Complete Sequence, Multichromosomal Architecture and Transcriptome Analysis of the Solanum tuberosum Mitochondrial Genome
Abstract
:1. Introduction
2. Results and Discussion
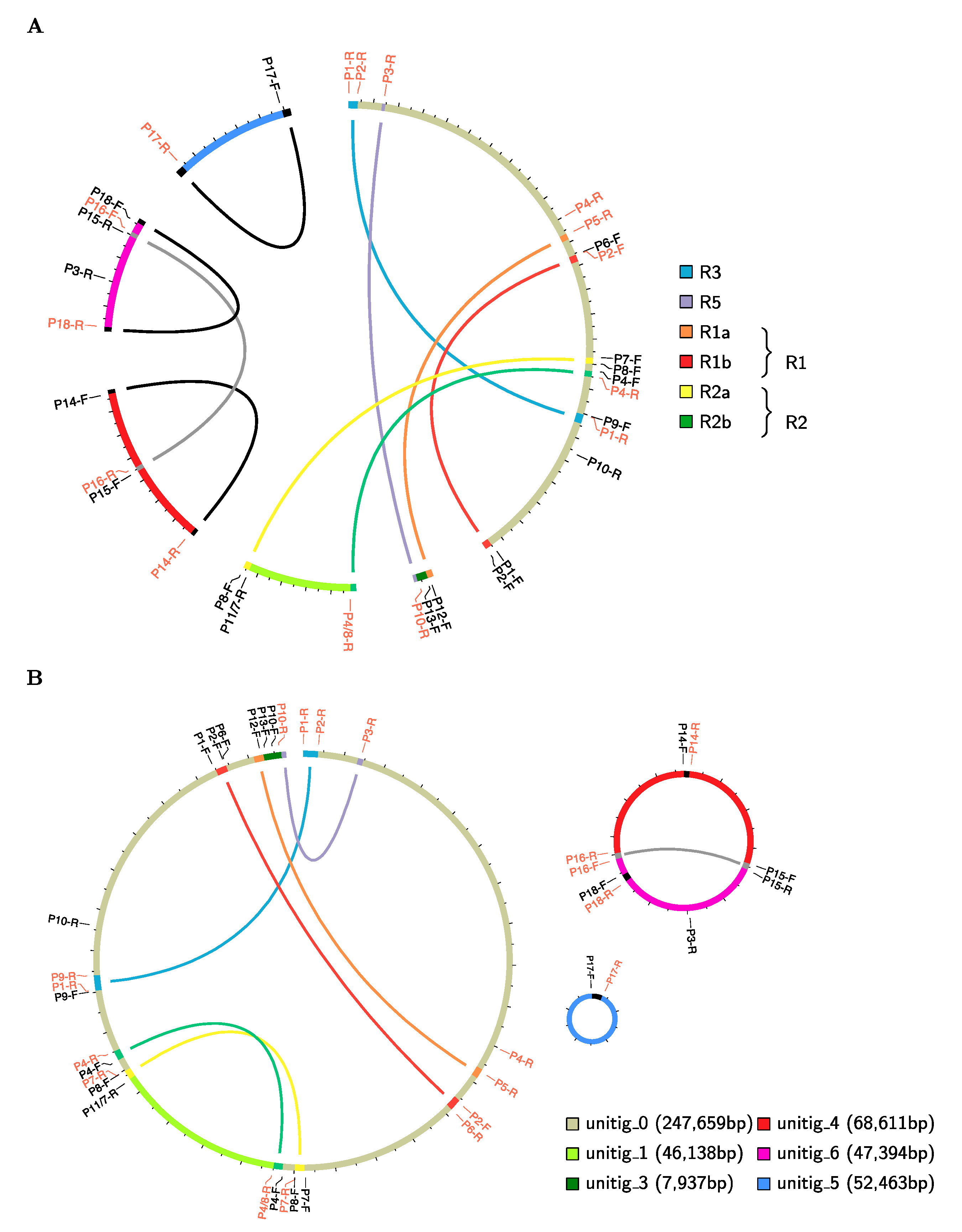
2.1. S. tuberosum Mitochondrial Genome Assembly
2.2. Gene Content of the Potato mtDNA
2.3. Transcriptome of the Potato mtDNA
2.4. RNA Editing
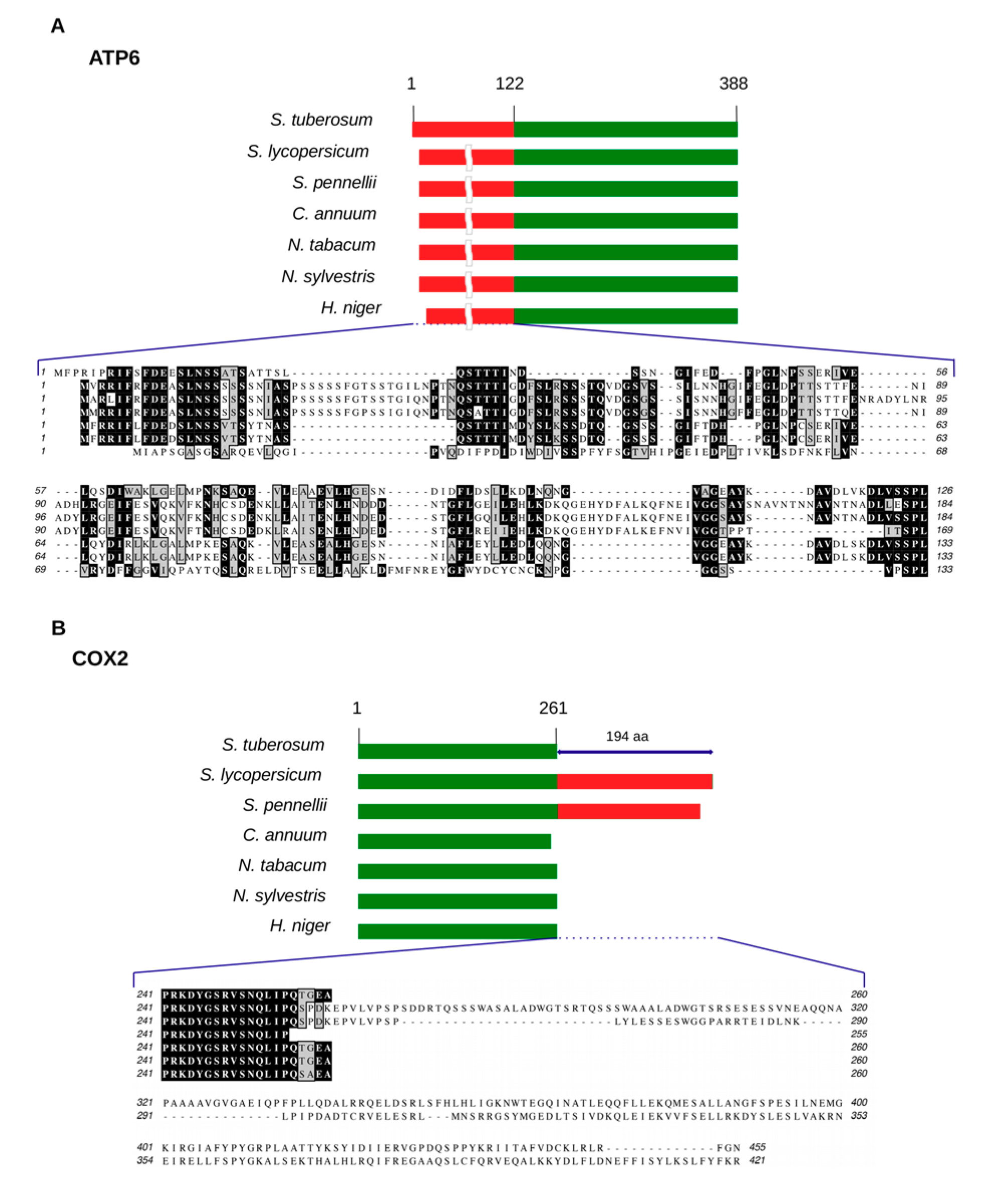
2.5. Comparative Analysis of Mitochondrial Genomes among Solanaceae
3. Materials and Methods
3.1. Plant Material
3.2. Isolation of mtDNA
3.3. Genome Sequencing and Assembly
3.4. Genome Annotation
3.5. Transcriptome Sequencing and Analysis
3.6. Detection of Repeats
3.7. Comparative and Evolutionary Analysis
3.8. Data Deposition
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Palmer, J.D.; Herbon, L.A. Unicircular structure of the Brassica hirta mitochondrial genome. Curr. Genet. 1987, 11, 565–570. [Google Scholar] [CrossRef] [PubMed]
- Sloan, D.B.; Alverson, A.J.; Chuckalovcak, J.P.; Wu, M.; McCauley, D.E.; Palmer, J.D.; Taylor, D.R. Rapid evolution of enormous, multichromosomal genomes in flowering plant mitochondria with exceptionally high mutation rates. PLoS Biol. 2012, 10, e1001241. [Google Scholar] [CrossRef] [PubMed]
- Alverson, A.J.; Wei, X.; Rice, D.W.; Stern, D.B.; Barry, K.; Palmer, J.D. Insights into the evolution of mitochondrial genome size from complete sequences of Citrullus lanatus and Cucurbita pepo (Cucurbitaceae). Mol. Biol. Evol. 2010, 27, 1436–1448. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Moreno, L.; González, V.M.; Benjak, A.; Martí, M.C.; Puigdomènech, P.; Aranda, M.A.; Garcia-Mas, J. Determination of the melon chloroplast and mitochondrial genome sequences reveals that the largest reported mitochondrial genome in plants contains a significant amount of DNA having a nuclear origin. BMC Genom. 2011, 12, 424. [Google Scholar] [CrossRef] [PubMed]
- Morley, S.A.; Nielsen, B.L. Plant mitochondrial DNA. Molecules 2017, 15, 17. [Google Scholar]
- Richardson, A.O.; Palmer, J.D. Horizontal gene transfer in plants. J. Exp. Bot. 2006, 58, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bendich, A.J. Reaching for the ring: The study of mitochondrial genome structure. Curr. Genet. 1993, 24, 279–290. [Google Scholar] [CrossRef]
- Oldenburg, D.J.; Bendich, A.J. Size and structure of replicating mitochondrial DNA in cultured tobacco cells. Plant Cell 1996, 8, 447–461. [Google Scholar] [CrossRef]
- Manchekar, M.; Scissum-Gunn, K.; Song, D.; Khazi, F.; McLean, S.L.; Nielsen, B.L. DNA recombination activity in soybean mitochondria. J. Mol. Biol. 2006, 356, 288–299. [Google Scholar] [CrossRef]
- Gualberto, J.M.; Mileshina, D.; Wallet, C.; Niazi, A.K.; Weber-Lotfi, F.; Dietrich, A. The plant mitochondrial genome: Dynamics and maintenance. Biochimie 2014, 100, 107–120. [Google Scholar] [CrossRef]
- Alverson, A.J.; Rice, D.W.; Dickinson, S.; Barry, K.; Palmer, J.D. Origins and recombination of the bacterial-sized multichromosomal mitochondrial genome of cucumber. Plant Cell 2011, 23, 2499–2513. [Google Scholar] [CrossRef] [PubMed]
- Sloan, D.B. One ring to rule them all? Genome sequencing provides new insights into the ‘master circle’ model of plant mitochondrial DNA structure. New Phytol. 2013, 200, 978–985. [Google Scholar] [CrossRef] [PubMed]
- Davila, J.I.; Arrieta-Montiel, M.P.; Wamboldt, Y.; Cao, J.; Hagmann, J.; Shedge, V.; Xu, Y.-Z.; Weigel, D.; Mackenzie, S.A. Double-strand break repair processes drive evolution of the mitochondrial genome in Arabidopsis. BMC Biol. 2011, 9, 64. [Google Scholar] [CrossRef] [PubMed]
- Shearman, J.R.; Sonthirod, C.; Naktang, C.; Pootakham, W.; Yoocha, T.; Sangsrakru, D.; Jomchai, N.; Tragoonrung, S.; Tangphatsornruang, S. The two chromosomes of the mitochondrial genome of a sugarcane cultivar: Assembly and recombination analysis using long PacBio reads. Sci. Rep. 2016, 6, 31533. [Google Scholar] [CrossRef] [PubMed]
- Kazama, T.; Toriyama, K. Whole mitochondrial genome sequencing and re-examination of a cytoplasmic male sterility-associated gene in Boro-taichung-type cytoplasmic male sterile rice. PLoS ONE 2016, 11, e0159379. [Google Scholar] [CrossRef] [PubMed]
- Tsujimura, M.; Kaneko, T.; Sakamoto, T.; Kimura, S.; Shigyo, M.; Yamagishi, H.; Terachi, T. Multichromosomal structure of the onion mitochondrial genome and a transcript analysis. Mitochondrion 2019, 46, 179–186. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Cuthbert, J.M.; Taylor, D.R.; Sloan, D.B. The massive mitochondrial genome of the angiosperm Silene noctiflora is evolving by gain or loss of entire chromosomes. Proc. Natl. Acad. Sci. USA 2015, 112, 10185–10191. [Google Scholar] [CrossRef]
