Migration of Small Ribosomal Subunits on the 5′ Untranslated Regions of Capped Messenger RNA
Abstract
1. Introduction
2. Results
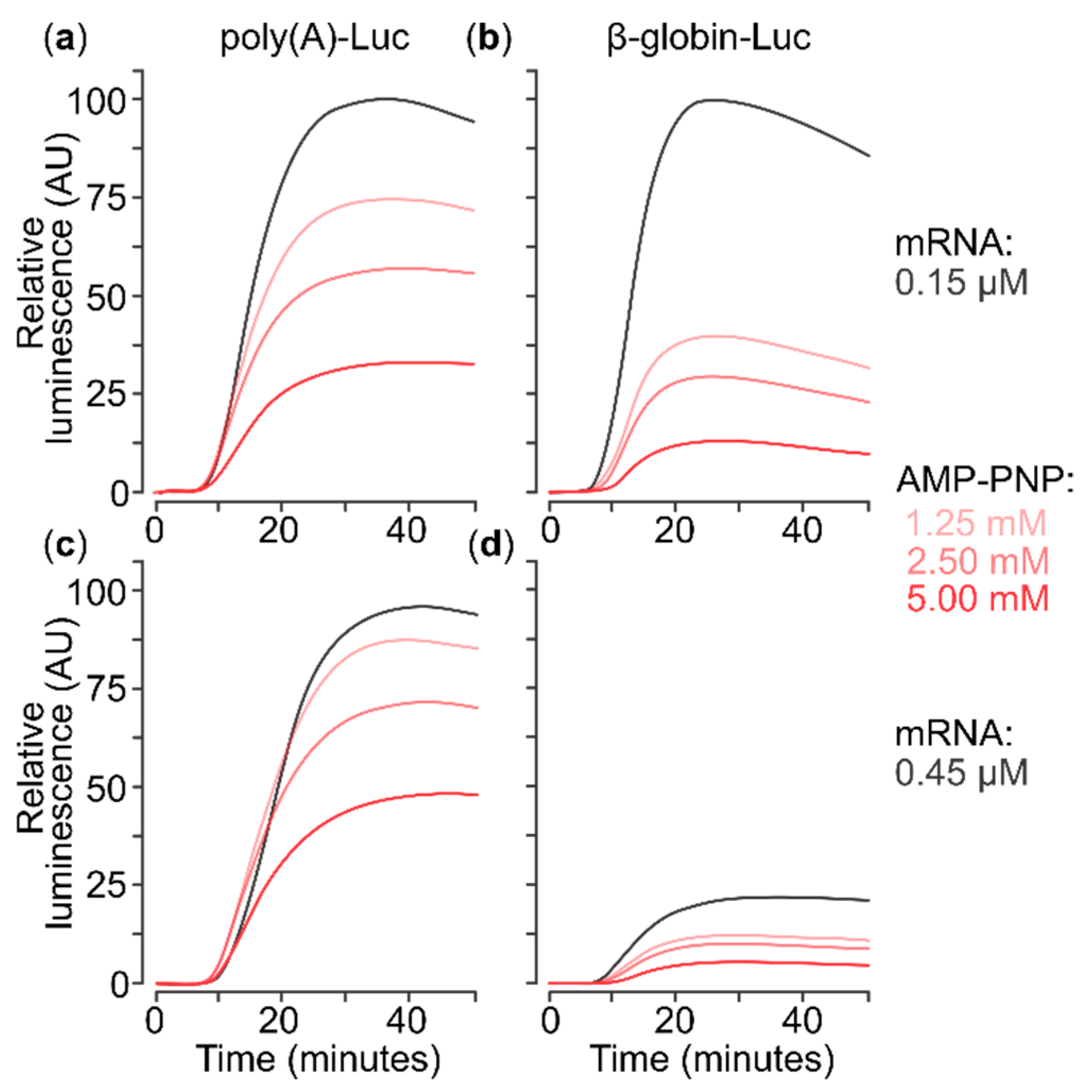
2.1. A Nonhydrolysable ATP Analogue Can Inhibit Translation of Capped Poly(A) 5′UTR mRNA
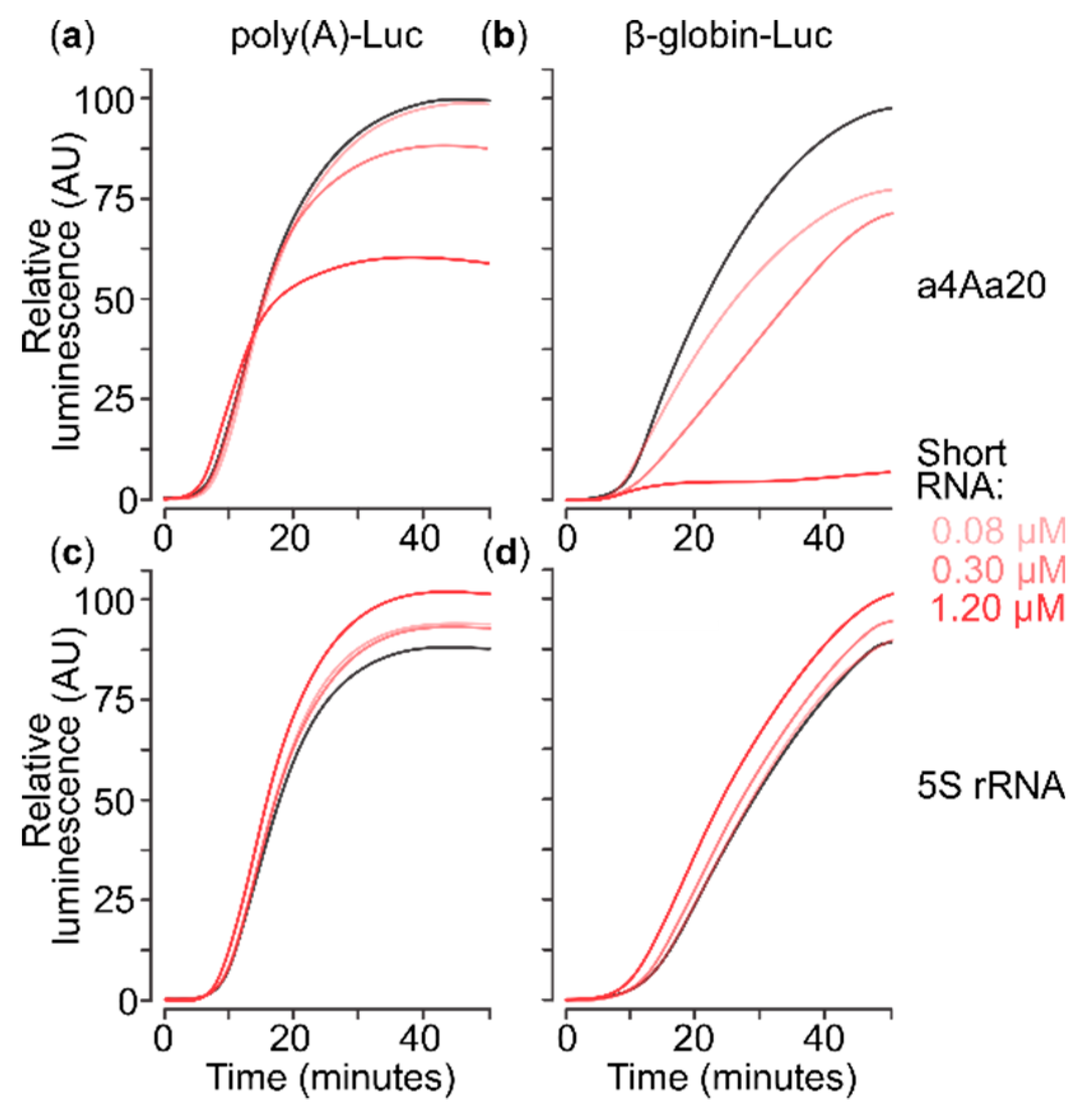
2.2. eIF4A-Blocking RNA Aptamer Does Not Affect Translation of Capped Poly(A) 5′UTR mRNA
2.3. Presence of AMP-PNP Decreases Efficiency of SSU Complex Assembly at the Start Codon of Capped Poly(A) 5′UTR mRNA
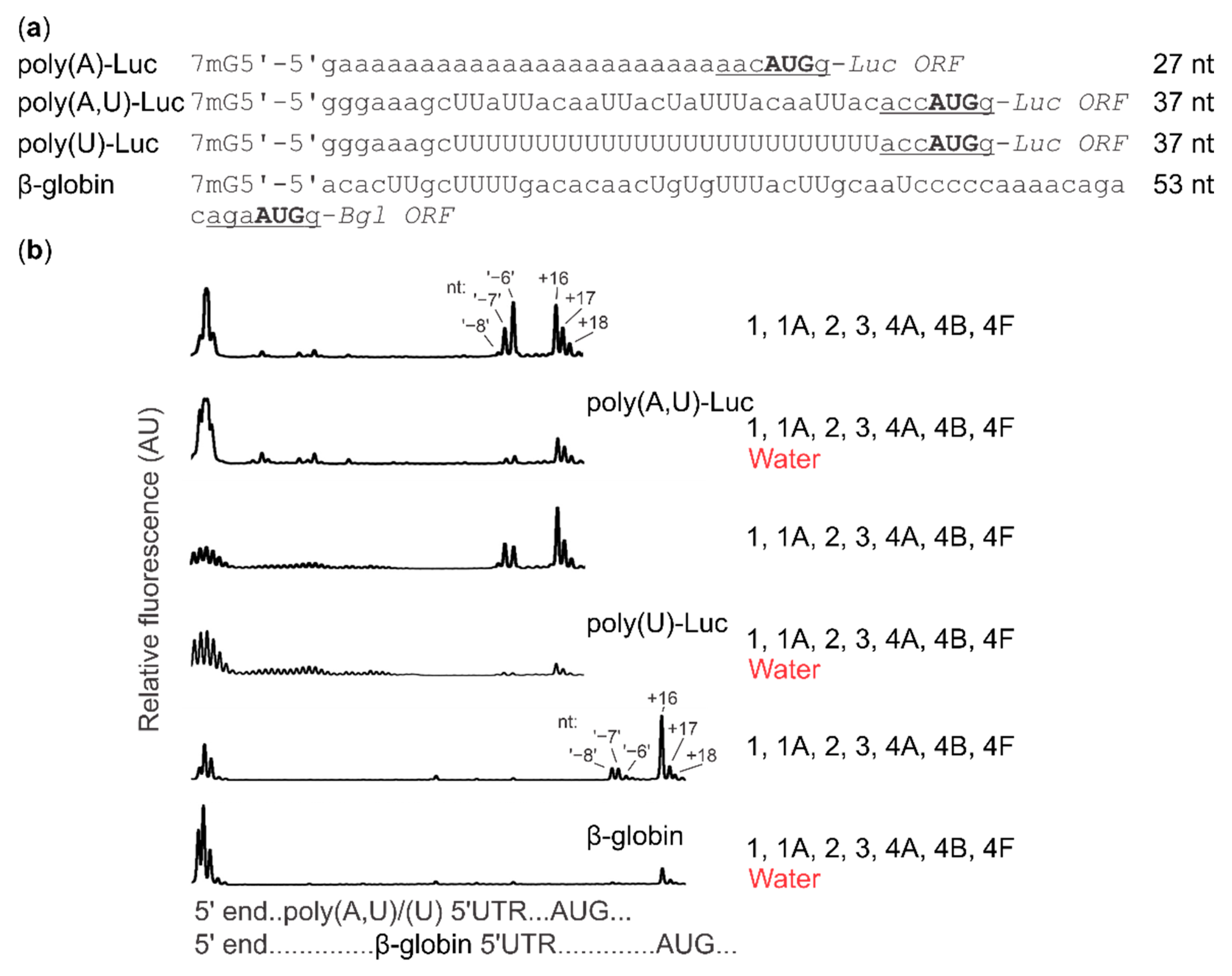
2.4. Cap-Guided Initiation Results in the Appearance of an mRNA:SSU Toeprint Upstream of the Usual Start Codon Signal
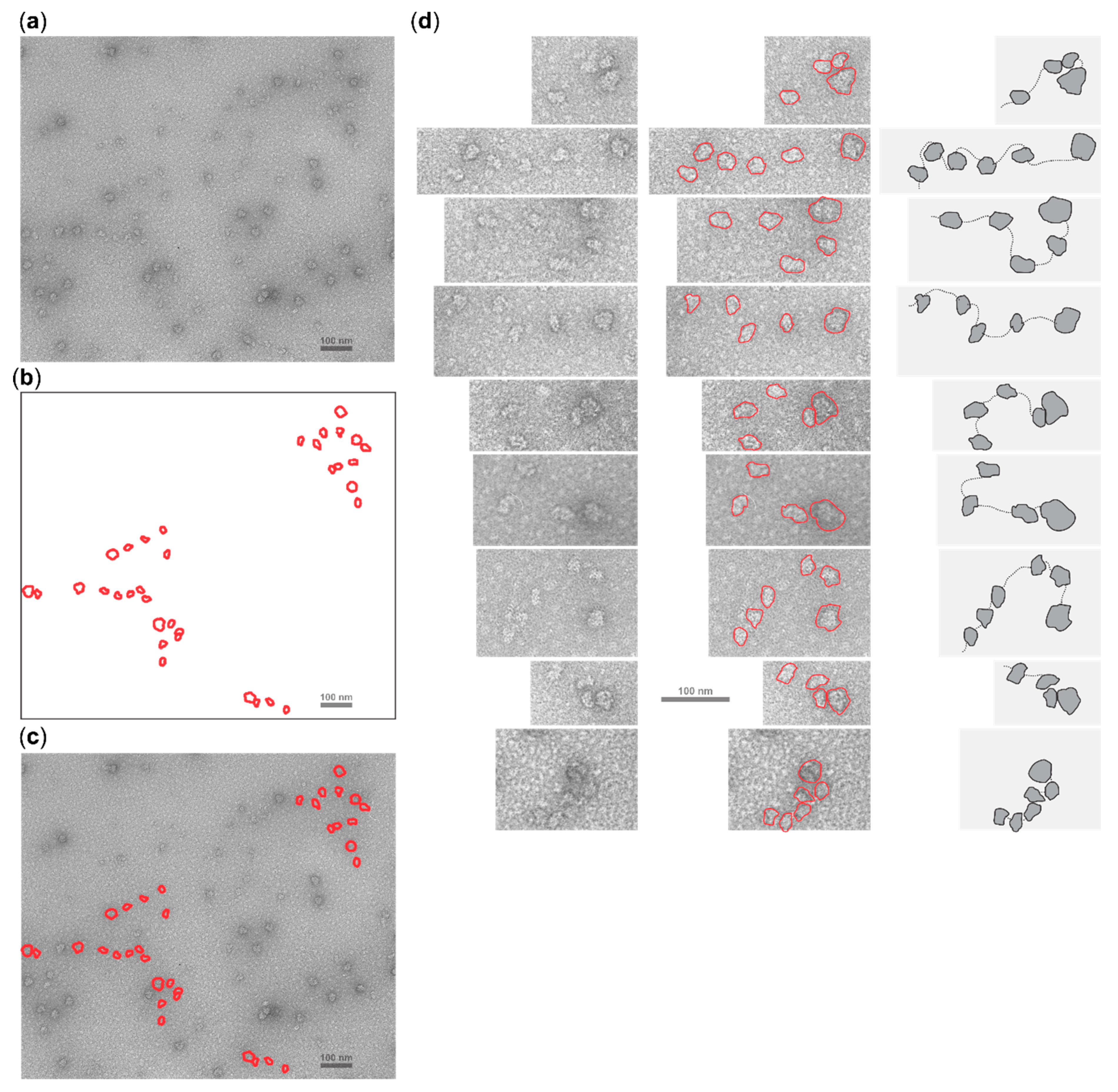
2.5. Cycloheximide Elongation Arrest Leads to the Accumulation of Multiple SSUs on the Long, Cap-and-Scanning Initiated LL1 5′UTR in the Presence of High Concentration of Magnesium Ions and Nonhydrolysable ATP Analogue AMP-PNP
3. Discussion
4. Materials and Methods
4.1. Construction of Plasmids for Run-Off Transcription of Anti-eIF4A Aptamer, Its Scrambled Control RNA and Poly(U)-Luc, Poly(A,U)-Luc mRNAs
4.2. In Vitro Synthesis of RNA
4.3. Cell-Free Translation
4.4. Toeprinting in a Reconstituted Translation System
4.5. Electron Microscopy
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| Ac | Acetate |
| AMP-PNP | Adenosine MonoPhosphate-PNP; (β,γ-imidoadenosine 5′-triphosphate) |
| AMP-PCP | Adenosine MonoPhosphate-PCP; (β,γ-methyleneadenosine 5′-triphosphate) |
| ARCA | Anti-Reverse Cap Analogue |
| ATP | Adenosine TriPhosphate |
| EDTA | EthyleneDiamineTetraacetic Acid |
| EGTA | Ethylene Glycol-bis(β-aminoethyl ether)-N,N,N′,N′-Tetraacetic Acid |
| eIF | Eukaryotic translation Initiation Factor |
| FAM | 6-carboxyfluorescein |
| fMet | N-Formylmethionyl |
| GTP | Guanosine TriPhosphate |
| HEPES | 4-(2-HydroxyEthyl)-1-PiperazineEthaneSulfonic acid |
| IRES | Interal Ribosome Entry Site |
| LINE-1 | Long Interspersed Nuclear Element 1 |
| LSU | (ribosomal) Large Subunit |
| m6A | N6-MethylAdenosine |
| Met | Methionyl |
| mRNA | Messenger RNA |
| NTP | Nucleoside TriPhosphate |
| ORF | Open Reading Frame |
| PAGE | PolyAcrylamide Gel Electrophoresis |
| RNase | RiboNuclease |
| rRNA | Ribosomal RNA |
| RT | Reverse Transcriptase |
| S | Svedberg unit (sedimentation coefficient) |
| SSU | (ribosomal) Small Subunit |
| TISU | Translation Initiator of Short 5′ UTR |
| Tris | Tris(hydroxymethyl) aminomethane |
| tRNA | Transfer RNA |
| UTR | Untranslated Region (of mRNA) |
References
- Kozak, M. How do eucaryotic ribosomes select initiation regions in messenger RNA? Cell 1978, 15, 1109–1123. [Google Scholar] [CrossRef]
- Kozak, M.; Shatkin, A.J. Migration of 40 S ribosomal subunits on messenger RNA in the presence of edeine. J. Biol. Chem. 1978, 253, 6568–6577. [Google Scholar] [PubMed]
- Kozak, M. Role of ATP in binding and migration of 40S ribosomal subunits. Cell 1980, 22, 459–467. [Google Scholar] [CrossRef]
- Kozak, M. Regulation of translation via mRNA structure in prokaryotes and eukaryotes. Gene 2005, 361, 13–37. [Google Scholar] [CrossRef] [PubMed]
- Pisarev, A.V.; Kolupaeva, V.G.; Pisareva, V.P.; Merrick, W.C.; Hellen, C.U.; Pestova, T.V. Specific functional interactions of nucleotides at key -3 and +4 positions flanking the initiation codon with components of the mammalian 48S translation initiation complex. Genes. Dev. 2006, 20, 624–636. [Google Scholar] [CrossRef] [PubMed]
- Lind, C.; Esguerra, M.; Aqvist, J. A close-up view of codon selection in eukaryotic initiation. RNA Biol. 2017, 14, 815–819. [Google Scholar] [CrossRef] [PubMed]
- Lacerda, R.; Menezes, J.; Romao, L. More than just scanning: The importance of cap-independent mRNA translation initiation for cellular stress response and cancer. Cell. Mol. Life Sci. 2017, 74, 1659–1680. [Google Scholar] [CrossRef]
- Walters, B.; Thompson, S.R. Cap-Independent Translational Control of Carcinogenesis. Front. Oncol. 2016, 6, 128. [Google Scholar] [CrossRef]
