The Roles of MicroRNA in Lung Cancer
Abstract
:1. Introduction
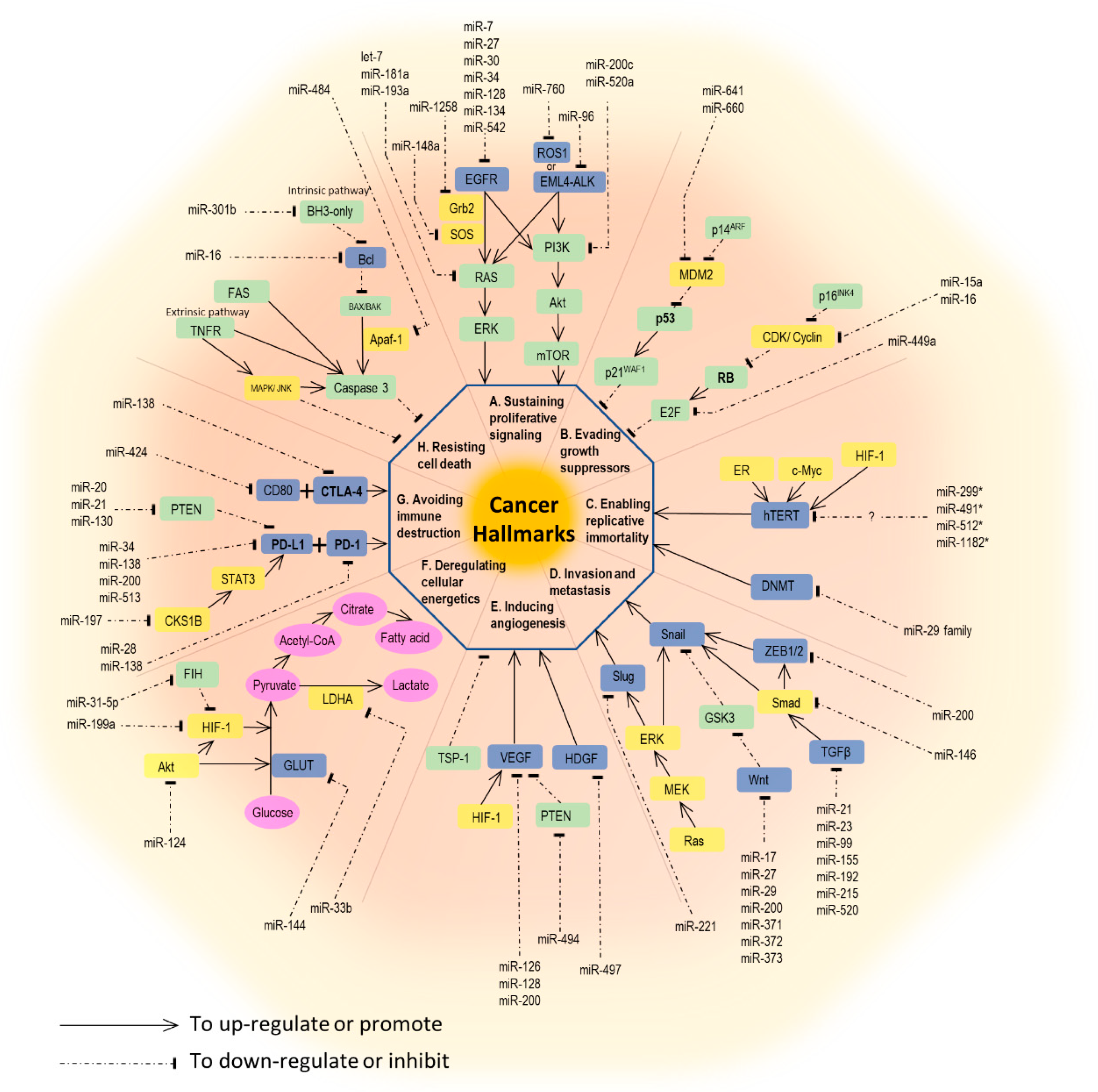
2. The Role of miRNA in Lung Cancer Development and Behavior
2.1. Sustaining Proliferative Signaling
2.2. Evading Growth Suppressors
2.3. Enabling Replicative Immortality
2.4. Activating Invasion and Metastasis
2.5. Inducing Angiogenesis
2.6. Resisting Cell Death (Apoptosis)
2.7. Deregulating Cellular Energetics
2.8. Avoiding Immune Destruction and Tumors Promoting Inflammation
2.9. Genome Instability and Mutation
3. The Interaction of Cigarette Smoke (CS) and miRNAs in Lung Cancer Tumorigenesis
4. MicroRNA and Tumor Microenvironment
4.1. Tumor-Associated Macrophages
4.2. Tumor-Associated Fibroblasts
4.3. Tumor Endothelial Cells and Tumor-Associated Angiogenesis
4.4. The Epithelial-To-Mesenchymal Transition
5. The Role of miRNAs in Lung Cancer Diagnosis
5.1. Histologic Subtype Identification
5.2. Distinguishing Primary from Metastatic Tumors
5.3. Lung Cancer Screen
6. The Role of miRNAs in Lung Cancer Prognosis
7. The Role of miRNAs in Lung Cancer Therapy
7.1. Chemotherapy
| Chemo-sensitive | Chemo-resistant | ||||
|---|---|---|---|---|---|
| miRNA | Target/pathway | Ref. | miRNA | Target/pathway | Ref. |
| let-7 | LIN28 | [112] | miR-15b | PEBP4 | [110] |
| miR-7 | EGFR | [113] | miR-21 | PTEN, SMAD7 | [114,115] |
| miR-17, 20a, 20b | TGFβR2 | [116] | |||
| miR-17 and miR-92 family | CDKN1A, RAD21 | [117] | miR-27a | RKIP | [111] |
| miR-34a | PEBP4 | [118] | miR-92a | PTEN | [119] |
| miR-101 | ROCK2 | [120] | miR-106a | ? | [121] |
| miR-106b | PKD2 | [107] | |||
| miR-135b | FZD1 | [122] | miR-181c | WIF1, Wnt/β-catenin pathway | [123] |
| miR-196 | up-regulation of MDR1, MRP1, ERCC1, Survivin and Bcl-2 | [109] | |||
| miR-137 | NUCKS1 | [124] | miR-205 | PTEN, Mcl-1 and Survivin | [125,126] |
| miR-138 | ZEB2 | [127] | miR-488 | eIF3a-mediated NER signaling pathway | [128] |
| miR-146a | CCNJ | [129] | |||
| miR-181b | TGFβR1/Smad signaling pathway | [130] | |||
| miR-184 | Bcl-2 | [131] | |||
| miR-218 | RUNX2 | [132] | |||
| mir-296 | CX3CR1 | [133] | |||
| miR-379 | EIF4G2 | [134] | |||
| miR-451 | c-Myc-survivin/rad-51 signaling | [135] | |||
| miR-503 | Down-regulation of MDR1, MRP1, ERCC1, Survivin and Bcl-2 | [108] | |||
| miR-9600 | STAT3 | [136] | |||
| Radio-sensitive | Radio-resistant | ||||
| miRNA | Target/pathway | Ref. | MiRNA | Target/pathway | Ref. |
| miR-29c | Bcl-2, Mcl-1 | [137] | miR-21 | PTEN, HIF1α, PDCD4 | [114,138] |
| miR-138 | SENP1 | [139] | miR-25 | BTG2 | [140] |
| miR-200c | PRDX2, GAPB/Nrf2, and SESN1 | [141] | miR-210 | HIF1 | [142] |
| miR-328 | γ-H2AX | [143] | miR-1323 | PRKDC | [144] |
| miR-449a | ? | [145] | |||
| miR-451 | c-Myc-survivin/rad-51 signaling | [135] | |||
| EGFR TKI-sensitive | EGFR TKI-resistant | ||||
| miRNA | Target/pathway | Ref. | miRNA | Target/pathway | Ref. |
| miR-126 | Akt and ERK pathways | [146] | miR-21 | PTEN, PDCD4 | [147] |
| miR-134/miR-487b/miR-655 cluster | MAGI2 | [148] | miR-23a | PTEN/PI3K/Akt pathway | [149] |
| miR-200c | PI3K/Akt pathway | [150] | miR-30 family | PI3K-SIAH2 | [151] |
| miR-223 | IGF1R/PI3K/Akt pathway | [152] | miR-214 | PTEN/AKT pathway | [153] |
| miR-483-3p | integrin β3/ FAK/Erk pathway | [154] | |||
| ALK TKI-sensitive | ALK TKI-resistant | ||||
| miRNA | Target/pathway | Ref. | miRNA | Target/pathway | Ref. |
| miR-200c | ZEB1 | [155] | |||
7.2. Radiation Therapy
7.3. Molecular Targeted Therapy
7.3.1. EGFR-Tyrosine Kinase Inhibitors (TKI)
7.3.2. ALK-TKI
7.4. Immunotherapy
7.5. miRNA-Targeted Therapeutics Using miRNA Mimics or Anti-miRNAs
8. Current Limitations and Future Perspectives
9. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviations
| ALK | anaplastic lymphoma kinase |
| Apaf-1 | apoptotic protease activating factor 1 |
| AXL | AXL receptor tyrosine kinase |
| Bcl | B-cell lymphoma |
| CDC42 | cell division control protein 42 homolog |
| CDK | cyclin-dependent kinase |
| CEBPB | CCAAT/enhancer binding protein - beta |
| CKS1B | CDC28 protein kinase regulatory subunit 1B |
| CTLA-4 | cytotoxic T-lymphocyte–associated antigen 4 |
| DNMT | DNA methyltransferases |
| E2F3 | E2F transcription factor 3 |
| EGFR | epidermal growth factor receptor |
| ER | estrogen receptor |
| ERCC1 | excision repair cross-complementing group 1 |
| ERK | extracellular signal-regulated kinase |
| FGFR3 | fibroblast growth factor receptor 3 |
| FIH | inhibitor of HIF-1α |
| GLUT | glucose transporter |
| Grb2 | growth factor receptor-bound protein 2 |
| GSK3 | glycogen synthase kinase-3 |
| H3K27ac | histone H3 lysine 27 acetylation |
| HDGF | hepatoma-derived growth factor |
| HIF-1 | hypoxia inducible factor-1 |
| hTERT | human telomerase reverse transcriptase |
| IGFR | insulin-like growth factor receptor |
| LDHA | lactate dehydrogenase A |
| MDM2 | mouse double minute 2 homolog |
| MDR1 | multidrug resistance p-glycoprotein 1 |
| MEK | mitogen-activated protein kinase |
| MRP1 | multidrug resistance protein 1 |
| mTOR | mammalian target of rapamycin |
| PD-1 | programmed cell death protein 1 |
| PDCD4 | programmed cell death 4 |
| PD-L1 | programmed death-ligand 1 |
| PEBP4 | phosphatidylethanolamine-binding protein 4 |
| PI3K | phosphoinositide 3-kinase |
| PIK3R1 | phosphatidylinositol 3-kinase regulatory subunit alpha |
| PPAR-γ | peroxisome proliferator-activated receptor gamma |
| PTEN | phosphatase and tensin homolog |
| Raf | rapidly accelerated fibrosarcoma |
| Ras | rat sarcoma |
| RB | retinoblastoma protein |
| RKIP | raf kinase inhibitor protein |
| siRNA | small interfering RNA |
| SOS | son of sevenless |
| SPTAN1 | spectrin alpha, non-erythrocytic 1 |
| STAT3 | signal transducer and activator of transcription 3 |
| TGF-β | transforming growth factor beta |
| TNFR | tumor necrosis factor receptor |
| TSP-1 | thrombospodin-1 |
| VEGF | vascular endothelial growth factors |
| Wnt | wingless-type MMTV integration site family |
| ZEB | zinc finger E-box-binding homeobox |
References
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Mathers, C.; Parkin, D.M.; Pineros, M.; Znaor, A.; Bray, F. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 2019, 144, 1941–1953. [Google Scholar] [CrossRef]
