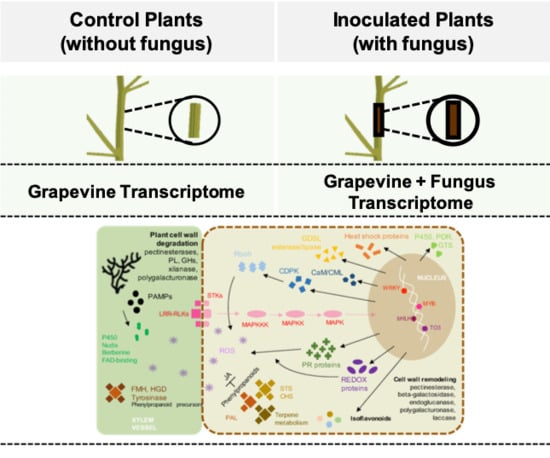
Dual RNA Sequencing of Vitis vinifera during Lasiodiplodia theobromae Infection Unveils Host–Pathogen Interactions
Abstract
1. Introduction
2. Results
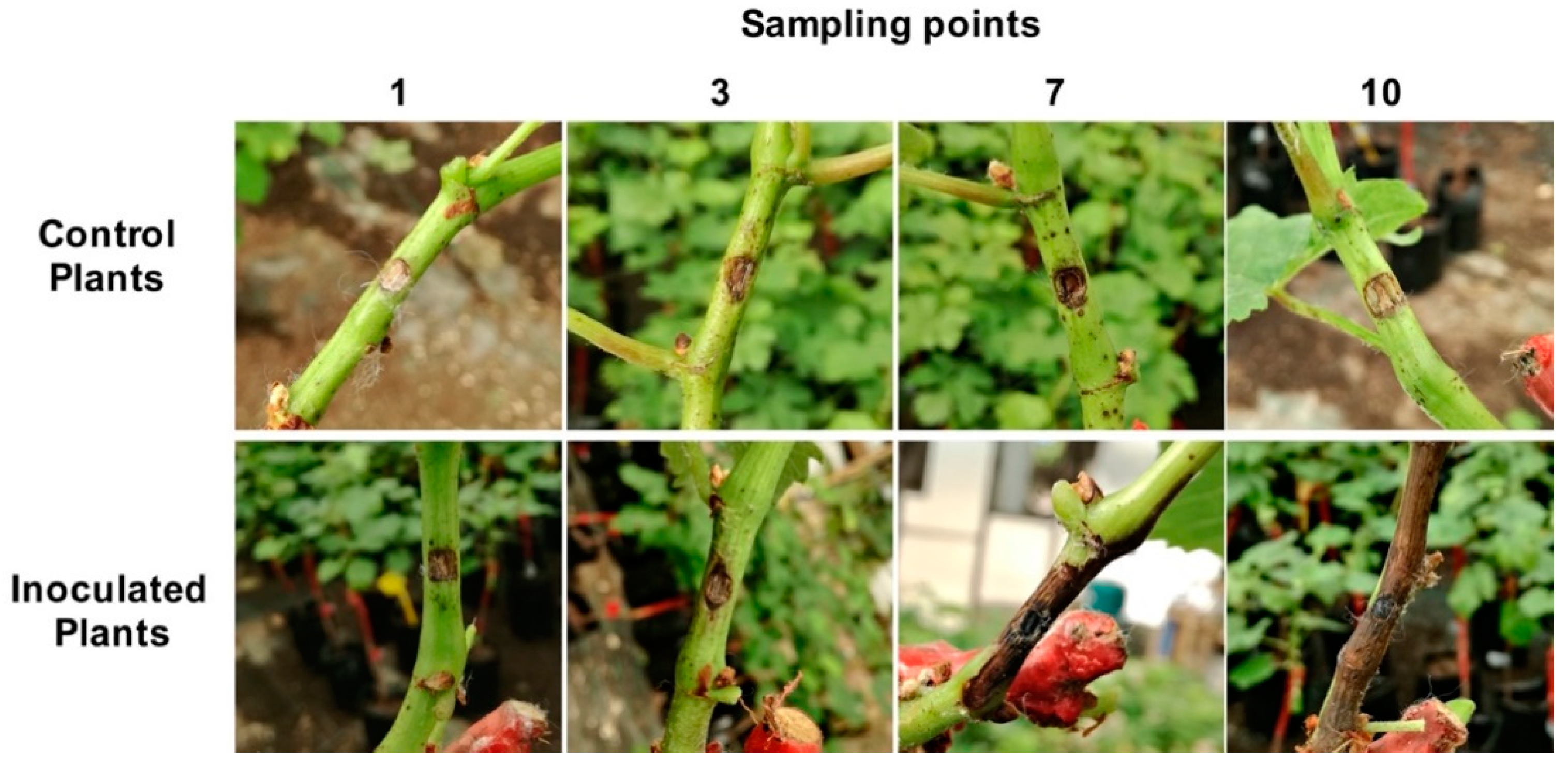
2.1. Macroscopic Analysis
2.2. Dual RNA-Seq Analysis
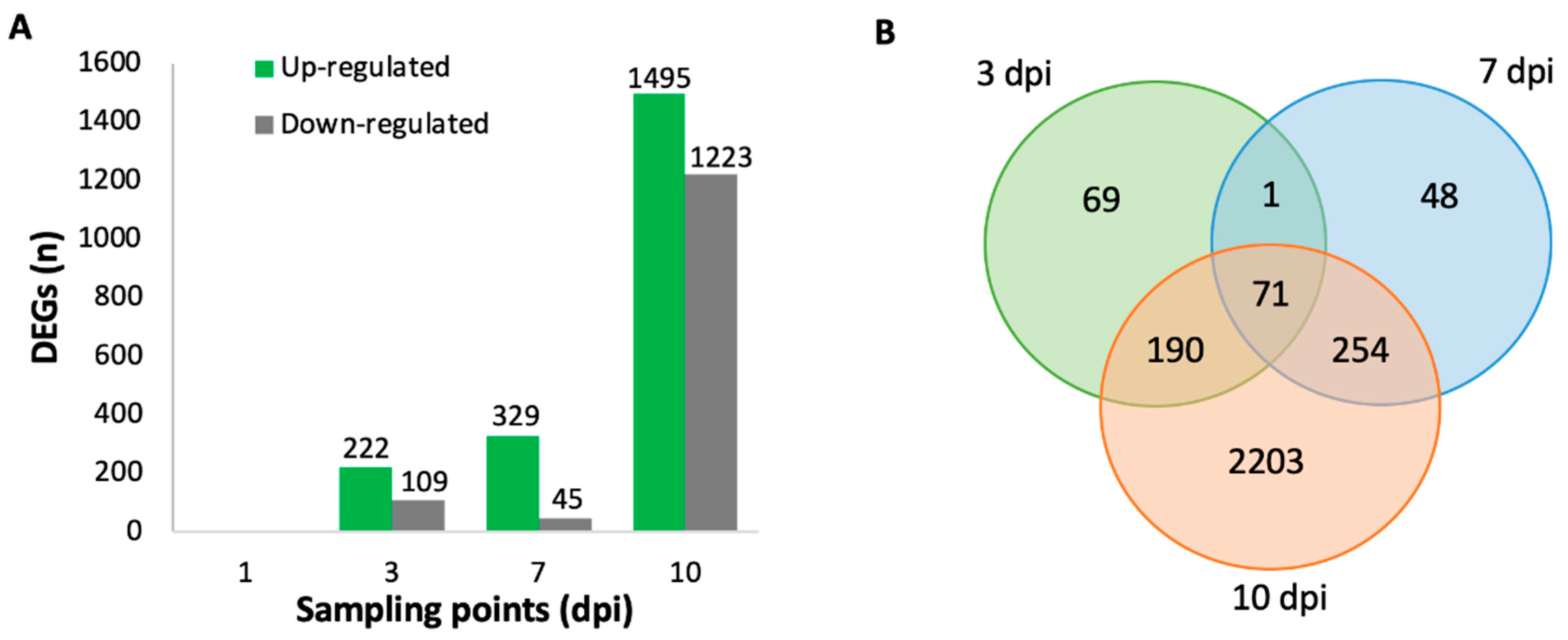
2.3. Differentially Expressed Genes (DEGs) in V. vinifera
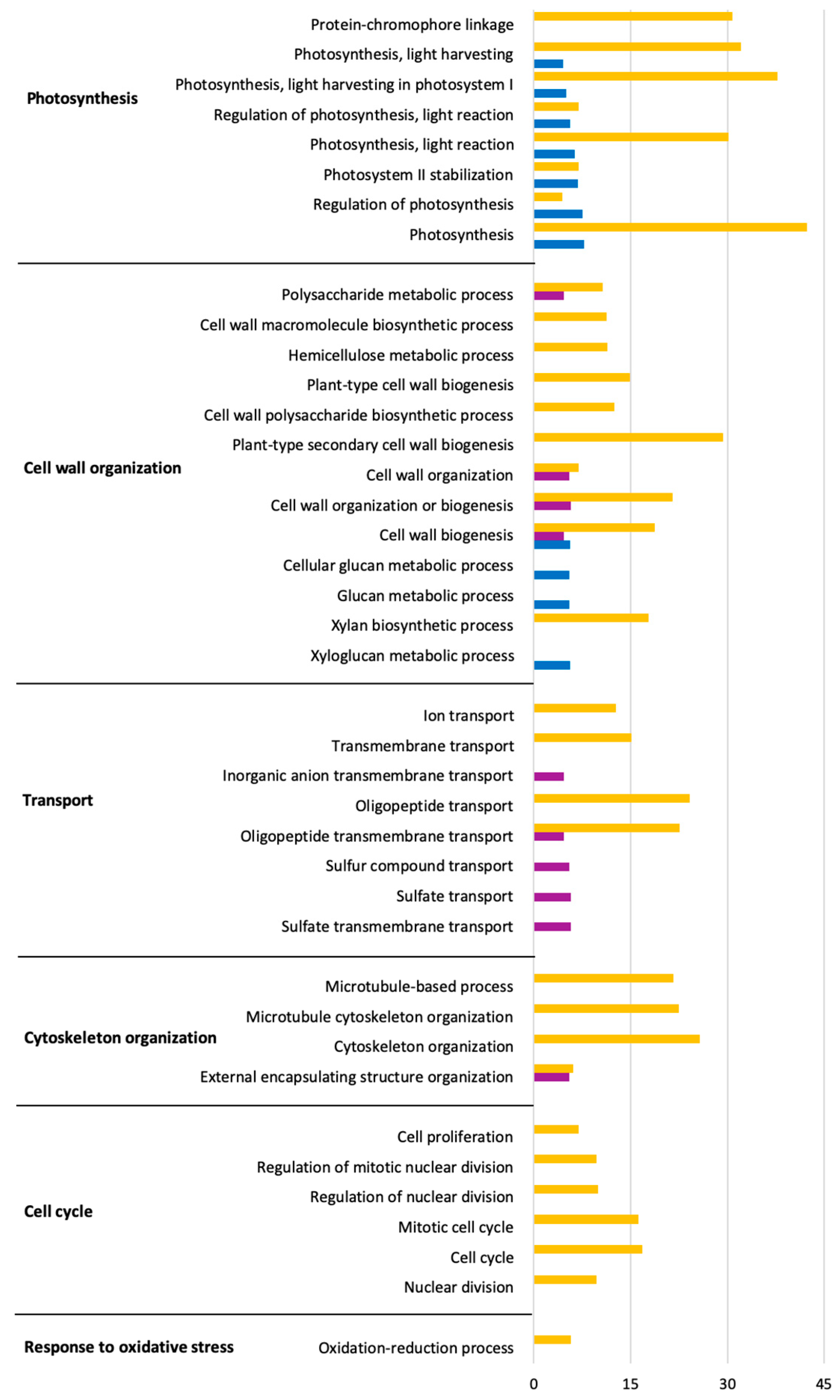
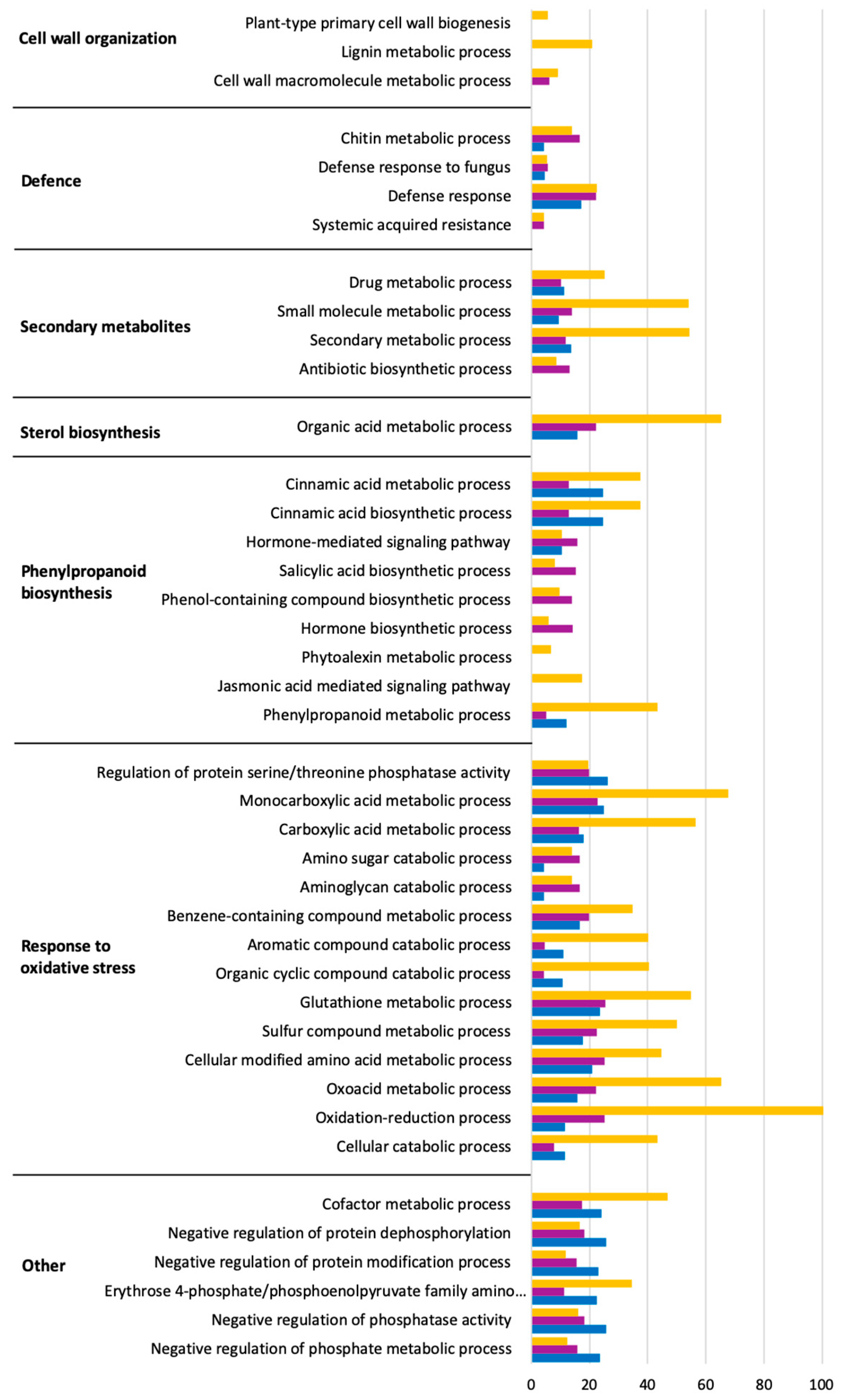
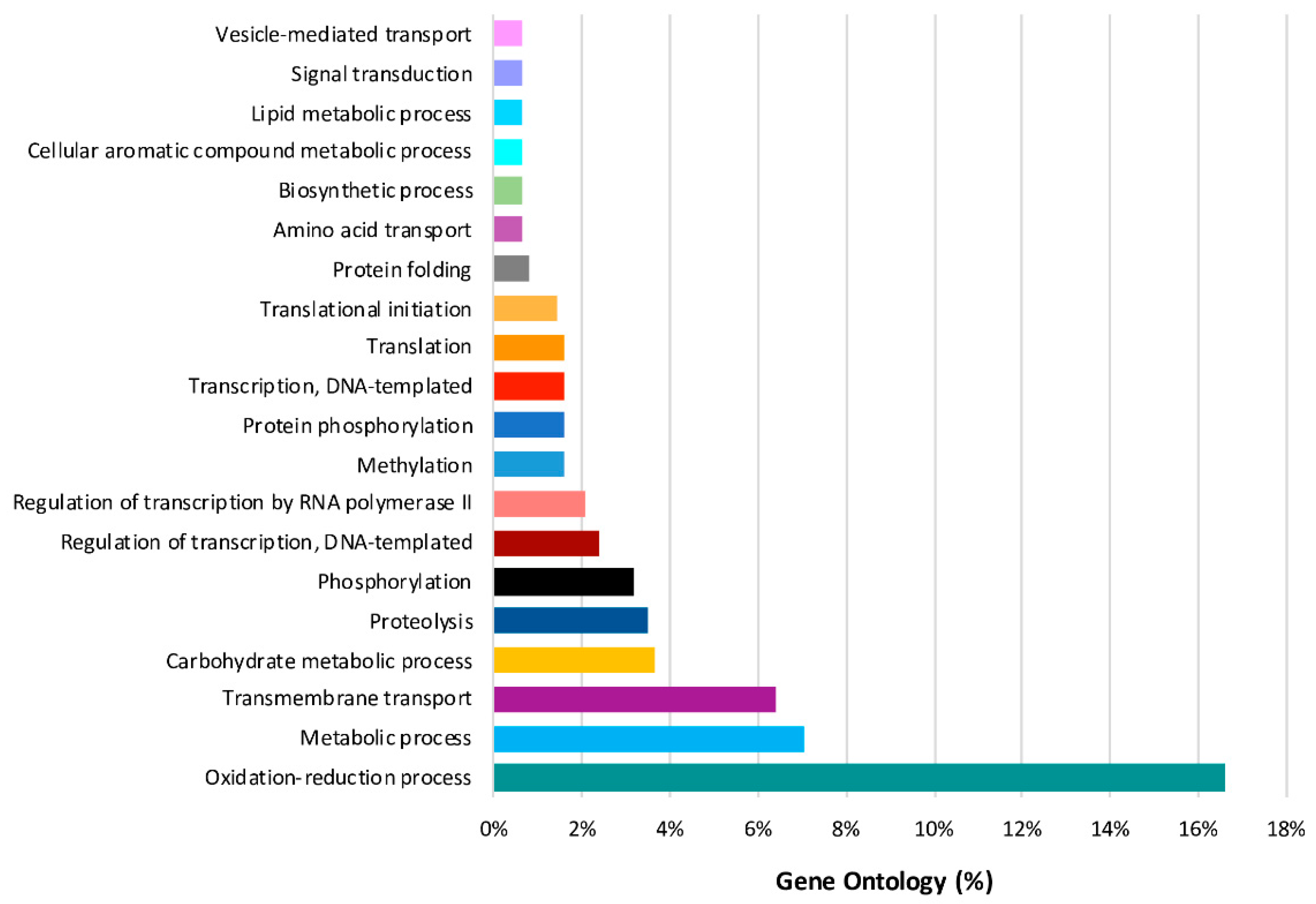
2.4. Predominant Functions (GO) of V. vinifera DEGs during L. theobromae Infection
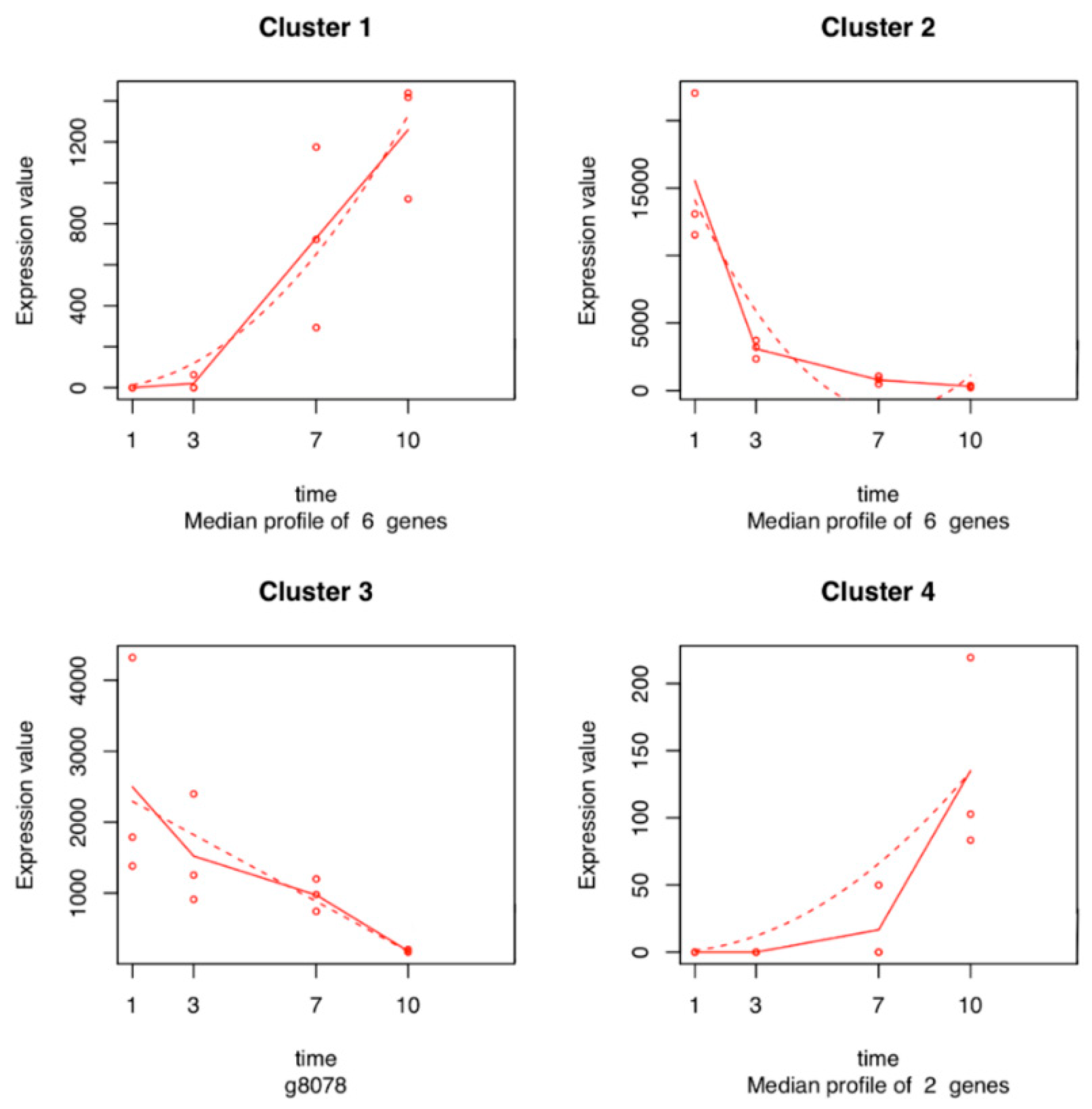
2.5. Time Series Analysis of V. vinifera Genes
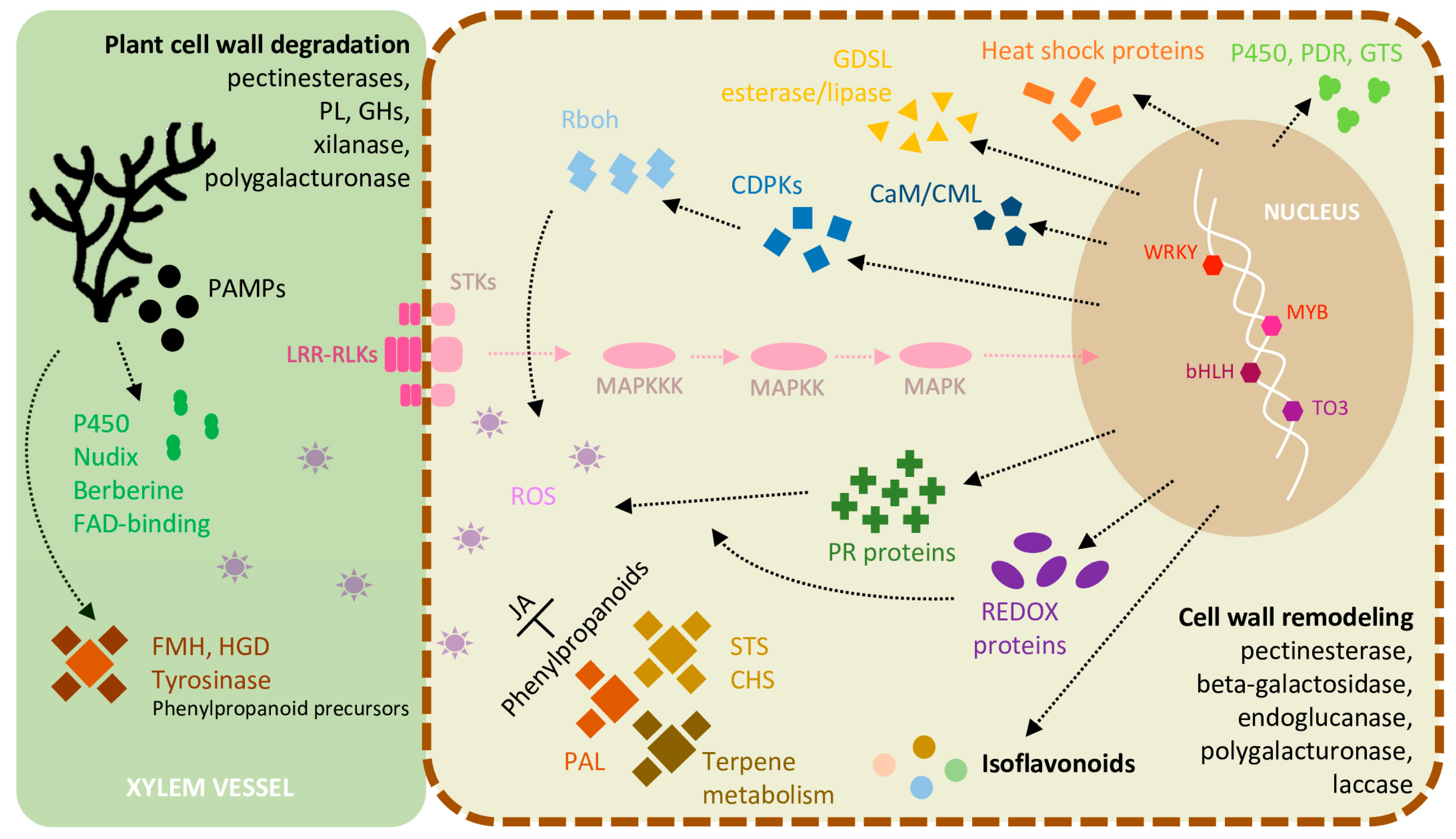
2.6. Inducible Defence Mechanisms during L. theobromae Infection
2.7. DEGs Involved in ET and JA Biosynthesis and Ca2+ Signaling in the Inoculated Green Shoots
2.8. Predominant GO Functions of L. theobromae DEGs in the Time Series Analysis
3. Discussion
4. Materials and Methods
4.1. Biological Material and Experimental Design
4.2. Sample collection
4.3. RNA Extraction, Library Preparation, and Sequencing
4.4. Pairwise Comparison of the Samples Based on V. vinifera Genes
4.5. Time Series Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| dpi | Days post-inoculation |
| hpi | Hours post-inoculation |
| GO | Gene Ontology |
| DEGs | Differentially expressed genes |
| PR | Pathogenesis related |
| PRRs | Pattern recognition receptors |
| GSTs | Glutathione transferases |
| PDR | Pleiotropic drug resistance |
| TLPs | Thaumatin-like proteins |
| STS | Stilbene synthase |
| CHS | Chalcone synthase |
| PAL | Phenylalanine ammonia lyase |
| ET/JA | Ethylene/Jasmonic acid |
| CDPKs | Ca2+-dependent protein kinases |
| FAD | Flavin adenine dinucleotide |
| CaM/CML | Calmodulin/Calmodulin-like transcripts |
| CTAB | Cetyltrimethyl ammonium bromide |
References
- Úrbez-Torres, J.R.; Gubler, W.D. Pathogenicity of Botryosphaeriaceae species isolated from grapevine cankers in California. Plant Dis. 2009, 93, 584–592. [Google Scholar] [CrossRef] [PubMed]
- Úrbez-Torres, J.R. The status of Botryosphaeriaceae species infecting grapevines. Phytopathol. Mediterr. 2011, 50, 5–45. [Google Scholar]
- Bertsch, C.; Ramírez-Suero, M.; Magnin-Robert, M.; Larignon, P.; Chong, J.; Abou-Mansour, E.; Spagnolo, A.; Clément, C.; Fontaine, F. Grapevine trunk diseases: Complex and still poorly understood. Plant Pathol. 2013, 62, 243–265. [Google Scholar] [CrossRef]
