Usefulness of Mesenchymal Cell Lines for Bone and Cartilage Regeneration Research
Abstract
1. Introduction
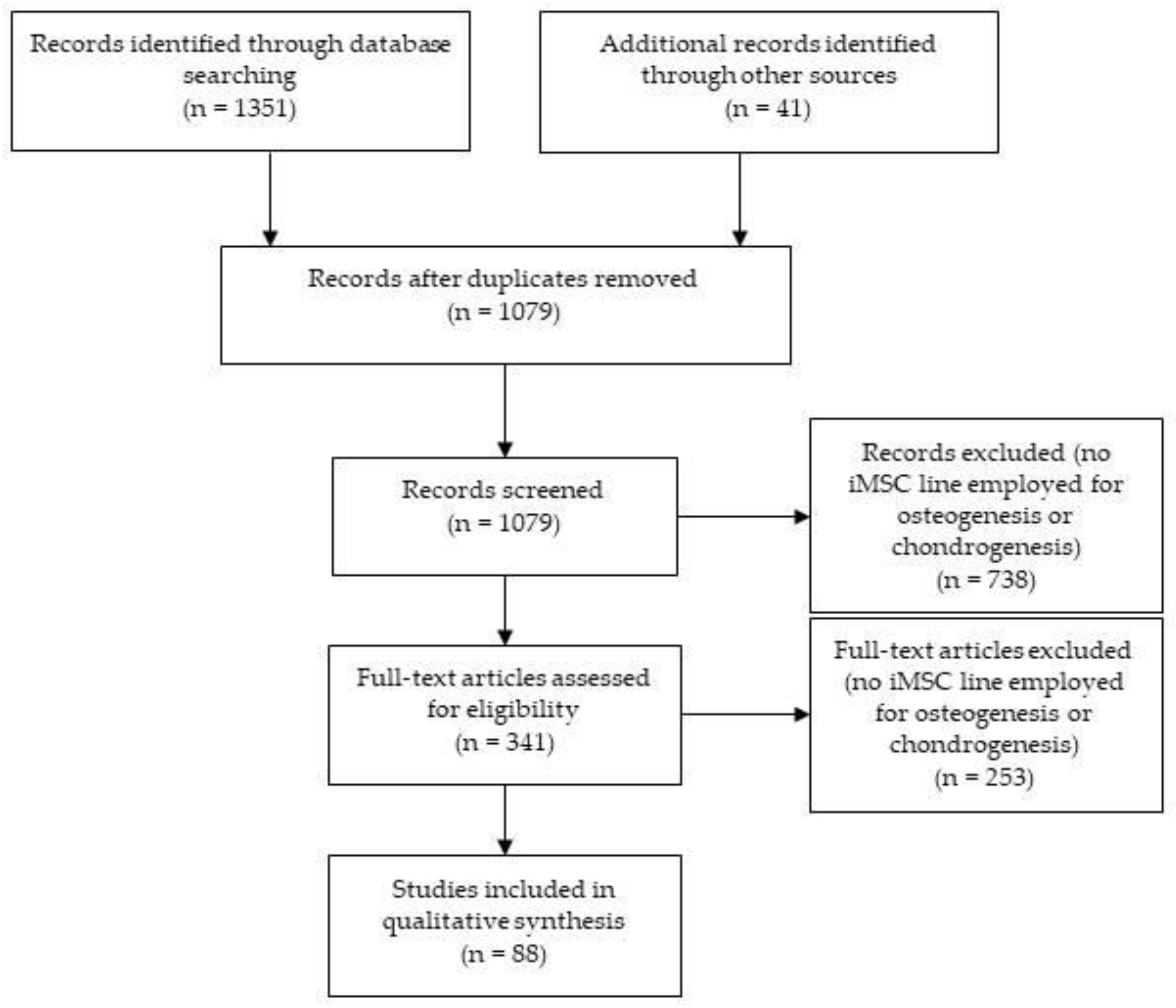
2. Methodology
3. Immortal Mesenchymal Stromal Cell (iMSC) Lines
3.1. Immortalizing Human Adult MSCs
3.2. Multidifferentiation Potential of iMSCs
3.2.1. Osteogenic Potential
3.2.2. Chondrogenic Potential
3.2.3. Adipogenic Potential
3.3. Surface Markers Expression of iMSCs
3.4. Clonality, Selection and Validation
3.5. Tumorigenicity
3.6. In Vivo Bone Formation Capacity
4. Conclusions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| MSC | Mesenchymal stromal cells |
| PDs | Population doublings |
| iMSC | Immortal mesenchymal stromal cells |
| hTERT | Human telomerase reverse transcriptase |
| SV40LT | Simian virus 40 large T antigen |
| HPV | Human papillomavirus |
| ARS | Alizarin Red Staining |
| VKS | Von Kossa Staining |
| APS | Alkaline Phosphatase Staining |
| ABS | Alcian Blue Staining |
| TBS | Toluidine Blue Staining |
| OROS PSR | Oil Red O Staining Picro-Sirius Red |
| STR | Short Tandem Repeat |
| IDM | Immunodeficient Mice |
References
- Grayson, W.L.; Bunnell, B.A.; Martin, E.; Frazier, T.; Hung, B.P.; Gimble, J.M. Stromal cells and stem cells in clinical bone regeneration. Nat. Rev. Endocrinol. 2015, 11, 140–150. [Google Scholar] [CrossRef] [PubMed]
- Tamaddon, M.; Wang, L.; Liu, Z.; Liu, C. Osteochondral tissue repair in osteoarthritic joints: Clinical challenges and opportunities in tissue engineering. Bio Des. Manuf. 2018, 1, 101–114. [Google Scholar] [CrossRef] [PubMed]
- Roseti, L.; Parisi, V.; Petretta, M.; Cavallo, C.; Desando, G.; Bartolotti, I.; Grigolo, B. Scaffolds for Bone Tissue Engineering: State of the art and new perspectives. Mater. Sci. Eng. C 2017, 78, 1246–1262. [Google Scholar] [CrossRef] [PubMed]
- Piñeiro-Ramil, M.; Castro-Viñuelas, R.; Sanjurjo-Rodríguez, C.; Hermida-Gómez, T.; Fuentes-Boquete, I.; Toro-Santos, F.J.; de Blanco-García, F.J.; Díaz-Prado, S.M. Cell Therapy and Tissue Engineering for Cartilage Repair. In Cartilage Repair and Regeneration; InTech: London, UK, 2018. [Google Scholar]
- Deng, C.; Zhu, H.; Li, J.; Feng, C.; Yao, Q.; Wang, L.; Chang, J.; Wu, C. Bioactive Scaffolds for Regeneration of Cartilage and Subchondral Bone Interface. Theranostics 2018, 8, 1940–1955. [Google Scholar] [CrossRef] [PubMed]
- Ghassemi, T.; Shahroodi, A.; Ebrahimzadeh, M.H.; Mousavian, A.; Movaffagh, J.; Moradi, A. Current Concepts in Scaffolding for Bone Tissue Engineering. Arch. Bone Jt. Surg. 2018, 6, 90–99. [Google Scholar] [PubMed]
- Samsonraj, R.M.; Raghunath, M.; Nurcombe, V.; Hui, J.H.; van Wijnen, A.J.; Cool, S.M. Concise Review: Multifaceted Characterization of Human Mesenchymal Stem Cells for Use in Regenerative Medicine. Stem Cells Transl. Med. 2017, 6, 2173–2185. [Google Scholar] [CrossRef] [PubMed]
- Szychlinska, M.A.; Stoddart, M.J.; D’Amora, U.; Ambrosio, L.; Alini, M.; Musumeci, G. Mesenchymal Stem Cell-Based Cartilage Regeneration Approach and Cell Senescence: Can We Manipulate Cell Aging and Function? Tissue Eng. Part B Rev. 2017, 23, 529–539. [Google Scholar] [CrossRef]
- Kouroupis, D.; Sanjurjo-Rodriguez, C.; Jones, E.; Correa, D. MSC functionalization for enhanced therapeutic applications. Tissue Eng. Part B Rev. 2018. [Google Scholar] [CrossRef]
- Richardson, S.M.; Kalamegam, G.; Pushparaj, P.N.; Matta, C.; Memic, A.; Khademhosseini, A.; Mobasheri, R.; Poletti, F.L.; Hoyland, J.A.; Mobasheri, A. Mesenchymal stem cells in regenerative medicine: Focus on articular cartilage and intervertebral disc regeneration. Methods 2016, 99, 69–80. [Google Scholar] [CrossRef]
- Yong, K.W.; Choi, J.R.; Dolbashid, A.S.; Wan Safwani, W.K.Z. Biosafety and bioefficacy assessment of human mesenchymal stem cells: What do we know so far? Regen Med. 2018, 13, 219–232. [Google Scholar] [CrossRef]
- Herberts, C.A.; Kwa, M.S.; Hermsen, H.P. Risk factors in the development of stem cell therapy. J. Transl. Med. 2011, 9, 29. [Google Scholar] [CrossRef] [PubMed]
- Koch, C.M.; Reck, K.; Shao, K.; Lin, Q.; Joussen, S.; Ziegler, P.; Walenda, G.; Drescher, W.; Opalka, B.; May, T.; et al. Pluripotent stem cells escape from senescence-associated DNA methylation changes. Genome Res. 2013, 23, 248–259. [Google Scholar] [CrossRef] [PubMed]
- Böcker, W.; Yin, Z.; Drosse, I.; Haasters, F.; Rossmann, O.; Wierer, M.; Popov, C.; Locher, M.; Mutschler, W.; Docheva, D.; et al. Introducing a single-cell-derived human mesenchymal stem cell line expressing hTERT after lentiviral gene transfer. J. Cell. Mol. Med. 2008, 12, 1347–1359. [Google Scholar] [CrossRef]
- Bourgine, P.; Le Magnen, C.; Pigeot, S.; Geurts, J.; Scherberich, A.; Martin, I. Combination of immortalization and inducible death strategies to generate a human mesenchymal stromal cell line with controlled survival. Stem Cell Res. 2014, 12, 584–598. [Google Scholar] [CrossRef]
- Baker, N.; Boyette, L.B.; Tuan, R.S. Characterization of bone marrow-derived mesenchymal stem cells in aging. Bone 2015, 70, 37–47. [Google Scholar] [CrossRef]
- Stölzel, K.; Schulze-Tanzil, G.; Olze, H.; Schwarz, S.; Feldmann, E.M.; Rotter, N. Immortalised human mesenchymal stem cells undergo chondrogenic differentiation in alginate and PGA/PLLA scaffolds. Cell Tissue Bank. 2015, 16, 159–170. [Google Scholar] [CrossRef]
- Okamoto, T.; Aoyama, T.; Nakayama, T.; Nakamata, T.; Hosaka, T.; Nishijo, K.; Nakamura, T.; Kiyono, T.; Toguchida, J. Clonal heterogeneity in differentiation potential of immortalized human mesenchymal stem cells. Biochem. Biophys. Res. Commun. 2002, 295, 354–361. [Google Scholar] [CrossRef]
- James, S.; Fox, J.; Afsari, F.; Lee, J.; Clough, S.; Knight, C.; Ashmore, J.; Ashton, P.; Preham, O.; Hoogduijn, M.; et al. Multiparameter Analysis of Human Bone Marrow Stromal Cells Identifies Distinct Immunomodulatory and Differentiation-Competent Subtypes. Stem Cell Rep. 2015, 4, 1004–1015. [Google Scholar] [CrossRef]
- Ahlfeld, T.; Cidonio, G.; Kilian, D.; Duin, S.; Akkineni, A.R.; Dawson, J.I.; Yang, S.; Lode, A.; Oreffo, R.O.C.; Gelinsky, M. Development of a clay based bioink for 3D cell printing for skeletal application. Biofabrication 2017, 9. [Google Scholar] [CrossRef]