- Terenin, I.M.; Andreev, D.E.; Dmitriev, S.E.; Shatsky, I.N. A novel mechanism of eukaryotic translation initiation that is neither m7G-cap-, nor IRES-dependent. Nucleic Acids Res. 2013, 41, 1807–1816. [Google Scholar] [CrossRef]
- Terenin, I.M.; Smirnova, V.V.; Andreev, D.E.; Dmitriev, S.E.; Shatsky, I.N. A researcher’s guide to the galaxy of IRESs. Cell. Mol. Life Sci. 2017, 74, 1431–1455. [Google Scholar] [CrossRef]
- Yamamoto, H.; Unbehaun, A.; Spahn, C.M.T. Ribosomal Chamber Music: Toward an Understanding of IRES Mechanisms. Trends Biochem. Sci. 2017, 42, 655–668. [Google Scholar] [CrossRef] [PubMed]
- Johnson, A.G.; Grosely, R.; Petrov, A.N.; Puglisi, J.D. Dynamics of IRES-mediated translation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2017, 372. [Google Scholar] [CrossRef] [PubMed]
- Pisarev, A.V.; Shirokikh, N.E.; Hellen, C.U. Translation initiation by factor-independent binding of eukaryotic ribosomes to internal ribosomal entry sites. C. R. Biol. 2005, 328, 589–605. [Google Scholar] [CrossRef] [PubMed]
- Komar, A.A.; Hatzoglou, M. Cellular IRES-mediated translation: The war of ITAFs in pathophysiological states. Cell Cycle 2011, 10, 229–240. [Google Scholar] [CrossRef] [PubMed]
- May, J.; Johnson, P.; Saleem, H.; Simon, A.E. A Sequence-Independent, Unstructured Internal Ribosome Entry Site Is Responsible for Internal Expression of the Coat Protein of Turnip Crinkle Virus. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Shirokikh, N.E.; Spirin, A.S. Poly(A) leader of eukaryotic mRNA bypasses the dependence of translation on initiation factors. Proc. Natl. Acad. Sci. USA 2008, 105, 10738–10743. [Google Scholar] [CrossRef]
- Gilbert, W.V.; Zhou, K.; Butler, T.K.; Doudna, J.A. Cap-independent translation is required for starvation-induced differentiation in yeast. Science 2007, 317, 1224–1227. [Google Scholar] [CrossRef]
- Coots, R.A.; Liu, X.M.; Mao, Y.; Dong, L.; Zhou, J.; Wan, J.; Zhang, X.; Qian, S.B. m6A Facilitates eIF4F-Independent mRNA Translation. Mol. Cell 2017, 68, 504–514. [Google Scholar] [CrossRef]
- Meyer, K.D.; Patil, D.P.; Zhou, J.; Zinoviev, A.; Skabkin, M.A.; Elemento, O.; Pestova, T.V.; Qian, S.B.; Jaffrey, S.R. 5′ UTR m6A Promotes Cap-Independent Translation. Cell 2015, 163, 999–1010. [Google Scholar] [CrossRef] [PubMed]
- Shatsky, I.N.; Dmitriev, S.E.; Terenin, I.M.; Andreev, D.E. Cap- and IRES-independent scanning mechanism of translation initiation as an alternative to the concept of cellular IRESs. Mol. Cells 2010, 30, 285–293. [Google Scholar] [CrossRef]
- Akulich, K.A.; Andreev, D.E.; Terenin, I.M.; Smirnova, V.V.; Anisimova, A.S.; Makeeva, D.S.; Arkhipova, V.I.; Stolboushkina, E.A.; Garber, M.B.; Prokofjeva, M.M.; et al. Four translation initiation pathways employed by the leaderless mRNA in eukaryotes. Sci. Rep. 2016, 6, 37905. [Google Scholar] [CrossRef] [PubMed]
- Martin, F.; Menetret, J.F.; Simonetti, A.; Myasnikov, A.G.; Vicens, Q.; Prongidi-Fix, L.; Natchiar, S.K.; Klaholz, B.P.; Eriani, G. Ribosomal 18S rRNA base pairs with mRNA during eukaryotic translation initiation. Nat. Commun. 2016, 7, 12622. [Google Scholar] [CrossRef] [PubMed]
- Pelletier, J.; Sonenberg, N. The Organizing Principles of Eukaryotic Ribosome Recruitment. Annu. Rev. Biochem. 2019, 88, 307–335. [Google Scholar] [CrossRef] [PubMed]
- Shirokikh, N.E.; Preiss, T. Translation initiation by cap-dependent ribosome recruitment: Recent insights and open questions. Wiley Interdiscip. Rev. RNA 2018, 9, e1473. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, A.G. Structural Insights into the Mechanism of Scanning and Start Codon Recognition in Eukaryotic Translation Initiation. Trends Biochem. Sci. 2017, 42, 589–611. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, A.G.; Ivanov, I.P.; Sonenberg, N. Translational control by 5′-untranslated regions of eukaryotic mRNAs. Science 2016, 352, 1413–1416. [Google Scholar] [CrossRef] [PubMed]
