The Ubiquitin Proteasome System in Ischemic and Dilated Cardiomyopathy
Abstract
1. Introduction
2. Results
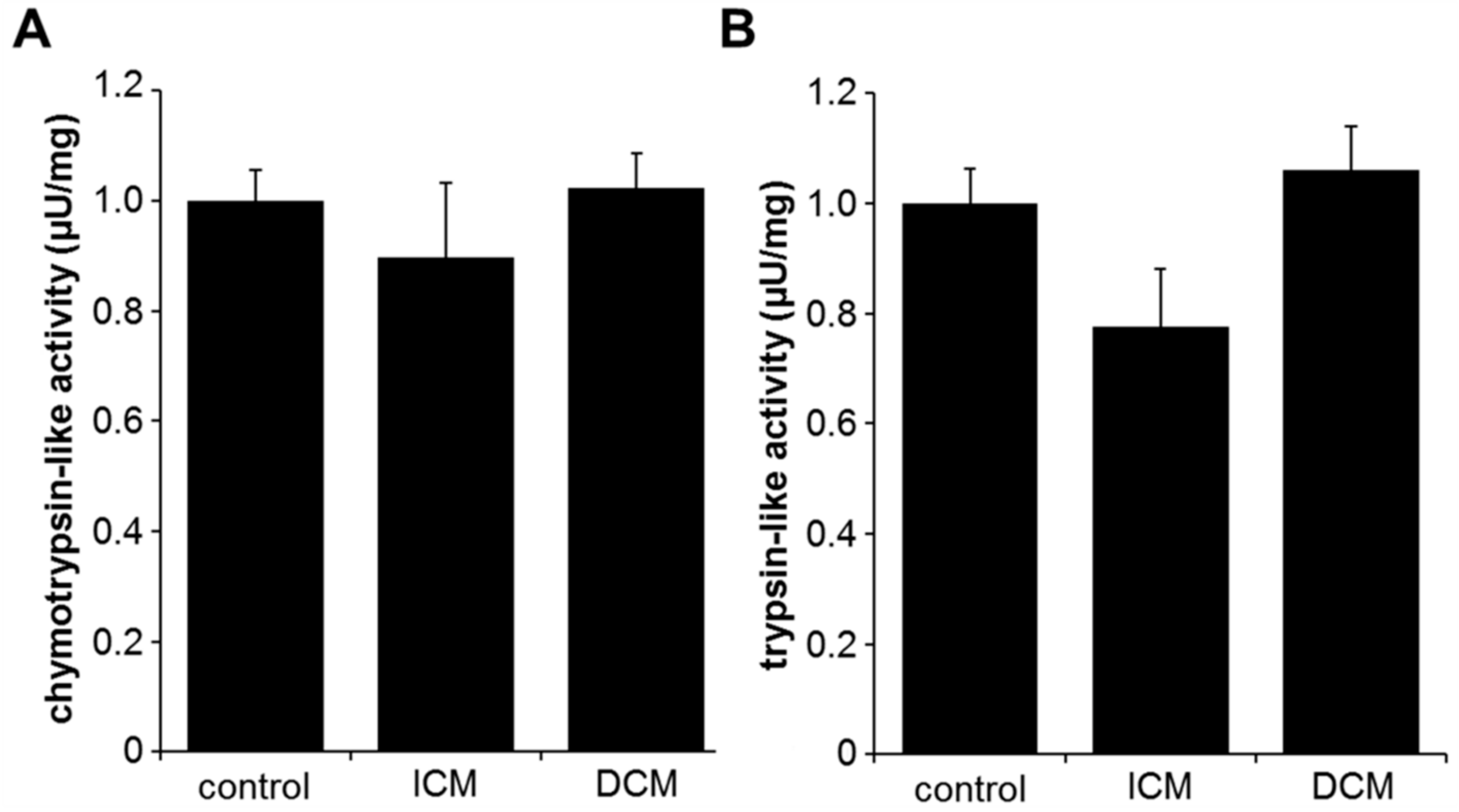
2.1. Proteasome Activity
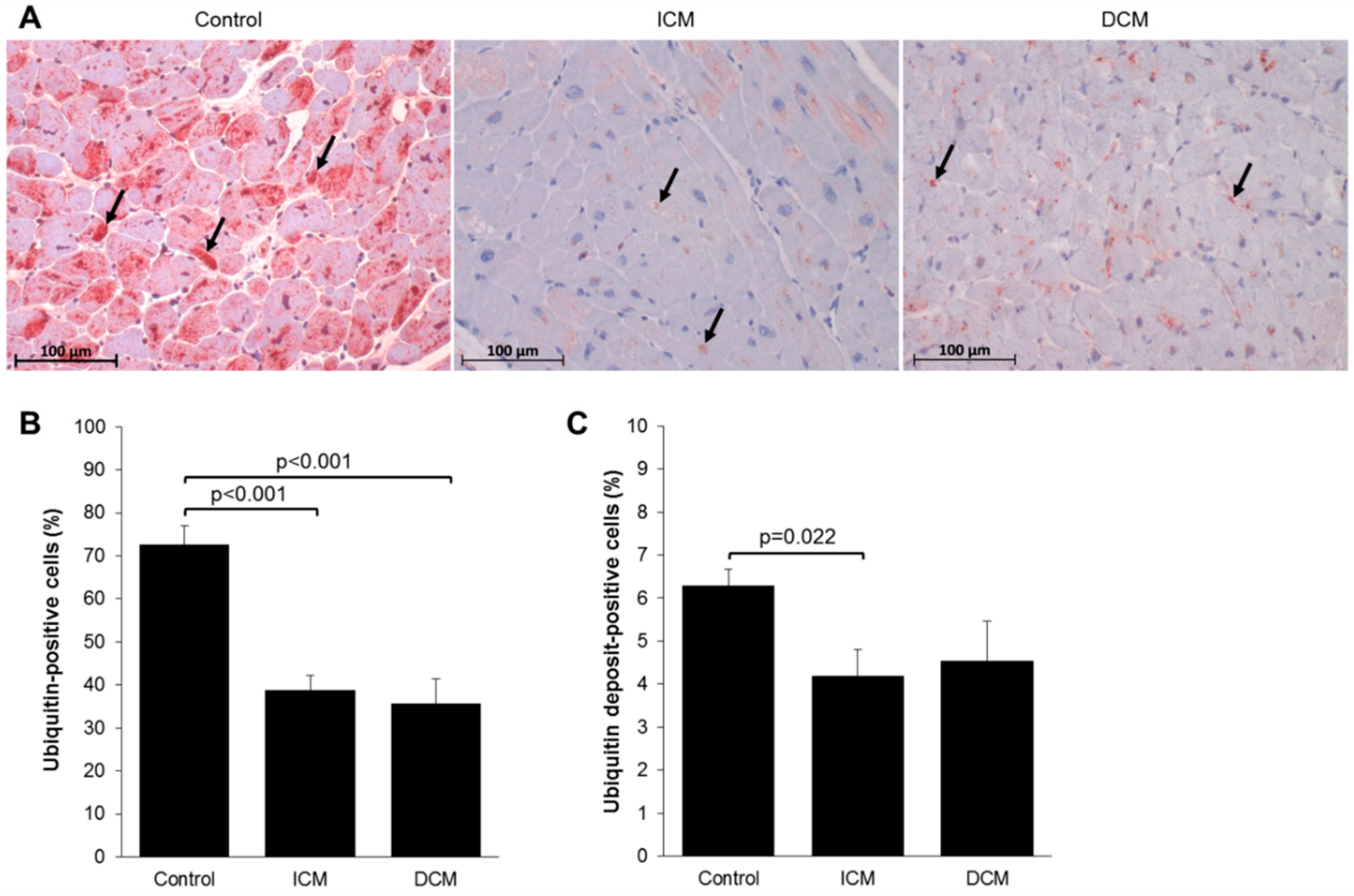
2.2. Myocardial Protein Ubiquitination
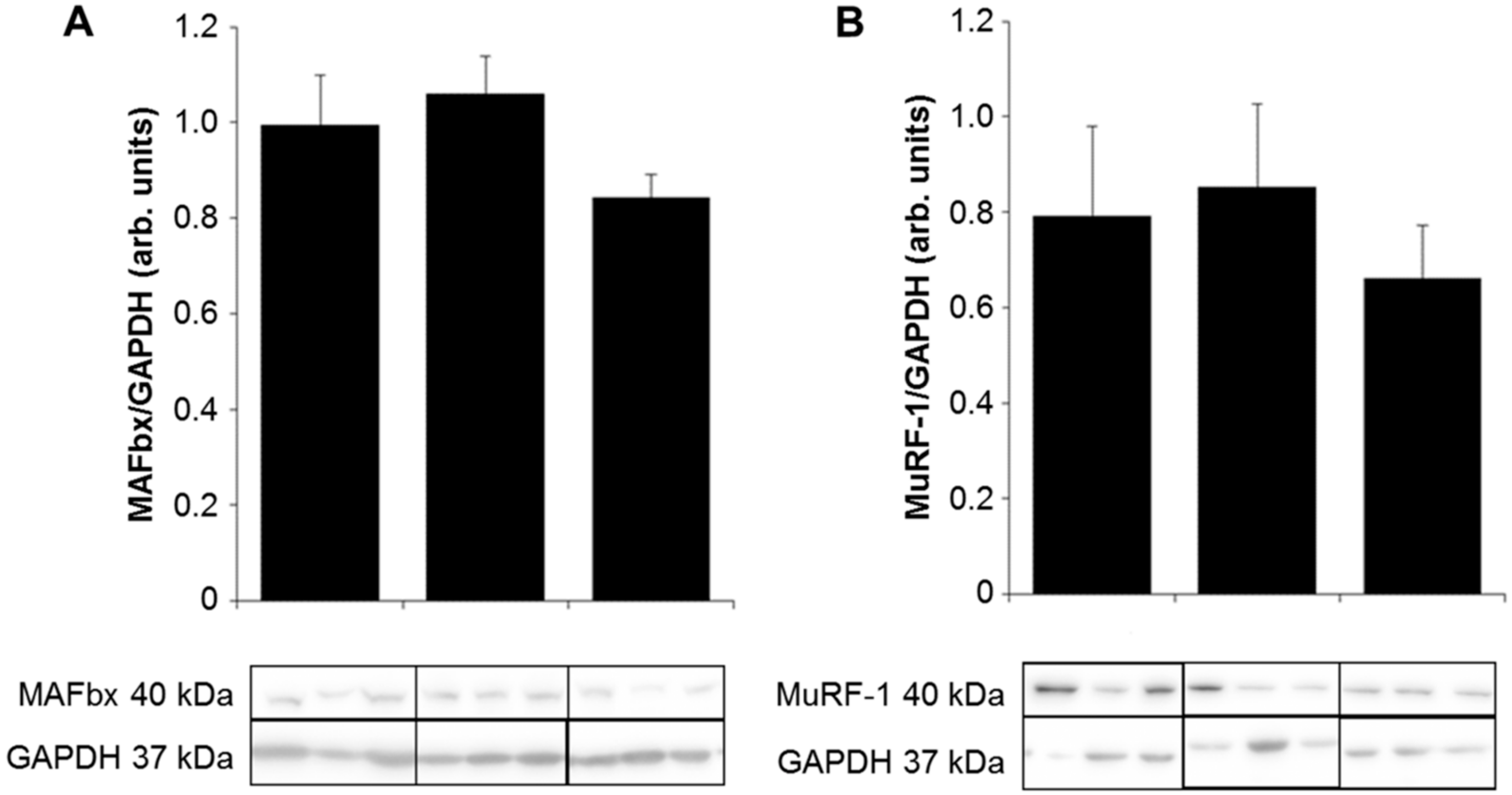
2.3. Expression of E3 Ligases
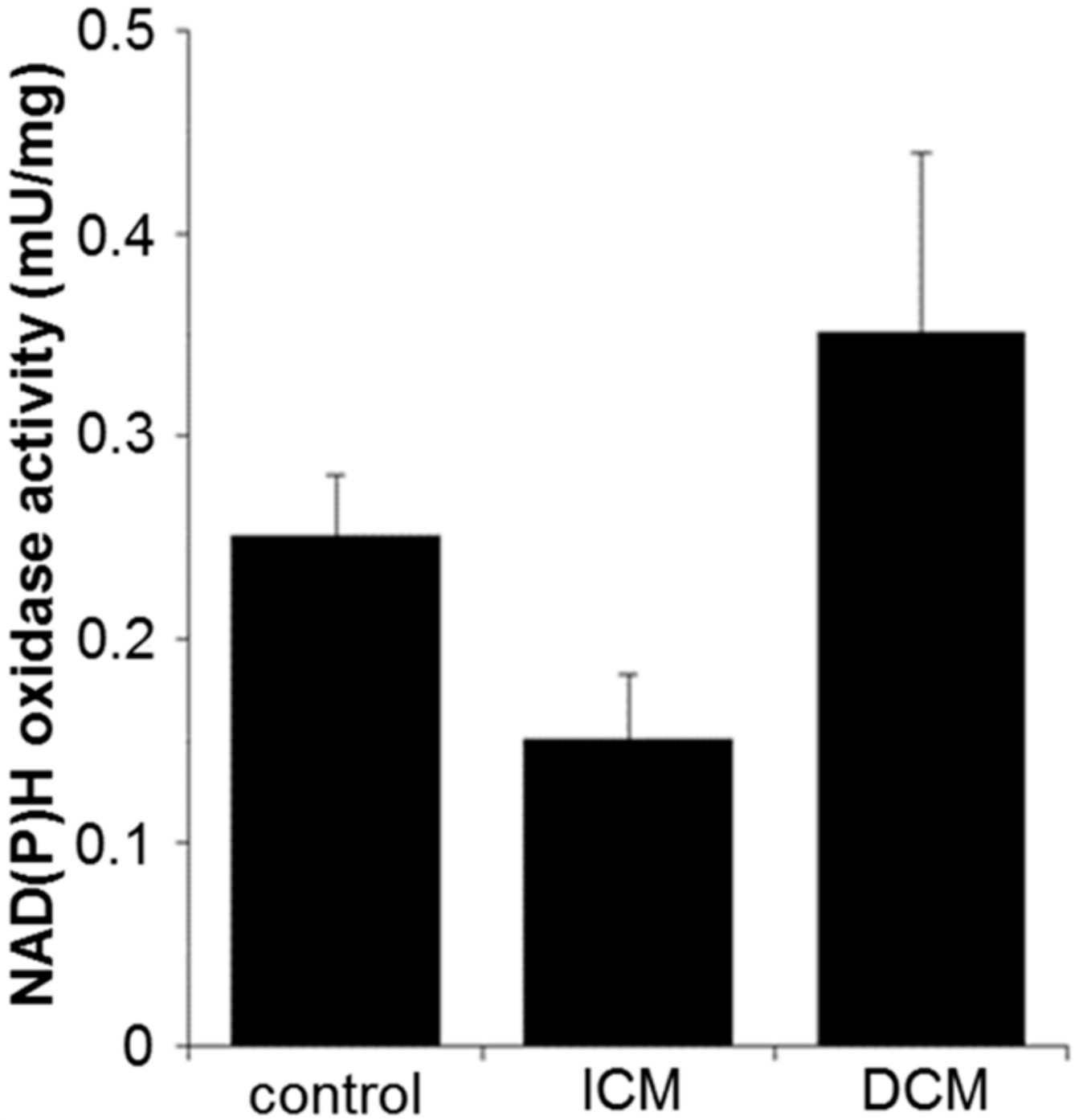
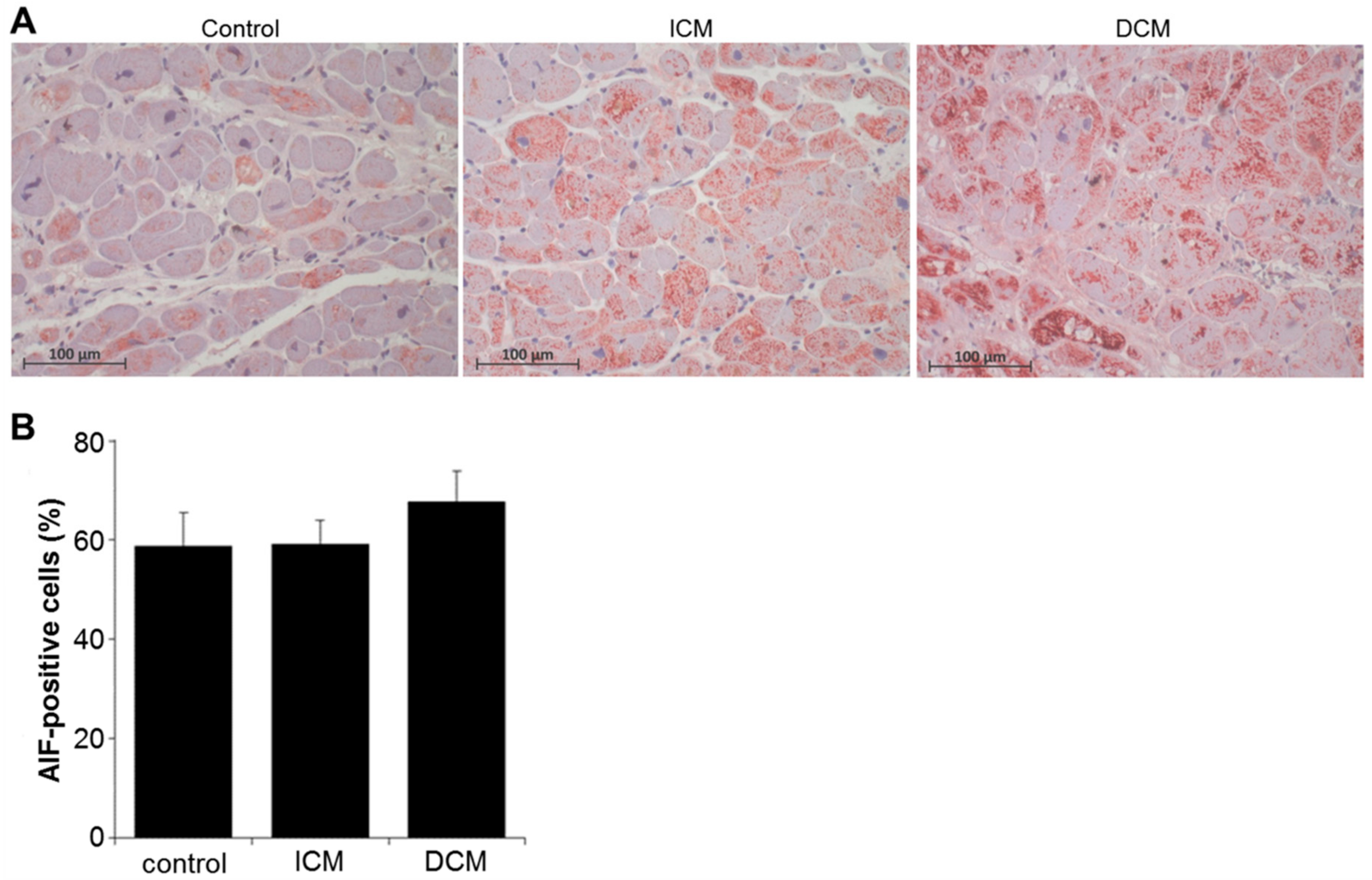
2.4. Oxidative Stress and Apoptosis
3. Discussion