- Akmammedov, R.; Huysal, M.; Isik, S.; Senel, M. Preparation and characterization of novel chitosan/zeolite scaffolds for bone tissue engineering applications. Int. J. Polym. Mater. Polym. Biomater. 2018, 67, 110–118. [Google Scholar] [CrossRef]
- Nürnberger, S.; Schneider, C.; van Osch, G.V.M.; Keibl, C.; Rieder, B.; Monforte, X.; Teuschl, A.H.; Mühleder, S.; Holnthoner, W.; Schädl, B.; et al. Repopulation of an auricular cartilage scaffold, AuriScaff, perforated with an enzyme combination. Acta Biomater. 2019, 86, 207–222. [Google Scholar] [CrossRef] [PubMed]
- Hossain, K.M.Z.; Patel, U.; Kennedy, A.R.; Macri-Pellizzeri, L.; Sottile, V.; Grant, D.M.; Scammell, B.E.; Ahmed, I. Porous calcium phosphate glass microspheres for orthobiologic applications. Acta Biomater. 2018, 72, 396–406. [Google Scholar] [CrossRef] [PubMed]
- Prasopthum, A.; Shakesheff, K.M.; Yang, J. Direct three-dimensional printing of polymeric scaffolds with nanofibrous topography. Biofabrication 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Tendulkar, G.; Sreekumar, V.; Rupp, F.; Teotia, A.K.; Athanasopulu, K.; Kemkemer, R.; Buck, A.; Buck, A.; Kaps, H.-P.; Geis-Gerstorfer, J.; et al. Characterisation of porous knitted titanium for replacement of intervertebral disc nucleus pulposus. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Bourgine, P.E.; Gaudiello, E.; Pippenger, B.; Jaquiery, C.; Klein, T.; Pigeot, S.; Todorov, A.; Feliciano, S.; Banfi, A.; Martin, I. Engineered Extracellular Matrices as Biomaterials of Tunable Composition and Function. Adv. Funct. Mater. 2017, 27. [Google Scholar] [CrossRef]
- da Conceicao Ribeiro, R.; Pal, D.; Ferreira, A.M.; Gentile, P.; Benning, M.; Dalgarno, K. Reactive jet impingement bioprinting of high cell density gels for bone microtissue fabrication. Biofabrication 2018, 11. [Google Scholar] [CrossRef]
- Aida, Y.; Kurihara, H.; Kato, K. Wnt3a promotes differentiation of human bone marrow-derived mesenchymal stem cells into cementoblast-like cells. Vitr. Cell. Dev. Biol. Anim. 2018, 54, 468–476. [Google Scholar] [CrossRef]
- Cesarz, Z.; Funnell, J.L.; Guan, J.; Tamama, K. Soft Elasticity-Associated Signaling and Bone Morphogenic Protein 2 Are Key Regulators of Mesenchymal Stem Cell Spheroidal Aggregates. Stem Cells Dev. 2016, 25, 622–635. [Google Scholar] [CrossRef]
- Elsafadi, M.; Manikandan, M.; Alajez, N.M.; Hamam, R.; Dawud, R.A.; Aldahmash, A.; Iqbal, Z.; Alfayez, M.; Kassem, M.; Mahmood, A. MicroRNA-4739 regulates osteogenic and adipocytic differentiation of immortalized human bone marrow stromal cells via targeting LRP3. Stem Cell Res. 2017, 20, 94–104. [Google Scholar] [CrossRef]
- Torii, D.; Tsutsui, T.W.; Watanabe, N.; Konishi, K. Bone morphogenetic protein 7 induces cementogenic differentiation of human periodontal ligament-derived mesenchymal stem cells. Odontology 2016, 104, 1–9. [Google Scholar] [CrossRef]
- Wilson, K.M.; Jagger, A.M.; Walker, M.; Seinkmane, E.; Fox, J.M.; Kröger, R.; Genever, P.; Ungar, D. Glycans modify mesenchymal stem cell differentiation to impact on the function of resulting osteoblasts. J. Cell Sci. 2018, 131. [Google Scholar] [CrossRef] [PubMed]
- Ali, D.; Hamam, R.; Alfayez, M.; Kassem, M.; Aldahmash, A.; Alajez, N.M. Epigenetic Library Screen Identifies Abexinostat as Novel Regulator of Adipocytic and Osteoblastic Differentiation of Human Skeletal (Mesenchymal) Stem Cells. Stem Cells Transl. Med. 2016, 5, 1036–1047. [Google Scholar] [CrossRef] [PubMed]
- Hossan, T.; Nagarajan, S.; Baumgart, S.J.; Xie, W.; Magallanes, R.T.; Hernandez, C.; Chiaroni, P.-M.; Indenbirken, D.; Spitzner, M.; Thomas-Chollier, M.; et al. Histone Chaperone SSRP1 is Essential for Wnt Signaling Pathway Activity During Osteoblast Differentiation. Stem Cells 2016, 34, 1369–1376. [Google Scholar] [CrossRef]
- Armbruster, N.; Krieg, J.; Weißenberger, M.; Scheller, C.; Steinert, A.F. Rescued Chondrogenesis of Mesenchymal Stem Cells under Interleukin 1 Challenge by Foamyviral Interleukin 1 Receptor Antagonist Gene Transfer. Front. Pharmacol. 2017, 8, 255. [Google Scholar] [CrossRef]
- Duan, P.; Toumpaniari, R.; Partridge, S.; Birch, M.A.; Genever, P.G.; Bull, S.J.; Dalgarno, K.W.; McCaskie, A.W.; Chen, J. How cell culture conditions affect the microstructure and nanomechanical properties of extracellular matrix formed by immortalized human mesenchymal stem cells: An experimental and modelling study. Mater. Sci. Eng. C 2018, 89, 149–159. [Google Scholar] [CrossRef]
- Hu, X.; Xie, P.; Li, W.; Li, Z.; Shan, H. Direct induction of hepatocyte-like cells from immortalized human bone marrow mesenchymal stem cells by overexpression of HNF4α. Biochem. Biophys. Res. Commun. 2016, 478, 791–797. [Google Scholar] [CrossRef]
- Tendulkar, G.; Ehnert, S.; Sreekumar, V.; Chen, T.; Kaps, H.-P.; Golombek, S.; Wendel, H.-P.; Nüssler, A.; Avci-Adali, M. Exogenous Delivery of Link N mRNA into Chondrocytes and MSCs—The Potential Role in Increasing Anabolic Response. Int. J. Mol. Sci. 2019, 20, 1716. [Google Scholar] [CrossRef]
- Lin, W.-H.; Yu, J.; Chen, G.; Tsai, W.-B. Fabrication of multi-biofunctional gelatin-based electrospun fibrous scaffolds for enhancement of osteogenesis of mesenchymal stem cells. Colloids Surf. B Biointerfaces 2016, 138, 26–31. [Google Scholar] [CrossRef]
- Harkness, L.; Zaher, W.; Ditzel, N.; Isa, A.; Kassem, M. CD146/MCAM defines functionality of human bone marrow stromal stem cell populations. Stem Cell Res. Ther. 2016, 7, 4. [Google Scholar] [CrossRef]
- Jayasuriya, C.T.; Hu, N.; Li, J.; Lemme, N.; Terek, R.; Ehrlich, M.G.; Chen, Q. Molecular characterization of mesenchymal stem cells in human osteoarthritis cartilage reveals contribution to the OA phenotype. Sci. Rep. 2018, 8, 7044. [Google Scholar] [CrossRef]