- Gyoba, J.; Shan, S.; Roa, W.; Bedard, E.L. Diagnosing Lung Cancers through Examination of Micro-RNA Biomarkers in Blood, Plasma, Serum and Sputum: A Review and Summary of Current Literature. Int. J. Mol. Sci. 2016, 17, 494. [Google Scholar] [CrossRef] [PubMed]
- Goulart, B.H.; Ramsey, S.D. Moving beyond the national lung screening trial: Discussing strategies for implementation of lung cancer screening programs. Oncologis 2013, 18, 941–946. [Google Scholar] [CrossRef] [PubMed]
- Morgensztern, D.; Ng, S.H.; Gao, F.; Govindan, R. Trends in Stage Distribution for Patients with Non-small Cell Lung Cancer: A National Cancer Database Survey. J. Thorac. Oncol. 2010, 5, 29–33. [Google Scholar] [CrossRef] [PubMed]
- Kanodra, N.M.; Silvestri, G.A.; Tanner, N.T. Screening and early detection efforts in lung cancer. Cancer 2015, 121, 1347–1356. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Hashemi, Z.S.; Khalili, S.; Forouzandeh Moghadam, M.; Sadroddiny, E. Lung cancer and miRNAs: A possible remedy for anti-metastatic, therapeutic and diagnostic applications. Expert Rev. Respir. Med. 2017, 11, 147–157. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Ahn, C.; Han, J.; Choi, H.; Kim, J.; Yim, J.; Lee, J.; Provost, P.; Radmark, O.; Kim, S.; et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003, 425, 415–419. [Google Scholar] [CrossRef]
- Lund, E.; Guttinger, S.; Calado, A.; Dahlberg, J.E.; Kutay, U. Nuclear export of microRNA precursors. Science 2004, 303, 95–98. [Google Scholar] [CrossRef] [PubMed]
- Gregory, R.I.; Yan, K.P.; Amuthan, G.; Chendrimada, T.; Doratotaj, B.; Cooch, N.; Shiekhattar, R. The Microprocessor complex mediates the genesis of microRNAs. Nature 2004, 432, 235–240. [Google Scholar] [CrossRef] [PubMed]
- Pasquinelli, A.E.; Ruvkun, G. Control of developmental timing by micrornas and their targets. Annu. Rev. Cell Dev. Biol. 2002, 18, 495–513. [Google Scholar] [CrossRef]
- Carmell, M.A.; Xuan, Z.; Zhang, M.Q.; Hannon, G.J. The Argonaute family: Tentacles that reach into RNAi, developmental control, stem cell maintenance, and tumorigenesis. Genes Dev. 2002, 16, 2733–2742. [Google Scholar] [CrossRef] [PubMed]
- Yang, N.; Ekanem, N.R.; Sakyi, C.A.; Ray, S.D. Hepatocellular carcinoma and microRNA: New perspectives on therapeutics and diagnostics. Adv. Drug Deliv. Rev. 2015, 81, 62–74. [Google Scholar] [CrossRef] [PubMed]
- Nana-Sinkam, S.P.; Geraci, M.W. MicroRNA in lung cancer. J. Thorac. Oncol. 2006, 1, 929–931. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Van Roosbroeck, K.; Calin, G.A. Chapter Four—Cancer Hallmarks and MicroRNAs: The Therapeutic Connection. In Advances in Cancer Research; Croce, C.M., Fisher, P.B., Eds.; Academic Press: Cambridge, MA, USA, 2017; Volume 135, pp. 119–149. [Google Scholar]
- Brambilla, E.; Gazdar, A. Pathogenesis of lung cancer signalling pathways: Roadmap for therapies. Eur. Respir. J. 2009, 33, 1485–1497. [Google Scholar] [CrossRef] [PubMed]
- Chan, L.W.; Wang, F.F.; Cho, W.C. Genomic sequence analysis of EGFR regulation by microRNAs in lung cancer. Curr. Top. Med. Chem. 2012, 12, 920–926. [Google Scholar] [CrossRef] [PubMed]
- Qin, Q.; Wei, F.; Zhang, J.; Wang, X.; Li, B. miR-134 inhibits non-small cell lung cancer growth by targeting the epidermal growth factor receptor. J. Cell. Mol. Med. 2016, 20, 1974–1983. [Google Scholar] [CrossRef] [PubMed]
- Zhu, K.; Ding, H.; Wang, W.; Liao, Z.; Fu, Z.; Hong, Y.; Zhou, Y.; Zhang, C.Y.; Chen, X. Tumor-suppressive miR-218-5p inhibits cancer cell proliferation and migration via EGFR in non-small cell lung cancer. Oncotarget 2016, 7, 28075–28085. [Google Scholar] [CrossRef] [PubMed]
- Karachaliou, N.; Rosell, R. Systemic treatment in EGFR-ALK NSCLC patients: Second line therapy and beyond. Cancer Biol. Med. 2014, 11, 173–181. [Google Scholar] [CrossRef]
- Vishwamitra, D.; Li, Y.; Wilson, D.; Manshouri, R.; Curry, C.V.; Shi, B.; Tang, X.M.; Sheehan, A.M.; Wistuba, I.I.; Shi, P.; et al. MicroRNA 96 is a post-transcriptional suppressor of anaplastic lymphoma kinase expression. Am. J. Pathol. 2012, 180, 1772–1780. [Google Scholar] [CrossRef]
- Sehgal, K.; Patell, R.; Rangachari, D.; Costa, D.B. Targeting ROS1 rearrangements in non-small cell lung cancer with crizotinib and other kinase inhibitors. Transl. Cancer Res. 2018, 7, S779–S786. [Google Scholar] [CrossRef]
- Davies, K.D.; Doebele, R.C. Molecular pathways: ROS1 fusion proteins in cancer. Clin. Cancer Res. 2013, 19, 4040–4045. [Google Scholar] [CrossRef]
- Yan, C.; Zhang, W.; Shi, X.; Zheng, J.; Jin, X.; Huo, J. MiR-760 suppresses non-small cell lung cancer proliferation and metastasis by targeting ROS1. Environ. Sci. Pollut. Res. Int. 2018, 25, 18385–18391. [Google Scholar] [CrossRef]
- Friday, B.B.; Adjei, A.A. K-ras as a target for cancer therapy. Biochim. Biophys. Acta 2005, 1756, 127–144. [Google Scholar] [CrossRef] [PubMed]
- Kopp, F.; Wagner, E.; Roidl, A. The proto-oncogene KRAS is targeted by miR-200c. Oncotarget 2014, 5, 185–195. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.Y.; Ren, T.; Cai, Y.Y.; He, X.Y. MicroRNA let-7a inhibits the proliferation and invasion of nonsmall cell lung cancer cell line 95D by regulating K-Ras and HMGA2 gene expression. Cancer Biother. Radiopharm. 2013, 28, 131–137. [Google Scholar] [CrossRef]
- He, X.Y.; Chen, J.X.; Zhang, Z.; Li, C.L.; Peng, Q.L.; Peng, H.M. The let-7a microRNA protects from growth of lung carcinoma by suppression of k-Ras and c-Myc in nude mice. J. Cancer Res. Clin. Oncol. 2010, 136, 1023–1028. [Google Scholar] [CrossRef] [PubMed]
- Seviour, E.G.; Sehgal, V.; Mishra, D.; Rupaimoole, R.; Rodriguez-Aguayo, C.; Lopez-Berestein, G.; Lee, J.S.; Sood, A.K.; Kim, M.P.; Mills, G.B.; et al. Targeting KRas-dependent tumour growth, circulating tumour cells and metastasis in vivo by clinically significant miR-193a-3p. Oncogene 2017, 36, 1339–1350. [Google Scholar] [CrossRef]
- Ma, Z.; Qiu, X.; Wang, D.; Li, Y.; Zhang, B.; Yuan, T.; Wei, J.; Zhao, B.; Zhao, X.; Lou, J.; et al. MiR-181a-5p inhibits cell proliferation and migration by targeting Kras in non-small cell lung cancer A549 cells. Acta Biochim. Biophys. Sin. 2015, 47, 630–638. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Yu, Z.; Lu, Y.; Fan, J.; Ni, Y.; Ma, L. microRNA-148a-3p inhibited the proliferation and epithelial-mesenchymal transition progression of non-small-cell lung cancer via modulating Ras/MAPK/Erk signaling. J. Cell. Physiol. 2018. [Google Scholar] [CrossRef]