- Haimov, O.; Sinvani, H.; Dikstein, R. Cap-dependent, scanning-free translation initiation mechanisms. Biochim. Biophys. Acta 2015, 1849, 1313–1318. [Google Scholar] [CrossRef] [PubMed]
- Elfakess, R.; Sinvani, H.; Haimov, O.; Svitkin, Y.; Sonenberg, N.; Dikstein, R. Unique translation initiation of mRNAs-containing TISU element. Nucleic Acids Res. 2011, 39, 7598–7609. [Google Scholar] [CrossRef]
- Archer, S.K.; Shirokikh, N.E.; Beilharz, T.H.; Preiss, T. Dynamics of ribosome scanning and recycling revealed by translation complex profiling. Nature 2016, 535, 570–574. [Google Scholar] [CrossRef]
- Soto-Rifo, R.; Rubilar, P.S.; Limousin, T.; de Breyne, S.; Decimo, D.; Ohlmann, T. DEAD-box protein DDX3 associates with eIF4F to promote translation of selected mRNAs. EMBO J. 2012, 31, 3745–3756. [Google Scholar] [CrossRef]
- Gupta, N.; Lorsch, J.R.; Hinnebusch, A.G. Yeast Ded1 promotes 48S translation pre-initiation complex assembly in an mRNA-specific and eIF4F-dependent manner. eLife 2018, 7, e38892. [Google Scholar] [CrossRef] [PubMed]
- Berthelot, K.; Muldoon, M.; Rajkowitsch, L.; Hughes, J.; McCarthy, J.E. Dynamics and processivity of 40S ribosome scanning on mRNA in yeast. Mol. Microbiol. 2004, 51, 987–1001. [Google Scholar] [CrossRef] [PubMed]
- Robichaud, N.; Sonenberg, N. Translational control and the cancer cell response to stress. Curr. Opin. Cell Biol. 2017, 45, 102–109. [Google Scholar] [CrossRef] [PubMed]
- Costello, J.L.; Kershaw, C.J.; Castelli, L.M.; Talavera, D.; Rowe, W.; Sims, P.F.G.; Ashe, M.P.; Grant, C.M.; Hubbard, S.J.; Pavitt, G.D. Dynamic changes in eIF4F-mRNA interactions revealed by global analyses of environmental stress responses. Genome Biol. 2017, 18, 201. [Google Scholar] [CrossRef] [PubMed]
- Starck, S.R.; Tsai, J.C.; Chen, K.; Shodiya, M.; Wang, L.; Yahiro, K.; Martins-Green, M.; Shastri, N.; Walter, P. Translation from the 5′ untranslated region shapes the integrated stress response. Science 2016, 351, aad3867. [Google Scholar] [CrossRef]
- Castelli, L.M.; Lui, J.; Campbell, S.G.; Rowe, W.; Zeef, L.A.; Holmes, L.E.; Hoyle, N.P.; Bone, J.; Selley, J.N.; Sims, P.F.; et al. Glucose depletion inhibits translation initiation via eIF4A loss and subsequent 48S preinitiation complex accumulation, while the pentose phosphate pathway is coordinately up-regulated. Mol. Biol. Cell 2011, 22, 3379–3393. [Google Scholar] [CrossRef] [PubMed]
- Janapala, Y.; Preiss, T.; Shirokikh, N.E. Control of Translation at the Initiation Phase During Glucose Starvation in Yeast. Int. J. Mol. Sci. 2019, 20, 4043. [Google Scholar] [CrossRef]
- Robichaud, N.; del Rincon, S.V.; Huor, B.; Alain, T.; Petruccelli, L.A.; Hearnden, J.; Goncalves, C.; Grotegut, S.; Spruck, C.H.; Furic, L.; et al. Phosphorylation of eIF4E promotes EMT and metastasis via translational control of SNAIL and MMP-3. Oncogene 2015, 34, 2032–2042. [Google Scholar] [CrossRef]
- Xie, J.; Merrett, J.E.; Jensen, K.B.; Proud, C.G. The MAP kinase-interacting kinases (MNKs) as targets in oncology. Expert. Opin. Targets 2019, 23, 187–199. [Google Scholar] [CrossRef]
- Steinberger, J.; Chu, J.; Maiga, R.I.; Sleiman, K.; Pelletier, J. Developing anti-neoplastic biotherapeutics against eIF4F. Cell. Mol. Life Sci. 2017, 74, 1681–1692. [Google Scholar] [CrossRef]
- Masvidal, L.; Hulea, L.; Furic, L.; Topisirovic, I.; Larsson, O. mTOR-sensitive translation: Cleared fog reveals more trees. RNA Biol. 2017, 14, 1299–1305. [Google Scholar] [CrossRef] [PubMed]
- Pestova, T.V.; Kolupaeva, V.G. The roles of individual eukaryotic translation initiation factors in ribosomal scanning and initiation codon selection. Genes Dev. 2002, 16, 2906–2922. [Google Scholar] [CrossRef] [PubMed]
- Jan, E.; Thompson, S.R.; Wilson, J.E.; Pestova, T.V.; Hellen, C.U.; Sarnow, P. Initiator Met-tRNA-independent translation mediated by an internal ribosome entry site element in cricket paralysis virus-like insect viruses. Cold Spring Harb. Symp. Quant. Biol. 2001, 66, 285–292. [Google Scholar] [CrossRef] [PubMed]
