Pyruvate Substitutions on Glycoconjugates
Abstract
1. Introduction
2. Enol-Pyruvylation in Peptidoglycan Biosynthesis
3. Ketal-Pyruvylated Glycoconjugates
3.1. Exopolysaccharides
3.1.1. Xanthan
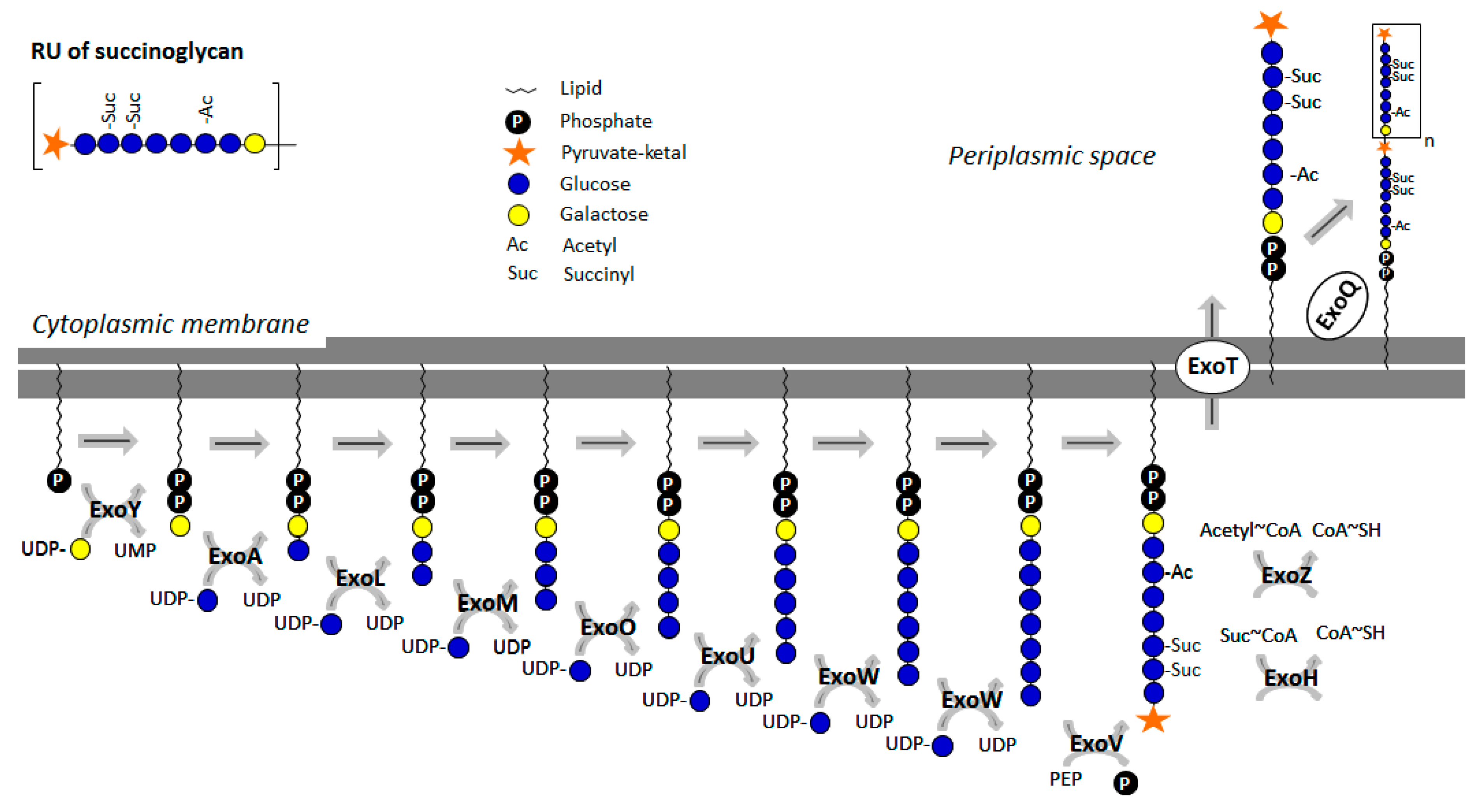
3.1.2. Succinoglycan
3.1.3. Salecan
3.1.4. Colonic Acid
3.1.5. Unclassified Pyruvylated EPS
3.2. Capsular Polysaccharides
3.2.1. Streptococcus pneumoniae CPS
3.2.2. Acinetobacter baumannii CPS
3.2.3. Klebsiella CPSs
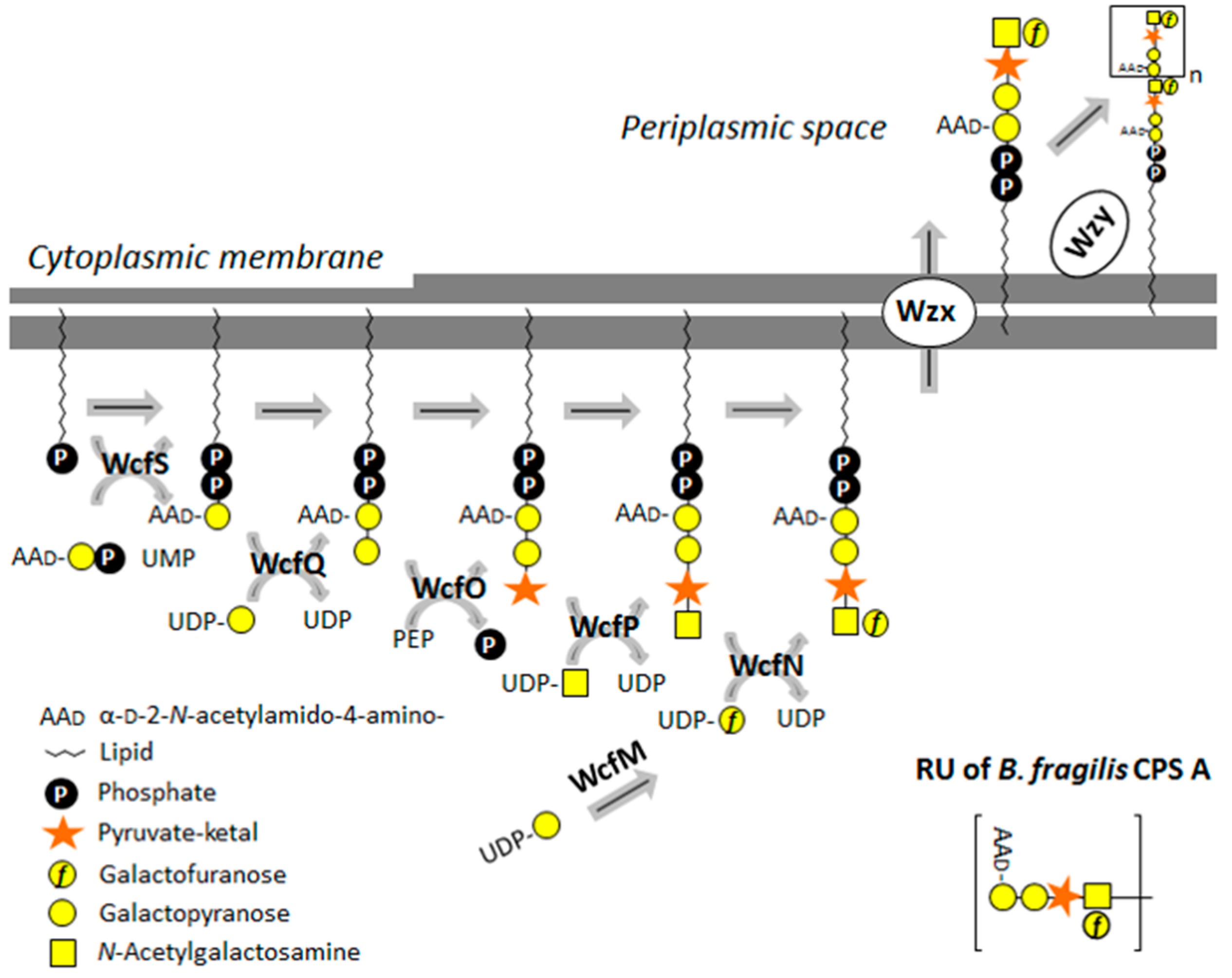
3.2.4. Bacteroides fragilis CPS A
3.2.5. Rhodococcus equi CPS
3.3. “Non-Classical” Secondary Cell Wall Glycopolymers with Pyruvylated β-d-ManNAc
3.3.1. Bacillus anthracis SCWP
3.3.2. Paenibacillus alvei SCWP
3.3.3. Cell Wall Polysaccharide of Paenibacillus polymyxa
3.4. SCWPs with Other Pyruvylated Sugar Epitopes
3.4.1. SCWPs from the Genus Promicromonospora
3.4.2. Teichoic Acids from the Genus Nocardiopsis
3.4.3. Teichoic Acids of Brevibacterium iodinum
3.5. Lipopolysaccharides and Lipooligosaccharides
3.5.1. Pseudomonas stutzeri
3.5.2. Providencia alcalifaciens
3.5.3. Shigella dysenteriae
3.5.4. Raoultella terrigena
3.5.5. Proteus mirabilis
3.5.6. Cobetia pacifica
3.6. Mycobacterial Glycolipids with Pyruvate
3.7. Pyruvylated Glycoconjugates in Eukaryotes
3.7.1. Eukaryotic Glycolipids
3.7.2. N-Linked Glycans in Yeast
3.7.3. Pyruvylated Galactans of Algae
3.7.4. Pyruvylated Proteoglycan
4. Methods for Research of Pyruvylated Glycoconjugates
4.1. Isolation of Pyruvylated Bacterial Glycoconjugates
4.2. Pyruvate Analytics
4.2.1. Lectin Approach
4.2.2. Biochemical Pyruvate Assays
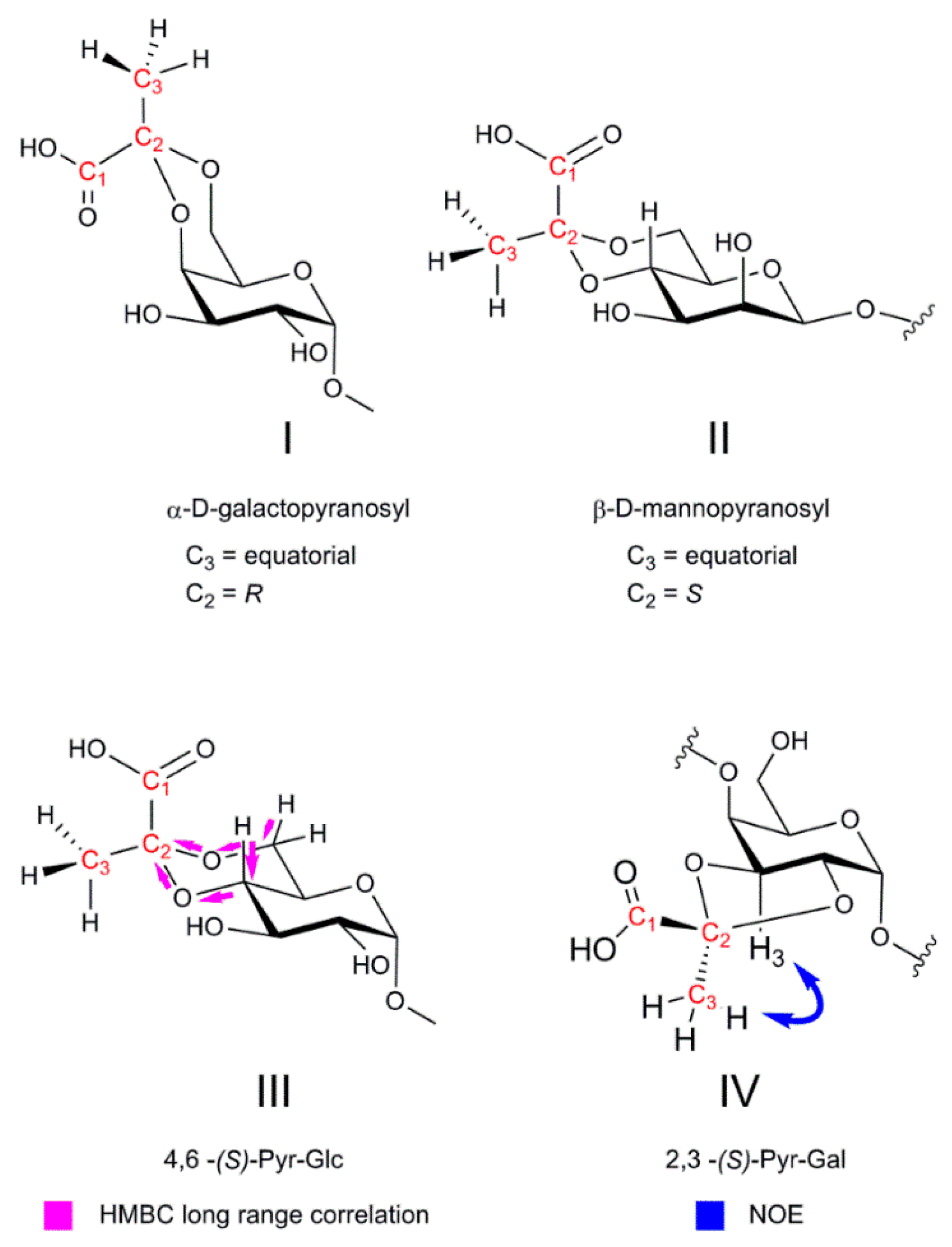
4.2.3. NMR Analysis of Pyruvylation
4.2.4. MS analysis of Pyruvylation
5. Ketal-Pyruvyltransferases
5.1. Substrate Specificity of Ketal-Pyruvyltransferases
5.1.1. CsaB from P. alvei
5.1.2. WcfO from B. fragilis
5.1.3. Pvg1p from S. pombe
5.2. Challenges in Research of Ketal-Pyruvyltransferases
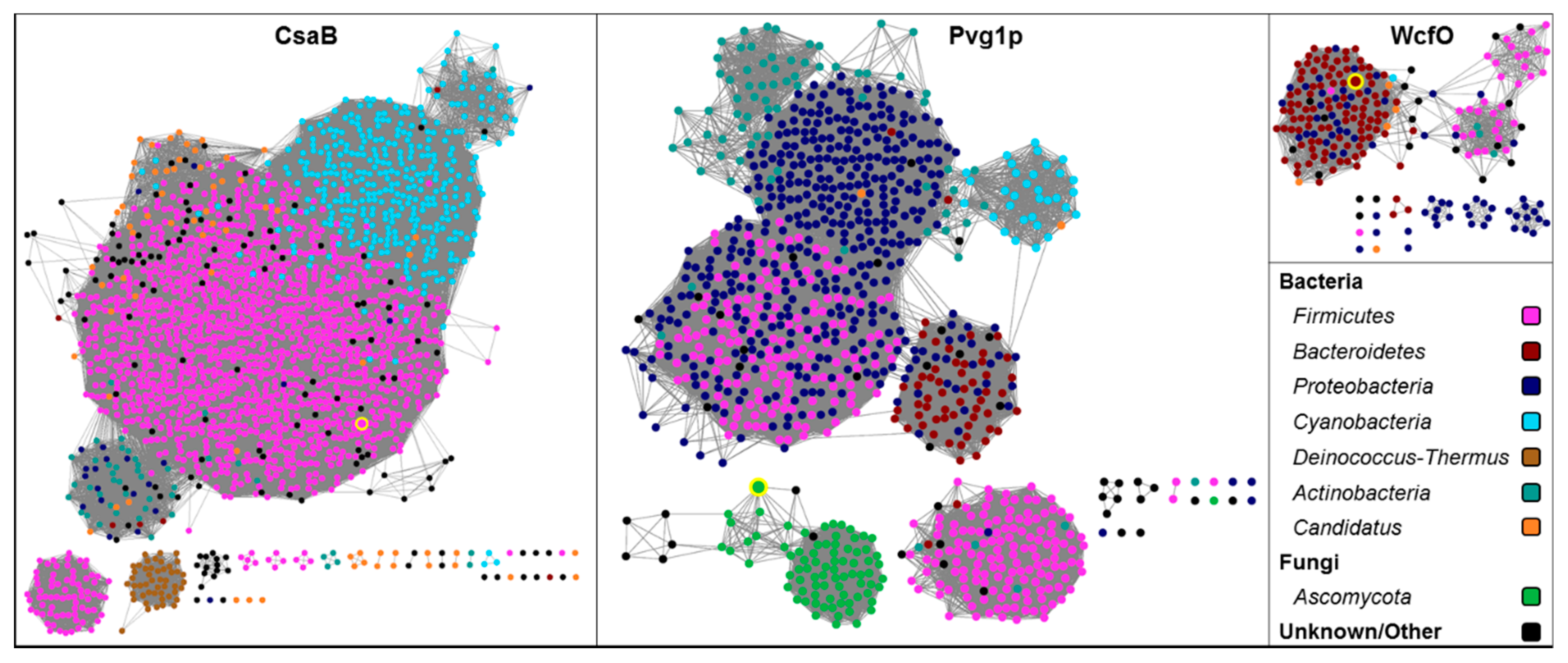
5.3. Sequence Space of Ketal-Pyruvyltransferases
5.3.1. Methods
5.3.2. Results
6. Discussion
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| 4,6Pyr | 4,6-ketal-pyruvate |
| AADGalp | α-d-2-N-acetylamido-4-amino-galactopyranose |
| Ac | acetyl |
| Am | aecetamido |
| CA | colonic acid |
| Cm | carbamoyl |
| CPS | capsular polysaccharide |
| CSDB | Carbohydrate Structure Database |
| Cys | cysteine |
| D | aspartic acid |
| DPNH | reduced diphosphopyridine nucleotide |
| EPS | exopolysaccharide |
| ER | endoplasmic reticulum |
| Ery-ol | erythritol |
| ESI-MS | electrospray ionization mass spectrometry |
| f | furanosidic |
| Fuc | fucose |
| Gal | galactose |
| GalN | galactosamine |
| GalNAc | N-acetylgalactosamine |
| GalNAcA | α-N-acetyl-d-galactosamine uronic acid |
| GC | gas chromatography |
| Glc | glucose |
| GlcA | glucuronic acid |
| GlcNAc | N-acetylglucosamine |
| GOESY | gradient nuclear Overhauser and exchange spectroscopy |
| GT | glycosyltransferase |
| H | histidine |
| HMBC | hetero multiple bond correlation |
| HMM | Hidden Markow Model |
| HPLC | high-performance liquid chromatography |
| Kdn | 2-keto-3-deoxy-d-glycero-d-galacto-nononic acid |
| Kdo | 3-deoxy-d-manno-oct-2-ulosonic acid |
| L | leucine |
| LOS | lipooligosaccharide |
| LPS | lipopolysaccharide |
| LTA | lipoteichoic acid |
| MAI complex | Mycobacterium avium-Mycobacterium intracellulare complex |
| Man | mannose |
| Man-ol | mannitol |
| ManNAc | N-acetylmannosamine |
| Me | methyl |
| MOPDG | 4,6Pyr-OMe-β-d-Galp |
| MurNAc | N-acetylmuramic acid |
| NOESY | nuclear Overhauser and exchange spectroscopy |
| p | pyranosidic |
| P | phosphate |
| PEP | phosphoenolpyruvate |
| PEtN | phosphoetanolamine |
| pNP | para-nitrophenyl |
| Pyr | pyruvate |
| pyrGal-ase | pyruvylated galactose-releasing enzyme |
| QuiNAc | N-acetylquinovosamine |
| R | arginine |
| Rha | rhamnose |
| Rho | 5-amino-3,5-dideoxynonulosonic (rhodaminic) acid |
| Rib-ol | ribitol |
| ROESY | rotating frame Overhauser enhancement spectroscopy |
| RP | reversed phase |
| RU | repeating unit |
| SAP | plasma glycoprotein serum amyloid P component |
| SCWP | secondary cell wall glycopolymer |
| Ser | serine |
| SLH | surface layer homology |
| sp. | species |
| spp. | species (plural) |
| TA | teichoic acid |
| undp-P | undecaprenylphosphate |
| WTA | wall teichoic acid |
References
- Bennet, L.G.; Bishop, C.T. The pyruvate ketal as a stereospecific immunodeterminant in the type XXVII Streptococcus pneumoniae (pneumococcal) capsular polysaccharide. Immunochemistry 1977, 14, 693–696. [Google Scholar] [CrossRef]
- Lovering, A.L.; Safadi, S.S.; Strynadka, N.C. Structural perspective of peptidoglycan biosynthesis and assembly. Annu. Rev. Biochem. 2012, 81, 451–478. [Google Scholar] [CrossRef] [PubMed]
- Andreishcheva, E.N.; Kunkel, J.P.; Gemmill, T.R.; Trimble, R.B. Five genes involved in biosynthesis of the pyruvylated Galβ1,3-epitope in Schizosaccharomyces pombe N-linked glycans. J. Biol. Chem. 2004, 279, 35644–35655. [Google Scholar] [CrossRef] [PubMed]
- Leone, S.; Izzo, V.; Silipo, A.; Sturiale, L.; Garozzo, D.; Lanzetta, R.; Parrilli, M.; Molinaro, A.; Di Donato, A. A novel type of highly negatively charged lipooligosaccharide from Pseudomonas stutzeri OX1 possessing two 4,6-O-(1-carboxy)-ethylidene residues in the outer core region. Eur. J. Biochem. 2004, 271, 2691–2704. [Google Scholar] [CrossRef] [PubMed]
- Toukach, P.V.; Egorova, K.S. Carbohydrate structure database merged from bacterial, archaeal, plant and fungal parts. Nucleic Acids Res. 2016, 44, D1229–D1236. [Google Scholar] [CrossRef]
- Toukach, P.V.; Egorova, K.S. Bacterial, plant, and fungal carbohydrate structure databases: Daily usage. Methods Mol. Biol. 2015, 1273, 55–85. [Google Scholar] [PubMed]
- Sharma, S.; Erickson, K.M.; Troutman, J.M. Complete tetrasaccharide repeat unit biosynthesis of the immunomodulatory Bacteroides fragilis capsular polysaccharide A. ACS Chem. Biol. 2017, 12, 92–101. [Google Scholar] [CrossRef]
- Geissner, A.; Pereira, C.L.; Leddermann, M.; Anish, C.; Seeberger, P.H. Deciphering antigenic determinants of Streptococcus pneumoniae serotype 4 capsular polysaccharide using synthetic oligosaccharides. ACS Chem. Biol. 2016, 11, 335–344. [Google Scholar] [CrossRef]
- Gemmill, T.R.; Trimble, R.B. Schizosaccharomyces pombe produces novel pyruvate-containing N-linked oligosaccharides. J. Biol. Chem. 1996, 271, 25945–25949. [Google Scholar] [CrossRef]
- Chiovittia, A.; Bacica, A.; Craik, D.J.; Munro, S.L.A.; Kraft, G.T.; Liao, M.-L. Cell-wall polysaccharides from Australian red algae of the family Solieriaceae (Gigartinales, Rhodophyta): Novel, highly pyruvated carrageenans from the genus Callophycus. Carbohydr. Res. 1997, 299, 229–243. [Google Scholar] [CrossRef]
- Arata, P.X.; Quintana, I.; Canelon, D.J.; Vera, B.E.; Compagnone, R.S.; Ciancia, M. Chemical structure and anticoagulant activity of highly pyruvylated sulfated galactans from tropical green seaweeds of the order Bryopsidales. Carbohydr. Polym. 2015, 122, 376–386. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Mao, W.; Liu, X.; Wang, S.; Xia, Z.; Cao, S.; Li, L.; Zhang, Q.; Liu, S. Sequence analysis of the pyruvylated galactan sulfate-derived oligosaccharides by negative-ion electrospray tandem mass spectrometry. Carbohydr. Res. 2016, 433, 80–88. [Google Scholar] [CrossRef] [PubMed]
