Sinorhizobium fredii HH103 RirA Is Required for Oxidative Stress Resistance and Efficient Symbiosis with Soybean
Abstract
:1. Introduction
2. Results
2.1. The Lack of RirA in S. fredii HH103 Confers a Siderophore-Independent Growth Advantage under Low Iron Conditions
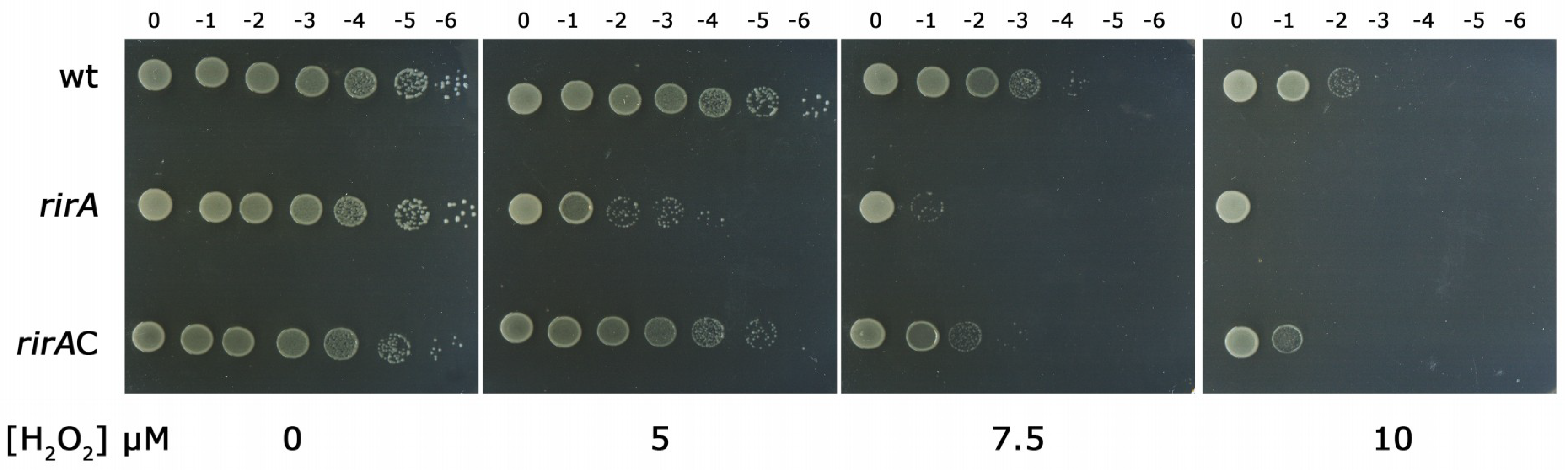
2.2. RirA Plays A Role in Oxidative Stress Resistance in HH103
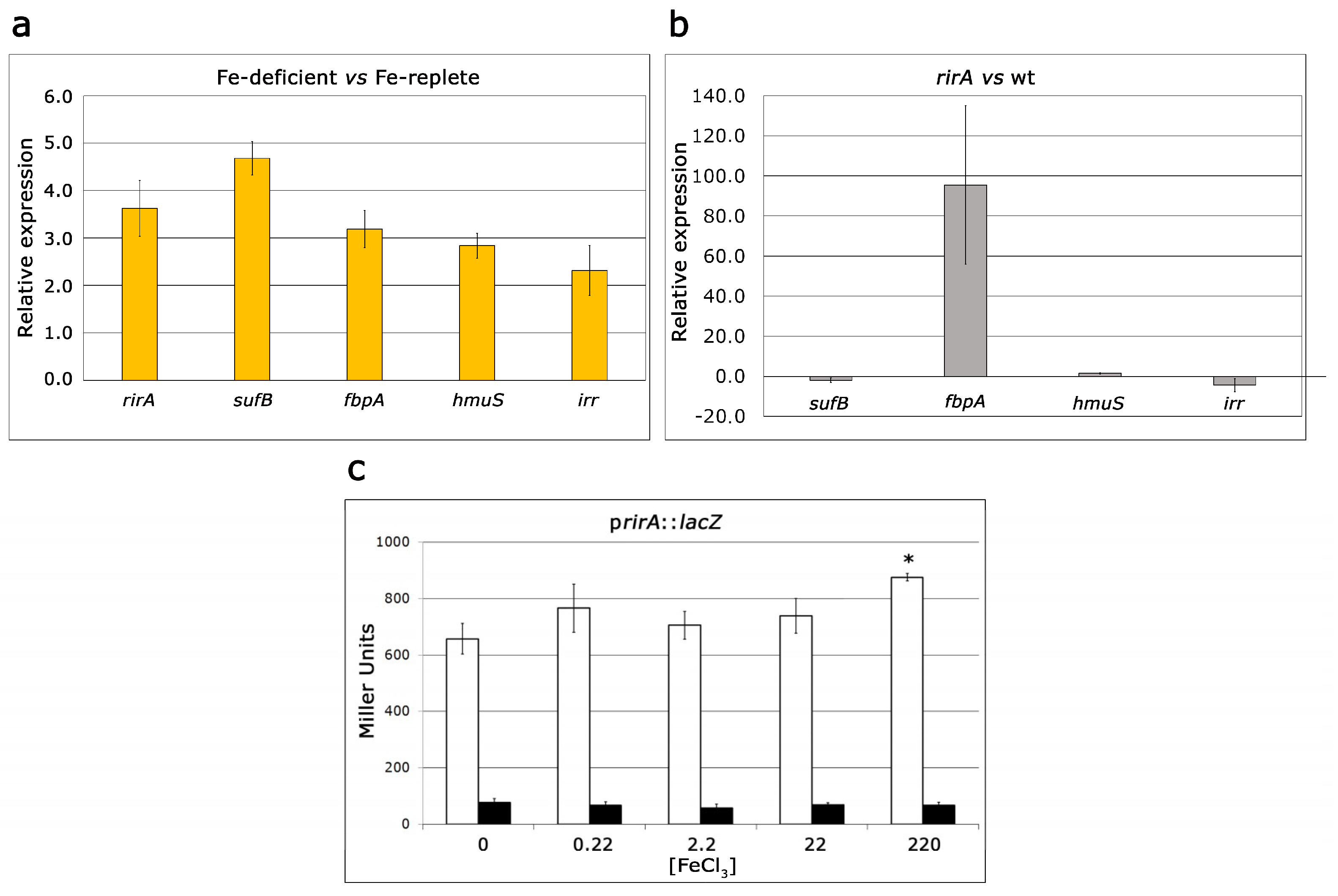
2.3. Role of RirA in Iron-Responsive Gene Expression
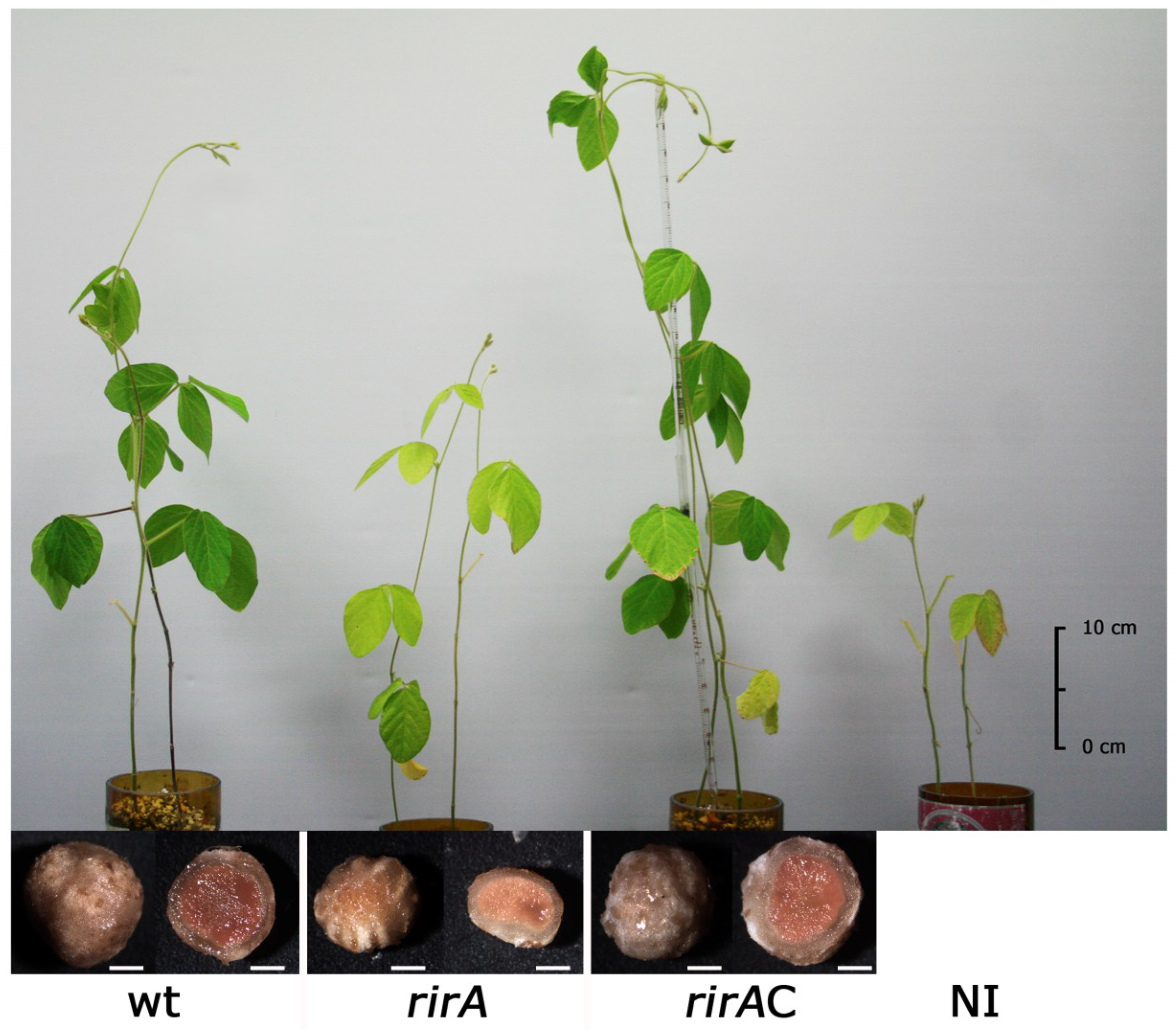
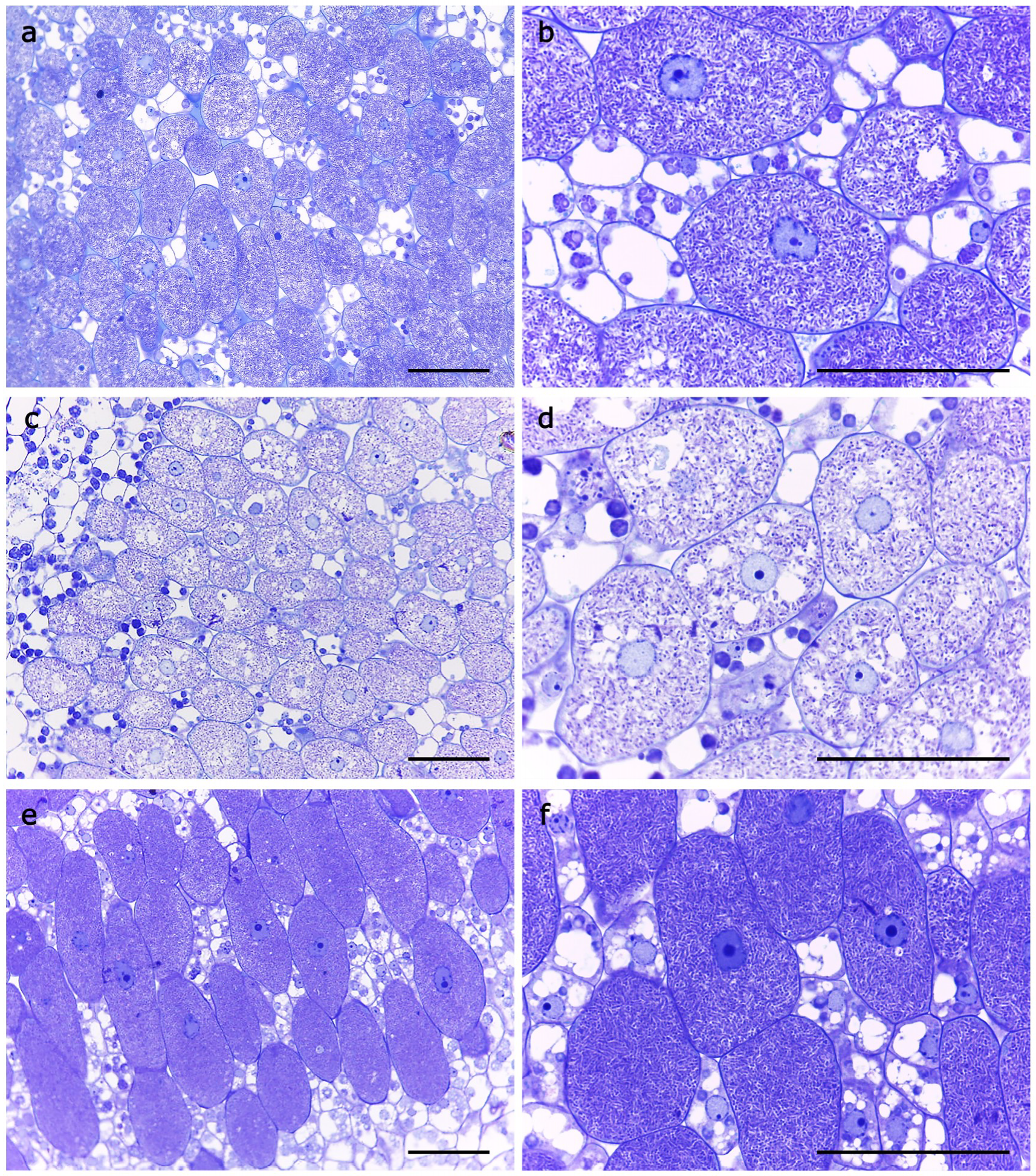
2.4. Nitrogen-Fixing Symbiosis with Soybean Plants Is Impaired in an S. fredii HH103 rirA Mutant
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains, Plasmids, and Growth Conditions
4.2. Construction of the S. fredii rirA Mutant and Complementation.
4.3. Siderophore Detection
4.4. Sensitivity to H2O2
4.5. Reverse Transcription Quantitative PCR (RT-qPCR)
4.6. Nodulation Assays
4.7. Microscopy Studies
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Andrews, S.C.; Robinson, A.K.; Rodríguez-Quiñones, F. Bacterial iron homeostasis. FEMS Microbiol. Rev. 2003, 27, 215–237. [Google Scholar] [CrossRef]
- Imlay, J.A. Pathways of oxidative damage. Annu. Rev. Microbiol. 2003, 57, 395–418. [Google Scholar] [CrossRef] [PubMed]
- Imlay, J.A. The mismetallation of enzymes during oxidative stress. J. Biol. Chem. 2014, 289, 28121–28128. [Google Scholar] [CrossRef] [PubMed]
- Saha, R.; Saha, N.; Donofrio, R.S.; Bestervelt, L.L. Microbial siderophores: A mini review. J. Basic Microbiol. 2013, 53, 303–317. [Google Scholar] [CrossRef] [PubMed]
- Frawley, E.R.; Fang, F.C. The ins and outs of bacterial iron metabolism. Mol. Microbiol. 2014, 93, 609–616. [Google Scholar] [CrossRef] [PubMed]
- Chandrangsu, P.; Rensing, C.; Helmann, J.D. Metal homeostasis and resistance in bacteria. Nat. Rev. Microbiol. 2017, 15, 338–350. [Google Scholar] [CrossRef] [PubMed]
- Pi, H.; Helmann, J.D. Ferrous iron efflux systems in bacteria. Metallomics 2017, 9, 840–851. [Google Scholar] [CrossRef] [PubMed]
