Systematic Enzyme Mapping of Cellular Metabolism by Phasor-Analyzed Label-Free NAD(P)H Fluorescence Lifetime Imaging
Abstract
1. Introduction
2. Results
2.1. Benchmarking of NAD(P)H-FLIM Data Evaluated Using the Phasor Approach
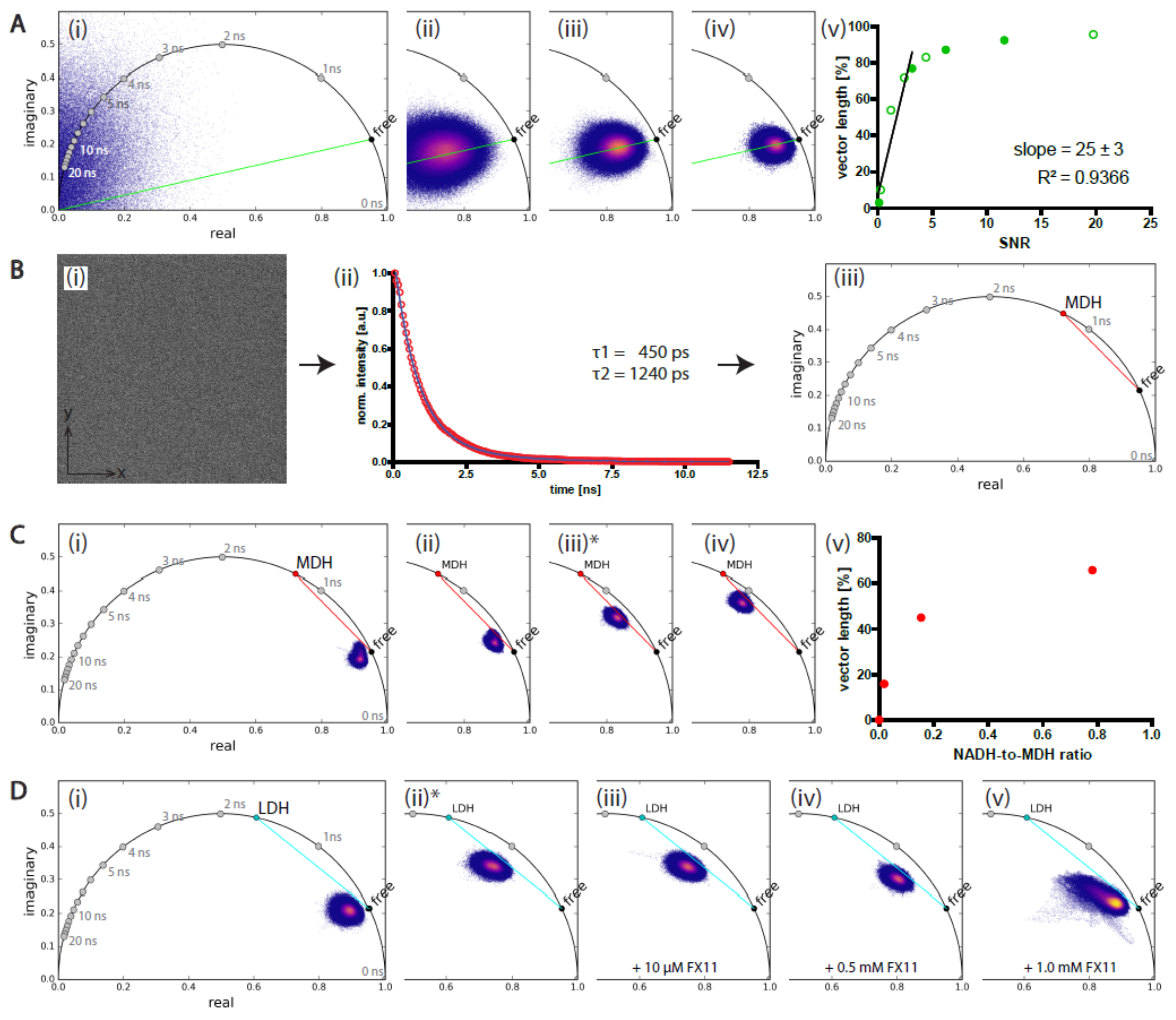
2.1.1. Low Signal-to-Noise Ratio of the Image Affects the NAD(P)H-FLIM Data
2.1.2. Validation of Phasor-Evaluated FLIM Data on Mixtures of NADH and MDH or LDH
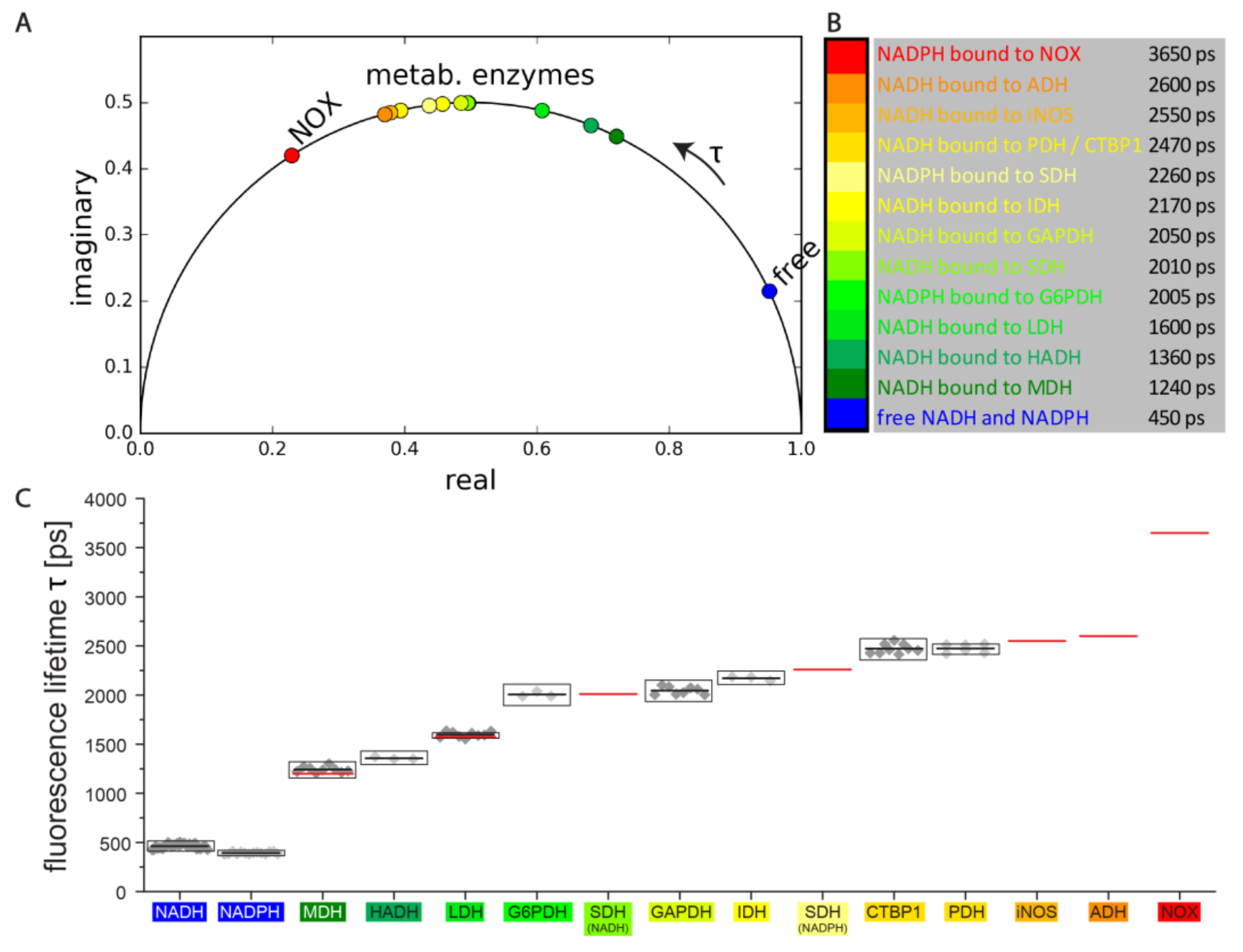
2.2. Enzyme-Based Reference System to Interpret Label-Free NAD(P)H-FLIM Data in Cells
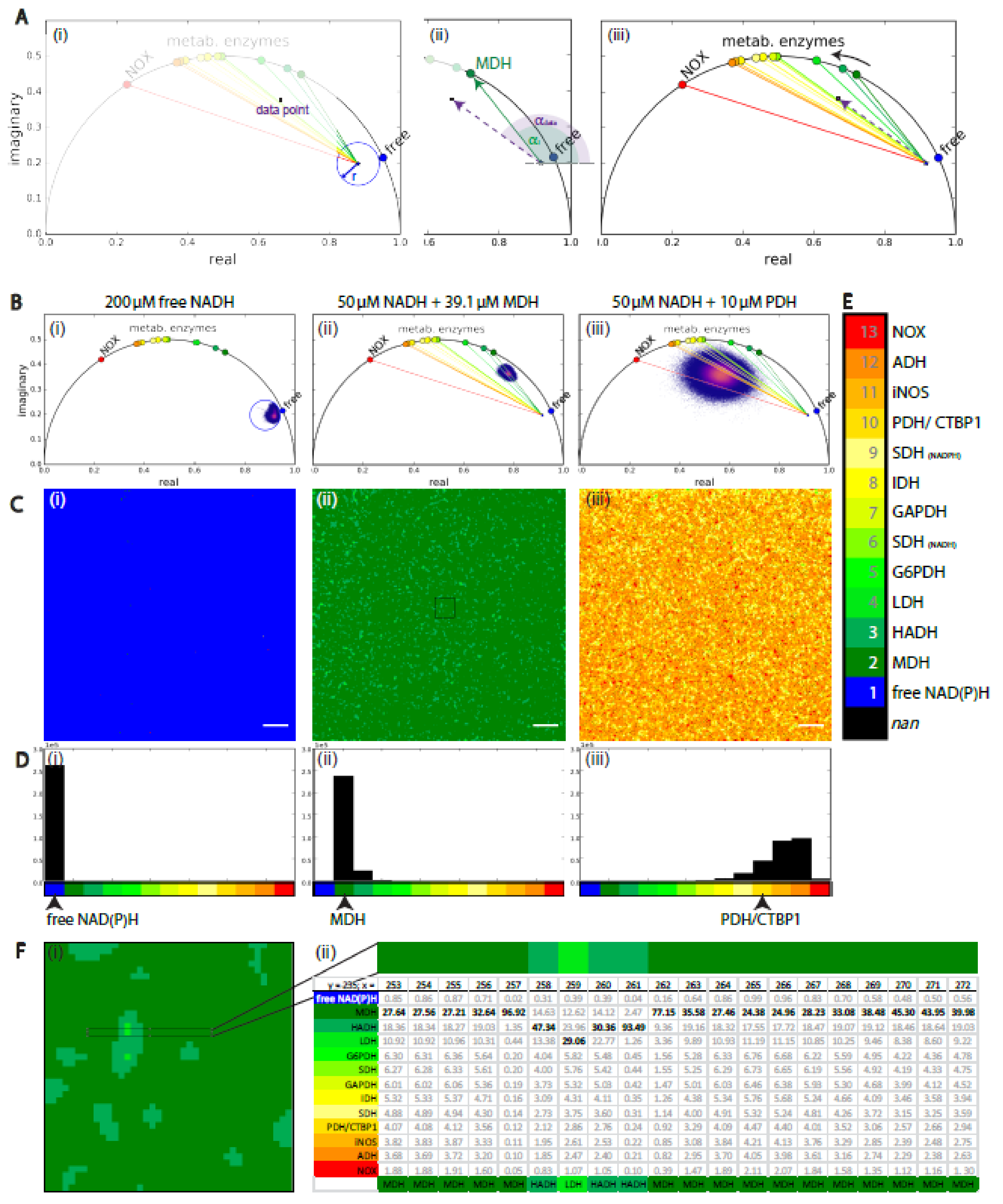
2.3. Analysis Framework of NAD(P)H-FLIM Data: Validation on Homogeneous NAD(P)H-Enzyme Mixtures
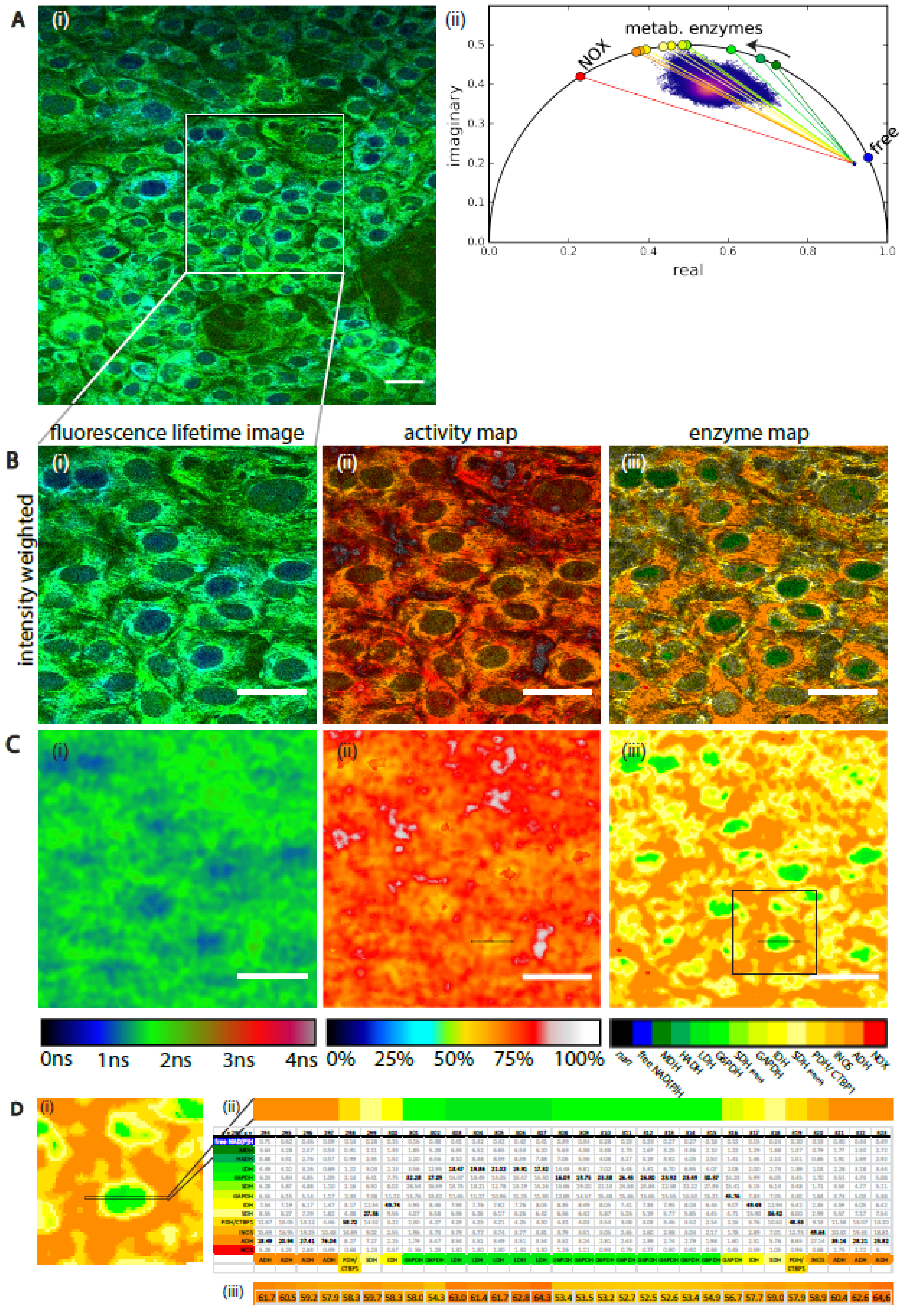
2.4. Validation of the NAD(P)H-FLIM Analysis Framework on Stromal-Like 3T3-L1 Cells
3. Discussion
4. Material and Methods
4.1. Two-Photon Microscope Setup Adequate for FLIM
4.2. Phasor Analysis of Time-Domain NAD(P)H-FLIM Data
4.3. Enzyme and NAD(P)H Solutions
4.4. T3-L1 Cell Culture and Preparation for Imaging
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jellusova, J.; Rickert, R.C. A Brake for B Cell Proliferation: Appropriate responses to metabolic stress are crucial to maintain B cell viability and prevent malignant outgrowth. Bioessays 2017, 39. [Google Scholar] [CrossRef]
- Caputa, G.; Castoldi, A.; Pearce, E.J. Metabolic adaptations of tissue-resident immune cells. Nat. Immunol. 2019, 20, 793–801. [Google Scholar] [CrossRef]
- Jellusova, J.; Cato, M.H.; Apgar, J.R.; Ramezani-Rad, P.; Leung, C.R.; Chen, C.; Richardson, A.D.; Conner, E.M.; Benschop, R.J.; Woodgett, J.R.; et al. Gsk3 is a metabolic checkpoint regulator in B cells. Nat. Immunol. 2017, 18, 303–312. [Google Scholar] [CrossRef]
- Huntosova, V.; Gay, S.; Nowak-Sliwinska, P.M.; Rajendran, S.K.; Zellweger, M.; Van Den Bergh, H.; Wagnières, G. In vivo measurement of tissue oxygenation by time-resolved luminescence spectroscopy: Advantageous properties of dichlorotris (1, 10-phenanthroline)-ruthenium (II) hydrate. J. Biomed. Opt. 2014, 19, 077004. [Google Scholar] [CrossRef] [PubMed]
- Papkovsky, D.B.; Dmitriev, R.I. Biological detection by optical oxygen sensing. Chem. Soc. Rev. 2013, 42, 8700–8732. [Google Scholar] [CrossRef] [PubMed]
- Lakowicz, J.R.; Szmacinski, H.; Nowaczyk, K.; Johnson, M.L. Fluorescence lifetime imaging of free and protein-bound NADH. Proc. Natl. Acad. Sci. USA 1992, 89, 1271–1275. [Google Scholar] [CrossRef] [PubMed]