- Halldén, C.; Lind, C.; Bryngelsson, T. Minicircle variation in Beta mitochondrial DNA. Theor. Appl. Genet. 1989, 77, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Warren, J.M.; Simmons, M.P.; Wu, Z.; Sloan, D.B. Linear plasmids and the rate of sequence evolution in plant mitochondrial genomes. Genome Biol. Evol. 2016, 8, 364–374. [Google Scholar] [CrossRef] [PubMed]
- Backert, S. R-loop-dependent rolling-circle replication and a new model for DNA concatemer resolution by mitochondrial plasmid mp1. EMBO J. 2002, 21, 3128–3136. [Google Scholar] [CrossRef] [Green Version]
- Christensen, A.C. Plant mitochondrial genome evolution can be explained by DNA repair mechanisms. Genome Biol. Evol. 2013, 5, 1079–1086. [Google Scholar] [CrossRef] [PubMed]
- Gualberto, J.M.; Newton, K.J. Plant mitochondrial genomes: Dynamics and mechanisms of mutation. Annu. Rev. Plant Biol. 2017, 68, 225–252. [Google Scholar] [CrossRef] [PubMed]
- Horn, R.; Gupta, K.J.; Colombo, N. Mitochondrion role in molecular basis of cytoplasmic male sterility. Mitochondrion 2014, 19, 198–205. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhao, N.; Li, S.; Grover, C.E.; Nie, H.; Wendel, J.F.; Hua, J. Plant mitochondrial genome evolution and cytoplasmic male sterility. Crit. Rev. Plant Sci. 2017, 36, 55–69. [Google Scholar] [CrossRef]
- Kim, Y.-J.; Zhang, D. Molecular control of male fertility for crop hybrid breeding. Trends Plant Sci. 2018, 23, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Takenaka, M.; van der Merwe, J.A.; Verbitskiy, D.; Neuwirt, J.; Zehrmann, A.; Brennicke, A. RNA Editing in Plant Mitochondria. In RNA Editing; Göringer, H.U., Ed.; Springer Berlin Heidelberg: Berlin/Heidelberg, Germany, 2008; pp. 105–122. [Google Scholar] [CrossRef]
- Takenaka, M.; Verbitskiy, D.; van der Merwe, J.A.; Zehrmann, A.; Brennicke, A. The process of RNA editing in plant mitochondria. Mitochondrion 2008, 8, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Bentolila, S.; Hanson, M.R. The Unexpected Diversity of Plant Organelle RNA Editosomes. Trends Plant Sci. 2016, 21, 962–973. [Google Scholar] [CrossRef] [Green Version]
- Tang, W.; Luo, C. Molecular and Functional Diversity of RNA Editing in Plant Mitochondria. Mol. Biotechnol. 2018, 60, 935–945. [Google Scholar] [CrossRef]
- Oldenkott, B.; Yang, Y.; Lesch, E.; Knoop, V.; Schallenberg-Rüdinger, M. Plant-type pentatricopeptide repeat proteins with a DYW domain drive C-to-U RNA editing in Escherichia coli. Commun. Biol. 2019, 2, 85. [Google Scholar] [CrossRef]
- Hardigan, M.A.; Laimbeer, F.P.E.; Newton, L.; Crisovan, E.; Hamilton, J.P.; Vaillancourt, B.; Wiegert-Rininger, K.; Wood, J.C.; Douches, D.S.; Farré, E.M.; et al. Genome diversity of tuber-bearing Solanum uncovers complex evolutionary history and targets of domestication in the cultivated potato. Proc. Natl. Acad. Sci. USA 2017, 114, E9999. [Google Scholar] [CrossRef]
- The Potato Genome Sequencing Consortium. Genome sequence and analysis of the tuber crop potato. Nature 2011, 475, 189. [Google Scholar] [CrossRef] [PubMed]
- McGregor, C.E.; Lambert, C.A.; Greyling, M.M.; Louw, J.H.; Warnich, L. A comparative assessment of DNA fingerprinting techniques (RAPD, ISSR, AFLP and SSR) in tetraploid potato (Solanum tuberosum L.) germplasm. Euphytica 2000, 113, 135–144. [Google Scholar] [CrossRef]
- Ghislain, M.; Andrade, D.; Rodríguez, F.; Hijmans, R.J.; Spooner, D.M. Genetic analysis of the cultivated potato Solanum tuberosum L. Phureja Group using RAPDs and nuclear SSRs. Theor. Appl. Genet. 2006, 113, 1515–1527. [Google Scholar] [CrossRef]
- Spooner, D.M.; Núñez, J.; Trujillo, G.; del Rosario Herrera, M.; Guzmán, F.; Ghislain, M. Extensive simple sequence repeat genotyping of potato landraces supports a major reevaluation of their gene pool structure and classification. Proc. Natl. Acad. Sci. USA 2007, 104, 19398–19403. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peters, S.A.; Bargsten, J.W.; Szinay, D.; van de Belt, J.; Visser, R.G.F.; Bai, Y.; de Jong, H. Structural homology in the Solanaceae: Analysis of genomic regions in support of synteny studies in tomato, potato and pepper. Plant J. 2012, 71, 602–614. [Google Scholar] [CrossRef]
