LJELSR: A Strengthened Version of JELSR for Feature Selection and Clustering
Abstract
:1. Introduction
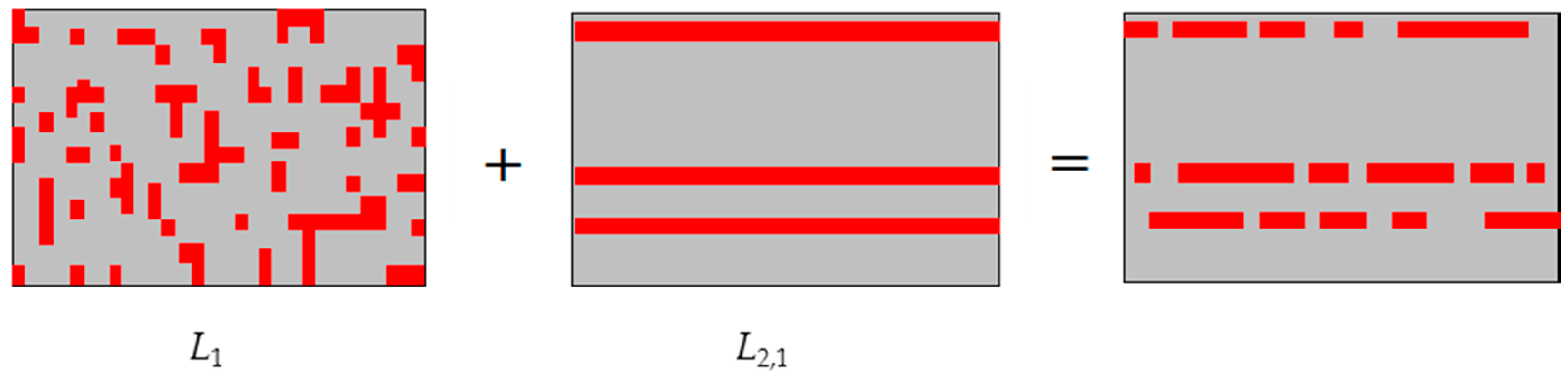
- More zero values are produced by adding an -norm constraint on the sparse regression matrix, such that we get more sparse results.
- The internal geometric structure of data is preserved along with dimensionality reduction by embedding learning to reduce the occurrence of inaccurate results.
- Although an exact solution cannot be obtained by our method, we provide a convergent iterative algorithm to get the optimal results.
2. Results
2.1. Datasets
2.2. Parameters Selection
2.3. Evaluation Metrics
2.4. Feature Selection Analysis
2.4.1. Experimental Results and Analysis on ALL_AML Dataset
2.4.2. Experimental Results and Analysis on Colon Cancer Dataset
2.4.3. Experimental Results and Analysis on ESCA Dataset
2.4.4. Differentially Expressed Genes Comparing by Methods
2.5. Clustering Analysis
3. Materials and Methods
3.1. Related Notations and Definitions
3.2. Joint Embedding Learning and Sparse Regression (JELSR)
3.3. The Proposed Method
3.4. Optimization
3.5. Feature Selection
| Algorithm 1. Procedure of LJELSR. |
| Input: Data matrix ; Neighborhood size ; Balance parameters ; Dimensionality of embedding ; Feature selection number . Output: Selected feature index set |
| Stage one: Graph construction Construct the weight matrix ; Compute the diagonal matrix , graph Laplacian matrix ; Stage two: Alternative optimization, Initialize ; Loop Update and fix , by (19), Update and fix , by (17), Update , and fix by (12) and (13). until convergence Stage three: Feature selection |
3.6. Convergence Analysis
4. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
Appendix A
References
- Church, G.M.; Gilbert, W. Genomic sequencing. Proc. Natl. Acad. Sci. USA 1984, 81, 1991–1995. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.C.; Boscolo, R.; Yang, Y.-L.; Tran, L.M.; Sabatti, C.; Roychowdhury, V.P. Network component analysis: Reconstruction of regulatory signals in biological systems. Proc. Natl. Acad. Sci. USA 2003, 100, 15522–15527. [Google Scholar] [CrossRef] [PubMed]
- Constantinopoulos, C.; Titsias, M.K.; Likas, A. Bayesian feature and model selection for gaussian mixture models. IEEE Trans. Pattern Anal. Mach. Intell. 2006, 28, 1013–1018. [Google Scholar] [CrossRef] [PubMed]
- Nie, F.; Zeng, Z.; Tsang, I.W.; Xu, D.; Zhang, C. Spectral embedded clustering: A framework for in-sample and out-of-sample spectral clustering. IEEE Trans. Neural Netw. 2011, 22, 1796–1808. [Google Scholar] [PubMed]
