Control of Translation at the Initiation Phase During Glucose Starvation in Yeast
Abstract
1. Introduction
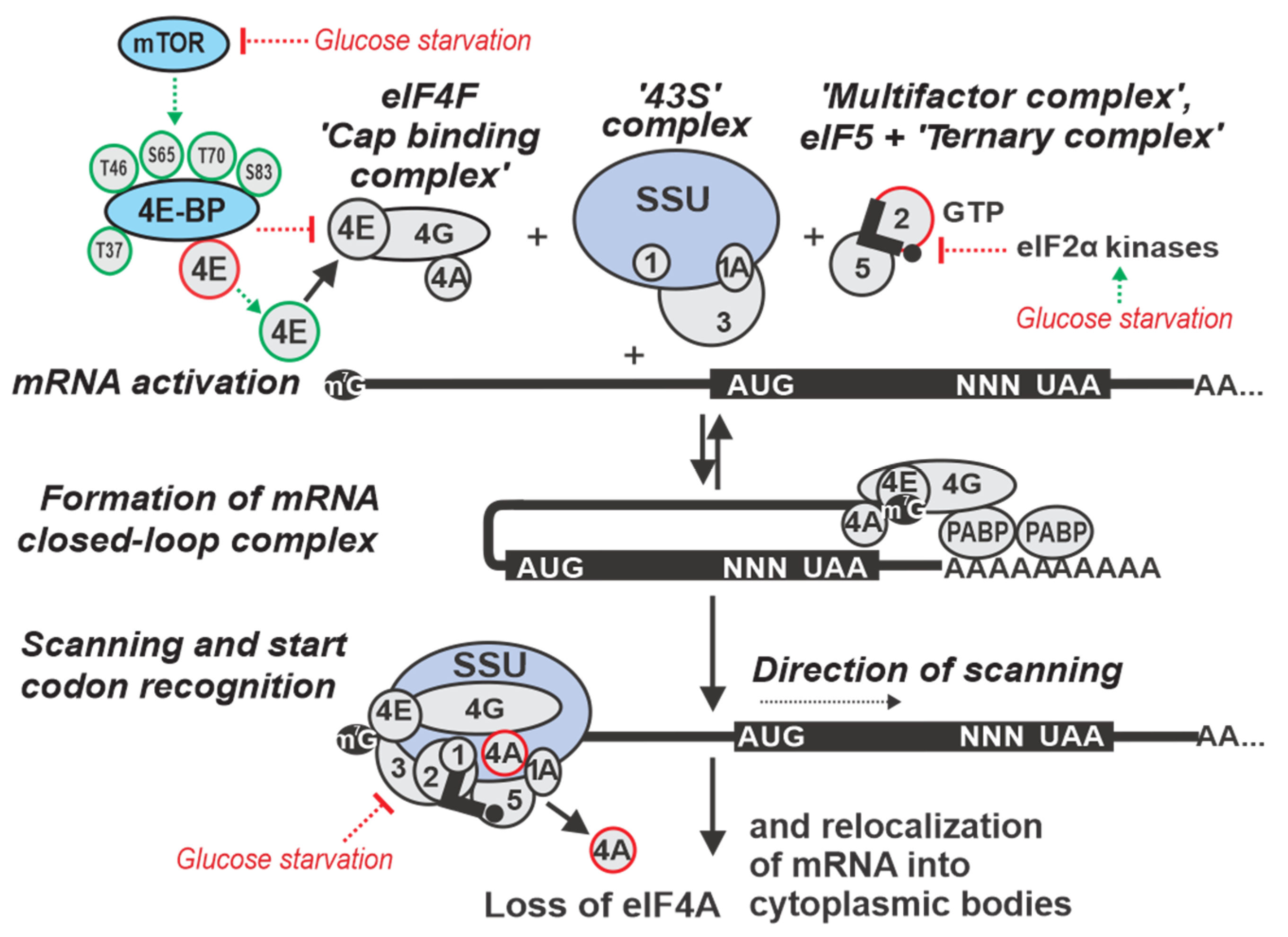
2. Overview of Eukaryotic Translation Initiation
3. Regulation by Targeting Ribosomal Attachment to mRNA
4. Regulation by Targeting the mRNA Scanning Step
5. Regulation by Targeting Start Codon Recognition
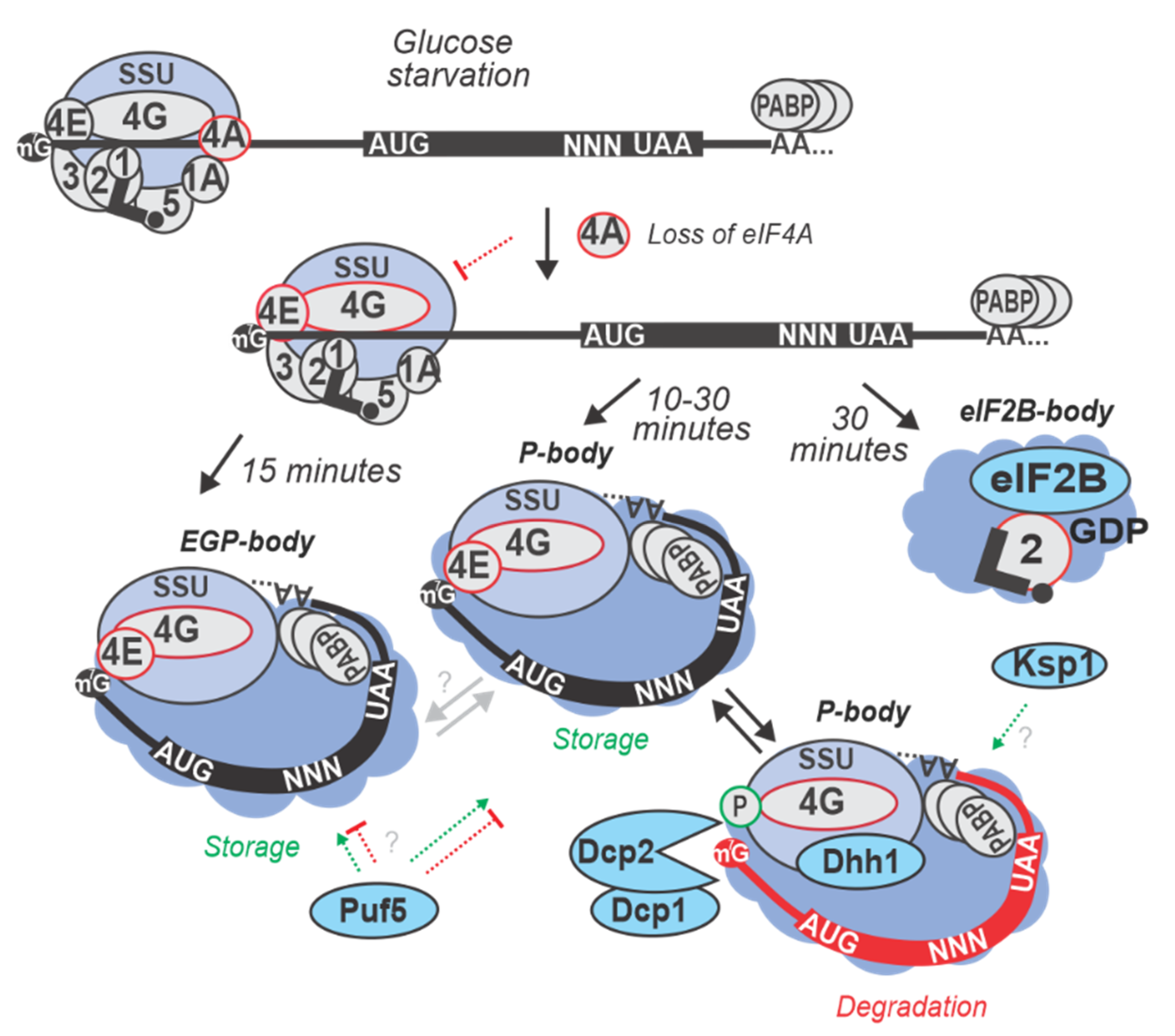
6. Formation of Cytoplasmic Foci and the Fate of Inhibited mRNA
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| 3T | Tracing Translation by T-cells |
| 4E-BP | eIF4E Binding Protein |
| acORF | Annotated main Coding ORF |
| ACT1 | Actin 1 |
| AMPK | Adenosine Monophosphate-Activated Protein Kinase |
| ATF4 | Activating Transcription Factor 4 |
| BCY1 | Bypass of CYclic-AMP requirement; catalytic subunit of PKA |
| BiP | Binding Immunoglobulin Protein |
| Caf20p | Cap-Associated Factor 20 Protein |
| CAT1 | Cationic amino acid transporter 1 |
| cCLAP | Chemical Crosslinking Coupled to Affinity Purification |
| CCR4 | Carbon Catabolite Repression Factor 4 |
| CLC | Closed- Loop Complex |
| DMEM | Dulbecco’s Modified Eagle Medium |
| Eap1p | eIF4E-Associated Protein 1 Protein |
| EGP-bodies | eIF4E/G, Pab1p-containing bodies |
| eIF | Eukaryotic translation Initiation Factor |
| ER | Endoplasmic Reticulum |
| fMet-tRNAfMet | Initiator Formyl-Methyl-tRNA |
| GCN4, GCN2 | General Control Protein 4 or 2 |
| GDP | Guanosine DiPhosphate |
| GEF | Guanine Exchange Factor |
| GST | Glutathione-S-Transferase |
| HBH | His6-Biotinylation sequence-His |
| HRI | Heme Regulated Inhibitor protein |
| HSP30 | Heat Shock Protein 30 |
| HXT2 | HeXose transmembrane Transporter 2 |
| IRES | Internal Ribosomal Entry Site |
| LC-MS | Liquid Chromatography-Mass Spectrophotometry |
| LSU | Large ribosomal SubUnit |
| MFC | MultiFactor Complex |
| mRNA | Messenger RiboNucleic Acid |
| mTOR | Mechanistic Target Of Rapamycin |
| Ngr1p | Negative Growth Regulatory 1 Protein |
| NMD | Nonsense-Mediated Decay |
| OGD | Oxygen Glucose Deprivation |
| ORF | Open Reading Frame |
| P13K | PhosphatidylInositol-3 Kinase |
| p20 | Protein 20 |
| Pab1p | Poly (A) Binding 1 Protein |
| P-bodies | Processing bodies |
| PEPCK | PhosphoEnolPyruvate CarboxyKinase |
| PERK | PKR-like Endoplasmic Reticulum Kinase |
| PGK1 | 3-PhosphoGlycerate Kinase 1 |
| PI-4,5-P2 | PhosphatidylInositol 4,5-biPhosphate |
| PI-5-P | PhosphatidylInositol 5-Phosphate |
| PIP4ks | PhosphatidylInositol 5-Phosphate 4-kinases |
| PKA | Protein Kinase A |
| PKR | Protein kinase R |
| PP | Pentose Phosphate pathway |
| PRAS40 | Proline-Rich AKT1 Substrate 40 |
| qRT-PCR | Quantitative Real-Time Polymerase Chain Reaction |
| RBPs | RNA-Binding Protein |
| RIP | RNA-ImmunoPurification |
| RocA | Rocaglamide A |
| SC | Synthetic Complete media |
| SD | Shine–Dalgarno sequence |
| siRNA | Small Interfering RNA |
| SK6 | S6 Kinase |
| SMAR1 | Scaffold/Matrix Attachment Region binding protein 1 |
| Snf1p | Sucrose Non-Fermenting 1 Protein |
| SSU | Small ribosomal SubUnit |
| TAP | Tandem Affinity Purification |
| TC | Ternary Complex |
| TCA | Tri-Carboxylic Acid |
| TOP | Tract of OligoPyrimidine |
| tRNA | Transfer RiboNucleic Acid |
| uORF | Upstream Open Reading Frame |
| UTR | UnTranslated Region |
| UV | UltraViolet (light) |
| VWMD | Vanishing White Matter Disease |
References
- Jona, G.; Choder, M.; Gileadi, O. Glucose starvation induces a drastic reduction in the rates of both transcription and degradation of mRNA in yeast. Biochim. Biophys. Acta 2000, 1491, 37–48. [Google Scholar] [CrossRef]
- Van Der Kelen, K.; Beyaert, R.; Inze, D.; De Veylder, L. Translational control of eukaryotic gene expression. Crit. Rev. Biochem. Mol. Biol. 2009, 44, 143–168. [Google Scholar] [CrossRef]
- Crawford, R.A.; Pavitt, G.D. Translational regulation in response to stress in Saccharomyces cerevisiae. Yeast 2019, 36, 5–21. [Google Scholar] [CrossRef] [PubMed]
- Simpson, C.E.; Ashe, M.P. Adaptation to stress in yeast: To translate or not? Biochem. Soc. Trans. 2012, 40, 794–799. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ferretti, M.B.; Barre, J.L.; Karbstein, K. Translational Reprogramming Provides a Blueprint for Cellular Adaptation. Cell Chem. Biol. 2018, 25, 1372–1379. [Google Scholar] [CrossRef]
- Lee, A.S. The glucose-regulated proteins: Stress induction and clinical applications. Trends Biochem. Sci. 2001, 26, 504–510. [Google Scholar] [CrossRef]
- Kafri, M.; Metzl-Raz, E.; Jona, G.; Barkai, N. The Cost of Protein Production. Cell Rep. 2016, 14, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, A.G. The scanning mechanism of eukaryotic translation initiation. Annu. Rev. Biochem. 2014, 83, 779–812. [Google Scholar] [CrossRef] [PubMed]
- Scheuner, D.; Song, B.; McEwen, E.; Liu, C.; Laybutt, R.; Gillespie, P.; Saunders, T.; Bonner-Weir, S.; Kaufman, R.J. Translational Control Is Required for the Unfolded Protein Response and In Vivo Glucose Homeostasis. Mol. Cell 2001, 7, 1165–1176. [Google Scholar] [CrossRef]
- Moon, H.W.; Han, H.G.; Jeon, Y.J. Protein Quality Control in the Endoplasmic Reticulum and Cancer. Int. J. Mol. Sci. 2018, 19. [Google Scholar] [CrossRef] [PubMed]
- Richter, K.; Haslbeck, M.; Buchner, J. The heat shock response: Life on the verge of death. Mol. Cell 2010, 40, 253–266. [Google Scholar] [CrossRef]
- Barraza, C.E.; Solari, C.A.; Marcovich, I.; Kershaw, C.; Galello, F.; Rossi, S.; Ashe, M.P.; Portela, P. The role of PKA in the translational response to heat stress in Saccharomyces cerevisiae. PLoS ONE 2017, 12, e0185416. [Google Scholar] [CrossRef]
- Manning, B.D. Adaptation to starvation: Translating a matter of life or death. Cancer Cell 2013, 23, 713–715. [Google Scholar] [CrossRef]
- Parenteau, J.; Maignon, L.; Berthoumieux, M.; Catala, M.; Gagnon, V.; Abou Elela, S. Introns are mediators of cell response to starvation. Nature 2019, 565, 612–617. [Google Scholar] [CrossRef]
- Iwama, R.; Ohsumi, Y. Analysis of autophagy activated during changes in carbon source availability in yeast cells. J. Biol. Chem. 2019, 294, 5590–5603. [Google Scholar] [CrossRef]
- Hinnebusch, A.G. Translational regulation of GCN4 and the general amino acid control of yeast. Annu. Rev. Microbiol. 2005, 59, 407–450. [Google Scholar] [CrossRef]
- Nienderberger, P.; Aebi, M.; Hutter, R. Influence of the General Control of Amino Acid Biosynthesis on Cell Growth and Cell Viability in Saccharomyces cerevisiae. Microbiology 1983, 129, 2571–2583. [Google Scholar] [CrossRef][Green Version]
- Duncan, C.D.S.; Rodriguez-Lopez, M.; Ruis, P.; Bahler, J.; Mata, J. General amino acid control in fission yeast is regulated by a nonconserved transcription factor, with functions analogous to Gcn4/Atf4. Proc. Natl. Acad. Sci. USA 2018, 115, E1829–E1838. [Google Scholar] [CrossRef]
- Bae, N.S.; Seberg, A.P.; Carroll, L.P.; Swanson, M.J. Identification of Genes in Saccharomyces cerevisiae that Are Haploinsufficient for Overcoming Amino Acid Starvation. G3 2017, 7, 1061–1084. [Google Scholar] [CrossRef]
- Secco, D.; Wang, C.; Shou, H.; Whelan, J. Phosphate homeostasis in the yeast Saccharomyces cerevisiae, the key role of the SPX domain-containing proteins. FEBS Lett. 2012, 586, 289–295. [Google Scholar] [CrossRef]
- Marzluf, G.A. Molecular Genetics of Sulphur Assimilation in Filamentous Fungi and Yeast. Annu. Rev. Microbiol. 1997, 51, 73–96. [Google Scholar] [CrossRef]
- Chen, Z.; Zhang, X.; Li, H.; Liu, H.; Xia, Y.; Xun, L. The Complete Pathway for Thiosulfate Utilization in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef]
- Miller, D.; Brandt, N.; Gresham, D. Systematic identification of factors mediating accelerated mRNA degradation in response to changes in environmental nitrogen. PLoS Genet. 2018, 14, e1007406. [Google Scholar] [CrossRef]
- Zhang, W.; Du, G.; Zhou, J.; Chen, J. Regulation of Sensing, Transportation, and Catabolism of Nitrogen Sources in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 2018, 82. [Google Scholar] [CrossRef]
- Costello, J.L.; Kershaw, C.J.; Castelli, L.M.; Talavera, D.; Rowe, W.; Sims, P.F.G.; Ashe, M.P.; Grant, C.M.; Hubbard, S.J.; Pavitt, G.D. Dynamic changes in eIF4F-mRNA interactions revealed by global analyses of environmental stress responses. Genome Biol. 2017, 18, 201. [Google Scholar] [CrossRef]
- Udom, N.; Chansongkrow, P.; Charoensawan, V.; Auesukaree, C. Coordination of the cell wall integrity and high osmolarity glycerol pathways in response to ethanol stress in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2019. [Google Scholar] [CrossRef]
- Gerbasi, V.R.; Browne, C.M.; Samir, P.; Shen, B.; Sun, M.; Hazelbaker, D.Z.; Galassie, A.C.; Frank, J.; Link, A.J. Critical Role for Saccharomyces cerevisiae Asc1p in Translational Initiation at Elevated Temperatures. Proteomics 2018, 18, e1800208. [Google Scholar] [CrossRef]
- Spriggs, K.A.; Bushell, M.; Willis, A.E. Translational regulation of gene expression during conditions of cell stress. Mol. Cell 2010, 40, 228–237. [Google Scholar] [CrossRef]