- Denk, W.; Strickler, J.H.; Webb, W.W. Two-photon laser scanning fluorescence microscopy. Science 1990, 248, 73–76. [Google Scholar] [CrossRef]
- Mossakowski, A.A.; Pohlan, J.; Bremer, D.; Lindquist, R.; Millward, J.M.; Bock, M.; Pollok, K.; Mothes, R.; Viohl, L.; Radbruch, M.; et al. Tracking CNS and systemic sources of oxidative stress during the course of chronic neuroinflammation. Acta Neuropathol. 2015, 130, 799–814. [Google Scholar] [CrossRef]
- Radbruch, H.; Bremer, D.; Guenther, R.; Cseresnyes, Z.; Lindquist, R.; Hauser, A.E.; Niesner, R. Ongoing Oxidative Stress Causes Subclinical Neuronal Dysfunction in the Recovery Phase of EAE. Front. Immunol. 2016, 7, 92. [Google Scholar] [CrossRef]
- Radbruch, H.; Bremer, D.; Mothes, R.; Gunther, R.; Rinnenthal, J.L.; Pohlan, J.; Ulbricht, C.; Hauser, A.E.; Niesner, R. Intravital FRET: Probing Cellular and Tissue Function in Vivo. Int. J. Mol. Sci. 2015, 16, 11713–11727. [Google Scholar] [CrossRef]
- Rinnenthal, J.L.; Bornchen, C.; Radbruch, H.; Andresen, V.; Mossakowski, A.; Siffrin, V.; Seelemann, T.; Spiecker, H.; Moll, I.; Herz, J.; et al. Parallelized TCSPC for dynamic intravital fluorescence lifetime imaging: Quantifying neuronal dysfunction in neuroinflammation. PLoS ONE 2013, 8, e60100. [Google Scholar] [CrossRef] [PubMed]
- Rakymzhan, A.; Radbruch, H.; Niesner, R.A. Quantitative Imaging of Ca(2+) by 3D-FLIM in Live Tissues. Adv. Exp. Med. Biol. 2017, 1035, 135–141. [Google Scholar] [CrossRef] [PubMed]
- Radbruch, H.; Mothes, R.; Bremer, D.; Seifert, S.; Kohler, R.; Pohlan, J.; Ostendorf, L.; Gunther, R.; Leben, R.; Stenzel, W.; et al. Analyzing Nicotinamide Adenine Dinucleotide Phosphate Oxidase Activation in Aging and Vascular Amyloid Pathology. Front. Immunol. 2017, 8, 844. [Google Scholar] [CrossRef] [PubMed]
- Bayerl, S.H.; Niesner, R.; Cseresnyes, Z.; Radbruch, H.; Pohlan, J.; Brandenburg, S.; Czabanka, M.A.; Vajkoczy, P. Time lapse in vivo microscopy reveals distinct dynamics of microglia-tumor environment interactions-a new role for the tumor perivascular space as highway for trafficking microglia. Glia 2016, 64, 1210–1226. [Google Scholar] [CrossRef]
- Tomkova, S.; Misuth, M.; Lenkavska, L.; Miskovsky, P.; Huntosova, V. In vitro identification of mitochondrial oxidative stress production by time-resolved fluorescence imaging of glioma cells. Biochim. Biophys. Acta 2018, 1865, 616–628. [Google Scholar] [CrossRef]