- Kaplan, J.; Travadon, R.; Cooper, M.; Hillis, V.; Lubell, M.; Baumgartner, K. Identifying economic hurdles to early adoption of preventative practices: The case of trunk diseases in California winegrape vineyards. Wine Econ. Pol. 2016, 5, 127–141. [Google Scholar] [CrossRef]
- Slippers, B.; Wingfield, M.J. Botryosphaeriaceae as endophytes and latent pathogens of woody plants: Diversity, ecology and impact. Fungal Biol. Rev. 2007, 21, 90–106. [Google Scholar] [CrossRef]
- Eastburn, D.M.; McElrone, A.J.; Bilgin, D.D. Influence of atmospheric and climatic change on plant-pathogen interactions. Plant Pathol. 2011, 60, 54–69. [Google Scholar] [CrossRef]
- Gallana, M.; Ryser-Degiorgis, M.P.; Wahli, T.; Segner, H. Climate change and infectious diseases of wildlife: Altered interactions between pathogens, vectors and hosts. Curr. Zool. 2013, 59, 427–437. [Google Scholar] [CrossRef]
- King, B.C.; Waxman, K.D.; Nenni, N.V.; Walker, L.P.; Bergstrom, G.C.; Gibson, D.M. Arsenal of plant cell wall degrading enzymes reflects host preference among plant pathogenic fungi. Biotechnol. Biofuels 2011, 4, 4. [Google Scholar] [CrossRef]
- Taylor, A.; Hardy, G.E.S.J.; Wood, P.; Burgess, T. Identification and pathogenicity of Botryosphaeria species associated with grapevine decline in Western Australia. Australas. Plant Pathol. 2005, 34, 187–195. [Google Scholar]
- Úrbez-Torres, J.R.; Leavitt, G.M.; Guerrero, J.C.; Guevara, J.; Gubler, W.D. Identification and pathogenicity of Lasiodiplodia theobromae and Diplodia seriata, the causal agents of bot canker disease of grapevines in Mexico. Plant Dis. 2008, 92, 519–529. [Google Scholar] [CrossRef]
- Rodríguez-Gálvez, E.; Maldonado, E.; Alves, A. Identification and pathogenicity of Lasiodiplodia theobromae causing dieback of table grapes in Peru. Eur. J. Plant Pathol. 2015, 141, 477–489. [Google Scholar] [CrossRef]
- Alves, A.; Crous, P.W.; Correia, A.; Phillips, A.J.L. Morphological and molecular data reveal cryptic speciation in Lasiodiplodia theobromae. Fungal Divers. 2008, 28, 1–13. [Google Scholar]
- Phillips, A.J.L.; Alves, A.; Abdollahzadeh, J.; Slippers, B.; Wingfield, M.J.; Groenewald, J.Z.; Crous, P.W. The Botryosphaeriaceae: Genera and species known from culture. Stud. Mycol. 2013, 76, 51–167. [Google Scholar] [CrossRef] [PubMed]
- Félix, C.; Duarte, A.S.; Vitorino, R.; Guerreiro, A.C.; Domingues, P.; Correia, A.; Alves, A.; Esteves, A.C. Temperature modulates the secretome of the phytopathogenic fungus Lasiodiplodia theobromae. Front. Plant Sci. 2016, 7, 1096. [Google Scholar] [CrossRef]
- Correia, K.; Câmara, M.P.S.; Barbosa, M.A.G.; Sales, R., Jr.; Agustí-Brisach, C.; Gramaje, D.; León, M.; Garcia-Jiménez, J.; Abad-Campos, P.; Armengol, J.; et al. Fungal trunk pathogens associated with table grape decline in Northeastern Brazil. Phytopathol. Mediterr. 2013, 52, 380–387. [Google Scholar]