- Hu, N.; Gao, Y.; Jayasuriya, C.T.; Liu, W.; Du, H.; Ding, J.; Feng, M.; Chen, Q. Chondrogenic induction of human osteoarthritic cartilage-derived mesenchymal stem cells activates mineralization and hypertrophic and osteogenic gene expression through a mechanomiR. Arthr. Res. Ther. 2019, 21, 167. [Google Scholar] [CrossRef] [PubMed]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G. Preferred Reporting Items for Systematic Reviews and Meta-Analyses: The PRISMA Statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.S.; Shim, J.S.; Paik, M.J.; Joo, W.H.; Kim, S.H.; Lee, G.; Kim, D.W. Characterization of a growth-elevated cell line of human bone marrow-derived mesenchymal stem cells by SV40 T-antigen. Biotechnol. Bioprocess. Eng. 2015, 20, 498–505. [Google Scholar] [CrossRef]
- Harigaya, K.; Handa, H. Generation of functional clonal cell lines from human bone marrow stroma. Proc. Natl. Acad. Sci. USA 1985, 82, 3477–3480. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Glowacki, J.; Hahne, J.; Xie, L.; LeBoff, M.S.; Zhou, S. Dehydroepiandrosterone Stimulation of Osteoblastogenesis in Human MSCs Requires IGF-I Signaling. J. Cell. Biochem. 2016, 117, 1769–1774. [Google Scholar] [CrossRef]
- Thalmeier, K.; Meissner, P.; Reisbach, G.; Falk, M.; Brechtel, A.; Dörmer, P. Establishment of two permanent human bone marrow stromal cell lines with long-term post irradiation feeder capacity. Blood 1994, 83, 1799–1807. [Google Scholar] [CrossRef]
- Conrad, C.; Gottgens, B.; Kinston, S.; Ellwart, J.; Huss, R. GATA transcription in a small rhodamine 123(low)CD34(+) subpopulation of a peripheral blood-derived CD34(-)CD105(+) mesenchymal cell line. Exp. Hematol. 2002, 30, 887–895. [Google Scholar] [CrossRef]
- Shu, Y.; Yang, C.; Ji, X.; Zhang, L.; Bi, Y.; Yang, K.; Gong, M.; Liu, X.; Guo, Q.; Su, Y.; et al. Reversibly immortalized human umbilical cord-derived mesenchymal stem cells (UC-MSCs) are responsive to BMP9-induced osteogenic and adipogenic differentiation. J. Cell. Biochem. 2018, 119, 8872–8886. [Google Scholar] [CrossRef]
- Song, D.; Zhang, F.; Reid, R.R.; Ye, J.; Wei, Q.; Liao, J.; Zou, Y.; Fan, J.; Ma, C.; Hu, X.; et al. BMP9 induces osteogenesis and adipogenesis in the immortalized human cranial suture progenitors from the patent sutures of craniosynostosis patients. J. Cell. Mol. Med. 2017, 21, 2782–2795. [Google Scholar] [CrossRef]
- Alexander, D.; Biller, R.; Rieger, M.; Ardjomandi, N.; Reinert, S. Phenotypic Characterization of a Human Immortalized Cranial Periosteal Cell Line. Cell. Physiol. Biochem. 2015, 35, 2244–2254. [Google Scholar] [CrossRef]
- Wu, Y.; Feng, G.; Song, J.; Zhang, Y.; Yu, Y.; Huang, L.; Zheng, L.; Deng, F. TrAmplification of Human Dental Follicle Cells by piggyBac Transposon—Mediated Reversible Immortalization System. PLoS ONE 2015, 10, e0130937. [Google Scholar] [CrossRef] [PubMed]
- Dou, L.; Wu, Y.; Yan, Q.; Wang, J.; Zhang, Y.; Ji, P. Secretome profiles of immortalized dental follicle cells using iTRAQ-based proteomic analysis. Sci. Rep. 2017, 7, 7300. [Google Scholar] [CrossRef] [PubMed]
- Hung, S.-C.; Yang, D.-M.; Chang, C.-F.; Lin, R.-J.; Wang, J.-S.; Low-Tone Ho, L.; Yang, W.K. Immortalization without neoplastic transformation of human mesenchymal stem cells by transduction with HPV16E6/E7 genes. Int. J. Cancer 2004, 110, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.-C.; Chen, C.-L.; Liu, H.-C.; Lee, Y.-T.; Wang, H.-W.; Hou, L.-T.; Hung, S.-C. Overexpression of hTERT increases stem-like properties and decreases spontaneous differentiation in human mesenchymal stem cell lines. J. Biomed. Sci. 2010, 17, 64. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Moreno, M.; Hermida-Gómez, T.; Gallardo, M.E.; Dalmao-Fernández, A.; Rego-Pérez, I.; Garesse, R.; Blanco, F.J. Generating Rho-0 Cells Using Mesenchymal Stem Cell Lines. PLoS ONE 2016, 11, e0164199. [Google Scholar] [CrossRef]
- Chen, Y.-C.; Wu, K.-C.; Huang, B.-M.; So, E.C.; Wang, Y.-K. Midazolam inhibits chondrogenesis via peripheral benzodiazepine receptor in human mesenchymal stem cells. J. Cell. Mol. Med. 2018, 22, 2896–2907. [Google Scholar] [CrossRef]
- Mori, T.; Kiyono, T.; Imabayashi, H.; Takeda, Y.; Tsuchiya, K.; Miyoshi, S.; Makino, H.; Matsumoto, K.; Saito, H.; Ogawa, S.; et al. Combination of hTERT and bmi-1, E6, or E7 Induces Prolongation of the Life Span of Bone Marrow Stromal Cells from an Elderly Donor without Affecting Their Neurogenic Potential. Mol. Cell. Biol. 2005, 25, 5183–5195. [Google Scholar] [CrossRef]
- Kusuyama, J.; Kamisono, A.; ChangHwan, S.; Amir, M.S.; Bandow, K.; Eiraku, N.; Ohnishi, T.; Matsuguchi, T. Spleen tyrosine kinase influences the early stages of multilineage differentiation of bone marrow stromal cell lines by regulating phospholipase C gamma activities. J. Cell. Physiol. 2018, 233, 2549–2559. [Google Scholar] [CrossRef]
- Roecklein, B.A.; Torok-Storb, B. Functionally distinct human marrow stromal cell lines immortalized by transduction with the human papilloma virus E6/E7 genes. Blood 1995, 85, 997–1005. [Google Scholar] [CrossRef]
- Liu, B.; Wu, S.; Han, L.; Zhang, C. β-catenin signaling induces the osteoblastogenic differentiation of human pre-osteoblastic and bone marrow stromal cells mainly through the upregulation of osterix expression. Int. J. Mol. Med. 2015, 36, 1572–1582. [Google Scholar] [CrossRef][Green Version]
- Wei, K.; Xie, Y.; Chen, T.; Fu, B.; Cui, S.; Wang, Y.; Cai, G.; Chen, X. ERK1/2 signaling mediated naringin-induced osteogenic differentiation of immortalized human periodontal ligament stem cells. Biochem. Biophys. Res. Commun. 2017, 489, 319–325. [Google Scholar] [CrossRef] [PubMed]
- Simonsen, J.L.; Rosada, C.; Serakinci, N.; Justesen, J.; Stenderup, K.; Rattan, S.I.S.; Jensen, T.G.; Kassem, M. Telomerase expression extends the proliferative life-span and maintains the osteogenic potential of human bone marrow stromal cells. Nat. Biotechnol. 2002, 20, 592–596. [Google Scholar] [CrossRef] [PubMed]
- Leber, J.; Barekzai, J.; Blumenstock, M.; Pospisil, B.; Salzig, D.; Czermak, P. Microcarrier choice and bead-to-bead transfer for human mesenchymal stem cells in serum-containing and chemically defined media. Process. Biochem. 2017, 59, 255–265. [Google Scholar] [CrossRef]