- Kumar, S.; Dunsby, C.; De Beule, P.A.; Owen, D.M.; Anand, U.; Lanigan, P.M.; Benninger, R.K.; Davis, D.M.; Neil, M.A.; Anand, P.; et al. Multifocal multiphoton excitation and time correlated single photon counting detection for 3-D fluorescence lifetime imaging. Opt. Express 2007, 15, 12548–12561. [Google Scholar] [CrossRef]
- Stringari, C.; Cinquin, A.; Cinquin, O.; Digman, M.A.; Donovan, P.J.; Gratton, E. Phasor approach to fluorescence lifetime microscopy distinguishes different metabolic states of germ cells in a live tissue. Proc. Natl. Acad. Sci. USA 2011, 108, 13582–13587. [Google Scholar] [CrossRef]
- Stringari, C.; Nourse, J.L.; Flanagan, L.A.; Gratton, E. Phasor fluorescence lifetime microscopy of free and protein-bound NADH reveals neural stem cell differentiation potential. PLoS ONE 2012, 7, e48014. [Google Scholar] [CrossRef]
- Wright, B.K.; Andrews, L.M.; Jones, M.R.; Stringari, C.; Digman, M.A.; Gratton, E. Phasor-FLIM analysis of NADH distribution and localization in the nucleus of live progenitor myoblast cells. Microsc. Res. Tech. 2012, 75, 1717–1722. [Google Scholar] [CrossRef]
- Wright, B.K.; Andrews, L.M.; Markham, J.; Jones, M.R.; Stringari, C.; Digman, M.A.; Gratton, E. NADH distribution in live progenitor stem cells by phasor-fluorescence lifetime image microscopy. Biophys. J. 2012, 103, L7–L9. [Google Scholar] [CrossRef]
- Guo, H.W.; Chen, C.T.; Wei, Y.H.; Lee, O.K.; Gukassyan, V.; Kao, F.J.; Wang, H.W. Reduced nicotinamide adenine dinucleotide fluorescence lifetime separates human mesenchymal stem cells from differentiated progenies. J. Biomed. Opt. 2008, 13, 050505. [Google Scholar] [CrossRef] [PubMed]
- Gratton, E.; Breusegem, S.; Sutin, J.; Ruan, Q.; Barry, N. Fluorescence lifetime imaging for the two-photon microscope: Time-domain and frequency-domain methods. J. Biomed. Opt. 2003, 8, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Elson, D.; Requejo-Isidro, J.; Munro, I.; Reavell, F.; Siegel, J.; Suhling, K.; Tadrous, P.; Benninger, R.; Lanigan, P.; McGinty, J.; et al. Time-domain fluorescence lifetime imaging applied to biological tissue. Photochem. Photobiol. Sci. 2004, 3, 795–801. [Google Scholar] [CrossRef] [PubMed]
- Soloviev, V.Y.; Tahir, K.B.; McGinty, J.; Elson, D.S.; Neil, M.A.; French, P.M.; Arridge, S.R. Fluorescence lifetime imaging by using time-gated data acquisition. Appl. Opt. 2007, 46, 7384–7391. [Google Scholar] [CrossRef] [PubMed]
