Answer to Controversy: miR-10a Replacement Approaches Do Not Offer Protection against Chemotherapy-Induced Gonadotoxicity in Mouse Model
Abstract
1. Introduction
2. Results
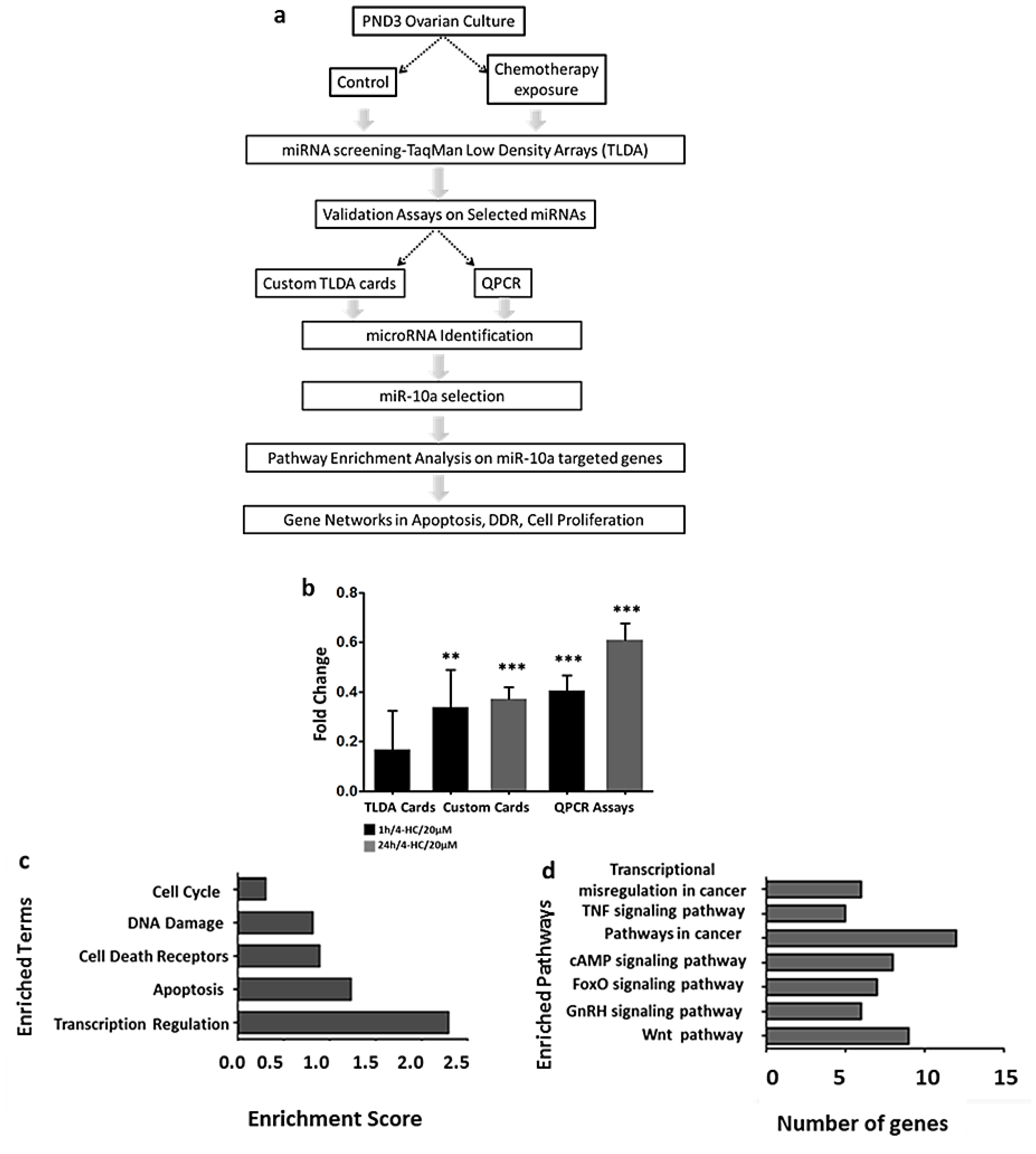
2.1. MiR-10a is Significantly Downregulated in Mouse Postnatal Ovaries in Response to 4-Hydroxyperoxycyclophosphamide (4-HC)
2.2. MiR-10a Can Regulate a Big Network of Genes Involved in DNA Damage Response, Apoptosis, and Follicle Activation
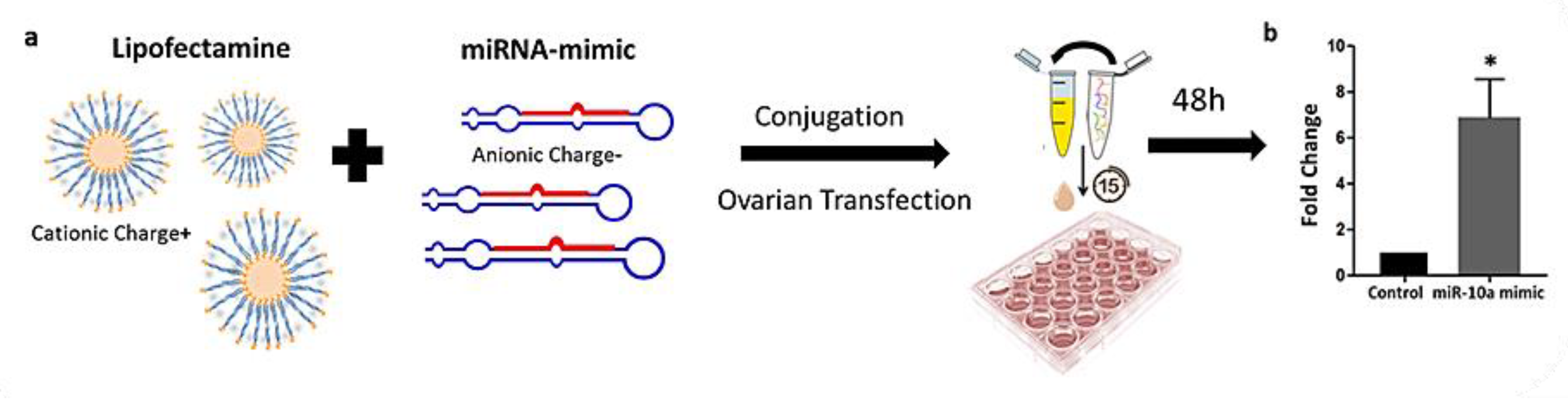
2.3. Liposome-Based Transfection of PND3 Ovaries with miR-10 Mimic
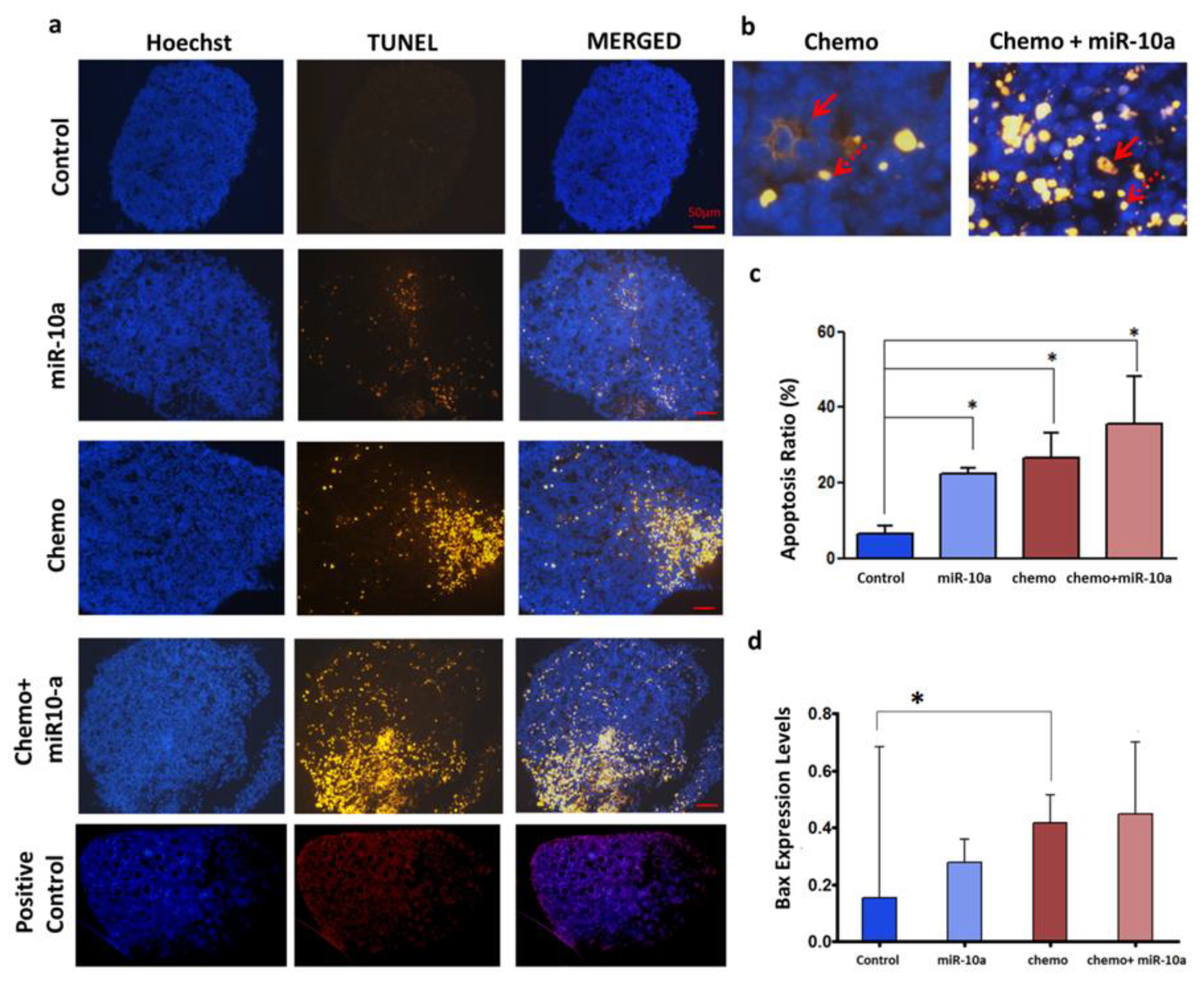
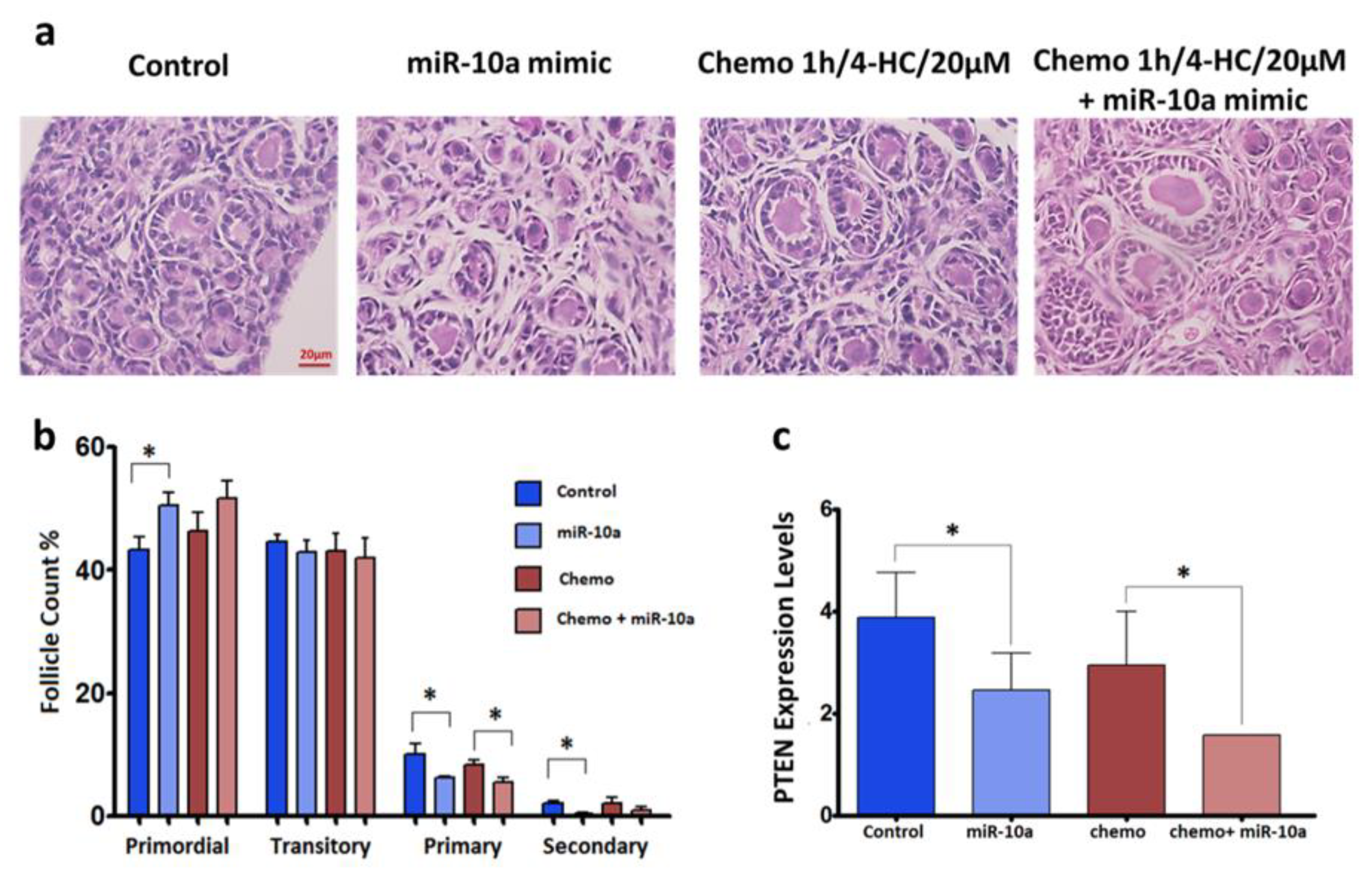
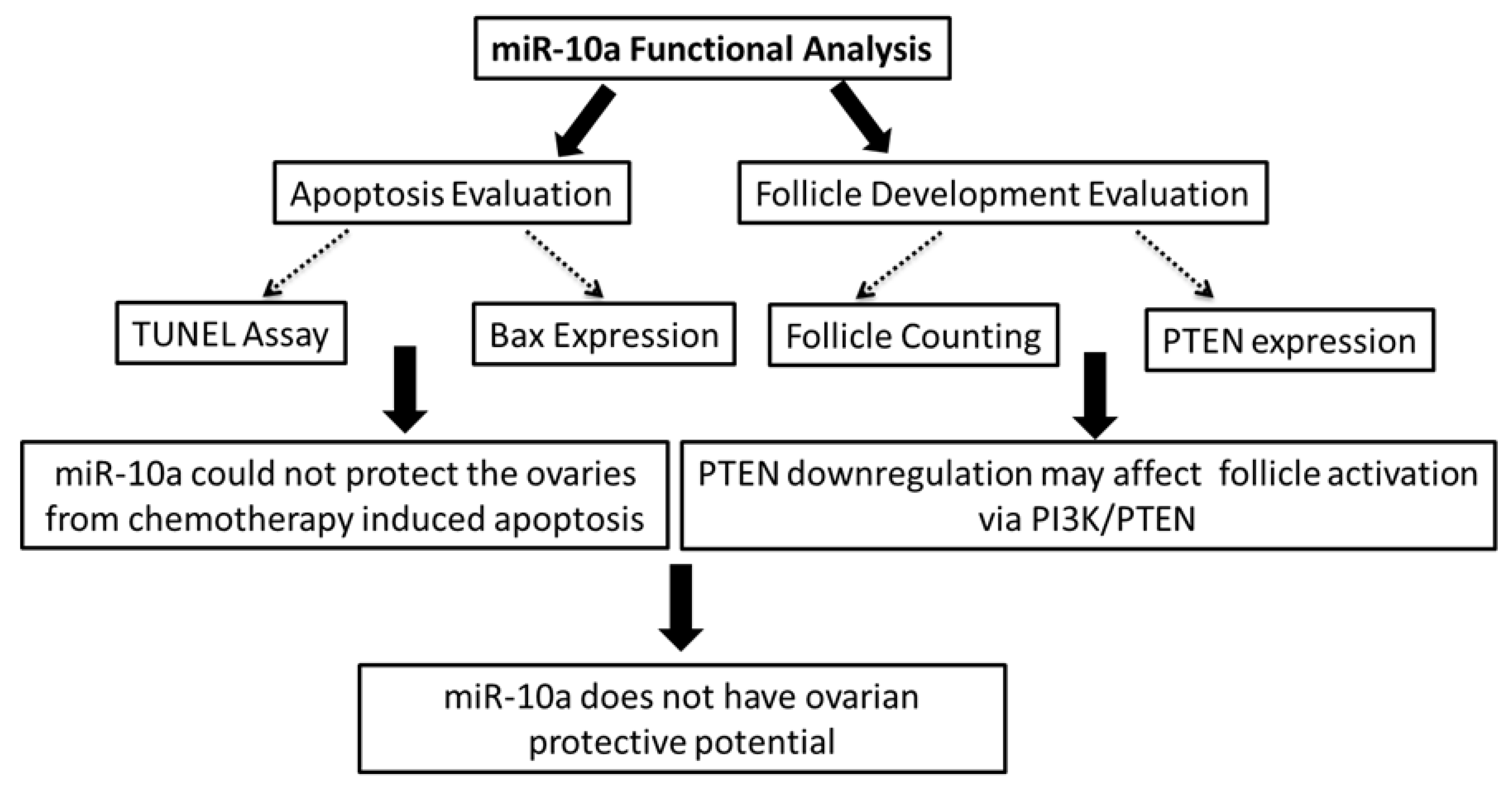
2.4. Mir-10a Restoration Does Not Protect the PND3 Mouse Ovaries after Exposure to 4-HC
2.5. MiR-10a is Involved in the PI3K/PTEN Signaling Pathway
3. Discussion
4. Materials and Methods
4.1. Mice
4.2. Culture of Mouse Ovaries in Vitro
4.3. Liposome-Mediated Transfection with Synthetic miR-10a Mimic
4.4. In Silico Gene Expression Analysis
4.5. Histological Studies and Follicle Counts
4.6. TUNEL Assay
4.7. MicroRNA Extraction and Reverse Transcription
4.8. MicroRNA Quantification and Expression Analysis
4.9. Gene Expression Quantification by QPCR
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AFSCs | Amniotic Fluid Stem Cells |
| PND3 | Post-natal day 3 |
| PTEN | Phosphatase and Tensin Homolog |
| 4-HC | 4-hydroxyperoxycyclophosphamide |
| QPCR | Quantitive Polymerase Chain Reaction |
| TLDA | TaqMan Low-Density Arrays |
