MicroRNA-15a Regulates the Differentiation of Intramuscular Preadipocytes by Targeting ACAA1, ACOX1 and SCP2 in Chickens
Abstract
1. Introduction
2. Results
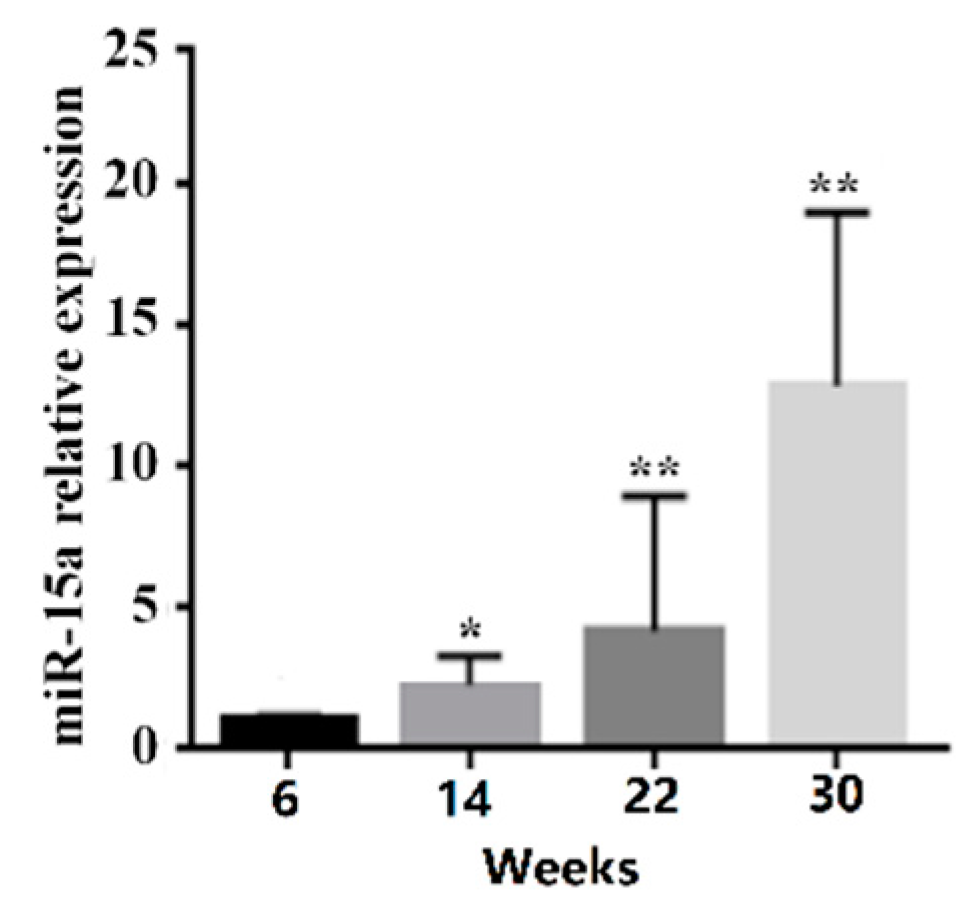
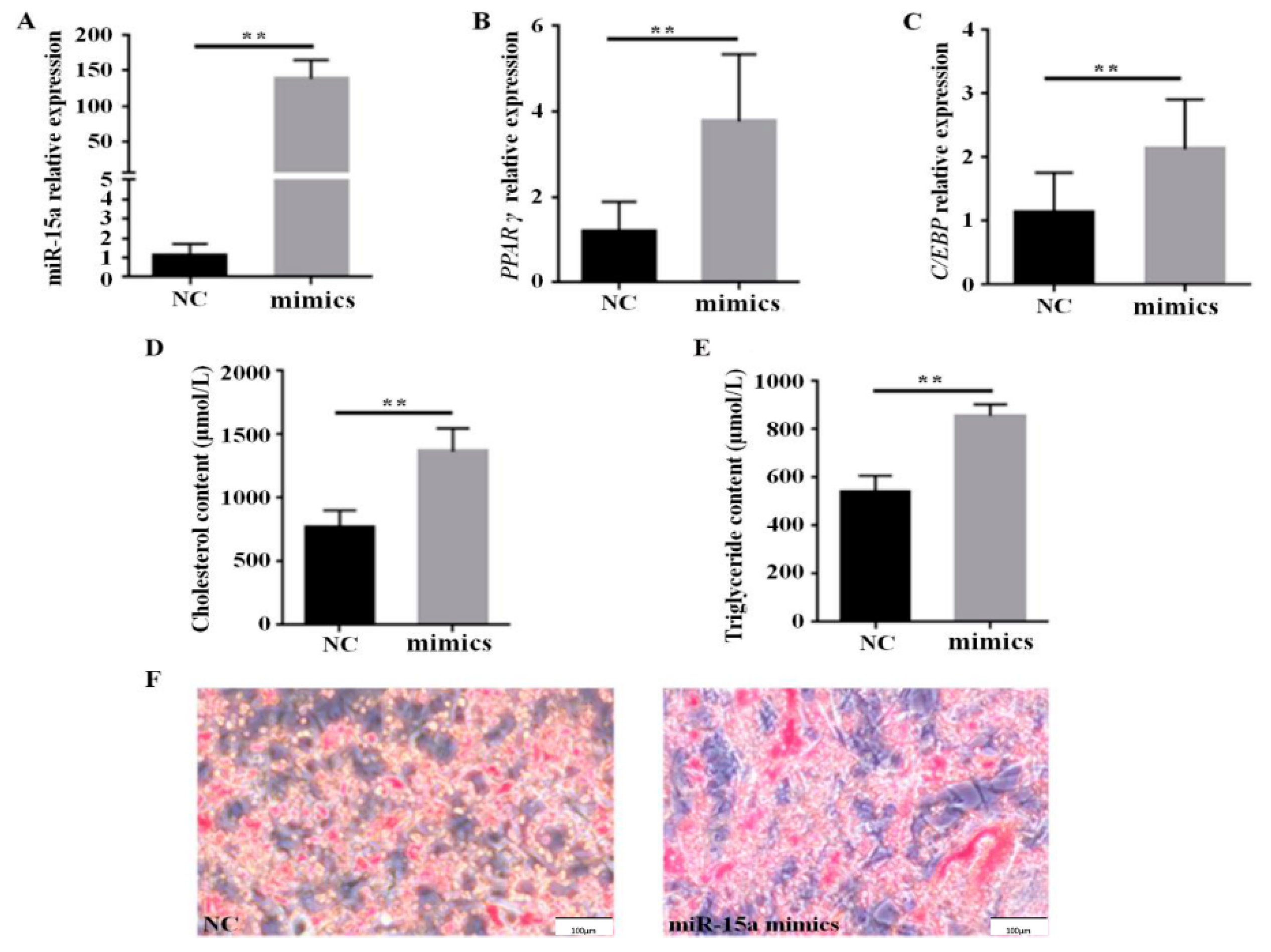
2.1. Over-Expression of Mir-15a Promotes the Differentiation of Intramuscular Preadipocytes From Chicken Breast Muscle Tissue
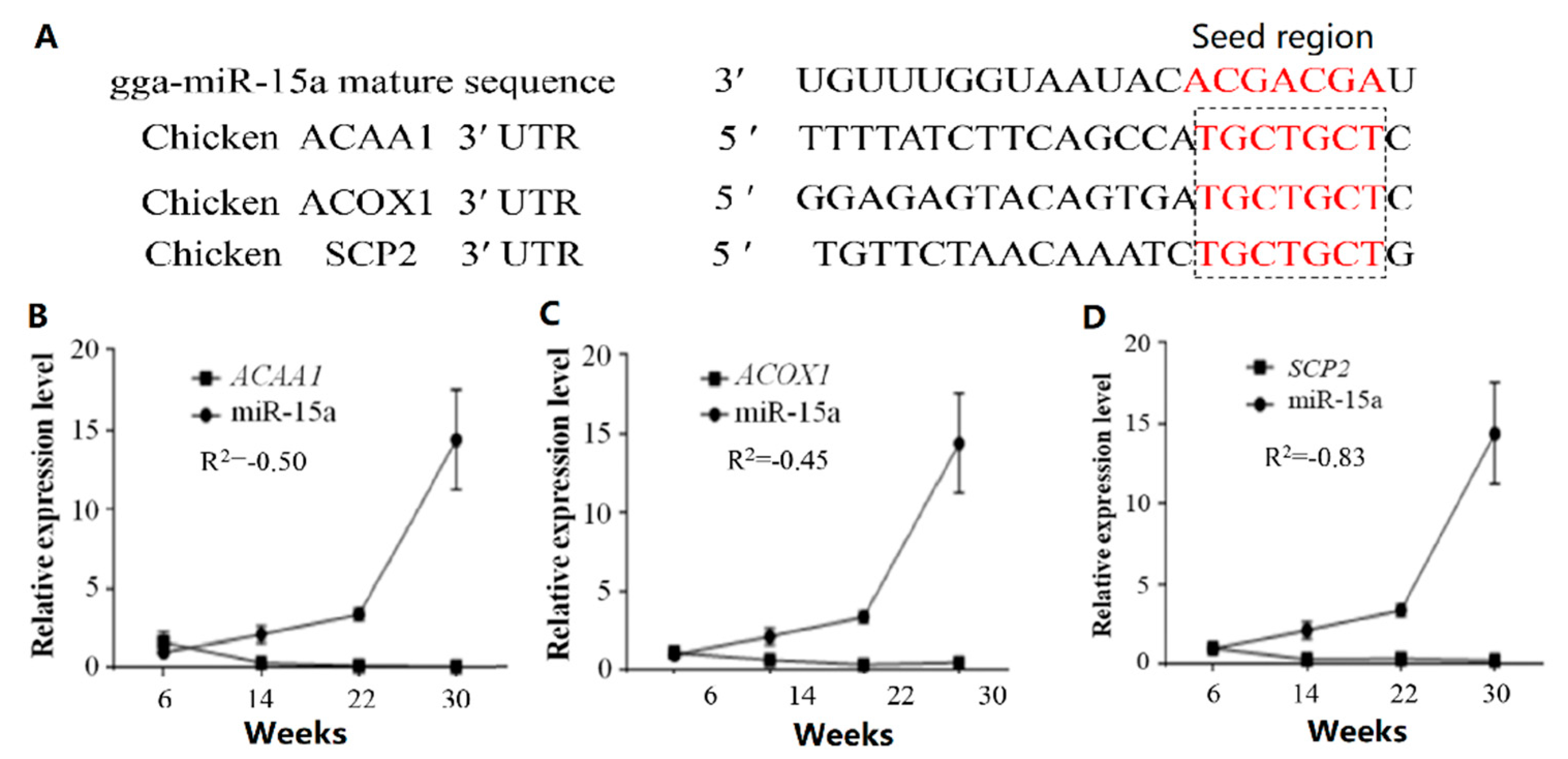
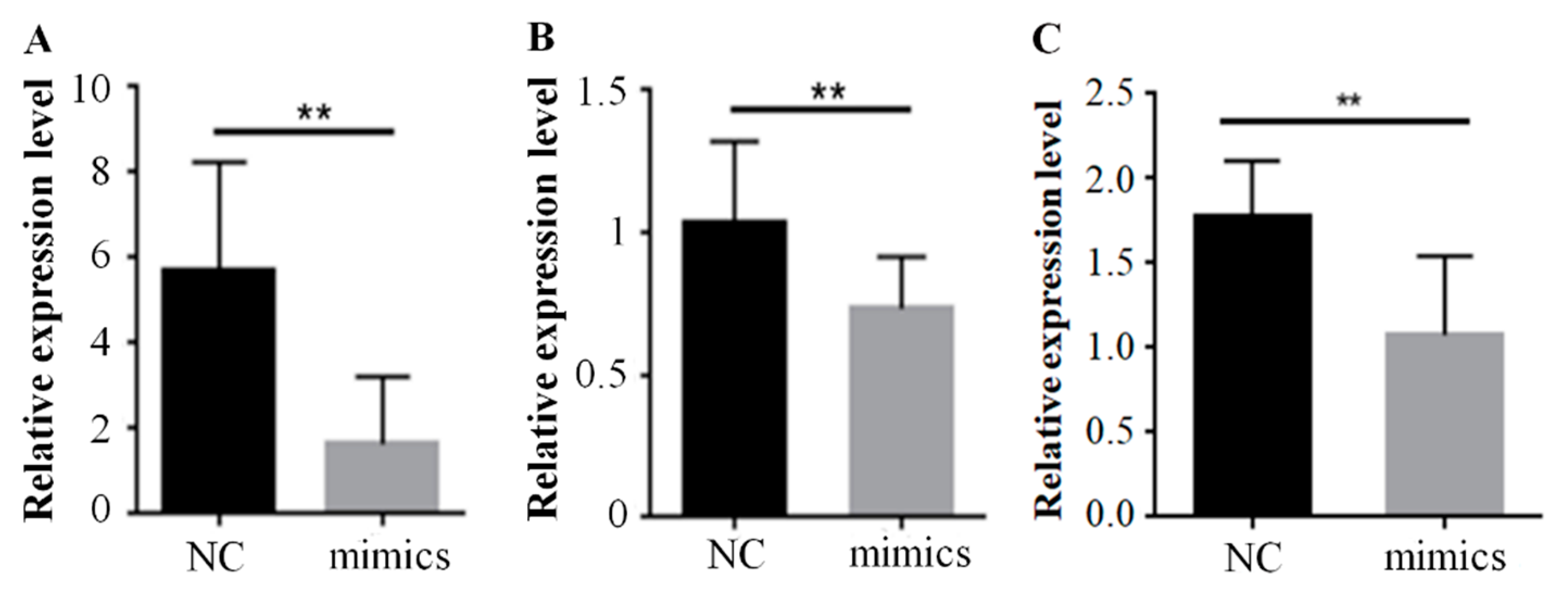
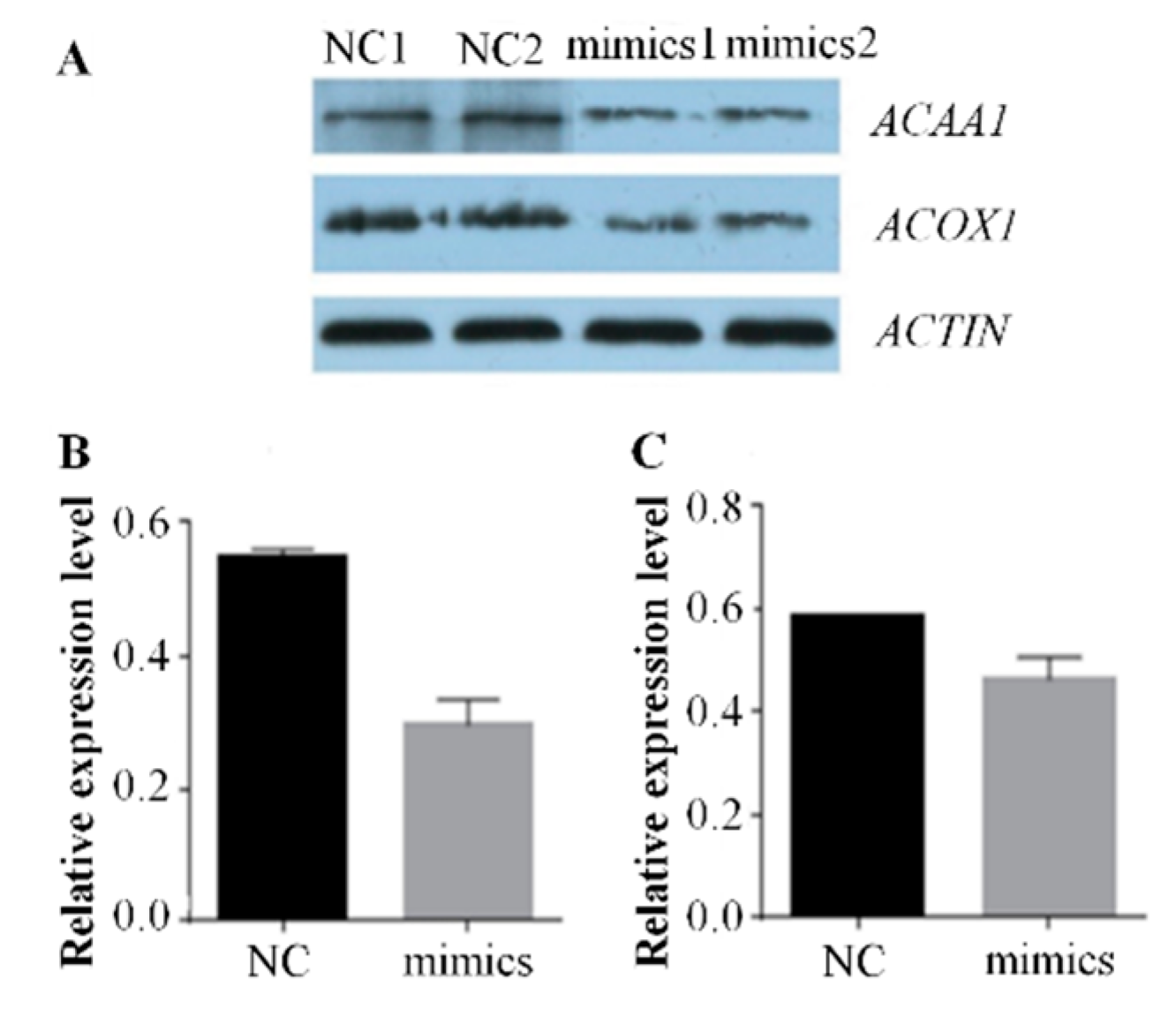
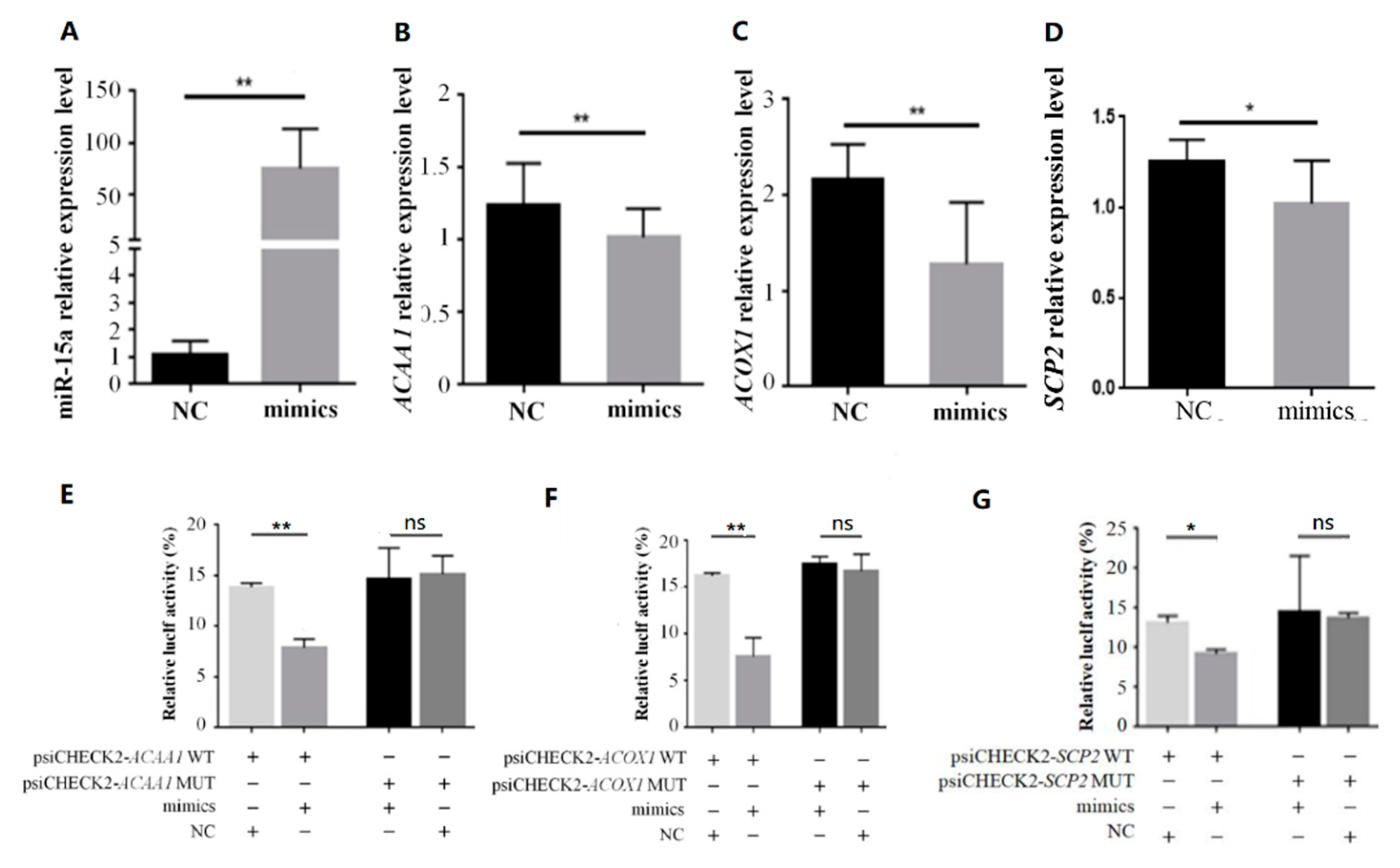
2.2. Over-Expression of Mir-15a Inhibits ACAA1, ACOX1 And SCP2 Gene Expression During Differentiation of Chicken Intramuscular Preadipocytes
2.3. MiR-15a Can Directly Target ACAA1, ACOX1 and SCP2 in Chicken Intramuscular Preadipocytes
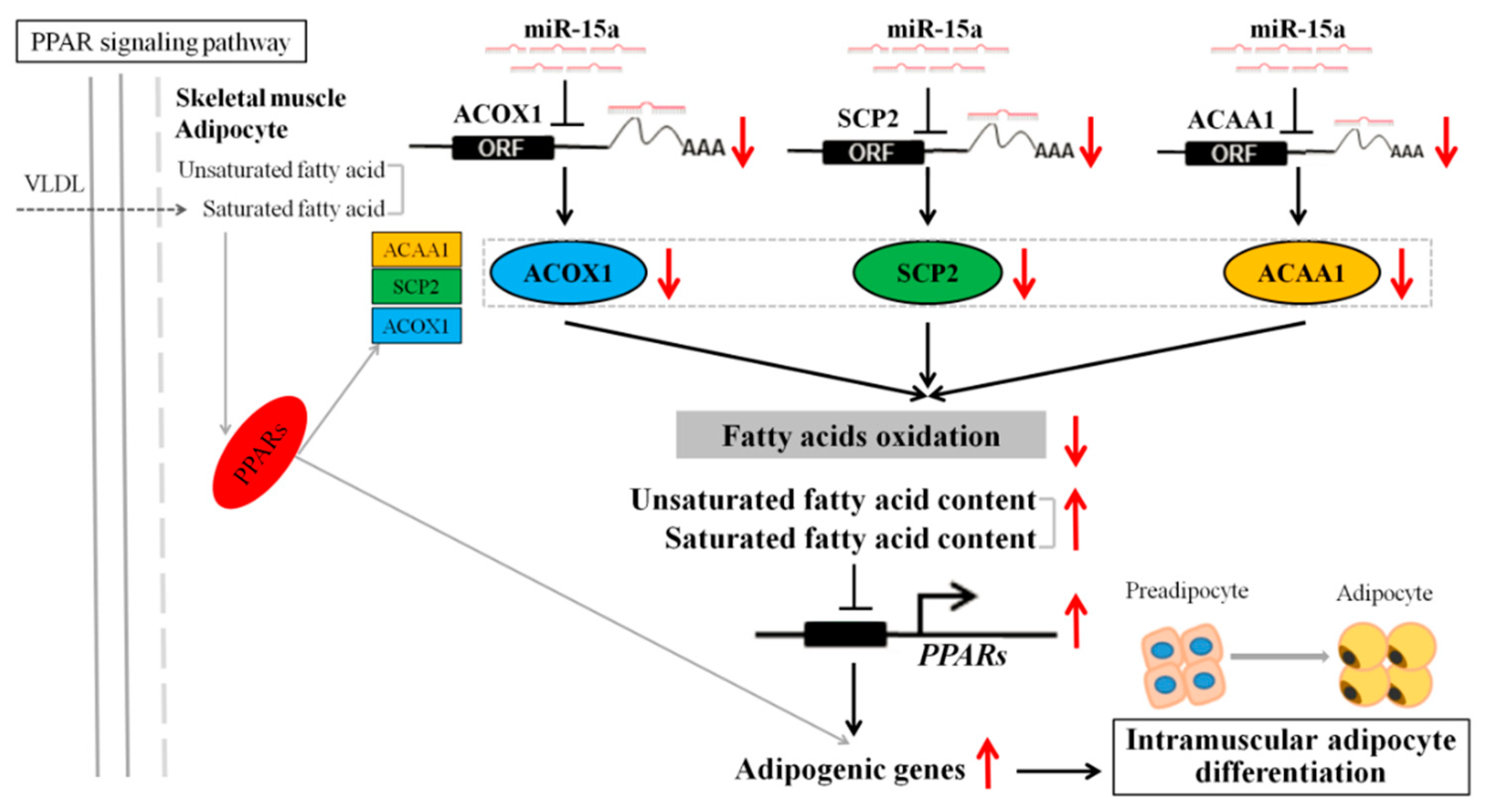
3. Discussion
4. Materials and Methods
4.1. Sample Collection
4.2. Isolation, Culture, and Differentiation Induction of Intramuscular Preadipocytes
4.3. Transfection of miR-15a Mimics
4.4. Construction of Dual-Luciferase Reporter Vector and Luciferase Activity Assay
4.5. Triglyceride and Cholesterol Assay
4.6. Oil Red O Staining
4.7. Quantitative Real-Time PCR (qPCR)
4.8. Western Blot Analysis
4.9. Statistical Analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wang, X.G.; Yu, J.F.; Zhang, Y.; Gong, D.Q.; Gu, Z.L. Identification and characterization of microRNA from chicken adipose tissue and skeletal muscle. Poult. Sci. 2012, 91, 139–149. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Deng, L.; Liang, M.; Xu, L.; Zhang, L.; Ma, Y.; Li, Y. MicroRNAs expression profiles in adipose tissues and liver from sex-linked dwarf and normal chickens. Acta Biochim. Biophys. Sin. 2014, 46, 723–726. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.Y.; Liu, R.R.; Zhao, G.P.; Li, Q.H.; Zheng, M.Q.; Zhang, J.J.; Li, S.F.; Liang, Z.; Wen, J. Integrated analysis of microRNA and mRNA expression profiles in abdominal adipose tissues in chickens. Sci. Rep. 2015, 5, 16132. [Google Scholar] [CrossRef] [PubMed]
