Expression of Two α-Type Expansins from Ammopiptanthus nanus in Arabidopsis thaliana Enhance Tolerance to Cold and Drought Stresses
Abstract
1. Introduction
2. Results
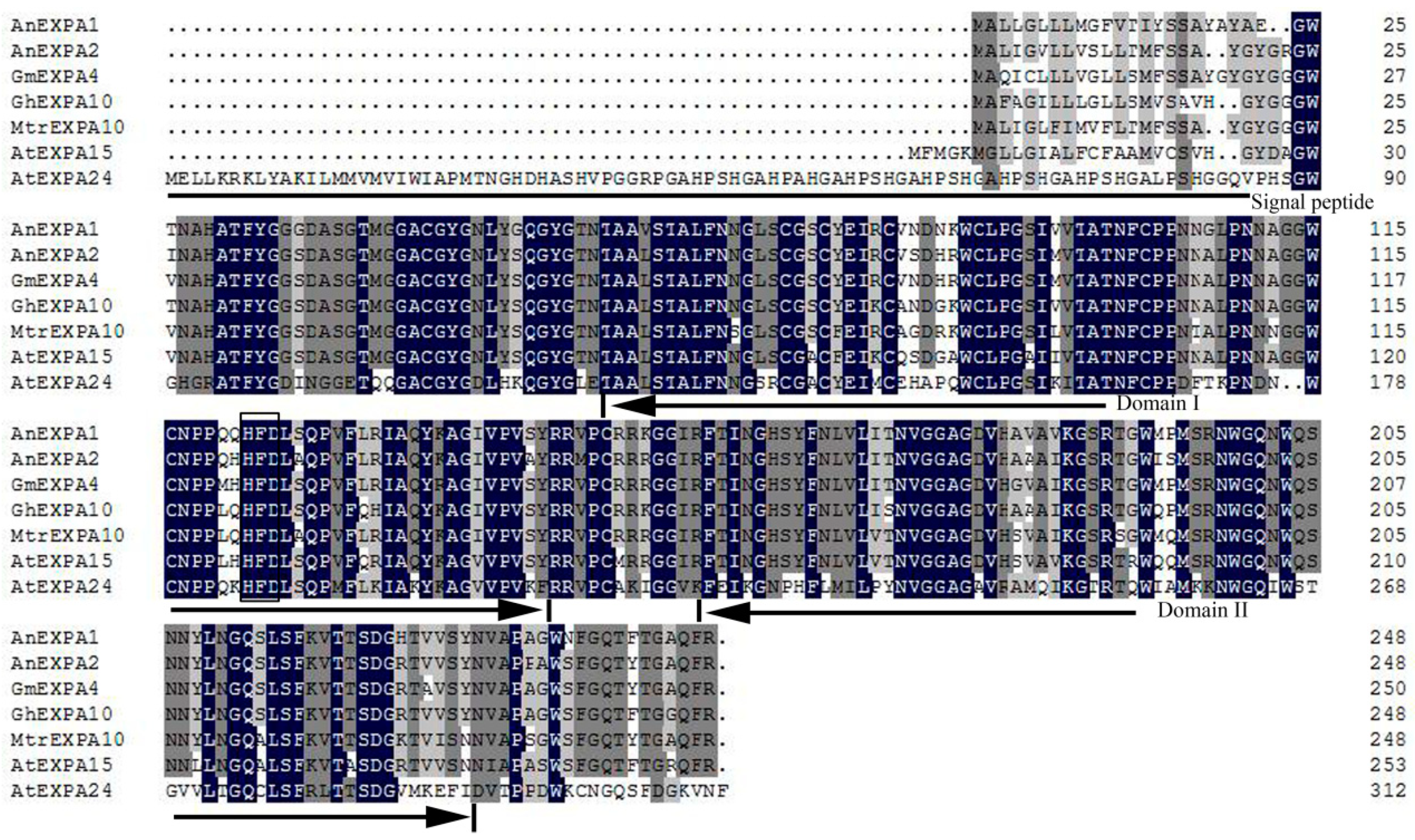
2.1. Cloning and Characterization of AnEXPA1 and AnEXPA2
2.2. AnEXPA1 and AnEXPA2 Showed Similar 3D Structure
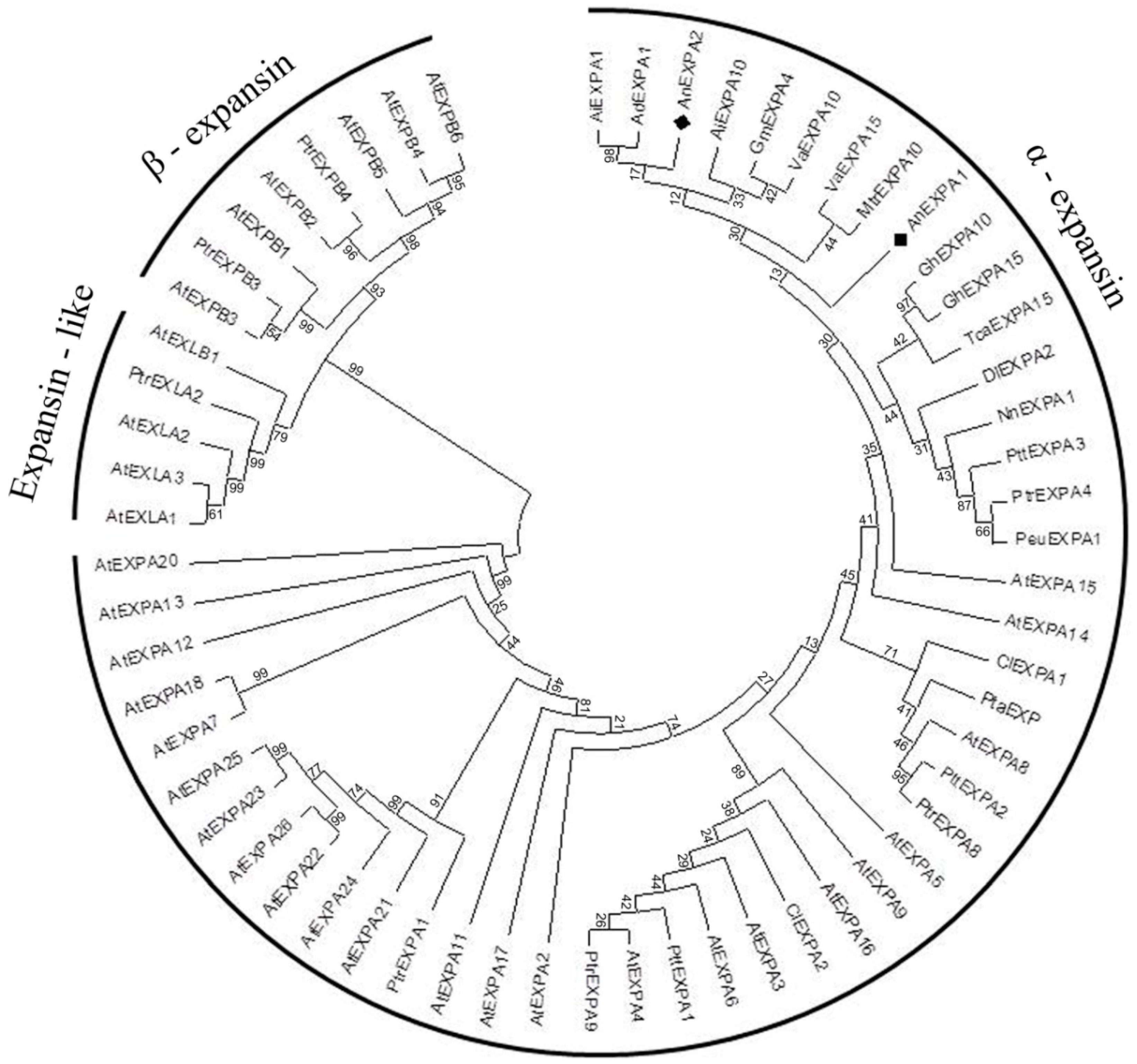
2.3. Phylogenetic Analysis
2.4. Cis-Element Analysis of AnEXPA1 and AnEXPA2 Promoters
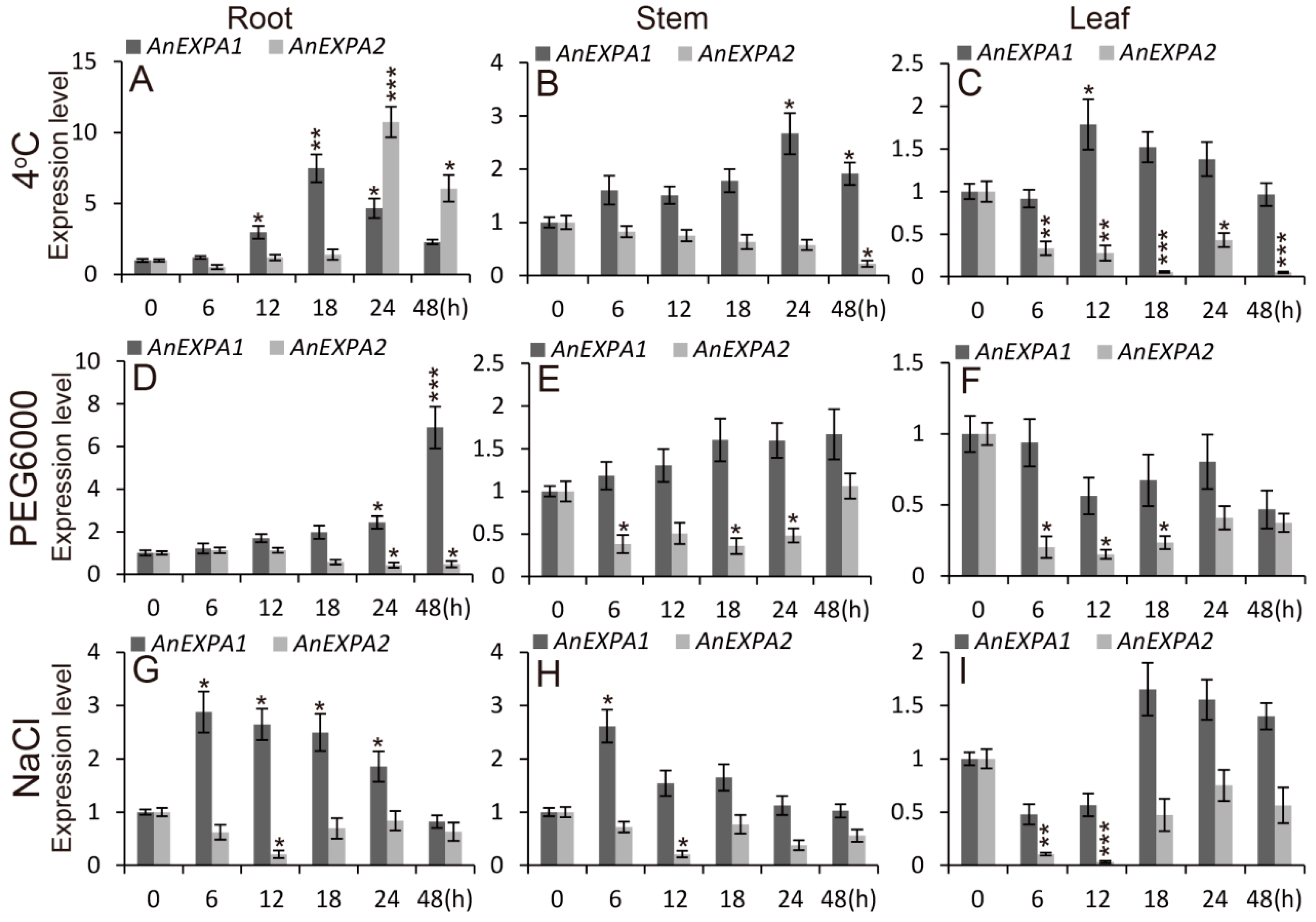
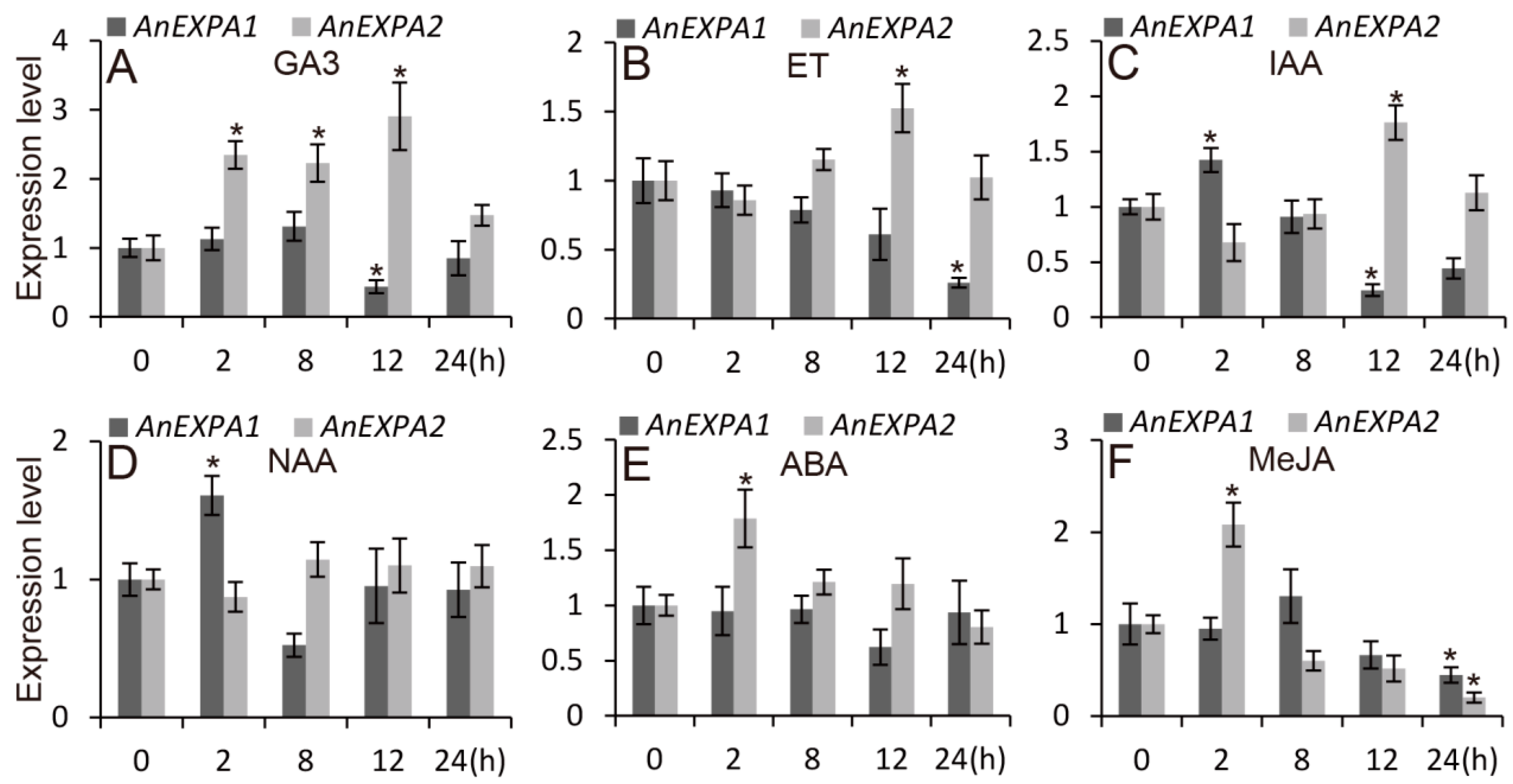
2.5. AnEXPA1 and AnEXPA2 Response to Different Abiotic Stresses and Hormone Induction
2.6. Promoters of AnEXPA1 and AnEXPA2 Showed Different Effects during Abiotic Stress and Hormone Induction
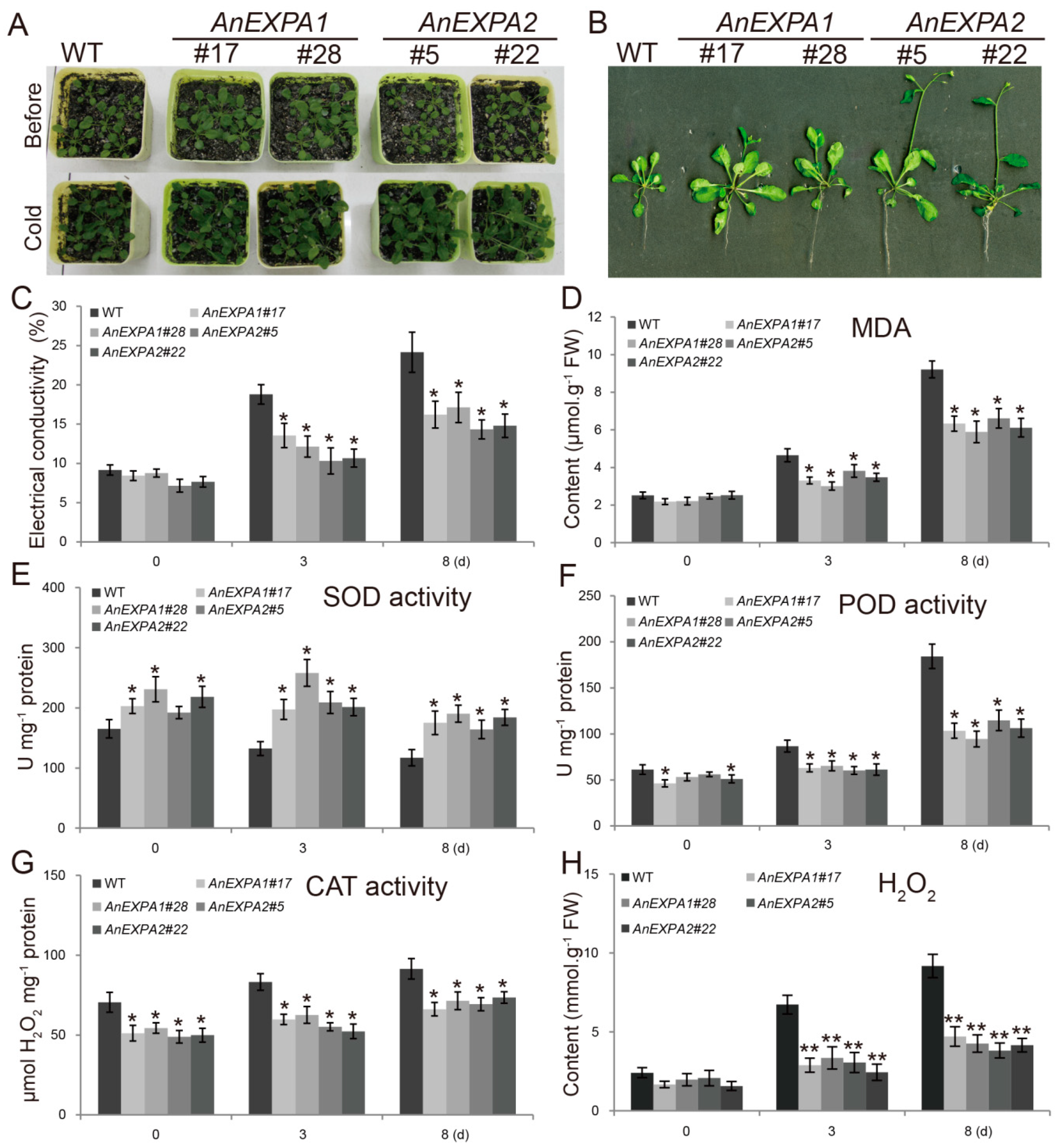
2.7. Overexpression of AnEXPA1 and AnEXPA2 in Arabidopsis Enhanced Tolerance to Cold Stress
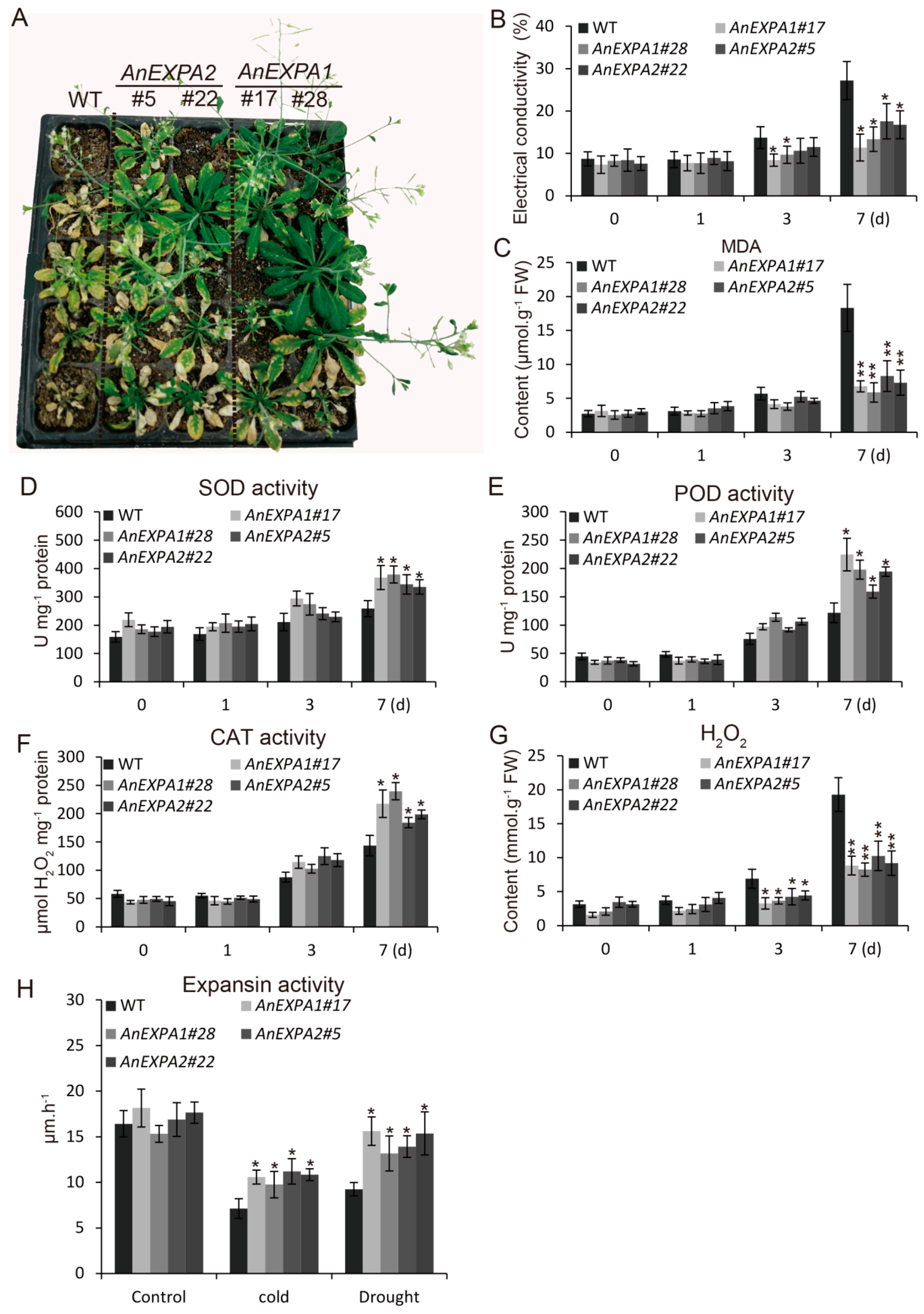
2.8. Overexpression of AnEXPA1 and AnEXPA2 Enhanced Transgenic Plant Tolerance to Drought Stress
3. Discussion
3.1. AnEXPA1 and AnEXPA2 Responded to Different Abiotic Stresses and Hormone Signals
3.2. Overexpression of AnEXPA1 and AnEXPA2 in Arabidopsis Enhanced Tolerance to Cold and Drought Stresses
4. Materials and Methods
4.1. Plant Materials
4.2. RNA and DNA Extraction
4.3. Genome Walking
4.4. Generation of Plant Expression Vectors
4.5. GUS Dyeing and Protein Activity Assays
4.6. Arabidopsis Transgenic Plant Isolation
4.7. Cold and Drought Tolerance of Transgenic Plant Assays
4.8. Bioinformatics Analysis
4.9. qRT-PCR
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Madhava Rao, K.; Raghavendra, A.; Janardhan Reddy, K. Physiology and Molecular Biology of Stress Tolerance in Plants; Springer Science & Business Media: Berlin, Germany, 2006; pp. 1–335. [Google Scholar]