- Thomas, J.D.; Dias, L.M.; Johannes, G.J. Translational repression during chronic hypoxia is dependent on glucose levels. RNA 2008, 14, 771–781. [Google Scholar] [CrossRef][Green Version]
- Jamar, N.H.; Kritsiligkou, P.; Grant, C.M. The non-stop decay mRNA surveillance pathway is required for oxidative stress tolerance. Nucleic Acids Res. 2017, 45, 6881–6893. [Google Scholar] [CrossRef]
- Ruis, H.; Schuller, C. Stress signaling in yeast. Bioessays 1995, 17, 959–965. [Google Scholar] [CrossRef]
- Giannattasio, S.; Guaragnella, N.; Zdralevic, M.; Marra, E. Molecular mechanisms of Saccharomyces cerevisiae stress adaptation and programmed cell death in response to acetic acid. Front. Microbiol. 2013, 4, 33. [Google Scholar] [CrossRef]
- Folch-Mallol, J.L.; Garay-Arroyo, A.; Lledias, F.; Covarrubias Robles, A.A. The stress response in the yeast Saccharomyces cerevisiae. Rev. Latinoam. Microbiol. 2004, 46, 24–46. [Google Scholar]
- Walker, G.; Stewart, G. Saccharomyces cerevisiae in the Production of Fermented Beverages. Beverages 2016, 2. [Google Scholar] [CrossRef]
- Tokpohozin, S.E.; Fischer, S.; Becker, T. Selection of a new Saccharomyces yeast to enhance relevant sorghum beer aroma components, higher alcohols and esters. Food Microbiol. 2019, 83, 181–186. [Google Scholar] [CrossRef]
- Pinheiro, T.; Coelho, E.; Romani, A.; Domingues, L. Intensifying ethanol production from brewer’s spent grain waste: Use of whole slurry at high solid loadings. New Biotechnol. 2019. [Google Scholar] [CrossRef]
- Oliveira, I.; Ferreira, V. Modulating Fermentative, Varietal and Aging Aromas of Wine Using non-Saccharomyces Yeasts in a Sequential Inoculation Approach. Microorganisms 2019, 7. [Google Scholar] [CrossRef]
- Li, Z.; Song, K.; Li, H.; Ma, R.; Cui, M. Effect of mixed Saccharomyces cerevisiae Y10 and Torulaspora delbrueckii Y22 on dough fermentation for steamed bread making. Int. J. Food Microbiol. 2019, 303, 58–64. [Google Scholar] [CrossRef]
- Kayikci, O.; Nielsen, J. Glucose repression in Saccharomyces cerevisiae. FEMS Yeast Res. 2015, 15. [Google Scholar] [CrossRef]
- Ashe, M.P.; De Long, S.K.; Sachs, A.B. Glucose depletion rapidly inhibits translation initiation in yeast. Mol. Biol. Cell 2000, 11, 833–848. [Google Scholar] [CrossRef]
- Kovatchev, B.; Cobelli, C. Glucose Variability: Timing, Risk Analysis, and Relationship to Hypoglycemia in Diabetes. Diabetes Care 2016, 39, 502–510. [Google Scholar] [CrossRef]
- Castelli, L.M.; Lui, J.; Campbell, S.G.; Rowe, W.; Zeef, L.A.H.; Holmes, L.E.A.; Hoyle, N.P.; Bone, J.; Selley, J.N.; Sims, P.F.G.; et al. Glucose depletion inhibits translation initiation via eIF4A loss and subsequent 48S preinitiation complex accumulation, while the pentose phosphate pathway is coordinately up-regulated. Mol. Biol. Cell 2011, 22, 3379–3393. [Google Scholar] [CrossRef]
- Lui, J.; Campbell, S.G.; Ashe, M.P. Inhibition of translation initiation following glucose depletion in yeast facilitates a rationalization of mRNA content. Biochem. Soc. Trans. 2010, 38, 1131–1136. [Google Scholar] [CrossRef]
- Holmes, L.E.; Campbell, S.G.; De Long, S.K.; Sachs, A.B.; Ashe, M.P. Loss of translational control in yeast compromised for the major mRNA decay pathway. Mol. Cell. Biol. 2004, 24, 2998–3010. [Google Scholar] [CrossRef]
- Alberti, S. Phase separation in biology. Curr. Biol. 2017, 27, R1097–R1102. [Google Scholar] [CrossRef]
- Hyman, A.A.; Weber, C.A.; Julicher, F. Liquid-liquid phase separation in biology. Annu. Rev. Cell Dev. Biol. 2014, 30, 39–58. [Google Scholar] [CrossRef]
- Forman-Kay, J.D.; Kriwacki, R.W.; Seydoux, G. Phase Separation in Biology and Disease. J. Mol. Biol. 2018, 430, 4603–4606. [Google Scholar] [CrossRef]
- Boeynaems, S.; Alberti, S.; Fawzi, N.L.; Mittag, T.; Polymenidou, M.; Rousseau, F.; Schymkowitz, J.; Shorter, J.; Wolozin, B.; Van Den Bosch, L.; et al. Protein Phase Separation: A New Phase in Cell Biology. Trends Cell Biol. 2018, 28, 420–435. [Google Scholar] [CrossRef]
- Protter, D.S.W.; Parker, R. Principles and Properties of Stress Granules. Trends Cell Biol 2016, 26, 668–679. [Google Scholar] [CrossRef]
- Van Treeck, B.; Parker, R. Principles of Stress Granules Revealed by Imaging Approaches. Cold Spring Harb. Perspect. Biol. 2019, 11. [Google Scholar] [CrossRef]
- Moon, S.L.; Parker, R. Analysis of eIF2B bodies and their relationships with stress granules and P-bodies. Sci. Rep. 2018, 8, 12264. [Google Scholar] [CrossRef]
- Kedersha, N.; Anderson, P. Mammalian stress granules and processing bodies. Methods Enzymol. 2007, 431, 61–81. [Google Scholar] [CrossRef]
- Anderson, P.; Kedersha, N. RNA granules. J. Cell Biol. 2006, 172, 803–808. [Google Scholar] [CrossRef]
- Anderson, P.; Kedersha, N.; Ivanov, P. Stress granules, P-bodies and cancer. Biochim. Biophys. Acta 2015, 1849, 861–870. [Google Scholar] [CrossRef]
- Arribere, J.A.; Doudna, J.A.; Gilbert, W.V. Reconsidering Movement of Eukaryotic mRNAs between Polysomes and P Bodies. Mol. Cell 2011, 44, 745–758. [Google Scholar] [CrossRef]
- Zid, B.M.; O’Shea, E.K. Promoter sequences direct cytoplasmic localization and translation of mRNAs during starvation in yeast. Nature 2014, 514, 117–121. [Google Scholar] [CrossRef]
- Andreev, D.E.; O’Connor, P.B.F.; Zhdanov, A.V.; Dmitriev, R.I.; Shatsky, I.N.; Papkovsky, D.B.; Baranov, P.V. Oxygen and glucose deprivation induces widespread alterations in mRNA translation within 20 minutes. Genome Biol. 2015, 16. [Google Scholar] [CrossRef]
- Chang, Y.; Huh, W.K. Ksp1-dependent phosphorylation of eIF4G modulates post-transcriptional regulation of specific mRNAs under glucose deprivation conditions. Nucleic Acids Res. 2018, 46, 3047–3060. [Google Scholar] [CrossRef]
- Khan, D.; Katoch, A.; Das, A.; Sharathchandra, A.; Lal, R.; Roy, P.; Das, S.; Chattopadhyay, S.; Das, S. Reversible induction of translational isoforms of p53 in glucose deprivation. Cell Death Differ. 2015, 22, 1203–1218. [Google Scholar] [CrossRef]
- Yang, R.; Wek, S.A.; Wek, R.C. Glucose Limitation Induces GCN4 Translation by Activation of Gcn2 Protein Kinase. Mol. Cell. Biol. 2000, 20, 2706–2717. [Google Scholar] [CrossRef]
- Parker, R.; Sheth, U. P bodies and the control of mRNA translation and degradation. Mol. Cell 2007, 25, 635–646. [Google Scholar] [CrossRef]
- Teixeira, D.; Parker, R. Analysis of P-Body Assembly in Saccharomyces cerevisiae. Mol. Biol. Cell 2007, 18, 2274–2287. [Google Scholar] [CrossRef]
- Hilliker, A.; Gao, Z.; Jankowsky, E.; Parker, R. The DEAD-box protein Ded1 modulates translation by the formation and resolution of an eIF4F-mRNA complex. Mol. Cell 2011, 43, 962–972. [Google Scholar] [CrossRef]
- Hoyle, N.P.; Castelli, L.M.; Campbell, S.G.; Holmes, L.E.A.; Ashe, M.P. Stress-dependent relocalization of translationally primed mRNPs to cytoplasmic granules that are kinetically and spatially distinct from P-bodies. J. Cell Biol. 2007, 179, 65–74. [Google Scholar] [CrossRef]
- Mergenthaler, P.; Lindauer, U.; Dienel, G.A.; Meisel, A. Sugar for the brain: The role of glucose in physiological and pathological brain function. Trends Neurosci. 2013, 36, 587–597. [Google Scholar] [CrossRef]
- Gilbert, W.V.; Zhou, K.; Butler, T.K.; Doudna, J.A. Cap-independent translation is required for starvation-induced differentiation in yeast. Science 2007, 317, 1224–1227. [Google Scholar] [CrossRef]
- Bhat, M.; Robichaud, N.; Hulea, L.; Sonenberg, N.; Pelletier, J.; Topisirovic, I. Targeting the translation machinery in cancer. Nat. Rev. Drug Discov. 2015, 14, 261–278. [Google Scholar] [CrossRef]
- Scheper, G.C.; van der Knaap, M.S.; Proud, C.G. Translation matters: Protein synthesis defects in inherited disease. Nat. Rev. Genet. 2007, 8, 711–723. [Google Scholar] [CrossRef]
- Schuller, A.P.; Green, R. Roadblocks and resolutions in eukaryotic translation. Nat. Rev. Mol. Cell Biol. 2018, 19, 526–541. [Google Scholar] [CrossRef]
- Jackson, R.J.; Hellen, C.U.; Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 2010, 11, 113–127. [Google Scholar] [CrossRef]
- Hinnebusch, A.G. Structural Insights into the Mechanism of Scanning and Start Codon Recognition in Eukaryotic Translation Initiation. Trends Biochem. Sci. 2017, 42, 589–611. [Google Scholar] [CrossRef]
- Aylett, C.H.; Ban, N. Eukaryotic aspects of translation initiation brought into focus. Philos. Trans. R. Soc. B Biol. Sci. 2017, 372. [Google Scholar] [CrossRef]
- Voigts-Hoffmann, F.; Klinge, S.; Ban, N. Structural insights into eukaryotic ribosomes and the initiation of translation. Curr. Opin. Struct. Biol. 2012, 22, 768–777. [Google Scholar] [CrossRef]
- Dever, T.E.; Kinzy, T.G.; Pavitt, G.D. Mechanism and Regulation of Protein Synthesis. Genetics 2016, 203, 65–107. [Google Scholar] [CrossRef]
- Shirokikh, N.E.; Preiss, T. Translation initiation by cap-dependent ribosome recruitment: Recent insights and open questions. Wiley Interdiscip. Rev. RNA 2018, 9, e1473. [Google Scholar] [CrossRef]
- Browning, K.S.; Bailey-Serres, J. Mechanism of cytoplasmic mRNA translation. Arab. Book 2015, 13, e0176. [Google Scholar] [CrossRef]