- Yao, J.; Wang, Y.; Wang, W.; Wang, N.; Li, H. Solexa sequencing analysis of chicken pre-adipocyte microRNAs. Biosci. Biotechnol. Biochem. 2011, 75, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Du, Z.Q.; Cheng, B.; Wang, Y.; Yao, J.; Li, Y.; Cao, Z.; Luan, P.; Wang, N.; Li, H. Expression profiling of preadipocyte microRNAs by deep sequencing on chicken lines divergently selected for abdominal fatness. PLoS ONE 2015, 10, e0117843. [Google Scholar] [CrossRef]
- Wang, W.; Cheng, M.; Qiao, S.; Wang, Y.; Li, H.; Wang, N. Gga-miR-21 inhibits chicken pre-adipocyte proliferation in part by down-regulating Kruppel-like factor 5. Poult. Sci. 2017, 96, 200–210. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Li, D.H.; Li, F.; Sun, J.W.; Jiang, R.R.; Li, Z.J.; Han, R.L.; Li, G.X.; Liu, X.J.; Kang, X.T.; et al. Integrated Analysis of MiRNA and Genes Associated with Meat Quality Reveals that Gga-MiR-140-5p Affects Intramuscular Fat Deposition in Chickens. Cell. Physiol. Biochem. 2018, 46, 2421–2433. [Google Scholar] [CrossRef]
- Li, F.; Li, D.; Zhang, M.; Sun, J.; Li, W.; Jiang, R.; Han, R.; Wang, Y.; Tian, Y.; Kang, X.; et al. miRNA-223 targets the GPAM gene and regulates the differentiation of intramuscular adipocytes. Gene 2019, 685, 106–113. [Google Scholar] [CrossRef]
- Zhang, M.; Li, F.; Sun, J.W.; Li, D.H.; Li, W.T.; Jiang, R.R.; Li, Z.J.; Liu, X.J.; Han, R.L.; Li, G.X.; et al. LncRNA IMFNCR Promotes Intramuscular Adipocyte Differentiation by Sponging miR-128-3p and miR-27b-3p. Front. Genet. 2019, 10, 42. [Google Scholar] [CrossRef]