Limitations
4. Materials and Methods
4.1. Sample Collection and Preparation
4.2. Quantification of Protein Expression
4.3. Quantification of Proteasome Activity
4.4. Quantification of Activity of A Reactive Oxygen Species-Producing Enzyme
4.5. Immunohistological Analysis
4.6. Statistical Analysis
Author Contributions
Funding
Conflicts of Interest
References
- Drews, O.; Taegtmeyer, H. Targeting the Ubiquitin-Proteasome System in Heart Disease: The Basis for New Therapeutic Strategies. Antioxid. Redox Signal. 2014, 21, 2322–2343. [Google Scholar] [CrossRef] [PubMed]
- Schaun, M.I.; Eibel, B.; Kristocheck, M.; Sausen, G.; Machado, L.; Koche, A.; Markoski, M.M. Cell Therapy in Ischemic Heart Disease: Interventions That Modulate Cardiac Regeneration. Stem Cells Int. 2016, 2016, 2171035. [Google Scholar] [CrossRef] [PubMed]
- Cave, A.; Grieve, D.; Johar, S.; Zhang, M.; Shah, A.M. NADPH oxidase-derived reactive oxygen species in cardiac pathophysiology. Philos. Trans. R. Soc. B Biol. Sci. 2005, 360, 2327–2334. [Google Scholar] [CrossRef] [PubMed]
- Bulteau, A.-L.; Lundberg, K.C.; Humphries, K.M.; Sadek, H.A.; Szweda, P.A.; Friguet, B.; Szweda, L.I. Oxidative Modification and Inactivation of the Proteasome during Coronary Occlusion/Reperfusion. J. Biol. Chem. 2001, 276, 30057–30063. [Google Scholar] [CrossRef]
- Candell-Riera, J.; Romero-Farina, G.; Aguadé-Bruix, S.; Castell-Conesa, J. Ischemic cardiomyopathy: A clinical nuclear cardiology perspective. Rev. Española Cardiol. 2009, 62, 903–917. [Google Scholar] [CrossRef]
- Jameson, J.L.; Kasper, D.L.; Longo, D.L.; Dan, L.; Fauci, A.S.; Hauser, S.L.; Loscalzo, J. Harrison’s Principles of Internal Medicine, 20th ed.; Mcgraw Education Hill: New York City, NY, USA, 2018; Available online: https://accessmedicine.mhmedical.com/book.aspx?bookID=2129 (accessed on 12 December 2019).