- Jiang, W.; Wei, K.; Pan, C.; Li, H.; Cao, J.; Han, X.; Tang, Y.; Zhu, S.; Yuan, W.; He, Y.; et al. MicroRNA-1258 suppresses tumour progression via GRB2/Ras/Erk pathway in non-small-cell lung cancer. Cell Prolif. 2018, 51, e12502. [Google Scholar] [CrossRef]
- Lv, X.; Li, C.Y.; Han, P.; Xu, X.Y. MicroRNA-520a-3p inhibits cell growth and metastasis of non-small cell lung cancer through PI3K/AKT/mTOR signaling pathway. Eur. Rev. Med Pharmacol. Sci. 2018, 22, 2321–2327. [Google Scholar] [CrossRef]
- Bandi, N.; Zbinden, S.; Gugger, M.; Arnold, M.; Kocher, V.; Hasan, L.; Kappeler, A.; Brunner, T.; Vassella, E. miR-15a and miR-16 are implicated in cell cycle regulation in a Rb-dependent manner and are frequently deleted or down-regulated in non-small cell lung cancer. Cancer Res 2009, 69, 5553–5559. [Google Scholar] [CrossRef] [PubMed]
- Ren, X.S.; Yin, M.H.; Zhang, X.; Wang, Z.; Feng, S.P.; Wang, G.X.; Luo, Y.J.; Liang, P.Z.; Yang, X.Q.; He, J.X.; et al. Tumor-suppressive microRNA-449a induces growth arrest and senescence by targeting E2F3 in human lung cancer cells. Cancer Lett. 2014, 344, 195–203. [Google Scholar] [CrossRef] [PubMed]
- Kong, Q.; Shu, N.; Li, J.; Xu, N. miR-641 Functions as a Tumor Suppressor by Targeting MDM2 in Human Lung Cancer. Oncol. Res. 2018, 26, 735–741. [Google Scholar] [CrossRef] [PubMed]
- Fortunato, O.; Boeri, M.; Moro, M.; Verri, C.; Mensah, M.; Conte, D.; Caleca, L.; Roz, L.; Pastorino, U.; Sozzi, G. Mir-660 is downregulated in lung cancer patients and its replacement inhibits lung tumorigenesis by targeting MDM2-p53 interaction. Cell Death Dis. 2014, 5, e1564. [Google Scholar] [CrossRef] [PubMed]
- Kyo, S.; Takakura, M.; Fujiwara, T.; Inoue, M. Understanding and exploiting hTERT promoter regulation for diagnosis and treatment of human cancers. Cancer Sci. 2008, 99, 1528–1538. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lei, H.; Xu, Y.; Tao, Z.Z. miR-512-5p suppresses tumor growth by targeting hTERT in telomerase positive head and neck squamous cell carcinoma in vitro and in vivo. PLoS ONE 2015, 10, e0135265. [Google Scholar] [CrossRef]
- Kasiappan, R.; Shen, Z.; Tse, A.K.; Jinwal, U.; Tang, J.; Lungchukiet, P.; Sun, Y.; Kruk, P.; Nicosia, S.V.; Zhang, X.; et al. 1,25-Dihydroxyvitamin D3 suppresses telomerase expression and human cancer growth through microRNA-498. J. Biol. Chem. 2012, 287, 41297–41309. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, M.; Garzon, R.; Cimmino, A.; Liu, Z.; Zanesi, N.; Callegari, E.; Liu, S.; Alder, H.; Costinean, S.; Fernandez-Cymering, C.; et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc. Natl. Acad. Sci. USA 2007, 104, 15805–15810. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Pertsemlidis, A. microRNAs and lung cancer: Tumors and 22-mers. Cancer Metastasis Rev. 2010, 29, 109–122. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Gibbons, D.L.; Goswami, S.; Cortez, M.A.; Ahn, Y.H.; Byers, L.A.; Zhang, X.; Yi, X.; Dwyer, D.; Lin, W.; et al. Metastasis is regulated via microRNA-200/ZEB1 axis control of tumour cell PD-L1 expression and intratumoral immunosuppression. Nat. Commun. 2014, 5, 5241. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, L.; Chen, C.; Chu, X. New insights into the regulatory role of microRNA in tumor angiogenesis and clinical implications. Mol. Cancer 2018, 17, 22. [Google Scholar] [CrossRef]
- Yamakuchi, M.; Lotterman, C.D.; Bao, C.; Hruban, R.H.; Karim, B.; Mendell, J.T.; Huso, D.; Lowenstein, C.J. P53-induced microRNA-107 inhibits HIF-1 and tumor angiogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 6334–6339. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.C.; Yoon, S.; Jeong, Y.; Yoon, J.; Baek, K. Regulation of vascular endothelial growth factor signaling by miR-200b. Mol. Cells 2011, 32, 77–82. [Google Scholar] [CrossRef] [PubMed]
- Pecot, C.V.; Rupaimoole, R.; Yang, D.; Akbani, R.; Ivan, C.; Lu, C.; Wu, S.; Han, H.D.; Shah, M.Y.; Rodriguez-Aguayo, C.; et al. Tumour angiogenesis regulation by the miR-200 family. Nat. Commun. 2013, 4, 2427. [Google Scholar] [CrossRef]
- Liu, B.; Peng, X.C.; Zheng, X.L.; Wang, J.; Qin, Y.W. MiR-126 restoration down-regulate VEGF and inhibit the growth of lung cancer cell lines in vitro and in vivo. Lung Cancer 2009, 66, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Cheng, Y.; Li, Y.; Jin, Z.; Pan, Y.; Liu, G.; Fu, S.; Zhang, Y.; Feng, K.; Feng, Y. microRNA-128 plays a critical role in human non-small cell lung cancer tumourigenesis, angiogenesis and lymphangiogenesis by directly targeting vascular endothelial growth factor-C. Eur. J. Cancer 2014, 50, 2336–2350. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.Y.; Wang, Y.; An, Z.J.; Shi, C.G.; Zhu, G.A.; Wang, B.; Lu, M.Y.; Pan, C.K.; Chen, P. Downregulation of miR-497 promotes tumor growth and angiogenesis by targeting HDGF in non-small cell lung cancer. Biochem. Biophys. Res. Commun. 2013, 435, 466–471. [Google Scholar] [CrossRef]
- Mao, G.; Liu, Y.; Fang, X.; Liu, Y.; Fang, L.; Lin, L.; Liu, X.; Wang, N. Tumor-derived microRNA-494 promotes angiogenesis in non-small cell lung cancer. Angiogenesis 2015, 18, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.L.; Hung, J.Y.; Chang, W.A.; Lin, Y.S.; Pan, Y.C.; Tsai, P.H.; Wu, C.Y.; Kuo, P.L. Hypoxic lung cancer-secreted exosomal miR-23a increased angiogenesis and vascular permeability by targeting prolyl hydroxylase and tight junction protein ZO-1. Oncogene 2017, 36, 4929–4942. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Chen, J.; Dai, J.; Zhang, B.; Wang, F.; Sun, Y. MicroRNA-16-1 Inhibits Tumor Cell Proliferation and Induces Apoptosis in A549 Non-Small Cell Lung Carcinoma Cells. Oncol. Res. 2016, 24, 345–351. [Google Scholar] [CrossRef]
- Tian, J.; Hu, L.; Li, X.; Geng, J.; Dai, M.; Bai, X. MicroRNA-130b promotes lung cancer progression via PPARgamma/VEGF-A/BCL-2-mediated suppression of apoptosis. J. Exp. Clin. Cancer Res. 2016, 35, 105. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Gao, J.; Huang, Q.; Jin, Y.; Wei, Z. Downregulating microRNA-144 mediates a metabolic shift in lung cancer cells by regulating GLUT1 expression. Oncol. Lett. 2016, 11, 3772–3776. [Google Scholar] [CrossRef] [PubMed]