- Han, Y.; Li, A.X.; Li, F.; Zhao, M.R.; Wang, W. Characterization of a wheat (Triticum aestivum L.) expansin gene, TaEXPB23, involved in the abiotic stress response and phytohormone regulation. Plant Physiol. Biochem. 2012, 54, 49–58. [Google Scholar] [CrossRef]
- Chen, L.J.; Zou, W.S.; Wu, G.; Lin, H.H.; Xi, D.H. Tobacco alpha-expansin EXPA4 plays a role in Nicotiana benthamiana defence against Tobacco mosaic virus. Planta 2018, 247, 355–368. [Google Scholar] [CrossRef]
- Martin, C.; Bhatt, K.; Baumann, K. Shaping in plant cells. Curr. Opin. Plant Biol. 2001, 4, 540–549. [Google Scholar] [CrossRef]
- Smith, L.G. Cytoskeletal control of plant cell shape: Getting the fine points. Curr. Opin. Plant Biol. 2003, 6, 63–73. [Google Scholar] [CrossRef]
- Cosgrove, D.J.; Li, L.C.; Cho, H.T.; Hoffmann-Benning, S.; Moore, R.C.; Blecker, D. The growing world of expansins. Plant Cell Physiol. 2002, 43, 1436–1444. [Google Scholar] [CrossRef]
- Sampedro, J.; Cosgrove, D.J. The expansin superfamily. Genome Biol. 2005, 6, 242. [Google Scholar] [CrossRef][Green Version]
- Ishimaru, M.; Smith, D.L.; Gross, K.C.; Kobayashi, S. Expression of three expansin genes during development and maturation of Kyoho grape berries. J. Plant Physiol. 2007, 164, 1675–1682. [Google Scholar] [CrossRef]
- Kasai, S.; Hayama, H.; Kashimura, Y.; Kudo, S.; Osanai, Y. Relationship between fruit cracking and expression of the expansin gene MdEXPA3 in ‘Fuji’ apples (Malus domestica Borkh.). Sci. Hortic. 2008, 116, 194–198. [Google Scholar] [CrossRef]
- Kumar, M.; Singh, P.; Sukla, L. Addition of expansin to cellulase enhanced bioethanol production. Process. Biochem. 2016, 51, 2097–2103. [Google Scholar] [CrossRef]
- Lei, X.Y.; Wang, Q.J.; Wang, J.W.; Zheng, L.P. Cloning and characterization of an expansin gene AbEXP from Achyranthes bidentata. Plant Growth Regul. 2017, 83, 479–487. [Google Scholar] [CrossRef]