- Malys, N.; McCarthy, J.E. Translation initiation: Variations in the mechanism can be anticipated. Cell. Mol. Life Sci. 2011, 68, 991–1003. [Google Scholar] [CrossRef]
- Pestova, T.V.; Kolupaeva, V.G. The roles of individual eukaryotic translation initiation factors in ribosomal scanning and initiation codon selection. Genes Dev. 2002, 16, 2906–2922. [Google Scholar] [CrossRef]
- Hinnebusch, A.G. Molecular mechanism of scanning and start codon selection in eukaryotes. Microbiol. Mol. Biol. Rev. 2011, 75, 434–467. [Google Scholar] [CrossRef]
- Schmitt, E.; Coureux, P.D.; Monestier, A.; Dubiez, E.; Mechulam, Y. Start Codon Recognition in Eukaryotic and Archaeal Translation Initiation: A Common Structural Core. Int. J. Mol. Sci. 2019, 20. [Google Scholar] [CrossRef]
- Eliseev, B.; Yeramala, L.; Leitner, A.; Karuppasamy, M.; Raimondeau, E.; Huard, K.; Alkalaeva, E.; Aebersold, R.; Schaffitzel, C. Structure of a human cap-dependent 48S translation pre-initiation complex. Nucleic Acids Res. 2018, 46, 2678–2689. [Google Scholar] [CrossRef]
- Haimov, O.; Sehrawat, U.; Tamarkin-Ben Harush, A.; Bahat, A.; Uzonyi, A.; Will, A.; Hiraishi, H.; Asano, K.; Dikstein, R. Dynamic Interaction of Eukaryotic Initiation Factor 4G1 (eIF4G1) with eIF4E and eIF1 Underlies Scanning-Dependent and -Independent Translation. Mol. Cell. Biol. 2018, 38. [Google Scholar] [CrossRef]
- Merrick, W.C.; Pavitt, G.D. Protein Synthesis Initiation in Eukaryotic Cells. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef]
- Dever, T.E.; Green, R. The elongation, termination, and recycling phases of translation in eukaryotes. Cold Spring Harb. Perspect. Biol. 2012, 4, a013706. [Google Scholar] [CrossRef]
- Dever, T.E.; Dinman, J.D.; Green, R. Translation Elongation and Recoding in Eukaryotes. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef]
- Archer, S.K.; Shirokikh, N.E.; Beilharz, T.H.; Preiss, T. Dynamics of ribosome scanning and recycling revealed by translation complex profiling. Nature 2016, 535, 570–574. [Google Scholar] [CrossRef]
- Alkalaeva, E.; Mikhailova, T. Reassigning stop codons via translation termination: How a few eukaryotes broke the dogma. Bioessays 2017, 39. [Google Scholar] [CrossRef]
- Valasek, L.S. ‘Ribozoomin’—Translation initiation from the perspective of the ribosome-bound eukaryotic initiation factors (eIFs). Curr. Protein Pept. Sci. 2012, 13, 305–330. [Google Scholar] [CrossRef]
- Hinnebusch, A.G.; Ivanov, I.P.; Sonenberg, N. Translational control by 5’-untranslated regions of eukaryotic mRNAs. Science 2016, 352, 1413–1416. [Google Scholar] [CrossRef]
- Kozak, M. The scanning model for translation: An update. J. Cell Biol. 1989, 108, 229–241. [Google Scholar] [CrossRef]
- Kozak, M. Pushing the limits of the scanning mechanism for initiation of translation. Gene 2002, 299, 1–34. [Google Scholar] [CrossRef]
- Milon, P.; Maracci, C.; Filonava, L.; Gualerzi, C.O.; Rodnina, M.V. Real-time assembly landscape of bacterial 30S translation initiation complex. Nat. Struct. Mol. Biol. 2012, 19, 609–615. [Google Scholar] [CrossRef]
- Nakagawa, S.; Niimura, Y.; Gojobori, T. Comparative genomic analysis of translation initiation mechanisms for genes lacking the Shine-Dalgarno sequence in prokaryotes. Nucleic Acids Res. 2017, 45, 3922–3931. [Google Scholar] [CrossRef]
- Senicar, L.; Accetto, T. The 5’ untranslated mRNA region base content can greatly affect translation initiation in the absence of secondary structures in Prevotella bryantii TC1-1. FEMS Microbiol. Lett. 2015, 362, 1–4. [Google Scholar] [CrossRef][Green Version]
- Ramakrishnan, V. Ribosome structure and the mechanism of translation. Cell 2002, 108, 557–572. [Google Scholar] [CrossRef]
- Abolbaghaei, A.; Silke, J.R.; Xia, X. How Changes in Anti-SD Sequences Would Affect SD Sequences in Escherichia coli and Bacillus subtilis. G3 2017, 7, 1607–1615. [Google Scholar] [CrossRef]
- Rodnina, M.V. Translation in Prokaryotes. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef]
- Shine, J.; Dalgarno, L. The 3’-terminal sequence of Escherichia coli 16S ribosomal RNA: Complementarity to nonsense triplets and ribosome binding sites. Proc. Natl. Acad. Sci. USA 1974, 71, 1342–1346. [Google Scholar] [CrossRef]
- Pisarev, A.V.; Kolupaeva, V.G.; Yusupov, M.M.; Hellen, C.U.; Pestova, T.V. Ribosomal position and contacts of mRNA in eukaryotic translation initiation complexes. EMBO J. 2008, 27, 1609–1621. [Google Scholar] [CrossRef]
- Lomakin, I.B.; Shirokikh, N.E.; Yusupov, M.M.; Hellen, C.U.; Pestova, T.V. The fidelity of translation initiation: Reciprocal activities of eIF1, IF3 and YciH. EMBO J. 2006, 25, 196–210. [Google Scholar] [CrossRef]
- Simonetti, A.; Marzi, S.; Myasnikov, A.G.; Fabbretti, A.; Yusupov, M.; Gualerzi, C.O.; Klaholz, B.P. Structure of the 30S translation initiation complex. Nature 2008, 455, 416–420. [Google Scholar] [CrossRef]
- Asano, K. Why is start codon selection so precise in eukaryotes? Translation 2014, 2, e28387. [Google Scholar] [CrossRef]
- Pelletier, J.; Sonenberg, N. The Organizing Principles of Eukaryotic Ribosome Recruitment. Annu. Rev. Biochem. 2019, 88, 307–335. [Google Scholar] [CrossRef]
- Asano, K.; Clayton, J.; Shalev, A.; Hinnebusch, A.G. A multifactor complex of eukaryotic initiation factors, eIF1, eIF2, eIF3, eIF5, and initiator tRNA(Met) is an important translation initiation intermediate in vivo. Genes Dev. 2000, 14, 2534–2546. [Google Scholar] [CrossRef]
- Lomakin, I.B.; Steitz, T.A. The initiation of mammalian protein synthesis and mRNA scanning mechanism. Nature 2013, 500, 307–311. [Google Scholar] [CrossRef]
- Nag, N.; Lin, K.Y.; Edmonds, K.A.; Yu, J.; Nadkarni, D.; Marintcheva, B.; Marintchev, A. eIF1A/eIF5B interaction network and its functions in translation initiation complex assembly and remodeling. Nucleic Acids Res. 2016, 44, 7441–7456. [Google Scholar] [CrossRef]
- Merrick, W.C. eIF4F: A retrospective. J. Biol. Chem. 2015, 290, 24091–24099. [Google Scholar] [CrossRef]
- Archer, S.K.; Shirokikh, N.E.; Hallwirth, C.V.; Beilharz, T.H.; Preiss, T. Probing the closed-loop model of mRNA translation in living cells. RNA Biol. 2015, 12, 248–254. [Google Scholar] [CrossRef]
- Preiss, T.; Hentze, M.W. Dual function of the messenger RNA cap structure in poly(A)-tail-promoted translation in yeast. Nature 1998, 392, 516–520. [Google Scholar] [CrossRef]
- Tarun, S.Z., Jr.; Sachs, A.B. Association of the yeast poly(A) tail binding protein with translation initiation factor eIF-4G. EMBO J. 1996, 15, 7168–7177. [Google Scholar] [CrossRef]
- Wells, S.E.; Hillner, P.E.; Vale, R.D.; Sachs, A.B. Circularization of mRNA by eukaryotic translation initiation factors. Mol. Cell 1998, 2, 135–140. [Google Scholar] [CrossRef]
- Kahvejian, A.; Roy, G.; Sonenberg, N. The mRNA closed-loop model: The function of PABP and PABP-interacting proteins in mRNA translation. Cold Spring Harb. Symp. Quant. Biol. 2001, 66, 293–300. [Google Scholar] [CrossRef]
- Kahvejian, A.; Svitkin, Y.V.; Sukarieh, R.; M’Boutchou, M.N.; Sonenberg, N. Mammalian poly(A)-binding protein is a eukaryotic translation initiation factor, which acts via multiple mechanisms. Genes Dev. 2005, 19, 104–113. [Google Scholar] [CrossRef]
- Vicens, Q.; Kieft, J.S.; Rissland, O.S. Revisiting the Closed-Loop Model and the Nature of mRNA 5′–3′ Communication. Mol. Cell 2018, 72, 805–812. [Google Scholar] [CrossRef]
- Borman, A.M.; Michel, Y.M.; Kean, K.M. Biochemical characterisation of cap-poly(A) synergy in rabbit reticulocyte lysates: The eIF4G-PABP interaction increases the functional affinity of eIF4E for the capped mRNA 5’-end. Nucleic Acids Res. 2000, 28, 4068–4075. [Google Scholar] [CrossRef]
- Wang, X.; Xi, W.; Toomey, S.; Chiang, Y.C.; Hasek, J.; Laue, T.M.; Denis, C.L. Stoichiometry and Change of the mRNA Closed-Loop Factors as Translating Ribosomes Transit from Initiation to Elongation. PLoS ONE 2016, 11, e0150616. [Google Scholar] [CrossRef]
- Shatsky, I.N.; Terenin, I.M.; Smirnova, V.V.; Andreev, D.E. Cap-Independent Translation: What’s in a Name? Trends Biochem. Sci. 2018, 43, 882–895. [Google Scholar] [CrossRef]
- Gilbert, W.V. Alternative ways to think about cellular internal ribosome entry. J. Biol. Chem. 2010, 285, 29033–29038. [Google Scholar] [CrossRef]
- Gritsenko, A.A.; Weingarten-Gabbay, S.; Elias-Kirma, S.; Nir, R.; de Ridder, D.; Segal, E. Sequence features of viral and human Internal Ribosome Entry Sites predictive of their activity. PLoS Comput. Biol. 2017, 13, e1005734. [Google Scholar] [CrossRef]
- Komar, A.A.; Mazumder, B.; Merrick, W.C. A new framework for understanding IRES-mediated translation. Gene 2012, 502, 75–86. [Google Scholar] [CrossRef]
- Komar, A.A.; Hatzoglou, M. Cellular IRES-mediated translation: The war of ITAFs in pathophysiological states. Cell Cycle 2011, 10, 229–240. [Google Scholar] [CrossRef]
- Johnson, A.G.; Grosely, R.; Petrov, A.N.; Puglisi, J.D. Dynamics of IRES-mediated translation. Philos. Trans. R. Soc. B Biol. Sci. 2017, 372. [Google Scholar] [CrossRef]
- Terenin, I.M.; Smirnova, V.V.; Andreev, D.E.; Dmitriev, S.E.; Shatsky, I.N. A researcher’s guide to the galaxy of IRESs. Cell. Mol. Life Sci. 2017, 74, 1431–1455. [Google Scholar] [CrossRef]