- Smyda-Dajmund, P.; Śliwka, J.; Wasilewicz-Flis, I.; Jakuczun, H.; Zimnoch-Guzowska, E. Genetic composition of interspecific potato somatic hybrids and autofused 4x plants evaluated by DArT and cytoplasmic DNA markers. Plant Cell Rep. 2016, 35, 1345–1358. [Google Scholar] [CrossRef] [PubMed]
- Berdugo-Cely, J.; Valbuena, R.I.; Sánchez-Betancourt, E.; Barrero, L.S.; Yockteng, R. Genetic diversity and association mapping in the Colombian Central Collection of Solanum tuberosum L. Andigenum group using SNPs markers. PLoS ONE 2017, 12, e0173039. [Google Scholar] [CrossRef] [PubMed]
- Cardi, T.; Bastia, T.; Monti, L.; Earle, E.D. Organelle DNA and male fertility variation in Solanum spp. and interspecific somatic hybrids. Theor. Appl. Genet. 1999, 99, 819–828. [Google Scholar] [CrossRef]
- Lössl, A.; Adler, N.; Horn, R.; Frei, U.; Wenzel, G. Chondriome-type characterization of potato: Mt α, β, γδɛ and novel plastid-mitochondrial configurations in somatic hybrids. Theor. Appl. Genet. 1999, 99, 1–10. [Google Scholar] [CrossRef]
- Scotti, N.; Monti, L.; Cardi, T. Organelle DNA variation in parental Solanum spp. genotypes and nuclear-cytoplasmic interactions in Solanum tuberosum (+) S. commersonii somatic hybrid-backcross progeny. Theor. Appl. Genet. 2003, 108, 87–94. [Google Scholar] [CrossRef]
- Scotti, N.; Cozzolino, S.; Cardi, T. Mitochondrial DNA variation in cultivated and wild potato species (Solanum spp.). Genome 2007, 50, 706–713. [Google Scholar] [CrossRef] [PubMed]
- Lössl, A.; Götz, M.; Braun, A.; Wenzel, G. Molecular markers for cytoplasm in potato: Male sterility and contribution of different plastid-mitochondrial configurations to starch production. Euphytica 2000, 116, 221–230. [Google Scholar] [CrossRef]
- Brandt, P.; Unseld, M.; Eckert-Ossenkopp, U.; Brennicke, A. An rps14 pseudogene is transcribed and edited in Arabidopsis mitochondria. Curr. Genet. 1993, 24, 330–336. [Google Scholar] [CrossRef] [PubMed]
- Quiñones, V.; Zanlungo, S.; Moenne, A.; Gómez, I.; Holuigue, L.; Litvak, S.; Jordana, X. The rpl5-rps 14-cob gene arrangement in Solanum tuberosum: rps14 is a transcribed and unedited pseudogene. Plant Mol. Biol. 1996, 31, 937–943. [Google Scholar] [CrossRef] [PubMed]
- Scotti, N.; Maréchal-Drouard, L.; Cardi, T. The rpl5–rps14 mitochondrial region: A hot spot for DNA rearrangements in Solanum spp. somatic hybrids. Curr. Genet. 2004, 45, 378–382. [Google Scholar] [CrossRef] [PubMed]
- Figueroa, P.; Gómez, I.; Holuigue, L.; Araya, A.; Jordana, X. Transfer of rps14 from the mitochondrion to the nucleus in maize implied integration within a gene encoding the iron–sulphur subunit of succinate dehydrogenase and expression by alternative splicing. Plant J. 1999, 18, 601–609. [Google Scholar] [CrossRef]
- Ong, H.C.; Palmer, J.D. Pervasive survival of expressed mitochondrial rps14 pseudogenes in grasses and their relatives for 80 million years following three functional transfers to the nucleus. BMC Evol. Biol. 2006, 6, 55. [Google Scholar] [CrossRef]
- Bastia, T.; Scotti, N.; Cardi, T. Organelle DNA analysis of Solanum and Brassica somatic hybrids by PCR with‘universal primers’. Theor. Appl. Genet. 2001, 102, 1265–1272. [Google Scholar] [CrossRef]
- Maréchal-Drouard, L.; Guillemaut, P.; Cosset, A.; Arbogast, M.; Weber, F.; Weil, J.H.; Dietrich, A. Transfer RNAs of potato (Solanum tuberosum) mitochondria have different genetic origins. Nucleic Acids Res. 1990, 18, 3689–3696. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, A.; Wallet, C.; Iqbal, R.K.; Gualberto, J.M.; Lotfi, F. Organellar non-coding RNAs: Emerging regulation mechanisms. Biochimie 2015, 117, 48–62. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Kang, J.G.; Kim, B.-D. Isolation and characterization of the cytoplasmic male sterility-associated orf456 gene of chili pepper (Capsicum annuum L.). Plant Mol. Biol. 2007, 63, 519–532. [Google Scholar] [CrossRef] [PubMed]
- Fey, J.; Maréchal-Drouard, L. Compilation and Analysis of Plant Mitochondrial Promoter Sequences: An Illustration of a Divergent Evolution between Monocot and Dicot Mitochondria. Biochem. Biophys. Res. Commun. 1999, 256, 409–414. [Google Scholar] [CrossRef] [PubMed]
- Kühn, K.; Weihe, A.; Börner, T. Multiple promoters are a common feature of mitochondrial genes in Arabidopsis. Nucleic Acids Res. 2005, 33, 337–346. [Google Scholar] [CrossRef] [Green Version]
- Kühn, K.; Bohne, A.-V.; Liere, K.; Weihe, A.; Börner, T. Arabidopsis phage-type RNA polymerases: Accurate in vitro transcription of organellar genes. Plant Cell 2007, 19, 959–971. [Google Scholar] [CrossRef] [PubMed]