- Zhai, S.; Zhao, L.; Lin, T.; Wang, W. Downregulation of miR-33b promotes non-small cell lung cancer cell growth through reprogramming glucose metabolism miR-33b regulates non-small cell lung cancer cell growth. J. Cell. Biochem. 2019, 120, 6651–6660. [Google Scholar] [CrossRef] [PubMed]
- Ding, G.; Huang, G.; Liu, H.D.; Liang, H.X.; Ni, Y.F.; Ding, Z.H.; Ni, G.Y.; Hua, H.W. MiR-199a suppresses the hypoxia-induced proliferation of non-small cell lung cancer cells through targeting HIF1alpha. Mol. Cell. Biochem. 2013, 384, 173–180. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Cao, X.; Zhang, W.; Pan, G.; Yi, Q.; Zhong, W.; Yan, D. MicroRNA-31-5p enhances the Warburg effect via targeting FIH. FASEB J. 2018, 33, 545–556. [Google Scholar] [CrossRef] [PubMed]
- Cortez, M.A.; Ivan, C.; Valdecanas, D.; Wang, X.; Peltier, H.J.; Ye, Y.; Araujo, L.; Carbone, D.P.; Shilo, K.; Giri, D.K.; et al. PDL1 Regulation by p53 via miR-34. J. Natl. Cancer Inst. 2015, 108, djv303. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Yagishita, S.; Hagiwara, K.; Yoshioka, Y.; Kosaka, N.; Takeshita, F.; Fujiwara, T.; Tsuta, K.; Nokihara, H.; Tamura, T.; et al. The clinical relevance of the miR-197/CKS1B/STAT3-mediated PD-L1 network in chemoresistant non-small-cell lung cancer. Mol. Ther. 2015, 23, 717–727. [Google Scholar] [CrossRef]
- Yao, Y.; Dai, W. Genomic Instability and Cancer. J. Carcinog. Mutagenesis 2014, 5. [Google Scholar] [CrossRef]
- Vincent, K.; Pichler, M.; Lee, G.W.; Ling, H. MicroRNAs, genomic instability and cancer. Int. J. Mol. Sci. 2014, 15, 14475–14491. [Google Scholar] [CrossRef]
- Calin, G.A.; Sevignani, C.; Dumitru, C.D.; Hyslop, T.; Noch, E.; Yendamuri, S.; Shimizu, M.; Rattan, S.; Bullrich, F.; Negrini, M.; et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl. Acad. Sci. USA 2004, 101, 2999–3004. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Liu, C.; Niu, Y.; Jing, Y.; Zhang, H.; Wang, J.; Yang, J.; Zen, K.; Zhang, J.; Zhang, C.Y.; et al. MicroRNA-128-3p regulates mitomycin C-induced DNA damage response in lung cancer cells through repressing SPTAN1. Oncotarget 2017, 8, 58098–58107. [Google Scholar] [CrossRef] [PubMed]
- Gandini, S.; Botteri, E.; Iodice, S.; Boniol, M.; Lowenfels, A.B.; Maisonneuve, P.; Boyle, P. Tobacco smoking and cancer: A meta-analysis. Int. J. Cancer 2008, 122, 155–164. [Google Scholar] [CrossRef]
- Leon, M.E.; Peruga, A.; McNeill, A.; Kralikova, E.; Guha, N.; Minozzi, S.; Espina, C.; Schüz, J. European Code against Cancer, 4th Edition: Tobacco and cancer. Cancer Epidemiol. 2015, 39, S20–S33. [Google Scholar] [CrossRef] [PubMed]
- Hecht, S.S. Lung carcinogenesis by tobacco smoke. Int. J. Cancer 2012, 131, 2724–2732. [Google Scholar] [CrossRef] [PubMed]
- Buro-Auriemma, L.J.; Salit, J.; Hackett, N.R.; Walters, M.S.; Strulovici-Barel, Y.; Staudt, M.R.; Fuller, J.; Mahmoud, M.; Stevenson, C.S.; Hilton, H.; et al. Cigarette smoking induces small airway epithelial epigenetic changes with corresponding modulation of gene expression. Hum. Mol. Genet. 2013, 22, 4726–4738. [Google Scholar] [CrossRef]
- Wang, T.-H.; Hsia, S.-M.; Shih, Y.-H.; Shieh, T.-M. Association of Smoking, Alcohol Use, and Betel Quid Chewing with Epigenetic Aberrations in Cancers. Int. J. Mol. Sci. 2017, 18, 1210. [Google Scholar] [CrossRef] [PubMed]
- Sundar, I.K.; Nevid, M.Z.; Friedman, A.E.; Rahman, I. Cigarette smoke induces distinct histone modifications in lung cells: Implications for the pathogenesis of COPD and lung cancer. J. Proteome Res. 2014, 13, 982–996. [Google Scholar] [CrossRef]
- Advani, J.; Subbannayya, Y.; Patel, K.; Khan, A.A.; Patil, A.H.; Jain, A.P.; Solanki, H.S.; Radhakrishnan, A.; Pinto, S.M.; Sahasrabuddhe, N.A.; et al. Long-Term Cigarette Smoke Exposure and Changes in MiRNA Expression and Proteome in Non-Small-Cell Lung Cancer. OMICS 2017, 21, 390–403. [Google Scholar] [CrossRef]
- Lu, L.; Xu, H.; Yang, P.; Xue, J.; Chen, C.; Sun, Q.; Yang, Q.; Lu, J.; Shi, A.; Liu, Q. Involvement of HIF-1alpha-regulated miR-21, acting via the Akt/NF-kappaB pathway, in malignant transformation of HBE cells induced by cigarette smoke extract. Toxicol. Lett. 2018, 289, 14–21. [Google Scholar] [CrossRef]
- Du, X.; Qi, F.; Lu, S.; Li, Y.; Han, W. Nicotine upregulates FGFR3 and RB1 expression and promotes non-small cell lung cancer cell proliferation and epithelial-to-mesenchymal transition via downregulation of miR-99b and miR-192. Biomed. Pharmacother. 2018, 101, 656–662. [Google Scholar] [CrossRef] [PubMed]
- Kuninty, P.R.; Schnittert, J.; Storm, G.; Prakash, J. MicroRNA Targeting to Modulate Tumor Microenvironment. Front. Oncol. 2016, 6, 3. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.I.; Katsura, A.; Matsuyama, H.; Miyazono, K. MicroRNA regulons in tumor microenvironment. Oncogene 2015, 34, 3085–3094. [Google Scholar] [CrossRef] [PubMed]
- Fortunato, O.; Borzi, C.; Milione, M.; Centonze, G.; Conte, D.; Boeri, M.; Verri, C.; Moro, M.; Facchinetti, F.; Andriani, F.; et al. Circulating mir-320a promotes immunosuppressive macrophages M2 phenotype associated with lung cancer risk. Int. J. Cancer 2018. [Google Scholar] [CrossRef]
- Lin, L.; Lin, H.; Wang, L.; Wang, B.; Hao, X.; Shi, Y. miR-130a regulates macrophage polarization and is associated with non-small cell lung cancer. Oncol. Rep. 2015, 34, 3088–3096. [Google Scholar] [CrossRef]
- Kunita, A.; Morita, S.; Irisa, T.U.; Goto, A.; Niki, T.; Takai, D.; Nakajima, J.; Fukayama, M. MicroRNA-21 in cancer-associated fibroblasts supports lung adenocarcinoma progression. Sci. Rep. 2018, 8, 8838. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Liu, J.; Liu, Y.; Wu, W.; Li, X.; Wu, Y.; Chen, H.; Zhang, K.; Gu, L. miR-101 represses lung cancer by inhibiting interaction of fibroblasts and cancer cells by down-regulating CXCL12. Biomed. Pharmacother. 2015, 74, 215–221. [Google Scholar] [CrossRef]
- Li, J.; Guan, J.; Long, X.; Wang, Y.; Xiang, X. mir-1-mediated paracrine effect of cancer-associated fibroblasts on lung cancer cell proliferation and chemoresistance. Oncol. Rep. 2016, 35, 3523–3531. [Google Scholar] [CrossRef]
- Baluk, P.; Hashizume, H.; McDonald, D.M. Cellular abnormalities of blood vessels as targets in cancer. Curr. Opin. Genet. Dev. 2005, 15, 102–111. [Google Scholar] [CrossRef]
- Wu, F.; Yang, Z.; Li, G. Role of specific microRNAs for endothelial function and angiogenesis. Biochem. Biophys. Res. Commun. 2009, 386, 549–553. [Google Scholar] [CrossRef] [PubMed]