- Yamamoto, H.; Unbehaun, A.; Spahn, C.M.T. Ribosomal Chamber Music: Toward an Understanding of IRES Mechanisms. Trends Biochem. Sci. 2017, 42, 655–668. [Google Scholar] [CrossRef]
- Pisarev, A.V.; Shirokikh, N.E.; Hellen, C.U. Translation initiation by factor-independent binding of eukaryotic ribosomes to internal ribosomal entry sites. C. R. Biol. 2005, 328, 589–605. [Google Scholar] [CrossRef]
- James, C.C.; Smyth, J.W. Alternative mechanisms of translation initiation: An emerging dynamic regulator of the proteome in health and disease. Life Sci. 2018, 212, 138–144. [Google Scholar] [CrossRef]
- Lacerda, R.; Menezes, J.; Romao, L. More than just scanning: The importance of cap-independent mRNA translation initiation for cellular stress response and cancer. Cell. Mol. Life Sci. 2017, 74, 1659–1680. [Google Scholar] [CrossRef]
- Spriggs, K.A.; Stoneley, M.; Bushell, M.; Willis, A.E. Re-programming of translation following cell stress allows IRES-mediated translation to predominate. Biol. Cell 2008, 100, 27–38. [Google Scholar] [CrossRef]
- Pestova, T.V.; Kolupaeva, V.G.; Lomakin, I.B.; Pilipenko, E.V.; Shatsky, I.N.; Agol, V.I.; Hellen, C.U. Molecular mechanisms of translation initiation in eukaryotes. Proc. Natl. Acad. Sci. USA 2001, 98, 7029–7036. [Google Scholar] [CrossRef]
- Jackson, R.J. The current status of vertebrate cellular mRNA IRESs. Cold Spring Harb. Perspect. Biol. 2013, 5. [Google Scholar] [CrossRef]
- de la Parra, C.; Ernlund, A.; Alard, A.; Ruggles, K.; Ueberheide, B.; Schneider, R.J. A widespread alternate form of cap-dependent mRNA translation initiation. Nat. Commun. 2018, 9, 3068. [Google Scholar] [CrossRef]
- des Georges, A.; Dhote, V.; Kuhn, L.; Hellen, C.U.; Pestova, T.V.; Frank, J.; Hashem, Y. Structure of mammalian eIF3 in the context of the 43S preinitiation complex. Nature 2015, 525, 491–495. [Google Scholar] [CrossRef]
- Jivotovskaya, A.V.; Valasek, L.; Hinnebusch, A.G.; Nielsen, K.H. Eukaryotic translation initiation factor 3 (eIF3) and eIF2 can promote mRNA binding to 40S subunits independently of eIF4G in yeast. Mol. Cell. Biol. 2006, 26, 1355–1372. [Google Scholar] [CrossRef]
- Akulich, K.A.; Andreev, D.E.; Terenin, I.M.; Smirnova, V.V.; Anisimova, A.S.; Makeeva, D.S.; Arkhipova, V.I.; Stolboushkina, E.A.; Garber, M.B.; Prokofjeva, M.M.; et al. Four translation initiation pathways employed by the leaderless mRNA in eukaryotes. Sci. Rep. 2016, 6, 37905. [Google Scholar] [CrossRef]
- Shirokikh, N.E.; Spirin, A.S. Poly(A) leader of eukaryotic mRNA bypasses the dependence of translation on initiation factors. Proc. Natl. Acad. Sci. USA 2008, 105, 10738–10743. [Google Scholar] [CrossRef]
- Merrick, W.C. Cap-dependent and cap-independent translation in eukaryotic systems. Gene 2004, 332, 1–11. [Google Scholar] [CrossRef]
- Lockard, R.E.; Lane, C. Requirement for 7-methylguanosine in translation of globin mRNA in vivo. Nucleic Acids Res. 1978, 5, 3237–3247. [Google Scholar] [CrossRef][Green Version]
- Alekhina, O.M.; Vassilenko, K.S.; Spirin, A.S. Translation of non-capped mRNAs in a eukaryotic cell-free system: Acceleration of initiation rate in the course of polysome formation. Nucleic Acids Res. 2007, 35, 6547–6559. [Google Scholar] [CrossRef]
- Erzberger, J.P.; Stengel, F.; Pellarin, R.; Zhang, S.; Schaefer, T.; Aylett, C.H.S.; Cimermancic, P.; Boehringer, D.; Sali, A.; Aebersold, R.; et al. Molecular architecture of the 40SeIF1eIF3 translation initiation complex. Cell 2014, 158, 1123–1135. [Google Scholar] [CrossRef]
- Schmitt, E.; Panvert, M.; Lazennec-Schurdevin, C.; Coureux, P.D.; Perez, J.; Thompson, A.; Mechulam, Y. Structure of the ternary initiation complex aIF2-GDPNP-methionylated initiator tRNA. Nat. Struct. Mol. Biol. 2012, 19, 450–454. [Google Scholar] [CrossRef]
- Jennings, M.D.; Kershaw, C.J.; Adomavicius, T.; Pavitt, G.D. Fail-safe control of translation initiation by dissociation of eIF2alpha phosphorylated ternary complexes. Elife 2017, 6. [Google Scholar] [CrossRef]
- Sokabe, M.; Fraser, C.S. Human eukaryotic initiation factor 2 (eIF2)-GTP-Met-tRNAi ternary complex and eIF3 stabilize the 43 S preinitiation complex. J. Biol. Chem. 2014, 289, 31827–31836. [Google Scholar] [CrossRef]
- Arndt, N.; Ross-Kaschitza, D.; Kojukhov, A.; Komar, A.A.; Altmann, M. Properties of the ternary complex formed by yeast eIF4E, p20 and mRNA. Sci. Rep. 2018, 8, 6707. [Google Scholar] [CrossRef]
- Valásek, L.; Nielsen, K.H.; Hinnebusch, A.G. Direct eIF2-eIF3 contact in the multifactor complex is important for translation initiation in vivo. EMBO J. 2002, 21, 5886–5898. [Google Scholar] [CrossRef]
- Valasek, L.S.; Zeman, J.; Wagner, S.; Beznoskova, P.; Pavlikova, Z.; Mohammad, M.P.; Hronova, V.; Herrmannova, A.; Hashem, Y.; Gunisova, S. Embraced by eIF3: Structural and functional insights into the roles of eIF3 across the translation cycle. Nucleic Acids Res. 2017, 45, 10948–10968. [Google Scholar] [CrossRef]
- Simonetti, A.; Brito Querido, J.; Myasnikov, A.G.; Mancera-Martinez, E.; Renaud, A.; Kuhn, L.; Hashem, Y. eIF3 Peripheral Subunits Rearrangement after mRNA Binding and Start-Codon Recognition. Mol. Cell 2016, 63, 206–217. [Google Scholar] [CrossRef]
- Valasek, L.; Nielsen, K.H.; Zhang, F.; Fekete, C.A.; Hinnebusch, A.G. Interactions of eukaryotic translation initiation factor 3 (eIF3) subunit NIP1/c with eIF1 and eIF5 promote preinitiation complex assembly and regulate start codon selection. Mol. Cell. Biol. 2004, 24, 9437–9455. [Google Scholar] [CrossRef]
- Sokabe, M.; Fraser, C.S.; Hershey, J.W. The human translation initiation multi-factor complex promotes methionyl-tRNAi binding to the 40S ribosomal subunit. Nucleic Acids Res. 2012, 40, 905–913. [Google Scholar] [CrossRef]
- Wang, X.Q.; Rothnagel, J.A. 5’-untranslated regions with multiple upstream AUG codons can support low-level translation via leaky scanning and reinitiation. Nucleic Acids Res. 2004, 32, 1382–1391. [Google Scholar] [CrossRef]
- Haimov, O.; Sinvani, H.; Dikstein, R. Cap-dependent, scanning-free translation initiation mechanisms. Biochim. Biophys. Acta 2015, 1849, 1313–1318. [Google Scholar] [CrossRef]
- Gunisova, S.; Beznoskova, P.; Mohammad, M.P.; Vlckova, V.; Valasek, L.S. In-depth analysis of cis-determinants that either promote or inhibit reinitiation on GCN4 mRNA after translation of its four short uORFs. RNA 2016, 22, 542–558. [Google Scholar] [CrossRef]
- Mohammad, M.P.; Munzarova Pondelickova, V.; Zeman, J.; Gunisova, S.; Valasek, L.S. In vivo evidence that eIF3 stays bound to ribosomes elongating and terminating on short upstream ORFs to promote reinitiation. Nucleic Acids Res. 2017, 45, 2658–2674. [Google Scholar] [CrossRef]
- Lageix, S.; Rothenburg, S.; Dever, T.E.; Hinnebusch, A.G. Enhanced interaction between pseudokinase and kinase domains in Gcn2 stimulates eIF2alpha phosphorylation in starved cells. PLoS Genet. 2014, 10, e1004326. [Google Scholar] [CrossRef]
- Visweswaraiah, J.; Lee, S.J.; Hinnebusch, A.G.; Sattlegger, E. Overexpression of eukaryotic translation elongation factor 3 impairs Gcn2 protein activation. J. Biol. Chem. 2012, 287, 37757–37768. [Google Scholar] [CrossRef]
- Zhang, F.; Hinnebusch, A.G. An upstream ORF with non-AUG start codon is translated in vivo but dispensable for translational control of GCN4 mRNA. Nucleic Acids Res. 2011, 39, 3128–3140. [Google Scholar] [CrossRef]
- Szamecz, B.; Rutkai, E.; Cuchalova, L.; Munzarova, V.; Herrmannova, A.; Nielsen, K.H.; Burela, L.; Hinnebusch, A.G.; Valasek, L. eIF3a cooperates with sequences 5’ of uORF1 to promote resumption of scanning by post-termination ribosomes for reinitiation on GCN4 mRNA. Genes Dev. 2008, 22, 2414–2425. [Google Scholar] [CrossRef]
- Hronova, V.; Mohammad, M.P.; Wagner, S.; Panek, J.; Gunisova, S.; Zeman, J.; Poncova, K.; Valasek, L.S. Does eIF3 promote reinitiation after translation of short upstream ORFs also in mammalian cells? RNA Biol. 2017, 14, 1660–1667. [Google Scholar] [CrossRef]
- Gunisova, S.; Valasek, L.S. Fail-safe mechanism of GCN4 translational control--uORF2 promotes reinitiation by analogous mechanism to uORF1 and thus secures its key role in GCN4 expression. Nucleic Acids Res. 2014, 42, 5880–5893. [Google Scholar] [CrossRef]
- Rozen, F.; Edery, I.; Meerovitch, K.; Dever, T.E.; Merrick, W.C.; Sonenberg, N. Bidirectional RNA helicase activity of eucaryotic translation initiation factors 4A and 4F. Mol. Cell. Biol. 1990, 10, 1134–1144. [Google Scholar] [CrossRef]
- Sen, N.D.; Zhou, F.; Harris, M.S.; Ingolia, N.T.; Hinnebusch, A.G. eIF4B stimulates translation of long mRNAs with structured 5’ UTRs and low closed-loop potential but weak dependence on eIF4G. Proc. Natl. Acad. Sci. USA 2016, 113, 10464–10472. [Google Scholar] [CrossRef]
- Sun, Y.; Atas, E.; Lindqvist, L.M.; Sonenberg, N.; Pelletier, J.; Meller, A. Single-molecule kinetics of the eukaryotic initiation factor 4AI upon RNA unwinding. Structure 2014, 22, 941–948. [Google Scholar] [CrossRef]