| DDR | DNA Damage Response |
| TUNEL | Terminal deoxynucleotidyl transferase dUTP nick end labeling |
References
- Demeestere, I.; Basso, O.; Moffa, F.; Peccatori, F.; Poirot, C.; Shalom-Paz, E. Fertility preservation in female cancer patients. Obstet. Gynecol. 2012, 2012, 695041. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rodriguez-Wallberg, K.A.; Oktay, K. Options on fertility preservation in female cancer patients. Cancer Treat. Rev. 2012, 38, 354–361. [Google Scholar] [CrossRef] [PubMed]
- Wahid, F.; Shehzad, A.; Khan, T.; Kim, Y.Y. MicroRNAs: Synthesis, mechanism, function, and recent clinical trials. Biochim Biophys Acta - Mol. Cell Res. 2010, 1803, 1231–1243. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol 2018, 9, 402. [Google Scholar] [CrossRef] [PubMed]
- Mohr, A.; Mott, J. Overview of MicroRNA Biology. Semin. Liver Dis. 2015, 35, 003–011. [Google Scholar] [CrossRef] [PubMed]
- Reza, A.M.M.T.; Choi, Y.-J.; Han, S.G.; Song, H.; Park, C.; Hong, K.; Kim, J.H. Roles of microRNAs in mammalian reproduction: from the commitment of germ cells to peri-implantation embryos. Biol. Rev. 2019, 94, 415–438. [Google Scholar] [CrossRef] [PubMed]
- McBride, D.; Carré, W.; Sontakke, S.D.; Hogg, C.O.; Law, A.; Donadeu, F.X.; Clinton, M. Identification of miRNAs associated with the follicular–luteal transition in the ruminant ovary. Reproduction 2012, 144, 221–233. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Sun, J.; Lai, D. Role of microRNAs in premature ovarian insufficiency. Reprod. Biol. Endocrinol. 2017, 15, 38. [Google Scholar] [CrossRef]