- Forner, J.; Weber, B.; Thuss, S.; Wildum, S.; Binder, S. Mapping of mitochondrial mRNA termini in Arabidopsis thaliana: T-elements contribute to 5’ and 3’ end formation. Nucleic Acids Res. 2007, 35, 3676–3692. [Google Scholar] [CrossRef]
- Palmer, J.D.; Herbon, L.A. Plant mitochondrial DNA evolved rapidly in structure, but slowly in sequence. J. Mol. Evol. 1988, 28, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Zanlungo, S.; Quinones, V.; Moenne, A.; Holuigue, L.; Jordana, X. Splicing and editing of rps10 transcripts in potato mitochondria. Curr. Genet. 1995, 27, 565–571. [Google Scholar] [CrossRef] [PubMed]
- Zanlungo, S.; Quiñones, V.; Holuigue, L.; Jordana, X. Identification of the RPS10 polypeptide encoded by the mitochondrial genome in Solanum tuberosum: Translation initiates at a genomic-encoded AUG and not at a conserved AUG codon created by RNA editing. Physiol. Plant 2000, 110, 256–261. [Google Scholar] [CrossRef]
- Quiñones, V.; Zanlungo, S.; Holuigue, L.; Litvak, S.; Jordana, X. The cox1 initiation codon is created by RNA editing in potato mitochondria. Plant Physiol. 1995, 108, 1327. [Google Scholar] [CrossRef]
- Dell’Orto, P.; Moenne, A.; Graves, P.V.; Jordana, X. The potato mitochondrial ATP synthase subunit 9: Gene structure, RNA editing and partial protein sequence. Plant Sci. 1993, 88, 45–53. [Google Scholar] [CrossRef]
- Hunt, M.D.; Newton, K.J. The NCS3 mutation: Genetic evidence for the expression of ribosomal protein genes in Zea mays mitochondria. EMBO J. 1991, 10, 1045–1052. [Google Scholar] [CrossRef] [PubMed]
- Takemura, M.; Oda, K.; Yamato, K.; Ohta, E.; Nakamura, Y.; Nozato, N.; Akashi, K.; Ohyama, K. Gene clusters for ribosomal proteins in the mitochondrial genome of a liverwort, Marchantia polymorpha. Nucleic Acids Res. 1992, 20, 3199–3205. [Google Scholar] [CrossRef] [PubMed]
- Giegé, P.; Brennicke, A. RNA editing in Arabidopsis mitochondria effects 441 C to U changes in ORFs. Proc. Natl. Acad. Sci. USA 1999, 96, 15324–15329. [Google Scholar] [CrossRef] [PubMed]
- Marechal-Drouard, L.; Cosset, A.; Remacle, C.; Ramamonjisoa, D.; Dietrich, A. A single editing event is a prerequisite for efficient processing of potato mitochondrial phenylalanine tRNA. Mol. Cell. Biol. 1996, 16, 3504–3510. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fey, J.; Tomita, K.; Bergdoll, M.; MarÉChal-Drouard, L. Evolutionary and functional aspects of C-to-U editing at position 28 of tRNACys(GCA) in plant mitochondria. RNA 2000, 6, 470–474. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ngu, M.; Massel, K.; Bonen, L. Group II introns in wheat mitochondria have degenerate structural features and varied splicing pathways. Int. J. Biochem. Cell Biol. 2017, 91, 156–167. [Google Scholar] [CrossRef]
- Cognat, V.; Morelle, G.; Megel, C.; Lalande, S.; Molinier, J.; Vincent, T.; Small, I.; Duchêne, A.-M.; Maréchal-Drouard, L. The nuclear and organellar tRNA-derived RNA fragment population in Arabidopsis thaliana is highly dynamic. Nucleic Acids Res. 2016, 45, 3460–3472. [Google Scholar] [CrossRef]
- Guo, W.; Zhu, A.; Fan, W.; Mower, J.P. Complete mitochondrial genomes from the ferns Ophioglossum californicum and Psilotum nudum are highly repetitive with the largest organellar introns. New Phytol. 2017, 213, 391–403. [Google Scholar] [CrossRef]
- Adams, K.L.; Daley, D.O.; Whelan, J.; Palmer, J.D. Genes for two mitochondrial ribosomal proteins in flowering plants are derived from their chloroplast or cytosolic counterparts. Plant Cell 2002, 14, 931–943. [Google Scholar] [CrossRef]
- Notsu, Y.; Masood, S.; Nishikawa, T.; Kubo, N.; Akiduki, G.; Nakazono, M.; Hirai, A.; Kadowaki, K. The complete sequence of the rice (Oryza sativa L.) mitochondrial genome: Frequent DNA sequence acquisition and loss during the evolution of flowering plants. Mol. Genet. Genom. 2002, 268, 434–445. [Google Scholar] [CrossRef]
- Aubert, D.; Bisanz-Seyer, C.; Herzog, M. Mitochondrial rps14 is a transcribed and edited pseudogene in Arabidopsis thaliana. Plant Mol. Biol. 1992, 20, 1169–1174. [Google Scholar] [CrossRef] [PubMed]
- Unseld, M.; Marienfeld, J.R.; Brandt, P.; Brennicke, A. The mitochondrial genome of Arabidopsis thaliana contains 57 genes in 366,924 nucleotides. Nat. Genet. 1997, 15, 57–61. [Google Scholar] [CrossRef] [PubMed]