- Sun, Y.; Atas, E.; Lindqvist, L.; Sonenberg, N.; Pelletier, J.; Meller, A. The eukaryotic initiation factor eIF4H facilitates loop-binding, repetitive RNA unwinding by the eIF4A DEAD-box helicase. Nucleic Acids Res. 2012, 40, 6199–6207. [Google Scholar] [CrossRef]
- Andreou, A.Z.; Harms, U.; Klostermeier, D. Single-stranded regions modulate conformational dynamics and ATPase activity of eIF4A to optimize 5’-UTR unwinding. Nucleic Acids Res. 2019, 47, 5260–5275. [Google Scholar] [CrossRef]
- Gupta, N.; Lorsch, J.R.; Hinnebusch, A.G. Yeast Ded1 promotes 48S translation pre-initiation complex assembly in an mRNA-specific and eIF4F-dependent manner. Elife 2018, 7. [Google Scholar] [CrossRef]
- Sharma, D.; Jankowsky, E. The Ded1/DDX3 subfamily of DEAD-box RNA helicases. Crit. Rev. Biochem. Mol. Biol. 2014, 49, 343–360. [Google Scholar] [CrossRef]
- Gao, Z.; Putnam, A.A.; Bowers, H.A.; Guenther, U.P.; Ye, X.; Kindsfather, A.; Hilliker, A.K.; Jankowsky, E. Coupling between the DEAD-box RNA helicases Ded1p and eIF4A. Elife 2016, 5. [Google Scholar] [CrossRef]
- Abaeva, I.S.; Marintchev, A.; Pisareva, V.P.; Hellen, C.U.; Pestova, T.V. Bypassing of stems versus linear base-by-base inspection of mammalian mRNAs during ribosomal scanning. EMBO J. 2011, 30, 115–129. [Google Scholar] [CrossRef]
- Dhote, V.; Sweeney, T.R.; Kim, N.; Hellen, C.U.; Pestova, T.V. Roles of individual domains in the function of DHX29, an essential factor required for translation of structured mammalian mRNAs. Proc. Natl. Acad. Sci. USA 2012, 109, E3150–E3159. [Google Scholar] [CrossRef]
- Marintchev, A. Roles of helicases in translation initiation: A mechanistic view. Biochim. Biophys. Acta 2013, 1829, 799–809. [Google Scholar] [CrossRef]
- Obayashi, E.; Luna, R.E.; Nagata, T.; Martin-Marcos, P.; Hiraishi, H.; Singh, C.R.; Erzberger, J.P.; Zhang, F.; Arthanari, H.; Morris, J.; et al. Molecular Landscape of the Ribosome Pre-initiation Complex during mRNA Scanning: Structural Role for eIF3c and Its Control by eIF5. Cell Rep. 2017, 18, 2651–2663. [Google Scholar] [CrossRef]
- Parsyan, A.; Svitkin, Y.; Shahbazian, D.; Gkogkas, C.; Lasko, P.; Merrick, W.C.; Sonenberg, N. mRNA helicases: The tacticians of translational control. Nat. Rev. Mol. Cell Biol. 2011, 12, 235–245. [Google Scholar] [CrossRef]
- Pisareva, V.P.; Pisarev, A.V. DHX29 reduces leaky scanning through an upstream AUG codon regardless of its nucleotide context. Nucleic Acids Res. 2016, 44, 4252–4265. [Google Scholar] [CrossRef]
- Sen, N.D.; Zhou, F.; Ingolia, N.T.; Hinnebusch, A.G. Genome-wide analysis of translational efficiency reveals distinct but overlapping functions of yeast DEAD-box RNA helicases Ded1 and eIF4A. Genome Res. 2015, 25, 1196–1205. [Google Scholar] [CrossRef]
- Soto-Rifo, R.; Rubilar, P.S.; Limousin, T.; de Breyne, S.; Decimo, D.; Ohlmann, T. DEAD-box protein DDX3 associates with eIF4F to promote translation of selected mRNAs. EMBO J. 2012, 31, 3745–3756. [Google Scholar] [CrossRef]
- Garcia-Garcia, C.; Frieda, K.L.; Feoktistova, K.; Fraser, C.S.; Block, S.M. RNA BIOCHEMISTRY. Factor-dependent processivity in human eIF4A DEAD-box helicase. Science 2015, 348, 1486–1488. [Google Scholar] [CrossRef]
- Sharifulin, D.E.; Bartuli, Y.S.; Meschaninova, M.I.; Ven’yaminova, A.G.; Graifer, D.M.; Karpova, G.G. Exploring accessibility of structural elements of the mammalian 40S ribosomal mRNA entry channel at various steps of translation initiation. Biochim. Biophys. Acta 2016, 1864, 1328–1338. [Google Scholar] [CrossRef]
- Pisareva, V.P.; Pisarev, A.V.; Komar, A.A.; Hellen, C.U.; Pestova, T.V. Translation initiation on mammalian mRNAs with structured 5’UTRs requires DExH-box protein DHX29. Cell 2008, 135, 1237–1250. [Google Scholar] [CrossRef]
- Asnani, M.; Pestova, T.V.; Hellen, C.U. Initiation on the divergent Type I cadicivirus IRES: Factor requirements and interactions with the translation apparatus. Nucleic Acids Res. 2016, 44, 3390–3407. [Google Scholar] [CrossRef][Green Version]
- Hussain, T.; Llacer, J.L.; Fernandez, I.S.; Munoz, A.; Martin-Marcos, P.; Savva, C.G.; Lorsch, J.R.; Hinnebusch, A.G.; Ramakrishnan, V. Structural changes enable start codon recognition by the eukaryotic translation initiation complex. Cell 2014, 159, 597–607. [Google Scholar] [CrossRef]
- Kozak, M. Downstream secondary structure facilitates recognition of initiator codons by eukaryotic ribosomes. Proc. Natl. Acad. Sci. USA 1990, 87, 8301–8305. [Google Scholar] [CrossRef]
- Thakur, A.; Marler, L.; Hinnebusch, A.G. A network of eIF2beta interactions with eIF1 and Met-tRNAi promotes accurate start codon selection by the translation preinitiation complex. Nucleic Acids Res. 2019, 47, 2574–2593. [Google Scholar] [CrossRef]
- Llacer, J.L.; Hussain, T.; Saini, A.K.; Nanda, J.S.; Kaur, S.; Gordiyenko, Y.; Kumar, R.; Hinnebusch, A.G.; Lorsch, J.R.; Ramakrishnan, V. Translational initiation factor eIF5 replaces eIF1 on the 40S ribosomal subunit to promote start-codon recognition. Elife 2018, 7. [Google Scholar] [CrossRef]
- Kearse, M.G.; Wilusz, J.E. Non-AUG translation: A new start for protein synthesis in eukaryotes. Genes Dev. 2017, 31, 1717–1731. [Google Scholar] [CrossRef]
- Starck, S.R.; Tsai, J.C.; Chen, K.; Shodiya, M.; Wang, L.; Yahiro, K.; Martins-Green, M.; Shastri, N.; Walter, P. Translation from the 5’ untranslated region shapes the integrated stress response. Science 2016, 351, aad3867. [Google Scholar] [CrossRef]
- Hellen, C.U.T. Translation Termination and Ribosome Recycling in Eukaryotes. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef]
- Weisser, M.; Schafer, T.; Leibundgut, M.; Bohringer, D.; Aylett, C.H.S.; Ban, N. Structural and Functional Insights into Human Re-initiation Complexes. Mol. Cell 2017, 67, 447–456. [Google Scholar] [CrossRef]
- Buskirk, A.R.; Green, R. Ribosome pausing, arrest and rescue in bacteria and eukaryotes. Philos. Trans. R. Soc. B Biol. Sci. 2017, 372. [Google Scholar] [CrossRef]
- Saini, A.K.; Nanda, J.S.; Martin-Marcos, P.; Dong, J.; Zhang, F.; Bhardwaj, M.; Lorsch, J.R.; Hinnebusch, A.G. Eukaryotic translation initiation factor eIF5 promotes the accuracy of start codon recognition by regulating Pi release and conformational transitions of the preinitiation complex. Nucleic Acids Res. 2014, 42, 9623–9640. [Google Scholar] [CrossRef]
- Jennings, M.D.; Pavitt, G.D. A new function and complexity for protein translation initiation factor eIF2B. Cell Cycle 2014, 13, 2660–2665. [Google Scholar] [CrossRef]
- Wortham, N.C.; Proud, C.G. eIF2B: Recent structural and functional insights into a key regulator of translation. Biochem. Soc. Trans. 2015, 43, 1234–1240. [Google Scholar] [CrossRef]
- Bogorad, A.M.; Lin, K.Y.; Marintchev, A. Novel mechanisms of eIF2B action and regulation by eIF2alpha phosphorylation. Nucleic Acids Res. 2017, 45, 11962–11979. [Google Scholar] [CrossRef]
- Schuller, A.P.; Wu, C.C.; Dever, T.E.; Buskirk, A.R.; Green, R. eIF5A Functions Globally in Translation Elongation and Termination. Mol. Cell 2017, 66, 194–205. [Google Scholar] [CrossRef]
- Rubio, C.A.; Weisburd, B.; Holderfield, M.; Arias, C.; Fang, E.; DeRisi, J.L.; Fanidi, A. Transcriptome-wide characterization of the eIF4A signature highlights plasticity in translation regulation. Genome Biol. 2014, 15, 476. [Google Scholar] [CrossRef]
- Sharma, D.K.; Bressler, K.; Patel, H.; Balasingam, N.; Thakor, N. Role of Eukaryotic Initiation Factors during Cellular Stress and Cancer Progression. J. Nucleic Acids 2016, 2016, 8235121. [Google Scholar] [CrossRef]
- Kozak, M. Regulation of translation via mRNA structure in prokaryotes and eukaryotes. Gene 2005, 361, 13–37. [Google Scholar] [CrossRef]
- Sonenberg, N.; Hinnebusch, A.G. Regulation of translation initiation in eukaryotes: Mechanisms and biological targets. Cell 2009, 136, 731–745. [Google Scholar] [CrossRef]
- Wu, C.C.; Zinshteyn, B.; Wehner, K.A.; Green, R. High-Resolution Ribosome Profiling Defines Discrete Ribosome Elongation States and Translational Regulation during Cellular Stress. Mol. Cell 2019, 73, 959–970. [Google Scholar] [CrossRef]
- Khitun, A.; Ness, T.J.; Slavoff, S.A. Small open reading frames and cellular stress responses. Mol. Omics 2019, 15, 108–116. [Google Scholar] [CrossRef]
- Harding, H.P.; Novoa, I.; Zhang, Y.; Zeng, H.; Wek, R.; Schapira, M.; Ron, D. Regulated translation initiation controls stress-induced gene expression in mammalian cells. Mol. Cell 2000, 6, 1099–1108. [Google Scholar] [CrossRef]
- Leprivier, G.; Rotblat, B.; Khan, D.; Jan, E.; Sorensen, P.H. Stress-mediated translational control in cancer cells. Biochim. Biophys. Acta 2015, 1849, 845–860. [Google Scholar] [CrossRef]
- Chen, J.; Choi, J.; O’Leary, S.E.; Prabhakar, A.; Petrov, A.; Grosely, R.; Puglisi, E.V.; Puglisi, J.D. The molecular choreography of protein synthesis: Translational control, regulation, and pathways. Q. Rev. Biophys. 2016, 49, e11. [Google Scholar] [CrossRef]
- Breaker, R.R. Riboswitches and Translation Control. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef]
- Hor, J.; Gorski, S.A.; Vogel, J. Bacterial RNA Biology on a Genome Scale. Mol. Cell 2018, 70, 785–799. [Google Scholar] [CrossRef]