- Hou, C.; Nie, F.; Yi, D.; Wu, Y. Feature selection via joint embedding learning and sparse regression. In Proceedings of the International Joint Conference on Artificial Intelligence (IJCAI 2011), Barcelona, Spain, 16–22 July 2011; pp. 1324–1329. [Google Scholar]
- D’Addabbo, A.; Papale, M.; Di Paolo, S.; Magaldi, S.; Colella, R.; d’Onofrio, V.; Di Palma, A.; Ranieri, E.; Gesualdo, L.; Ancona, N. Svd based feature selection and sample classification of proteomic data. In Knowledge-Based Intelligent Information and Engineering Systems; Springer: Berlin/Heidelberg, Germany, 2008; pp. 556–563. [Google Scholar]
- Cai, D.; He, X.; Han, J. Spectral regression for efficient regularized subspace learning. Proceedings 2007, 149, 1–8. [Google Scholar]
- Zhao, Z.; Wang, L.; Liu, H. Efficient spectral feature selection with minimum redundancy. In Proceedings of the Twenty-Fourth AAAI Conference on Artificial Intelligence (AAAI-10), Atlanta, GA, USA, 11–15 July 2010; pp. 673–678. [Google Scholar]
- Cai, D.; Zhang, C.; He, X. Unsupervised feature selection for multi-cluster data. In Proceedings of the ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Washington, DC, USA, 25–28 July 2010; pp. 333–342. [Google Scholar]
- Tibshirani, R.J. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Wang, H.; Nie, F.; Huang, H.; Risacher, S.; Ding, C.; Saykin, A.J.; Shen, L. Sparse multi-task regression and feature selection to identify brain imaging predictors for memory performance. In Proceedings of the International Conference on Computer Vision, Barcelona, Spain, 6–13 November 2011; pp. 557–562. [Google Scholar]
- Zhao, Q.; Meng, D.; Xu, Z. A recursive divide-and-conquer approach for sparse principal component analysis. arXiv, 2012; arXiv:1211.7219. [Google Scholar]
- Brunet, J.P.; Tamayo, P.; Golub, T.R.; Mesirov, J.P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl. Acad. Sci. USA 2004, 101, 4164–4169. [Google Scholar] [CrossRef] [PubMed]
- Alon, U.; Barkai, N.; Notterman, D.A.; Gish, K.; Ybarra, S.; Mack, D.; Levine, A.J. Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc. Natl. Acad. Sci. USA 1999, 96, 6745–6750. [Google Scholar] [CrossRef] [PubMed]
- Cai, D.; He, X.; Han, J.; Huang, T.S. Graph regularized nonnegative matrix factorization for data representation. IEEE Trans. Pattern Anal. Mach. Intell. 2011, 33, 1548–1560. [Google Scholar] [PubMed]
- Shang, R.; Wang, W.; Stolkin, R.; Jiao, L. Non-negative spectral learning and sparse regression-based dual-graph regularized feature selection. IEEE Trans. Cybern. 2017, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Schölkopf, B. A local learning approach for clustering. In Proceedings of the International Conference on Neural Information Processing Systems, Vancouver, BC, Canada, 4–7 December 2006; pp. 1529–1536. [Google Scholar]