- Hantke, K. Regulation of ferric iron transport in Escherichia coli K12: Isolation of a constitutive mutant. Mol. Gen. Genet. 1981, 182, 288–292. [Google Scholar] [CrossRef] [PubMed]
- Bsat, N.; Herbig, A.; Casillas-Martinez, L.; Setlow, P.; Helmann, J.D. Bacillus subtilis contains multiple Fur homologues: Identification of the iron uptake (Fur) and peroxide regulon (PerR) repressors. Mol. Microbiol. 1998, 29, 189–198. [Google Scholar] [CrossRef]
- Fleischhacker, A.S.; Kiley, P.J. Iron-containing transcription factors and their roles as sensors. Curr. Opin. Chem. Biol. 2011, 15, 335–341. [Google Scholar] [CrossRef]
- Poole, P.; Ramachandran, V.; Terpolilli, J. Rhizobia: From saprophytes to endosymbionts. Nat. Rev. Microbiol. 2018, 16, 291–303. [Google Scholar] [CrossRef] [PubMed]
- Chao, T.C.; Becker, A.; Buhrmester, J.; Pühler, A.; Weidner, S. The Sinorhizobium meliloti fur gene regulates, with dependence on Mn(II), transcription of the sitABCD operon, encoding a metal-type transporter. J. Bacteriol. 2004, 186, 3609–3620. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Mireles, E.; Wexler, M.; Sawers, G.; Bellini, D.; Todd, J.D.; Johnston, A.W. The Fur-like protein Mur of Rhizobium leguminosarum is a Mn(2+)-responsive transcriptional regulator. Microbiology 2004, 150, 1447–1456. [Google Scholar] [CrossRef] [PubMed]
- Rodionov, D.A.; Gelfand, M.S.; Todd, J.D.; Curson, A.R.; Johnston, A.W. Computational reconstruction of iron- and manganese-responsive transcriptional networks in alpha-proteobacteria. PLoS Comput. Biol. 2006, 2, e163. [Google Scholar] [CrossRef] [PubMed]
- Rudolph, G.; Hennecke, H.; Fischer, H.M. Beyond the Fur paradigm: Iron-controlled gene expression in rhizobia. FEMS Microbiol. Rev. 2006, 30, 631–648. [Google Scholar] [CrossRef] [PubMed]
- Johnston, A.W.; Todd, J.D.; Curson, A.R.; Lei, S.; Nikolaidou-Katsaridou, N.; Gelfand, M.S.; Rodionov, D.A. Living without Fur: The subtlety and complexity of iron-responsive gene regulation in the symbiotic bacterium Rhizobium and other alpha-proteobacteria. Biometals 2007, 20, 501–511. [Google Scholar] [CrossRef] [PubMed]