- Kirchner, M.; Schneider, S. Gene expression control by Bacillus anthracis purine riboswitches. RNA 2017, 23, 762–769. [Google Scholar] [CrossRef]
- Zoschke, R.; Bock, R. Chloroplast Translation: Structural and Functional Organization, Operational Control, and Regulation. Plant Cell 2018, 30, 745–770. [Google Scholar] [CrossRef]
- Oruganty-Das, A.; Ng, T.; Udagawa, T.; Goh, E.L.; Richter, J.D. Translational control of mitochondrial energy production mediates neuron morphogenesis. Cell Metab. 2012, 16, 789–800. [Google Scholar] [CrossRef]
- Ingolia, N.T.; Ghaemmaghami, S.; Newman, J.R.; Weissman, J.S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 2009, 324, 218–223. [Google Scholar] [CrossRef]
- Couvillion, M.T.; Soto, I.C.; Shipkovenska, G.; Churchman, L.S. Synchronized mitochondrial and cytosolic translation programs. Nature 2016, 533, 499–503. [Google Scholar] [CrossRef]
- Falcone, C.; Agostinelli, M.; Frontali, L. Mitochondrial Translation Products during Release from Glucose Repression in Saccharomyces cerevisiae. J. Bacteriol. 1983, 153, 1125–1132. [Google Scholar]
- Richter, J.D.; Sonenberg, N. Regulation of cap-dependent translation by eIF4E inhibitory proteins. Nature 2005, 433, 477–480. [Google Scholar] [CrossRef]
- Gingras, A.C.; Gygi, S.P.; Raught, B.; Polakiewicz, R.D.; Abraham, R.T.; Hoekstra, M.F.; Aebersold, R.; Sonenberg, N. Regulation of 4E-BP1 phosphorylation: A novel two-step mechanism. Genes Dev. 1999, 13, 1422–1437. [Google Scholar] [CrossRef]
- Scheper, G.C.; van Kollenburg, B.; Hu, J.; Luo, Y.; Goss, D.J.; Proud, C.G. Phosphorylation of eukaryotic initiation factor 4E markedly reduces its affinity for capped mRNA. J. Biol. Chem. 2002, 277, 3303–3309. [Google Scholar] [CrossRef]
- Qin, X.; Jiang, B.; Zhang, Y. 4E-BP1, a multifactor regulated multifunctional protein. Cell Cycle 2016, 15, 781–786. [Google Scholar] [CrossRef]
- Wullschleger, S.; Loewith, R.; Hall, M.N. TOR signaling in growth and metabolism. Cell 2006, 124, 471–484. [Google Scholar] [CrossRef]
- Xie, J.; Proud, C.G. Signaling crosstalk between the mTOR complexes. Translation 2014, 2, e28174. [Google Scholar] [CrossRef]
- Thoreen, C.C.; Chantranupong, L.; Keys, H.R.; Wang, T.; Gray, N.S.; Sabatini, D.M. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 2012, 485, 109–113. [Google Scholar] [CrossRef]
- Hsieh, A.C.; Liu, Y.; Edlind, M.P.; Ingolia, N.T.; Janes, M.R.; Sher, A.; Shi, E.Y.; Stumpf, C.R.; Christensen, C.; Bonham, M.J.; et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 2012, 485, 55–61. [Google Scholar] [CrossRef]
- Reiling, J.H.; Sabatini, D.M. Stress and mTORture signaling. Oncogene 2006, 25, 6373–6383. [Google Scholar] [CrossRef]
- Saxton, R.A.; Sabatini, D.M. mTOR Signaling in Growth, Metabolism, and Disease. Cell 2017, 168, 960–976. [Google Scholar] [CrossRef]
- Gingras, A.C.; Raught, B.; Sonenberg, N. Regulation of translation initiation by FRAP/mTOR. Genes Dev. 2001, 15, 807–826. [Google Scholar] [CrossRef]
- Altmann, M.; Schmitz, N.; Berset, C.; Trachsel, H. A novel inhibitor of cap-dependent translation initiation in yeast: p20 competes with eIF4G for binding to eIF4E. EMBO J. 1997, 16, 1114–1121. [Google Scholar] [CrossRef]
- Cosentino, G.P.; Schmelzle, T.; Haghighat, A.; Helliwell, S.B.; Hall, M.N.; Sonenberg, N. Eap1p, a Novel Eukaryotic Translation Initiation Factor 4E-Associated Protein in Saccharomyces cerevisiae. Mol. Cell. Biol. 2000, 20, 4604–4613. [Google Scholar] [CrossRef]
- Gruner, S.; Weber, R.; Peter, D.; Chung, M.Y.; Igreja, C.; Valkov, E.; Izaurralde, E. Structural motifs in eIF4G and 4E-BPs modulate their binding to eIF4E to regulate translation initiation in yeast. Nucleic Acids Res. 2018, 46, 6893–6908. [Google Scholar] [CrossRef]
- Castelli, L.M.; Talavera, D.; Kershaw, C.J.; Mohammad-Qureshi, S.S.; Costello, J.L.; Rowe, W.; Sims, P.F.; Grant, C.M.; Hubbard, S.J.; Ashe, M.P.; et al. The 4E-BP Caf20p Mediates both eIF4E-Dependent and Independent Repression of Translation. PLoS Genet. 2015, 11, e1005233. [Google Scholar] [CrossRef]
- Masvidal, L.; Hulea, L.; Furic, L.; Topisirovic, I.; Larsson, O. mTOR-sensitive translation: Cleared fog reveals more trees. RNA Biol. 2017, 14, 1299–1305. [Google Scholar] [CrossRef]
- Nakamura, A.; Sato, K.; Hanyu-Nakamura, K. Drosophila cup is an eIF4E binding protein that associates with Bruno and regulates oskar mRNA translation in oogenesis. Dev. Cell 2004, 6, 69–78. [Google Scholar] [CrossRef]
- Nelson, M.R.; Leidal, A.M.; Smibert, C.A. Drosophila Cup is an eIF4E-binding protein that functions in Smaug-mediated translational repression. EMBO J. 2004, 23, 150–159. [Google Scholar] [CrossRef]
- Le Bacquer, O.; Petroulakis, E.; Paglialunga, S.; Poulin, F.; Richard, D.; Cianflone, K.; Sonenberg, N. Elevated sensitivity to diet-induced obesity and insulin resistance in mice lacking 4E-BP1 and 4E-BP2. J. Clin. Investig. 2007, 117, 387–396. [Google Scholar] [CrossRef]
- Rapley, J.; Oshiro, N.; Ortiz-Vega, S.; Avruch, J. The mechanism of insulin-stimulated 4E-BP protein binding to mammalian target of rapamycin (mTOR) complex 1 and its contribution to mTOR complex 1 signaling. J. Biol. Chem. 2011, 286, 38043–38053. [Google Scholar] [CrossRef]
- Mackey, A.M.; Sarkes, D.A.; Bettencourt, I.; Asara, J.M.; Rameh, L.E. PIP4kgamma is a substrate for mTORC1 that maintains basal mTORC1 signaling during starvation. Sci. Signal. 2014, 7, ra104. [Google Scholar] [CrossRef]
- Lecompte, O.; Poch, O.; Laporte, J. PtdIns5P regulation through evolution: Roles in membrane trafficking? Trends Biochem. Sci. 2008, 33, 453–460. [Google Scholar] [CrossRef]
- Itoh, T.; Ijuin, T.; Takenawa, T. A novel phosphatidylinositol-5-phosphate 4-kinase (phosphatidylinositol-phosphate kinase IIgamma) is phosphorylated in the endoplasmic reticulum in response to mitogenic signals. J. Biol. Chem. 1998, 273, 20292–20299. [Google Scholar] [CrossRef]
- Teleman, A.A. Molecular mechanisms of metabolic regulation by insulin in Drosophila. Biochem. J. 2009, 425, 13–26. [Google Scholar] [CrossRef]
- Bjordal, M.; Arquier, N.; Kniazeff, J.; Pin, J.P.; Leopold, P. Sensing of amino acids in a dopaminergic circuitry promotes rejection of an incomplete diet in Drosophila. Cell 2014, 156, 510–521. [Google Scholar] [CrossRef]
- Marshall, L.; Rideout, E.J.; Grewal, S.S. Nutrient/TOR-dependent regulation of RNA polymerase III controls tissue and organismal growth in Drosophila. EMBO J. 2012, 31, 1916–1930. [Google Scholar] [CrossRef]
- Nagarajan, S.; Grewal, S.S. An investigation of nutrient-dependent mRNA translation in Drosophila larvae. Biol. Open 2014, 3, 1020–1031. [Google Scholar] [CrossRef]
- Binda, M.; Peli-Gulli, M.P.; Bonfils, G.; Panchaud, N.; Urban, J.; Sturgill, T.W.; Loewith, R.; De Virgilio, C. The Vam6 GEF controls TORC1 by activating the EGO complex. Mol. Cell 2009, 35, 563–573. [Google Scholar] [CrossRef]
- Reineke, L.C.; Cao, Y.; Baus, D.; Hossain, N.M.; Merrick, W.C. Insights into the role of yeast eIF2A in IRES-mediated translation. PLoS ONE 2011, 6, e24492. [Google Scholar] [CrossRef]
- Miura, F.; Kawaguchi, N.; Sese, J.; Toyoda, A.; Hattori, M.; Morishita, S.; Ito, T. A large-scale full-length cDNA analysis to explore the budding yeast transcriptome. Proc. Natl. Acad. Sci. USA 2006, 103, 17846–17851. [Google Scholar] [CrossRef]
- Iizuka, N.; Najita, L.; Franzusoff, A.; Sarnow, P. Cap-dependent and cap-independent translation by internal initiation of mRNAs in cell extracts prepared from Saccharomyces cerevisiae. Mol. Cell. Biol. 1994, 14, 7322–7330. [Google Scholar] [CrossRef]
- Searfoss, A.; Dever, T.E.; Wickner, R. Linking the 3’ poly(A) tail to the subunit joining step of translation initiation: Relations of Pab1p, eukaryotic translation initiation factor 5b (Fun12p), and Ski2p-Slh1p. Mol. Cell. Biol. 2001, 21, 4900–4908. [Google Scholar] [CrossRef]
- Zampar, G.G.; Kummel, A.; Ewald, J.; Jol, S.; Niebel, B.; Picotti, P.; Aebersold, R.; Sauer, U.; Zamboni, N.; Heinemann, M. Temporal system-level organization of the switch from glycolytic to gluconeogenic operation in yeast. Mol. Syst. Biol. 2013, 9, 651. [Google Scholar] [CrossRef]
- Fernandez, J.; Bode, B.; Koromilas, A.; Diehl, J.A.; Krukovets, I.; Snider, M.D.; Hatzoglou, M. Translation mediated by the internal ribosome entry site of the cat-1 mRNA is regulated by glucose availability in a PERK kinase-dependent manner. J. Biol. Chem. 2002, 277, 11780–11787. [Google Scholar] [CrossRef]
- Ray, P.S.; Grover, R.; Das, S. Two internal ribosome entry sites mediate the translation of p53 isoforms. EMBO Rep. 2006, 7, 404–410. [Google Scholar] [CrossRef]
- Singh, S.; Sreenath, K.; Pavithra, L.; Roy, S.; Chattopadhyay, S. SMAR1 regulates free radical stress through modulation of AKR1a4 enzyme activity. Int. J. Biochem. Cell Biol. 2010, 42, 1105–1114. [Google Scholar] [CrossRef]
- Khan, D.; Chattopadhyay, S.; Das, S. Influence of metabolic stress on translation of p53 isoforms. Mol. Cell Oncol. 2016, 3, e1039689. [Google Scholar] [CrossRef]
