RBM20 Regulates CaV1.2 Surface Expression by Promoting Exon 9* Inclusion of CACNA1C in Neonatal Rat Cardiomyocytes
Abstract
1. Introduction
2. Results
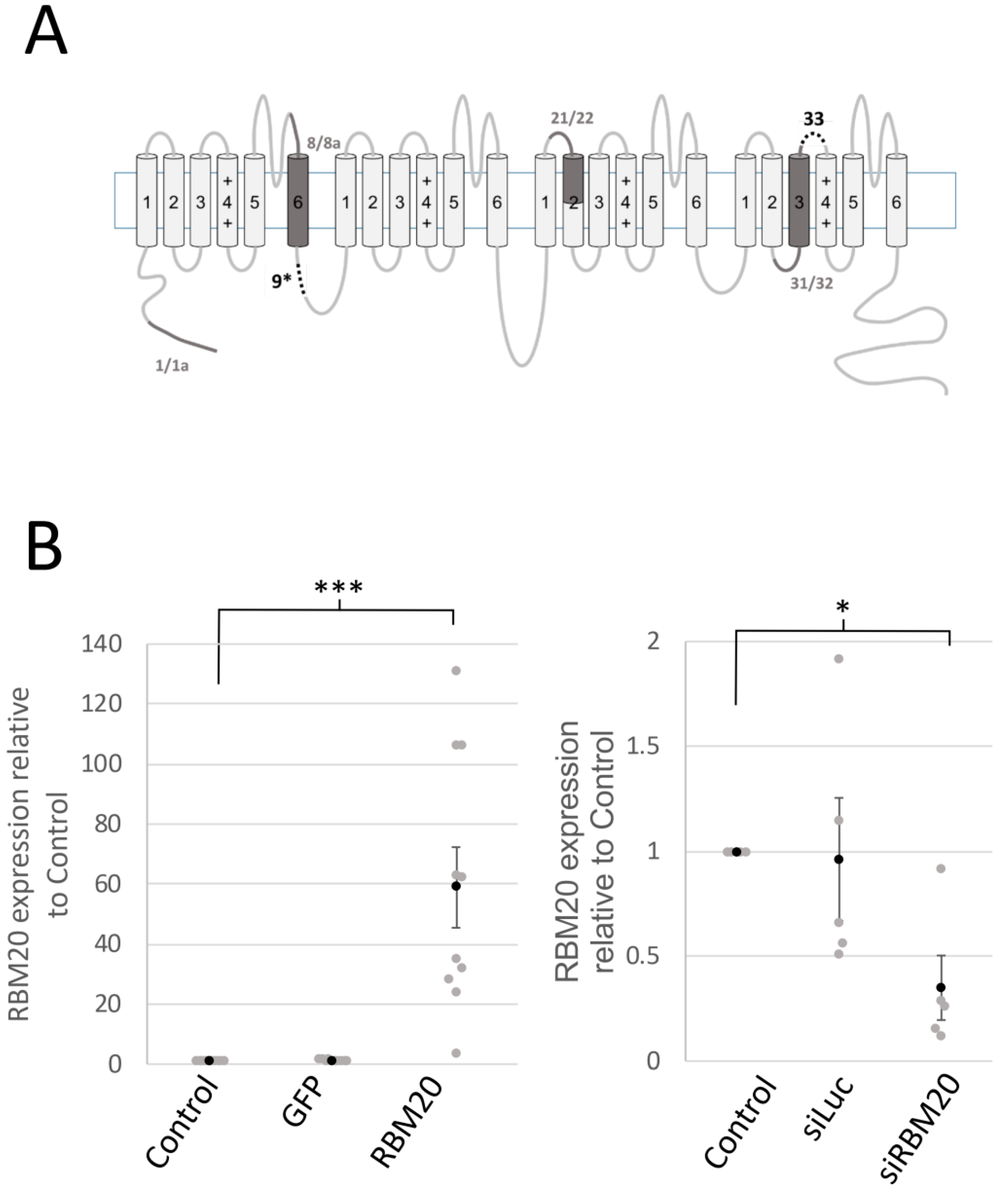
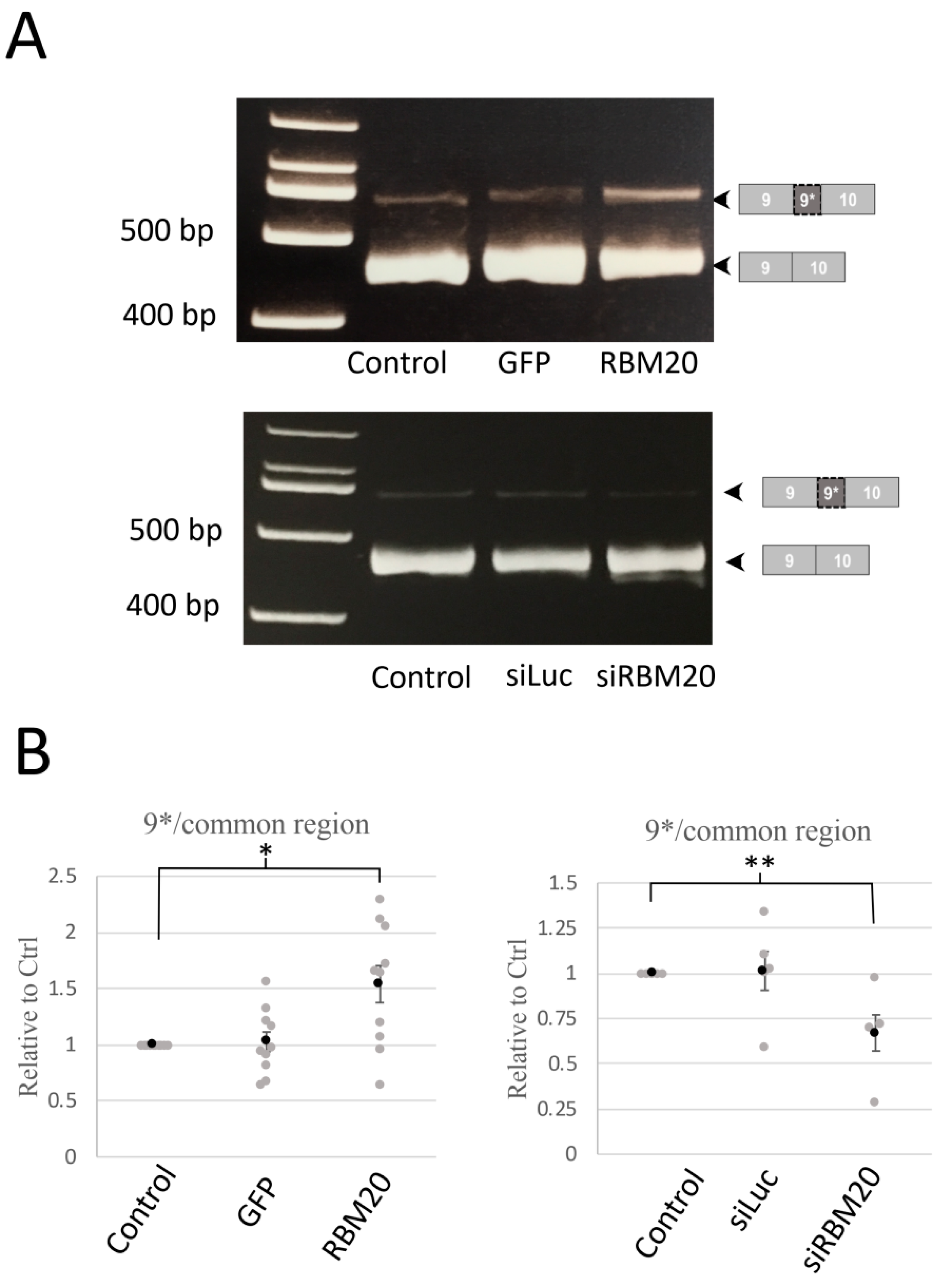
2.1. RBM20 Promotes CACNA1C Exon 9* Inclusion in Cardiomyocytes
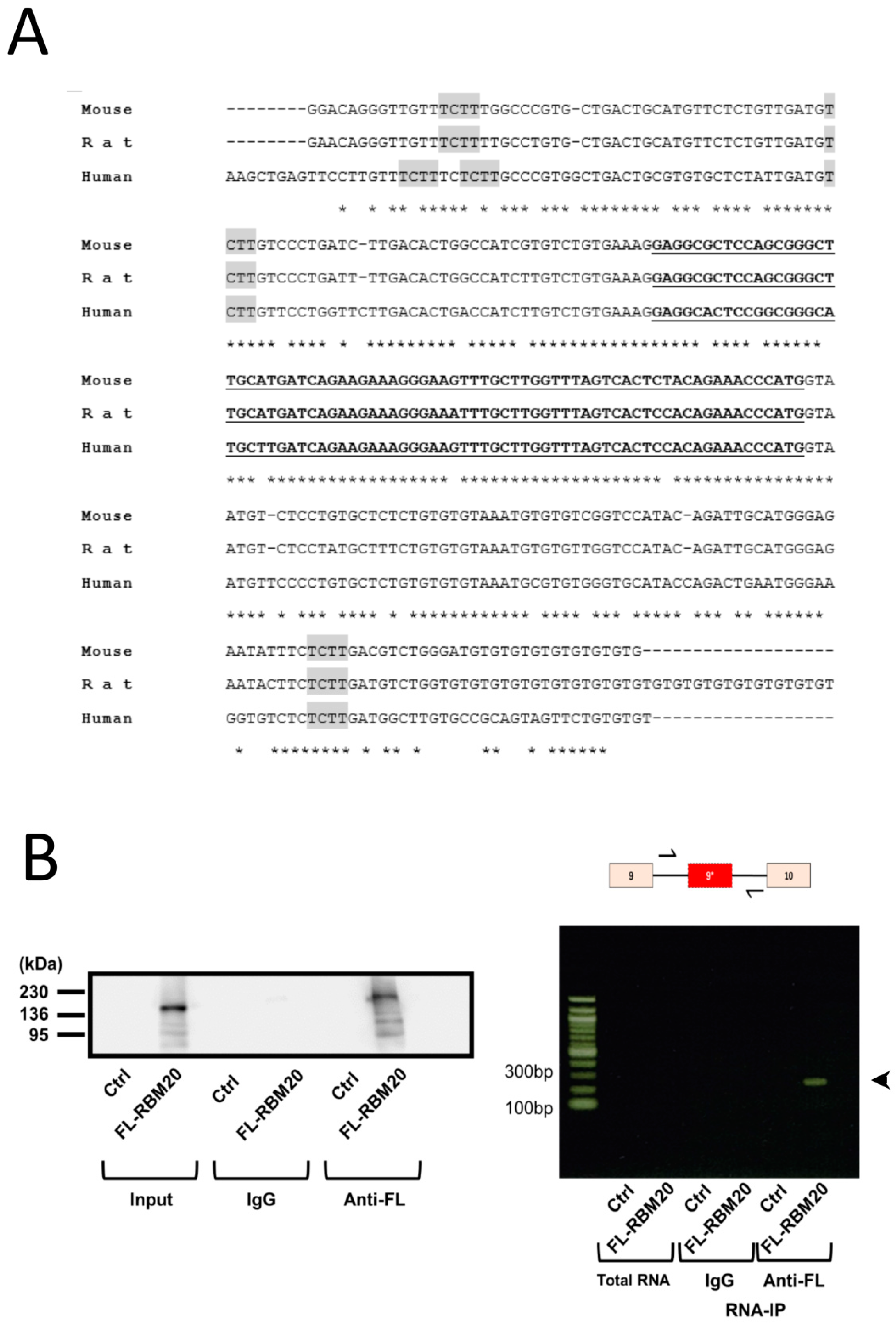
2.2. RBM20 Binds the Intronic Region Surrounding Exon 9*
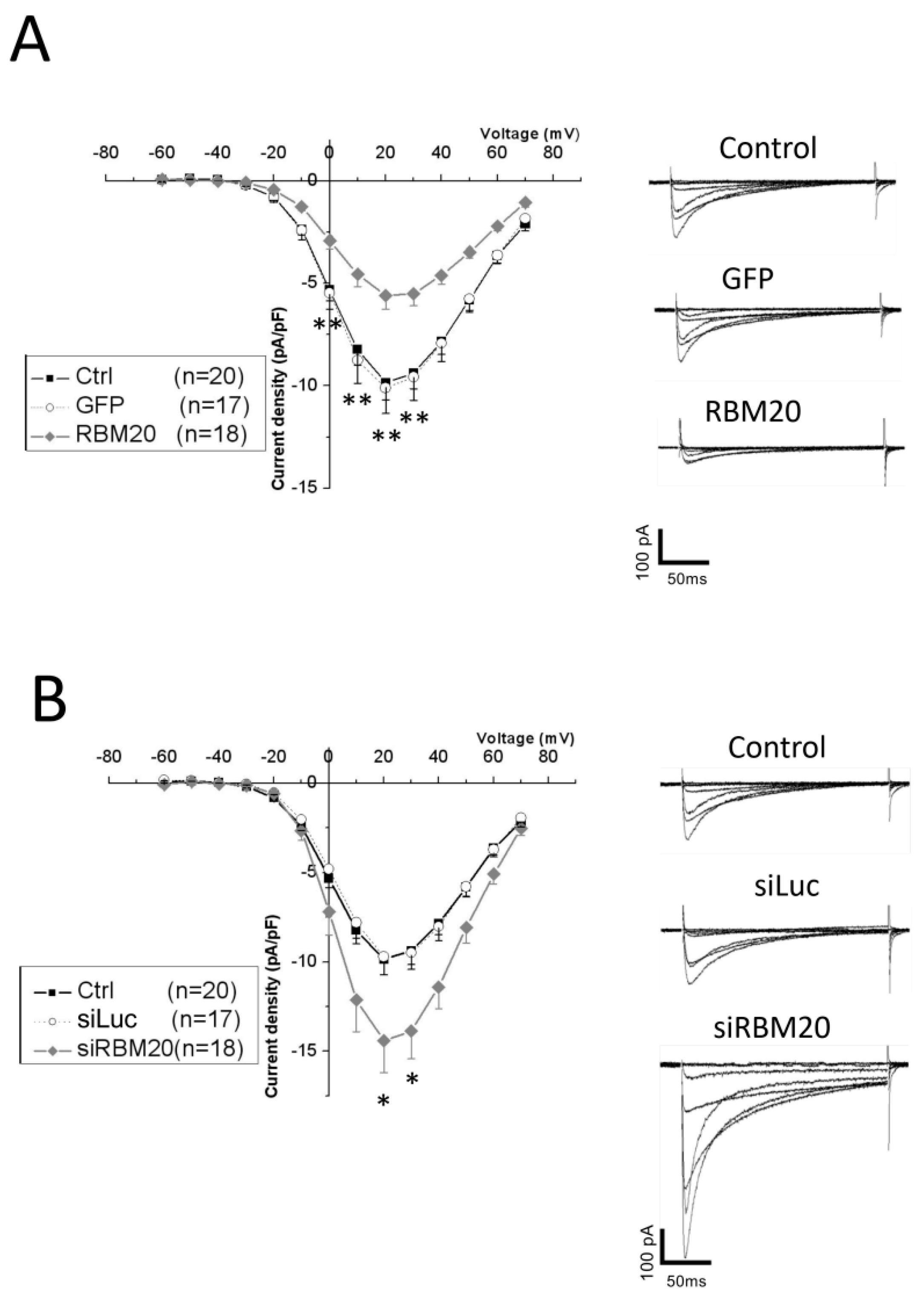
2.3. Manipulation of RBM20 Expression Alters l-tType Ca2+ Currents in Cardiomyocytes
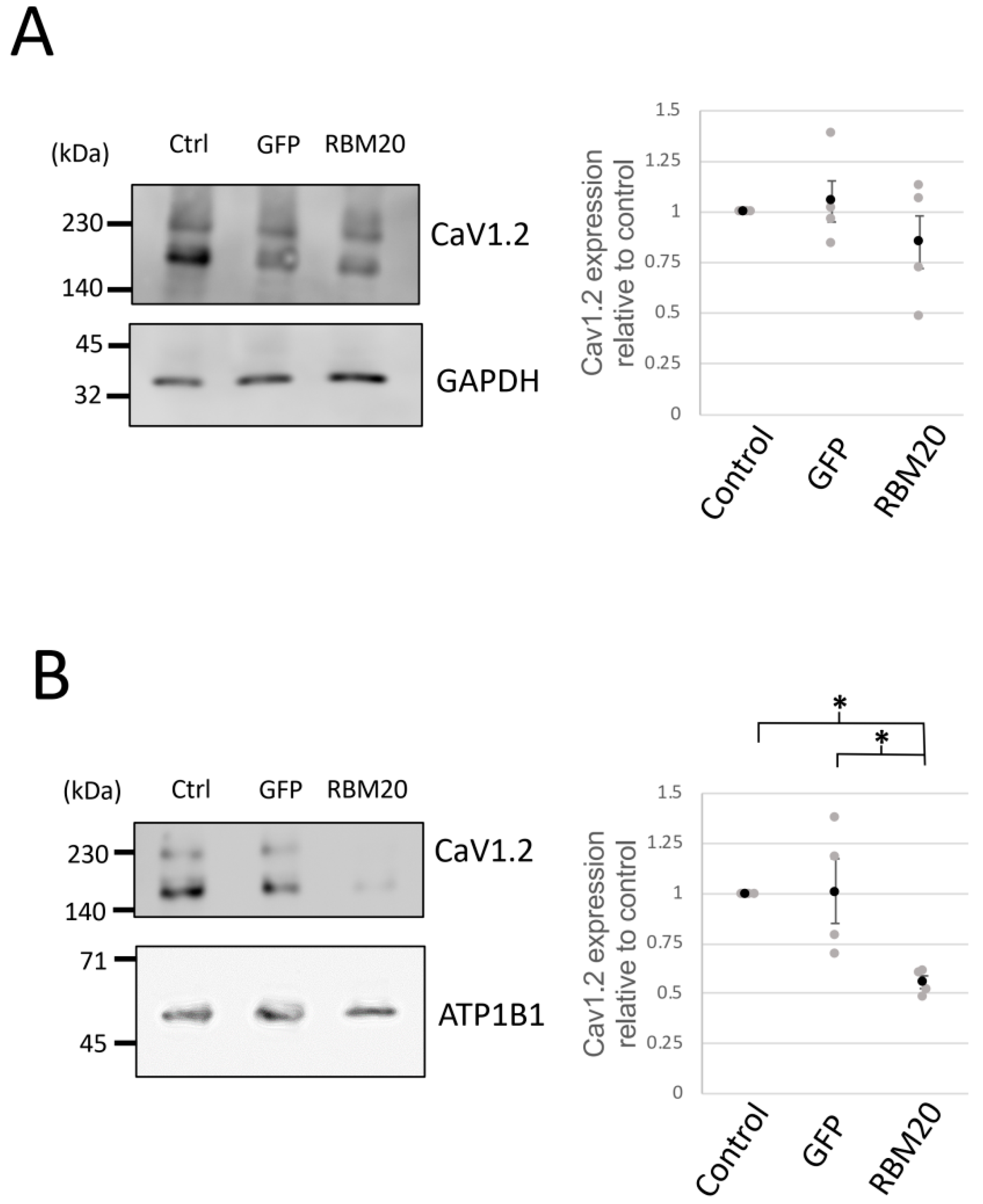
2.4. RBM20 Represses the Membrane Surface Expression of CaV1.2
3. Discussion
4. Materials and Methods
4.1. Experimental Animals
4.2. Cell Culture
4.3. RNA Isolation, Reverse Transcription, PCR, and Electrophoresis
4.4. Real-Time RT-PCR
4.5. Cell-Surface Biotinylation, Protein Isolation, and Western Blot Analysis
4.6. Patch Clamp