- Aruffo, A.; Seed, B. Molecular cloning of two cd7 (T-cell leukemia antigen) cdnas by a cos cell expression system. EMBO J. 1987, 6, 3313–3316. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.Y.; Chen, C.Y.; Tien, H.F.; Lin, C.W. Loss of cd7, independent of galectin-3 expression, implies a worse prognosis in adult T-cell leukaemia/lymphoma. Histopathology 2009, 54, 214–220. [Google Scholar] [CrossRef] [PubMed]
- Lahortiga, I.; De Keersmaecker, K.; Van Vlierberghe, P.; Graux, C.; Cauwelier, B.; Lambert, F.; Mentens, N.; Beverloo, H.B.; Pieters, R.; Speleman, F. Duplication of the myb oncogene in t cell acute lymphoblastic leukemia. Nat. Genet. 2007, 39, 593–595. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Liu, S.; Wang, J.; Sun, M.-Z.; Greenaway, F.T. Actb in cancer. Clin. Chim. Acta 2013, 417, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Andersen, C.L.; Jensen, J.L.; Ørntoft, T.F. Normalization of real-time quantitative reverse transcription-pcr data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Nowakowska, M.; Pospiech, K.; Lewandowska, U.; Piastowska-Ciesielska, A.W.; Bednarek, A.K. Diverse effect of wwox overexpression in ht29 and sw480 colon cancer cell lines. Tumor Biol. 2014, 35, 9291–9301. [Google Scholar] [CrossRef] [PubMed]
- Dahlberg, P.S.; Jacobson, B.A.; Dahal, G.; Fink, J.M.; Kratzke, R.A.; Maddaus, M.A.; Ferrin, L.J. Erbb2 amplifications in esophageal adenocarcinoma. Ann. Thorac. Surg. 2004, 78, 1790–1800. [Google Scholar] [CrossRef] [PubMed]
- Bolling, M.; Lemmink, H.; Jansen, G.; Jonkman, M. Mutations in krt5 and krt14 cause epidermolysis bullosa simplex in 75% of the patients. Br. J. Dermatol. 2011, 164, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Liu, H. Web user clustering analysis based on kmeans algorithm. In Proceedings of the International Conference on Information NETWORKING and Automation, Kunming, China, 18–19 October 2010. [Google Scholar]
- Zhou, D.; Huang, J.; Schölkopf, B. Learning with hypergraphs: Clustering, classification, and embedding. In Proceedings of the Advances in Neural Information Processing Systems, Vancouver, BC, Canada, 3–6 December 2007; pp. 1601–1608. [Google Scholar]
- Nie, F.; Huang, H.; Cai, X.; Ding, C.H. Efficient and robust feature selection via joint ℓ2, 1-norms minimization. In Proceedings of the Advances in Neural Information Processing Systems, Vancouver, BC, Canada, 6–9 December 2010; pp. 1813–1821. [Google Scholar]