- O’Brian, M.R. Perception and homeostatic control of iron in the Rhizobia and related bacteria. Annu. Rev. Microbiol. 2015, 69, 229–245. [Google Scholar] [CrossRef]
- Hamza, I.; Chauhan, S.; Hassett, R.; O’Brian, M.R. The bacterial Irr protein is required for coordination of heme biosynthesis with iron availability. J. Biol. Chem. 1998, 273, 21669–21674. [Google Scholar] [CrossRef]
- Yeoman, K.H.; Curson, A.R.; Todd, J.D.; Sawers, G.; Johnston, A.W. Evidence that the Rhizobium regulatory protein RirA binds to cis-acting iron-responsive operators (IROs) at promoters of some Fe-regulated genes. Microbiology 2004, 150, 4065–4074. [Google Scholar] [CrossRef]
- Pellicer Martinez, M.T.; Martinez, A.B.; Crack, J.C.; Holmes, J.D.; Svistunenko, D.A.; Johnston, A.W.B.; Cheesman, M.R.; Todd, J.D.; Le Brun, N.E. Sensing iron availability via the fragile [4Fe-4S] cluster of the bacterial transcriptional repressor RirA. Chem. Sci. 2017, 8, 8451–8463. [Google Scholar] [CrossRef]
- Todd, J.D.; Wexler, M.; Sawers, G.; Yeoman, K.H.; Poole, P.S.; Johnston, A.W. RirA, an iron-responsive regulator in the symbiotic bacterium Rhizobium leguminosarum. Microbiology 2002, 148, 4059–4071. [Google Scholar] [CrossRef] [PubMed]
- Todd, J.D.; Sawers, G.; Johnston, A.W. Proteomic analysis reveals the wide-ranging effects of the novel, iron-responsive regulator RirA in Rhizobium leguminosarum bv. viciae. Mol. Genet. Genom. 2005, 273, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Chao, T.C.; Buhrmester, J.; Hansmeier, N.; Pühler, A.; Weidner, S. Role of the regulatory gene rirA in the transcriptional response of Sinorhizobium meliloti to iron limitation. Appl. Environ. Microbiol. 2005, 71, 5969–5982. [Google Scholar] [CrossRef] [PubMed]