- Donnem, T.; Fenton, C.G.; Lonvik, K.; Berg, T.; Eklo, K.; Andersen, S.; Stenvold, H.; Al-Shibli, K.; Al-Saad, S.; Bremnes, R.M.; et al. MicroRNA signatures in tumor tissue related to angiogenesis in non-small cell lung cancer. PLoS ONE 2012, 7, e29671. [Google Scholar] [CrossRef]
- Rupaimoole, R.; Calin, G.A.; Lopez-Berestein, G.; Sood, A.K. miRNA Deregulation in Cancer Cells and the Tumor Microenvironment. Cancer Discov. 2016, 6, 235–246. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; Neilson, E.G. Epithelial-mesenchymal transition and its implications for fibrosis. J. Clin. Investig. 2003, 112, 1776–1784. [Google Scholar] [CrossRef]
- Ke, Y.; Zhao, W.; Xiong, J.; Cao, R. miR-149 Inhibits Non-Small-Cell Lung Cancer Cells EMT by Targeting FOXM1. Biochem. Res. Int. 2013, 2013, 506731. [Google Scholar] [CrossRef]
- Yang, L.; Yang, J.; Li, J.; Shen, X.; Le, Y.; Zhou, C.; Wang, S.; Zhang, S.; Xu, D.; Gong, Z. MircoRNA-33a inhibits epithelial-to-mesenchymal transition and metastasis and could be a prognostic marker in non-small cell lung cancer. Sci. Rep. 2015, 5, 13677. [Google Scholar] [CrossRef]
- Bishop, J.A.; Benjamin, H.; Cholakh, H.; Chajut, A.; Clark, D.P.; Westra, W.H. Accurate classification of non-small cell lung carcinoma using a novel microRNA-based approach. Clin Cancer Res. 2010, 16, 610–619. [Google Scholar] [CrossRef] [PubMed]
- Lebanony, D.; Benjamin, H.; Gilad, S.; Ezagouri, M.; Dov, A.; Ashkenazi, K.; Gefen, N.; Izraeli, S.; Rechavi, G.; Pass, H.; et al. Diagnostic assay based on hsa-miR-205 expression distinguishes squamous from nonsquamous non-small-cell lung carcinoma. J. Clin. Oncol. 2009, 27, 2030–2037. [Google Scholar] [CrossRef] [PubMed]
- Lujambio, A.; Ropero, S.; Ballestar, E.; Fraga, M.F.; Cerrato, C.; Setien, F.; Casado, S.; Suarez-Gauthier, A.; Sanchez-Cespedes, M.; Git, A.; et al. Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res. 2007, 67, 1424–1429. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.K.; Zhu, W.Y.; He, J.Y.; Chen, D.D.; Huang, Y.Y.; Le, H.B.; Liu, X.G. miRNAs expression profiling to distinguish lung squamous-cell carcinoma from adenocarcinoma subtypes. J. Cancer Res. Clin. Oncol. 2012, 138, 1641–1650. [Google Scholar] [CrossRef] [PubMed]
- Nishikawa, E.; Osada, H.; Okazaki, Y.; Arima, C.; Tomida, S.; Tatematsu, Y.; Taguchi, A.; Shimada, Y.; Yanagisawa, K.; Yatabe, Y.; et al. miR-375 Is Activated by ASH1 and Inhibits YAP1 in a Lineage-Dependent Manner in Lung Cancer. Cancer Res. 2011, 71, 6165–6173. [Google Scholar] [CrossRef] [PubMed]
- Demes, M.; Aszyk, C.; Bartsch, H.; Schirren, J.; Fisseler-Eckhoff, A. Differential miRNA-Expression as an Adjunctive Diagnostic Tool in Neuroendocrine Tumors of the Lung. Cancers 2016, 8, 38. [Google Scholar] [CrossRef]
- Nadal, E.; Zhong, J.; Lin, J.; Reddy, R.M.; Ramnath, N.; Orringer, M.B.; Chang, A.C.; Beer, D.G.; Chen, G. A MicroRNA Cluster at 14q32 Drives Aggressive Lung Adenocarcinoma. Clin. Cancer Res. 2014, 20, 3107–3117. [Google Scholar] [CrossRef]
- Gilad, S.; Lithwick-Yanai, G.; Barshack, I.; Benjamin, S.; Krivitsky, I.; Edmonston, T.B.; Bibbo, M.; Thurm, C.; Horowitz, L.; Huang, Y.; et al. Classification of the Four Main Types of Lung Cancer Using a MicroRNA-Based Diagnostic Assay. J. Mol. Diagn. 2012, 14, 510–517. [Google Scholar] [CrossRef]
- Rosenwald, S.; Gilad, S.; Benjamin, S.; Lebanony, D.; Dromi, N.; Faerman, A.; Benjamin, H.; Tamir, R.; Ezagouri, M.; Goren, E.; et al. Validation of a microRNA-based qRT-PCR test for accurate identification of tumor tissue origin. Mod. Pathol. 2010, 23, 814. [Google Scholar] [CrossRef]
- Barshack, I.; Lithwick-Yanai, G.; Afek, A.; Rosenblatt, K.; Tabibian-Keissar, H.; Zepeniuk, M.; Cohen, L.; Dan, H.; Zion, O.; Strenov, Y.; et al. MicroRNA expression differentiates between primary lung tumors and metastases to the lung. Pathology 2010, 206, 578–584. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Lim, N.J.; Jang, S.G.; Kim, H.K.; Lee, G.K. miR-592 and miR-552 can distinguish between primary lung adenocarcinoma and colorectal cancer metastases in the lung. Anticancer Res. 2014, 34, 2297–2302. [Google Scholar] [PubMed]
- Han, Y.; Li, H. miRNAs as biomarkers and for the early detection of non-small cell lung cancer (NSCLC). J. Thorac. Dis. 2018, 10, 3119–3131. [Google Scholar] [CrossRef]
- Montani, F.; Marzi, M.J.; Dezi, F.; Dama, E.; Carletti, R.M.; Bonizzi, G.; Bertolotti, R.; Bellomi, M.; Rampinelli, C.; Maisonneuve, P.; et al. miR-Test: A blood test for lung cancer early detection. J. Natl. Cancer Inst. 2015, 107, djv063. [Google Scholar] [CrossRef]
- Sozzi, G.; Boeri, M.; Rossi, M.; Verri, C.; Suatoni, P.; Bravi, F.; Roz, L.; Conte, D.; Grassi, M.; Sverzellati, N.; et al. Clinical utility of a plasma-based miRNA signature classifier within computed tomography lung cancer screening: A correlative MILD trial study. J. Clin. Oncol. 2014, 32, 768–773. [Google Scholar] [CrossRef] [PubMed]
- Takamizawa, J.; Konishi, H.; Yanagisawa, K.; Tomida, S.; Osada, H.; Endoh, H.; Harano, T.; Yatabe, Y.; Nagino, M.; Nimura, Y.; et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 2004, 64, 3753–3756. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Zhu, Y.; Zhou, X.; Chen, Y. Low expression of let-7 predicts poor prognosis in patients with multiple cancers: A meta-analysis. Tumour Biol. 2014, 35, 5143–5148. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Shen, H.; Qiu, C.; Ni, Y.; Wang, L.; Dong, W.; Liao, Y.; Du, J. High expression of miR-21 and miR-155 predicts recurrence and unfavourable survival in non-small cell lung cancer. Eur. J. Cancer 2013, 49, 604–615. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Zhan, Y.; Feng, J.; Luo, J.; Fan, S. MicroRNAs associated with therapy of non-small cell lung cancer. Int. J. Biol. Sci. 2018, 14, 390–397. [Google Scholar] [CrossRef]