- Suhling, K.; French, P.M.; Phillips, D. Time-resolved fluorescence microscopy. Photochem. Photobiol. Sci. 2005, 4, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Poland, S.P.; Krstajic, N.; Monypenny, J.; Coelho, S.; Tyndall, D.; Walker, R.J.; Devauges, V.; Richardson, J.; Dutton, N.; Barber, P.; et al. A high speed multifocal multiphoton fluorescence lifetime imaging microscope for live-cell FRET imaging. Biomed. Opt. Express 2015, 6, 277–296. [Google Scholar] [CrossRef] [PubMed]
- Agronskaia, A.V.; Tertoolen, L.; Gerritsen, H.C. Fast fluorescence lifetime imaging of calcium in living cells. J. Biomed. Opt. 2004, 9, 1230–1237. [Google Scholar] [CrossRef] [PubMed]
- Gerritsen, H.C.; Asselbergs, M.A.; Agronskaia, A.V.; Van Sark, W.G. Fluorescence lifetime imaging in scanning microscopes: Acquisition speed, photon economy and lifetime resolution. J. Microsc. 2002, 206, 218–224. [Google Scholar] [CrossRef]
- Scott, T.G.; Spencer, R.D.; Leonard, N.J.; Weber, G. Synthetic spectroscopic models related to coenzymes and base pairs. V. Emission properties of NADH. Studies of fluorescence lifetimes and quantum efficiencies of NADH, AcPyADH,[reduced acetylpyridineadenine dinucleotide] and simplified synthetic models. J. Am. Chem. Soc. 1970, 92, 687–695. [Google Scholar] [CrossRef]
- Lakowicz, J.R.; Szmacinski, H.; Nowaczyk, K.; Berndt, K.W.; Johnson, M. Fluorescence lifetime imaging. Anal. Biochem. 1992, 202, 316–330. [Google Scholar] [CrossRef]
- Squire, A.; Verveer, P.J.; Bastiaens, P.I. Multiple frequency fluorescence lifetime imaging microscopy. J. Microsc. 2000, 197, 136–149. [Google Scholar] [CrossRef] [PubMed]
- Verveer, P.J.; Rocks, O.; Harpur, A.G.; Bastiaens, P.I. FLIM measurements and frequency domain FLIM data analysis. CSH Protoc. 2006, 2006. [Google Scholar] [CrossRef] [PubMed]
- Leben, R.; Ostendorf, L.; van Koppen, S.; Rakhymzhan, A.; Hauser, A.E.; Radbruch, H.; Niesner, R.A. Phasor-Based Endogenous NAD(P)H Fluorescence Lifetime Imaging Unravels Specific Enzymatic Activity of Neutrophil Granulocytes Preceding NETosis. Int. J. Mol. Sci. 2018, 19, 1018. [Google Scholar] [CrossRef] [PubMed]
- Chance, B. Mitochondrial NADH redox state, monitoring discovery and deployment in tissue. Methods Enzymol 2004, 385, 361–370. [Google Scholar] [CrossRef] [PubMed]