- Pestova, T.V.; Borukhov, S.I.; Hellen, C.U. Eukaryotic ribosomes require initiation factors 1 and 1A to locate initiation codons. Nature 1998, 394, 854–859. [Google Scholar] [CrossRef] [PubMed]
- Dhote, V.; Sweeney, T.R.; Kim, N.; Hellen, C.U.; Pestova, T.V. Roles of individual domains in the function of DHX29, an essential factor required for translation of structured mammalian mRNAs. Proc. Natl. Acad. Sci. USA 2012, 109, E3150–E3159. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; You, J.; Wang, X.; Weber, J. The DHX33 RNA Helicase Promotes mRNA Translation Initiation. Mol. Cell. Biol. 2015, 35, 2918–2931. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Parsyan, A.; Svitkin, Y.; Shahbazian, D.; Gkogkas, C.; Lasko, P.; Merrick, W.C.; Sonenberg, N. mRNA helicases: The tacticians of translational control. Nat. Rev. Mol. Cell Biol. 2011, 12, 235–245. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Hellen, C.U.; Pestova, T.V. Toward the mechanism of eIF4F-mediated ribosomal attachment to mammalian capped mRNAs. Genes Dev. 2016, 30, 1573–1588. [Google Scholar] [CrossRef] [PubMed]
- Skabkin, M.A.; Skabkina, O.V.; Hellen, C.U.; Pestova, T.V. Reinitiation and other unconventional posttermination events during eukaryotic translation. Mol. Cell 2013, 51, 249–264. [Google Scholar] [CrossRef]
- Skabkin, M.A.; Skabkina, O.V.; Dhote, V.; Komar, A.A.; Hellen, C.U.; Pestova, T.V. Activities of Ligatin and MCT-1/DENR in eukaryotic translation initiation and ribosomal recycling. Genes Dev. 2010, 24, 1787–1801. [Google Scholar] [CrossRef]
- Terenin, I.M.; Akulich, K.A.; Andreev, D.E.; Polyanskaya, S.A.; Shatsky, I.N.; Dmitriev, S.E. Sliding of a 43S ribosomal complex from the recognized AUG codon triggered by a delay in eIF2-bound GTP hydrolysis. Nucleic Acids Res. 2016, 44, 1882–1893. [Google Scholar] [CrossRef] [PubMed]
- Vassilenko, K.S.; Alekhina, O.M.; Dmitriev, S.E.; Shatsky, I.N.; Spirin, A.S. Unidirectional constant rate motion of the ribosomal scanning particle during eukaryotic translation initiation. Nucleic Acids Res. 2011, 39, 5555–5567. [Google Scholar] [CrossRef] [PubMed]
- Yourik, P.; Aitken, C.E.; Zhou, F.; Gupta, N.; Hinnebusch, A.G.; Lorsch, J.R. Yeast eIF4A enhances recruitment of mRNAs regardless of their structural complexity. eLife 2017, 6, e31476. [Google Scholar] [CrossRef] [PubMed]
- Sen, N.D.; Zhou, F.; Harris, M.S.; Ingolia, N.T.; Hinnebusch, A.G. eIF4B stimulates translation of long mRNAs with structured 5′ UTRs and low closed-loop potential but weak dependence on eIF4G. Proc. Natl. Acad. Sci. USA 2016, 113, 10464–10472. [Google Scholar] [CrossRef] [PubMed]
- Merrick, W.C. eIF4F: A retrospective. J. Biol. Chem. 2015, 290, 24091–24099. [Google Scholar] [CrossRef] [PubMed]
- Marintchev, A. Roles of helicases in translation initiation: A mechanistic view. Biochim. Biophys. Acta 2013, 1829, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Atas, E.; Lindqvist, L.; Sonenberg, N.; Pelletier, J.; Meller, A. The eukaryotic initiation factor eIF4H facilitates loop-binding, repetitive RNA unwinding by the eIF4A DEAD-box helicase. Nucleic Acids Res. 2012, 40, 6199–6207. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.H.; Behrens, M.A.; He, Y.; Oliveira, C.L.; Jensen, L.S.; Hoffmann, S.V.; Pedersen, J.S.; Andersen, G.R. Synergistic activation of eIF4A by eIF4B and eIF4G. Nucleic Acids Res. 2011, 39, 2678–2689. [Google Scholar] [CrossRef]
- Andreou, A.Z.; Harms, U.; Klostermeier, D. Single-stranded regions modulate conformational dynamics and ATPase activity of eIF4A to optimize 5′-UTR unwinding. Nucleic Acids Res. 2019, 47, 5260–5275. [Google Scholar] [CrossRef]
- Andreou, A.Z.; Klostermeier, D. eIF4B and eIF4G jointly stimulate eIF4A ATPase and unwinding activities by modulation of the eIF4A conformational cycle. J. Mol. Biol. 2014, 426, 51–61. [Google Scholar] [CrossRef]