- Finnerty, J.R.; Wang, W.X.; Hébert, S.S.; Wilfred, B.R.; Mao, G.; Nelson, P.T. The miR-15/107 group of microRNA genes: Evolutionary biology, cellular functions, and roles in human diseases. J. Mol. Biol. 2010, 402, 491–509. [Google Scholar] [CrossRef]
- Bargaje, R.; Hariharan, M.; Scaria, V.; Pillai, B. Consensus miRNA expression profiles derived from interplatform normalization of microarraydata. RNA 2010, 16, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Li, H.M.; Wang, C.M.; Li, Q.Z.; Gao, X.J. MiR-15a decreases bovine mammary epithelial cell viability and lactation and regulates growth hormone receptor expression. Molecules 2012, 17, 12037–12048. [Google Scholar] [CrossRef] [PubMed]
- Ofir, M.; Hacohen, D.; Ginsberg, D. MiR-15 and miR-16 are direct transcriptional targets of E2F1 that limit E2F-induced proliferation by targeting cyclin E. Mol. Cancer Res. 2011, 9, 440–447. [Google Scholar] [CrossRef] [PubMed]
- Lindner, S.E.; Lohmüller, M.; Kotkamp, B.; Schuler, F.; Knust, Z.; Villunger, A.; Herzog, S. The miR-15 family reinforces the transition from proliferation to differentiation in pre-B cells. EMBO Rep. 2017, 18, 1604–1617. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, R.P.; Leong, J.W.; Schneider, S.E.; Ireland, A.R.; Berrien-Elliott, M.M.; Singh, A.; Schappe, T.; Jewell, B.A.; Sexl, V.; Fehniger, T.A. MicroRNA-15/16 Antagonizes Myb To Control NK Cell Maturation. J. Immunol. 2015, 195, 2806–2817. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef]
- Pekarsky, Y.; Balatti, V.; Croce, C.M. BCL2 and miR-15/16: From gene discovery to treatment. Cell Death Differ. 2018, 25, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Aqeilan, R.I.; Calin, G.A.; Croce, C.M. miR-15a and miR-16-1 in cancer: Discovery, function and future perspectives. Cell Death Differ. 2010, 17, 215–220. [Google Scholar] [CrossRef]