- Toloubeydokhti, T.; Bukulmez, O.; Chegini, N. Potential Regulatory Functions of MicroRNAs in the Ovary. Semin. Reprod. Med. 2008, 26, 469–478. [Google Scholar] [CrossRef]
- Chakraborty, C.; Sharma, A.R.; Sharma, G.; Sarkar, B.K.; Lee, S.-S.; Chakraborty, C. The novel strategies for next-generation cancer treatment: miRNA combined with chemotherapeutic agents for the treatment of cancer. Oncotarget 2018, 9, 10164–10174. [Google Scholar]
- Viteri, S.; Rosell, R. An innovative mesothelioma treatment based on miR-16 mimic loaded EGFR targeted minicells (TargomiRs). Transl. Lung Cancer Res. 2018, 7, 1–4. [Google Scholar] [CrossRef]
- Penso-Dolfin, L.; Moxon, S.; Haerty, W.; Di Palma, F. The evolutionary dynamics of microRNAs in domestic mammals. Sci. Rep. 2018, 8, 17050. [Google Scholar] [CrossRef] [PubMed]
- Friedman, R.C.; Farh, K.K.-H.; Burge, C.B.; Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009, 19, 92–105. [Google Scholar] [CrossRef] [PubMed]
- Tulay, P.; Naja, R.P.; Cascales-Roman, O.; Doshi, A.; Serhal, P.; SenGupta, S.B. Investigation of microRNA expression and DNA repair gene transcripts in human oocytes and blastocysts. J. Assist. Reprod. Genet. 2015, 32, 1757–1764. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, J.; Xu, Y.; Liu, H.; Pan, Z. MicroRNAs in ovarian follicular atresia and granulosa cell apoptosis. Reprod. Biol. Endocrinol. 2019, 17, 9. [Google Scholar] [CrossRef] [PubMed]