- Andreou, A.Z.; Harms, U.; Klostermeier, D. eIF4B stimulates eIF4A ATPase and unwinding activities by direct interaction through its 7-repeats region. RNA Biol. 2017, 14, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Garcia, C.; Frieda, K.L.; Feoktistova, K.; Fraser, C.S.; Block, S.M. RNA BIOCHEMISTRY. Factor-dependent processivity in human eIF4A DEAD-box helicase. Science 2015, 348, 1486–1488. [Google Scholar] [CrossRef] [PubMed]
- Park, E.H.; Walker, S.E.; Zhou, F.; Lee, J.M.; Rajagopal, V.; Lorsch, J.R.; Hinnebusch, A.G. Yeast eukaryotic initiation factor 4B (eIF4B) enhances complex assembly between eIF4A and eIF4G in vivo. J. Biol. Chem. 2013, 288, 2340–2354. [Google Scholar] [CrossRef] [PubMed]
- Ozes, A.R.; Feoktistova, K.; Avanzino, B.C.; Fraser, C.S. Duplex unwinding and ATPase activities of the DEAD-box helicase eIF4A are coupled by eIF4G and eIF4B. J. Mol. Biol. 2011, 412, 674–687. [Google Scholar] [CrossRef] [PubMed]
- Walker, S.E.; Zhou, F.; Mitchell, S.F.; Larson, V.S.; Valasek, L.; Hinnebusch, A.G.; Lorsch, J.R. Yeast eIF4B binds to the head of the 40S ribosomal subunit and promotes mRNA recruitment through its N-terminal and internal repeat domains. Nucleic Acids Res. 2013, 19, 191–207. [Google Scholar] [CrossRef] [PubMed]
- Sokabe, M.; Fraser, C.S. A helicase-independent activity of eIF4A in promoting mRNA recruitment to the human ribosome. Proc. Natl. Acad. Sci. USA 2017, 114, 6304–6309. [Google Scholar] [CrossRef] [PubMed]
- Spirin, A.S. How does a scanning ribosomal particle move along the 5′-untranslated region of eukaryotic mRNA? Brownian Ratchet model. Biochemistry 2009, 48, 10688–10692. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, I.P.; Shin, B.S.; Loughran, G.; Tzani, I.; Young-Baird, S.K.; Cao, C.; Atkins, J.F.; Dever, T.E. Polyamine Control of Translation Elongation Regulates Start Site Selection on Antizyme Inhibitor mRNA via Ribosome Queuing. Mol. Cell 2018, 70, 254–264. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.G.; Goldman, D.H.; Choi, J.; Nwaezeapu, C.; Liang, D.; Green, K.M.; Goldstrohm, A.C.; Todd, P.K.; Green, R.; Wilusz, J.E. Ribosome queuing enables non-AUG translation to be resistant to multiple protein synthesis inhibitors. Genes Dev. 2019, 33, 871–885. [Google Scholar] [CrossRef] [PubMed]
- Pisareva, V.P.; Pisarev, A.V.; Komar, A.A.; Hellen, C.U.; Pestova, T.V. Translation initiation on mammalian mRNAs with structured 5′UTRs requires DExH-box protein DHX29. Cell 2008, 135, 1237–1250. [Google Scholar] [CrossRef]
- Iwasaki, S.; Iwasaki, W.; Takahashi, M.; Sakamoto, A.; Watanabe, C.; Shichino, Y.; Floor, S.N.; Fujiwara, K.; Mito, M.; Dodo, K.; et al. The Translation Inhibitor Rocaglamide Targets a Bimolecular Cavity between eIF4A and Polypurine RNA. Mol. Cell 2019, 73, 738–748. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, S.; Floor, S.N.; Ingolia, N.T. Rocaglates convert DEAD-box protein eIF4A into a sequence-selective translational repressor. Nature 2016, 534, 558–561. [Google Scholar] [CrossRef] [PubMed]
- Ricci, E.P.; Limousin, T.; Soto-Rifo, R.; Rubilar, P.S.; Decimo, D.; Ohlmann, T. miRNA repression of translation in vitro takes place during 43S ribosomal scanning. Nucleic Acids Res. 2013, 41, 586–598. [Google Scholar] [CrossRef] [PubMed]
- Shirokikh, N.E.; Alkalaeva, E.Z.; Vassilenko, K.S.; Afonina, Z.A.; Alekhina, O.M.; Kisselev, L.L.; Spirin, A.S. Quantitative analysis of ribosome-mRNA complexes at different translation stages. Nucleic Acids Res. 2010, 38, e15. [Google Scholar] [CrossRef] [PubMed]
- Schnierle, B.S.; Moss, B. Vaccinia virus-mediated inhibition of host protein synthesis involves neither degradation nor underphosphorylation of components of the cap-binding eukaryotic translation initiation factor complex eIF-4F. Virology 1992, 188, 931–933. [Google Scholar] [CrossRef]