- Kubo, N.; Harada, K.; Hirai, A.; Kadowaki, K. A single nuclear transcript encoding mitochondrial RPS14 and SDHB of rice is processed by alternative splicing: Common use of the same mitochondrial targeting signal for different proteins. Proc. Natl. Acad. Sci. USA 1999, 96, 9207–9211. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mohanty, A.; Martín, J.P.; González, L.M.; Aguinagalde, I. Association between chloroplast DNA and mitochondrial DNA haplotypes in Prunus spinosa L. (Rosaceae) populations across Europe. Ann. Bot. 2003, 92, 749–755. [Google Scholar] [CrossRef] [PubMed]
- Bartoszewski, G.; Malepszy, S.; Havey, M.J. Mosaic (MSC) cucumbers regenerated from independent cell cultures possess different mitochondrial rearrangements. Curr. Genet. 2004, 45, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Sandoval, P.; León, G.; Gómez, I.; Carmona, R.; Figueroa, P.; Holuigue, L.; Araya, A.; Jordana, X. Transfer of RPS14 and RPL5 from the mitochondrion to the nucleus in grasses. Gene 2004, 324, 139–147. [Google Scholar] [CrossRef]
- Adams, K.L.; Qiu, Y.-L.; Stoutemyer, M.; Palmer, J.D. Punctuated evolution of mitochondrial gene content: High and variable rates of mitochondrial gene loss and transfer to the nucleus during angiosperm evolution. Proc. Natl. Acad. Sci. USA 2002, 99, 9905–9912. [Google Scholar] [CrossRef] [Green Version]
- Bergthorsson, U.; Adams, K.L.; Thomason, B.; Palmer, J.D. Widespread horizontal transfer of mitochondrial genes in flowering plants. Nature 2003, 424, 197–201. [Google Scholar] [CrossRef] [Green Version]
- Park, S.; Grewe, F.; Zhu, A.; Ruhlman, T.A.; Sabir, J.; Mower, J.P.; Jansen, R.K. Dynamic evolution of Geranium mitochondrial genomes through multiple horizontal and intracellular gene transfers. New Phytol. 2015, 208, 570–583. [Google Scholar] [CrossRef]
- Handa, H. The complete nucleotide sequence and RNA editing content of the mitochondrial genome of rapeseed (Brassica napus L.): Comparative analysis of the mitochondrial genomes of rapeseed and Arabidopsis thaliana. Nucleic Acids Res. 2003, 31, 5907–5916. [Google Scholar] [CrossRef]
- Clifton, S.W.; Minx, P.; Fauron, C.M.R.; Gibson, M.; Allen, J.O.; Sun, H.; Thompson, M.; Barbazuk, W.B.; Kanuganti, S.; Tayloe, C.; et al. Sequence and comparative analysis of the maize NB mitochondrial genome. Plant Physiol. 2004, 136, 3486–3503. [Google Scholar] [CrossRef] [PubMed]
- Luo, D.; Xu, H.; Liu, Z.; Guo, J.; Li, H.; Chen, L.; Fang, C.; Zhang, Q.; Bai, M.; Yao, N.; et al. A detrimental mitochondrial-nuclear interaction causes cytoplasmic male sterility in rice. Nat. Genet. 2013, 45, 573. [Google Scholar] [CrossRef] [PubMed]
- Osman, C.; Wilmes, C.; Tatsuta, T.; Langer, T. Prohibitins interact genetically with Atp23, a novel processing peptidase and chaperone for the F1Fo-ATP synthase. Mol. Biol. Cell 2007, 18, 627–635. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Neupert, W.; Tzagoloff, A. The metalloprotease encoded by ATP23 has a dual function in processing and assembly of subunit 6 of mitochondrial ATPase. Mol. Biol. Cell 2007, 18, 617–626. [Google Scholar] [CrossRef] [PubMed]
- Fox, T.D. Mitochondrial protein synthesis, import, and assembly. Genetics 2012, 192, 1203–1234. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M.; Ohta, T. On some principles governing molecular evolution. Proc. Natl. Acad. Sci. USA 1974, 71, 2848–2852. [Google Scholar] [CrossRef] [PubMed]
- Echave, J.; Spielman, S.J.; Wilke, C.O. Causes of evolutionary rate variation among protein sites. Nat. Rev. Genet. 2016, 17, 109–121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Darracq, A.; Varré, J.-S.; Maréchal-Drouard, L.; Courseaux, A.; Castric, V.; Saumitou-Laprade, P.; Oztas, S.; Lenoble, P.; Vacherie, B.; Barbe, V. Structural and content diversity of mitochondrial genome in beet: A comparative genomic analysis. Genome Biol. Evol. 2011, 3, 723–736. [Google Scholar] [CrossRef]
- Aljohi, H.A.; Liu, W.; Lin, Q.; Zhao, Y.; Zeng, J.; Alamer, A.; Alanazi, I.O.; Alawad, A.O.; Al-Sadi, A.M.; Hu, S.; et al. Complete Sequence and Analysis of Coconut Palm (Cocos nucifera) Mitochondrial Genome. PLoS ONE 2016, 11, e0163990. [Google Scholar] [CrossRef]
- Bi, C.; Paterson, A.H.; Wang, X.; Xu, Y.; Wu, D.; Qu, Y.; Jiang, A.; Ye, Q.; Ye, N. Analysis of the Complete Mitochondrial Genome Sequence of the Diploid Cotton Gossypium raimondii by Comparative Genomics Approaches. BioMed Res. Int. 2016, 2016. [Google Scholar] [CrossRef]
- Wang, X.; Cheng, F.; Rohlsen, D.; Bi, C.; Wang, C.; Xu, Y.; Wei, S.; Ye, Q.; Yin, T.; Ye, N. Organellar genome assembly methods and comparative analysis of horticultural plants. Hortic. Rese. 2018, 5, 3. [Google Scholar] [CrossRef] [PubMed]