- Zibetti, R.G.; Duarte, M.E.; Noseda, M.D.; Colodi, F.G.; Ducatti, D.R.; Ferreira, L.G.; Cardoso, M.A.; Cerezo, A.S. Galactans from Cryptonemia species. Part II: Studies on the system of galactans of Cryptonemia seminervis (Halymeniales) and on the structure of major fractions. Carbohydr. Res. 2009, 344, 2364–2374. [Google Scholar] [CrossRef] [PubMed]
- Forsberg, L.S.; Abshire, T.G.; Friedlander, A.; Quinn, C.P.; Kannenberg, E.L.; Carlson, R.W. Localization and structural analysis of a conserved pyruvylated epitope in Bacillus anthracis secondary cell wall polysaccharides and characterization of the galactose-deficient wall polysaccharide from avirulent B. anthracis CDC 684. Glycobiology 2012, 22, 1103–1117. [Google Scholar] [CrossRef] [PubMed]
- Schäffer, C.; Müller, N.; Mandal, P.K.; Christian, R.; Zayni, S.; Messner, P. A pyrophosphate bridge links the pyruvate-containing secondary cell wall polymer of Paenibacillus alvei CCM 2051 to muramic acid. Glycoconj. J. 2000, 17, 681–690. [Google Scholar] [CrossRef] [PubMed]
- Khatun, F.; Stephenson, R.J.; Toth, I. An overview of structural features of antibacterial glycoconjugate vaccines that influence their immunogenicity. Chemistry 2017, 23, 4233–4254. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.T.; Oh, S.Y.; Hendrickx, A.P.; Lunderberg, J.M.; Schneewind, O. Bacillus cereus G9241 S-layer assembly contributes to the pathogenesis of anthrax-like disease in mice. J. Bacteriol. 2013, 195, 596–605. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bianco, M.I.; Toum, L.; Yaryura, P.M.; Mielnichuk, N.; Gudesblat, G.E.; Roeschlin, R.; Marano, M.R.; Ielpi, L.; Vojnov, A.A. Xanthan pyruvilation is essential for the virulence of Xanthomonas campestris pv. campestris. Mol. Plant. Microbe Interact. 2016, 29, 688–699. [Google Scholar] [CrossRef]
- Silver, L.L. Fosfomycin: Mechanism and resistance. Cold Spring Harb Perspect. Med. 2017, 7, a025262. [Google Scholar] [CrossRef]
- Eschenburg, S.; Priestman, M.; Schönbrunn, E. Evidence that the fosfomycin target Cys115 in UDP-N-acetylglucosamine enolpyruvyl transferase (MurA) is essential for product release. J. Biol. Chem. 2005, 280, 3757–3763. [Google Scholar] [CrossRef]
- Sonkar, A.; Shukla, H.; Shukla, R.; Kalita, J.; Pandey, T.; Tripathi, T. UDP-N-Acetylglucosamine enolpyruvyl transferase (MurA) of Acinetobacter baumannii (AbMurA): Structural and functional properties. Int. J. Biol. Macromol. 2017, 97, 106–114. [Google Scholar] [CrossRef] [PubMed]
- Falagas, M.E.; Vouloumanou, E.K.; Samonis, G.; Vardakas, K.Z. Fosfomycin. Clin. Microbiol. Rev. 2016, 29, 321–347. [Google Scholar] [CrossRef] [PubMed]
- Oberdorfer, G.; Binter, A.; Ginj, C.; Macheroux, P.; Gruber, K. Structural and functional characterization of NikO, an enolpyruvyl transferase essential in nikkomycin biosynthesis. J. Biol. Chem. 2012, 287, 31427–31436. [Google Scholar] [CrossRef] [PubMed]
- Cescutti, P. Chapter 6—Bacterial capsular polysaccharides and exopolysaccharides. In Microbial Glycobiology; Holst, O., Brennan, P.J., Itzstein, M.V., Moran, A.P., Eds.; Academic Press: San Diego, CA, USA, 2010; pp. 93–108. [Google Scholar]
- Stewart, P.S.; Costerton, J.W. Antibiotic resistance of bacteria in biofilms. Lancet 2001, 358, 135–138. [Google Scholar] [CrossRef]
- Freitas, F.; Alves, V.D.; Reis, M.A. Advances in bacterial exopolysaccharides: From production to biotechnological applications. Trends Biotechnol. 2011, 29, 388–398. [Google Scholar] [CrossRef] [PubMed]
- Aslam, S.N.; Newman, M.A.; Erbs, G.; Morrissey, K.L.; Chinchilla, D.; Boller, T.; Jensen, T.T.; De Castro, C.; Ierano, T.; Molinaro, A.; et al. Bacterial polysaccharides suppress induced innate immunity by calcium chelation. Curr. Biol. 2008, 18, 1078–1083. [Google Scholar] [CrossRef]
- Gansbiller, M.; Schmid, J.; Sieber, V. In-depth rheological characterization of genetically modified xanthan-variants. Carbohydr. Polym. 2019, 213, 236–246. [Google Scholar] [CrossRef]
- Becker, A.; Katzen, F.; Puhler, A.; Ielpi, L. Xanthan gum biosynthesis and application: A biochemical/genetic perspective. Appl. Microbiol. Biotechnol. 1998, 50, 145–152. [Google Scholar] [CrossRef]
- Jansson, P.E.; Kenne, L.; Lindberg, B. Structure of extracellular polysaccharide from Xanthomonas campestris. Carbohydr. Res. 1975, 45, 275–282. [Google Scholar] [CrossRef]
- Marzocca, M.P.; Harding, N.E.; Petroni, E.A.; Cleary, J.M.; Ielpi, L. Location and cloning of the ketal pyruvate transferase gene of Xanthomonas campestris. J. Bacteriol. 1991, 173, 7519–7524. [Google Scholar] [CrossRef]
- Schmid, J.; Sieber, V.; Rehm, B. Bacterial exopolysaccharides: Biosynthesis pathways and engineering strategies. Front. Microbiol. 2015, 6, 496. [Google Scholar] [CrossRef] [PubMed]
- Barreras, M.; Abdian, P.L.; Ielpi, L. Functional characterization of GumK, a membrane-associated beta-glucuronosyltransferase from Xanthomonas campestris required for xanthan polysaccharide synthesis. Glycobiology 2004, 14, 233–241. [Google Scholar] [CrossRef] [PubMed]
- Vorhölter, F.J.; Schneiker, S.; Goesmann, A.; Krause, L.; Bekel, T.; Kaiser, O.; Linke, B.; Patschkowski, T.; Ruckert, C.; Schmid, J.; et al. The genome of Xanthomonas campestris pv. campestris B100 and its use for the reconstruction of metabolic pathways involved in xanthan biosynthesis. J. Biotechnol. 2008, 134, 33–45. [Google Scholar]
- Nankai, H.; Hashimoto, W.; Miki, H.; Kawai, S.; Murata, K. Microbial system for polysaccharide depolymerization: Enzymatic route for xanthan depolymerization by Bacillus sp. strain GL1. Appl. Environ. Microbiol. 1999, 65, 2520–2526. [Google Scholar]
- Hashimoto, W.; Miki, H.; Tsuchiya, N.; Nankai, H.; Murata, K. Xanthan lyase of Bacillus sp. strain GL1 liberates pyruvylated mannose from xanthan side chains. Appl. Env. Microbiol. 1998, 64, 3765–3768. [Google Scholar]
- Thompson, M.A.; Onyeziri, M.C.; Fuqua, C. Function and regulation of Agrobacterium tumefaciens cell surface structures that promote attachment. Curr. Top. Microbiol. Immunol. 2018, 418, 143–184. [Google Scholar] [PubMed]
- Matthysse, A.G. Exopolysaccharides of Agrobacterium tumefaciens. Curr. Top. Microbiol. Immunol. 2018, 418, 111–141. [Google Scholar]
- Meade, M.J.; Tanenbaum, S.W.; Nakas, J.P. Production and rheological properties of a succinoglycan from Pseudomonas sp. 31260 grown on wood hydrolysates. Can. J. Microbiol. 1995, 41, 1147–1152. [Google Scholar] [CrossRef] [PubMed]
- Andhare, P.; Delattre, C.; Pierre, G.; Michaud, P.; Pathak, H. Characterization and rheological behaviour analysis of the succinoglycan produced by Rhizobium radiobacter strain CAS from curd sample. Food Hydrocoll. 2017, 64, 1–8. [Google Scholar] [CrossRef]
- Becker, A.; Rüberg, S.; Baumgarth, B.; Bertram-Drogatz, P.A.; Quester, I.; Pühler, A. Regulation of succinoglycan and galactoglucan biosynthesis in Sinorhizobium meliloti. J. Mol. Microbiol. Biotechnol. 2002, 4, 187–190. [Google Scholar]
- Stredansky, M. Succinoglycan. In Biopolymers Online; Wiley: New York, NY, USA, 2005. [Google Scholar]
- Becker, A. Challenges and perspectives in combinatorial assembly of novel exopolysaccharide biosynthesis pathways. Front. Microbiol. 2015, 6, 687. [Google Scholar] [CrossRef] [PubMed]