- Zhang, J.F.; Xu, Y.Q.; Dong, J.M.; Peng, L.N.; Feng, X.; Wang, X.; Li, F.; Miao, Y.; Yao, S.K.; Zhao, Q.Q. Genome-wide identification of wheat (Triticum aestivum) expansins and expansin expression analysis in cold-tolerant and cold-sensitive wheat cultivars. PLoS ONE 2018, 13, e0195138. [Google Scholar] [CrossRef]
- Goh, H.H.; Sloan, J.; Malinowski, R.; Fleming, A. Variable expansin expression in Arabidopsis leads to different growth responses. J. Plant Physiol. 2014, 171, 329–339. [Google Scholar] [CrossRef]
- He, D.; Lei, Z.; Xing, H.; Tang, B.; Zhao, J.; Lu, B. Exp2 polymorphisms associated with variation for fiber quality properties in cotton (Gossypium spp.). Crop. J. 2014, 2, 315–328. [Google Scholar] [CrossRef][Green Version]
- Park, S.H.; Li, F.; Renaud, J.; Shen, W.; Li, Y.; Guo, L.; Cui, H.; Sumarah, M.; Wang, A. NbEXPA1, an α-expansin, is plasmodesmata-specific and a novel host factor for potyviral infection. Plant J. 2017, 92, 846–861. [Google Scholar] [CrossRef]
- Dotto, M.C.; Pombo, M.A.; Martínez, G.A.; Civello, P.M. Heat treatments and expansin gene expression in strawberry fruit. Sci. Hortic. 2011, 130, 775–780. [Google Scholar] [CrossRef]
- Nardi, C.F.; Villarreal, N.M.; Rossi, F.R.; Martínez, S.; Martínez, G.A.; Civello, P.M. Overexpression of the carbohydrate binding module of strawberry expansin 2 in Arabidopsis thaliana modifies plant growth and cell wall metabolism. Plant Mol. Biol. 2015, 88, 101–117. [Google Scholar] [CrossRef]
- Han, Y.; Chen, Y.; Yin, S.; Zhang, M.; Wang, W. Over-expression of TaEXPB23, a wheat expansin gene, improves oxidative stress tolerance in transgenic tobacco plants. J. Plant Physiol. 2015, 173, 62–71. [Google Scholar] [CrossRef]