- Olmstead, R.G.; Bohs, L.; Migid, H.A.; Santiago-Valentin, E.; Garcia, V.F.; Collier, S.M. A Molecular Phylogeny of the Solanaceae. Taxon 2008, 57, 1159–1181. [Google Scholar] [CrossRef]
- Pujol, C.; Bailly, M.; Kern, D.; Maréchal-Drouard, L.; Becker, H.; Duchêne, A.-M. Dual-targeted tRNA-dependent amidotransferase ensures both mitochondrial and chloroplastic Gln-tRNAGln synthesis in plants. Proc. Natl. Acad. Sci. USA 2008, 105, 6481–6485. [Google Scholar] [CrossRef] [PubMed]
- Scotti, N.; Cardi, T.; Marechaldrouard, L. Mitochondrial DNA and RNA isolation from small amounts of potato tissue. Plant Mol. Biol. Rep. 2001, 19, 67. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics (Oxford, UK) 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Zerbino, D.R.; Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008, 18, 821–829. [Google Scholar] [CrossRef] [PubMed]
- Bashir, A.; Klammer, A.; Robins, W.P.; Chin, C.-S.; Webster, D.; Paxinos, E.; Hsu, D.; Ashby, M.; Wang, S.; Peluso, P.; et al. A hybrid approach for the automated finishing of bacterial genomes. Nat. Biotechnol. 2012, 30, 701–707. [Google Scholar] [CrossRef] [PubMed]
- Luo, R.; Liu, B.; Xie, Y.; Li, Z.; Huang, W.; Yuan, J.; He, G.; Chen, Y.; Pan, Q.; Liu, Y.; et al. SOAPdenovo2: An empirically improved memory-efficient short-read de novo assembler. GigaScience 2012, 1, 18. [Google Scholar] [CrossRef] [PubMed]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef]
- Holt, K.E.; Schultz, M.B.; Wick, R.R.; Zobel, J. Bandage: Interactive visualization of de novo genome assemblies. Bioinformatics (Oxford, UK) 2015, 31, 3350–3352. [Google Scholar]
- Vincent, T.; Vingadassalon, A.; Ubrig, E.; Azeredo, K.; Srour, O.; Cognat, V.; Graindorge, S.; Salinas, T.; Maréchal-Drouard, L.; Duchêne, A.-M. A genome-scale analysis of mRNAs targeting to plant mitochondria: Upstream AUGs in 5’ untranslated regions reduce mitochondrial association. Plant J. 2017, 92, 1132–1142. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics (Oxford, UK) 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
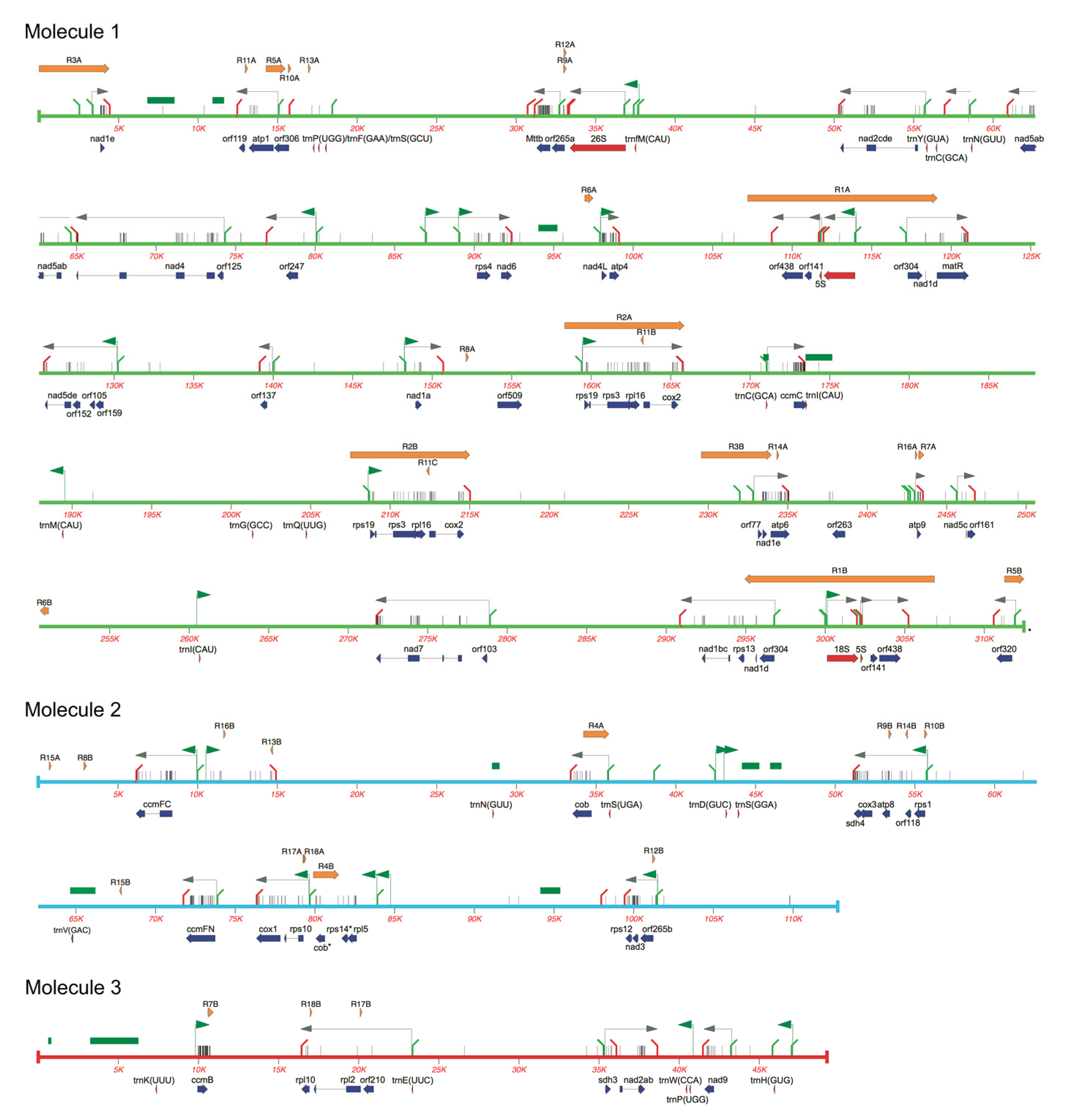


| Molecule 1 | ||||
| Gene | Start | Stop | Strand | Comments |
| nad1e | 3860 | 4118 | + | |
| orf119 | 12570 | 12929 | – | |
| atp1 | 13197 | 14732 | – | |
| Mttb | 31292 | 32122 | – | |
| orf265a | 32244 | 33041 | – | N-term atp8 (20 codons), ELF-domain (pfam03317). Co-transcribed with MttB |
| 26S | 33385 | 36879 | – | |
| nad2cde | 50394 | 50581 | – | |
| 52050 | 52622 | – | ||
| 55094 | 55254 | – | ||
| nad5ab | 61695 | 62910 | – | |
| 63755 | 63984 | – | ||
| nad4 | 64989 | 65077 | – | |
| 67705 | 68127 | – | ||
| 71259 | 71773 | – | ||
| 73185 | 73645 | – | ||
| orf125 | 73849 | 74226 | – | |
| orf247 | 78169 | 78912 | – | |
| rps4 | 90192 | 91013 | + | No evidence of TAG created by editing. Possible non-canonical initiation at GTG codon |
| nad6 | 91707 | 92360 | + | Transcript is processed upstream of stop codon |
| nad4L | 98027 | 98329 | + | |
| atp4 | 98518 | 99114 | + | |
| orf438 | 109344 | 110660 | – | RdRp-like |
| orf141 | 110784 | 111209 | – | |
| 5S | 111720 | 111838 | – | |
| 18S | 112001 | 113946 | – | |
| orf304 | 117275 | 118189 | + | N-term pfam12725. C-term has 59% identity to hypothetical protein RirG_027070 |
| nad1d | 118391 | 118449 | + | |
| matR | 119111 | 121087 | + | |
| nad5de | 125647 | 125793 | – | |
| 126889 | 127283 | – | ||
| orf152 | 127397 | 127855 | – | |
| orf105 | 128459 | 128776 | – | |
| orf159 | 128839 | 129318 | – | RdRp-like |
| orf137 | 139197 | 139610 | – | CMS-associated protein |
| nad1a | 148955 | 149339 | + | |
| rps19 | 159585 | 159869 | + | |
| rps3 | 159883 | 159956 | + | |
| 161026 | 162643 | + | ||
| rpl16 | 162534 | 163049 | + | Editing site 162570 (96%) creates internal stop codon. Editing is conserved in Arabidopsis. It implies that there is no re-initiation of translation inside rps3. Possible initiation at GTG codon. |
| cox2 | 163299 | 163680 | + | |
| 165066 | 165466 | + | ||
| ccmC | 172753 | 173583 | + | ORF overlaps with tRNA and transcript is processd at the tRNA 5’, without stop codon. |
| rps19 | 208754 | 209038 | + | |
| rps3 | 209052 | 209125 | + | |
| 210195 | 211812 | + | ||
| rpl16 | 211703 | 212218 | + | |
| cox2 | 212468 | 212849 | + | |
| 214235 | 214635 | + | ||
| orf77 | 233142 | 233375 | + | |
| nad1e | 233431 | 233689 | + | |
| atp6 | 233940 | 235106 | + | Editing site 235065 creates stop codon making orf 13 aa shorter, with the same C-term as Arabidopsis atp6. |