- Kalich-Philosoph, L.; Roness, H.; Carmely, A.; Fishel-Bartal, M.; Ligumsky, H.; Paglin, S.; Wolf, I.; Kanety, H.; Sredni, B.; Meirow, D.; et al. Cyclophosphamide Triggers Follicle Activation and “Burnout”; AS101 Prevents Follicle Loss and Preserves Fertility. Sci. Transl. Med. 2013, 15, 185ra62. [Google Scholar] [CrossRef] [PubMed]
- Kong, Y.W.; Ferland-McCollough, D.; Jackson, T.J.; Bushell, M. microRNAs in cancer management. Lancet Oncol. 2012, 13, 249–258. [Google Scholar] [CrossRef]
- Yu, X.; Chen, Y.; Tian, R.; Li, J.; Li, H.; Lv, T.; Yao, Q. miRNA-21 enhances chemoresistance to cisplatin in epithelial ovarian cancer by negatively regulating PTEN. Oncol. Lett. 2017, 14, 1807–1810. [Google Scholar] [CrossRef] [PubMed]
- Blower, P.E.; Chung, J.-H.; Verducci, J.S.; Lin, S.; Park, J.-K.; Dai, Z.; Liu, C.G.; Schmittgen, T.D.; Reinhold, W.C.; Croce, C.M.; et al. MicroRNAs modulate the chemosensitivity of tumor cells. Mol. Cancer Ther. 2008, 7, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Tehler, D.; Høyland-Kroghsbo, N.M.; Lund, A.H. The miR-10 microRNA precursor family. RNA Biol. 2011, 8, 728–734. [Google Scholar] [CrossRef] [PubMed]
- Tanzer, A.; Amemiya, C.T.; Kim, C.-B.; Stadler, P.F. Evolution of microRNAs located within Hox gene clusters. J. Exp. Zool Part. B Mol. Dev. Evol. 2005, 304B, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Lemons, D.; McGinnis, W. Genomic Evolution of Hox Gene Clusters. Science 2006, 313, 1918–1922. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Teruya-Feldstein, J.; Weinberg, R.A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 2007, 449, 682–688. [Google Scholar] [CrossRef]
