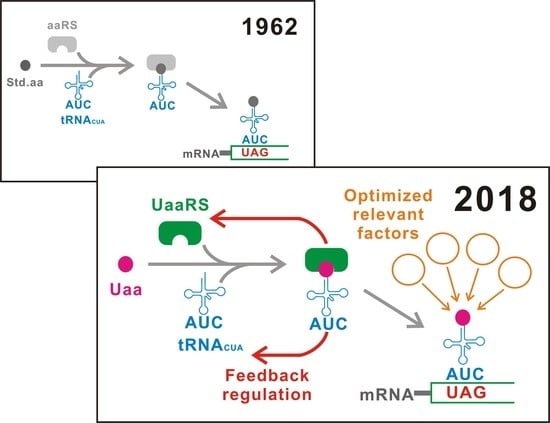
Translational Control using an Expanded Genetic Code
Abstract
1. Introduction
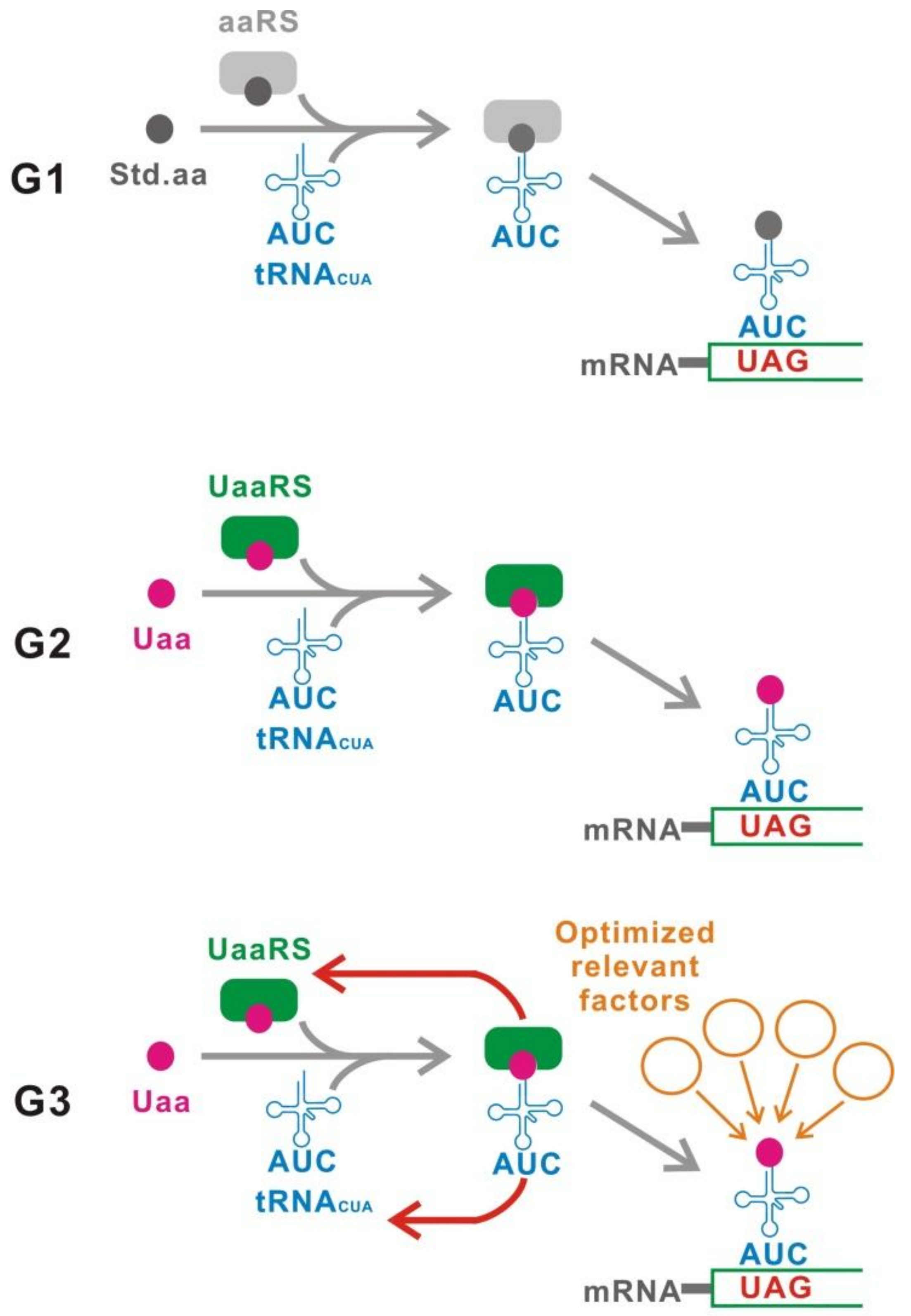
2. Classifications
3. G1
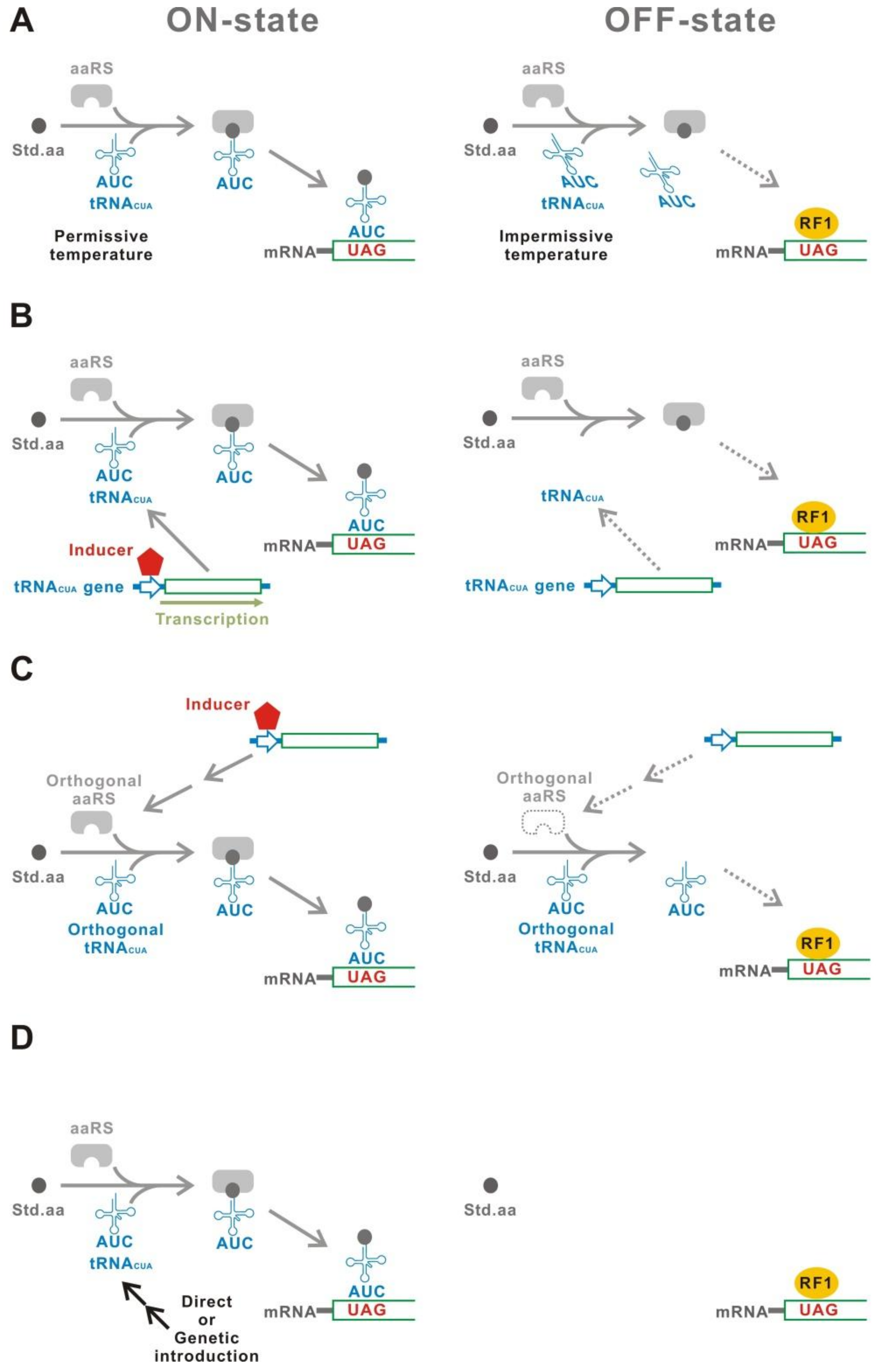
3.1. Thermolabile tRNACUA
3.2. Inducible Transcription of tRNACUA
3.3. Inducible Aminoacyl-tRNA Synthetase
3.4. Introduction of the tRNACUA Gene
4. G2
5. G3
5.1. Inducible UaaRS/tRNACUA
5.2. Optimization of the UaaRS/tRNACUA Expression Level
5.3. Multiplexed TAG Insertion in a Target Gene
5.4. Location of the UAG Insertion
5.5. Uaa-Residue Conversion to Standard Amino Acid Residue
5.6. Suppression of Peptide Release Factor
5.7. Uaa-Permissive Elongation Factor
5.8. Host Selection
5.9. Improved UaaRS/tRNACUA
6. Uaa Transcriptional Switch
7. Applications
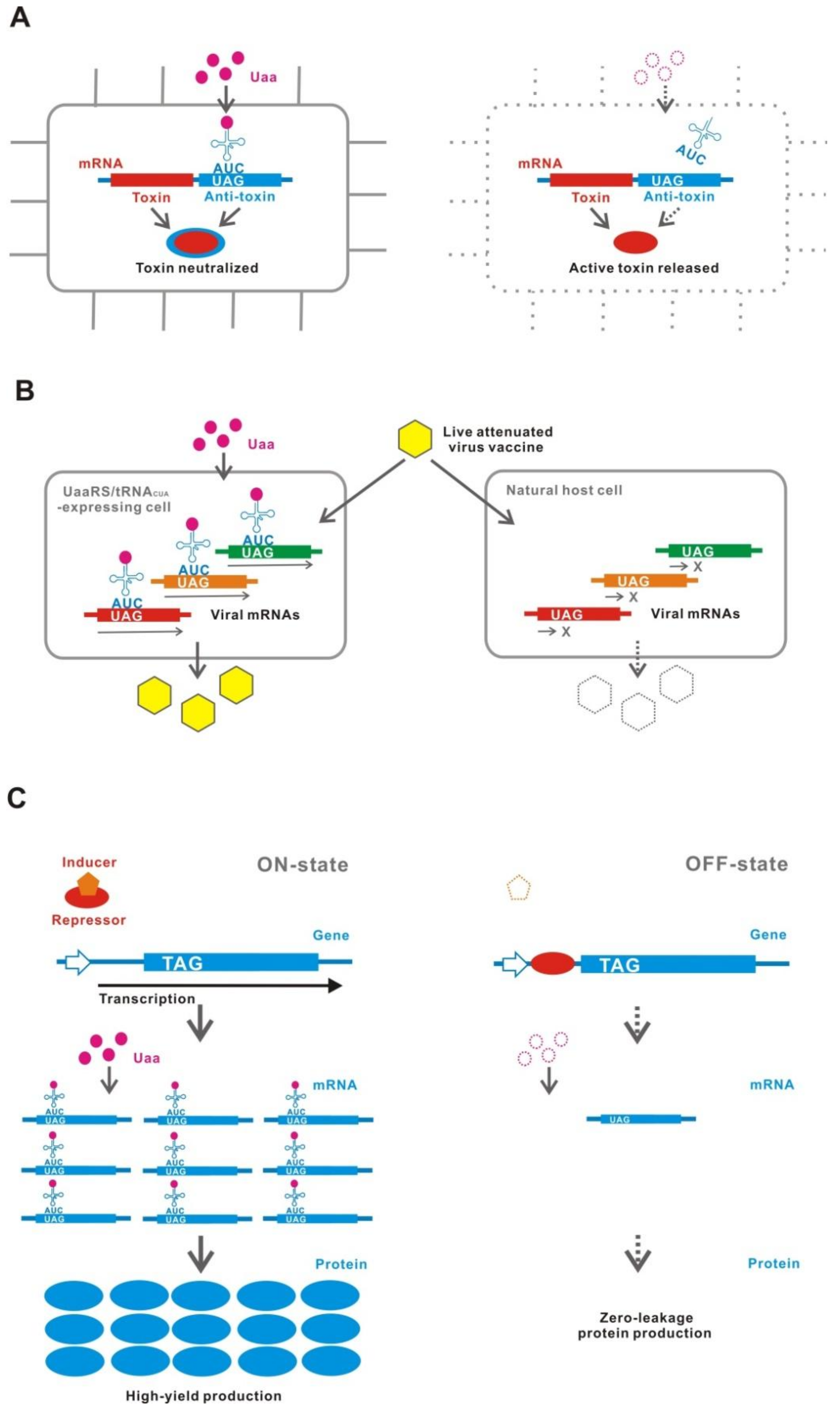
7.1. Biological Containment
7.2. Live Attenuated-Virus Vaccine
7.3. High-yield and Zero-Leakage Expression System (HYZEL)
8. Conclusions
Funding
Conflicts of Interest
References
- Dabrowski, M.; Bukowy-Bieryllo, Z.; Zietkiewicz, E. Advances in therapeutic use of a drug-stimulated translational readthrough of premature termination codons. Mol. Med. 2018, 24, 25. [Google Scholar] [CrossRef] [PubMed]
- Benzer, S.; Champe, S.P. A change from nonsense to sense in the genetic code. Proc. Natl. Acad. Sci. USA 1962, 48, 1114–1121. [Google Scholar] [CrossRef]
- Garen, A.; Siddiqi, O. Suppression of mutations in the alkaline phosphatase structural cistron of E. coli. Proc. Natl. Acad. Sci. USA 1962, 48, 1121–1127. [Google Scholar] [CrossRef] [PubMed]
- Sarabhai, A.S.; Stretton, A.O.W.; Brenner, S. Co-linearity of the gene with the polypeptide chain. Nature 1964, 201, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Gorini, L. Informational suppression. Annu. Rev. Genet. 1970, 4, 107–134. [Google Scholar] [CrossRef] [PubMed]
- Eggertsson, G.; Söll, D. Transfer ribonucleic acid-mediated suppression of termination codons in Escherichia coli. Microbiol. Rev. 1988, 52, 354–374. [Google Scholar] [PubMed]
- Garen, A.; Garen, S.; Wilhelm, R.C. Suppressor genes for nonsense mutations. I. The Su-1, Su-2 and Su-3 genes of Escherichia coli. J. Mol. Biol. 1965, 14, 167–178. [Google Scholar] [CrossRef]
- Weigert, M.G.; Lanka, E.; Garen, A. Amino acid substitutions resulting from suppression of nonsense mutations. II. Glutamine insertion by the Su-2 gene; tyrosine insertion by Su-3 gene. J. Mol. Biol. 1965, 14, 522–527. [Google Scholar] [CrossRef]
- Kimura, M.; Miki, T.; Hiraga, S.; Nagata, T.; Yura, T. Conditionally lethal amber mutations in the dnaA region of the Escherichia coli chromosome that affect chromosome replication. J. Bacteriol. 1979, 140, 825–834. [Google Scholar]
- Herring, C.D.; Blattner, F. Conditional lethal amber mutations in essential Escherichia coli genes. J. Bacteriol. 2004, 186, 2673–2681. [Google Scholar] [CrossRef]
- Tian, H.; Deng, D.; Huang, J.; Yao, D.; Xiaowei, X.; Gao, X. Screening system for orthogonal suppressor tRNAs based on the species-specific toxicity of suppressor tRNAs. Biochimie 2013, 95, 881–888. [Google Scholar] [CrossRef] [PubMed]
- Gallucci, E.; Pacchetti, G.; Zangrossi, S. Genetic studies on temperature sensitive nonsense suppression. Mol. Gen. Genet. 1970, 106, 362–370. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.D.; Anderson, K.; Cashmore, A.; hooper, M.L.; Russell, R.L. Studies on the structure and synthesis of Escherichia coli tyrosine transfer RNA. Cold Spring Harb. Symp. Quant. Biol. 1970, 35, 21–27. [Google Scholar] [CrossRef]
- Rasse-Messenguy, F.; Fink, G.R. Temperature-sensitive nonsense suppressors in yeast. Genetics 1973, 75, 459–464. [Google Scholar] [PubMed]