- Zhou, S.; Han, Y.Y.; Chen, Y.; Kong, X.; Wang, W. The involvement of expansins in response to water stress during leaf development in wheat. J. Plant Physiol. 2015, 183, 64–74. [Google Scholar] [CrossRef]
- Cosgrove, D.J. Plant expansins: Diversity and interactions with plant cell walls. Curr. Opin. Plant Biol. 2015, 25, 162–172. [Google Scholar] [CrossRef]
- Fahad, S.; Hussain, S.; Bano, A.; Saud, S.; Hassan, S.; Shan, D.; Khan, F.A.; Khan, F.; Chen, Y.; Wu, C. Potential role of phytohormones and plant growth-promoting rhizobacteria in abiotic stresses: Consequences for changing environment. Environ. Sci. Pollut. Res. 2015, 22, 4907–4921. [Google Scholar] [CrossRef]
- Kohli, A.; Sreenivasulu, N.; Lakshmanan, P.; Kumar, P.P. The phytohormone crosstalk paradigm takes center stage in understanding how plants respond to abiotic stresses. Plant Cell Rep. 2013, 32, 945–957. [Google Scholar] [CrossRef]
- Ochiai, M.; Matsumoto, S.; Yamada, K. Methyl jasmonate treatment promotes flower opening of cut Eustoma by inducing cell wall loosening proteins in petals. Postharvest Biol. Technol. 2013, 82, 1–5. [Google Scholar] [CrossRef]
- Liu, Y.D.; Zhang, L.; Chen, L.J.; Ma, H.; Ruan, Y.Y.; Xu, T.; Xu, C.Q.; He, Y.; Qi, M.F. Molecular cloning and expression of an encoding galactinol synthase gene (AnGolS1) in seedling of Ammopiptanthus nanus. Sci. Rep. 2016, 6, 36113. [Google Scholar] [CrossRef]
- Valenzuela-Riffo, F.; Gaete-Eastman, C.; Stappung, Y.; Lizana, R.; Herrera, R.; Moya-León, M.A.; Morales-Quintana, L. Comparative in silico study of the differences in the structure and ligand interaction properties of three alpha-expansin proteins from Fragaria chiloensis fruit. J. Biomol. Struct. Dyn. 2019, 37, 3245–3258. [Google Scholar] [CrossRef]
