi, W.; Smyth, G. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Chunlei, W.; Mark, A.; Su, A. MyGene.info: Gene Annotation Query as a Service. 2014. Available online: http://biorxiv.org/content/biorxiv/early/2014/09/17/009332.full.pdf (accessed on 30 March 2019).
- Oliveros, J. VENNY. An Interactive Tool for Comparing Lists with Venn Diagrams. 2007. Available online: http://bioinfogp.cnb.csic.es/tools/venny/index.html2github.com/benfred/venn.js (accessed on 30 March 2019).
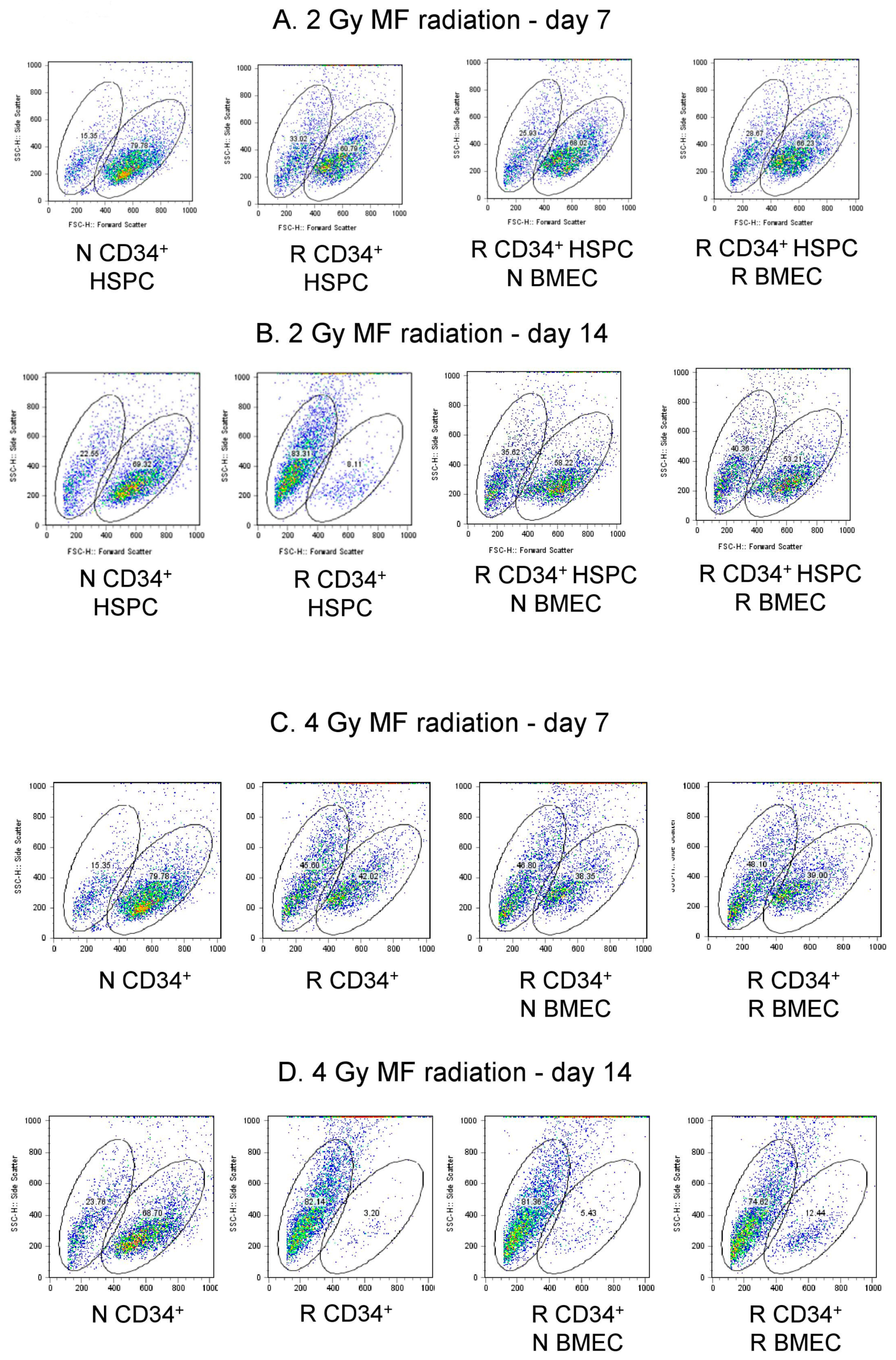
| 2 Gy | N CD34+ HSPC | R CD34+ HSPC | R CD34+ HSPC + N BMEC | R CD34+ HSPC + R BMEC |
|---|---|---|---|---|
| Scatter Population | Small Large | Small Large | Small Large | Small Large |
| Day 7 | 15.35 79.78 | 33.02 60.79 | 25.93 68.02 | 28.67 66.23 |
| Day 14 | 22.55 69.22 | 63.31 8.11 | 35.02 58.22 | 40.36 53.21 |
| 4 Gy | N CD34+ HSPC | R CD34+ HSPC | R CD34+ HSPC + N BMEC | R CD34+ HSPC + R BMEC |
|---|---|---|---|---|
| Scatter Population | Small Large | Small Large | Small Large | Small Large |
| Day 7 | 15.35 79.78 | 45.60 42.02 | 46.80 38.35 | 48.10 39.00 |
| Day 14 | 23.76 68.70 | 62.14 3.20 | 81.36 5.43 | 74.82 12.44 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cary, L.; Noutai, D.; Salber, R.; Fadiyimu, O.; Gross, A.; Almeida-Porada, G.; Kidane, Y.; Whitnall, M. Bone Marrow Endothelial Cells Influence Function and Phenotype of Hematopoietic Stem and Progenitor Cells after Mixed Neutron/Gamma Radiation. Int. J. Mol. Sci. 2019, 20, 1795. https://doi.org/10.3390/ijms20071795
Cary L, Noutai D, Salber R, Fadiyimu O, Gross A, Almeida-Porada G, Kidane Y, Whitnall M. Bone Marrow Endothelial Cells Influence Function and Phenotype of Hematopoietic Stem and Progenitor Cells after Mixed Neutron/Gamma Radiation. International Journal of Molecular Sciences. 2019; 20(7):1795. https://doi.org/10.3390/ijms20071795
Chicago/Turabian StyleCary, Lynnette, Daniel Noutai, Rudolph Salber, Opeyemi Fadiyimu, Arthur Gross, Graca Almeida-Porada, Yared Kidane, and Mark Whitnall. 2019. "Bone Marrow Endothelial Cells Influence Function and Phenotype of Hematopoietic Stem and Progenitor Cells after Mixed Neutron/Gamma Radiation" International Journal of Molecular Sciences 20, no. 7: 1795. https://doi.org/10.3390/ijms20071795
APA StyleCary, L., Noutai, D., Salber, R., Fadiyimu, O., Gross, A., Almeida-Porada, G., Kidane, Y., & Whitnall, M. (2019). Bone Marrow Endothelial Cells Influence Function and Phenotype of Hematopoietic Stem and Progenitor Cells after Mixed Neutron/Gamma Radiation. International Journal of Molecular Sciences, 20(7), 1795. https://doi.org/10.3390/ijms20071795