- Viguier, C.; Cuiv, O.; Clarke, P.; O’Connell, M. RirA is the iron response regulator of the rhizobactin 1021 biosynthesis and transport genes in Sinorhizobium meliloti 2011. FEMS Microbiol. Lett. 2005, 246, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Costa, D.; Amarelle, V.; Valverde, C.; O’Brian, M.R.; Fabiano, E. The Irr and RirA proteins participate in a complex regulatory circuit and act in concert to modulate bacterioferritin expression in Ensifer meliloti 1021. Appl. Environ. Microbiol. 2017, 83. [Google Scholar] [CrossRef]
- Ngok-Ngam, P.; Ruangkiattikul, N.; Mahavihakanont, A.; Virgem, S.S.; Sukchawalit, R.; Mongkolsuk, S. Roles of Agrobacterium tumefaciens RirA in iron regulation, oxidative stress response, and virulence. J. Bacteriol. 2009, 191, 2083–2090. [Google Scholar] [CrossRef]
- Hibbing, M.E.; Fuqua, C. Antiparallel and interlinked control of cellular iron levels by the Irr and RirA regulators of Agrobacterium tumefaciens. J. Bacteriol. 2011, 193, 3461–3472. [Google Scholar] [CrossRef]
- Dokpikul, T.; Chaoprasid, P.; Saninjuk, K.; Sirirakphaisarn, S.; Johnrod, J.; Nookabkaew, S.; Sukchawalit, R.; Mongkolsuk, S. Regulation of the Cobalt/Nickel Efflux Operon dmeRF in Agrobacterium tumefaciens and a Link between the Iron-Sensing Regulator RirA and Cobalt/Nickel Resistance. Appl. Environ. Microbiol. 2016, 82, 4732–4742. [Google Scholar] [CrossRef]
- Margaret, I.; Becker, A.; Blom, J.; Bonilla, I.; Goesmann, A.; Gottfert, M.; Lloret, J.; Mittard-Runte, V.; Ruckert, C.; Ruiz-Sainz, J.E.; et al. Symbiotic properties and first analyses of the genomic sequence of the fast growing model strain Sinorhizobium fredii HH103 nodulating soybean. J. Biotechnol. 2011, 155, 11–19. [Google Scholar] [CrossRef]
- López-Baena, F.J.; Ruiz-Sainz, J.E.; Rodríguez-Carvajal, M.A.; Vinardell, J.M. Bacterial Molecular Signals in the Sinorhizobium fredii-Soybean Symbiosis. Int. J. Mol. Sci. 2016, 17, 755. [Google Scholar] [CrossRef]
- Weidner, S.; Becker, A.; Bonilla, I.; Jaenicke, S.; Lloret, J.; Margaret, I.; Puhler, A.; Ruiz-Sainz, J.E.; Schneiker-Bekel, S.; Szczepanowski, R.; et al. Genome sequence of the soybean symbiont Sinorhizobium fredii HH103. J. Bacteriol. 2012, 194, 1617–1618. [Google Scholar] [CrossRef]