- Steege, D.A.; Horabin, J.I. Temperature-inducible amber suppressor: Construction of plasmids containing the Escherichia coli serU-(supD-) gene under control of the bacteriophage lambda pL promoter. J. Bacteriol. 1983, 155, 1417–1425. [Google Scholar] [PubMed]
- Sedivy, J.M.; Capone, J.P.; RajBhandary, U.L.; Sharp, P.A. An inducible mammalian amber suppressor: Propagation of a poliovirus mutant. Cell 1987, 50, 379–389. [Google Scholar] [CrossRef]
- Bouet, J.-Y.; Campo, N.J.; Krisch, H.M.; Louarn, J.-M. The effects on Escherichia coli of expression of the cloned bacteriophage T4 nucleoid disruption (ndd) gene. Mol. Microbiol. 1996, 20, 519–528. [Google Scholar] [CrossRef]
- Dingermann, T.; Werner, H.; Schutz, L.; Zundorf, I.; Nerke, K.; Knecht, D.; Marschalek, R. Establishment of a system for conditional gene expression using an iducible tRNA suppressor gene. Mol. Cell. Biol. 1992, 12, 4038–4045. [Google Scholar] [CrossRef]
- Grundy, F.J.; Henkin, T.M. Inducible amber suppressor for Bacillus subtilis. J. Bacteriol. 1994, 176, 2108–2110. [Google Scholar] [CrossRef]
- Herring, C.D.; Glasner, J.D.; Blattner, F.R. Gene replacement without selection: Regulated suppression of amber mutations in Escherichia coli. Gene 2003, 331, 153–163. [Google Scholar] [CrossRef]
- Park, H.-J.; RajBhandary, U.L. Tetracycline-regulated suppression of amber codons in mammalian cells. Mol. Cell Biol. 1998, 18, 4418–4425. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Edwards, H.; Schimmel, P. A bacterial amber suppressor in Saccharomyces cerevisiae is selectively recognized by a bacterial aminoacyl-tRNA synthetase. Mol. Cell. Biol. 1990, 10, 1633–1641. [Google Scholar] [CrossRef] [PubMed]
- Drabkin, H.J.; Park, H.-J.; RajBhandary, U.L. Amber suppression in mammalian cells dependent upon expression of an Escherichia coli aminoacyl-tRNA synthetase gene. Mol. Cell. Biol. 1996, 16, 907–913. [Google Scholar] [CrossRef] [PubMed]
- Carneiro, V.T.C.; Pelletier, G.; Small, I. Transfer RNA-mediated suppression of stop codons in protoplasts and transgenic plants. Plant Mol. Biol 1993, 22, 681–690. [Google Scholar] [CrossRef] [PubMed]
- Kunes, S.; Steller, H. Ablation of Drosophila photoreceptor cells by conditional expression of a toxin gene. Genes Dev. 1991, 5, 970–983. [Google Scholar] [CrossRef]
- Betzner, A.S.; Oakes, M.P.; Huttner, E. Transfer RNA-mediated suppression of amber stop codons in transgenic Arabidopsis thaliana. Plant J. 1997, 11, 587–595. [Google Scholar] [CrossRef] [PubMed]
- Levine, M.; Tarver, H. Studies on ethionine. III. Incorporation of ethionine into rat proteins. J. Biol. Chem. 1951, 192, 835–850. [Google Scholar] [PubMed]
- Cowie, D.B.; Cohen, G.N. Biosynthesis by Escherichia coli of active altered proteins containing selenium instead of sulfur. Biochim. Biophys. Acta 1957, 26, 252–261. [Google Scholar] [CrossRef]
- Munier, R.; Cohen, G. Incorporation of structural analogues of amino acid into the bacterial proteins during their synthesis in vivo. Biochim. Biophys. Acta 1959, 31, 378–391. [Google Scholar] [CrossRef]
- Richmond, M.H. Random replacement of phenylalanine by p-fluorophenylalanine in alkaline phosphatase(s) formed during biosynthesis by E. coli. J. Mol. Biol. 1963, 6, 284–294. [Google Scholar] [CrossRef]
- Pratt, E.A.; Ho, C. Incorporation of fluorotryptophans into proteins of Escherichia coli. Biochemistry 1975, 14, 3035–3040. [Google Scholar] [CrossRef] [PubMed]
- Hoagland, M.B.; Keller, E.B.; Zamecnik, P.C. Enzymatic carboxyl activation of amino acids. J. Biol. Chem. 1956, 218, 345–358. [Google Scholar] [PubMed]
- Hoagland, M.B.; Zamecnik, P.C.; Stephenson, M.L. Intermediate reactions in protein biosynthesis. Biochim. Biophys. Acta 1957, 24, 215–216. [Google Scholar] [CrossRef]