- Yu, S.; Qin, X.; Chen, T.; Zhou, L.; Xu, X.; Feng, J. MicroRNA-106b-5p regulates cisplatin chemosensitivity by targeting polycystic kidney disease-2 in non-small-cell lung cancer. Anti-Cancer Drugs 2017, 28, 852–860. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Guo, L.; Liu, J.; Liu, R.; Liu, M.; Chen, J. The reversing and molecular mechanisms of miR-503 on the drug-resistance to cisplatin in A549/DDP cells. Zhongguo Fei Ai Za Zhi 2014, 17, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Li, J.H.; Luo, N.; Zhong, M.Z.; Xiao, Z.Q.; Wang, J.X.; Yao, X.Y.; Peng, Y.; Cao, J. Inhibition of microRNA-196a might reverse cisplatin resistance of A549/DDP non-small-cell lung cancer cell line. Tumour Biol. 2016, 37, 2387–2394. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Zhang, L.; Yao, Q.; Tao, Z. miR-15b regulates cisplatin resistance and metastasis by targeting PEBP4 in human lung adenocarcinoma cells. Cancer Gene Ther. 2015, 22, 108–114. [Google Scholar] [CrossRef]
- Li, J.; Wang, Y.; Song, Y.; Fu, Z.; Yu, W. miR-27a regulates cisplatin resistance and metastasis by targeting RKIP in human lung adenocarcinoma cells. Mol. Cancer 2014, 13, 193. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Zhao, J.; Hu, W.; Yang, G.; Yu, H.; Wang, R.; Wang, L.; Zhang, G.; Fu, W.; Dai, L.; et al. Disturbance of the let-7/LIN28 double-negative feedback loop is associated with radio- and chemo-resistance in non-small cell lung cancer. PLoS ONE 2017, 12, e0172787. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Liu, X.; Zheng, Y.; Gu, J.; Xiong, S.; Jiang, P.; Jiang, X.; Huang, E.; Yang, Y.; Ge, D.; et al. MicroRNA-7 sensitizes non-small cell lung cancer cells to paclitaxel. Oncol. Lett. 2014, 8, 2193–2200. [Google Scholar] [CrossRef] [PubMed]
- Bica-Pop, C.; Cojocneanu-Petric, R.; Magdo, L.; Raduly, L.; Gulei, D.; Berindan-Neagoe, I. Overview upon miR-21 in lung cancer: Focus on NSCLC. Cell. Mol. Life Sci. 2018, 75, 3539–3551. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Tu, H.B.; Wu, L.; Liu, M.; Jiang, G.N. MicroRNA-21 Regulates Non-Small Cell Lung Cancer Cell Invasion and Chemo-Sensitivity through SMAD7. Cell. Physiol. Biochem. 2016, 38, 2152–2162. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Yin, J.; Fu, W.; Mo, Y.; Pan, Y.; Dai, L.; Huang, H.; Li, S.; Zhao, J. MiRNA 17 family regulates cisplatin-resistant and metastasis by targeting TGFbetaR2 in NSCLC. PLoS ONE 2014, 9, e94639. [Google Scholar] [CrossRef]
- Zhao, J.; Fu, W.; Liao, H.; Dai, L.; Jiang, Z.; Pan, Y.; Huang, H.; Mo, Y.; Li, S.; Yang, G.; et al. The regulatory and predictive functions of miR-17 and miR-92 families on cisplatin resistance of non-small cell lung cancer. BMC Cancer 2015, 15, 731. [Google Scholar] [CrossRef]
- Yu, G.; Zhong, N.; Chen, G.; Huang, B.; Wu, S. Downregulation of PEBP4, a target of miR-34a, sensitizes drug-resistant lung cancer cells. Tumour Biol. 2014, 35, 10341–10349. [Google Scholar] [CrossRef]
- Ren, P.; Gong, F.; Zhang, Y.; Jiang, J.; Zhang, H. MicroRNA-92a promotes growth, metastasis, and chemoresistance in non-small cell lung cancer cells by targeting PTEN. Tumour Biol. 2016, 37, 3215–3225. [Google Scholar] [CrossRef] [PubMed]
- Ye, Z.; Yin, S.; Su, Z.; Bai, M.; Zhang, H.; Hei, Z.; Cai, S. Downregulation of miR-101 contributes to epithelial-mesenchymal transition in cisplatin resistance of NSCLC cells by targeting ROCK2. Oncotarget 2016, 7, 37524–37535. [Google Scholar] [CrossRef]
- Tian, Y.; Sun, C.; Zhang, L.; Pan, Y. Clinical significance of miRNA—106a in non-small cell lung cancer patients who received cisplatin combined with gemcitabine chemotherapy. Cancer Biol. Med. 2018, 15, 157–164. [Google Scholar] [CrossRef]
- Su, W.; Mo, Y.; Wu, F.; Guo, K.; Li, J.; Luo, Y.; Ye, H.; Guo, H.; Li, D.; Yang, Z. miR-135b reverses chemoresistance of non-small cell lung cancer cells by downregulation of FZD1. Biomed. Pharmacother. 2016, 84, 123–129. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Hu, B.; Wang, Z.; Zhang, F.; Wei, H.; Li, L. miR-181c contributes to cisplatin resistance in non-small cell lung cancer cells by targeting Wnt inhibition factor 1. Cancer Chemother. Pharmacol. 2017, 80, 973–984. [Google Scholar] [CrossRef]
- Shen, H.; Wang, L.; Ge, X.; Jiang, C.F.; Shi, Z.M.; Li, D.M.; Liu, W.T.; Yu, X.; Shu, Y.Q. MicroRNA-137 inhibits tumor growth and sensitizes chemosensitivity to paclitaxel and cisplatin in lung cancer. Oncotarget 2016, 7, 20728–20742. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Huang, Y.; Gong, W. miR-205 promotes the growth, metastasis and chemoresistance of NSCLC cells by targeting PTEN. Oncol. Rep. 2013, 30, 2897–2902. [Google Scholar] [CrossRef]
- Zarogoulidis, P.; Petanidis, S.; Kioseoglou, E.; Domvri, K.; Anestakis, D.; Zarogoulidis, K. MiR-205 and miR-218 expression is associated with carboplatin chemoresistance and regulation of apoptosis via Mcl-1 and Survivin in lung cancer cells. Cell. Signal. 2015, 27, 1576–1588. [Google Scholar] [CrossRef] [PubMed]
- Jin, Z.; Guan, L.; Song, Y.; Xiang, G.M.; Chen, S.X.; Gao, B. MicroRNA-138 regulates chemoresistance in human non-small cell lung cancer via epithelial mesenchymal transition. Eur. Rev. Med Pharmacol. Sci. 2016, 20, 1080–1086. [Google Scholar] [PubMed]
- Fang, C.; Chen, Y.X.; Wu, N.Y.; Yin, J.Y.; Li, X.P.; Huang, H.S.; Zhang, W.; Zhou, H.H.; Liu, Z.Q. MiR-488 inhibits proliferation and cisplatin sensibility in non-small-cell lung cancer (NSCLC) cells by activating the eIF3a-mediated NER signaling pathway. Sci. Rep. 2017, 7, 40384. [Google Scholar] [CrossRef]
- Shi, L.; Xu, Z.; Wu, G.; Chen, X.; Huang, Y.; Wang, Y.; Jiang, W.; Ke, B. Up-regulation of miR-146a increases the sensitivity of non-small cell lung cancer to DDP by downregulating cyclin J. BMC Cancer 2017, 17, 138. [Google Scholar] [CrossRef]
- Wang, X.; Chen, X.; Meng, Q.; Jing, H.; Lu, H.; Yang, Y.; Cai, L.; Zhao, Y. MiR-181b regulates cisplatin chemosensitivity and metastasis by targeting TGFbetaR1/Smad signaling pathway in NSCLC. Sci. Rep. 2015, 5, 17618. [Google Scholar] [CrossRef]
- Tung, M.C.; Lin, P.L.; Cheng, Y.W.; Wu, D.W.; Yeh, S.D.; Chen, C.Y.; Lee, H. Reduction of microRNA-184 by E6 oncoprotein confers cisplatin resistance in lung cancer via increasing Bcl-2. Oncotarget 2016, 7, 32362–32374. [Google Scholar] [CrossRef]
- Xie, J.; Yu, F.; Li, D.; Zhu, X.; Zhang, X.; Lv, Z. MicroRNA-218 regulates cisplatin (DPP) chemosensitivity in non-small cell lung cancer by targeting RUNX2. Tumour Biol. 2016, 37, 1197–1204. [Google Scholar] [CrossRef]
- Luo, W.; Lin, Y.; Meng, S.; Guo, Y.; Zhang, J.; Zhang, W. miRNA-296-3p modulates chemosensitivity of lung cancer cells by targeting CX3CR1. Am. J. Transl. Res. 2016, 8, 1848–1856. [Google Scholar] [PubMed]