- Sasayama, T.; Nishihara, M.; Kondoh, T.; Hosoda, K.; Kohmura, E. MicroRNA-10b is overexpressed in malignant glioma and associated with tumor invasive factors, uPAR and RhoC. Int. J. Cancer 2009, 125, 1407–1413. [Google Scholar] [CrossRef]
- Chai, G.; Liu, N.; Ma, J.; Li, H.; Oblinger, J.L.; Prahalad, A.K.; Gong, M.; Chang, L.-S.; Wallace, M.; Muir, D.; et al. MicroRNA-10b regulates tumorigenesis in neurofibromatosis type 1. Cancer Sci. 2010, 101, 1997–2004. [Google Scholar] [CrossRef]
- Tian, Y.; Luo, A.; Cai, Y.; Su, Q.; Ding, F.; Chen, H.; Liu, Z. MicroRNA-10b Promotes Migration and Invasion through KLF4 in Human Esophageal Cancer Cell Lines. J. Biol. Chem. 2010, 285, 7986–7994. [Google Scholar] [CrossRef] [PubMed]
- Nakata, K.; Ohuchida, K.; Mizumoto, K.; Kayashima, T.; Ikenaga, N.; Sakai, H.; Lin, C.; Fujita, H.; Otsuka, T.; Aishima, S.; et al. MicroRNA-10b is overexpressed in pancreatic cancer, promotes its invasiveness, and correlates with a poor prognosis. Surgery 2011, 150, 916–922. [Google Scholar] [CrossRef] [PubMed]
- Balatti, V.; Oghumu, S.; Bottoni, A.; Maharry, K.; Cascione, L.; Fadda, P.; Parwani, A.; Croce, C.; Iwenofu, O.H. MicroRNA Profiling of Salivary Duct Carcinoma Versus Her2/Neu Overexpressing Breast Carcinoma Identify miR-10a as a Putative Breast Related Oncogene. Head Neck Pathol. 2018, 13, 344–354. [Google Scholar] [CrossRef]