- Aroca, A.; Raposo, R.; Gramaje, D.; Armengol, J.; Martos, S.; Luque, J. First report of Lasiodiplodia theobromae associated with decline of grapevine rootstock mother plants in Spain. Plant Dis. 2008, 92, 832. [Google Scholar] [CrossRef]
- Burruano, S.; Mondello, V.; Conigliaro, G.; Alfonzo, A.; Spagnolo, A.; Mugnai, L. Grapevine decline in Italy caused by Lasiodiplodia theobromae. Phytopathol. Mediterr. 2008, 47, 132–136. [Google Scholar]
- Úrbez-Torres, J.R.; Leavitt, G.M.; Voegel, T.M.; Gubler, W.D. Identification and distribution of Botryosphaeria spp. associated with grapevine cankers in California. Plant Dis. 2006, 90, 1490–1503. [Google Scholar] [CrossRef]
- Naidoo, S.; Visser, E.A.; Zwart, L.; Du Toit, Y.; Bhadauria, V.; Shuey, L.S. Dual RNA-seq to elucidate the plant– pathogen duel. Curr. Issues Mol. Biol. 2017, 27, 127–142. [Google Scholar]
- Dou, D.; Zhou, J.M. Phytopathogen effectors subverting host immunity: Different foes, similar battleground. Cell Host Microbe 2012, 12, 484–495. [Google Scholar] [CrossRef]
- Meyer, F.E.; Shuey, L.S.; Naidoo, S.; Mamni, T.; Berger, D.K.; Myburg, A.A.; van den Berg, N.; Naidoo, S. Dual RNA-sequencing of Eucalyptus nitens during Phytophthora cinnamomi challenge reveals pathogen and host factors influencing compatibility. Front. Plant Sci. 2016, 7, 191. [Google Scholar] [CrossRef] [PubMed]
- Westermann, A.J.; Gorski, S.A.; Vogel, J. Dual RNA-seq of pathogen and host. Nat. Rev. Microbiol. 2012, 10, 618. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Yan, J.; Li, X.; Xing, Q.; Chethana, K.T.; Zhao, W. Transcriptional response of grapevine to infection with the fungal pathogen Lasiodiplodia theobromae. Sci. Rep. 2019, 9, 5387. [Google Scholar] [CrossRef] [PubMed]
- Massonnet, M.; Figueroa-Balderas, R.; Galarneau, E.R.; Miki, S.; Lawrence, D.P.; Sun, Q.; Wallisc, C.M.; Baumgartner, K.; Cantu, D. Neofusicoccum parvum colonization of the grapevine woody stem triggers asynchronous host responses at the site of infection and in the leaves. Front. Plant Sci. 2017, 8, 1117. [Google Scholar] [CrossRef] [PubMed]
- Lowe, R.G.T.; Cassin, A.; Grandaubert, J.; Clark, B.L.; Van de Wouw, A.P.; Rouxel, T.; Howlett, B.J. Genomes and transcriptomes of partners in plant- fungal- interactions between canola (Brassica napus) and two Leptosphaeria species. PLoS ONE 2014, 9, e103098. [Google Scholar] [CrossRef]