- Glucksmann, M.A.; Reuber, T.L.; Walker, G.C. Genes needed for the modification, polymerization, export, and processing of succinoglycan by Rhizobium meliloti: A model for succinoglycan biosynthesis. J. Bacteriol. 1993, 175, 7045–7055. [Google Scholar] [CrossRef] [PubMed]
- Varki, A.; Cummings, R.D.; Aebi, M.; Packer, N.H.; Seeberger, P.H.; Esko, J.D.; Stanley, P.; Hart, G.; Darvill, A.; Kinoshita, T.; et al. Symbol nomenclature for graphical representations of glycans. Glycobiology 2015, 25, 1323–1324. [Google Scholar] [CrossRef] [PubMed]
- Ivashina, T.V.; Fedorova, E.E.; Ashina, N.P.; Kalinchuk, N.A.; Druzhinina, T.N.; Shashkov, A.S.; Shibaev, V.N.; Ksenzenko, V.N. Mutation in the pssM gene encoding ketal pyruvate transferase leads to disruption of Rhizobium leguminosarum bv. viciae-Pisum sativum symbiosis. J. Appl. Microbiol. 2010, 109, 731–742. [Google Scholar] [PubMed]
- Xiu, A.; Kong, Y.; Zhou, M.; Zhu, B.; Wang, S.; Zhang, J. The chemical and digestive properties of a soluble glucan from Agrobacterium sp. ZX09. Carbohydr. Polym. 2010, 82, 623–628. [Google Scholar] [CrossRef]
- Xu, L.; Cheng, R.; Li, J.; Wang, Y.; Zhu, B.; Ma, S.; Zhang, W.; Dong, W.; Wang, S.; Zhang, J. Identification of substituent groups and related genes involved in salecan biosynthesis in Agrobacterium sp. ZX09. Appl. Microbiol. Biotechnol. 2017, 101, 585–598. [Google Scholar] [CrossRef] [PubMed]
- Meredith, T.C.; Mamat, U.; Kaczynski, Z.; Lindner, B.; Holst, O.; Woodard, R.W. Modification of lipopolysaccharide with colanic acid (M-antigen) repeats in Escherichia coli. J. Biol. Chem. 2007, 282, 7790–7798. [Google Scholar] [CrossRef] [PubMed]
- Scott, P.M.; Erickson, K.M.; Troutman, J.M. Identification of the functional roles of six key proteins in the biosynthesis of Enterobacteriaceae colanic acid. Biochemistry 2019, 58, 1818–1830. [Google Scholar] [CrossRef]
- Stevenson, G.; Andrianopoulos, K.; Hobbs, M.; Reeves, P.R. Organization of the Escherichia coli K-12 gene cluster responsible for production of the extracellular polysaccharide colanic acid. J. Bacteriol. 1996, 178, 4885–4893. [Google Scholar] [CrossRef]
- Zhang, J.; Poh, C.L. Regulating exopolysaccharide gene wcaF allows control of Escherichia coli biofilm formation. Sci. Rep. 2018, 8, 13127. [Google Scholar] [CrossRef]
- Cerantola, S.; Marty, N.; Montrozier, H. Structural studies of the acidic exopolysaccharide produced by a mucoid strain of Burkholderia cepacia, isolated from cystic fibrosis. Carbohydr. Res. 1996, 285, 59–67. [Google Scholar] [CrossRef]
- Cescutti, P.; Toffanin, R.; Pollesello, P.; Sutherland, I.W. Structural determination of the acidic exopolysaccharide produced by a Pseudomonas sp. strain 1.15. Carbohydr. Res. 1999, 315, 159–168. [Google Scholar] [CrossRef]
- Cescutti, P.; Kallioinen, A.; Impallomeni, G.; Toffanin, R.; Pollesello, P.; Leisola, M.; Eerikainen, T. Structure of the exopolysaccharide produced by Enterobacter amnigenus. Carbohydr. Res. 2005, 340, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Kuttel, M.; Ravenscroft, N.; Foschiatti, M.; Cescutti, P.; Rizzo, R. Conformational properties of two exopolysaccharides produced by Inquilinus limosus, a cystic fibrosis lung pathogen. Carbohydr. Res. 2012, 350, 40–48. [Google Scholar] [CrossRef]
- Herasimenka, Y.; Cescutti, P.; Impallomeni, G.; Rizzo, R. Exopolysaccharides produced by Inquilinus limosus, a new pathogen of cystic fibrosis patients: Novel structures with usual components. Carbohydr. Res. 2007, 342, 2404–2415. [Google Scholar] [CrossRef] [PubMed]
- Savant, A.P.; McColley, S.A. Cystic fibrosis year in review 2016. Pediatr. Pulmonol. 2017, 52, 1092–1102. [Google Scholar] [CrossRef]
- D’Haeze, W.; Glushka, J.; De Rycke, R.; Holsters, M.; Carlson, R.W. Structural characterization of extracellular polysaccharides of Azorhizobium caulinodans and importance for nodule initiation on Sesbania rostrata. Mol. Microbiol. 2004, 52, 485–500. [Google Scholar] [CrossRef]
- Lelchat, F.; Cerantola, S.; Brandily, C.; Colliec-Jouault, S.; Baudoux, A.C.; Ojima, T.; Boisset, C. The marine bacteria Cobetia marina DSMZ 4741 synthesizes an unexpected K-antigen-like exopolysaccharide. Carbohydr. Polym. 2015, 124, 347–356. [Google Scholar] [CrossRef]
- Yasutake, T.; Kumagai, T.; Inoue, A.; Kobayashi, K.; Noda, M.; Orikawa, A.; Matoba, Y.; Sugiyama, M. Characterization of the LP28 strain-specific exopolysaccharide biosynthetic gene cluster found in the whole circular genome of Pediococcus pentosaceus. Biochem. Biophys. Rep. 2016, 5, 266–271. [Google Scholar] [CrossRef]
- Van Zyl, L.M.; Coutinho, T.A.; Wingfield, M.J.; Pongpanich, K.; Wingfield, B.D. Morphological and molecular relatedness of geographically diverse isolates of Coniothyrium zuluense from South Africa and Thailand. Mycol. Res. 2002, 106, 51–59. [Google Scholar] [CrossRef]
- Yang, B.Y.; Ding, Q.; Montgomery, R. Extracellular polysaccharides of Erwinia futululu, a bacterium associated with a fungal canker disease of Eucalyptus spp. Carbohydr. Res. 2002, 337, 2469–2480. [Google Scholar] [CrossRef]
- O’Neill, M.A.; Robison, P.D.; Chou, K.J.; Darvill, A.G.; Albersheim, P. Evidence that the acidic polysaccharide secreted by Agrobacterium radiobacter (ATCC 53271) has a seventeen glycosyl-residue repeating unit. Carbohydr. Res. 1992, 226, 131–154. [Google Scholar] [CrossRef]
- Verhoef, R.; de Waard, P.; Schols, H.A.; Siika-aho, M.; Voragen, A.G. Methylobacterium sp. isolated from a Finnish paper machine produces highly pyruvated galactan exopolysaccharide. Carbohydr. Res. 2003, 338, 1851–1859. [Google Scholar] [CrossRef]
- Rougeaux, H.; Talaga, P.; Carlson, R.W.; Guezennec, J. Structural studies of an exopolysaccharide produced by Alteromonas macleodii subsp. fijiensis originating from a deep-sea hydrothermal vent. Carbohydr. Res. 1998, 312, 53–59. [Google Scholar] [CrossRef]
- Poli, A.; Anzelmo, G.; Nicolaus, B. Bacterial exopolysaccharides from extreme marine habitats: Production, characterization and biological activities. Mar. Drugs 2010, 8, 1779–1802. [Google Scholar] [CrossRef] [PubMed]
- Willis, L.M.; Whitfield, C. Structure, biosynthesis, and function of bacterial capsular polysaccharides synthesized by ABC transporter-dependent pathways. Carbohydr Res. 2013, 378, 35–44. [Google Scholar] [CrossRef] [PubMed]
- Kay, E.; Cuccui, J.; Wren, B.W. Recent advances in the production of recombinant glycoconjugate vaccines. npj Vaccines 2019, 4, 16. [Google Scholar] [CrossRef] [PubMed]
- Whitfield, C. Biosynthesis and assembly of capsular polysaccharides in Escherichia coli. Annu. Rev. Biochem. 2006, 75, 39–68. [Google Scholar] [CrossRef] [PubMed]
- Hausdorff, W.P.; Hoet, B.; Adegbola, R.A. Predicting the impact of new pneumococcal conjugate vaccines: Serotype composition is not enough. Expert Rev. Vaccines 2015, 14, 413–428. [Google Scholar] [CrossRef] [PubMed]
- Harding, C.M.; Nasr, M.A.; Scott, N.E.; Goyette-Desjardins, G.; Nothaft, H.; Mayer, A.E.; Chavez, S.M.; Huynh, J.P.; Kinsella, R.L.; Szymanski, C.M.; et al. A platform for glycoengineering a polyvalent pneumococcal bioconjugate vaccine using E. coli as a host. Nat. Commun. 2019, 10, 891. [Google Scholar] [CrossRef] [PubMed]
- Kaplonek, P.; Khan, N.; Reppe, K.; Schumann, B.; Emmadi, M.; Lisboa, M.P.; Xu, F.F.; Calow, A.D.J.; Parameswarappa, S.G.; Witzenrath, M.; et al. Improving vaccines against Streptococcus pneumoniae using synthetic glycans. Proc. Natl. Acad. Sci. USA 2018, 115, 13353–13358. [Google Scholar] [CrossRef] [PubMed]