- Hoagland, M.B.; Stephenson, M.L.; Scott, J.F.; Hecht, L.I.; Zamecnik, P.C. A soluble ribonucleic acid intermediate in protein synthesis. J. Biol. Chem. 1958, 231, 241–257. [Google Scholar] [PubMed]
- Browne, D.T.; Kenyon, G.L.; Hegeman, G.D. Incorporation of monofluorotryptophans into protein during the growth of Escherichia coli. Biochem. Biophys. Res. Commun. 1970, 39, 13–19. [Google Scholar] [CrossRef]
- Sykes, B.D.; Weingarten, H.I.; Schlesinger, M.J. Fluorotyrosine alkaline phosphatase from Escherichia coli: Preparation, properties, and fluorine-19 nuclear magnetic resonance spectrum. Proc. Natl. Acad. Sci. USA 1974, 71, 469–473. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.A.; Nakashima, Y.; Coleman, J.E. Chemical modifications of functional residues of fd gene 5 DNA-binding protein. Biochemistry 1975, 14, 907–917. [Google Scholar] [CrossRef]
- Lu, P.; Jarema, M.; Mosser, K.; Danier, W.E., Jr. lac repressor: 3-fluorotyrosine substitution for nuclear magnetic resonance studies. Proc. Natl. Acad. Sci. USA 1976, 73, 3471–3475. [Google Scholar] [CrossRef]
- Browne, D.T.; Otvos, J.D. 4-Fluorotryptphan alkaline phosphatase from E. coli: Preparation, properties, and 19F NMR spectrum. Biochem. Biophys. Res. Commun. 1976, 68, 907–913. [Google Scholar] [CrossRef]
- Koide, H.; Yokoyama, S.; Kawai, G.; Ha, J.-M.; Oka, T.; Kawai, S.; Miyake, T.; Fuwa, T.; Miyazawa, T. Biosynthesis of a protein containing a nonprotein amino acid by Escherichia coli: L-2-aminohexanoic acid at position 21 in human epidermal growth factor. Proc. Natl. Acad. Sci. USA 1988, 85, 6237–6241. [Google Scholar] [CrossRef]
- Noren, C.J.; Anthony-Cahill, S.J.; Griffith, M.C.; Schultz, P.G. A general method for site-specific incorporation of unnatural amino acids into proteins. Science 1989, 244, 182–188. [Google Scholar] [CrossRef]
- Liu, D.R.; Magliery, T.J.; Pastrnak, M.; Schultz, P.G. Engineering a tRNA and aminoacyl-tRNA synthetase for the site-specific incorporation of unnatural amino acids into proteins in vivo. Proc. Natl. Acad. Sci. USA 1997, 94, 10092–10097. [Google Scholar] [CrossRef] [PubMed]
- Ohno, S.; Yokogawa, T.; Fujii, I.; Asahara, H.; Inokuchi, H.; Nishikawa, K. Co-expression of yeast amber suppressor tRNATyr and tyrosyl-tRNA synthetase in Escherichia coli: Possibility to expand the genetic code. J. Biochem. 1998, 124, 1065–1068. [Google Scholar] [CrossRef]
- Furter, R. Expansion of the genetic code: Site-directed p-fluoro-phenylalanine incorporation in Escherichia coli. Prot. Sci. 1998, 7, 419–426. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Brock, A.; Herberich, B.; Schultz, P.G. Expanding the genetic code of Escherichia coli. Science 2001, 292, 498–500. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, K.; Hayashi, A.; Sakamoto, A.; Kiga, D.; Nakayama, H.; Soma, A.; Kobayashi, T.; Kitabatake, M.; Takio, K.; Saito, K.; et al. Site-specific incorporation of an unnatural amino acid into proteins in mammalian cells. Nucl. Acids Res. 2002, 30, 4692–4699. [Google Scholar] [CrossRef]
- Chin, J.W.; Cropp, A.; Anderson, J.C.; Mukherji, M.; Zhang, Z.; Schultz, P.G. An expanded eukaryotic genetic code. Science 2003, 301, 964–967. [Google Scholar] [CrossRef]
- Minaba, M.; Kato, Y. High-yield, zero-leakage expression system with a translational switch using site-specific unnatural amino acid incorporation. Appl. Environ. Microbiol. 2014, 80, 1718–1725. [Google Scholar] [CrossRef]
- Kato, Y. Tunable translational control using site-specific unnatural amino acid incorporation in Escherichia coli. PeerJ 2015, 3, e904. [Google Scholar] [CrossRef]
- Kato, Y. Tight translational control using site-specific unnatural amino acid incorporation with positive feedback gene circuits. ACS Synth. Biol. 2018, 7, 1956–1963. [Google Scholar] [CrossRef]