- Xiao, G.-Y.; Cheng, C.-C.; Chiang, Y.-S.; Cheng, W.T.-K.; Liu, I.-H.; Wu, S.-C. Exosomal miR-10a derived from amniotic fluid stem cells preserves ovarian follicles after chemotherapy. Scientific Reports 2016, 6, 23120. [Google Scholar] [CrossRef] [PubMed]
- Jiajie, T.; Yanzhou, Y.; Hoi-Hung, A.C.; Zi-Jiang, C.; Wai-Yee, C. Conserved miR-10 family represses proliferation and induces apoptosis in ovarian granulosa cells. Sci. Rep. 2017, 7, 41304. [Google Scholar] [CrossRef] [PubMed]
- Sirotkin, A.V.; Marcela, L.; Dmitriy, O.; Pauline, B.; Miloš, M. Identification of MicroRNAs controlling human ovarian cell proliferation and apoptosis. J. Cell Physiol. 2010, 223, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Alexandri, C.; Stamatopoulos, B.; Rothé, F.; Bareche, Y.; Devos, M.; Demeestere, I. MicroRNA profiling and identification of let-7a as a target to prevent chemotherapy-induced primordial follicles apoptosis in mouse ovaries. Sci. Rep. 2019, 9, 9636. [Google Scholar] [CrossRef] [PubMed]
- Desmeules, P.; Devine, P.J. Characterizing the Ovotoxicity of Cyclophosphamide Metabolites on Cultured Mouse Ovaries. Toxicol. Sci. 2006, 90, 500–509. [Google Scholar] [CrossRef] [PubMed]
- Horicks, F.; Van Den Steen, G.; Houben, S.; Englert, Y.; Demeestere, I. Folliculogenesis Is Not Fully Inhibited during GnRH Analogues Treatment in Mice Challenging Their Efficiency to Preserve the Ovarian Reserve during Chemotherapy in This Model. PLoS ONE 2015, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Wu, W.J.; Zhou, X.L.; Xiao, P.; Wang, Y.; Liu, H.L. Expression and Preliminary Functional Profiling of the let-7 Family during Porcine Ovary Follicle Atresia. Mol. Cells 2015, 38, 304–311. [Google Scholar] [CrossRef] [PubMed]