- Davison, A.J.; Moss, B. Structure of vaccinia virus late promoters. J. Mol. Biol. 1989, 210, 771–784. [Google Scholar] [CrossRef]
- Ahn, B.Y.; Moss, B. Capped poly(A) leaders of variable lengths at the 5′ ends of vaccinia virus late mRNAs. J. Virol. 1989, 63, 226–232. [Google Scholar] [PubMed]
- Patel, D.D.; Pickup, D.J. Messenger RNAs of a strongly-expressed late gene of cowpox virus contain 5′-terminal poly(A) sequences. EMBO J. 1987, 6, 3787–3794. [Google Scholar] [CrossRef] [PubMed]
- Bertholet, C.; Van Meir, E.; ten Heggeler-Bordier, B.; Wittek, R. Vaccinia virus produces late mRNAs by discontinuous synthesis. Cell 1987, 50, 153–162. [Google Scholar] [CrossRef]
- Schwer, B.; Stunnenberg, H.G. Vaccinia virus late transcripts generated in vitro have a poly(A) head. EMBO J. 1988, 7, 1183–1190. [Google Scholar] [CrossRef]
- Broyles, S.S. Vaccinia virus transcription. J. Gen. Virol. 2003, 84, 2293–2303. [Google Scholar] [CrossRef] [PubMed]
- Wright, C.F.; Moss, B. In vitro synthesis of vaccinia virus late mRNA containing a 5′ poly(A) leader sequence. Proc. Natl. Acad. Sci. USA 1987, 84, 8883–8887. [Google Scholar] [CrossRef] [PubMed]
- Silver, M.; McFadden, G.; Wilton, S.; Dales, S. Biogenesis of poxviruses: Role for the DNA-dependent RNA polymerase II of the host during expression of late functions. Proc. Natl. Acad. Sci. USA 1979, 76, 4122–4125. [Google Scholar] [CrossRef] [PubMed]
- Paoletti, E.; Grady, L.J. Transcriptional complexity of vaccinia virus in vivo and in vitro. J. Virol. 1977, 23, 608–615. [Google Scholar] [PubMed]
- Sebring, E.D.; Salzman, N.P. Metabolic properties of early and late vaccinia virus messenger ribonucleic acid. J. Virol. 1967, 1, 550–558. [Google Scholar] [PubMed]
- Liu, S.W.; Katsafanas, G.C.; Liu, R.; Wyatt, L.S.; Moss, B. Poxvirus decapping enzymes enhance virulence by preventing the accumulation of dsRNA and the induction of innate antiviral responses. Cell Host Microbe 2015, 17, 320–331. [Google Scholar] [CrossRef] [PubMed]
- Bablanian, R.; Goswami, S.K.; Esteban, M.; Banerjee, A.K.; Merrick, W.C. Mechanism of selective translation of vaccinia virus mRNAs: Differential role of poly(A) and initiation factors in the translation of viral and cellular mRNAs. J. Virol. 1991, 65, 4449–4460. [Google Scholar]
- Vassef, A.; Ben-Hamida, F.; Dru, A.; Beaud, G. Translational control of early protein synthesis at the late stage of vaccinia virus infection. Virology 1982, 118, 45–53. [Google Scholar] [CrossRef]
- Mulder, J.; Robertson, M.E.; Seamons, R.A.; Belsham, G.J. Vaccinia virus protein synthesis has a low requirement for the intact translation initiation factor eIF4F, the cap-binding complex, within infected cells. J. Virol. 1998, 72, 8813–8819. [Google Scholar]
- Dhungel, P.; Cao, S.; Yang, Z. The 5′-poly(A) leader of poxvirus mRNA confers a translational advantage that can be achieved in cells with impaired cap-dependent translation. PLoS Pathog. 2017, 13, e1006602. [Google Scholar] [CrossRef]
- Gudkov, A.T.; Ozerova, M.V.; Shiryaev, V.M.; Spirin, A.S. 5′-poly(A) sequence as an effective leader for translation in eukaryotic cell-free systems. Biotechnol. Bioeng. 2005, 91, 468–473. [Google Scholar] [CrossRef] [PubMed]
- Oguro, A.; Ohtsu, T.; Svitkin, Y.V.; Sonenberg, N.; Nakamura, Y. RNA aptamers to initiation factor 4A helicase hinder cap-dependent translation by blocking ATP hydrolysis. RNA 2003, 9, 394–407. [Google Scholar] [CrossRef] [PubMed]
- Kozak, M. Primer extension analysis of eukaryotic ribosome-mRNA complexes. Nucleic Acids Res. 1998, 26, 4853–4859. [Google Scholar] [CrossRef] [PubMed]
- Egorova, T.; Sokolova, E.; Shuvalova, E.; Matrosova, V.; Shuvalov, A.; Alkalaeva, E. Fluorescent toeprinting to study the dynamics of ribosomal complexes. Methods 2019, 162–163, 54–59. [Google Scholar] [CrossRef] [PubMed]
- Wilson, J.E.; Chin, A. Chelation of divalent cations by ATP, studied by titration calorimetry. Anal. Biochem. 1991, 193, 16–19. [Google Scholar] [CrossRef]