- Taylor, M.R.G.; Carniel, E.; Mestroni, L. Cardiomyopathy, familial dilated. Orphanet J. Rare Dis. 2006, 1, 27. [Google Scholar] [CrossRef]
- Maron, B.J.; Towbin, J.A.; Thiene, G.; Antzelevitch, C.; Corrado, D.; Arnett, D.; Moss, A.J.; Seidman, C.E.; Young, J.B. Contemporary Definitions and Classification of the Cardiomyopathies. Circulation 2006, 113, 1807–1816. [Google Scholar] [CrossRef]
- Pagan, J.; Seto, T.; Pagano, M.; Cittadini, A. Role of the ubiquitin proteasome system in the heart. Circ. Res. 2013, 112, 1046–1058. [Google Scholar] [CrossRef]
- Calise, J.; Powell, S.R. The ubiquitin proteasome system and myocardial ischemia. Am. J. Physiol. Heart Circ. Physiol. 2013, 304, H337–H349. [Google Scholar] [CrossRef]
- Groll, M.; Huber, R. Substrate access and processing by the 20S proteasome core particle. Int. J. Biochem. Cell Biol. 2003, 35, 606–616. [Google Scholar] [CrossRef]
- Haglund, K.; Dikic, I. Ubiquitylation and cell signaling. EMBO J. 2005, 24, 3353–3359. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Kang, J.; Zhang, L.; Liang, Z.; Tang, X.; Yan, Y.; Qian, H.; Zhang, X.; Xu, W.; Mao, F. Ubiquitination regulation of inflammatory responses through NF-κB pathway. Am. J. Transl. Res. 2018, 10, 881–891. [Google Scholar] [PubMed]
- Stewart, M.D.; Ritterhoff, T.; Klevit, R.E.; Brzovic, P.S. E2 enzymes: More than just middle men. Cell Res. 2016, 26, 423–440. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, S. Structural mechanisms of HECT-type ubiquitin ligases. Biol. Chem. 2018, 399, 127–145. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Ye, Y. Polyubiquitin chains: Functions, structures, and mechanisms. Cell. Mol. Life Sci. 2008, 65, 2397–2406. [Google Scholar] [CrossRef]
- Sadowski, M.; Sarcevic, B. Mechanisms of mono- and poly-ubiquitination: Ubiquitination specificity depends on compatibility between the E2 catalytic core and amino acid residues proximal to the lysine. Cell Div. 2010, 5, 19. [Google Scholar] [CrossRef]
- Grice, G.L.; Nathan, J.A. The recognition of ubiquitinated proteins by the proteasome. Cell. Mol. Life Sci. 2016, 73, 3497–3506. [Google Scholar] [CrossRef]
- Shaid, S.; Brandts, C.H.; Serve, H.; Dikic, I. Ubiquitination and selective autophagy. Cell Death Differ. 2013, 20, 21–30. [Google Scholar] [CrossRef]
- Lilienbaum, A. Relationship between the proteasomal system and autophagy. Int. J. Biochem. Mol. Biol. 2013, 4, 1. [Google Scholar]
- Ji, C.H.; Kwon, Y.T. Crosstalk and Interplay between the Ubiquitin-Proteasome System and Autophagy. Mol. Cells 2017, 40, 441. [Google Scholar]
- Dwane, L.; Gallagher, W.M.; Chonghaile, T.N.; O’Connor, D.P. The Emerging Role of Non-traditional Ubiquitination in Oncogenic Pathways. J. Biol. Chem. 2017, 292, 3543. [Google Scholar] [CrossRef] [PubMed]