- Vinardell, J.M.; Acosta-Jurado, S.; Zehner, S.; Gottfert, M.; Becker, A.; Baena, I.; Blom, J.; Crespo-Rivas, J.C.; Goesmann, A.; Jaenicke, S.; et al. The Sinorhizobium fredii HH103 genome: A comparative analysis with S. fredii strains differing in their symbiotic behavior with soybean. Mol. Plant Microbe Interact. 2015, 28, 811–824. [Google Scholar] [CrossRef] [PubMed]
- Carter, R.A.; Yeoman, K.H.; Klein, A.; Hosie, A.H.; Sawers, G.; Poole, P.S.; Johnston, A.W. dpp genes of Rhizobium leguminosarum specify uptake of delta-aminolevulinic acid. Mol. Plant Microbe Interact. 2002, 15, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Novichkov, P.S.; Kazakov, A.E.; Ravcheev, D.A.; Leyn, S.A.; Kovaleva, G.Y.; Sutormin, R.A.; Kazanov, M.D.; Riehl, W.; Arkin, A.P.; Dubchak, I.; et al. RegPrecise 3.0—a resource for genome-scale exploration of transcriptional regulation in bacteria. BMC Genom. 2013, 14, 745. [Google Scholar] [CrossRef]
- Parker Siburt, C.J.; Mietzner, T.A.; Crumbliss, A.L. FbpA--a bacterial transferrin with more to offer. Biochim. Biophys. Acta 2012, 1820, 379–392. [Google Scholar] [CrossRef] [PubMed]
- Guerinot, M.L.; Meidl, E.J.; Plessner, O. Citrate as a siderophore in Bradyrhizobium japonicum. J. Bacteriol. 1990, 172, 3298–3303. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.H.; Chen, W.H.; Wang, E.T.; Chen, W.F.; Yan, J.; Han, X.Z.; Tian, C.F.; Sui, X.H.; Singh, R.P.; Jiang, G.M.; et al. Rhizobial biogeography and inoculation application to soybean in four regions across China. J. Appl. Microbiol. 2018, 125, 853–866. [Google Scholar] [CrossRef]
- Jiao, J.; Wu, L.J.; Zhang, B.; Hu, Y.; Li, Y.; Zhang, X.X.; Guo, H.J.; Liu, L.X.; Chen, W.X.; Zhang, Z.; et al. MucR is required for transcriptional activation of conserved ion transporters to support nitrogen fixation of Sinorhizobium fredii in soybean nodules. Mol. Plant Microbe Interact. 2016, 29, 352–361. [Google Scholar] [CrossRef]