- Liu, C.C.; Schultz, P.G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 2010, 79, 413–444. [Google Scholar] [CrossRef] [PubMed]
- Xiao, H.; Schultz, P.G. At the interface of chemical and biological synthesis: An expanded genetic code. Cold Spring Harbor Perspect. Biol. 2016, 8, a023945. [Google Scholar] [CrossRef] [PubMed]
- Chin, J.W. Expanding and reprogramming the genetic code. Nature 2017, 550, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Khazaie, K.; Buchanan, J.H.; Rosenberger, R.F. The accuracy of Qβ RNA translation: 1. Errors during the synthesis of Qβ proteins by intact Escherichia coli cells. Eur. J. Biochem. 1984, 144, 485–489. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, M.; Rydén-Aulin, M. Glutamine is incorporated at the nonsense codons UAG and UAA in a suppressor-free Escherichia coli strain. Biochim. Biophys. Acta 2003, 1627, 1–6. [Google Scholar] [CrossRef]
- O’Donghue, P.; Prat, L.; Heinemann, I.U.; Ling, J.; Odoi, K.; Liu, W.R.; Söll, D. Near-cognate suppression of amber, opal and quadruplet codons competes with aminoacyl-tRNAPyl for genetic code expansion. FEBS Lett. 2012, 586, 3931–3937. [Google Scholar] [CrossRef] [PubMed]
- Guzman, L.; Belin, D.; Carson, M.J.; Beckwith, J. Thight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 1995, 177, 4121–4130. [Google Scholar] [CrossRef]
- Zheng, Y.; Lewis, T.L.; Igo, P.; Polleux, F.; Chatterjee, A. Virus-enabled optimization and delivery of the genetic machinery for efficient unnatural amio acid mutagenesis in mammalian cells and tissues. ACS Synth. Biol. 2017, 6, 13–18. [Google Scholar] [CrossRef]
- Swanson, R.; Hoben, P.; Sumner-Smith, M.; Uemura, H.; Watson, L.; Söll, D. Accuracy of in vivo aminoacylation requires proper balance of tRNA and aminoacyl-tRNA synthetase. Science 1988, 242, 1548–1551. [Google Scholar] [CrossRef]
- Varshney, U.; RajBhandary, U.L. Role of methionine and formylation of initiator tRNA in initiation of protein synthesis in E. coli. J. Bacteriol. 1992, 174, 7819–7826. [Google Scholar] [CrossRef]
- Kiick, K.L.; van Hest, J.C.M.; Tirrell, D.A. Expanding the scope of protein biosynthesis by altering the methionyl-tRNA synthetase activity of a bacterial expression host. Angew. Chem. Int. Ed. 2000, 39, 2148–2152. [Google Scholar] [CrossRef]
- Yarus, M.; Cline, S.W.; Wier, P.; Breeden, L.; Thompson, R.C. Actions of the anticodon arm in translation on the phenotypes of RNA mutants. J. Mol. Biol. 1986, 192, 235–255. [Google Scholar] [CrossRef]
- Kudla, G.; Murray, A.W.; Tollervey, D.; Plotkin, J.B. Coding-sequence determinants of gene expression in Escherichia coli. Science 2009, 324, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Bentele, K.; Saffert, P.; Rauscher, R.; Ignatova, Z.; Bluthgen, N. Efficient translation initiation dictates codon usage at gene start. Mol. Syst. Biol. 2013, 9, 675. [Google Scholar] [CrossRef] [PubMed]
- Goodman, D.B.; Church, G.M.; Kosuri, S. Causes and effects of N-terminal codon bias in bacterial genes. Science 2013, 342, 475–479. [Google Scholar] [CrossRef] [PubMed]
- Hirel, P.-H.; Schmitter, J.-M.; Dessen, P.; Fayat, G.; Blanquet, S. Extent of N-terminal methionine excision from Escherichia coli proteins is governed by the side-chain length of the penultimate amino acid. Proc. Natl. Acad. Sci. USA 1989, 86, 8247–8251. [Google Scholar] [CrossRef]
- Tobias, J.W.; Shrader, T.E.; Rocap, G.; Varshavsky, A. The N-end rule in bacteria. Science 1991, 254, 1374–1377. [Google Scholar] [CrossRef]
- Schuenemann, V.J.; Kralik, S.M.; Albrecht, R.; Spall, S.K.; Truscott, K.N.; Dougan, D.A.; Zeth, K. Structural basis of N-end rule substrate recognition in Escherichia coli by the ClpAP adaptor protein ClpS. EMBO Rep. 2009, 10, 508–514. [Google Scholar] [CrossRef]