- Hao, G.J.; Hao, H.J.; Ding, Y.H.; Wen, H.; Li, X.F.; Wang, Q.R.; Zhang, B.B. Suppression of EIF4G2 by miR-379 potentiates the cisplatin chemosensitivity in nonsmall cell lung cancer cells. FEBS Lett. 2017, 591, 636–645. [Google Scholar] [CrossRef]
- Wang, R.; Chen, D.Q.; Huang, J.Y.; Zhang, K.; Feng, B.; Pan, B.Z.; Chen, J.; De, W.; Chen, L.B. Acquisition of radioresistance in docetaxel-resistant human lung adenocarcinoma cells is linked with dysregulation of miR-451/c-Myc-survivin/rad-51 signaling. Oncotarget 2014, 5, 6113–6129. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.C.; Li, S.J.; Zhang, F.; Zhang, Y.D.; Zuo, Z.Y.; Xi, Y.Y.; Wang, L.; Li, D.J. The Novel miR-9600 Suppresses Tumor Progression and Promotes Paclitaxel Sensitivity in Non-small-cell Lung Cancer Through Altering STAT3 Expression. Mol. Therapy. Nucleic Acids 2016, 5, e387. [Google Scholar] [CrossRef] [PubMed]
- Arechaga-Ocampo, E.; Lopez-Camarillo, C.; Villegas-Sepulveda, N.; Gonzalez-De la Rosa, C.H.; Perez-Anorve, I.X.; Roldan-Perez, R.; Flores-Perez, A.; Pena-Curiel, O.; Angeles-Zaragoza, O.; Rangel Corona, R.; et al. Tumor suppressor miR-29c regulates radioresistance in lung cancer cells. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef]
- Jiang, S.; Wang, R.; Yan, H.; Jin, L.; Dou, X.; Chen, D. MicroRNA-21 modulates radiation resistance through upregulation of hypoxia-inducible factor-1alpha-promoted glycolysis in non-small cell lung cancer cells. Mol. Med. Rep. 2016, 13, 4101–4107. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Tang, Y.; Guo, W.; Du, Y.; Wang, Y.; Li, P.; Zang, W.; Yin, X.; Wang, H.; Chu, H.; et al. Up-regulation of microRNA-138 induce radiosensitization in lung cancer cells. Tumour Biol. 2014, 35, 6557–6565. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Liu, Y.; Xiao, B.; Qian, X. miR-25 modulates NSCLC cell radio-sensitivity through directly inhibiting BTG2 expression. Biochem. Biophys. Res. Commun. 2015, 457, 235–241. [Google Scholar] [CrossRef] [PubMed]
- Cortez, M.A.; Valdecanas, D.; Zhang, X.; Zhan, Y.; Bhardwaj, V.; Calin, G.A.; Komaki, R.; Giri, D.K.; Quini, C.C.; Wolfe, T.; et al. Therapeutic delivery of miR-200c enhances radiosensitivity in lung cancer. Mol. Ther. J. Am. Soc. Gene Ther. 2014, 22, 1494–1503. [Google Scholar] [CrossRef]
- Grosso, S.; Doyen, J.; Parks, S.K.; Bertero, T.; Paye, A.; Cardinaud, B.; Gounon, P.; Lacas-Gervais, S.; Noel, A.; Pouyssegur, J.; et al. MiR-210 promotes a hypoxic phenotype and increases radioresistance in human lung cancer cell lines. Cell Death Dis. 2013, 4, e544. [Google Scholar] [CrossRef]
- Ma, W.; Ma, C.N.; Zhou, N.N.; Li, X.D.; Zhang, Y.J. Up- regulation of miR-328-3p sensitizes non-small cell lung cancer to radiotherapy. Sci. Rep. 2016, 6, 31651. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Han, W.; Ni, T.T.; Lu, L.; Huang, M.; Zhang, Y.; Cao, H.; Zhang, H.Q.; Luo, W.; Li, H. Knockdown of microRNA-1323 restores sensitivity to radiation by suppression of PRKDC activity in radiation-resistant lung cancer cells. Oncol. Rep. 2015, 33, 2821–2828. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.J.; Lin, Y.F.; Chen, Y.F.; Luo, E.C.; Sher, Y.P.; Tsai, M.H.; Chuang, E.Y.; Lai, L.C. MicroRNA-449a enhances radiosensitivity in CL1-0 lung adenocarcinoma cells. PLoS ONE 2013, 8, e62383. [Google Scholar] [CrossRef] [PubMed]
- Zhong, M.; Ma, X.; Sun, C.; Chen, L. MicroRNAs reduce tumor growth and contribute to enhance cytotoxicity induced by gefitinib in non-small cell lung cancer. Chem. Biol. Interact. 2010, 184, 431–438. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Ren, S.; Li, X.; Wang, Y.; Garfield, D.; Zhou, S.; Chen, X.; Su, C.; Chen, M.; Kuang, P.; et al. MiR-21 overexpression is associated with acquired resistance of EGFR-TKI in non-small cell lung cancer. Lung Cancer 2014, 83, 146–153. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, K.; Seike, M.; Okano, T.; Matsuda, K.; Miyanaga, A.; Mizutani, H.; Noro, R.; Minegishi, Y.; Kubota, K.; Gemma, A. MiR-134/487b/655 cluster regulates TGF-beta-induced epithelial-mesenchymal transition and drug resistance to gefitinib by targeting MAGI2 in lung adenocarcinoma cells. Mol. Cancer Ther. 2014, 13, 444–453. [Google Scholar] [CrossRef]
- Han, Z.; Zhou, X.; Li, S.; Qin, Y.; Chen, Y.; Liu, H. Inhibition of miR-23a increases the sensitivity of lung cancer stem cells to erlotinib through PTEN/PI3K/Akt pathway. Oncol. Rep. 2017, 38, 3064–3070. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.; Zhang, F.; Guo, Y.; Huang, J.; Xie, Y.; Yue, S.; Chen, M.; Jiang, H.; Li, M. miR-200c enhances sensitivity of drug-resistant non-small cell lung cancer to gefitinib by suppression of PI3K/Akt signaling pathway and inhibites cell migration via targeting ZEB1. Biomed. Pharmacother. 2017, 85, 113–119. [Google Scholar] [CrossRef]
- Chan, L.W.; Wang, F.; Meng, F.; Wang, L.; Wong, S.C.; Au, J.S.; Yang, S.; Cho, W.C. MiR-30 Family Potentially Targeting PI3K-SIAH2 Predicted Interaction Network Represents a Novel Putative Theranostic Panel in Non-small Cell Lung Cancer. Front. Genet. 2017, 8, 8. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Zhao, F.; Zhang, J.; Zhu, H.; Ma, H.; Li, X.; Peng, L.; Sun, J.; Chen, Z. miR-223 reverses the resistance of EGFR-TKIs through IGF1R/PI3K/Akt signaling pathway. Int. J. Oncol. 2016, 48, 1855–1867. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.S.; Wang, Y.H.; Xia, H.P.; Zhou, S.W.; Schmid-Bindert, G.; Zhou, C.C. MicroRNA-214 regulates the acquired resistance to gefitinib via the PTEN/AKT pathway in EGFR-mutant cell lines. Asian Pac. J. Cancer Prev. 2012, 13, 255–260. [Google Scholar] [CrossRef] [PubMed]
- Yue, J.; Lv, D.; Wang, C.; Li, L.; Zhao, Q.; Chen, H.; Xu, L. Epigenetic silencing of miR-483-3p promotes acquired gefitinib resistance and EMT in EGFR-mutant NSCLC by targeting integrin beta3. Oncogene 2018, 37, 4300–4312. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.X.; Yan, L.; Li, C.; Zhao, L.M.; Liu, W. miR-200c regulates crizotinib-resistant ALK-positive lung cancer cells by reversing epithelial-mesenchymal transition via targeting ZEB1. Mol. Med. Rep. 2016, 14, 4135–4143. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Liao, Z.; O’Reilly, M.S.; Levy, L.B.; Welsh, J.W.; Wang, L.E.; Lin, S.H.; Komaki, R.; Liu, Z.; Wei, Q.; et al. Serum inflammatory miRNAs predict radiation esophagitis in patients receiving definitive radiochemotherapy for non-small cell lung cancer. Radiother. Oncol. 2014, 113, 379–384. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Xu, Y.; Liao, X.; Liao, R.; Zhang, L.; Niu, K.; Li, T.; Li, D.; Chen, Z.; Duan, Y.; et al. Plasma miRNAs in predicting radiosensitivity in non-small cell lung cancer. Tumour Biol. 2016, 37, 11927–11936. [Google Scholar] [CrossRef]