| atp9 | 243146 | 243379 | + | Editing site 243368 creates stop codon making protein 3 aa shorter. With editing it is the same C-term as Arabidopsis atp9. |
| nad5c | 246225 | 246246 | + | |
| orf161 | 246314 | 246799 | + | |
| nad7 | 271765 | 272026 | – | |
| 273770 | 274477 | – | ||
| 275936 | 276004 | – | ||
| 276920 | 277062 | – | ||
| orf103 | 278419 | 278730 | – | |
| nad1bc | 292250 | 292441 | – | |
| 293925 | 294007 | – | ||
| rps13 | 294547 | 294897 | – | |
| nad1d | 295623 | 295681 | – | |
| orf304 | 295883 | 296797 | – | hypothetical protein RirG (Rhizophagus irregularis) |
| 18S | 300126 | 302071 | + | |
| 5S | 302234 | 302352 | + | |
| orf141 | 302863 | 303288 | + | Similarities to region 3’ UTR of orf247 |
| orf438 | 303412 | 304728 | + | RdRp-like |
| orf320 | 310786 | 311748 | – | Chimeric orf: 5’ of atp1, 3’ region upstream nad5c. Promoter of atp1 that is present in repeat R5 |
| Molecule 2 | ||||
| Gene | Start | Stop | Strand | Comments |
| ccmFC | 6136 | 6685 | – | |
| 7635 | 8401 | – | ||
| Cob | 33524 | 34705 | – | |
| sdh4 | 51176 | 51589 | – | Overlaps cox3. Real ATG might be at codon 24. The transcript is processed about 8 codons before stop codon. Internal stop codon created by partial editing (26%) at codon 93. |
| cox3 | 51517 | 52314 | – | |
| atp8 | 52947 | 53417 | – | |
| orf118 | 54391 | 54747 | – | Chimeric orf: C-term is from atp6 |
| rps1 | 54964 | 55635 | – | |
| ccmFN | 71926 | 73737 | – | |
| cox1 | 76329 | 77825 | – | Initiation codon created by editing |
| rps10 | 78075 | 78187 | – | Stop codon created by editing (78107) |
| 78963 | 79212 | – | Initiation codon created by editing (79211). | |
| rps14 * | 81682 | 82051 | – | Pseudo-gene |
| rpl5 | 82053 | 82613 | – | |
| rps12 | 99496 | 99867 | – | |
| nad3 | 99916 | 100272 | – | |
| orf265b | 100423 | 101220 | – | N-term atp8, ELF-domain (pfam03317) |
| Molecule 3 | ||||
| Gene | Start | Stop | Strand | Comments |
| ccmB | 9945 | 10565 | + | Editing site 10238 creates stop codon in about 50% of the transcripts, in the middle of the ORF. |
| rpl10 | 16440 | 16919 | – | |
| rpl2 | 17204 | 17320 | – | |
| 19230 | 20114 | – | ||
| orf210 | 20294 | 20926 | – | |
| sdh3 | 35400 | 35726 | + | |
| nad2ab | 36302 | 36453 | + | |
| 37470 | 37862 | + | ||
| nad9 | 41589 | 42161 | – | |
| tRNA Gene | Start | Stop | Strand | Editing NGS | Expression |
|---|---|---|---|---|---|
| Molecule 1 | |||||
| trnP(UGG) | 17220 | 17294 | - | + | |
| trnF(GAA) | 17545 | 17618 | - | + | + |
| trnS(GCU) | 17981 | 18068 | - | + | |
| trnMf(CAU) | 37458 | 37531 | - | + | |
| trnY(GUA) | 55772 | 55854 | - | + | |
| trnN(GUU) * | 56390 | 56461 | - | + | |
| trnC(GCA) | 58606 | 58676 | - | + | + |
| trnC(GCA) * | 171005 | 171076 | + | - | |
| trnI(CAU) * | 173488 | 173561 | + | nd | |
| trnMe(CAU) * | 189363 | 189435 | - | + | |
| trnG(GCC) | 201326 | 201397 | + | + | |
| trnQ(UUG) | 204702 | 204773 | + | + | |
| trnI(CAU) | 260636 | 260709 | + | + | |
| Molecule 2 | |||||
| trnN(GUU) * | 28511 | 28582 | + | + | |
| trnS(UGA) | 35798 | 35884 | - | + | |
| trnD(GUC) * | 43141 | 43214 | + | + | |
| trnS(GGA) * | 43901 | 43987 | + | + | |
| trnV(GAC) * | 64733 | 64804 | - | - | |
| Molecule 3 | |||||
| trnK(UUU) | 7326 | 7398 | - | + | |
| trnE(UUC) | 23306 | 23377 | - | + | |
| trnW(CCA) * | 40411 | 40484 | - | + | |
| trnP(UGG) | 40642 | 40715 | - | + | |
| trnH(GUG) * | 45897 | 45971 | - | + |
| Arabidopsis thaliana | ||||
| Gene | mRNA End | Kind of Secondary Structure | Putative Nuclease | Conservation in Potato |
| ccmFC | 5’ | trnG | RNAseZ | No |
| rps3 | 5’ | trnK | RNAseZ | No |
| rps4 | 5’ | t-element | RNAseZ | No |
| ccmFN1 | 5’ | t-element | RNAseZ | No |
| cox1 | 5’ | t-element | RNAseZ | No |
| rpl5 | 5’ | Acceptor stem-like stem–loop | RNAseZ | No |
| rpl5 | 5’ | Acceptor stem-like stem–loop | RNAseZ | No |
| atp6-2 | 5’ | Acceptor stem-like stem–loop | RNAseZ | No |
| nad7 | 5’ | Acceptor stem-like stem–loop | RNAseP | No |
| atp6-1 | 3’ | trnS | RNAseP | No |
| atp6-2 | 3’ | trnS | RNAseP | No |
| atp9 | 3’ | Double stem-loop | RNAseZ | No |
| nad1e | 3’ | Double stem-loop | RNAseZ | No |
| cox2 | 3’ | Stem-loop | RNAseZ | No |
| ccmC | 3’ | t-element | RNAseP | trnI |
| nad6 | 3’ | t-element | RNAseP | Yes |
| Solanum tuberosum | ||||
| rrrn26S | 5’ precursor | trnfM | RNAseZ | |
| nad2cde | 5’ | trnY | RNAseZ | |
| non-coding RNA | 5’ | trnC | RNAseZ | |
| Cob | 5’ | trnS | RNAseZ | |
| ccmC | 5’ | trnC | RNAseZ | |
| ccmC | 3’ | trnI | RNaseP | |