- Yazawa, T.; Kawahigashi, H.; Matsumoto, T.; Mizuno, H. Simultaneous transcriptome analysis of sorghum and Bipolaris sorghicola by using RNA-seq in combination with de novo transcriptome assembly. PLoS ONE 2013, 8, e62460. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Wang, Y.; Li, Z.; Zhang, H.; Ye, J.; Li, G. Gene expression changes during the gummosis development of peach shoots in response to Lasiodiplodia theobromae infection using RNA-Seq. Front. Physiol. 2016, 7, 170. [Google Scholar] [CrossRef]
- Teixeira, P.J.; Thomazella, D.P.; Reis, O.; do Prado, P.F.V.; do Rio, M.C.S.; Fiorin, G.L.; José, J.; Costa, G.G.L.; Negri, V.A.; Mondego, J.M.C.; et al. High-resolution transcript profiling of the atypical biotrophic interaction between Theobroma cacao and the fungal pathogen Moniliophthora perniciosa. Plant Cell 2014, 26, 4245–4269. [Google Scholar] [CrossRef]
- Enault, F.; Suhre, K.; Claverie, J.M. Phydbac “Gene Function Predictor”: A gene annotation tool based on genomic context analysis. BMC Bioinform. 2005, 6, 247. [Google Scholar] [CrossRef]
- Singh, H.; Rai, K.M.; Upadhyay, S.K.; Pant, P.; Verma, P.C.; Singh, A.P.; Singh, P.K. Transcriptome sequencing of a thalloid bryophyte; Dumortiera hirsuta (Sw) Nees: Assembly, annotation, and marker discovery. Sci. Rep. 2015, 5, 15350. [Google Scholar] [CrossRef]
- Rasmussen, M.W.; Roux, M.; Petersen, M.; Mundy, J. MAP kinase cascades in Arabidopsis innate immunity. Front. Plant Sci. 2012, 3, 169. [Google Scholar] [CrossRef] [PubMed]
- Reignault, P.; Valette-Collet, O.; Boccara, M. The importance of fungal pectinolytic enzymes in plant invasion, host adaptability and symptom type. Eur. J. Plant Pathol. 2008, 120, 1–11. [Google Scholar] [CrossRef]
- Kubicek, C.P.; Starr, T.L.; Glass, N.L. Plant cell wall–degrading enzymes and their secretion in plant-pathogenic fungi. Annu. Rev. Phytopathol. 2014, 52, 427–451. [Google Scholar] [CrossRef] [PubMed]
- Alves da Cunha, M.A.; Barbosa, A.M.; Giese, E.C.; Dekker, R.F. The effect of carbohydrate carbon sources on the production of constitutive and inducible laccases by Botryosphaeria sp. J. Basic Microbiol. 2003, 43, 385–392. [Google Scholar] [CrossRef]
- Félix, C.; Libório, S.; Nunes, M.; Félix, R.; Duarte, A.; Alves, A.; Esteves, A. Lasiodiplodia theobromae as a producer of biotechnologically relevant enzymes. Int. J. Mol. Sci. 2018, 19, 29. [Google Scholar] [CrossRef]
- Morales-Cruz, A.; Amrine, K.C.; Blanco-Ulate, B.; Lawrence, D.P.; Travadon, R.; Rolshausen, P.E.; Baumgartner, K.; Cantu, D. Distinctive expansion of gene families associated with plant cell wall degradation, secondary metabolism, and nutrient uptake in the genomes of grapevine trunk pathogens. BMC Genom. 2015, 16, 469. [Google Scholar] [CrossRef]