- Neumann, H.; Peak-Chew, S.Y.; Chin, J.W. Genetically encoding Nε-acetyllysine in recombinant proteins. Nat. Chem. Biol. 2008, 4, 232–234. [Google Scholar] [CrossRef]
- Umehara, T.; Kim, J.; Lee, S.; Guo, L.T.; Söll, D.; Park, H.S. N-acetyl lysyl-tRNA synthetases evolved by a CcdB-based selection possess N-acetyl lysine specificity in vitro and in vivo. FEBS Lett. 2012, 586, 729–733. [Google Scholar] [CrossRef]
- Xuan, W.; Schultz, P.G. A strategy for creating organisms dependent on noncanonical amino acids. Angew. Chem. Int. Ed. 2017, 56, 9170–9173. [Google Scholar] [CrossRef] [PubMed]
- Volkwein, W.; Maier, C.; Krafczyk, R.; Jung, K.; Lassak, J. A versatile toolbox for the control of protein levels using Nε-acetyl-L-lysine dependent amber suppression. ACS Synth. Biol. 2017, 6, 1892–1902. [Google Scholar] [CrossRef] [PubMed]
- Walsh, C.T.; Garneau-Tsodikova, S.; Gatto, G.J., Jr. Protein posttranslational modifications: The chemistry of proteome diversifications. Angew. Chem. Int. Ed. 2005, 44, 7342–7372. [Google Scholar] [CrossRef] [PubMed]
- De Ruijter, A.J.M.; van Gennip, A.H.; Caron, H.N.; Kemp, S.; van Kuilenburg, A.B.P. Histone deacetylases (HDACs): Characterization of the classical HDAC family. Biochem. J. 2003, 370, 737–749. [Google Scholar] [CrossRef] [PubMed]
- Blander, G.; Guarente, L. The Sir2 family of protein deacetylases. Annu. Rev. Biochem. 2004, 73, 417–435. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.B.F.; Wang, C.; Xu, J.; Schultz, M.D.; Schmitz, R.J.; Ecker, J.R.; Wang, L. Release factor one is nonessential in Escherichia coli. ACS Chem. Biol. 2012, 7, 1337–1344. [Google Scholar] [CrossRef] [PubMed]
- Lajoie, M.J.; Rovner, A.J.; Goodman, D.B.; Aerni, H.-R.; Haimovich, A.D.; Kuznetsov, G.; Mercer, J.A.; Wang, H.H.; Carr, P.A.; Mosberg, J.A.; et al. Genomically recoded organisms expand biological functions. Science 2013, 342, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Pott, M.; Schmidt, M.J.; Summerer, D. Evolved sequence contexts for highly efficient amber suppression with noncanonical amino acids. ACS Chem. Biol. 2014, 9, 2815–2822. [Google Scholar] [CrossRef] [PubMed]
- Hayes, C.S.; Bose, B.; Sauer, R.T. Proline residues at the C terminus of nascent chains induce SsrA tagging during translation termination. J. Biol. Chem. 2002, 277, 33825–33832. [Google Scholar] [CrossRef]
- Tanner, D.R.; Cariello, D.A.; Woolstenhulme, C.J.; Broadbent, M.A.; Buskirk, A.R. Genetic identification of nascent peptides that induce ribosome stalling. J. Biol. Chem. 2009, 284, 34809–34818. [Google Scholar] [CrossRef]
- Ude, S.; Lassak, J.; Starosta, A.; Kraxenberger, T.; Wilson, D.N.; Jung, K. Translation elongation factor EF-P alleviates ribosome stalling at polyproline stretches. Science 2013, 339, 82–85. [Google Scholar] [CrossRef] [PubMed]
- Phillips-Jones, M.K.; Watson, F.J.; Martin, R. The 3′ codon context effect on UAG suppressor tRNA is different in Escherichia coli and human cells. J. Mol. Biol. 1993, 233, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Schmied, W.H.; Elsässer, S.J.; Uttamapinant, C.; Chin, J.W. Efcient multisite unnatural amino acid incorporation in mammalian cells via optimized pyrrolysyl tRNA synthetase/tRNA expression and engineered eRF1. J. Am. Chem. Soc. 2014, 136, 15577–15583. [Google Scholar] [CrossRef]
- Wu, I.L.; Patterson, M.A.; Desai, H.E.C.; Mehl, R.A.; Giorgi, G.; Conticello, V.P. Multiple site-selective insertions of noncanonical amino acids into sequence-repetitive polypeptides. ChemBioChem 2013, 14, 968–978. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mukai, T.; Hayashi, A.; Iraha, F.; Sato, A.; Ohtake, K.; Yokoyama, S.; Sakamoto, K. Codon-reassignment in the Escherichia coli genetic code. Nucl. Acids Res. 2010, 38, 8188–8195. [Google Scholar] [CrossRef]