- Hou, C.; Nie, F.; Li, X.; Yi, D.; Wu, Y. Joint embedding learning and sparse regression: A framework for unsupervised feature selection. IEEE Trans. Cybern. 2014, 44, 793–804. [Google Scholar] [PubMed]
| Datasets | Genes | Samples | Classes | Description |
|---|---|---|---|---|
| ALL_AML | 5000 | 38 | 3 | acute lymphoblastic leukemia and acute myelogenous leukemia |
| colon | 2000 | 62 | 2 | colon cancer |
| ESCA | 20,502 | 192 | 2 | esophageal carcinoma |
| ID | LJELSR | JELSR | ReDac | SMART |
|---|---|---|---|---|
| GO:0006955 | 2.249 × 10−20 | 3.157 × 10−18 | 8.447 × 10−14 | 1.900 × 10−10 |
| GO:0050776 | 8.768 × 10−18 | 1.312 × 10−15 | 7.228 × 10−12 | 1.000 × 10−9 |
| GO:0045321 | 1.306 × 10−16 | 1.808 × 10−14 | 1.154 × 10−11 | 2.500 × 10−10 |
| GO:0001775 | 1.548 × 10−15 | 1.604 × 10−13 | 6.621 × 10−11 | 1.590 × 10−10 |
| GO:0051251 | 2.005 × 10−15 | 4.157 × 10−14 | 9.117 × 10−12 | 4.380 × 10−12 |
| GO:0007159 | 2.098 × 10−15 | 2.754 × 10−15 | 3.712 × 10−11 | 1.510 × 10−10 |
| GO:0002682 | 2.477 × 10−15 | 2.725 × 10−14 | 7.817 × 10−12 | 5.870 × 10−8 |
| GO:0046649 | 3.964 × 10−15 | 5.426 × 10−14 | 3.164 × 10−10 | 1.540 × 10−11 |
| GO:0016337 | 7.034 × 10−15 | 8.993 × 10−14 | 5.253 × 10−11 | 1.940 × 10−11 |
| GO:0070486 | 7.220 × 10−15 | 9.350 × 10−15 | 1.299 × 10−11 | 5.400 × 10−10 |
| Gene | Gene Official Name | Related Diseases |
|---|---|---|
| CD34 | CD34 Molecule | Dermatofibrosarcoma Protuberans and Hypercalcemic Type Ovarian Small Cell Carcinoma |
| CD7 | CD7 Molecule | Pityriasis Lichenoides Et Varioliformis Acuta and T-Cell Leukemia |
| MYB | MYB Proto-Oncogene, Transcription Factor | Acute Basophilic Leukemia and Angiocentric Glioma |
| CXCR4 | C-X-C Motif Chemokine Receptor 4 | Whim Syndrome and Human Immunodeficiency Virus Infectious Disease |
| CTSG | Cathepsin G | Papillon-Lefevre Syndrome and Cutaneous Mastocytosis |
| ID | LJELSR | JELSR | ReDac | SMART |
|---|---|---|---|---|
| GO:0006614 | 1.612 × 10−17 | 9.836 × 10−14 | 2.677 × 10−14 | 1.016 × 10−12 |
| GO:0006613 | 3.970 × 10−17 | 2.060 × 10−13 | 5.617 × 10−14 | 1.970 × 10−12 |
| GO:0045047 | 5.074 × 10−17 | 2.519 × 10−13 | 6.872 × 10−14 | 2.360 × 10−12 |
| GO:0072599 | 8.161 × 10−17 | 3.720 × 10−13 | 1.016 × 10−13 | 3.348 × 10−12 |
| GO:0022626 | 3.104 × 10−16 | 8.999 × 10−13 | 2.465 × 10−13 | 1.125 × 10−11 |
| GO:0000184 | 3.751 × 10−16 | 1.301 × 10−12 | 3.568 × 10−13 | 1.029 × 10−11 |
| GO:0003735 | 5.428 × 10−16 | 6.787 × 10−13 | 1.412 × 10−13 | 3.310 × 10−12 |
| GO:0070972 | 6.180 × 10−16 | 1.960 × 10−12 | 5.384 × 10−13 | 1.488 × 10−11 |