- Acosta-Jurado, S.; Alias-Villegas, C.; Navarro-Gómez, P.; Zehner, S.; Murdoch, P.D.; Rodríguez-Carvajal, M.A.; Soto, M.J.; Ollero, F.J.; Ruiz-Sainz, J.E.; Gottfert, M.; et al. The Sinorhizobium fredii HH103 MucR1 global regulator is connected with the nod regulon and is required for efficient symbiosis with Lotus burttii and Glycine max cv. Williams. Mol. Plant Microbe Interact. 2016, 29, 700–712. [Google Scholar] [CrossRef]
- Santos, R.; Herouart, D.; Sigaud, S.; Touati, D.; Puppo, A. Oxidative burst in alfalfa-Sinorhizobium meliloti symbiotic interaction. Mol. Plant Microbe Interact. 2001, 14, 86–89. [Google Scholar] [CrossRef]
- Bueno, P.; Soto, M.J.; Rodríguez-Rosales, M.P.; Sanjuan, J.; Olivares, J.; Donaire, J.P. Time-course of lipoxygenase, antioxidant enzyme activities and H2O2 accumulation during the early stages of Rhizobium-legume symbiosis. New Phytol. 2001, 152, 91–96. [Google Scholar] [CrossRef]
- Ramu, S.K.; Peng, H.M.; Cook, D.R. Nod factor induction of reactive oxygen species production is correlated with expression of the early nodulin gene rip1 in Medicago truncatula. Mol. Plant Microbe Interact. 2002, 15, 522–528. [Google Scholar] [CrossRef] [PubMed]
- Shaw, S.L.; Long, S.R. Nod factor inhibition of reactive oxygen efflux in a host legume. Plant Physiol. 2003, 132, 2196–2204. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.; Damiani, I.; Puppo, A.; Frendo, P. Redox changes during the legume-rhizobium symbiosis. Mol. Plant 2009, 2, 370–377. [Google Scholar] [CrossRef] [PubMed]
- Damiani, I.; Pauly, N.; Puppo, A.; Brouquisse, R.; Boscari, A. Reactive oxygen species and nitric oxide control early steps of the legume-Rhizobium symbiotic interaction. Front. Plant Sci. 2016, 7, 454. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1989. [Google Scholar]
- Beringer, J.E. R factor transfer in Rhizobium leguminosarum. J. Gen. Microbiol. 1974, 84, 188–198. [Google Scholar] [CrossRef] [PubMed]