- Chacko, J.V.; Eliceiri, K.W. Autofluorescence lifetime imaging of cellular metabolism: Sensitivity toward cell density, pH, intracellular, and intercellular heterogeneity. Cytometry A 2019, 95, 56–69. [Google Scholar] [CrossRef]
- Xu, H.N.; Tchou, J.; Chance, B.; Li, L.Z. Imaging the redox states of human breast cancer core biopsies. Adv. Exp. Med. Biol. 2013, 765, 343–349. [Google Scholar] [CrossRef]
- Bird, D.K.; Yan, L.; Vrotsos, K.M.; Eliceiri, K.W.; Vaughan, E.M.; Keely, P.J.; White, J.G.; Ramanujam, N. Metabolic mapping of MCF10A human breast cells via multiphoton fluorescence lifetime imaging of the coenzyme NADH. Cancer Res. 2005, 65, 8766–8773. [Google Scholar] [CrossRef]
- Skala, M.C.; Riching, K.M.; Bird, D.K.; Gendron-Fitzpatrick, A.; Eickhoff, J.; Eliceiri, K.W.; Keely, P.J.; Ramanujam, N. In vivo multiphoton fluorescence lifetime imaging of protein-bound and free nicotinamide adenine dinucleotide in normal and precancerous epithelia. J. Biomed. Opt. 2007, 12, 024014. [Google Scholar] [CrossRef]
- Skala, M.C.; Riching, K.M.; Gendron-Fitzpatrick, A.; Eickhoff, J.; Eliceiri, K.W.; White, J.G.; Ramanujam, N. In vivo multiphoton microscopy of NADH and FAD redox states, fluorescence lifetimes, and cellular morphology in precancerous epithelia. Proc. Natl. Acad. Sci. USA 2007, 104, 19494–19499. [Google Scholar] [CrossRef]
- Provenzano, P.P.; Eliceiri, K.W.; Keely, P.J. Multiphoton microscopy and fluorescence lifetime imaging microscopy (FLIM) to monitor metastasis and the tumor microenvironment. Clin. Exp. Metastasis 2009, 26, 357–370. [Google Scholar] [CrossRef]
- Meleshina, A.V.; Cherkasova, E.I.; Shirmanova, M.V.; Klementieva, N.V.; Kiseleva, E.V.; Snopova Lcapital Ve, C.; Prodanets, N.N.; Zagaynova, E.V. Influence of mesenchymal stem cells on metastasis development in mice in vivo. Stem Cell Res. Ther. 2015, 6, 15. [Google Scholar] [CrossRef] [PubMed]
- Meleshina, A.V.; Dudenkova, V.V.; Bystrova, A.S.; Kuznetsova, D.S.; Shirmanova, M.V.; Zagaynova, E.V. Two-photon FLIM of NAD(P)H and FAD in mesenchymal stem cells undergoing either osteogenic or chondrogenic differentiation. Stem Cell Res. Ther. 2017, 8, 15. [Google Scholar] [CrossRef] [PubMed]
- Meleshina, A.V.; Dudenkova, V.V.; Shirmanova, M.V.; Shcheslavskiy, V.I.; Becker, W.; Bystrova, A.S.; Cherkasova, E.I.; Zagaynova, E.V. Probing metabolic states of differentiating stem cells using two-photon FLIM. Sci. Rep. 2016, 6, 21853. [Google Scholar] [CrossRef] [PubMed]