- Ziouti, F.; Ebert, R.; Rummler, M.; Krug, M.; Müller-Deubert, S.; Lüdemann, M.; Jakob, F.; Willie, B.M.; Jundt, F. NOTCH Signaling Is Activated through Mechanical Strain in Human Bone Marrow-Derived Mesenchymal Stromal Cells. Stem Cells Int. 2019, 2019, 1–13. [Google Scholar] [CrossRef]
- Müller-Deubert, S.; Seefried, L.; Krug, M.; Jakob, F.; Ebert, R. Epidermal growth factor as a mechanosensitizer in human bone marrow stromal cells. Stem Cell Res. 2017, 24, 69–76. [Google Scholar] [CrossRef]
- Salzig, D.; Leber, J.; Merkewitz, K.; Lange, M.C.; Köster, N.; Czermak, P. Attachment, Growth, and Detachment of Human Mesenchymal Stem Cells in a Chemically Defined Medium. Stem Cells Int. 2016, 2016. [Google Scholar] [CrossRef]
- Larsen, K.H.; Frederiksen, C.M.; Burns, J.S.; Abdallah, B.M.; Kassem, M. Identifying A Molecular Phenotype for Bone Marrow Stromal Cells With In Vivo Bone Forming Capacity. J. Bone Miner. Res. 2009, 25. [Google Scholar] [CrossRef]
- Elsafadi, M.; Manikandan, M.; Atteya, M.; Abu Dawud, R.; Almalki, S.; Ali Kaimkhani, Z.; Aldahmash, A.; Alajez, N.M.; Alfayez, M.; Kassem, M.; et al. SERPINB2 is a novel TGFβ-responsive lineage fate determinant of human bone marrow stromal cells. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Abdallah, B.M.; Haack-Sørensen, M.; Burns, J.S.; Elsnab, B.; Jakob, F.; Hokland, P.; Kassem, M. Maintenance of differentiation potential of human bone marrow mesenchymal stem cells immortalized by human telomerase reverse transcriptase gene despite of extensive proliferation. Biochem. Biophys. Res. Commun. 2005, 326, 527–538. [Google Scholar] [CrossRef]
- Skårn, M.; Noordhuis, P.; Wang, M.-Y.; Veuger, M.; Kresse, S.H.; Egeland, E.V.; Micci, F.; Namløs, H.M.; Håkelien, A.-M.; Olafsrud, S.M.; et al. Generation and Characterization of an Immortalized Human Mesenchymal Stromal Cell Line. Stem Cells Dev. 2014, 23, 2377–2389. [Google Scholar] [CrossRef]
- Wu, L.; Leijten, J.C.H.; Georgi, N.; Post, J.N.; van Blitterswijk, C.A.; Karperien, M. Trophic effects of mesenchymal stem cells increase chondrocyte proliferation and matrix formation. Tissue Eng. Part A 2011, 17, 1425–1436. [Google Scholar] [CrossRef] [PubMed]
- Fayyad, A.; Khan, A.; Abdallah, S.; Alomran, S.; Bajou, K.; Khattak, M. Rosiglitazone Enhances Browning Adipocytes in Association with MAPK and PI3-K Pathways During the Differentiation of Telomerase-Transformed Mesenchymal Stromal Cells into Adipocytes. Int. J. Mol. Sci. 2019, 20, 1618. [Google Scholar] [CrossRef] [PubMed]
- Dale, T.P.; de Castro, A.; Kuiper, N.J.; Parkinson, E.K.; Forsyth, N.R. Immortalisation with hTERT Impacts on Sulphated Glycosaminoglycan Secretion and Immunophenotype in a Variable and Cell Specific Manner. PLoS ONE 2015, 10, e0133745. [Google Scholar] [CrossRef] [PubMed]
- Dale, T.P.; Forsyth, N.R. Ectopic Telomerase Expression Fails to Maintain Chondrogenic Capacity in Three-Dimensional Cultures of Clinically Relevant Cell Types. Biores. Open Access 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Blaschke, M.; Koepp, R.; Lenz, C.; Kruppa, J.; Jung, K.; Siggelkow, H. Crohn’s disease patient serum changes protein expression in a human mesenchymal stem cell model in a linear relationship to patients’ disease stage and to bone mineral density. J. Clin. Transl. Endocrinol. 2018, 13, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Schmid, J.; Schwarz, S.; Meier-Staude, R.; Sudhop, S.; Clausen-Schaumann, H.; Schieker, M.; Huber, R. A Perfusion Bioreactor System for Cell Seeding and Oxygen-Controlled Cultivation of Three-Dimensional Cell Cultures. Tissue Eng. Part C. Methods 2018, 24, 585–595. [Google Scholar] [CrossRef]
- Sreekumar, V.; Aspera-Werz, R.; Ehnert, S.; Strobel, J.; Tendulkar, G.; Heid, D.; Schreiner, A.; Arnscheidt, C.; Nussler, A.K. Resveratrol protects primary cilia integrity of human mesenchymal stem cells from cigarette smoke to improve osteogenic differentiation in vitro. Arch. Toxicol. 2018, 92, 1525–1538. [Google Scholar] [CrossRef]
- Westphal, I.; Jedelhauser, C.; Liebsch, G.; Wilhelmi, A.; Aszodi, A.; Schieker, M. Oxygen mapping: Probing a novel seeding strategy for bone tissue engineering. Biotechnol. Bioeng. 2017, 114, 894–902. [Google Scholar] [CrossRef]
- Aspera-Werz, R.H.; Chen, T.; Ehnert, S.; Zhu, S.; Fröhlich, T.; Nussler, A.K. Cigarette Smoke Induces the Risk of Metabolic Bone Diseases: Transforming Growth Factor Beta Signaling Impairment via Dysfunctional Primary Cilia Affects Migration, Proliferation, and Differentiation of Human Mesenchymal Stem Cells. Int. J. Mol. Sci. 2019, 20, 2915. [Google Scholar] [CrossRef]
- Duan, P.; Chen, J. Nanomechanical and microstructure analysis of extracellular matrix layer of immortalized cell line Y201 from human mesenchymal stem cells. Surf. Coat. Technol. 2015, 284, 417–421. [Google Scholar] [CrossRef]
- Galarza Torre, A.; Shaw, J.E.; Wood, A.; Gilbert, H.T.J.; Dobre, O.; Genever, P.; Brennan, K.; Richardson, S.M.; Swift, J. An immortalised mesenchymal stem cell line maintains mechano-responsive behaviour and can be used as a reporter of substrate stiffness. Sci. Rep. 2018, 8, 8981. [Google Scholar] [CrossRef] [PubMed]
- Vanderburgh, J.; Sterling, J.A.; Guelcher, S.A. 3D Printing of Tissue Engineered Constructs for In Vitro Modeling of Disease Progression and Drug Screening. Ann. Biomed. Eng. 2017, 45, 164–179. [Google Scholar] [CrossRef] [PubMed]
- Wolbank, S.; Stadler, G.; Peterbauer, A.; Gillich, A.; Karbiener, M.; Streubel, B.; Wieser, M.; Katinger, H.; van Griensven, M.; Redl, H.; et al. Telomerase Immortalized Human Amnion- and Adipose-Derived Mesenchymal Stem Cells: Maintenance of Differentiation and Immunomodulatory Characteristics. Tissue Eng. Part A 2009, 15, 1843–1854. [Google Scholar] [CrossRef] [PubMed]
- Balducci, L.; Blasi, A.; Saldarelli, M.; Soleti, A.; Pessina, A.; Bonomi, A.; Coccè, V.; Dossena, M.; Tosetti, V.; Ceserani, V.; et al. Immortalization of human adipose-derived stromal cells: Production of cell lines with high growth rate, mesenchymal marker expression and capability to secrete high levels of angiogenic factors. Stem Cell Res. Ther. 2014, 5, 63. [Google Scholar] [CrossRef]