- Langevin, S.M.; Kratzke, R.A.; Kelsey, K.T. Epigenetics of lung cancer. Transl. Res. 2015, 165, 74–90. [Google Scholar] [CrossRef] [PubMed]
- Garofalo, M.; Romano, G.; Di Leva, G.; Nuovo, G.; Jeon, Y.J.; Ngankeu, A.; Sun, J.; Lovat, F.; Alder, H.; Condorelli, G.; et al. EGFR and MET receptor tyrosine kinase-altered microRNA expression induces tumorigenesis and gefitinib resistance in lung cancers. Nat. Med. 2011, 18, 74–82. [Google Scholar] [CrossRef]
- Shen, H.; Zhu, F.; Liu, J.; Xu, T.; Pei, D.; Wang, R.; Qian, Y.; Li, Q.; Wang, L.; Shi, Z.; et al. Alteration in Mir-21/PTEN expression modulates gefitinib resistance in non-small cell lung cancer. PLoS ONE 2014, 9, e103305. [Google Scholar] [CrossRef]
- Bisagni, A.; Pagano, M.; Maramotti, S.; Zanelli, F.; Bonacini, M.; Tagliavini, E.; Braglia, L.; Paci, M.; Mozzarelli, A.; Croci, S. Higher expression of miR-133b is associated with better efficacy of erlotinib as the second or third line in non-small cell lung cancer patients. PLoS ONE 2018, 13, e0196350. [Google Scholar] [CrossRef]
- Zhao, J.; Kelnar, K.; Bader, A.G. In-depth analysis shows synergy between erlotinib and miR-34a. PLoS ONE 2014, 9, e89105. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Guerrero, A.; Kelnar, K.; Peltier, H.J.; Bader, A.G. Synergy between next generation EGFR tyrosine kinase inhibitors and miR-34a in the inhibition of non-small cell lung cancer. Lung Cancer 2017, 108, 96–102. [Google Scholar] [CrossRef]
- Kim, J.Y.; Lee, W.J.; Park, H.Y.; Kim, A.; Shin, D.H.; Lee, C.H. Differential MicroRNA Expression between EGFR T790M and L858R Mutated Lung Cancer. J. Pathol. Transl. Med. 2018, 52, 275–282. [Google Scholar] [CrossRef]
- Shaw, A.T.; Yeap, B.Y.; Mino-Kenudson, M.; Digumarthy, S.R.; Costa, D.B.; Heist, R.S.; Solomon, B.; Stubbs, H.; Admane, S.; McDermott, U.; et al. Clinical features and outcome of patients with non-small-cell lung cancer who harbor EML4-ALK. J. Clin. Oncol. 2009, 27, 4247–4253. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Yang, J.M.; Jin, Y.; Jheon, S.; Kim, K.; Lee, C.T.; Chung, J.H.; Paik, J.H. MicroRNA expression profiles and clinicopathological implications in lung adenocarcinoma according to EGFR, KRAS, and ALK status. Oncotarget 2017, 8, 8484–8498. [Google Scholar] [CrossRef] [PubMed]
- Li, L.L.; Qu, L.L.; Fu, H.J.; Zheng, X.F.; Tang, C.H.; Li, X.Y.; Chen, J.; Wang, W.X.; Yang, S.X.; Wang, L.; et al. Circulating microRNAs as novel biomarkers of ALK-positive nonsmall cell lung cancer and predictors of response to crizotinib therapy. Oncotarget 2017, 8, 45399–45414. [Google Scholar] [CrossRef] [PubMed]
- Yun, M.R.; Lim, S.M.; Kim, S.K.; Choi, H.M.; Pyo, K.H.; Kim, S.K.; Lee, J.M.; Lee, Y.W.; Choi, J.W.; Kim, H.R.; et al. Enhancer Remodeling and MicroRNA Alterations Are Associated with Acquired Resistance to ALK Inhibitors. Cancer Res. 2018, 78, 3350–3362. [Google Scholar] [CrossRef] [PubMed]
- Halvorsen, A.R.; Sandhu, V.; Sprauten, M.; Flote, V.G.; Kure, E.H.; Brustugun, O.T.; Helland, A. Circulating microRNAs associated with prolonged overall survival in lung cancer patients treated with nivolumab. Acta Oncol. 2018, 57, 1225–1231. [Google Scholar] [CrossRef] [PubMed]
- Beg, M.S.; Brenner, A.J.; Sachdev, J.; Borad, M.; Kang, Y.K.; Stoudemire, J.; Smith, S.; Bader, A.G.; Kim, S.; Hong, D.S. Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Investig. New Drugs 2017, 35, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Van Zandwijk, N.; Pavlakis, N.; Kao, S.C.; Linton, A.; Boyer, M.J.; Clarke, S.; Huynh, Y.; Chrzanowska, A.; Fulham, M.J.; Bailey, D.L.; et al. Safety and activity of microRNA-loaded minicells in patients with recurrent malignant pleural mesothelioma: A first-in-man, phase 1, open-label, dose-escalation study. Lancet. Oncol. 2017, 18, 1386–1396. [Google Scholar] [CrossRef]
- Larrea, E.; Sole, C.; Manterola, L.; Goicoechea, I.; Armesto, M.; Arestin, M.; Caffarel, M.M.; Araujo, A.M.; Araiz, M.; Fernandez-Mercado, M.; et al. New Concepts in Cancer Biomarkers: Circulating miRNAs in Liquid Biopsies. Int. J. Mol. Sci. 2016, 17, 627. [Google Scholar] [CrossRef] [PubMed]
- Yang, N. An overview of viral and nonviral delivery systems for microRNA. Int. J. Pharm. Investig. 2015, 5, 179–181. [Google Scholar] [CrossRef]
- Xue, J.; Yang, J.; Luo, M.; Cho, W.C.; Liu, X. MicroRNA-targeted therapeutics for lung cancer treatment. Expert Opin. Drug Discov. 2017, 12, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Janssen, H.L.A.; Reesink, H.W.; Lawitz, E.J.; Zeuzem, S.; Rodriguez-Torres, M.; Patel, K.; van der Meer, A.J.; Patick, A.K.; Chen, A.; Zhou, Y.; et al. Treatment of HCV Infection by Targeting MicroRNA. New Engl. J. Med. 2013, 368, 1685–1694. [Google Scholar] [CrossRef] [PubMed]
- Titze-de-Almeida, R.; David, C.; Titze-de-Almeida, S.S. The Race of 10 Synthetic RNAi-Based Drugs to the Pharmaceutical Market. Pharm. Res. 2017, 34, 1339–1363. [Google Scholar] [CrossRef]
- Stenvang, J.; Petri, A.; Lindow, M.; Obad, S.; Kauppinen, S. Inhibition of microRNA function by antimiR oligonucleotides. Silence 2012, 3, 1. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, K.-L.; Tsai, Y.-M.; Lien, C.-T.; Kuo, P.-L.; Hung, J.-Y. The Roles of MicroRNA in Lung Cancer. Int. J. Mol. Sci. 2019, 20, 1611. https://doi.org/10.3390/ijms20071611
Wu K-L, Tsai Y-M, Lien C-T, Kuo P-L, Hung J-Y. The Roles of MicroRNA in Lung Cancer. International Journal of Molecular Sciences. 2019; 20(7):1611. https://doi.org/10.3390/ijms20071611
Chicago/Turabian StyleWu, Kuan-Li, Ying-Ming Tsai, Chi-Tun Lien, Po-Lin Kuo, and Jen-Yu Hung. 2019. "The Roles of MicroRNA in Lung Cancer" International Journal of Molecular Sciences 20, no. 7: 1611. https://doi.org/10.3390/ijms20071611
APA StyleWu, K.-L., Tsai, Y.-M., Lien, C.-T., Kuo, P.-L., & Hung, J.-Y. (2019). The Roles of MicroRNA in Lung Cancer. International Journal of Molecular Sciences, 20(7), 1611. https://doi.org/10.3390/ijms20071611