- Evers, M.; Salma, N.; Osseiran, S.; Casper, M.; Birngruber, R.; Evans, C.L.; Manstein, D. Enhanced quantification of metabolic activity for individual adipocytes by label-free FLIM. Sci. Rep. 2018, 8, 8757. [Google Scholar] [CrossRef] [PubMed]
- Lindquist, R.L.; Bayat-Sarmadi, J.; Leben, R.; Niesner, R.; Hauser, A.E. NAD(P)H Oxidase Activity in the Small Intestine Is Predominantly Found in Enterocytes, Not Professional Phagocytes. Int. J. Mol. Sci. 2018, 19, 1365. [Google Scholar] [CrossRef] [PubMed]
- Alturkistany, F.; Nichani, K.; Houston, K.D.; Houston, J.P. Fluorescence lifetime shifts of NAD(P)H during apoptosis measured by time-resolved flow cytometry. Cytometry A 2019, 95, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Sharick, J.T.; Favreau, P.F.; Gillette, A.A.; Sdao, S.M.; Merrins, M.J.; Skala, M.C. Protein-bound NAD(P)H Lifetime is Sensitive to Multiple Fates of Glucose Carbon. Sci. Rep. 2018, 8, 5456. [Google Scholar] [CrossRef]
- Niesner, R.; Peker, B.; Schlusche, P.; Gericke, K.H. Noniterative biexponential fluorescence lifetime imaging in the investigation of cellular metabolism by means of NAD(P)H autofluorescence. Chemphyschem 2004, 5, 1141–1149. [Google Scholar] [CrossRef]
- Strickler, S.J.; Robert, A.B. Relationship between Absorption Intensity and Fluorescence Lifetime of Molecules. J. Chem. Phys. 1962, 37, 814–822. [Google Scholar] [CrossRef]
- Behne, M.J.; Meyer, J.W.; Hanson, K.M.; Barry, N.P.; Murata, S.; Crumrine, D.; Clegg, R.W.; Gratton, E.; Holleran, W.M.; Elias, P.M.; et al. NHE1 regulates the stratum corneum permeability barrier homeostasis. Microenvironment acidification assessed with fluorescence lifetime imaging. J. Biol. Chem. 2002, 277, 47399–47406. [Google Scholar] [CrossRef]
- Niesner, R.; Peker, B.; Schlusche, P.; Gericke, K.H.; Hoffmann, C.; Hahne, D.; Muller-Goymann, C. 3D-resolved investigation of the pH gradient in artificial skin constructs by means of fluorescence lifetime imaging. Pharm. Res. 2005, 22, 1079–1087. [Google Scholar] [CrossRef] [PubMed]
- Celli, A.; Sanchez, S.; Behne, M.; Hazlett, T.; Gratton, E.; Mauro, T. The epidermal Ca(2+) gradient: Measurement using the phasor representation of fluorescent lifetime imaging. Biophys. J. 2010, 98, 911–921. [Google Scholar] [CrossRef]
- Blacker, T.S.; Mann, Z.F.; Gale, J.E.; Ziegler, M.; Bain, A.J.; Szabadkai, G.; Duchen, M.R. Separating NADH and NADPH fluorescence in live cells and tissues using FLIM. Nat. Commun. 2014, 5, 3936. [Google Scholar] [CrossRef] [PubMed]