- Andersen, D.C.; Jensen, C.H.; Schneider, M.; Nossent, A.Y.; Eskildsen, T.; Hansen, J.L.; Teisner, B.; Sheikh, S.P. MicroRNA-15a fine-tunes the level of Delta-like 1 homolog (DLK1) in proliferating 3T3-L1preadipocytes. Exp. Cell Res. 2010, 316, 1681–1691. [Google Scholar] [CrossRef]
- Dong, P.; Mai, Y.; Zhang, Z.; Mi, L.; Wu, G.; Chu, G.; Yang, G.; Sun, S. MiR-15a/b promote adipogenesis in porcine pre-adipocyte via repressing FoxO1. Acta Biochim. Biophys. Sin. 2014, 46, 565–571. [Google Scholar] [CrossRef]
- Chen, Z.; Qiu, H.; Ma, L.; Luo, J.; Sun, S.; Kang, K.; Gou, D.; Loor, J.J. miR-30e-5p and miR-15a Synergistically Regulate Fatty Acid Metabolism in Goat Mammary Epithelial Cells via LRP6 and YAP1. Int. J. Mol. Sci. 2016, 17, 1909. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, X.; Shi, J.; Cao, P.; Wan, M.; Zhang, Q.; Wang, Y.; Kridel, S.J.; Liu, W.; Xu, J.; et al. Fatty acid synthase is a primary target of MiR-15a and MiR-16-1 in breast cancer. Oncotarget 2016, 7, 78566–78576. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.L.; Jiang, B.G.; Li, W.T.; Zou, J.J.; Shi, Y.Q.; Liu, Z.M. MicroRNA-15a positively regulates insulin synthesis by inhibiting uncoupling protein-2 expression. Diabetes Res. Clin. Pract. 2011, 91, 94–100. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.; Zhao, Y.; Li, Y.; Li, G.; Chen, Y.; Li, Z.; Sun, G.; Li, H.; Kang, X.; Yan, F. Characterization of miRNA transcriptome profiles related to breast muscle development and intramuscular fat deposition in chickens. J. Cell. Biochem. 2018, 119, 7063–7079. [Google Scholar] [CrossRef] [PubMed]