- Santiago, T.R.; Pereira, V.M.; de Souza, W.R.; Steindorff, A.S.; Cunha, B.A.; Gaspar, M.; Favaro, L.C.; Formighieri, E.F.; Kobayashi, A.K.; Molinari, H.B. Genome-wide identification, characterization and expression profile analysis of expansins gene family in sugarcane (Saccharum spp.). PLoS ONE 2018, 13, e0191081. [Google Scholar] [CrossRef]
- Hepler, N.K.; Cosgrove, D.J. Directed in vitro evolution of bacterial expansin BsEXLX1 for higher cellulose binding and its consequences for plant cell wall-loosening activities. FEBS Lett. 2019, 593, 2545–2555. [Google Scholar] [CrossRef]
- Basu, A.; Sarkar, A.; Maulik, U.; Basak, P. Three dimensional structure prediction and ligand-protein interaction study of expansin protein ATEXPA23 from Arabidopsis thaliana L. Indian J. Biochem. Biophys. 2019, 56, 20–27. [Google Scholar]
- Bouzarelou, D.; Billini, M.; Roumelioti, K.; Sophianopoulou, V. EglD, a putative endoglucanase, with an expansin like domain is localized in the conidial cell wall of Aspergillus nidulans. Fungal Genet. Biol. 2008, 45, 839–850. [Google Scholar] [CrossRef]
- Bunterngsook, B.; Eurwilaichitr, L.; Thamchaipenet, A.; Champreda, V. Binding characteristics and synergistic effects of bacterial expansins on cellulosic and hemicellulosic substrates. Bioresour. Technol. 2015, 176, 129–135. [Google Scholar] [CrossRef]
- Georgelis, N.; Nikolaidis, N.; Cosgrove, D.J. Biochemical analysis of expansin-like proteins from microbes. Carbohydr. Polym. 2014, 100, 17–23. [Google Scholar] [CrossRef]
- Cheng, M.C.; Liao, P.M.; Kuo, W.W.; Lin, T.P. The Arabidopsis ETHYLENE RESPONSE FACTOR1 regulates abiotic stress-responsive gene expression by binding to different cis-acting elements in response to different stress signals. Plant Physiol. 2013, 162, 1566–1582. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.H.; Peng, T.; Dai, W. Critical cis-acting elements and interacting transcription factors: Key players associated with abiotic stress responses in plants. Plant Mol. Biol. Rep. 2014, 32, 303–317. [Google Scholar] [CrossRef]