4.7. Data Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Zamponi, G.W.; Striessnig, J.; Koschak, A.; Dolphin, A.C. The Physiology, Pathology, and Pharmacology of Voltage-Gated Calcium Channels and Their Future Therapeutic Potential. Pharm. Rev. 2015, 67, 821–870. [Google Scholar] [CrossRef] [PubMed]
- Rougier, J.S.; Abriel, H. Cardiac voltage-gated calcium channel macromolecular complexes. Biochim. Biophys. Acta 2016, 1863, 1806–1812. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Liang, M.C.; Soong, T.W. Alternative Splicing of l-type CaV1.2 Calcium Channels: Implications in Cardiovascular Diseases. Genes (Basel) 2017, 8, 344. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.Z.; Zheng, S.; Nikolic, J.; Black, D.L. Developmental control of CaV1.2 l-type calcium channel splicing by Fox proteins. Mol. Cell. Biol. 2009, 29, 4757–4765. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Fan, J.; Zhu, H.; Ji, L.; Fan, W.; Kapoor, I.; Wang, Y.; Wang, Y.; Zhu, G.; Wang, J. Aberrant Splicing Induced by Dysregulated Rbfox2 Produces Enhanced Function of CaV1.2 Calcium Channel and Vascular Myogenic Tone in Hypertension. Hypertension 2017, 70, 1183–1192. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.Z.; Sharma, S.; Zheng, S.; Chawla, G.; Nikolic, J.; Black, D.L. Regulation of the mutually exclusive exons 8a and 8 in the CaV1.2 calcium channel transcript by polypyrimidine tract-binding protein. J. Biol. Chem. 2011, 286, 10007–10016. [Google Scholar] [CrossRef] [PubMed]
- Rexiati, M.; Sun, M.; Guo, W. Muscle-Specific Mis-Splicing and Heart Disease Exemplified by RBM20. Genes (Basel) 2018, 9, 18. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Kimura, A.; Kuroyanagi, H. Alternative Splicing Regulator RBM20 and Cardiomyopathy. Front. Mol. Biosci. 2018, 5, 105. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Schafer, S.; Greaser, M.L.; Radke, M.H.; Liss, M.; Govindarajan, T.; Maatz, H.; Schulz, H.; Li, S.; Parrish, A.M.; et al. RBM20, a gene for hereditary cardiomyopathy, regulates titin splicing. Nat. Med. 2012, 18, 766–773. [Google Scholar] [CrossRef] [PubMed]
- Beraldi, R.; Li, X.; Martinez-Fernandez, A.; Reyes, S.; Secreto, F.; Terzic, A.; Olson, T.M.; Nelson, T.J. Rbm20-deficient cardiogenesis reveals early disruption of RNA processing and sarcomere remodeling establishing a developmental etiology for dilated cardiomyopathy. Hum. Mol. Genet. 2014, 23, 3779–3791. [Google Scholar] [CrossRef] [PubMed]
- Ito, J.; Iijima, M.; Yoshimoto, N.; Niimi, T.; Kuroda, S.; Maturana, A.D. RBM20 and RBM24 cooperatively promote the expression of short enh splice variants. FEBS Lett. 2016, 590, 2262–2274. [Google Scholar] [CrossRef] [PubMed]
- Maatz, H.; Jens, M.; Liss, M.; Schafer, S.; Heinig, M.; Kirchner, M.; Adami, E.; Rintisch, C.; Dauksaite, V.; Radke, M.H.; et al. RNA-binding protein RBM20 represses splicing to orchestrate cardiac pre-mRNA processing. J. Clin. Invest. 2014, 124, 3419–3430. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, X.; MacLoed, J.; Liu, J. Discerning novel splice junctions derived from RNA-seq alignment: A deep learning approach. BMC Genom. 2018, 19, 971. [Google Scholar] [CrossRef] [PubMed]