| GO:0019083 | 9.571 × 10−16 | 1.932 × 10−12 | 1.547 × 10−11 | 3.005 × 10−10 |
| GO:0044445 | 1.372 × 10−15 | 1.487 × 10−12 | 3.106 × 10−13 | 8.772 × 10−12 |
| Gene | Gene Official Name | Related Diseases |
|---|---|---|
| MUC3A | Mucin 3A, Cell Surface Associated | Cap Polyposis and Hypertrichotic Osteochondrodysplasia |
| ACTB | Actin Beta | Dystonia, Juvenile-Onset and Baraitser-Winter Syndrome 1 |
| WWOX | WW Domain Containing Oxidoreductase | Spinocerebellar Ataxia, Autosomal Recessive 12andEpileptic Encephalopathy, Early Infantile, 28 |
| SPI1 | Spi-1 Proto-Oncogene | Inflammatory Diarrhea and Interdigitating Dendritic Cell Sarcoma |
| RPS24 | Ribosomal Protein S24 | Diamond-Blackfan Anemia 3 and Diamond-Blackfan Anemia |
| ID | LJELSR | JELSR | ReDac | SMART |
|---|---|---|---|---|
| GO:0005198 | 2.772 × 10−27 | 8.941 × 10−18 | 1.096 × 10−18 | 1.036 × 10−4 |
| GO:0070161 | 7.504 × 10−21 | 3.347 × 10−14 | 2.733 × 10−23 | 6.527 × 10−14 |
| GO:0030055 | 1.400 × 10−19 | 1.111 × 10−10 | 3.576 × 10−17 | 1.619 × 10−12 |
| GO:0005912 | 6.655 × 10−19 | 1.946 × 10−11 | 4.401 × 10−20 | 3.498 × 10−12 |
| GO:0005925 | 1.319 × 10−18 | 7.671 × 10−11 | 2.103 × 10−17 | 1.061 × 10−12 |
| GO:0005924 | 1.746 × 10−18 | 9.246 × 10−11 | 2.747 × 10−17 | 1.313 × 10−12 |
| GO:0005615 | 5.538 × 10−16 | 8.395 × 10−20 | 3.985 × 10−15 | 2.117 × 10−17 |
| GO:0030054 | 5.243 × 10−14 | 4.156 × 10−10 | 5.243 × 10−14 | 4.510 × 10−9 |
| GO:0005200 | 3.076 × 10−13 | 1.301 × 10−10 | 2.062 × 10−13 | 2.358 × 10−4 |
| GO:0042060 | 7.442 × 10−13 | 2.650 × 10−9 | 4.536 × 10−12 | 8.790 × 10−11 |
| Gene | Gene Official Name | Related Diseases |
|---|---|---|
| ERBB2 | Erb-B2 Receptor Tyrosine Kinase 2 | Glioma Susceptibility 1andOvarian Cancer, Somatic |
| KRT14 | Keratin 14 | Epidermolysis Bullosa Simplex, Koebner Type and Epidermolysis Bullosa Simplex, Recessive 1 |
| KRT5 | Keratin 5 | Epidermolysis Bullosa Simplex, Dowling-Meara Type and Epidermolysis Bullosa Simplex, Weber-Cockayne Type |
| KRT19 | Keratin 19 | Anal Canal Adenocarcinoma and Thyroid Cancer |
| KRT4 | Keratin 4 | White Sponge Nevus 1andWhite Sponge Nevus Of Cannon, Krt4-Related |
| Methods | Differentially Expressed Genes | Number |
|---|---|---|
| LJELSR | ERBB2, KRT14, KRT5, KRT19, KRT4, TFF1,FSCN1, KRT13,ITGB4, ANXA1, MUC6,LAMC2,HLA-B, KRT16, JUP, KRT17,LAMB3, ATP4A, DSP,LAMA3, FOS, FN1, CTSB, MYH11,HLA-C, LDHA, PKM,SLC7A5, PSCA, SERPINA1, S100A7, S100A9, CRNN, S100A8, B2M, DMBT1, CD24,ENO1, TNC,KRT15, GLUL,HSPA1A, NDRG1, LCN2,COL17A1,CEACAM6, REG1A, PLEC, GAPDH, PIGR, AGR2,ANPEP, PKP1, ACTB, FLNA,PI3, FTL, CTSE,PABPC1, PGC, ALDOA, EEF2,KRT6B, LYZ, CLDN18, SPRR1B,KRT6C,PPP1R1B, PGA3,COL3A1,C3,REG1B,PERP, KRT6A,PGA5,CES1,PGA4,EIF1 | 78 |