- Nogales, J.; Domínguez-Ferreras, A.; Amaya-Gómez, C.V.; van Dillewijn, P.; Cuéllar, V.; Sanjuán, J.; Olivares, J.; Soto, M.J. Transcriptome profiling of a Sinorhizobium meliloti fadD mutant reveals the role of rhizobactin 1021 biosynthesis and regulation genes in the control of swarming. BMC Genom. 2010, 11, 157. [Google Scholar] [CrossRef]
- Becker, A.; Schmidt, M.; Jäger, W.; Pühler, A. New gentamicin-resistance and lacZ promoter-probe cassettes suitable for insertion mutagenesis and generation of transcriptional fusions. Gene 1995, 162, 37–39. [Google Scholar] [CrossRef]
- Schäfer, A.; Tauch, A.; Jager, W.; Kalinowski, J.; Thierbach, G.; Pühler, A. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: Selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 1994, 145, 69–73. [Google Scholar] [CrossRef]
- Schwyn, B.; Neilands, J.B. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 1987, 160, 47–56. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Hidalgo, A.; Margaret, I.; Crespo-Rivas, J.C.; Parada, M.; Murdoch Pdel, S.; López, A.; Buendía-Clavería, A.M.; Moreno, J.; Albareda, M.; Gil-Serrano, A.M.; et al. The rkpU gene of Sinorhizobium fredii HH103 is required for bacterial K-antigen polysaccharide production and for efficient nodulation with soybean but not with cowpea. Microbiology 2010, 156, 3398–3411. [Google Scholar] [CrossRef] [PubMed]
- Buendía-Clavería, A.M.; Moussaid, A.; Ollero, F.J.; Vinardell, J.M.; Torres, A.; Moreno, J.; Gil-Serrano, A.M.; Rodríguez-Carvajal, M.A.; Tejero-Mateo, P.; Peart, J.L.; et al. A purL mutant of Sinorhizobium fredii HH103 is symbiotically defective and altered in its lipopolysaccharide. Microbiology 2003, 149, 1807–1818. [Google Scholar] [CrossRef] [PubMed]
- Buendía-Clavería, A.M.; Chamber, M.; Ruiz-Sainz, J.E. A comparative study of the physiological characteristics, plasmid content and symbiotic properties of different Rhizobium fredii strains. Syst. Appl. Microbiol. 1989, 12, 203–209. [Google Scholar] [CrossRef]


| Glycine max cv.Williams 1 | |||||
|---|---|---|---|---|---|