- Wei, R.; Liu, X.; Yu, W.; Yang, T.; Cai, W.; Liu, J.; Huang, X.; Xu, G.; Zhao, S.; Yang, J.; et al. Deubiquitinases in cancer. Oncotarget 2015, 6, 12872–12889. [Google Scholar] [CrossRef] [PubMed]
- Predmore, J.M.; Wang, P.; Davis, F.; Bartolone, S.; Westfall, M.V.; Dyke, D.B.; Pagani, F.; Powell, S.R.; Day, S.M. Ubiquitin Proteasome Dysfunction in Human Hypertrophic and Dilated Cardiomyopathies. Circulation 2010, 121, 997–1004. [Google Scholar] [CrossRef] [PubMed]
- Zolk, O.; Schenke, C.; Sarikas, A. The ubiquitin-proteasome system: Focus on the heart. Cardiovasc. Res. 2006, 70, 410–421. [Google Scholar] [CrossRef]
- Sitte, N.; Huber, M.; Grune, T.; Ladhoff, A.; Doecke, W.D.; Von Zglinicki, T.; Davies, K.J. Proteasome inhibition by lipofuscin/ceroid during postmitotic aging of fibroblasts. FASEB J. 2000, 14, 1490–1498. [Google Scholar] [CrossRef]
- Bence, N.F.; Sampat, R.M.; Kopito, R.R. Impairment of the Ubiquitin-Proteasome System by Protein Aggregation. Science 2001, 292, 1552–1555. [Google Scholar] [CrossRef]
- Kostin, S.; Pool, L.; Elsasser, A.; Hein, S.; Drexler, H.C.A.; Arnon, E.; Hayakawa, Y.; Zimmermann, R.; Bauer, E.; Klövekorn, W.-P.; et al. Myocytes Die by Multiple Mechanisms in Failing Human Hearts. Circ. Res. 2003, 92, 715–724. [Google Scholar] [CrossRef]
- Li, H.H.; Kedar, V.; Zhang, C.; McDonough, H.; Arya, R.; Wang, D.-Z.; Patterson, C. Atrogin-1/muscle atrophy F-box inhibits calcineurin-dependent cardiac hypertrophy by participating in an SCF ubiquitin ligase complex. J. Clin. Investig. 2004, 114, 1058–1071. [Google Scholar] [CrossRef]
- Stitt, T.N.; Drujan, D.; Clarke, B.A.; Panaro, F.; Timofeyva, Y.; Kline, W.O.; Gonzalez, M.; Yancopoulos, G.D.; Glass, D.J. The IGF-1/PI3K/Akt pathway prevents expression of muscle atrophy-induced ubiquitin ligases by inhibiting FOXO transcription factors. Mol. Cell 2004, 14, 395–403. [Google Scholar] [CrossRef]
- Sandri, M.; Sandri, C.; Gilbert, A.; Skurk, C.; Calabria, E.; Picard, A.; Walsh, K.; Schiaffino, S.; Lecker, S.H.; Goldberg, A.L. Foxo transcription factors induce the atrophy-related ubiquitin ligase atrogin-1 and cause skeletal muscle atrophy. Cell 2004, 117, 399–412. [Google Scholar] [CrossRef]
- Schisler, J.C.; Willis, M.S.; Patterson, C. You spin me round: MafBx/Atrogin-1 feeds forward on FOXO transcription factors (like a record). Cell Cycle 2008, 7, 440–443. [Google Scholar] [CrossRef] [PubMed]
- Conraads, V.M.; Vrints, C.J.; Rodrigus, I.E.; Hoymans, V.Y.; Van Craenenbroeck, E.M.; Bosmans, J.; Claeys, M.J.; Van Herck, P.; Linke, A.; Schuler, G.; et al. Depressed expression of MuRF1 and MAFbx in areas remote of recent myocardial infarction: A mechanism contributing to myocardial remodeling? Basic Res. Cardiol. 2010, 105, 219–226. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanian, S.; Mani, S.; Shiraishi, H.; Johnston, R.K.; Yamane, K.; Willey, C.D.; Cooper, G., IV; Tuxworth, W.J.; Kuppuswamy, D. Enhanced ubiquitination of cytoskeletal proteins in pressure overloaded myocardium is accompanied by changes in specific E3 ligases. J. Mol. Cell. Cardiol. 2006, 41, 669–679. [Google Scholar] [CrossRef] [PubMed]