- Agarwal, P.K.; Agarwal, P.; Reddy, M.; Sopory, S.K. Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep. 2006, 25, 1263–1274. [Google Scholar] [CrossRef]
- Kuluev, B.; Avalbaev, A.; Mikhaylova, E.; Nikonorov, Y.; Berezhneva, Z.; Chemeris, A. Expression profiles and hormonal regulation of tobacco expansin genes and their involvement in abiotic stress response. J. Plant Physiol. 2016, 206, 1–12. [Google Scholar] [CrossRef]
- Peng, L.N.; Xu, Y.; Wang, X.; Feng, X.; Zhao, Q.; Feng, S.; Zhao, Z.; Hu, B.; Li, F. Overexpression of paralogues of the wheat expansin gene TaEXPA8 improves low-temperature tolerance in Arabidopsis. Plant Biol. 2019. [Google Scholar] [CrossRef]
- Fujiwara, T.; Beachy, R.N. Tissue-specific and temporal regulation of a beta-conglycinin gene: Roles of the RY repeat and other cis-acting elements. Plant Mol. Biol. 1994, 24, 261–272. [Google Scholar] [CrossRef]
- Zhang, T.; Zhao, X.Q.; Huang, L.Y.; Liu, X.Y.; Ying, Z.; Zhu, L.H.; Yang, D.C.; Fu, B.Y. Tissue-specific transcriptomic profiling of Sorghum propinquum using a rice genome array. PLoS ONE 2013, 8, e60202. [Google Scholar] [CrossRef]
- Nishiuchi, T.; Shinshi, H.; Suzuki, K. Rapid and transient activation of transcription of the ERF3 gene by wounding in tobacco leaves: Possible involvement of NtWRKYs and autorepression. J. Biol. Chem. 2004, 279, 55355. [Google Scholar] [CrossRef]
- Yu, T.; Zhao, W.; Fu, J.; Liu, Y.W.; Chen, M.; Zhou, Y.; Ma, Y.Z.; Xu, Z.S.; Xi, Y. Genome-wide analysis of CDPK family in Foxtail millet and determination of SiCDPK24 functions in drought stress. Front. Plant Sci. 2018, 9, 651. [Google Scholar] [CrossRef]