- Pereira, C.L.; Geissner, A.; Anish, C.; Seeberger, P.H. Chemical synthesis elucidates the immunological importance of a pyruvate modification in the capsular polysaccharide of Streptococcus pneumoniae serotype 4. Angew Chem. Int. Ed. Engl. 2015, 54, 10016–10019. [Google Scholar] [CrossRef] [PubMed]
- Falagas, M.E.; Karveli, E.A.; Siempos, I.I.; Vardakas, K.Z. Acinetobacter infections: A growing threat for critically ill patients. Epidemiol. Infect. 2008, 136, 1009–1019. [Google Scholar] [CrossRef] [PubMed]
- Antunes, L.C.; Visca, P.; Towner, K.J. Acinetobacter baumannii: Evolution of a global pathogen. Pathog. Dis. 2014, 71, 292–301. [Google Scholar] [CrossRef] [PubMed]
- Kenyon, J.J.; Hall, R.M. Variation in the complex carbohydrate biosynthesis loci of Acinetobacter baumannii genomes. PLoS ONE 2013, 8, e62160. [Google Scholar] [CrossRef] [PubMed]
- Kenyon, J.J.; Speciale, I.; Hall, R.M.; De Castro, C. Structure of repeating unit of the capsular polysaccharide from Acinetobacter baumannii D78 and assignment of the K4 gene cluster. Carbohydr. Res. 2016, 434, 12–17. [Google Scholar] [CrossRef]
- Yang, C.C.; Yen, C.H.; Ho, M.W.; Wang, J.H. Comparison of pyogenic liver abscess caused by non-Klebsiella pneumoniae and Klebsiella pneumoniae. J. Microbiol. Immunol. Infect. 2004, 37, 176–184. [Google Scholar]
- Yang, F.L.; Yang, Y.L.; Liao, P.C.; Chou, J.C.; Tsai, K.C.; Yang, A.S.; Sheu, F.; Lin, T.L.; Hsieh, P.F.; Wang, J.T.; et al. Structure and immunological characterization of the capsular polysaccharide of a pyrogenic liver abscess caused by Klebsiella pneumoniae: Activation of macrophages through Toll-like receptor 4. J. Biol. Chem. 2011, 286, 21041–22151. [Google Scholar] [CrossRef]
- Rao, A.S.; Liao, J.; Kabat, E.A.; Osserman, E.F.; Harboe, M.; Nimmich, W. Immunochemical studies on human monoclonal macroglobulins with specificities for 3,4-pyruvylated d-galactose and 4,6-pyruvylated d-glucose. J. Biol. Chem. 1984, 259, 1018–1026. [Google Scholar]
- Dutton, G.G.; Parolis, H.; Parolis, L.A. A structural investigation of the capsular polysaccharide of Klebsiella K14. Carbohydr. Res. 1985, 140, 263–275. [Google Scholar] [CrossRef]
- Beurret, M.; Joseleau, J.P.; Vignon, M.; Dutton, G.G.; Savage, A.V. Proof of the occurrence of 5,6-O-(1-carboxyethylidene)-d-galactofuranose units in the capsular polysaccharide of Klebsiella K12. Carbohydr. Res. 1989, 189, 247–260. [Google Scholar] [CrossRef]
- Dutton, G.G.; Mackie, K.L. Structural investigation of Klebsiella serotype K70 polysaccharide. Carbohydr. Res. 1978, 62, 321–335. [Google Scholar] [CrossRef]
- Merrifield, E.H.; Stephen, A.M. Structural studies on the capsular polysaccharide from Klebsiella serotype K64. Carbohydr. Res. 1979, 74, 241–257. [Google Scholar] [CrossRef]
- Okutani, K.; Dutton, G.G. Structural investigation of Klebsiella serotype K46 polysaccharide. Carbohydr. Res. 1980, 86, 259–271. [Google Scholar] [CrossRef]
- Rao, A.S.; Kabat, E.A.; Nilsson, B.; Zopf, D.A.; Nimmich, W. Isolation and characterization of 4-O-[3,4-O-(1-carboxyethylidene)-β-d-galactopyranosyl]erythritol from Klebsiella K33 polysaccharide. Carbohydr. Res. 1983, 121, 205–209. [Google Scholar] [CrossRef]
- Kabat, E.A.; Liao, J.; Bretting, H.; Franklin, E.C.; Geltner, D.; Frangione, B.; Koshland, M.E.; Shyong, J.; Osserman, E.F. Human monoclonal macroglobulins with specificity for Klebsiella K polysaccharides that contain 3,4-pyruvylated-d-galactose and 4,6-pyruvylated-d-galactose. J. Exp. Med. 1980, 152, 979–995. [Google Scholar] [CrossRef] [PubMed]
- Dutton, G.G.; Parolis, H.; Joseleau, J.P.; Marais, M.F. The use of bacteriophage depolymerization in the structural investigation of the capsular polysaccharide from Klebsiella serotype K3. Carbohydr. Res. 1986, 149, 411–423. [Google Scholar] [CrossRef]
- Zamze, S.; Martinez-Pomares, L.; Jones, H.; Taylor, P.R.; Stillion, R.J.; Gordon, S.; Wong, S.Y. Recognition of bacterial capsular polysaccharides and lipopolysaccharides by the macrophage mannose receptor. J. Biol. Chem. 2002, 277, 41613–41623. [Google Scholar] [CrossRef] [PubMed]
- De Silva, R.A.; Wang, Q.; Chidley, T.; Appulage, D.K.; Andreana, P.R. Immunological response from an entirely carbohydrate antigen: Design of synthetic vaccines based on Tn-PS A1 conjugates. J. Am. Chem. Soc. 2009, 131, 9622–9623. [Google Scholar] [CrossRef]
- Trabbic, K.R.; De Silva, R.A.; Andreana, P.R. Elucidating structural features of an entirely carbohydrate cancer vaccine construct employing circular dichroism and fluorescent labeling. Medchemcomm 2014, 5, 1143–1149. [Google Scholar] [CrossRef]
- Coyne, M.J.; Tzianabos, A.O.; Mallory, B.C.; Carey, V.J.; Kasper, D.L.; Comstock, L.E. Polysaccharide biosynthesis locus required for virulence of Bacteroides fragilis. Infect. Immun. 2001, 69, 4342–4350. [Google Scholar] [CrossRef] [PubMed]
- Baumann, H.; Tzianabos, A.O.; Brisson, J.R.; Kasper, D.L.; Jennings, H.J. Structural elucidation of two capsular polysaccharides from one strain of Bacteroides fragilis using high-resolution NMR spectroscopy. Biochemistry 1992, 31, 4081–40899. [Google Scholar] [CrossRef] [PubMed]
- Severn, W.B.; Richards, J.C. The structure of the specific capsular polysaccharide of Rhodococcus equi serotype 4. Carbohydr. Res. 1999, 320, 209–222. [Google Scholar] [CrossRef]
- Percy, M.G.; Gründling, A. Lipoteichoic acid synthesis and function in gram-positive bacteria. Annu. Rev. Microbiol. 2014, 68, 81–100. [Google Scholar] [CrossRef] [PubMed]
- Brown, S.; Santa Maria, J.P., Jr.; Walker, S. Wall teichoic acids of Gram-positive bacteria. Annu. Rev. Microbiol. 2013, 67, 313–336. [Google Scholar] [CrossRef]
- Pasquina, L.W.; Santa Maria, J.P.; Walker, S. Teichoic acid biosynthesis as an antibiotic target. Curr. Opin. Microbiol. 2013, 16, 531–537. [Google Scholar] [CrossRef]
- Mesnage, S.; Fontaine, T.; Mignot, T.; Delepierre, M.; Mock, M.; Fouet, A. Bacterial SLH domain proteins are non-covalently anchored to the cell surface via a conserved mechanism involving wall polysaccharide pyruvylation. EMBO J. 2000, 19, 4473–4484. [Google Scholar] [CrossRef]
- Schäffer, C.; Messner, P. The structure of secondary cell wall polymers: How Gram-positive bacteria stick their cell walls together. Microbiology 2005, 151, 643–651. [Google Scholar] [CrossRef]
- Fouet, A. The surface of Bacillus anthracis. Mol. Asp. Med. 2009, 30, 374–385. [Google Scholar] [CrossRef]
- Choudhury, B.; Leoff, C.; Saile, E.; Wilkins, P.; Quinn, C.P.; Kannenberg, E.L.; Carlson, R.W. The structure of the major cell wall polysaccharide of Bacillus anthracis is species-specific. J. Biol. Chem. 2006, 281, 27932–27941. [Google Scholar] [CrossRef]
- Sychantha, D.; Chapman, R.N.; Bamford, N.C.; Boons, G.J.; Howell, P.L.; Clarke, A.J. Molecular basis for the attachment of S-layer proteins to the cell wall of Bacillus anthracis. Biochemistry 2018, 57, 1949–1953. [Google Scholar] [CrossRef] [PubMed]
- Blackler, R.J.; López-Guzmán, A.; Hager, F.F.; Janesch, B.; Martinz, G.; Gagnon, S.M.L.; Haji-Ghassemi, O.; Kosma, P.; Messner, P.; Schäffer, C.; et al. Structural basis of cell wall anchoring by SLH domains in Paenibacillus alvei. Nat. Commun. 2018, 9, 3120. [Google Scholar] [CrossRef] [PubMed]