- Johnson, D.B.F.; Xu, J.; Shen, Z.; Takimoto, J.K.; Schultz, M.D.; Schmitz, R.J.; Ecker, J.R.; Briggs, S.P.; Wang, L. RF1 knockout allows ribosomal incorporation of unnatural amino acids at multiple sites. Nat. Chem. Biol. 2012, 7, 779–786. [Google Scholar] [CrossRef] [PubMed]
- Ohtake, K.; Sato, A.; Mukai, T.; Hino, N.; Yokoyama, S.; Sakamoto, K. Efficient decoding of the UAG triplet as a full-fledged sense codon enhances the growth of a prfA-deficient strain of Escherichia coli. J. Bacteriol. 2012, 194, 2606–2613. [Google Scholar] [CrossRef]
- Amiram, M.; Haimovich, A.D.; Fan, C.; Wang, Y.-S.; Aerni, H.-R.; Ntai, I.; Moonan, D.W.; Ma, N.J.; Rovner, A.J.; Hong, S.H.; et al. Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nat. Biotechnol. 2015, 33, 1272–1279. [Google Scholar] [CrossRef]
- Wang, K.; Neumann, H.; Peak-Chew, S.Y.; Chin, J.W. Evolved orthogonal ribosomes enhance the efficiency of synthetic genetic code expansion. Nat. Biotechnol. 2007, 25, 770–777. [Google Scholar] [CrossRef]
- LaRiviere, F.J.; Wolfson, A.D.; Uhlenbeck, O.C. Uniform binding of aminoacyl-tRNAs to elongation factor Tu by thermodynamic compensation. Science 2001, 294, 165–168. [Google Scholar] [CrossRef]
- Park, H.S.; Hohn, M.J.; Umehara, T.; Guo, L.-T.; Osborne, E.M.; Benner, J.; Noren, C.J.; Rinehart, J.; Söll, D. Expanding the genetic code of Escherichia coli with phosphoserine. Science 2011, 333, 1151–1154. [Google Scholar] [CrossRef] [PubMed]
- Fan, C.; Ip, K.; Söll, D. Expanding the genetic code of Escherichia coli with phosphotyrosine. FEBS Lett. 2016, 590, 3040–3047. [Google Scholar] [CrossRef] [PubMed]
- Nehring, S.; Budisa, N.; Wiltschi, B. Performance analysis of orthogonal pairs designed for an expanded eukaryotic genetic code. PLoS ONE 2012, 7, e31992. [Google Scholar] [CrossRef] [PubMed]
- Tanrikulu, I.C.; Schmitt, E.; Mechulam, Y.; Goddard, W.A., III.; Tirrell, D.A. Discovery of Escherichia coli methionyl-tRNA synthetase mutants for efficient labeling of proteins with azidonorleucine in vivo. Proc. Natl. Acad. Sci. USA 2009, 106, 15285–15290. [Google Scholar] [CrossRef] [PubMed]
- Boniecki, M.T.; Vu, M.T.; Betha, A.K.; Martinis, S.A. CP1-dependent partitioning of pretransfer and posttransfer editing in leucyl-tRNA synthetase. Proc. Natl. Acad. Sci. USA 2008, 105, 19223–19228. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, N.M.; Ling, J.; Roy, H.; Banerjee, R.; Repasky, S.E.; Hamel, P.; Ibba, M. Cell-specific differences in the requirements for translation quality control. Proc. Natl. Acad. Sci. USA 2010, 107, 4063–4068. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.S.; Brunner, S.F.; Huguenin-Dezot, N.; Liang, A.D.; Schmied, W.H.; Rogerson, D.T.; Chin, J.W. Biosynthesis and genetic encoding of phosphothreonine through parallel selection and deep sequencing. Nat. Methods 2017, 14, 729–736. [Google Scholar] [CrossRef] [PubMed]
- Ellefson, J.W.; Meyer, A.J.; Hughes, R.A.; Cannon, J.R.; Brodbelt, J.S.; Ellington, A.D. Directed evolution of genetic parts and circuits by compartmentalized partnered replication. Nat. Biotechnol. 2014, 32, 97–101. [Google Scholar] [CrossRef] [PubMed]
- Bryson, D.I.; Fan, C.; Guo, L.-T.; Miller, C.; Söll, D.; Liu, D.R. Continuous directed evolution of aminoacyl-tRNAsynthetases. Nat. Chem. Biol. 2017, 13, 1253–1260. [Google Scholar] [CrossRef] [PubMed]
- Kunjapura, A.M.; Stork, D.A.; Kuru, E.; Vargas-Rodriguez, O.; Landon, M.; Söll, D.; Church, G.M. Engineering posttranslational proofreading to discriminate nonstandard amino acids. Proc. Natl. Acad. Sci. USA 2018, 115, 619–624. [Google Scholar] [CrossRef]