- Bieging, K.T.; Mello, S.S.; Attardi, L.D. Unravelling mechanisms of p53-mediated tumour suppression. Nat. Rev. Cancer 2014, 14, 359–370. [Google Scholar] [CrossRef]
- Humpton, T.J.; Vousden, K.H. Regulation of Cellular Metabolism and Hypoxia by p53. Cold Spring Harb. Perspect. Med. 2016, 6. [Google Scholar] [CrossRef]
- Malonia, S.K.; Sinha, S.; Lakshminarasimhan, P.; Singh, K.; Jalota-Badhwar, A.; Rampalli, S.; Kaul-Ghanekar, R.; Chattopadhyay, S. Gene regulation by SMAR1: Role in cellular homeostasis and cancer. Biochim. Biophys. Acta 2011, 1815, 1–12. [Google Scholar] [CrossRef]
- Knoch, K.P.; Nath-Sain, S.; Petzold, A.; Schneider, H.; Beck, M.; Wegbrod, C.; Sonmez, A.; Munster, C.; Friedrich, A.; Roivainen, M.; et al. PTBP1 is required for glucose-stimulated cap-independent translation of insulin granule proteins and Coxsackieviruses in beta cells. Mol. Metab. 2014, 3, 518–530. [Google Scholar] [CrossRef]
- Chen, C.H.; Lin, H.Y.; Pan, C.L.; Chen, F.C. The genomic features that affect the lengths of 5’ untranslated regions in multicellular eukaryotes. BMC Bioinform. 2011, 12 (Suppl. 9), S3. [Google Scholar] [CrossRef]
- Walker, S.E.; Zhou, F.; Mitchell, S.F.; Larson, V.S.; Valasek, L.; Hinnebusch, A.G.; Lorsch, J.R. Yeast eIF4B binds to the head of the 40S ribosomal subunit and promotes mRNA recruitment through its N-terminal and internal repeat domains. RNA 2013, 19, 191–207. [Google Scholar] [CrossRef]
- Shirokikh, N.E.; Archer, S.K.; Beilharz, T.H.; Powell, D.; Preiss, T. Translation complex profile sequencing to study the in vivo dynamics of mRNA-ribosome interactions during translation initiation, elongation and termination. Nat. Protoc. 2017, 12, 697–731. [Google Scholar] [CrossRef]
- Ashe, M.P.; Slaven, J.W.; De Long, S.K.; Ibrahimo, S.; Sachs, A.B. A novel eIF2B-dependent mechanism of translational control in yeast as a response to fusel alcohols. EMBO J. 2001, 20, 6464–6474. [Google Scholar] [CrossRef]
- Iwasaki, S.; Iwasaki, W.; Takahashi, M.; Sakamoto, A.; Watanabe, C.; Shichino, Y.; Floor, S.N.; Fujiwara, K.; Mito, M.; Dodo, K.; et al. The Translation Inhibitor Rocaglamide Targets a Bimolecular Cavity between eIF4A and Polypurine RNA. Mol. Cell 2019, 73, 738–748.e9. [Google Scholar] [CrossRef]
- Iwasaki, S.; Floor, S.N.; Ingolia, N.T. Rocaglates convert DEAD-box protein eIF4A into a sequence-selective translational repressor. Nature 2016, 534, 558–561. [Google Scholar] [CrossRef]
- Jennings, M.D.; Zhou, Y.; Mohammad-Qureshi, S.S.; Bennett, D.; Pavitt, G.D. eIF2B promotes eIF5 dissociation from eIF2*GDP to facilitate guanine nucleotide exchange for translation initiation. Genes Dev. 2013, 27, 2696–2707. [Google Scholar] [CrossRef]
- Krishnamoorthy, T.; Pavitt, G.D.; Zhang, F.; Dever, T.E.; Hinnebusch, A.G. Tight binding of the phosphorylated alpha subunit of initiation factor 2 (eIF2alpha) to the regulatory subunits of guanine nucleotide exchange factor eIF2B is required for inhibition of translation initiation. Mol. Cell. Biol. 2001, 21, 5018–5030. [Google Scholar] [CrossRef]
- Dey, M.; Trieselmann, B.; Locke, E.G.; Lu, J.; Cao, C.; Dar, A.C.; Krishnamoorthy, T.; Dong, J.; Sicheri, F.; Dever, T.E. PKR and GCN2 kinases and guanine nucleotide exchange factor eukaryotic translation initiation factor 2B (eIF2B) recognize overlapping surfaces on eIF2alpha. Mol. Cell. Biol. 2005, 25, 3063–3075. [Google Scholar] [CrossRef]
- Vattem, K.M.; Wek, R.C. Reinitiation involving upstream ORFs regulates ATF4 mRNA translation in mammalian cells. Proc. Natl. Acad. Sci. USA 2004, 101, 11269–11274. [Google Scholar] [CrossRef]
- Kilberg, M.S.; Shan, J.; Su, N. ATF4-dependent transcription mediates signaling of amino acid limitation. Trends Endocrinol. Metab. 2009, 20, 436–443. [Google Scholar] [CrossRef]
- Broer, S.; Broer, A. Amino acid homeostasis and signalling in mammalian cells and organisms. Biochem. J. 2017, 474, 1935–1963. [Google Scholar] [CrossRef]
- Sonenberg, N.; Newgard, C.B. The Perks of Balancing Glucose. Science 2001, 293, 818. [Google Scholar] [CrossRef]
- Balsa, E.; Soustek, M.S.; Thomas, A.; Cogliati, S.; Garcia-Poyatos, C.; Martin-Garcia, E.; Jedrychowski, M.; Gygi, S.P.; Enriquez, J.A.; Puigserver, P. ER and Nutrient Stress Promote Assembly of Respiratory Chain Supercomplexes through the PERK-eIF2alpha Axis. Mol. Cell 2019, 74, 877–890. [Google Scholar] [CrossRef]
- Cnop, M.; Toivonen, S.; Igoillo-Esteve, M.; Salpea, P. Endoplasmic reticulum stress and eIF2alpha phosphorylation: The Achilles heel of pancreatic beta cells. Mol. Metab. 2017, 6, 1024–1039. [Google Scholar] [CrossRef]
- Marc Delépine, M.N.; Barrett, T.; Golamaully, M.; Lathrop, M.; Julier, C. EIF2AK3, encoding translation initiation factor 2-α kinase 3, is mutated in patients with Wolcott-Rallison syndrome. Nat. Genet. 2000, 25, 406–409. [Google Scholar] [CrossRef]
- Calkhoven, C.F.; Muller, C.; Leutz, A. Translational control of gene expression and disease. Trends Mol. Med. 2002, 8, 577–583. [Google Scholar] [CrossRef]
- Harding, H.P.; Zhang, Y.; Jungries, R.; Chung, P.; Plesken, H.; Sabatini, D.D.; Ron, D. Diabetes Mellitus and Exocrine Pancreatic Dysfunction in Perk-/- Mice Reveals a Role for Translational Control in Secretory Cell Survival. Mol. Cell 2001, 7, 1153–1163. [Google Scholar] [CrossRef]
- Zhang, P.; McGrath, B.; Li, S.A.; Frank, A.; Zambito, F.; Reinert, J.; Gannon, M.; Ma, K.; McNaughton, K.; Cavener, D.R. The PERK Eukaryotic Initiation Factor 2 Kinase Is Required for the Development of the Skeletal System, Postnatal Growth, and the Function and Viability of the Pancreas. Mol. Cell. Biol. 2002, 22, 3864–3874. [Google Scholar] [CrossRef]
- Moore, M.J. From birth to death: The complex lives of eukaryotic mRNAs. Science 2005, 309, 1514–1518. [Google Scholar] [CrossRef]
- Van Treeck, B.; Parker, R. Emerging Roles for Intermolecular RNA-RNA Interactions in RNP Assemblies. Cell 2018, 174, 791–802. [Google Scholar] [CrossRef]
- Keene, J.D. RNA regulons: Coordination of post-transcriptional events. Nat. Rev. Genet. 2007, 8, 533–543. [Google Scholar] [CrossRef]
- Buchan, J.R. mRNP granules. Assembly, function, and connections with disease. RNA Biol. 2014, 11, 1019–1030. [Google Scholar] [CrossRef]
- Dash, S.; Siddam, A.D.; Barnum, C.E.; Janga, S.C.; Lachke, S.A. RNA-binding proteins in eye development and disease: Implication of conserved RNA granule components. Wiley Interdiscip. Rev. RNA 2016, 7, 527–557. [Google Scholar] [CrossRef]
- Johnson, M.E.; Grassetti, A.V.; Taroni, J.N.; Lyons, S.M.; Schweppe, D.; Gordon, J.K.; Spiera, R.F.; Lafyatis, R.; Anderson, P.J.; Gerber, S.A.; et al. Stress granules and RNA processing bodies are novel autoantibody targets in systemic sclerosis. Arthritis Res. Ther. 2016, 18, 27. [Google Scholar] [CrossRef]
- Mahboubi, H.; Stochaj, U. Cytoplasmic stress granules: Dynamic modulators of cell signaling and disease. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 884–895. [Google Scholar] [CrossRef]
- Stoecklin, G.; Kedersha, N. Relationship of GW/P-bodies with stress granules. Adv. Exp. Med. Biol. 2013, 768, 197–211. [Google Scholar] [CrossRef]
- Kshirsagar, M.; Parker, R. Identification of Edc3p as an Enhancer of mRNA Decapping in Saccharomyces cerevisiae. Genet. Soc. Am. 2004, 166, 729–739. [Google Scholar] [CrossRef]
- Nissan, T.; Rajyaguru, P.; She, M.; Song, H.; Parker, R. Decapping activators in Saccharomyces cerevisiae act by multiple mechanisms. Mol. Cell 2010, 39, 773–783. [Google Scholar] [CrossRef]
- Decker, C.J.; Parker, R. P-bodies and stress granules: Possible roles in the control of translation and mRNA degradation. Cold Spring Harb. Perspect. Biol. 2012, 4, a012286. [Google Scholar] [CrossRef]
- Teixeira, D.; Sheth, U.; Valencia-Sanchez, M.A.; Brengues, M.; Parker, R. Processing bodies require RNA for assembly and contain nontranslating mRNAs. RNA 2005, 11, 371–382. [Google Scholar] [CrossRef]
- Pizzinga, M.; Ashe, M.P. Yeast mRNA localization: Protein asymmetry, organelle localization and response to stress. Biochem. Soc. Trans. 2014, 42, 1256–1260. [Google Scholar] [CrossRef]
- Wang, C.; Schmich, F.; Srivatsa, S.; Weidner, J.; Beerenwinkel, N.; Spang, A. Context-dependent deposition and regulation of mRNAs in P-bodies. Elife 2018, 7. [Google Scholar] [CrossRef]
- Standart, N.; Weil, D. P-Bodies: Cytosolic Droplets for Coordinated mRNA Storage. Trends Genet. 2018, 34, 612–626. [Google Scholar] [CrossRef]
- Sheth, U.; Parker, R. Decapping and Decay of Messenger RNA Occur in Cytoplasmic Processing Bodies. Science 2003, 300, 805–808. [Google Scholar] [CrossRef]
- Sheth, U.; Parker, R. Targeting of aberrant mRNAs to cytoplasmic processing bodies. Cell 2006, 125, 1095–1109. [Google Scholar] [CrossRef]
- Bhattacharyya, S.N.; Habermacher, R.; Martine, U.; Closs, E.I.; Filipowicz, W. Stress-induced reversal of microRNA repression and mRNA P-body localization in human cells. Cold Spring Harb. Symp. Quant. Biol. 2006, 71, 513–521. [Google Scholar] [CrossRef]
- Pillai, R.S.; Bhattacharyya, S.N.; Artus, C.G.; Zoller, T.; Cougot, N.; Basyuk, E.; Bertrand, E.; Filipowicz, W. Inhibition of Translational Initiation by Let-7 MicroRNA in Human Cells. Science 2005, 309, 1573–1576. [Google Scholar] [CrossRef]