- Messner, P.; Schäffer, C.; Egelseer, E.M.; Sleytr, U.B. Occurrence, structure, chemistry, genetics, morphogenesis and functions of S-layers. In Prokaryotic Cell Wall Compounds - Structure and Biochemistry; König, H., Claus, H., Varma, A., Eds.; Springer: Berlin, Germany, 2010; pp. 53–109. [Google Scholar]
- Chung, S.; Shin, S.H.; Bertozzi, C.R.; De Yoreo, J.J. Self-catalyzed growth of S layers via an amorphous-to-crystalline transition limited by folding kinetics. Proc. Natl. Acad. Sci. USA 2010, 107, 16536–16541. [Google Scholar] [CrossRef]
- Messner, P.; Allmaier, G.; Schäffer, C.; Wugeditsch, T.; Lortal, S.; König, H.; Niemetz, R.; Dorner, M. Biochemistry of S-layers. FEMS Microbiol. Rev. 1997, 20, 25–46. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Schuster, B.; Egelseer, E.M.; Pum, D. S-layers: Principles and applications. FEMS Microbiol. Rev. 2014, 38, 823–864. [Google Scholar] [CrossRef] [PubMed]
- Hager, F.F.; Lopez-Guzman, A.; Krauter, S.; Blaukopf, M.; Polter, M.; Brockhausen, I.; Kosma, P.; Schäffer, C. Functional characterization of enzymatic steps involved in pyruvylation of bacterial secondary cell wall polymer fragments. Front. Microbiol. 2018, 9, 1356. [Google Scholar] [CrossRef]
- Forsberg, L.S.; Choudhury, B.; Leoff, C.; Marston, C.K.; Hoffmaster, A.R.; Saile, E.; Quinn, C.P.; Kannenberg, E.L.; Carlson, R.W. Secondary cell wall polysaccharides from Bacillus cereus strains G9241, 03BB87 and 03BB102 causing fatal pneumonia share similar glycosyl structures with the polysaccharides from Bacillus anthracis. Glycobiology 2011, 21, 934–948. [Google Scholar] [CrossRef]
- Mesnage, S.; Tosi-Couture, E.; Mock, M.; Fouet, A. The S-layer homology domain as a means for anchoring heterologous proteins on the cell surface of Bacillus anthracis. J. Appl. Microbiol. 1999, 87, 256–260. [Google Scholar] [CrossRef]
- Kern, J.; Ryan, C.; Faull, K.; Schneewind, O. Bacillus anthracis surface-layer proteins assemble by binding to the secondary cell wall polysaccharide in a manner that requires csaB and tagO. J. Mol. Biol. 2010, 401, 757–775. [Google Scholar] [CrossRef]
- Missiakas, D.; Schneewind, O. Assembly and function of the Bacillus anthracis S-layer. Annu. Rev. Microbiol. 2017, 71, 79–98. [Google Scholar] [CrossRef]
- Oh, S.Y.; Lunderberg, J.M.; Chateau, A.; Schneewind, O.; Missiakas, D. Genes required for Bacillus anthracis secondary cell wall polysaccharide synthesis. J. Bacteriol. 2017, 199, e00613–e00616. [Google Scholar] [CrossRef]
- Chateau, A.; Lunderberg, J.M.; Oh, S.Y.; Abshire, T.; Friedlander, A.; Quinn, C.P.; Missiakas, D.M.; Schneewind, O. Galactosylation of the secondary cell wall polysaccharide of Bacillus anthracis and Its contribution to anthrax pathogenesis. J. Bacteriol. 2018, 200, e00562-17. [Google Scholar] [CrossRef] [PubMed]
- Lunderberg, J.M.; Liszewski Zilla, M.; Missiakas, D.; Schneewind, O. Bacillus anthracis tagO is required for vegetative growth and secondary cell wall polysaccharide synthesis. J. Bacteriol. 2015, 197, 3511–3520. [Google Scholar] [CrossRef] [PubMed]
- Sychantha, D.; Little, D.J.; Chapman, R.N.; Boons, G.J.; Robinson, H.; Howell, P.L.; Clarke, A.J. PatB1 is an O-acetyltransferase that decorates secondary cell wall polysaccharides. Nat. Chem. Biol. 2018, 14, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Schneewind, O.; Missiakas, D.M. Protein secretion and surface display in Gram-positive bacteria. Philos Trans. R. Soc. Lond. B Biol. Sci. 2012, 367, 1123–1139. [Google Scholar] [CrossRef] [PubMed]
- Garufi, G.; Hendrickx, A.P.; Beeri, K.; Kern, J.W.; Sharma, A.; Richter, S.G.; Schneewind, O.; Missiakas, D. Synthesis of lipoteichoic acids in Bacillus anthracis. J. Bacteriol. 2012, 194, 4312–4321. [Google Scholar] [CrossRef] [PubMed]
- Janesch, B.; Messner, P.; Schäffer, C. Are the surface layer homology domains essential for cell surface display and glycosylation of the S-layer protein from Paenibacillus alvei CCM 2051T? J. Bacteriol. 2013, 195, 565–575. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ilk, N.; Kosma, P.; Puchberger, M.; Egelseer, E.M.; Mayer, H.F.; Sleytr, U.B.; Sára, M. Structural and functional analyses of the secondary cell wall polymer of Bacillus sphaericus CCM 2177 that serves as an S-layer-specific anchor. J. Bacteriol. 1999, 181, 7643–7646. [Google Scholar] [PubMed]
- Zarschler, K.; Janesch, B.; Kainz, B.; Ristl, R.; Messner, P.; Schäffer, C. Cell surface display of chimeric glycoproteins via the S-layer of Paenibacillus alvei. Carbohydr. Res. 2010, 345, 1422–1431. [Google Scholar] [CrossRef] [PubMed][Green Version]
- D’Elia, M.A.; Pereira, M.P.; Chung, Y.S.; Zhao, W.; Chau, A.; Kenney, T.J.; Sulavik, M.C.; Black, T.A.; Brown, E.D. Lesions in teichoic acid biosynthesis in Staphylococcus aureus lead to a lethal gain of function in the otherwise dispensable pathway. J. Bacteriol. 2006, 188, 4183–4189. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.H.; Ginsberg, C.; Yuan, Y.; Walker, S. Acceptor substrate selectivity and kinetic mechanism of Bacillus subtilis TagA. Biochemistry 2006, 45, 10895–10904. [Google Scholar] [CrossRef] [PubMed][Green Version]
- D’Elia, M.A.; Henderson, J.A.; Beveridge, T.J.; Heinrichs, D.E.; Brown, E.D. The N-acetylmannosamine transferase catalyzes the first committed step of teichoic acid assembly in Bacillus subtilis and Staphylococcus aureus. J. Bacteriol. 2009, 191, 4030–4034. [Google Scholar] [CrossRef] [PubMed]
- Ginsberg, C.; Zhang, Y.H.; Yuan, Y.; Walker, S. In vitro reconstitution of two essential steps in wall teichoic acid biosynthesis. ACS Chem. Biol. 2006, 1, 25–28. [Google Scholar] [CrossRef] [PubMed]
- Swoboda, J.G.; Campbell, J.; Meredith, T.C.; Walker, S. Wall teichoic acid function, biosynthesis, and inhibition. Chembiochem 2010, 11, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Schirner, K.; Marles-Wright, J.; Lewis, R.J.; Errington, J. Distinct and essential morphogenic functions for wall- and lipo-teichoic acids in Bacillus subtilis. EMBO J. 2009, 28, 830–842. [Google Scholar] [CrossRef] [PubMed]
- Chapot-Chartier, M.P.; Kulakauskas, S. Cell wall structure and function in lactic acid bacteria. Microb Cell Fact. 2014, 13 (Suppl. 1), S9. [Google Scholar] [CrossRef]
- Kojima, N.; Kaya, S.; Araki, Y.; Ito, E. Pyruvic-acid-containing polysaccharide in the cell wall of Bacillus polymyxa AHU 1385. Eur. J. Biochem. 1988, 174, 255–260. [Google Scholar] [CrossRef]
- Dmitrenok, A.S.; Streshinskaya, G.M.; Tul’skaya, E.M.; Potekhina, N.V.; Senchenkova, S.N.; Shashkov, A.S.; Bilan, M.I.; Starodumova, I.P.; Bueva, O.V.; Evtushenko, L.I. Pyruvylated cell wall glycopolymers of Promicromonospora citrea VKM A succeeds, similar-665(T) and Promicromonospora sp. VKM A succeeds, similar-1028. Carbohydr. Res. 2017, 449, 134–142. [Google Scholar]
- Schleifer, K.H.; Kandler, O. Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol. Rev. 1972, 36, 407–477. [Google Scholar]
- Tul’skaya, E.M.; Shashkov, A.S.; Streshinskaya, G.M.; Potekhina, N.V.; Evtushenko, L.I. New structures and composition of cell wall teichoic acids from Nocardiopsis synnemataformans, Nocardiopsis halotolerans, Nocardiopsis composta and Nocardiopsis metallicus: A chemotaxonomic value. Antonie Van Leeuwenhoek 2014, 106, 1105–1117. [Google Scholar] [CrossRef]