- Siska, E.K.; Weisman, I.; Romano, J.; Ivics, Z.; Izsvák, Z.; Barkai, U.; Petrakis, S.; Koliakos, G. Generation of an immortalized mesenchymal stem cell line producing a secreted biosensor protein for glucose monitoring. PLoS ONE 2017, 12, e0185498. [Google Scholar] [CrossRef]
- Akazawa, Y.; Hasegawa, T.; Yoshimura, Y.; Chosa, N.; Asakawa, T.; Ueda, K.; Sugimoto, A.; Kitamura, T.; Nakagawa, H.; Ishisaki, A.; et al. Recruitment of mesenchymal stem cells by stromal cell-derived factor 1α in pulp cells from deciduous teeth. Int. J. Mol. Med. 2015, 36, 442–448. [Google Scholar] [CrossRef]
- Sugimoto, A.; Miyazaki, A.; Kawarabayashi, K.; Shono, M.; Akazawa, Y.; Hasegawa, T.; Ueda-Yamaguchi, K.; Kitamura, T.; Yoshizaki, K.; Fukumoto, S.; et al. Piezo type mechanosensitive ion channel component 1 functions as a regulator of the cell fate determination of mesenchymal stem cells. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Tsutsui, T.; Kumakura, S.-I.; Yamamoto, A.; Kanai, H.; Tamura, Y.; Kato, T.; Anpo, M.; Tahara, H.; Barrett, J.C. Association of p16INK4a and pRb inactivation with immortalization of human cells. Carcinogenesis 2002, 23, 2111–2117. [Google Scholar] [CrossRef]
- Torii, D.; Konishi, K.; Watanabe, N.; Goto, S.; Tsutsui, T. Cementogenic potential of multipotential mesenchymal stem cells purified from the human periodontal ligament. Odontology 2015, 103, 27–35. [Google Scholar] [CrossRef]
- Qin, S.Q.; Kusuma, G.D.; Al-Sowayan, B.; Pace, R.A.; Isenmann, S.; Pertile, M.D.; Gronthos, S.; Abumaree, M.H.; Brennecke, S.P.; Kalionis, B. Establishment and characterization of fetal and maternal mesenchymal stem/stromal cell lines from the human term placenta. Placenta 2016, 39, 134–146. [Google Scholar] [CrossRef]
- Kusuma, G.D.; Brennecke, S.P.; O’Connor, A.J.; Kalionis, B.; Heath, D.E. Decellularized extracellular matrices produced from immortal cell lines derived from different parts of the placenta support primary mesenchymal stem cell expansion. PLoS ONE 2017, 12, e0171488. [Google Scholar] [CrossRef] [PubMed]
- Kusuma, G.D.; Yang, M.C.; Brennecke, S.P.; O’Connor, A.J.; Kalionis, B.; Heath, D.E. Transferable Matrixes Produced from Decellularized Extracellular Matrix Promote Proliferation and Osteogenic Differentiation of Mesenchymal Stem Cells and Facilitate Scale-Up. ACS Biomater. Sci. Eng. 2018, 8, 744–752. [Google Scholar] [CrossRef]
- Matta, C.; Fodor, J.; Miosge, N.; Takács, R.; Juhász, T.; Rybaltovszki, H.; Tóth, A.; Csernoch, L.; Zákány, R. Purinergic signalling is required for calcium oscillations in migratory chondrogenic progenitor cells. Pflüg. Arch. Eur. J. Physiol. 2015, 467, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Koelling, S.; Kruegel, J.; Irmer, M.; Path, J.R.; Sadowski, B.; Miro, X.; Miosge, N. Migratory Chondrogenic Progenitor Cells from Repair Tissue during the Later Stages of Human Osteoarthritis. Cell Stem Cell 2009, 4, 324–335. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, M.; Higashino, A.; Takeuchi, K.; Hori, Y.; Koshiba-Takeuchi, K.; Makino, H.; Monobe, Y.; Kishida, M.; Adachi, J.; Takeuchi, J.; et al. Transcriptional Dynamics of Immortalized Human Mesenchymal Stem Cells during Transformation. PLoS ONE 2015, 10, e0126562. [Google Scholar]
- Takeuchi, M.; Takeuchi, K.; Kohara, A.; Satoh, M.; Shioda, S.; Ozawa, Y.; Ohtani, A.; Morita, K.; Hirano, T.; Terai, M.; et al. Chromosomal instability in human mesenchymal stem cells immortalized with human papilloma virus E6, E7, and hTERT genes. Vitr. Cell. Dev. Biol. Anim. 2007, 43, 129–138. [Google Scholar] [CrossRef]
- Kajiyama, S.; Ujiie, Y.; Nishikawa, S.; Inoue, K.; Shirakawa, S.; Hanada, N.; Liddell, R.; Davies, J.E.; Gomi, K. Bone formation by human umbilical cord perivascular cells. J. Biomed. Mater. Res. Part A 2015, 103, 2807–2814. [Google Scholar] [CrossRef]
- Okita, N.; Honda, Y.; Kishimoto, N.; Liao, W.; Azumi, E.; Hashimoto, Y.; Matsumoto, N. Supplementation of strontium to a chondrogenic medium promotes chondrogenic differentiation of human dedifferentiated fat cells. Tissue Eng. Part A 2015, 21, 1695–1704. [Google Scholar] [CrossRef]
- Toda, H.; Yamamoto, M.; Uyama, H.; Tabata, Y. Fabrication of hydrogels with elasticity changed by alkaline phosphatase for stem cell culture. Acta Biomater. 2016, 29, 215–227. [Google Scholar] [CrossRef]
- Funes, J.M.; Quintero, M.; Henderson, S.; Martinez, D.; Qureshi, U.; Westwood, C.; Clements, M.O.; Bourboulia, D.; Pedley, R.B.; Moncada, S.; et al. Transformation of human mesenchymal stem cells increases their dependency on oxidative phosphorylation for energy production. Proc. Natl. Acad. Sci. USA 2007, 104, 6223–6228. [Google Scholar] [CrossRef]
- Abarrategi, A.; Gambera, S.; Alfranca, A.; Rodriguez-Milla, M.A.; Perez-Tavarez, R.; Rouault-Pierre, K.; Waclawiczek, A.; Chakravarty, P.; Mulero, F.; Trigueros, C.; et al. c-Fos induces chondrogenic tumor formation in immortalized human mesenchymal progenitor cells. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Gambera, S.; Abarrategi, A.; Rodríguez-Milla, M.A.; Mulero, F.; Menéndez, S.T.; Rodriguez, R.; Navarro, S.; García-Castro, J. Role of Activator Protein-1 Complex on the Phenotype of Human Osteosarcomas Generated from Mesenchymal Stem Cells. Stem Cells 2018, 36, 1487–1500. [Google Scholar] [CrossRef] [PubMed]
- Takeda, Y.; Mori, T.; Imabayashi, H.; Kiyono, T.; Gojo, S.; Miyoshi, S.; Hida, N.; Ita, M.; Segawa, K.; Ogawa, S.; et al. Can the life span of human marrow stromal cells be prolonged by bmi-1, E6, E7, and/or telomerase without affecting cardiomyogenic differentiation? J. Gene Med. 2004, 6, 833–845. [Google Scholar] [CrossRef] [PubMed]
- Nemoto, A.; Chosa, N.; Kyakumoto, S.; Yokota, S.; Kamo, M.; Noda, M.; Ishisaki, A. Water-soluble factors eluated from surface pre-reacted glass-ionomer filler promote osteoblastic differentiation of human mesenchymal stem cells. Mol. Med. Rep. 2017, 17, 3448–3454. [Google Scholar] [CrossRef] [PubMed]