| atp1 | 3’ | Double stem-loop | RNAseZ | 12397–12441 |
| mttB | 3’ | Stem-loop | RNAseZ | 31111–31148 |
| nad5ab | 3’ | Double stem-loop | RNAseZ | 60956–60986 |
| orf247 | 3’ | Double stem-loop | RNAseZ | 76895–76938 |
| nad6 | 3’ | t-element | RNAseZ | 92309–92353 |
| atp4 | 3’ | Stem-loop | RNAseZ | 99142–99166 |
| nad1a | 3’ | Stem-loop | RNAseZ | 150600–150637 |
| orf438 | 3’ | Stem-loop | RNAseZ | 305247–305291 |
| Gene | S. tuberosum | S. commersonii | S. lycopersicum | S. pennellii | C. annuum | N. tabacum | N. sylvestris | H. niger |
|---|---|---|---|---|---|---|---|---|
| atp1 | ● | ● | ● | ● | ● | ● | ● | ● |
| atp4 | ● | ● | ● | ● | ● | ● | ● | ● |
| atp6 | ● | ● a | ● | ● | ● | ● | ● | ● |
| atp8 | ● | ● | ● | ● | ● | ● | ● | ● |
| atp9 | ● | ● | ● | ● | ● | ● | ● | ● |
| ccmB | ● | ● | ● | ● | ● | ● | ● | ● |
| ccmC | ● | ● | ● | ● | ● | ● | ● | ● |
| ccmFc | ● | ● | ● | ● | ● | ● | ● | ● |
| ccmFN | ● | ● | ● | ● | ● | ● | ● | ● |
| cob | ● | ● | ● | ● | ● | ● | ● | ● |
| cox1 | ● | ● | ● | ● | ● | ● | ● | ● |
| cox2 | ● | ● | ● | ● | ● | ● | ● | ● |
| cox3 | ● | ● | ● | ● | ● | ● | ● | ● |
| matR | ● | ● | ● | ● | ● | ● | ● | ● |
| mttB | ● | ● | ● | ● | ● | ● | ● | ● |
| nad1 | ● | ● | ● | ● | ● | ● | ● | ● |
| nad2 | ● | ● | ● | ● | ● | ● | ● | ● |
| nad3 | ● | ● | ● | ● | ● | ● | ● | ● |
| nad4 | ● | ● | ● | ● | ● | ● | ● | ● |
| nad4L | ● | ● | ● | ● | ● | ● | ● | ● |
| nad5 | ● | ● | ● | ● | ● | ● | ● | ● |
| nad6 | ● | ● | ● | ● | ● | ● | ● | ● |
| nad7 | ● | ● | ● | ● | ● | ● | ● | ● |
| nad9 | ● | ● | ● | ● | ● | ● | ● | ● |
| rpl2 | ● | ● | ● | ● | ● | ● | ● | ● |
| rpl5 | ● | ● | ● | ● | ● | ● | ● | ● |
| rpl10 | ● | ● | ● | ● | ● | ● | ● | ● |
| rpl16 | ● | ● | ● | ● | ● | ● | ● | ● |
| rps1 | ● | ● | ● | ● | ● | ● | ● | ● |
| rps3 | ● | ● | ● | ● | ● | ● | ● | ● |
| rps4 | ● | ● | ● | ● | ● | ● | ● | ● |
| rps10 | ● | ● | ● | ● | ● | ● | ● | ● |
| rps12 | ● | ● | ● | ● | ● | ● | ● | ● |
| rps13 | ● | ● | ● | ● | ● | ● | ● | ● |
| rps14 | ψ | ψ | ψ | ψ | ψ | ψ | ψ | ψ |
| rps19 | ● | ● | ● | ● | ● | ● | ● | ● |
| sdh3 | ● | ● | ● | ● | ● | ● | ● | ● |
| sdh4 | ● | ● | ● | ● | ● | ● | ● | ● |
| orf77 | ● | ● | ● | ● | ● | ● | ● | - |
| orf103 | ● | ● | ● | ● | ● | ● | ● | ● |
| orf105 | ● | ● | ● | ● | ● | ● | ● | - |
| orf118 | ● | ● | ψ | ψ | - | - | - | - |
| orf119 | ● | ● | ● | ● | ● | ● | ● | ● |
| orf125 | ● | - | - | - | - | - | - | - |
| orf137 | ● | - | - | - | - | - | - | - |
| orf141 | ● | ● | ● | ● | - | - | - | - |
| orf152 | ● | ● | ● | ● | ● | ● | ● | - |
| orf159 | ● | ● | ● | ● | ψ | - | - | - |
| orf161 | ● | ● | ● | ● | ● | ● | ● | ψ |
| orf210 | ● | ● | ● | ● | - | - | - | - |
| orf247 | ● | ● | ● | ● | - | - | - | - |
| orf265a | ● | ● | ● | ● | ● | ● | ● | ψ |
| orf265b | ● | ● | ● | ● | ● | ● | ● | ψ |
| orf304 | ● | ● | ● | ● | ψ | - | - | - |
| orf320 | ● | ● | ● | ● | ● | ● | ● | ● |
| orf438 | ● | ● | ● | ● | - | - | - | - |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Varré, J.-S.; D’Agostino, N.; Touzet, P.; Gallina, S.; Tamburino, R.; Cantarella, C.; Ubrig, E.; Cardi, T.; Drouard, L.; Gualberto, J.M.; et al. Complete Sequence, Multichromosomal Architecture and Transcriptome Analysis of the Solanum tuberosum Mitochondrial Genome. Int. J. Mol. Sci. 2019, 20, 4788. https://doi.org/10.3390/ijms20194788
Varré J-S, D’Agostino N, Touzet P, Gallina S, Tamburino R, Cantarella C, Ubrig E, Cardi T, Drouard L, Gualberto JM, et al. Complete Sequence, Multichromosomal Architecture and Transcriptome Analysis of the Solanum tuberosum Mitochondrial Genome. International Journal of Molecular Sciences. 2019; 20(19):4788. https://doi.org/10.3390/ijms20194788
Chicago/Turabian StyleVarré, Jean-Stéphane, Nunzio D’Agostino, Pascal Touzet, Sophie Gallina, Rachele Tamburino, Concita Cantarella, Elodie Ubrig, Teodoro Cardi, Laurence Drouard, José Manuel Gualberto, and et al. 2019. "Complete Sequence, Multichromosomal Architecture and Transcriptome Analysis of the Solanum tuberosum Mitochondrial Genome" International Journal of Molecular Sciences 20, no. 19: 4788. https://doi.org/10.3390/ijms20194788
APA StyleVarré, J.-S., D’Agostino, N., Touzet, P., Gallina, S., Tamburino, R., Cantarella, C., Ubrig, E., Cardi, T., Drouard, L., Gualberto, J. M., & Scotti, N. (2019). Complete Sequence, Multichromosomal Architecture and Transcriptome Analysis of the Solanum tuberosum Mitochondrial Genome. International Journal of Molecular Sciences, 20(19), 4788. https://doi.org/10.3390/ijms20194788