- Črešnar, B.; Petrič, Š. Cytochrome P450 enzymes in the fungal kingdom. Biochim. Biophys. Acta 2011, 1814, 29–35. [Google Scholar] [CrossRef]
- Nierman, W.C.; Pain, A.; Anderson, M.J.; Wortman, J.R.; Kim, H.S.; Arroyo, J.; Bennett, J.; Bowyer, P.; Chen, D.; Collins, M.; et al. Genomic sequence of the pathogenic and allergenic filamentous fungus Aspergillus fumigatus. Nature 2005, 438, 1151. [Google Scholar] [CrossRef]
- Eswaramoorthy, S.; Bonanno, J.B.; Burley, S.K.; Swaminathan, S. Mechanism of action of a flavin-containing monooxygenase. Proc. Natl. Acad. Sci. USA 2006, 103, 9832–9837. [Google Scholar] [CrossRef]
- Daniel, B.; Konrad, B.; Toplak, M.; Lahham, M.; Messenlehner, J.; Winkler, A.; Macheroux, P. The family of berberine bridge enzyme-like enzymes: A treasure-trove of oxidative reactions. Arch. Biochem. Biophys. 2017, 632, 88–103. [Google Scholar] [CrossRef]
- Cregut, M.; Piutti, S.; Slezack-Deschaumes, S.; Benizri, E. Compartmentalization and regulation of arylsulfatase activities in Streptomyces sp., Microbacterium sp. and Rhodococcus sp. soil isolates in response to inorganic sulfate limitation. Microbiol. Res. 2013, 168, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Paolinelli-Alfonso, M.; Villalobos-Escobedo, J.M.; Rolshausen, P.; Herrera-Estrella, A.; Galindo-Sánchez, C.; López-Hernández, J.F.; Hernandez-Martinez, R. Global transcriptional analysis suggests Lasiodiplodia theobromae pathogenicity factors involved in modulation of grapevine defensive response. BMC Genom. 2016, 17, 615. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Wang, Y. Nudix Effectors: A common weapon in the arsenal of plant pathogens. PLoS Pathog. 2016, 12, e1005704. [Google Scholar] [CrossRef] [PubMed]
- Félix, C.; Meneses, R.; Gonçalves, M.F.; Tilleman, L.; Duarte, A.S.; Jorrín-Novo, J.V.; Van de Peer, Y.; Deforce, D.; Van Nieuwerburgh, F.; Esteves, A.C.; et al. A multi-omics analysis of the grapevine pathogen Lasiodiplodia theobromae reveals that temperature affects the expression of virulence-and pathogenicity-related genes. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef]
- Chong, J.; Le Henanff, G.; Bertsch, C.; Walter, B. Identification, expression analysis and characterization of defense and signaling genes in Vitis vinifera. Plant Physiol. Biochem. 2008, 46, 469–481. [Google Scholar] [CrossRef]
- Armijo, G.; Schlechter, R.; Agurto, M.; Muñoz, D.; Nuñez, C.; Arce-Johnson, P. Grapevine pathogenic microorganisms: Understanding infection strategies and host response scenarios. Front. Plant Sci. 2016, 7, 382. [Google Scholar] [CrossRef]