- Carnero, A.; Blanco-Aparicio, C.; Kondoh, H.; Lleonart, M.E.; Martinez-Leal, J.F.; Mondello, C.; Scovassi, A.I.; Bisson, W.H.; Amedei, A.; Roy, R.; et al. Disruptive chemicals, senescence and immortality. Carcinogenesis 2015, 36, S19–S37. [Google Scholar] [CrossRef] [PubMed]
- An, P.; Sáenz Robles, M.T.; Pipas, J.M. Large T Antigens of Polyomaviruses: Amazing Molecular Machines. Annu. Rev. Microbiol. 2012, 66, 213–236. [Google Scholar] [CrossRef]
- Ozono, E.; Yamaoka, S.; Ohtani, K. To Grow, Stop or Die?—Novel Tumor-Suppressive Mechanism Regulated by the Transcription Factor E2F. In Future Aspects of Tumor Suppressor Gene; InTech: London, UK, 2013. [Google Scholar]
- Tomaić, V. Functional Roles of E6 and E7 Oncoproteins in HPV-Induced Malignancies at Diverse Anatomical Sites. Cancers 2016, 8, 95. [Google Scholar] [CrossRef]
- Tátrai, P.; Szepesi, Á.; Matula, Z.; Szigeti, A.; Buchan, G.; Mádi, A.; Uher, F.; Német, K. Combined introduction of Bmi-1 and hTERT immortalizes human adipose tissue-derived stromal cells with low risk of transformation. Biochem. Biophys. Res. Commun. 2012, 422, 28–35. [Google Scholar] [CrossRef]
- Somoza, R.A.; Welter, J.F.; Correa, D.; Caplan, A.I. Chondrogenic differentiation of mesenchymal stem cells: Challenges and unfulfilled expectations. Tissue Eng. Part B Rev. 2014, 20, 596–608. [Google Scholar] [CrossRef]
- Castro-Viñuelas, R.; Sanjurjo-Rodríguez, C.; Piñeiro-Ramil, M.; Hermida-Gómez, T.; Fuentes-Boquete, I.; de Toro-Santos, F.; Blanco-García, F.; Díaz-Prado, S. Induced pluripotent stem cells for cartilage repair: Current status and future perspectives. Eur. Cells Mater. 2018, 36, 96–109. [Google Scholar] [CrossRef]
- Finger, F.; Schörle, C.; Zien, A.; Gebhard, P.; Goldring, M.B.; Aigner, T. Molecular phenotyping of human chondrocyte cell lines T/C-28a2, T/C-28a4, and C-28/I2. Arthr. Rheumatol. 2003, 48, 3395–3403. [Google Scholar] [CrossRef] [PubMed]
- Dominici, M.; Le Blanc, K.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.; Krause, D.S.; Deans, R.J.; Keating, A.; Prockop, D.J.; Horwitz, E.M. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 2006, 8, 315–317. [Google Scholar] [CrossRef] [PubMed]
- Uder, C.; Brückner, S.; Winkler, S.; Tautenhahn, H.-M.; Christ, B. Mammalian MSC from selected species: Features and applications. Cytom. Part A 2018, 93, 32–49. [Google Scholar] [CrossRef] [PubMed]
- Halfon, S.; Abramov, N.; Grinblat, B.; Ginis, I. Markers Distinguishing Mesenchymal Stem Cells from Fibroblasts Are Downregulated with Passaging. Stem Cells Dev. 2011, 20, 53–66. [Google Scholar] [CrossRef] [PubMed]
- Cleary, M.A.; Narcisi, R.; Focke, K.; van der Linden, R.; Brama, P.A.J.; van Osch, G.J.V.M. Expression of CD105 on expanded mesenchymal stem cells does not predict their chondrogenic potential. Osteoarthr. Cartil. 2016, 24, 868–872. [Google Scholar] [CrossRef] [PubMed]
- Burns, J.S.; Harkness, L.; Aldahmash, A.; Gautier, L.; Kassem, M. Chromosome copy number variation in telomerized human bone marrow stromal cells; insights for monitoring safe ex-vivo expansion of adult stem cells. Stem Cell Res. 2017, 25, 6–17. [Google Scholar] [CrossRef] [PubMed]
- Bianco, P.; Robey, P.G. Skeletal stem cells. Development 2015, 142, 1023–1027. [Google Scholar] [CrossRef]
| MSC Line | Immortalization Genes | Immortalization Method | Tissue | Donor Characteristics | STR Genotyping | Clonality | Tumorigenicity | References |
|---|---|---|---|---|---|---|---|---|
| hMSC-T | SV40LT | Transfection | Bone marrow | Unknown | No | Unclear 1 | No (tested by soft agar) | [44] |
| KM101 | SV40LT | Transfection | Bone marrow | 48-year-old male | No | Yes | Not tested | [45,46] |
| L87/4 | SV40LT | Transfection | Bone marrow | 70-year-old male | No | Yes | Not tested/shown | [17,47] |
| V54/2 | SV40LT | Transfection | Peripheral blood | Healthy donor | No | Yes | Not tested/shown | [17,48] |
| iUC-MSCs | SV40LT | Retroviral transduction | Umbilical cord | Unknown | No | No | No (tested in IDM) | [49] |
| iSuPs | SV40LT | Retroviral transduction | Coronal sutures | 15 to 17-month-old males | No | No | No (tested in IDM) | [50] |
| TAg cells | SV40LT | Lentiviral transduction | Cranial periosteum | Healthy (fracture patient) | No | No | Not tested | [51] |
| iDFCs | SV40LT | Retroviral transduction | Dental follicle | Three young adults (18–20 years old) | No | Yes | Not tested | [52,53] |
| OA-MSCs | SV40LT | Retroviral transduction | Articular cartilage (knee) | Osteoarthritic 61-year-old male and 69-year-old female | Yes | Yes 2 | Not tested/shown | [41,42] |
| KP | E6/E7 | Retroviral transduction | Bone marrow | 61-year-old female | No | No | No (tested in IDM) | [54,55,56,57] |
| UE6E7-16 3 | E6/E7 | Retroviral transduction | Bone marrow | 91-year-old female | Yes | Yes | Not shown | [58,59] |
| HS-27 | E6/E7 | Retroviral transduction | Bone marrow | Adult donor | No | Yes | Not tested | [60,61] |
| PDLSC-Bmi1 4 | Bmi1 | Retroviral transduction | Periodontal ligament | 15 to 20-year-old donors | No | No | Not tested | [62] |
| hMSC-hTERT | hTERT | Retroviral transduction | Bone marrow | Healthy 33-year-old male | No | No 5 | No (tested in IDM) | [30,33,34,63,64,65,66,67,68,69] |
| TERT4 (hMSC-hTERT derived) | hTERT | Retroviral transduction | Bone marrow | Healthy 33-year-old male | No | No | No (tested in IDM) 6 | [35,40,70] |
| iMSC#3 | hTERT | Retroviral transduction | Bone marrow | Healthy male | No | Yes | No (tested in IDM) | [71,72,73] |
| BMA13H 7 | hTERT | Retroviral transduction | Bone marrow | Unknown | No | No | Not tested | [74,75] |
| SCP-1 | hTERT | Lentiviral transduction | Bone marrow | Unknown | No | Yes | No (tested in IMD and by soft agar assay) | [14,20,24,25,38,76,77,78,79,80] |
| Y201 | hTERT | Lentiviral transduction | Bone marrow | Unknown | No | Yes | No (tested in IDM) | [19,36,81,82,83] |
| Y101 | hTERT | Lentiviral transduction | Bone marrow | Unknown | [19,32] | |||
| MSOD | hTERT | Lentiviral transduction | Bone marrow | Healthy 55-year-old female | Yes | Yes | No (tested in IDM) | [15,26] |
| ASC/TERT1 | hTERT | Retroviral transduction | Adipose tissue | Unknown | Yes | No | No (soft agar assay) | [22,84] |