- Lomakin, I.B.; Hellen, C.U.; Pestova, T.V. Physical association of eukaryotic initiation factor 4G (eIF4G) with eIF4A strongly enhances binding of eIF4G to the internal ribosomal entry site of encephalomyocarditis virus and is required for internal initiation of translation. Mol. Cell Biol. 2000, 20, 6019–6029. [Google Scholar] [CrossRef] [PubMed]
- Kolupaeva, V.G.; Pestova, T.V.; Hellen, C.U. Ribosomal binding to the internal ribosomal entry site of classical swine fever virus. RNA 2000, 6, 1791–1807. [Google Scholar] [CrossRef] [PubMed]
- Unbehaun, A.; Borukhov, S.I.; Hellen, C.U.; Pestova, T.V. Release of initiation factors from 48S complexes during ribosomal subunit joining and the link between establishment of codon-anticodon base-pairing and hydrolysis of eIF2-bound GTP. Genes Dev. 2004, 18, 3078–3093. [Google Scholar] [CrossRef]
- Fekete, C.A.; Applefield, D.J.; Blakely, S.A.; Shirokikh, N.; Pestova, T.; Lorsch, J.R.; Hinnebusch, A.G. The eIF1A C-terminal domain promotes initiation complex assembly, scanning and AUG selection in vivo. EMBO J. 2005, 24, 3588–3601. [Google Scholar] [CrossRef]
- Lomakin, I.B.; Shirokikh, N.E.; Yusupov, M.M.; Hellen, C.U.; Pestova, T.V. The fidelity of translation initiation: reciprocal activities of eIF1, IF3 and YciH. EMBO J. 2006, 25, 196–210. [Google Scholar] [CrossRef]
- Cheung, Y.N.; Maag, D.; Mitchell, S.F.; Fekete, C.A.; Algire, M.A.; Takacs, J.E.; Shirokikh, N.; Pestova, T.; Lorsch, J.R.; Hinnebusch, A.G. Dissociation of eIF1 from the 40S ribosomal subunit is a key step in start codon selection in vivo. Genes Dev. 2007, 21, 1217–1230. [Google Scholar] [CrossRef] [PubMed]
- Abaeva, I.S.; Marintchev, A.; Pisareva, V.P.; Hellen, C.U.T.; Pestova, T.V. Bypassing of stems versus linear base-by-base inspection of mammalian mRNAs during ribosomal scanning. EMBO J. 2010, 30, 115–129. [Google Scholar] [CrossRef] [PubMed]
- Pisareva, V.P.; Pisarev, A.V. eIF5 and eIF5B together stimulate 48S initiation complex formation during ribosomal scanning. Nucleic Acids Res. 2014, 42, 12052–12069. [Google Scholar] [CrossRef] [PubMed]
- Pisareva, V.P.; Pisarev, A.V. DHX29 reduces leaky scanning through an upstream AUG codon regardless of its nucleotide context. Nucleic Acids Res. 2016, 44, 4252–4265. [Google Scholar] [CrossRef] [PubMed]
- Llacer, J.L.; Hussain, T.; Marler, L.; Aitken, C.E.; Thakur, A.; Lorsch, J.R.; Hinnebusch, A.G.; Ramakrishnan, V. Conformational Differences between Open and Closed States of the Eukaryotic Translation Initiation Complex. Mol. Cell 2015, 59, 399–412. [Google Scholar] [CrossRef] [PubMed]
- Dmitriev, S.E.; Andreev, D.E.; Ad’ianova, Z.V.; Terenin, I.M.; Shatskii, I.N. [Efficient cap-dependent in vitro and in vivo translation of mammalian mRNAs with long and highly structured 5′-untranslated regions]. Mol. Biol. (Mosk.) 2009, 43, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Dmitriev, S.E.; Andreev, D.E.; Terenin, I.M.; Olovnikov, I.A.; Prassolov, V.S.; Merrick, W.C.; Shatsky, I.N. Efficient translation initiation directed by the 900-nucleotide-long and GC-rich 5′ untranslated region of the human retrotransposon LINE-1 mRNA is strictly cap dependent rather than internal ribosome entry site mediated. Mol. Cell. Biol. 2007, 27, 4685–4697. [Google Scholar] [CrossRef]
- Khaleghpour, K.; Svitkin, Y.V.; Craig, A.W.; DeMaria, C.T.; Deo, R.C.; Burley, S.K.; Sonenberg, N. Translational repression by a novel partner of human poly(A) binding protein, Paip2. Mol. Cell 2001, 7, 205–216. [Google Scholar] [CrossRef]
- Andreev, D.E.; O’Connor, P.B.; Loughran, G.; Dmitriev, S.E.; Baranov, P.V.; Shatsky, I.N. Insights into the mechanisms of eukaryotic translation gained with ribosome profiling. Nucleic Acids Res. 2017, 45, 513–526. [Google Scholar] [CrossRef]
- Gerashchenko, M.V.; Gladyshev, V.N. Translation inhibitors cause abnormalities in ribosome profiling experiments. Nucleic Acids Res. 2014, 42, e134. [Google Scholar] [CrossRef]