- Hocquette, J.F.; Gondret, F.; Baéza, E.; Médale, F.; Jurie, C.; Pethick, D.W. Intramuscular fat content in meat-producing animals: Development, genetic and nutritional control, and identification of putative markers. Animal 2010, 4, 303–319. [Google Scholar] [CrossRef] [PubMed]
- Erson-Bensan, A.E. Introduction to microRNAs in biological systems. Methods Mol Biol. 2014, 1107, 1–14. [Google Scholar]
- Hao, R.; Hu, X.; Wu, C.; Li, N. Hypoxia-induced miR-15a promotes mesenchymal ablation and adaptation to hypoxia during lung development in chicken. PLoS ONE 2014, 9, e98868. [Google Scholar] [CrossRef]
- Yuan, J.; Chen, S.; Shi, F.; Wu, G.; Liu, A.; Yang, N.; Sun, C. Genome-wide association study reveals putative role of gga-miR-15a in controlling feed conversion ratio in layer chickens. BMC Genom. 2017, 18, 699. [Google Scholar] [CrossRef]
- Cheng, Z.; White, M.F. Targeting Forkhead box O1 from the concept to metabolic diseases: Lessons from mouse models. Antioxid. Redox Signal. 2011, 14, 649–661. [Google Scholar] [CrossRef]
- Fournier, B.; Saudubray, J.M.; Benichou, B.; Lyonnet, S.; Munnich, A.; Clevers, H.; Poll-The, B.T. Large deletion of the peroxisomal acyl-CoA oxidase gene in pseudoneonatal adrenoleukodystrophy. J. Clin. Investig. 1994, 94, 526–531. [Google Scholar] [CrossRef]
- Li, Y.; Tharappel, J.C.; Cooper, S.; Glenn, M.; Glauert, H.P.; Spear, B.T. Expression of the hydrogen peroxide-generating enzyme fatty acyl CoA oxidase activates NF-kappaB. DNA Cell Biol. 2000, 19, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Morais, S.; Knoll-Gellida, A.; André, M.; Barthe, C.; Babin, P.J. Conserved expression of alternative splicing variants of peroxisomal acyl-CoA oxidase 1 in vertebrates and developmental and nutritional regulation in fish. Physiol. Genom. 2007, 28, 239–252. [Google Scholar] [CrossRef] [PubMed]