| hASCs-T 7 | hTERT | Lentiviral transduction | Adipose tissue | Two males and two females (21 to 59 years old) | No | No | No (soft agar assay) | [85] |
| GB/hTERT MSCs | hTERT | Transfection | Umbilical cord | Unknown | No | No | No (soft agar assay) | [86] |
| SDP11 | hTERT | Transfection | Dental pulp | 6 to 8-year-old donors | No | Yes | Not tested | [87,88] |
| Pelt cells | hTERT | Retroviral transduction | Periodontal ligament | Adult donor | No | No | Not tested/shown | [31,89,90] |
| CMSC29 | hTERT | Retroviral transduction | Placenta (Chorionic Villi) | Unknown | No | Yes | No (tested by soft agar assay) | [91,92,93] |
| DMSC23 | hTERT | Retroviral transduction | Placenta (Decidua Basalis) | Unknown | No | Yes | No (tested by soft agar assay) | [91,92,93] |
| CPC531 | hTERT | Lentiviral transduction | Articular cartilage (knee) | 65 to 75-year-old patients | No | Unclear 1 | Not tested/shown | [94,95] |
| hASCs-TS (same parental cells as hASCs-T) | hTERT and SV40LT | Lentiviral transduction | Adipose tissue | Two males and two females (21 to 59 years old) | No | No | No (soft agar assay) | [85] |
| 3A6 (KP-derived) | hTERT and E6/E7 | Transfection (hTERT) | Bone marrow | 61-year-old female | No | Yes | Not tested | [39,55,56] |
| hASCs-TE (same parental cells as hASCs-T) | hTERT and E6/E7 | Lentiviral transduction | Adipose tissue | Two males and two females (21 to 59 years old) | No | No | No (soft agar assay) | [85] |
| UE6E7T-3 (same parental cells as UE6E7-16) | hTERT and E6/E7 | Retroviral transduction | Bone marrow | 91-year-old female | Yes | Yes | Tested in soft agar at “low” (PDs ≤ 200) and high (PDs = 252) passages, with only high passage UE6E7T-3 being capable of forming colonies; high passage UE6E7T-3 formed sarcomas in IDM | [28,96,97] |
| UE6E7T-11 (same parental cells as UE6E7-16) | hTERT and E6/E7 | Retroviral transduction | Bone marrow | 91-year-old female | Yes | Yes | Not shown | [58,98] |
| UE6E7T-2 (same parental cells as UE6E7-16) | hTERT and E6/E7 | Retroviral transduction | Bone marrow | 91-year-old female | Yes | Yes | Not shown | [99] |
| imhMSCs | hTERT and E6/E7 | Retroviral transduction | Bone marrow | Unknown | No | Unclear 1 | No (tested in IDM) | [18,23,29,100] |
| 3 Hits hMPC | hTERT and E6/E7 | Retroviral transduction | Bone marrow | Healthy 34-year-old male | Yes | No | No (tested in IDM; only c-Fos-transduced cells were tumorigenic) | [101,102,103] |
| UE7T-13 (same parental cells as UE6E7-16) | hTERT and E7 | Retroviral transduction | Bone marrow | 91-year-old female | Yes | Yes | Not shown | [21,37,88,104,105] |
| MSC Line | Osteogenic Potential | Chondrogenic Potential | Adipogenic Potential |
|---|---|---|---|
| hMSC-T | Positive for VKS and osteocalcin upregulation (increased compared with primary MSCs) [44] | Not tested | Not tested |
| KM101 | Positive for ALP activity [46] | Not tested | Tested and no adipogenic differentiation potential was found (also not shown) [45] |
| L87/4 | Not tested/shown | Positive for ABS and ColII immunostaining in 3D alginate and PGA/PLLA scaffolds [17] | Not tested/shown |
| V54/2 | Not tested/shown | Positive for ABS and ColII immunostaining in 3D alginate and PGA/PLLA scaffolds [17] | Not tested/shown |
| iUC-MSCs | Positive for Runx2 and Osteocalcin upregulation [49] | Positive for Sox9 upregulation [49] | Positive for PPARγ upregulation [49] |
| iSuPs | Positive for ARS (increased if SV40LT is removed) and osteogenesis-related genes upregulation [50] | No chondrogenic differentiation potential was found (also not shown) [50] | Positive for OROS (increased if SV40LT is removed) [50] |
| TAg cells | Positive for hydroxyapatite formation (showing earlier and stronger mineralization than parental cells) and upregulation of osteogenesis-related genes (increased compared with primary cells) [51] | Not tested | Not tested |
| iDFCs | Positive for ARS, APS, and osteogenesis-related genes upregulation; osteogenic potential similar to primary cells [52] | Positive for ABS and SOX9 upregulation in 2D culture [52] | Positive for OROS and adipogenesis-related genes upregulation (PPARγ and LPL) [52] |
| OA-MSCs | Positive for ARS and ALP upregulation [41,42] | Positive for SOS [41], ABS [42] and upregulation of Sox9, Col2A1, ACAN and COL10A1 [41,42] in pellet [41] and 2D culture [42] | Positive for OROS (weak staining) and LPL upregulation [41] |
| KP | Positive for APS, ARS, and VKS [54] | Proved by ABS [54,57] and ColII immunostaining [57] in pellet culture | Positive for OROS [54] |
| UE6E7-16 | Positive for osteocalcin production [59] | Not tested/shown | Positive for PPARγ production [59] |
| HS-27 | Positive for ALP activity, calcium deposition and osterix upregulation [61] | Not tested/shown | Positive for OROS in presence of steroids [60] |
| PDLSC-Bmi1 | Positive for ARS, ALP activity, and osteogenesis-related genes upregulation [62] | Not tested | Positive for OROS [62] |
| hMSC-hTERT | Positive for ARS [64], ALP activity [33], upregulation of osteogenesis-related genes [33,63,68], and in vivo bone formation [63,68] | Positive for ABS [64] and ColII immunostaining [63,67] in 2D culture | Positive for OROS and upregulation of adipogenesis-related genes [33] |
| TERT4 (hMSC-hTERT derived) | Positive for ARS [40,70], ALP activity [40] and upregulation of osteogenesis-related genes [40] | Positive for ABS [35,70], GAG assay [35], and upregulation of ColII [35,70] but also ColX [35], in pellet culture; reduced compared with primary MSCs | Positive for OROS [40,70] and upregulation of adipogenesis-related genes [40] |
| iMSC#3 | Positive for ARS, APS, and Runx2 upregulation [71] | Positive for ABS, TBS, and GAG assay in pellet culture [72]; low chondrogenic potential but stimulation of chondrocyte differentiation | Positive for OROS, adipogenesis-related genes upregulation [71,73], and NRS [73] |