- Arjun, S.; Khaled, M.; Salma, A.A.; Ahmed, A.A.; Rabah, I.; Synan, A.Q. Identification of Arabidopsis candidate genes in response to biotic and abiotic stresses using comparative microarrays. PLoS ONE 2015, 10, e0125666. [Google Scholar]
- Sun, T.; Busta, L.; Zhang, Q.; Ding, P.; Jetter, R.; Zhang, Y. TGACG-BINDING FACTOR 1 (TGA1) and TGA4 regulate salicylic acid and pipecolic acid biosynthesis by modulating the expression of SYSTEMIC ACQUIRED RESISTANCE DEFICIENT 1 (SARD1) and CALMODULIN-BINDING PROTEIN 60g (CBP60g). New Phytol. 2017, 217, 344. [Google Scholar] [CrossRef] [PubMed]
- Li, A.X.; Han, Y.Y.; Wang, X.; Chen, Y.H.; Zhao, M.R.; Zhou, S.M.; Wang, W. Root-specific expression of wheat expansin gene TaEXPB23 enhances root growth and water stress tolerance in tobacco. Environ. Exp. Bot. 2014, 110, 73–84. [Google Scholar] [CrossRef]
- Hao, Z.; Qian, X.; Xiao, X.; Liu, H.; Zhi, J.; Xu, J. Transgenic tobacco plants expressing grass AstEXPA1 gene show improved performance to several stresses. Plant Biotechnol. Rep. 2017, 11, 1–7. [Google Scholar] [CrossRef]
- Chen, Y.; Ren, Y.; Zhang, G.; An, J.; Yang, J.; Wang, Y.; Wang, W. Overexpression of the wheat expansin gene TaEXPA2 improves oxidative stress tolerance in transgenic Arabidopsis plants. Plant Physiol. Biochem. 2018, 124, 190–198. [Google Scholar] [CrossRef]
- He, X.; Zeng, J.; Cao, F.; Ahmed, I.M.; Zhang, G.; Vincze, E.; Wu, F. HvEXPB7, a novel β-expansin gene revealed by the root hair transcriptome of Tibetan wild barley, improves root hair growth under drought stress. J. Exp. Bot. 2015, 66, 7405–7419. [Google Scholar] [CrossRef]
- Jefferson, R.A.; Kavanagh, T.A.; Bevan, M.W. GUS fusions: Beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar] [CrossRef]
- Ren, Y.; Chen, Y.; An, J.; Zhao, Z.; Zhang, G.; Wang, Y.; Wang, W. Wheat expansin gene TaEXPA2 is involved in conferring plant tolerance to Cd toxicity. Plant Sci. 2018, 270, 245. [Google Scholar] [CrossRef]