- Lizhong, D.; Ethan, R.; Yongsheng, B. Comparrison of alternative splicing junction detection tools using RNA-seq data. Curr. Genom. 2017, 18, 268–277. [Google Scholar]
- Liao, P.; Li, G.; Yu, D.J.; Yong, T.F.; Wang, J.J.; Wang, J.; Soong, T.W. Molecular alteration of Ca(v)1.2 calcium channel in chronic myocardial infarction. Pflug. Arch. 2009, 458, 701–711. [Google Scholar] [CrossRef] [PubMed]
- Liao, P.; Yu, D.; Lu, S.; Tang, Z.; Liang, M.C.; Zeng, S.; Lin, W.; Soong, T.W. Smooth muscle-selective alternatively spliced exon generates functional variation in Cav1.2 calcium channels. J. Biol. Chem. 2004, 279, 50329–50335. [Google Scholar] [CrossRef] [PubMed]
- Dolphin, A.C. Beta subunits of voltage-gated calcium channels. J. Bioenerg. Biomembr. 2003, 35, 599–620. [Google Scholar] [CrossRef] [PubMed]
- Dolphin, A.C. Calcium channel auxiliary α2δ and β subunits: Trafficking and one step beyond. Nat. Rev. Neurosci. 2012, 13, 542–555. [Google Scholar] [CrossRef] [PubMed]
| Gene | Oligonucleotide Sequences |
|---|---|
| GAPDH | Fw: 5′-CAACTCCCTCAAGATTGTCAGCAA-3′ |
| Rev: 5′-GGCATGGACTGTGGTCATGA-3′ | |