- Tu, J.; Cheung, H.-H.; Lu, G.; Chen, Z.; Chan, W.-Y. MicroRNA-10a promotes granulosa cells tumor development via PTEN-AKT/Wnt regulatory axis. Cell Death Dis. 2018, 9, 1076. [Google Scholar] [CrossRef] [PubMed]
- Morgan, S.; Anderson, R.A.; Gourley, C.; Wallace, W.H.; Spears, N. How do chemotherapeutic agents damage the ovary? Hum. Reprod. Update 2012, 18, 525–535. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Huang, Y.; Yu, Y.; Xin, X. GnRH antagonist cetrorelix inhibits mitochondria-dependent apoptosis triggered by chemotherapy in granulosa cells of rats. Gynecol. Oncol. 2010, 118, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Perez, G.I.; Knudson, C.M.; Leykin, L.; Korsmeyer, S.J.; Tilly, J.L. Apoptosis-associated signaling pathways are required for chemotherapy-mediated female germ cell destruction. Nat. Med. 1997, 3, 1228–1232. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, Q.-N.; Zerafa, N.; Liew, S.H.; Morgan, F.H.; Strasser, A.; Scott, C.L.; Findlay, J.K.; Hickey, M.; Hutt, K.J. Loss of PUMA protects the ovarian reserve during DNA-damaging chemotherapy and preserves fertility. Cell Death Dis. 2018, 9, 618. [Google Scholar] [CrossRef]
- Liu, S.; Sun, J.; Lan, Q. TGF-β-induced miR10a/b expression promotes human glioma cell migration by targeting PTEN. Mol. Med. Rep. 2013, 8, 1741–1746. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.-H.; Shrestha, S.; Yang, C.-D.; Chang, N.-W.; Lin, Y.-L.; Liao, K.-W.; Huang, W.-C.; Sun, T.-H.; Tu, S.-J.; Lee, W.-H.; et al. miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2018, 46, D296–D302. [Google Scholar] [CrossRef] [PubMed]