- Rédei, G.P. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar]
- Nicholas, K.B. GeneDoc: Analysis and visualization of genetic variation. Embnew News 1997, 4, 14. [Google Scholar]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol. Biol. Evol. 2007, 24, 1596. [Google Scholar] [CrossRef]
- Arnold, K.; Bordoli, L.; Kopp, J.; Schwede, T. The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics 2006, 22, 195–201. [Google Scholar] [CrossRef]
- Schwede, T.; Kopp, J.; Guex, N.; Peitsch, M.C. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res. 2003, 31, 3381–3385. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Zhang, L.; Hao, W.; Zhang, L.; Liu, Y.; Chen, L. Expression of Two α-Type Expansins from Ammopiptanthus nanus in Arabidopsis thaliana Enhance Tolerance to Cold and Drought Stresses. Int. J. Mol. Sci. 2019, 20, 5255. https://doi.org/10.3390/ijms20215255
Liu Y, Zhang L, Hao W, Zhang L, Liu Y, Chen L. Expression of Two α-Type Expansins from Ammopiptanthus nanus in Arabidopsis thaliana Enhance Tolerance to Cold and Drought Stresses. International Journal of Molecular Sciences. 2019; 20(21):5255. https://doi.org/10.3390/ijms20215255
Chicago/Turabian StyleLiu, Yanping, Li Zhang, Wenfang Hao, Ling Zhang, Yi Liu, and Longqing Chen. 2019. "Expression of Two α-Type Expansins from Ammopiptanthus nanus in Arabidopsis thaliana Enhance Tolerance to Cold and Drought Stresses" International Journal of Molecular Sciences 20, no. 21: 5255. https://doi.org/10.3390/ijms20215255
APA StyleLiu, Y., Zhang, L., Hao, W., Zhang, L., Liu, Y., & Chen, L. (2019). Expression of Two α-Type Expansins from Ammopiptanthus nanus in Arabidopsis thaliana Enhance Tolerance to Cold and Drought Stresses. International Journal of Molecular Sciences, 20(21), 5255. https://doi.org/10.3390/ijms20215255