- Varanasi, U.; Chu, R.; Chu, S.; Espinosa, R.; LeBeau, M.M.; Reddy, J.K. Isolation of the human peroxisomal acyl-CoA oxidase gene: Organization, promoter analysis, and chromosomal localization. Proc. Natl. Acad. Sci. USA 1994, 91, 3107–3111. [Google Scholar] [CrossRef] [PubMed]
- Baes, M.; Van Veldhoven, P.P. Mouse models for peroxisome biogenesis defects and β-oxidation enzyme deficiencies. Biochim. Biophys. Acta 2012, 1822, 1489–1500. [Google Scholar] [CrossRef] [PubMed]
- Bout, A.; Franse, M.M.; Collins, J.; Blonden, L.; Tager, J.M.; Benne, R. Characterization of the gene encoding human peroxisomal 3-oxoacyl-CoA thiolase (ACAA). No large DNA rearrangement in a thiolase-deficient patient. Biochim. Biophys. Acta 1991, 1090, 43–51. [Google Scholar] [CrossRef]
- Antonenkov, V.D.; Van Veldhoven, P.P.; Waelkens, E.; Mannaerts, G.P. Substrate specificities of 3-oxoacyl-CoA thiolase A and sterol carrier protein 2/3-oxoacyl-CoA thiolase purified from normal rat liver peroxisomes. Sterol carrier protein 2/3-oxoacyl-CoA thiolase is involved in the metabolism of 2-methyl-branched fatty acids and bile acid intermediates. J. Biol. Chem. 1997, 272, 26023–26031. [Google Scholar]
- Atshaves, B.P.; Storey, S.M.; Schroeder, F. Sterol carrier protein-2/sterol carrier protein-x expression differentially alters fatty acid metabolism in L cell fibroblasts. J. Lipid Res. 2003, 44, 1751–1762. [Google Scholar] [CrossRef]
- Wouters, F.S.; Bastiaens, P.I.; Wirtz, K.W.; Jovin, T.M. FRET microscopy demonstrates molecular association of non-specific lipid transfer protein (nsL-TP) with fatty acid oxidation enzymes in peroxisomes. EMBO J. 1998, 17, 7179–7189. [Google Scholar] [CrossRef]