- Ibrahim, A.H.; Desoukey, S.Y.; Fouad, M.A.; Kamel, M.S.; Gulder, T.A.M.; Abdelmohsen, U.R. Natural product potential of the genus Nocardiopsis. Mar. Drugs 2018, 16, 147. [Google Scholar] [CrossRef] [PubMed]
- Potekhina, N.V.; Evtushenko, L.I.; Senchenkova, S.N.; Shashkov, A.S.; Naumova, I.B. Structures of cell wall teichoic acids of Brevibacterium iodinum VKM Ac-2106. Biochem. (Mosc.) 2004, 69, 1353–1359. [Google Scholar] [CrossRef] [PubMed]
- Whitfield, C.; Trent, M.S. Biosynthesis and export of bacterial lipopolysaccharides. Annu. Rev. Biochem. 2014, 83, 99–128. [Google Scholar] [CrossRef] [PubMed]
- Kanipes, M.I.; Guerry, P. Chapter 44—Role of microbial glycosylation in host cell invasion. In Microbial Glycobiology; Holst, O., Brennan, P.J., Itzstein, M.V., Moran, A.P., Eds.; Academic Press: San Diego, CA, USA, 2010; pp. 871–883. [Google Scholar]
- Raetz, C.R.H.; Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 2002, 71, 635–700. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Meredith, T.C.; Kahne, D. On the essentiality of lipopolysaccharide to Gram-negative bacteria. Curr. Opin. Microbiol. 2013, 16, 779–785. [Google Scholar] [CrossRef] [PubMed]
- Broeker, N.K.; Barbirz, S. Not a barrier but a key: How bacteriophages exploit host’s O-antigen as an essential receptor to initiate infection. Mol. Microbiol. 2017, 105, 353–357. [Google Scholar] [CrossRef]
- Kocharova, N.A.; Vinogradov, E.; Kondakova, A.N.; Shashkov, A.S.; Rozalski, A.; Knirel, Y.A. The full structure of the carbohydrate chain of the lipopolysaccharide of Providencia alcalifaciens O19. J. Carbohydr. Chem. 2008, 27, 320–331. [Google Scholar] [CrossRef]
- Kondakova, A.N.; Vinogradov, E.; Lindner, B.; Kocharova, N.A.; Rozalski, A.; Knirel, Y.A. Elucidation of the lipopolysaccharide core Structures of bacteria of the genus Providencia. J. Carbohydr. Chem. 2006, 25, 499–520. [Google Scholar] [CrossRef]
- Kocharova, N.A.; Kondakova, A.N.; Vinogradov, E.; Ovchinnikova, O.G.; Lindner, B.; Shashkov, A.S.; Rozalski, A.; Knirel, Y.A. Full structure of the carbohydrate chain of the lipopolysaccharide of Providencia rustigianii O34. Chemistry 2008, 14, 6184–6191. [Google Scholar] [CrossRef]
- Perepelov, A.V.; Senchenkova, S.N.; Shashkov, A.S.; Lu, B.; Feng, L.; Vang, L.; Knirel, Y.A. Antigenic polysaccharides of bacteria: 40. The structures of O-specific polysaccharides from Shigella dysenteriae types 3 and 9 and S. boydii type 4 revised by NMR spectroscopy. Russ. J. Bioorgan. Chem. 2008, 34, 121–128. [Google Scholar] [CrossRef]
- Perepelov, A.V.; Senchenkova, S.N.; Shashkov, A.S.; Knirel, Y.A.; Lu, B.; Feng, L.; Wang, L. A completed structure of the O-polysaccharide from Shigella dysenteriae type 10. Rus. J. Bioorgan. Chem. 2009, 35, 131–133. [Google Scholar] [CrossRef]
- Leone, S.; Molinaro, A.; Dubery, I.; Lanzetta, R.; Parrilli, M. The O-specific polysaccharide structure from the lipopolysaccharide of the Gram-negative bacterium Raoultella terrigena. Carbohydr. Res. 2007, 342, 1514–1518. [Google Scholar] [CrossRef] [PubMed]
- Perepelov, A.V.; Rozalski, A.; Bartodziejska, B.; Senchenkova, S.N.; Knirel, Y.A. Structure of the O-polysaccharide of Proteus mirabilis O19 and reclassification of certain Proteus strains that were formerly classified in serogroup O19. Arch. Immunol. Ther. Exp. (Warsz.) 2004, 52, 1881–1896. [Google Scholar]
- Kokoulin, M.S.; Kalinovsky, A.I.; Komandrova, N.A.; Tomshich, S.V.; Romanenko, L.A.; Vaskovsky, V.E. The new sulfated O-specific polysaccharide from marine bacterium Cobetia pacifica KMM 3878, containing 3,4-O-[(S)-1-carboxyethylidene]-d-galactose and 2,3-O-disulfate-d-galactose. Carbohydr. Res. 2014, 397, 46–51. [Google Scholar] [CrossRef] [PubMed]
- Barksdale, L.; Kim, K.S. Mycobacterium. Bacteriol. Rev. 1977, 41, 217–372. [Google Scholar] [PubMed]
- Saadat, S.; Ballou, C.E. Pyruvylated glycolipids from Mycobacterium smegmatis. Structures of two oligosaccharide components. J. Biol. Chem. 1983, 258, 1813–1818. [Google Scholar] [PubMed]
- Thayer, W.R., Jr.; Bozic, C.M.; Camphausen, R.T.; McNeil, M. Implications of antibodies to pyruvylated glucose in healthy populations for mycobacterioses and other infectious diseases. J. Clin. Microbiol. 1990, 28, 714–718. [Google Scholar] [PubMed]
- Araki, S.; Abe, S.; Ando, S.; Kon, K.; Fujiwara, N.; Satake, M. Structure of phosphonoglycosphingolipid containing pyruvylated galactose in nerve fibers of Aplysia kurodai. J. Biol. Chem. 1989, 264, 19922–19927. [Google Scholar] [PubMed]
- Gemmill, T.R.; Trimble, R.B. Overview of N- and O-linked oligosaccharide structures found in various yeast species. Biochim. Biophys. Acta 1999, 1426, 227–237. [Google Scholar] [CrossRef]
- Cutler, J.E. N-glycosylation of yeast, with emphasis on Candida albicans. Med. Mycol. 2001, 39 (Suppl. 1), 75–86. [Google Scholar] [CrossRef]
- Parolis, L.A.; Duus, J.O.; Parolis, H.; Meldal, M.; Bock, K. The extracellular polysaccharide of Pichia (Hansenula) holstii NRRL Y-2448: The structure of the phosphomannan backbone. Carbohydr. Res. 1996, 293, 101–117. [Google Scholar] [CrossRef]
- Gemmill, T.R.; Trimble, R.B. All pyruvylated galactose in Schizosaccharomyces pombe N-glycans is present in the terminal disaccharide, 4, 6-O-[(R)-(1-carboxyethylidine)]-Galβ1,3Galα1. Glycobiology 1998, 8, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- Schüle, G.; Ziegler, T. Efficient block synthesis of a pyruvated tetrasaccharide 5-amino-pentyl glycoside related to Streptococcus pneumoniae type 27. Tetrahedron 1996, 52, 2925–2936. [Google Scholar] [CrossRef]
- Ziegler, F.D.; Cavanagh, J.; Lubowski, C.; Trimble, R.B. Novel Schizosaccharomyces pombe N-linked GalMan9GlcNAc isomers: Role of the Golgi GMA12 galactosyltransferase in core glycan galactosylation. Glycobiology 1999, 9, 497–505. [Google Scholar] [CrossRef] [PubMed]
- Ballou, C.E.; Ballou, L.; Ball, G. Schizosaccharomyces pombe glycosylation mutant with altered cell surface properties. Proc. Natl. Acad. Sci. USA 1994, 91, 9327–9331. [Google Scholar] [CrossRef]
- Yoritsune, K.; Matsuzawa, T.; Ohashi, T.; Takegawa, K. The fission yeast Pvg1p has galactose-specific pyruvyltransferase activity. FEBS Lett. 2013, 587, 917–921. [Google Scholar] [CrossRef]
- Ferreira, L.G.; Noseda, M.D.; Goncalves, A.G.; Ducatti, D.R.; Fujii, M.T.; Duarte, M.E. Chemical structure of the complex pyruvylated and sulfated agaran from the red seaweed Palisada flagellifera (Ceramiales, Rhodophyta). Carbohydr. Res. 2012, 347, 83–94. [Google Scholar] [CrossRef]
- Bondu, S.; Deslandes, E.; Fbre, M.S.; Berthou, C.; Guangli, A. Carrageenan from Solieria chordalis (Gigartinales): Structural analysis and immunological activities of the low molecular weight fractions. Carbohydr. Polym. 2010, 81, 448–460. [Google Scholar]
- Spillmann, D.; Thomas-Oates, J.E.; van Kuik, J.A.; Vliegenthart, J.F.; Misevic, G.; Burger, M.M.; Finne, J. Characterization of a novel sulfated carbohydrate unit implicated in the carbohydrate-carbohydrate-mediated cell aggregation of the marine sponge Microciona prolifera. J. Biol. Chem. 1995, 270, 5089–5097. [Google Scholar] [CrossRef]
- Nielsen, P.H.; Jahn, A. Extraction of EPS. In Microbial extracellular polymeric substances: Characterization, structure and function; Wingender, J., Neu, T.R., Flemming, H.-C., Eds.; Springer: Berlin/Heidelberg, Germany, 1999; pp. 49–72. [Google Scholar]