- Datta, R.; Alfonso-Garcia, A.; Cinco, R.; Gratton, E. Fluorescence lifetime imaging of endogenous biomarker of oxidative stress. Sci. Rep. 2015, 5, 9848. [Google Scholar] [CrossRef] [PubMed]
- Niesner, R.; Narang, P.; Spiecker, H.; Andresen, V.; Gericke, K.H.; Gunzer, M. Selective detection of NADPH oxidase in polymorphonuclear cells by means of NAD(P)H-based fluorescence lifetime imaging. J. Biophys. 2008, 2008, 602639. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.C.; Siegel, J.; Webb, S.E.; Leveque-Fort, S.; Cole, M.J.; Jones, R.; Dowling, K.; Lever, M.J.; French, P.M. Application of the stretched exponential function to fluorescence lifetime imaging. Biophys. J. 2001, 81, 1265–1274. [Google Scholar] [CrossRef]
- Zhang, Q.; Piston, D.W.; Goodman, R.H. Regulation of corepressor function by nuclear NADH. Science 2002, 295, 1895–1897. [Google Scholar] [CrossRef]
- Madhukar, N.S.; Warmoes, M.O.; Locasale, J.W. Organization of enzyme concentration across the metabolic network in cancer cells. PLoS ONE 2015, 10, e0117131. [Google Scholar] [CrossRef]
- Babior, B.M. NADPH oxidase. Curr. Opin. Immunol. 2004, 16, 42–47. [Google Scholar] [CrossRef]
- Ranjit, S.; Malacrida, L.; Jameson, D.M.; Gratton, E. Fit-free analysis of fluorescence lifetime imaging data using the phasor approach. Nat. Protoc. 2018, 13, 1979–2004. [Google Scholar] [CrossRef]
- Lindquist, R.L.; Niesner, R.A.; Hauser, A.E. In the Right Place, at the Right Time: Spatiotemporal Conditions Determining Plasma Cell Survival and Function. Front. Immunol. 2019, 10, 788. [Google Scholar] [CrossRef] [PubMed]
- Itkin, T.; Gur-Cohen, S.; Spencer, J.A.; Schajnovitz, A.; Ramasamy, S.K.; Kusumbe, A.P.; Ledergor, G.; Jung, Y.; Milo, I.; Poulos, M.G.; et al. Distinct bone marrow blood vessels differentially regulate haematopoiesis. Nature 2016, 532, 323–328. [Google Scholar] [CrossRef] [PubMed]
- Ambrosi, T.H.; Scialdone, A.; Graja, A.; Gohlke, S.; Jank, A.M.; Bocian, C.; Woelk, L.; Fan, H.; Logan, D.W.; Schurmann, A.; et al. Adipocyte Accumulation in the Bone Marrow during Obesity and Aging Impairs Stem Cell-Based Hematopoietic and Bone Regeneration. Cell Stem Cell 2017, 20, 771–778. [Google Scholar] [CrossRef] [PubMed]