- Lake, B.G. Mechanisms of hepatocarcinogenicity of peroxisome-proliferating drugs and chemicals. Annu. Rev. Pharmacol. Toxicol. 1995, 35, 483–507. [Google Scholar] [CrossRef]
- Bai, L.; Pang, W.J.; Yang, Y.J.; Yang, G.S. Modulation of Sirt1 by resveratrol and nicotinamide alters proliferation and differentiation of pig preadipocytes. Mol. Cell. Biochem. 2008, 307, 129–140. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Chapman, J.R.; Waldenström, J. With Reference to Reference Genes: A Systematic Review of Endogenous Controls in Gene Expression Studies. PLoS ONE 2015, 10, e0141853. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multipleinternal control genes. Genome Biol. 2002, 3, RESEARCH0034. [Google Scholar] [CrossRef] [PubMed]
| Primer Name | Primer Sequences (5′–3′) |
|---|---|
| ACAA1-WT | F: CCCTCGAGGCAGCTTGGCAAATGTCTTA |
| R: ATTTGCGGCCGCATTCAGGCATCCCAACAGTC | |
| ACAA1-ovelap | F: AGCCACTCAGACTTCCAGTGGGG |
| R: CCCCACTGGAAGTCTGAGTGGCT | |
| ACOX1-WT | F: CCCTCGAG CTTTCACTGCCCTGCAGAAG |
| R: ATTTGCGGCCGCAACAGTTAAAAGGGCAGAAAATC | |
| ACOX1-ovelap | F: GTACAGTGACTGAGGACA |
| R: TGTCCTCAGTCACTGTAC | |
| SCP2-WT | F: CCCTCGAGCCTCAGACAGCTCCTTGCTC |
| R: ATTTGCGGCCGC TCTGGAAAAGTGGTGGGTTC | |
| SCP2-ovelap | F: AATGTTGTCACCGGTATTG |
| R: CAATACCGGTGACAACATT |
| Primer Name | Primer Sequence (5′–3′) | Location | Product Length (bp) | Annealing Temperature (°C) | Accession Number |
|---|---|---|---|---|---|
| ACAA1 | F: CCAGCATACTGACAGCCCAA | 1316-1335 | 170 | 59 | NM_001197288.1 |
| R: TCCCACTTGCACATCAGACC | 1466-1485 | ||||
| ACOX1 | F: TTAATGACCCTGACTTCCAGC | 193-233 | 162 | 58 | NM_001006205.1 |
| R: CACGATGAACAAAGCTTTTAAACCA | 330-354 | ||||
| SCP2 | F: AGGAGGCAACCTGGGTAGT | 1416-1434 | 159 | 59 | NM_001305200.1 |
| R: ATTTGCCTTGAAAGAAGGCTGTC | 1552-1574 | ||||
| PPAR γ | F: CTCCTTCTCCTCCCTATTT | 245-263 | 227 | 60 | NM_001001460.1 |