- Yang, B.Y.; Brand, J.; Montgomery, R. Pyruvated galactose and oligosaccharides from Erwinia chrysanthemi Ech6 extracellular polysaccharide. Carbohydr. Res. 2001, 331, 59–67. [Google Scholar] [CrossRef]
- Domenico, P.; Diedrich, D.L.; Cunha, B.A. Quantitative extraction and purification of exopolysaccharides from Klebsiella pneumoniae. J. Microbiol. Methods 1989, 9, 211–219. [Google Scholar] [CrossRef]
- Brimacombe, C.A.; Beatty, J.T. Surface polysaccharide extraction and quantification. Bio-Protocol 2013, 3, e934. [Google Scholar] [CrossRef]
- Kho, K.; Meredith, T.C. Extraction and analysis of bacterial teichoic acids. Bio-Protocol 2018, 8, e3078. [Google Scholar] [CrossRef]
- Chateau, A.; Schneewind, O.; Missiakas, D. Extraction and purification of wall-bound polymers of Gram-positive bacteria. Methods Mol. Biol. 2019, 1954, 47–57. [Google Scholar] [PubMed]
- Whitfield, C.; Perry, M.B. 6.7 Isolation and purification of cell surface polysaccharides from Gram-negative bacteria. In Methods in Microbiology; Williams, P., Ketley, J., Salmond, G., Eds.; Academic Press: Cambridge, MA, USA, 1998; Volume 27, pp. 249–258. [Google Scholar]
- Grice, I.D.; Wilson, J.C. Chapter 13 - Analytical approaches towards the structural characterization of microbial wall glycopolymers. In Microbial Glycobiology; Holst, O., Brennan, P.J., Itzstein, M.V., Moran, A.P., Eds.; Academic Press: San Diego, CA, USA, 2010; pp. 233–252. [Google Scholar]
- Perepelov, A.V.; Weintraub, A.; Liu, B.; Senchenkova, S.N.; Shashkov, A.S.; Feng, L.; Wang, L.; Widmalm, G.; Knirel, Y.A. The O-polysaccharide of Escherichia coli O112ac has the same structure as that of Shigella dysenteriae type 2 but is devoid of O-acetylation: A revision of the S. dysenteriae type 2 O-polysaccharide structure. Carbohydr. Resc. 2008, 343, 977–981. [Google Scholar] [CrossRef] [PubMed]
- Westphal, O.; Jann, K. Bacterial lipopolysaccharides extraction with phenol-water and further applications of the procedure. Methods Carbohydr. Chem. 1965, 5, 83–91. [Google Scholar]
- Galanos, C.; Lüderitz, O.; Westphal, O. A new method for the extraction of R lipopolysaccharides. Eur. J. Biochem. 1969, 9, 245–249. [Google Scholar] [CrossRef] [PubMed]
- Darveau, R.P.; Hancock, R.E. Procedure for isolation of bacterial lipopolysaccharides from both smooth and rough Pseudomonas aeruginosa and Salmonella typhimurium strains. J. Bacteriol. 1983, 155, 831–838. [Google Scholar]
- Holst, O.; Broer, W.; Thomas-Oates, J.E.; Mamat, U.; Brade, H. Structural analysis of two oligosaccharide bisphosphates isolated from the lipopolysaccharide of a recombinant strain of Escherichia coli F515 (Re chemotype) expressing the genus-specific epitope of Chlamydia lipopolysaccharide. Eur. J. Biochem. 1993, 214, 703–710. [Google Scholar] [CrossRef]
- Hind, C.R.; Collins, P.M.; Renn, D.; Cook, R.B.; Caspi, D.; Baltz, M.L.; Pepys, M.B. Binding specificity of serum amyloid P component for the pyruvate acetal of galactose. J. Exp. Med. 1984, 159, 1058–1069. [Google Scholar] [CrossRef]
- Hind, C.R.; Collins, P.M.; Baltz, M.L.; Pepys, M.B. Human serum amyloid P component, a circulating lectin with specificity for the cyclic 4,6-pyruvate acetal of galactose. Interactions with various bacteria. Biochem. J. 1985, 225, 107–111. [Google Scholar] [CrossRef] [PubMed]
- Rühmann, B.; Schmid, J.; Sieber, V. Automated modular high throughput exopolysaccharide screening platform coupled with highly sensitive carbohydrate fingerprint analysis. J. Vis. Exp. 2016. [Google Scholar] [CrossRef] [PubMed]
- Bjerring, P.N.; Hauerberg, J.; Frederiksen, H.J.; Jorgensen, L.; Hansen, B.A.; Tofteng, F.; Larsen, F.S. Cerebral glutamine concentration and lactate-pyruvate ratio in patients with acute liver failure. Neurocrit. Care 2008, 9, 3–7. [Google Scholar] [CrossRef] [PubMed]
- Schwimmer, S.; Weston, W.J. Onion flavor and odor - Enzymatic development of pyruvic acid in onion as a measure of pungency. J. Agric. Food Chem. 1961, 9, 127–132. [Google Scholar] [CrossRef]
- Anthon, G.E.; Barrett, D.M. Modified method for the determination of pyruvic acid with dinitrophenylhydrazine in the assessment of onion pungency. J. Sci. Food Agric. 2003, 83, 1210–1213. [Google Scholar] [CrossRef]
- Ovodov, Y.S. Bacterial capsular antigens. Structural patterns of capsular antigens. Biochem. (Mosc.) 2006, 71, 937–954. [Google Scholar] [CrossRef] [PubMed]
- Garegg, P.J.; Jansson, P.E.; Lindberg, B.; Lindh, F.; Lönngren, J.; Kvarnström, I.; Nimmich, W. Configuration of the acetal carbon-atom of pyruvic-acid acetals in some bacterial polysaccharides. Carbohydr. Res. 1980, 78, 127–132. [Google Scholar] [CrossRef]
- Jansson, P.E.; Lindberg, J.; Widmalm, G. Syntheses and NMR-studies of pyruvic acid 4,6-acetals of some methyl hexopyranosides. Acta Chem. Scand. 1993, 47, 711–715. [Google Scholar] [CrossRef]
- Gorin, P.A.J.; Mazurek, M.; Duarte, H.S.; Iacomini, M.; Duarte, J.H. Properties of 13C-NMR spectra of O-(1-carboxyethylidene) derivatives of methyl-β-d-galactopyranoside - Models for determination of pyruvic-acid acetal structures in polysaccharides. Carbohydr. Res. 1982, 100, 1–15. [Google Scholar] [CrossRef]
- Lindberg, B. Components of bacterial polysaccharides. In Advances in Carbohydrate Chemistry and Biochemistry; Tipson, R.S., Horton, D., Eds.; Academic Press: Washington, DC, USA, 1990; Volume 48, pp. 279–318. [Google Scholar]
- Zhao, G.; Perepelov, A.V.; Senchenkova, S.N.; Shashkov, A.S.; Feng, L.; Li, X.M.; Knirel, Y.A.; Wang, L. Structural relation of the antigenic polysaccharides of Escherichia coli O40, Shigella dysenteriae type 9, and E. coli K47. Carbohydr. Res. 2007, 342, 1275–1279. [Google Scholar] [CrossRef]
- Ravenscroft, N.; Parolis, L.A.S.; Parolis, H. Bacteriophage degradation of Klebsiella K30 capsular polysaccharide - an NMR investigation of the 3,4-pyruvated galactose-containing repeating oligosaccharide. Carbohydr. Res. 1994, 254, 333–340. [Google Scholar] [CrossRef]
- Dudman, W.F.; Lacey, M.J. Identification of pyruvated monosaccharides in polysaccharides by gas-liquid chromatography mass spectrometry. Carbohydr. Res. 1986, 145, 175–191. [Google Scholar] [CrossRef]
- Fontaine, T.; Talmont, F.; Dutton, G.G.; Fournet, B. Analysis of pyruvic acid acetal containing polysaccharides by methanolysis and reductive cleavage methods. Anal. Biochem. 1991, 199, 154–161. [Google Scholar] [CrossRef]
- Kailemia, M.J.; Ruhaak, L.R.; Lebrilla, C.B.; Amster, I.J. Oligosaccharide analysis by mass spectrometry: A review of recent developments. Anal. Chem. 2014, 86, 196–212. [Google Scholar] [CrossRef] [PubMed]
- Harvey, D.J. Matrix-assisted laser desorption/ionization mass spectrometry of carbohydrates. Mass Spec. Rev. 1999, 18, 349–451. [Google Scholar] [CrossRef]
- Kern, J.; Wilton, R.; Zhang, R.; Binkowski, T.A.; Joachimiak, A.; Schneewind, O. Structure of surface layer homology (SLH) domains from Bacillus anthracis surface array protein. J. Biol. Chem. 2011, 286, 26042–26049. [Google Scholar] [CrossRef] [PubMed]
- Chapman, R.N.; Liu, L.; Boons, G.J. 4,6-O-pyruvyl ketal modified N-acetyl mannosamine of the secondary cell polysaccharide of Bacillus anthracis is the anchoring residue for its surface layer proteins. J. Am. Chem. Soc. 2018, 140, 49. [Google Scholar] [CrossRef]