- Ohno, S.; Yokogawa, T.; Nishikawa, K. Changing the amino acid specificity of yeast tyrosyl-tRNA synthetase by genetic engineering. J. Biochem. 2001, 130, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Kiga, D.; Sakamoto, K.; Kodama, K.; Kigawa, T.; Matsuda, T.; Yabuki, T.; Shirouzu, M.; Harada, Y.; Nakayama, H.; Tako, K.; et al. An engineered Escherichia coli tyrosyl-tRNA synthetase for site-specific incorporation of an unnatural amino acid into proteins in eukaryotic translation and its application in a wheat germ cell-free system. Proc. Natl. Acad. Sci. USA 2002, 99, 9715–9720. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, T.; Ishii, R.; Fukunaga, R.; Kobayashi, T.; Sakamoto, K.; Yokoyama, S. Multistep engineering of pyrrolysyl-tRNA synthetase to genetically encode Nε-(o-azidobenzyloxycarbonyl) lysine for site-specific protein modification. Chem Biol. 2008, 15, 1187–1197. [Google Scholar] [CrossRef] [PubMed]
- Oki, K.; Sakamoto, K.; Kobayashi, T.; Sasaki, H.M.; Yokoyama, S. Transplantation of a tyrosine editing domain into a tyrosyl-tRNA synthetase variant enhances its specificity for a tyrosine analog. Proc. Natl. Acad. Sci. USA 2008, 105, 13298–13303. [Google Scholar] [CrossRef] [PubMed]
- O’Donoghue, P.; Ling, J.; Wang, Y.S.; Söll, D. Upgrading protein synthesis for synthetic biology. Nat. Chem. Biol. 2013, 9, 594–598. [Google Scholar] [CrossRef] [PubMed]
- Vargas-Rodriguez, O.; Sevostyanova, A.; Söll, D.; Crnković, A. Upgrading aminoacyl-tRNA synthetases for genetic code expansion. Curr. Opin. Chem. Biol. 2018, 46, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.C.; Qi, L.; Yanofsky, C.; Arkin, A.P. Regulation of transcription by unnatural amino acids. Nat. Biotechnol. 2011, 29, 164–168. [Google Scholar] [CrossRef]
- Cruz-Vera, L.R.; Sachs, M.S.; Squires, C.L.; Yanofsky, C. Nascent polypeptide sequences that influence ribosome function. Curr. Opin. Microbiol. 2011, 14, 160–166. [Google Scholar] [CrossRef]
- Ito, K.; Chiba, S. Arrest peptides: Cis-acting modulators of translation. Annu. Rev. Biochem. 2013, 82, 171–202. [Google Scholar] [CrossRef]
- Bilezikian, J.; Kaempfer, R.O.R.; Magasanik, B. Mechanism of tryptophanase induction in Escherichia coli. J. Mol. Biol. 1967, 27, 495–506. [Google Scholar] [CrossRef]
- Stewart, V.; Landick, R.; Yanofsky, C. Rho-dependent transcription termination in the tryptophanase operon leader region of Escherichia coli K-12. J. Bacteriol. 1986, 166, 217–223. [Google Scholar] [CrossRef] [PubMed]
- Gong, F.; Ito, K.; Nakamura, Y.; Yanofsky, C. The mechanism of tryptophan induction of tryptophanase operon expression: Tryptophan inhibits release factor-mediated cleavage of TnaC-peptidyl-tRNA(Pro). Proc. Natl. Acad. Sci. USA 2001, 98, 8997–9001. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Vera, L.R.; Rajagopal, S.; Squires, C.; Yanofsky, C. Features of ribosome-peptidyl-tRNA interactions essential for tryptophan induction of tna operon expression. Mol. Cell 2005, 19, 333–343. [Google Scholar] [CrossRef] [PubMed]
- Kato, Y. An engineered bacterium auxotrophic for an unnatural amino acid: A novel biological containment system. PeerJ 2015, 3, e1247. [Google Scholar] [CrossRef] [PubMed]
- Whitford, C.M.; Dymek, S.; Kerkhoff, D.; März, C.; Schmidt, O.; Edich, M.; Droste, J.; Pucker, B.; Rückert, C.; Kalinowski, J. Auxotrophy to Xeno-DNA: An exploration of combinatorial mechanisms for a highfidelity biosafety system for synthetic biology applications. J. Biol. Eng. 2018, 12, 13. [Google Scholar] [CrossRef] [PubMed]
- Rovner, A.J.; Haimovich, A.D.; Katz, S.R.; Li, Z.; Grome, M.W.; Gassaway, B.M.; Amiram, M.; Patel, J.R.; Gallagher, R.R.; Rinehart, J.; et al. Recoded organisms engineered to depend on synthetic amino acids. Nature 2015, 518, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Mandell, D.J.; Lajoie, M.J.; Mee, M.T.; Takeuchi, R.; Kuznetsov, G.; Norville, J.E.; Gregg, C.J.; Stoddard, B.L.; Church, G.M. Biocontainment of genetically modifed organisms by synthetic protein design. Nature 2015, 518, 55–60. [Google Scholar] [CrossRef]