| R: TTTCTTATGGATGCGACA | 454-471 | ||||
| C/EBPα | F: GACAAGAACAGCAACGAGTACCGC | 503-536 | 195 | 56 | NM_001031459.1 |
| R: CCTGAAGATGCCCCGCAGAGT | 677-697 | ||||
| β-actin | F: GATATTGCTGCGCTCGTTG | 78-96 | 453 | 60 | NM_205518.1 |
| R: GTCCATCACAATACCAGTGG | 511-530 | ||||
| GADPH | F: TGATGGTCCACATGGCATCC | 1031-1050 | 141 | 60 | NM_204305.1 |
| R: GGGAACAGAACTGGCCTCTC | 1152-1171 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, G.; Fu, S.; Chen, Y.; Jin, W.; Zhai, B.; Li, Y.; Sun, G.; Han, R.; Wang, Y.; Tian, Y.; et al. MicroRNA-15a Regulates the Differentiation of Intramuscular Preadipocytes by Targeting ACAA1, ACOX1 and SCP2 in Chickens. Int. J. Mol. Sci. 2019, 20, 4063. https://doi.org/10.3390/ijms20164063
Li G, Fu S, Chen Y, Jin W, Zhai B, Li Y, Sun G, Han R, Wang Y, Tian Y, et al. MicroRNA-15a Regulates the Differentiation of Intramuscular Preadipocytes by Targeting ACAA1, ACOX1 and SCP2 in Chickens. International Journal of Molecular Sciences. 2019; 20(16):4063. https://doi.org/10.3390/ijms20164063
Chicago/Turabian StyleLi, Guoxi, Shouyi Fu, Yi Chen, Wenjiao Jin, Bin Zhai, Yuanfang Li, Guirong Sun, Ruili Han, Yanbin Wang, Yadong Tian, and et al. 2019. "MicroRNA-15a Regulates the Differentiation of Intramuscular Preadipocytes by Targeting ACAA1, ACOX1 and SCP2 in Chickens" International Journal of Molecular Sciences 20, no. 16: 4063. https://doi.org/10.3390/ijms20164063
APA StyleLi, G., Fu, S., Chen, Y., Jin, W., Zhai, B., Li, Y., Sun, G., Han, R., Wang, Y., Tian, Y., Li, H., & Kang, X. (2019). MicroRNA-15a Regulates the Differentiation of Intramuscular Preadipocytes by Targeting ACAA1, ACOX1 and SCP2 in Chickens. International Journal of Molecular Sciences, 20(16), 4063. https://doi.org/10.3390/ijms20164063