| RBM20 | Fw: 5′-GGCTTACACAGAAGCTGCTCAAG-3′ |
| Rev: 5′-GTGGATGTCCTGGATGATAGCAG-3′ | |
| Titin | Fw: 5′-CAGGAGCAGGTTTCTTTGGA-3′ |
| Rev: 5′-GAGCCGTATGAGGAACCGTA-3′ | |
| CACNA1C Exon 1a–4 | Fw: 5′-AGCGATAAGGCCGTATGAGA-3′ |
| Rev: 5′-GTTCCAGGTTGGAGTTGGTG-3′ | |
| CACNA1C Exon 1–4 | Fw: 5′-CAATGGTCAATGAAAACACGA-3′ |
| Rev: 5′-GTTCCAGGTTGGAGTTGGTG-3′ | |
| CACNA1C Exon 7–10 | Fw: 5′-CACCAACTTCGACAACTTCG-3′ |
| Rev: 5′-CCACAGTTTTCACCCTCGAT-3′ | |
| CACNA1C Exon 8–9 | Fw: 5′-GGGTCAATGATGCCGTAGGAA-3′ |
| Rev: 5′-TCTCCTCGAGCTTTGGCTTTC-3′ | |
| CACNA1C Exon 8a–9 | Fw: 5′-ATGCAAGACGCTATGGGCTAT-3′ |
| Rev: 5′-TCTCCTCGAGCTTTGGCTTTC-3′ | |
| CACNA1C Exon 19–21 | Fw: 5′-GCCGGAAGCCAGTGCATTTT-3′ |
| Rev: 5′-AACAATGTCAAAATAAAACAGA-3′ | |
| CACNA1C Exon 19–22 | Fw: 5′-GCCGGAAGCCAGTGCATTTT-3′ |
| Rev: 5′-ATAGTCTGCATTGCCTAGGAT-3′ | |
| CACNA1C Exon 30–35 | Fw: 5′-GCCTCTTCACGGTGGAG-3′ |