- Ramos, H.J.O.; Yates, M.G.; Pedrosa, F.O.; Souza, E.M. Strategies for bacterial tagging and gene expression in plant-host colonization studies. Soil Biol. Biochem. 2011, 43, 1626–1638. [Google Scholar] [CrossRef]
- Koh, M.; Nasertorabi, F.; Han, G.W.; Stevens, R.C.; Schultz, P.G. Generation of an orthogonal protein-protein interface with a noncanonical amino acid. J. Am. Chem. Soc. 2017, 139, 5728–5731. [Google Scholar] [CrossRef]
- Gan, F.; Liu, R.; Wang, F.; Schultz, P.G. Functional replacement of histidine in proteins to generate noncanonical amino acid dependent organisms. J. Am. Chem. Soc. 2018, 140, 3829–3832. [Google Scholar] [CrossRef]
- Wang, N.; Li, Y.; Niu, W.; Sun, M.; Cerny, R.; Li, Q.; Guo, J. Construction of a live-attenuated HIV-1 vaccine through genetic code expansion. Angew. Chem. Int. Ed. Engl. 2014, 53, 4867–4871. [Google Scholar] [CrossRef] [PubMed]
- Si, L.; Xu, H.; Zhou, X.; Zhang, Z.; Tian, Z.; Wang, Y.; Wu, Y.; Zhang, B.; Niu, Z.; Zhang, C.; et al. Generation of influenza A viruses as live but replication-incompetent virus vaccines. Science 2016, 354, 1170–1173. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Z.; Wang, N.; Kang, G.; Ziu, W.; Li, Q.; Guo, J. Controlling multicycle replication of live-attenuated HIV-1 using an unnatural genetic switch. ACS Synth. Biol. 2017, 6, 721–731. [Google Scholar] [CrossRef] [PubMed]
- Immerseel, F.V.; Methner, U.; Rychlik, I.; Nagy, B.; Velge, P.; Martin, G.; Foster, N.; Ducatelle, R.; Barrow, P.A. Vaccination and early protection against non-host-specific Salmonella serotypes in poultry: Exploitation of innate immunity and microbial activity. Epidemiol. Infect. 2005, 133, 959–978. [Google Scholar] [CrossRef] [PubMed]
- Crichton, M. Jurassic Park; Alfred A. Knopf: New York, NY, USA, 1990. [Google Scholar]
- Boeke, J.D.; Church, G.; Hessel, A.; Kelley, N.J.; Arkin, A.; Cai, Y.; Carlson, R.; Chakravarti, A.; Cornish, V.W.; Holt, L.; et al. The Genome Project–Write. Science 2016, 353, 126–127. [Google Scholar] [CrossRef] [PubMed]
- Annaluru, N.; Muller, H.; Mitchell, L.A.; Ramalingam, S.; Stracquadanio, G.; Richardson, S.M.; Dymond, J.S.; Kuang, Z.; Scheifele, L.Z.; Cooper, E.M.; et al. Total Synthesis of a Functional Designer Eukaryotic Chromosome. Science 2014, 344, 55–58. [Google Scholar] [CrossRef]
- Ostrov, N.; Landon, M.; Guell, M.; Kuznetsov, G.; Teramoto, J.; Cervantes, N.; Zhou, M.; Singh, K.; Napolitano, M.G.; Moosburner, M.; et al. Design, synthesis, and testing toward a 57-codon genome. Science 2016, 353, 819–822. [Google Scholar] [CrossRef]
- Wang, K.; Fredens, J.; Brunner, S.F.; Kim, S.H.; Chia, T.; Chin, J.W. Defining synonymous codon compression schemes by genome recoding. Nature 2016, 539, 59–64. [Google Scholar] [CrossRef]
- Neumann, H.; Wang, K.; Davis, L.; Garcia-Alai, M.; Chin, J.W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 2010, 464, 441–444. [Google Scholar] [CrossRef]


© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kato, Y. Translational Control using an Expanded Genetic Code. Int. J. Mol. Sci. 2019, 20, 887. https://doi.org/10.3390/ijms20040887
Kato Y. Translational Control using an Expanded Genetic Code. International Journal of Molecular Sciences. 2019; 20(4):887. https://doi.org/10.3390/ijms20040887
Chicago/Turabian StyleKato, Yusuke. 2019. "Translational Control using an Expanded Genetic Code" International Journal of Molecular Sciences 20, no. 4: 887. https://doi.org/10.3390/ijms20040887
APA StyleKato, Y. (2019). Translational Control using an Expanded Genetic Code. International Journal of Molecular Sciences, 20(4), 887. https://doi.org/10.3390/ijms20040887