| JELSR | MUC1, KRT14, KRT5, KRT19, KRT8, KRT4, TFF1, FN1, MUC6, S100A8, CTSD, SPRR3, ATP4A, HSPB1, FOS, CEACAM5, GJB2, H19, EZR, KRT16, KRT13, CTSB, JUP, ANXA1, TFF2, S100A9, SFN, KRT17, PSCA, S100A2, MUC5B, COL1A1, CD24, PKM, DSC3, GLUL, MALAT1, REG1A, ACTN4, DSG3, LCN2, DSP, S100A7, B2M, MYH9, PKP1, S100A11, PIGR, HSPA8, PLEC, TRIM29, EEF1A1, ATP1B1, AGR2, LYZ, ACTB, PGC, PABPC1, SPRR2A, SPRR1B, SPRR1A, CA2, REG4, P4HB, CLDN18, CTSE, EEF2, CREB3L1, KRT6A, A2M, PGA3 | 71 |
| ReDac | KRT14, KRT5, KRT4, FN1, CRNN, TGM3, MUC6, S100A8, IL1RN, SPRR3, ATP4A, HSPB1, MAL, KRT16, KRT13, JUP, THBS1, ANXA1, S100A9, PSCA, ECM1, CD24, PKM, FTL, HSPG2, DES, GLUL, MALAT1, PPL, EMP1, ACTN4, MYH11, CSTA, GAPDH, TAGLN, DSP, B2M, MYH9, GSN, PKP1, S100A11, PIGR, HSPA8, FLNA, PLEC, MYLK, CSTB, TRIM29, EEF1A1, RPL3, LYZ, PSAP, ACTB, ALDOA, PGC, PABPC1, SYNM, SPRR2A, SPRR1B, SPRR1A, REG4, P4HB, EEF2, KRT6A, ACTG2, PGA3 | 66 |
| SMART | ERBB2, CCND1, GSTP1, CD44, KRT19, MUC4, MUC2, GRB7, TFF1, HLA-A, FN1, SOD2, ITGA6, NDRG1, SPP1, SERPINA1, MUC6, CTSD, HSPB1, CEACAM5, H19, CTSB, F5, ITGB1, ANXA1, SDC1, DMBT1, CLU, LDHA, CD24, APP, PKM, FTL, HSPG2, TNC, GLUL, MALAT1, NTS, LCN2, MYH9, PIGR, FLNA, CD55, PLEC, TSPAN8, EEF1A1, AGR2, LYZ, GPX2, ACTB, DSG2, PABPC1, SPRR1B, REG4, SCD, CLDN18, FAT1 | 57 |
| Methods | ALL_AML | Colon | ESCA |
|---|---|---|---|
| LJELSR | 81.579 | 64.520 | 96.354 |
| JELSR | 81.579 | 61.290 | 95.833 |
| ReDac | 68.421 | 61.290 | 95.313 |
| SMART | 44.740 | 63.980 | 94.790 |
| Kmeans | 78.530 | 53.420 | 96.350 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, S.-S.; Hou, M.-X.; Feng, C.-M.; Liu, J.-X. LJELSR: A Strengthened Version of JELSR for Feature Selection and Clustering. Int. J. Mol. Sci. 2019, 20, 886. https://doi.org/10.3390/ijms20040886
Wu S-S, Hou M-X, Feng C-M, Liu J-X. LJELSR: A Strengthened Version of JELSR for Feature Selection and Clustering. International Journal of Molecular Sciences. 2019; 20(4):886. https://doi.org/10.3390/ijms20040886
Chicago/Turabian StyleWu, Sha-Sha, Mi-Xiao Hou, Chun-Mei Feng, and Jin-Xing Liu. 2019. "LJELSR: A Strengthened Version of JELSR for Feature Selection and Clustering" International Journal of Molecular Sciences 20, no. 4: 886. https://doi.org/10.3390/ijms20040886
APA StyleWu, S.-S., Hou, M.-X., Feng, C.-M., & Liu, J.-X. (2019). LJELSR: A Strengthened Version of JELSR for Feature Selection and Clustering. International Journal of Molecular Sciences, 20(4), 886. https://doi.org/10.3390/ijms20040886