- Czechowska, K.; Lannigan, J.; Wang, L.; Arcidiacono, J.; Ashhurst, T.M.; Barnard, R.M.; Bauer, S.; Bispo, C.; Bonilla, D.L.; Brinkman, R.R.; et al. Cyt-Geist: Current and Future Challenges in Cytometry: Reports of the CYTO 2018 Conference Workshops. Cytometry A 2019, 95, 598–644. [Google Scholar] [CrossRef] [PubMed]
- Jellusova, J. Cross-talk between signal transduction and metabolism in B cells. Immunol. Lett. 2018, 201, 1–13. [Google Scholar] [CrossRef]
- O’Neill, L.A.; Kishton, R.J.; Rathmell, J. A guide to immunometabolism for immunologists. Nat. Rev. Immunol. 2016, 16, 553–565. [Google Scholar] [CrossRef]
- Holzwarth, K.; Kohler, R.; Philipsen, L.; Tokoyoda, K.; Ladyhina, V.; Wahlby, C.; Niesner, R.A.; Hauser, A.E. Multiplexed fluorescence microscopy reveals heterogeneity among stromal cells in mouse bone marrow sections. Cytometry A 2018, 93, 876–888. [Google Scholar] [CrossRef]
- Zehentmeier, S.; Roth, K.; Cseresnyes, Z.; Sercan, O.; Horn, K.; Niesner, R.A.; Chang, H.D.; Radbruch, A.; Hauser, A.E. Static and dynamic components synergize to form a stable survival niche for bone marrow plasma cells. Eur J. Immunol. 2014, 44, 2306–2317. [Google Scholar] [CrossRef]
- Digman, M.A.; Caiolfa, V.R.; Zamai, M.; Gratton, E. The phasor approach to fluorescence lifetime imaging analysis. Biophys. J. 2008, 94, L14–L16. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Leben, R.; Köhler, M.; Radbruch, H.; Hauser, A.E.; Niesner, R.A. Systematic Enzyme Mapping of Cellular Metabolism by Phasor-Analyzed Label-Free NAD(P)H Fluorescence Lifetime Imaging. Int. J. Mol. Sci. 2019, 20, 5565. https://doi.org/10.3390/ijms20225565
Leben R, Köhler M, Radbruch H, Hauser AE, Niesner RA. Systematic Enzyme Mapping of Cellular Metabolism by Phasor-Analyzed Label-Free NAD(P)H Fluorescence Lifetime Imaging. International Journal of Molecular Sciences. 2019; 20(22):5565. https://doi.org/10.3390/ijms20225565
Chicago/Turabian StyleLeben, Ruth, Markus Köhler, Helena Radbruch, Anja E. Hauser, and Raluca A. Niesner. 2019. "Systematic Enzyme Mapping of Cellular Metabolism by Phasor-Analyzed Label-Free NAD(P)H Fluorescence Lifetime Imaging" International Journal of Molecular Sciences 20, no. 22: 5565. https://doi.org/10.3390/ijms20225565
APA StyleLeben, R., Köhler, M., Radbruch, H., Hauser, A. E., & Niesner, R. A. (2019). Systematic Enzyme Mapping of Cellular Metabolism by Phasor-Analyzed Label-Free NAD(P)H Fluorescence Lifetime Imaging. International Journal of Molecular Sciences, 20(22), 5565. https://doi.org/10.3390/ijms20225565