- Van Loon, L.C.; Rep, M.; Pieterse, C.M. Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef]
- Brandazza, A.; Angeli, S.; Tegoni, M.; Cambillau, C.; Pelosi, P. Plant stress proteins of the thaumatin-like family discovered in animals. FEBS Lett. 2004, 572, 3–7. [Google Scholar] [CrossRef]
- Jayasankar, S.; Li, Z.; Gray, D.J. Constitutive expression of Vitis vinifera thaumatin-like protein after in vitro selection and its role in anthracnose resistance. Funct. Plant Biol. 2003, 30, 1105–1115. [Google Scholar] [CrossRef]
- Monteiro, S.; Barakat, M.; Piçarra-Pereira, M.A.; Teixeira, A.R.; Ferreira, R.B. Osmotin and thaumatin from grape: A putative general defense mechanism against pathogenic fungi. Phytopathology 2003, 93, 1505–1512. [Google Scholar] [CrossRef]
- Menu-Bouaouiche, L.; Vriet, C.; Peumans, W.J.; Barre, A.; Van Damme, E.J.; Rougé, P. A molecular basis for the endo-β1, 3-glucanase activity of the thaumatin-like proteins from edible fruits. Biochimie 2003, 85, 123–131. [Google Scholar] [CrossRef]
- Derckel, J.P.; Baillieul, F.; Manteau, S.; Audran, J.C.; Haye, B.; Lambert, B.; Legendre, L. Differential induction of grapevine defenses by two strains of Botrytis cinerea. Phytopathology 1999, 89, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Dadakova, K.; Havelkova, M.; Kurkova, B.; Tlolkova, I.; Kasparovsky, T.; Zdrahal, Z.; Lochman, J. Proteome and transcript analysis of Vitis vinifera cell cultures subjected to Botrytis cinerea infection. J. Proteom. 2015, 119, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Giacomelli, L.; Nanni, V.; Lenzi, L.; Zhuang, J.; Serra, M.D.; Banfield, M.J.; Town, C.D.; Silverstein, K.A.T.; Baraldi, E.; Moser, C. Identification and characterization of the defensin-like gene family of grapevine. Mol. Plant Microbe Interact. 2012, 25, 1118–1131. [Google Scholar] [CrossRef]
- Bellée, A.; Comont, G.; Nivault, A.; Abou-Mansour, E.; Coppin, C.; Dufour, M.C.; Corio-Costet, M.F. Life traits of four Botryosphaeriaceae species and molecular responses of different grapevine cultivars or hybrids. Plant Pathol. 2017, 66, 763–776. [Google Scholar] [CrossRef]
- Girault, T.; François, J.; Rogniaux, H.; Pascal, S.; Delrot, S.; Coutos-Thévenot, P.; Gomès, E. Exogenous application of a lipid transfer protein-jasmonic acid complex induces protection of grapevine towards infection by Botrytis cinerea. Plant Physiol. Biochem. 2008, 46, 140–149. [Google Scholar] [CrossRef]
- Thaler, J.S.; Humphrey, P.T.; Whiteman, N.K. Evolution of jasmonate and salicylate signal crosstalk. Trends Plant Sci. 2012, 17, 260–270. [Google Scholar] [CrossRef]