- Willis, M.S.; Rojas, M.; Li, L.; Selzman, C.H.; Tang, R.-H.; Stansfield, W.E.; Rodriguez, J.E.; Glass, D.J.; Patterson, C. Muscle ring finger 1 mediates cardiac atrophy in vivo. Am. J. Physiol. Circ. Physiol. 2009, 296, H997–H1006. [Google Scholar] [CrossRef] [PubMed]
- Day, S.M. The ubiquitin proteasome system in human cardiomyopathies and heart failure. Am. J. Physiol. Heart Circ. Physiol. 2013, 304, H1283–H1293. [Google Scholar] [CrossRef] [PubMed]
- Orlowski, R.Z. The role of the ubiquitin-proteasome pathway in apoptosis. Cell Death Differ. 1999, 6, 303–313. [Google Scholar] [CrossRef]
- Tsukamoto, O.; Minamino, T.; Okada, K.; Shintani, Y.; Takashima, S.; Kato, H.; Liao, Y.; Okazaki, H.; Asai, M.; Hirata, A.; et al. Depression of proteasome activities during the progression of cardiac dysfunction in pressure-overloaded heart of mice. Biochem. Biophys. Res. Commun. 2006, 340, 1125–1133. [Google Scholar] [CrossRef]
- Di Meo, S.; Reed, T.T.; Venditti, P.; Victor, V.M. Role of ROS and RNS Sources in Physiological and Pathological Conditions. Oxidative Med. Cell. Longev. 2016, 2016, 1245049. [Google Scholar] [CrossRef]
- Bowen, T.S.; Adams, V.; Werner, S.; Fischer, T.; Vinke, P.; Brogger, M.N.; Mangner, N.; Linke, A.; Sehr, P.; Lewis, J.; et al. Small-molecule inhibition of MuRF1 attenuates skeletal muscle atrophy and dysfunction in cardiac cachexia. J. Cachexia Sarcopenia Muscle 2017, 8, 939–953. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Spänig, S.; Kellermann, K.; Dieterlen, M.-T.; Noack, T.; Lehmann, S.; Borger, M.A.; Garbade, J.; Barac, Y.D.; Emrich, F. The Ubiquitin Proteasome System in Ischemic and Dilated Cardiomyopathy. Int. J. Mol. Sci. 2019, 20, 6354. https://doi.org/10.3390/ijms20246354
Spänig S, Kellermann K, Dieterlen M-T, Noack T, Lehmann S, Borger MA, Garbade J, Barac YD, Emrich F. The Ubiquitin Proteasome System in Ischemic and Dilated Cardiomyopathy. International Journal of Molecular Sciences. 2019; 20(24):6354. https://doi.org/10.3390/ijms20246354
Chicago/Turabian StyleSpänig, Sabine, Kristina Kellermann, Maja-Theresa Dieterlen, Thilo Noack, Sven Lehmann, Michael A. Borger, Jens Garbade, Yaron D. Barac, and Fabian Emrich. 2019. "The Ubiquitin Proteasome System in Ischemic and Dilated Cardiomyopathy" International Journal of Molecular Sciences 20, no. 24: 6354. https://doi.org/10.3390/ijms20246354
APA StyleSpänig, S., Kellermann, K., Dieterlen, M.-T., Noack, T., Lehmann, S., Borger, M. A., Garbade, J., Barac, Y. D., & Emrich, F. (2019). The Ubiquitin Proteasome System in Ischemic and Dilated Cardiomyopathy. International Journal of Molecular Sciences, 20(24), 6354. https://doi.org/10.3390/ijms20246354