| BMA13H | Positive for ARS (reduced compared with primary cells) [74] | Positive for ABS and GAG assay in 2D culture [74]; also positive for TBS, PSR and aggrecan and ColII immunostaining in 3D culture [75]; chondrogenic potential reduced compared with primary cells [74] | Positive for OROS (reduced compared with primary cells) [74] |
| SCP-1 | Positive for VKS (increased compared with MSCs) [14], ARS [78], ALP activity [78], and upregulation of osteogenesis-related genes [76] | Positive for TBS in pellet culture [14]; ColII and GAG production in 3D printed scaffolds [24] | Positive for OROS [14] |
| Y201 | Positive for ARS [19], ALP activity [19,82,83], and Runx2 upregulation [19,82] | Positive for ABS, GAG assay, and Sox9 upregulation in pellet culture [19] | Positive for OROS (reduced compared to primary MSCs) and upregulation of adipogenesis-related genes [19,82] |
| Y101 (derived from the same donor than Y201) | Proved by ARS [19,32], VKS [32], ALP activity, and osteogenesis-related genes’ upregulation [19,32]; osteogenic potential similar to Y201 [19] | Positive for ABS, GAG assay and Sox9 upregulation in pellet culture; chondrogenic potential similar to Y201 [19] | Positive for OROS and upregulation of adipogenesis-related genes; adipogenic potential reduced compared to Y201 [19] |
| MSOD | Positive for ARS [15], upregulation of osteogenesis-related genes [15,26], and in vivo bone formation [15] | Weak positivity for ABS and upregulation of ColX but not ColII nor Sox9, similarly to primary parental cells; tested in pellet culture [15] | Positive for OROS and PPARγ upregulation [15] |
| ASC/TERT1 | Positive for VKS and ALP activity [84] | Positive for ABS, trichrome staining and ColII immunostaining in 3D scaffolds; reduced cartilage quality in comparison with chondrocytes [22] | Positive for OROS and PPARγ upregulation; adipogenic potential increased compared with primary cells [84] |
| hASCs-T | Positive for APS; reduced osteogenic potential in comparison with primary cells [85] | Not tested | Tested by OROS, but almost no lipid droplets detected [85] |
| GB/hTERT MSCs | Positive for ARS; reduced compared with primary cells [86] | Not tested | Positive for OROS [86] |
| SDP11 | Positive for BMP-2 and ALP upregulation [88] | Not tested | Positive for OROS but not shown [88] |
| Pelt cells | Positive for ARS (slightly reduced compared with primary cells) [90] and cementogenesis-related gene expression [31] | Not tested | Not tested |
| CMSC29 | Positive for ARS [91] | Positive for ABS in pellet culture [91,92] | Very weak positivity for OROS [91] |
| DMSC23 | Positive for ARS (increased compared with CMSC29) [91,93] | Positive for ABS in pellet culture [91,92] | Very weak positivity for OROS [91] |
| CPC531 | Positive for APS and upregulation of osteogenesis-related genes [95] | Spontaneous chondrogenesis in 3D alginate culture, proved by upregulation of ColII and Sox9 and downregulation of Runx2 and ColI [95] | Positive for OROS and upregulation of adipogenesis-related genes [95] |
| hASCs-TS (same parental cells as hASCs-T) | Tested by APS, but no mineralization detected [85] | Not tested | Positive for OROS; reduced adipogenic potential in comparison with primary cells [85] |
| 3A6 (KP-derived) | Positive for ARS and VKS (increased compared with KP) [55], and also ALP activity [39] | Positive for ABS [55] and ColII upregulation [56] in pellet culture | Positive for OROS (reduced compared with KP) [55] |
| hASCs-TE (same parental cells as hASCs-T) | Positive for APS; increased in comparison with primary cells [85] | Not tested | Positive for OROS; slightly reduced in comparison with primary cells [85] |
| UE6E7T-3 (same parental cells as UE6E7-16) | Positive for ALP activity [97], ARS and upregulation of osteogenesis-related genes [28] | Not tested/shown | Positive for OROS [97] |
| UE6E7T-11 (same parental cells as UE6E7-16) | Positive for APS and bone sialoprotein (BSP) upregulation [98] | Not tested/shown | Not tested/shown |
| UE6E7T-2 (same parental cells as UE6E7-16) | Not tested/shown | Tested by ABS in 2D culture; negative under employed conditions [99] | Not tested/shown |
| imhMSCs | Positive for VKS and upregulation of osteogenesis-related genes [18] | Weak positivity for ABS and with weak upregulation of chondrogenesis-related genes (similarly to primary parental cells); tested in pellet culture [18] | Positive for OROS and PPARγ upregulation [18] |
| 3 Hits hMPC | Positive for ARS [102,103], APS [101,102] and Runx2 upregulation [102] | Positive for ABS [102,103] and TBS [101] in pellet culture, but reduced compared with primary MSCs | Positive for OROS [101,102,103], but reduced compared with primary MSCs |
| UE7T-13 (same parental cells as UE6E7-16) | Positive for ARS [37,88,105] and ALP activity [88] | Not tested/shown | Positive for OROS [37,88] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Piñeiro-Ramil, M.; Sanjurjo-Rodríguez, C.; Castro-Viñuelas, R.; Rodríguez-Fernández, S.; Fuentes-Boquete, I.M.; Blanco, F.J.; Díaz-Prado, S.M. Usefulness of Mesenchymal Cell Lines for Bone and Cartilage Regeneration Research. Int. J. Mol. Sci. 2019, 20, 6286. https://doi.org/10.3390/ijms20246286
Piñeiro-Ramil M, Sanjurjo-Rodríguez C, Castro-Viñuelas R, Rodríguez-Fernández S, Fuentes-Boquete IM, Blanco FJ, Díaz-Prado SM. Usefulness of Mesenchymal Cell Lines for Bone and Cartilage Regeneration Research. International Journal of Molecular Sciences. 2019; 20(24):6286. https://doi.org/10.3390/ijms20246286
Chicago/Turabian StylePiñeiro-Ramil, M., C. Sanjurjo-Rodríguez, R. Castro-Viñuelas, S. Rodríguez-Fernández, I.M. Fuentes-Boquete, F.J. Blanco, and S.M. Díaz-Prado. 2019. "Usefulness of Mesenchymal Cell Lines for Bone and Cartilage Regeneration Research" International Journal of Molecular Sciences 20, no. 24: 6286. https://doi.org/10.3390/ijms20246286
APA StylePiñeiro-Ramil, M., Sanjurjo-Rodríguez, C., Castro-Viñuelas, R., Rodríguez-Fernández, S., Fuentes-Boquete, I. M., Blanco, F. J., & Díaz-Prado, S. M. (2019). Usefulness of Mesenchymal Cell Lines for Bone and Cartilage Regeneration Research. International Journal of Molecular Sciences, 20(24), 6286. https://doi.org/10.3390/ijms20246286