- MacDonald, M.J.; D’Cunha, G.B. A modern view of phenylalanine ammonia lyase. Biochem. Cell Biol. 2007, 85, 273–282. [Google Scholar] [CrossRef]
- Dixon, R.A.; Paiva, N.L. Stress-induced phenylpropanoid metabolism. Plant Cell 1995, 7, 1085. [Google Scholar] [CrossRef]
- Chong, J.; Poutaraud, A.; Hugueney, P. Metabolism and roles of stilbenes in plants. Plant Sci. 2009, 177, 143–155. [Google Scholar] [CrossRef]
- Birkenbihl, R.P.; Diezel, C.; Somssich, I.E. Arabidopsis WRKY33 is a key transcriptional regulator of hormonal and metabolic responses toward Botrytis cinerea infection. Plant Physiol. 2012, 159, 266–285. [Google Scholar] [CrossRef] [PubMed]
- Windram, O.; Madhou, P.; McHattie, S.; Hill, C.; Hickman, R.; Cooke, E.; Jenkins, D.J.; Penfold, C.A.; Baxter, L.; Breeze, E.; et al. Arabidopsis defense against Botrytis cinerea: Chronology and regulation deciphered by high-resolution temporal transcriptomic analysis. Plant Cell 2012, 24, 3530–3557. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, Y.; Kaya, H.; Hiraoka, G.; Yumoto, F.; Kimura, S.; Kadota, Y.; Hishinuma, H.; Senzaki, E.; Yamagoe, S.; Nagata, K.; et al. Synergistic activation of the Arabidopsis NADPH oxidase AtrbohD by Ca2+ and phosphorylation. J. Biol. Chem. 2008, 283, 8885–8892. [Google Scholar] [CrossRef] [PubMed]
- Tena, G.; Boudsocq, M.; Sheen, J. Protein kinase signaling networks in plant innate immunity. Curr. Opin. Plant Biol. 2011, 14, 519–529. [Google Scholar] [CrossRef] [PubMed]
- Boudsocq, M.; Sheen, J. CDPKs in immune and stress signaling. Trends Plant Sci. 2013, 18, 30–40. [Google Scholar] [CrossRef] [PubMed]
- Chiasson, D.; Ekengren, S.K.; Martin, G.B.; Dobney, S.L.; Snedden, W.A. Calmodulin-like proteins from Arabidopsis and tomato are involved in host defense against Pseudomonas syringae pv. tomato. Plant Mol. Biol. 2005, 58, 887–897. [Google Scholar] [CrossRef]
- Mzid, R.; Marchive, C.; Blancard, D.; Deluc, L.; Barrieu, F.; Corio-Costet, M.F.; Drira, D.; Hamdi, S.; Lauvergeat, V. Overexpression of VvWRKY2 in tobacco enhances broad resistance to necrotrophic fungal pathogens. Physiol. Plant 2007, 131, 434–447. [Google Scholar] [CrossRef]
- Guillaumie, S.; Mzid, R.; Méchin, V.; Léon, C.; Hichri, I.; Destrac-Irvine, A.; Trossat-Magnin, C.; Delrot, S.; Lauvergeat, V. The grapevine transcription factor WRKY2 influences the lignin pathway and xylem development in tobacco. Plant Mol. Biol. 2010, 72, 215–234. [Google Scholar] [CrossRef]