- Gougeon, A. Regulation of Ovarian Follicular Development in Primates: Facts and Hypotheses. Endocr. Rev. 1996, 17, 121–155. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alexandri, C.; Stratopoulou, C.-A.; Demeestere, I. Answer to Controversy: miR-10a Replacement Approaches Do Not Offer Protection against Chemotherapy-Induced Gonadotoxicity in Mouse Model. Int. J. Mol. Sci. 2019, 20, 4958. https://doi.org/10.3390/ijms20194958
Alexandri C, Stratopoulou C-A, Demeestere I. Answer to Controversy: miR-10a Replacement Approaches Do Not Offer Protection against Chemotherapy-Induced Gonadotoxicity in Mouse Model. International Journal of Molecular Sciences. 2019; 20(19):4958. https://doi.org/10.3390/ijms20194958
Chicago/Turabian StyleAlexandri, Chrysanthi, Christina-Anna Stratopoulou, and Isabelle Demeestere. 2019. "Answer to Controversy: miR-10a Replacement Approaches Do Not Offer Protection against Chemotherapy-Induced Gonadotoxicity in Mouse Model" International Journal of Molecular Sciences 20, no. 19: 4958. https://doi.org/10.3390/ijms20194958
APA StyleAlexandri, C., Stratopoulou, C.-A., & Demeestere, I. (2019). Answer to Controversy: miR-10a Replacement Approaches Do Not Offer Protection against Chemotherapy-Induced Gonadotoxicity in Mouse Model. International Journal of Molecular Sciences, 20(19), 4958. https://doi.org/10.3390/ijms20194958