| Inoculant | Number of Nodules | Nodules Fresh Weight (mg) | Plant-Top Dry Weight (mg) | ARA 2 (Nmoles Acetylene/Plant/Hour) | Nodules Size (mm) |
| SVQ269 | 26.38 ± 2.88 | 510.33 ± 65.71 | 1.08 ± 0.15 | 998.66 ± 175.56 | 4.04 ± 0.11 |
| SVQ780 | 23.80 ± 3.38 | 222.71 ± 16.04 ** | 0.62 ± 0.07 * | 167.99 ± 27.64 ** | 2.64 ± 0.10 ** |
| SVQ780C | 22.44 ± 2.50 | 453.48 ± 54.89 | 1.18 ± 0.12 | 1008.22 ± 118.18 | 4.37 ± 0.10 ** |
| NI 3 | 0 | 0 | 0.245 ± 0.02 ** | 0 | 0 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Crespo-Rivas, J.C.; Navarro-Gómez, P.; Alias-Villegas, C.; Shi, J.; Zhen, T.; Niu, Y.; Cuéllar, V.; Moreno, J.; Cubo, T.; Vinardell, J.M.; et al. Sinorhizobium fredii HH103 RirA Is Required for Oxidative Stress Resistance and Efficient Symbiosis with Soybean. Int. J. Mol. Sci. 2019, 20, 787. https://doi.org/10.3390/ijms20030787
Crespo-Rivas JC, Navarro-Gómez P, Alias-Villegas C, Shi J, Zhen T, Niu Y, Cuéllar V, Moreno J, Cubo T, Vinardell JM, et al. Sinorhizobium fredii HH103 RirA Is Required for Oxidative Stress Resistance and Efficient Symbiosis with Soybean. International Journal of Molecular Sciences. 2019; 20(3):787. https://doi.org/10.3390/ijms20030787
Chicago/Turabian StyleCrespo-Rivas, Juan Carlos, Pilar Navarro-Gómez, Cynthia Alias-Villegas, Jie Shi, Tao Zhen, Yanbo Niu, Virginia Cuéllar, Javier Moreno, Teresa Cubo, José María Vinardell, and et al. 2019. "Sinorhizobium fredii HH103 RirA Is Required for Oxidative Stress Resistance and Efficient Symbiosis with Soybean" International Journal of Molecular Sciences 20, no. 3: 787. https://doi.org/10.3390/ijms20030787
APA StyleCrespo-Rivas, J. C., Navarro-Gómez, P., Alias-Villegas, C., Shi, J., Zhen, T., Niu, Y., Cuéllar, V., Moreno, J., Cubo, T., Vinardell, J. M., Ruiz-Sainz, J. E., Acosta-Jurado, S., & Soto, M. J. (2019). Sinorhizobium fredii HH103 RirA Is Required for Oxidative Stress Resistance and Efficient Symbiosis with Soybean. International Journal of Molecular Sciences, 20(3), 787. https://doi.org/10.3390/ijms20030787