- Helser, T.L.; Baan, R.A.; Dahlberg, A.E. Characterization of a 40S ribosomal subunit complex in polyribosomes of Saccharomyces cerevisiae treated with cycloheximide. Mol. Cell. Biol. 1981, 1, 51–57. [Google Scholar] [CrossRef]
- Nielsen, K.H.; Szamecz, B.; Valasek, L.; Jivotovskaya, A.; Shin, B.S.; Hinnebusch, A.G. Functions of eIF3 downstream of 48S assembly impact AUG recognition and GCN4 translational control. EMBO J. 2004, 23, 1166–1177. [Google Scholar] [CrossRef] [PubMed]
- Sogorin, E.A.; Shirokikh, N.E.; Ibragimova, A.M.; Vasiliev, V.D.; Agalarov, S.; Spirin, A.S. Leader sequences of eukaryotic mRNA can be simultaneously bound to initiating 80S ribosome and 40S ribosomal subunit. Biochemistry (Mosc.) 2012, 77, 342–345. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.; Standart, N. The awesome power of ribosome profiling. Nucleic Acids Res. 2015, 21, 652–654. [Google Scholar] [CrossRef] [PubMed]
- Shirokikh, N.E.; Archer, S.K.; Beilharz, T.H.; Powell, D.; Preiss, T. Translation complex profile sequencing to study the in vivo dynamics of mRNA-ribosome interactions during translation initiation, elongation and termination. Nat. Protoc. 2017, 12, 697–731. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Wan, J.; Liu, B.; Ma, M.; Shen, B.; Qian, S.B. Quantitative profiling of initiating ribosomes in vivo. Nat. Methods 2015, 12, 147–153. [Google Scholar] [CrossRef] [PubMed]
- Andreev, D.E.; O’Connor, P.B.; Fahey, C.; Kenny, E.M.; Terenin, I.M.; Dmitriev, S.E.; Cormican, P.; Morris, D.W.; Shatsky, I.N.; Baranov, P.V. Translation of 5′ leaders is pervasive in genes resistant to eIF2 repression. eLife 2015, 4, e03971. [Google Scholar] [CrossRef] [PubMed]
- Battiste, J.L.; Pestova, T.V.; Hellen, C.U.; Wagner, G. The eIF1A solution structure reveals a large RNA-binding surface important for scanning function. Mol. Cell 2000, 5, 109–119. [Google Scholar] [CrossRef]
- De Breyne, S.; Yu, Y.; Pestova, T.V.; Hellen, C.U. Factor requirements for translation initiation on the Simian picornavirus internal ribosomal entry site. RNA 2008, 14, 367–380. [Google Scholar] [CrossRef]
- Pestova, T.V.; de Breyne, S.; Pisarev, A.V.; Abaeva, I.S.; Hellen, C.U. eIF2-dependent and eIF2-independent modes of initiation on the CSFV IRES: A common role of domain II. EMBO J. 2008, 27, 1060–1072. [Google Scholar] [CrossRef]
- Sweeney, T.R.; Abaeva, I.S.; Pestova, T.V.; Hellen, C.U. The mechanism of translation initiation on Type 1 picornavirus IRESs. EMBO J. 2014, 33, 76–92. [Google Scholar] [CrossRef] [PubMed]
- Abaeva, I.S.; Pestova, T.V.; Hellen, C.U. Attachment of ribosomal complexes and retrograde scanning during initiation on the Halastavi arva virus IRES. Nucleic Acids Res. 2016, 44, 2362–2377. [Google Scholar] [CrossRef] [PubMed]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shirokikh, N.E.; Dutikova, Y.S.; Staroverova, M.A.; Hannan, R.D.; Preiss, T. Migration of Small Ribosomal Subunits on the 5′ Untranslated Regions of Capped Messenger RNA. Int. J. Mol. Sci. 2019, 20, 4464. https://doi.org/10.3390/ijms20184464
Shirokikh NE, Dutikova YS, Staroverova MA, Hannan RD, Preiss T. Migration of Small Ribosomal Subunits on the 5′ Untranslated Regions of Capped Messenger RNA. International Journal of Molecular Sciences. 2019; 20(18):4464. https://doi.org/10.3390/ijms20184464
Chicago/Turabian StyleShirokikh, Nikolay E., Yulia S. Dutikova, Maria A. Staroverova, Ross D. Hannan, and Thomas Preiss. 2019. "Migration of Small Ribosomal Subunits on the 5′ Untranslated Regions of Capped Messenger RNA" International Journal of Molecular Sciences 20, no. 18: 4464. https://doi.org/10.3390/ijms20184464
APA StyleShirokikh, N. E., Dutikova, Y. S., Staroverova, M. A., Hannan, R. D., & Preiss, T. (2019). Migration of Small Ribosomal Subunits on the 5′ Untranslated Regions of Capped Messenger RNA. International Journal of Molecular Sciences, 20(18), 4464. https://doi.org/10.3390/ijms20184464