- Travadon, R.; Rolshausen, P.E.; Gubler, W.D.; Cadle-Davidson, L.; Baumgartner, K. Susceptibility of cultivated and wild Vitis spp. to wood infection by fungal trunk pathogens. Plant Dis. 2013, 97, 1529–1536. [Google Scholar] [CrossRef]
- Gambino, G.; Perrone, I.; Gribaudo, I. A rapid and effective method for RNA extraction from different tissues of grapevine and other woody plants. Phytochem. Anal. 2008, 19, 520–525. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 2 December 2019).
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Wingett, S.W.; Andrews, S. FastQ Screen: A tool for multi-genome mapping and quality control. F1000Research 2018, 7, 1338. [Google Scholar] [CrossRef] [PubMed]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Nueda, M.J.; Tarazona, S.; Conesa, A. Next maSigPro: Updating maSigPro bioconductor package for RNA-seq time series. Bioinformatics 2014, 30, 2598–2602. [Google Scholar] [CrossRef]
- Zaini, P.A.; Nascimento, R.; Gouran, H.; Cantu, D.; Chakraborty, S.; Phu, M.; Goulart, L.R.; Dandekar, A.M. Molecular profiling of Pierce’s disease outlines the response circuitry of Vitis vinifera to Xylella fastidiosa infection. Front. Plant Sci. 2018, 9, 771. [Google Scholar] [CrossRef]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gonçalves, M.F.M.; Nunes, R.B.; Tilleman, L.; Van de Peer, Y.; Deforce, D.; Van Nieuwerburgh, F.; Esteves, A.C.; Alves, A. Dual RNA Sequencing of Vitis vinifera during Lasiodiplodia theobromae Infection Unveils Host–Pathogen Interactions. Int. J. Mol. Sci. 2019, 20, 6083. https://doi.org/10.3390/ijms20236083
Gonçalves MFM, Nunes RB, Tilleman L, Van de Peer Y, Deforce D, Van Nieuwerburgh F, Esteves AC, Alves A. Dual RNA Sequencing of Vitis vinifera during Lasiodiplodia theobromae Infection Unveils Host–Pathogen Interactions. International Journal of Molecular Sciences. 2019; 20(23):6083. https://doi.org/10.3390/ijms20236083
Chicago/Turabian StyleGonçalves, Micael F. M., Rui B. Nunes, Laurentijn Tilleman, Yves Van de Peer, Dieter Deforce, Filip Van Nieuwerburgh, Ana C. Esteves, and Artur Alves. 2019. "Dual RNA Sequencing of Vitis vinifera during Lasiodiplodia theobromae Infection Unveils Host–Pathogen Interactions" International Journal of Molecular Sciences 20, no. 23: 6083. https://doi.org/10.3390/ijms20236083
APA StyleGonçalves, M. F. M., Nunes, R. B., Tilleman, L., Van de Peer, Y., Deforce, D., Van Nieuwerburgh, F., Esteves, A. C., & Alves, A. (2019). Dual RNA Sequencing of Vitis vinifera during Lasiodiplodia theobromae Infection Unveils Host–Pathogen Interactions. International Journal of Molecular Sciences, 20(23), 6083. https://doi.org/10.3390/ijms20236083