| Rev: 5′-TCCCAATCACTGCATAGATAA-3′ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morinaga, A.; Ito, J.; Niimi, T.; Maturana, A.D. RBM20 Regulates CaV1.2 Surface Expression by Promoting Exon 9* Inclusion of CACNA1C in Neonatal Rat Cardiomyocytes. Int. J. Mol. Sci. 2019, 20, 5591. https://doi.org/10.3390/ijms20225591
Morinaga A, Ito J, Niimi T, Maturana AD. RBM20 Regulates CaV1.2 Surface Expression by Promoting Exon 9* Inclusion of CACNA1C in Neonatal Rat Cardiomyocytes. International Journal of Molecular Sciences. 2019; 20(22):5591. https://doi.org/10.3390/ijms20225591
Chicago/Turabian StyleMorinaga, Akihito, Jumpei Ito, Tomoaki Niimi, and Andrés D. Maturana. 2019. "RBM20 Regulates CaV1.2 Surface Expression by Promoting Exon 9* Inclusion of CACNA1C in Neonatal Rat Cardiomyocytes" International Journal of Molecular Sciences 20, no. 22: 5591. https://doi.org/10.3390/ijms20225591
APA StyleMorinaga, A., Ito, J., Niimi, T., & Maturana, A. D. (2019). RBM20 Regulates CaV1.2 Surface Expression by Promoting Exon 9* Inclusion of CACNA1C in Neonatal Rat Cardiomyocytes. International Journal of Molecular Sciences, 20(22), 5591. https://doi.org/10.3390/ijms20225591