- Gardiol, A.E.; Dazzo, F.B. Biosynthesis of Rhizobium trifolii capsular polysaccharide: Enzymatic transfer of pyruvate substitutions into lipid-bound saccharide intermediates. J. Bacteriol. 1986, 168, 1459–1462. [Google Scholar] [CrossRef] [PubMed]
- Bossio, J.C.; de Iannino, N.I.; Dankert, M.A. In vitro synthesis of a lipid-linked acetylated and pyruvilated oligosaccharide in Rhizobium trifolii. Biochem. Biophys. Res. Commun. 1986, 134, 205–211. [Google Scholar] [CrossRef]
- Higuchi, Y.; Yoshinaga, S.; Yoritsune, K.; Tateno, H.; Hirabayashi, J.; Nakakita, S.; Kanekiyo, M.; Kakuta, Y.; Takegawa, K. A rationally engineered yeast pyruvyltransferase Pvg1p introduces sialylation-like properties in neo-human-type complex oligosaccharide. Sci. Rep. 2016, 6, 26349. [Google Scholar] [CrossRef]
- Higuchi, Y.; Matsufuji, H.; Tanuma, M.; Arakawa, T.; Mori, K.; Yamada, C.; Shofia, R.; Matsunaga, E.; Tashiro, K.; Fushinobu, S.; et al. Identification and characterization of a novel β-d-galactosidase that releases pyruvylated galactose. Sci. Rep. 2018, 8, 12013. [Google Scholar] [CrossRef] [PubMed]
- Cuthbertson, L.; Kos, V.; Whitfield, C. ABC transporters involved in export of cell surface glycoconjugates. Microbiol. Mol. Biol. Rev. 2010, 74, 341–362. [Google Scholar] [CrossRef] [PubMed]
- Workman, S.D.; Worrall, L.J.; Strynadka, N.C.J. Crystal structure of an intramembranal phosphatase central to bacterial cell-wall peptidoglycan biosynthesis and lipid recycling. Nat. Commun. 2018, 9, 1159. [Google Scholar] [CrossRef] [PubMed]
- Potter, S.C.; Luciani, A.; Eddy, S.R.; Park, Y.; Lopez, R.; Finn, R.D. HMMER web server: 2018 update. Nucleic Acids Res. 2018, 46, W200–W204. [Google Scholar] [CrossRef] [PubMed]
- Foley, G.; Sützl, L.; D’Cunha, S.A.; Gillam, E.M.; Boden, M. SeqScrub: A web tool for automatic cleaning and annotation of FASTA file headers for bioinformatic applications. Biotechniques 2019, 67, 50–54. [Google Scholar] [CrossRef]
- Gerlt, J.A.; Bouvier, J.T.; Davidson, D.B.; Imker, H.J.; Sadkhin, B.; Slater, D.R.; Whalen, K.L. Enzyme Function Initiative-Enzyme Similarity Tool (EFI-EST): A web tool for generating protein sequence similarity networks. Biochim. Biophys. Acta 2015, 1854, 1019–1037. [Google Scholar] [CrossRef]

| Organism | Saccharide (Stereochemistry) | Position of Pyr | Biological Significance * | References | |
|---|---|---|---|---|---|
| Pyr-Saccharide | Pyr-Glycoconjugate | ||||
| Exopolysaccharides | |||||
| Xanthomonas spp. | Man (β) | 4,6 | Plant pathogenesis | [18,28,32] | |
| Xanthomonas campestris | Man (β) | 4,6 | Virulence | [18,28,32] | |
| Sinorhizobium meliloti | Glc (?) | ? | Glycan polymerization and export | [41] | |
| Rhizobium leguminosarum | Gal; Glc (β) | 4,6 | Signalling | [46] | |
| Agrobacterium sp. ZX09 | Glc (?) | ? | Plant symbiosis | [47] | |
| Escherichia coli | Gal (α) | 4,6 | Biofilm | [32,50] | |
| Burkholderia cepacia | Gal (α) | 4,6 | Pathogenesis | [53] | |
| Pseudomonas strain 1.15 | Gal (α) | 4,6 | [54] | ||
| Enterobacter amnigenus | Man (α) | 4,6 | [55] | ||
| Inquilinus limosus | Man (α); Glc (β) | 4,6 | Pathogenesis | [56] | |
| Azorhizobiumcaulinodans | Gal (α) | 4,6 | Plant symbiosis | [59] | |
| Cobetia marina DSMZ 4741 | Kdo (α) | 7,8 | [60] | ||
| Pediococcus pentosaceus | hexose (?) | ? | [61] | ||
| Erwinia futululu | Gal (α) | 4,6 | Pathogenesis | [63] | |
| Agrobacterium radiobacter ATCC 53271 | Glc (α) | 4,6 | [64] | ||
| Methylobacterium sp. | Gal (α) | 4,6 | [65] | ||
| Alteromonas macleodii subsp. fijiensis | Man (?) | 4,6 | [66,67] | ||
| Capsular polysaccharides | |||||
| Streptococcus pneumoniae | Gal (α) | 2,3 | Immunostimulant | [74] | |
| Acinetobacter baumannii | GalNAc (α) | 4,6 | Virulence | [78] | |
| Klebsiella pneumoniae | GlcA (β) | 2,3 | Stimulation of immune system | [79] | |
| Klebsiella serotype K14 | Glc (β) | 4,6 | [82] | ||
| Klebsiella serotype K12 | Galf (α) | 5,6 | [83] | ||
| Klebsiella serotype K70 | Rha (α) | 3,4 | [84] | ||
| Klebsiella serotype K64 | Gal (β) | 4,6 | [85] | ||
| Klebsiella serotype K46 | Man (β) | 4,6 | [86] | ||
| Klebsiella serotype K33 | Gal (β) | 3,4 | [87] | ||
| Klebsiella serotype K3 | Man (α) | 4,6 | Immunogenicity; binding to human mannose receptor | [89] | |
| Bacteroides fragilis | Gal (β) | 4,6 | Virulence | [7] | |
| Rhodococcus equi serotype 4 | RhoAc (α) | 7,9 | Pathogenesis | [95] | |
| Non-classical secondary cell wall polymers | |||||
| Bacillus anthracis | ManNAc (β) | 4,6 | Binding of SLH-proteins | [102] | |
| Bacillus cereus | ManNAc (β) | 4,6 | Binding of SLH-proteins | [17] | |
| Paenibacillus alvei | ManNAc (β) | 4,6 | Binding of SLH-proteins | [15] | |
| Lysinibacillus sphaericus CCM 2177 | ManNAc (β) | 4,6 | Binding of SLH-proteins | [121] | |
| Paenibacillus (Bacillus) polymyxa | ManNAc (β) | 4,6 | [130] | ||
| Secondary cell wall glycopolymers | |||||
| Promicromonospora citrea 665T | Kdn (α); Gal (α) | 7,9; 4,6 | [131] | ||
| Promicromonospora sp. VKM Ac-1028 | GlcA (β) | 2,3 | [131] | ||
| Nocardiopsis metallicus VKM Ac-2522T | Rib-ol-P (?) | 2,4 | [133] | ||
| Nocardiopsis halotolerans | GalNAc (?) | 4,6 | [133] | ||
| Brevibacterium iodinum VKM Ac-2106 | Man-ol; GalNAc (α) | 4,5; 4,6 | [135] | ||
| Lipopolysaccharides and lipooligosaccharides | |||||
| Pseudomonas stutzeri | Glc (β); GlcNAc (β) | 4,6 | Glycoconjugate biosynthesis | [4] | |
| Providencia alcalifaciens | GlcNAc (α) | 4,6 | Pathogenesis | [141] | |
| Shigella dysenteriae | Man (β) | 4,6 | Pathogenesis | [145] | |
| Proteus mirabilis O16 | GalNAc (α) | 4,6 | [147] | ||
| Raoultella terrigena | Man (β) | 4,6 | [146] | ||
| Cobetia pacifica KMM 3878 | Gal (β) | 3,4 | [148] | ||
| Mycobacterial glycolipids | |||||
| Mycobacterium smegmatis | Me-Glc (α) | 4,6 | [150] | ||
| Mycobacterium avium-Mycobacterium intracellulare (MAI) serovar 8 | Me-Glc (α) | 4,6 | Immunostimulant | [151] | |
| MAI serovar 21 | Glc (α) | 4,6 | [151] | ||
| Pyruvylated glycoconjugates in eukaryotes | |||||
| Aplysia kurodai | Gal (β) | 3,4 | [152] | ||
| Schizosaccharomyces pombe | Gal (β) | 4,6 | Charge effect (mimic of sialylation?) | [9] | |
| Red algae | Gal (β) | 4,6 | Immunostimulant | [10,13,161,162] | |
| Green algae | Gal (?) | 3,4 | Anticoagulant | [11,12] | |
| Microciona prolifera | Gal (β) | 4,6 | Species-specific cell reaggregation | [163] | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hager, F.F.; Sützl, L.; Stefanović, C.; Blaukopf, M.; Schäffer, C. Pyruvate Substitutions on Glycoconjugates. Int. J. Mol. Sci. 2019, 20, 4929. https://doi.org/10.3390/ijms20194929
Hager FF, Sützl L, Stefanović C, Blaukopf M, Schäffer C. Pyruvate Substitutions on Glycoconjugates. International Journal of Molecular Sciences. 2019; 20(19):4929. https://doi.org/10.3390/ijms20194929
Chicago/Turabian StyleHager, Fiona F., Leander Sützl, Cordula Stefanović, Markus Blaukopf, and Christina Schäffer. 2019. "Pyruvate Substitutions on Glycoconjugates" International Journal of Molecular Sciences 20, no. 19: 4929. https://doi.org/10.3390/ijms20194929
APA StyleHager, F. F., Sützl, L., Stefanović, C., Blaukopf, M., & Schäffer, C. (2019). Pyruvate Substitutions on Glycoconjugates. International Journal of Molecular Sciences, 20(19), 4929. https://doi.org/10.3390/ijms20194929