- Buchan, J.R.; Yoon, J.H.; Parker, R. Stress-specific composition, assembly and kinetics of stress granules in Saccharomyces cerevisiae. J. Cell Sci. 2011, 124, 228–239. [Google Scholar] [CrossRef]
- Steiger, M.; Carr-Schmid, A.; Schwartz, D.C.; Kiledjian, M.; Parker, R. Analysis of recombinant yeast decapping enzyme. RNA 2003, 9, 231–238. [Google Scholar] [CrossRef]
- Carroll, J.S.; Munchel, S.E.; Weis, K. The DExD/H box ATPase Dhh1 functions in translational repression, mRNA decay, and processing body dynamics. J. Cell Biol. 2011, 194, 527–537. [Google Scholar] [CrossRef]
- Pelechano, V.; Wei, W.; Steinmetz, L.M. Widespread Co-translational RNA Decay Reveals Ribosome Dynamics. Cell 2015, 161, 1400–1412. [Google Scholar] [CrossRef]
- Sweet, T.; Kovalak, C.; Coller, J. The DEAD-box protein Dhh1 promotes decapping by slowing ribosome movement. PLoS Biol. 2012, 10, e1001342. [Google Scholar] [CrossRef]
- Buchan, J.R.; Muhlrad, D.; Parker, R. P bodies promote stress granule assembly in Saccharomyces cerevisiae. J. Cell Biol. 2008, 183, 441–455. [Google Scholar] [CrossRef]
- Aryanpur, P.P.; Regan, C.A.; Collins, J.M.; Mittelmeier, T.M.; Renner, D.M.; Vergara, A.M.; Brown, N.P.; Bolger, T.A. Gle1 Regulates RNA Binding of the DEAD-Box Helicase Ded1 in Its Complex Role in Translation Initiation. Mol. Cell. Biol. 2017, 37. [Google Scholar] [CrossRef]
- Johnstone, O.; Deuring, R.; Bock, R.; Linder, P.; Fuller, M.T.; Lasko, P. Belle is a Drosophila DEAD-box protein required for viability and in the germ line. Dev. Biol. 2005, 277, 92–101. [Google Scholar] [CrossRef]
- Lai, M.C.; Lee, Y.H.; Tarn, W.Y. The DEAD-box RNA helicase DDX3 associates with export messenger ribonucleoproteins as well as tip-associated protein and participates in translational control. Mol. Biol. Cell 2008, 19, 3847–3858. [Google Scholar] [CrossRef]
- Guenther, U.P.; Weinberg, D.E.; Zubradt, M.M.; Tedeschi, F.A.; Stawicki, B.N.; Zagore, L.L.; Brar, G.A.; Licatalosi, D.D.; Bartel, D.P.; Weissman, J.S.; et al. The helicase Ded1p controls use of near-cognate translation initiation codons in 5’ UTRs. Nature 2018, 559, 130–134. [Google Scholar] [CrossRef]
- Campbell, S.G.; Hoyle, N.P.; Ashe, M.P. Dynamic cycling of eIF2 through a large eIF2B-containing cytoplasmic body: Implications for translation control. J. Cell Biol. 2005, 170, 925–934. [Google Scholar] [CrossRef]
- Coller, J.M.; Tucker, M.; Sheth, U.; Valencia-Sanchez, M.A.; Parker, R. The DEAD box helicase, Dhh1p, functions in mRNA decapping and interacts with both the decapping and deadenylase complexes. RNA 2001, 7, 1717–1727. [Google Scholar] [CrossRef]
- Chang, L.C.; Lee, F.J. The RNA helicase Dhh1p cooperates with Rbp1p to promote porin mRNA decay via its non-conserved C-terminal domain. Nucleic Acids Res. 2012, 40, 1331–1344. [Google Scholar] [CrossRef]
- Kedersha, N.L.; Gupta, M.; Li, W.; Miller, I.; Anderson, P. RNA-Binding Proteins Tia-1 and Tiar Link the Phosphorylation of Eif-2α to the Assembly of Mammalian Stress Granules. J. Cell Biol. 1999, 147, 1431–1442. [Google Scholar] [CrossRef]
- Andrei, M.A.; Ingelfinger, D.; Heintzmann, R.; Achsel, T.; Rivera-Pomar, R.; Luhrmann, R. A role for eIF4E and eIF4E-transporter in targeting mRNPs to mammalian processing bodies. RNA 2005, 11, 717–727. [Google Scholar] [CrossRef]
- Wu, J.; Zhang, N.; Hayes, A.; Panoutsopoulou, K.; Oliver, S.G. Global analysis of nutrient control of gene expression in Saccharomyces cerevisiae during growth and starvation. Proc. Natl. Acad. Sci. USA 2004, 101, 3148–3153. [Google Scholar] [CrossRef]
| Conditions of Stress | Organism | Main Observations and Conclusions or Models | Reference |
|---|---|---|---|
| 10 min in YP lacking glucose (0% w/v added) at 30 °C | S. cerevisiae yAS2568 (W3031A) | Near-complete disassembly of polysomes observed after 10 min of starvation. Subsequent addition of glucose partially rescued polysomes within 1–2.5 min and completely restored polysomes in 10 min. | [40] |
| 10 min in YP with normal and low glucose (2.0, 0.7, 0.5, and 0.1% w/v added) at 30 °C | S. cerevisiae yMK37 (BY4741) | Disassembly of polysomes observed when glucose levels were lower than 0.5% w/v. High concentration of glucose was determined as necessary for a yeast batch culture to maintain efficient translation. | [42] |
| 10, 30, 60, 120 min in YP lacking glucose (0% w/v added) at 30 °C | S. cerevisiae (MATα ura3 leu2 trp1 his3) (1278b) | Polysomes remained low-abundant after 10 and 30 min of glucose starvation and showed partial recovery after 60 min, with further increased after 120 min. | [55] |
| 15 min in SC lacking glucose (0% w/v added) at 30 °C | S. cerevisiae (BY4741) and EY0690 (W303 MATa trp1-1 leu2-3 ura3-1 his3-11 can1-100) | Ribosome profiling revealed an inversed correlation between the change in ribosome occupancy upon glucose starvation and the change in mRNA expression levels. | [56] |
| 20, 40, and 60 min of Oxygen (0%) Glucose Deprivation (OGD); cells maintained in DMEM (lacking glucose), at 95% N2 and 5% CO2, 37 °C. | Rattus norvegicus PC12 (neural cell line) | Significant alteration of translation of approximately 3,000 genes occurred in the first 20 min of the stress (ribo-seq data). | [57] |
| Conditions of Stress | Organism | Main Observations and Conclusions or Models | Reference |
|---|---|---|---|
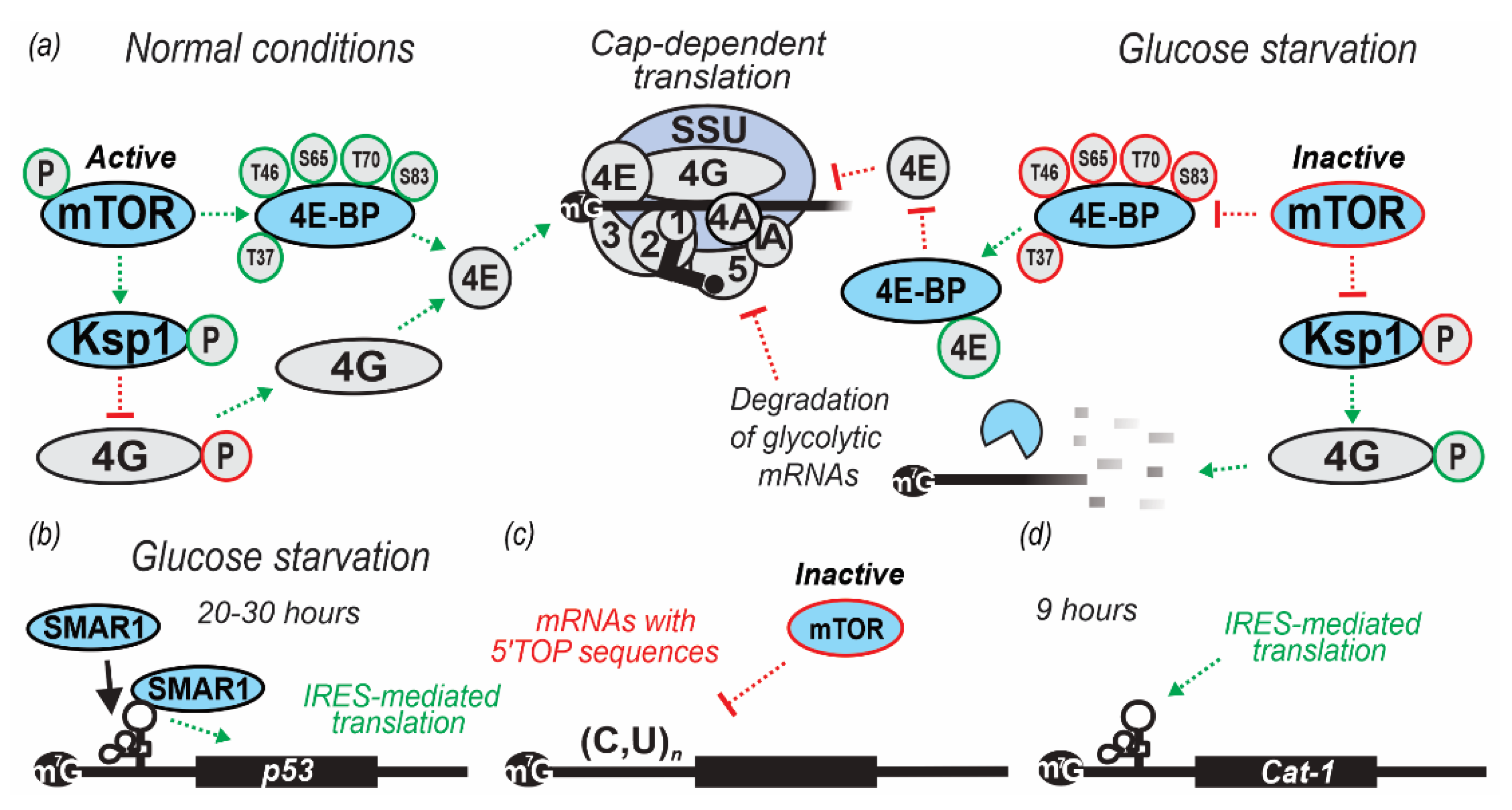
| 10 min in YP lacking glucose (0% w/v added) | S. cerevisiae yMK1751 (caf20Δ), yMK1752 (eap1Δ) and yMK1752 (caf20Δ eap1Δ) (BY4741) | Single and double mutant strains Caf20Δ and Eap1Δ exhibited polysome disassembly as a result of starvation. Therefore, Caf20Δ and Eap1Δ 4E-binding proteins concluded as not critical for inhibition of translation initiation during glucose starvation in yeast. | [42] |
| 30 min in YP lacking glucose (0% w/v added) | S. cerevisiae (BY4741) | During glucose starvation, eIF4G was phosphorylated by the serine/threonine protein kinase Ksp1p. Thirty-three phosphorylated residues of eIF4G were identified in WT cells and nine (S17, T138, S140, T183, T399, T400, S502, S503, T712) were identified in ksp1Δ cells during glucose starvation. This phosphorylation is regulated by both Snf1p/AMPK and TORC1 pathways. TOR promotes anabolic, and AMPK promotes catabolic responses when cells are high or low on nutrients, respectively. | [58] |
| 4, 8, 20 and 30 h in DMEM lacking glucose at 37 °C | Homo sapiens H1299 (p53-null lung cells derived from metastatic site lymph node) and A549 cells (lung-derived, expressing endogenous p53) | Glucose starvation was shown to induce cap-independent production of p53 and increase scaffold/matrix attachment region-binding protein 1 (SMAR1) abundance in the cytoplasm. SMAR1 was demonstrated to bind to the p53 IRES and suggested to control translation of mRNA encoding p53 isoform (Δ40p53). The increase in p53 isoform production was reversible, suggesting that transient glucose or dietary deprivation could impinge reversibly on p53 signaling, as also suggested by p53-target transactivation observed in the study. | [59] |
| Conditions of stress | Organism | Main observations and conclusions or models | Reference |
|---|---|---|---|
| 10 min in SC lacking (0% w/v) glucose | S. cerevisiae (BY4741) | An increase in the levels of eIF4G and Pab1p co-purifying with eIF3 was observed (transient build-up of eIF4E, eIF4G, Pab1p, and eIF3) in the first 10 min of the glucose starvation. After 30 min of starvation, this association weakened with possible re-localization of the corresponding closed-loop complex (CLC) factors and mRNAs into cytoplasmic granules. Severe reduction in the levels of eIF4G:eIF4A complex observed at both 10 and 30 min. | [42] |
| 20, 40, and 60 min of Oxygen (0%) Glucose Deprivation (OGD); cells maintained in DMEM (lacking glucose) at 95% N2 and 5% CO2, 37 °C. | Rattus norvegicus PC12 (neural cell line) | Widespread increase of ribosome density in mRNA 5’UTRs upstream of annotated main coding ORFs, increased efficiency of termination from 20–60 min of starvation as observed by ribosome profiling. | [57] |
| Conditions of Stress | Organism | Main Observations and Conclusions or Models | Reference |
|---|---|---|---|
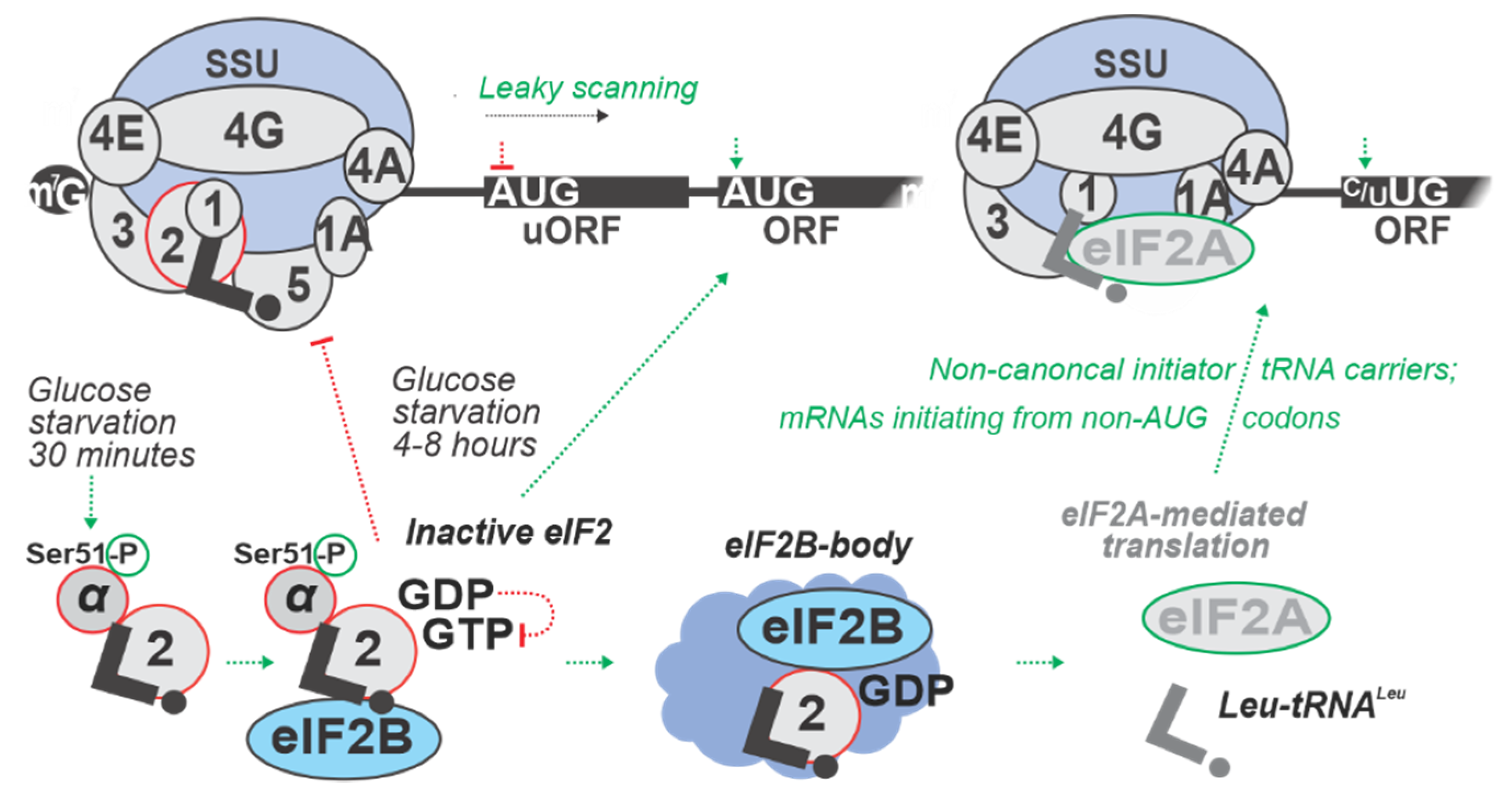
| 8 h in SC with minimal glucose (0.05% w/v added) | S. cerevisiae RY124 (MATα ura3-52 leu2 trp1 p180 [GCN4-lacZ, URA3]) | eIF2α phosphorylation by Gcn2p protein kinase is increased transiently 4 h into and stays increased up to 8 h of glucose starvation. Gcn2p function found to contribute to the maintenance of glycogen levels during prolonged glucose starvation, suggesting a link between amino acid control and glycogen metabolism. | [60] |
| 20, 40, and 60 min of Oxygen (0%) Glucose Deprivation (OGD); cells maintained in DMEM (lacking glucose), at 95% N2 and 5% CO2, 37 °C | Rattus norvegicus PC12 (neural cell line) | uORF-mediated inhibition of annotated coding ORFs (acORF) translation at all time points. E.g., Eif1a and Eef2k mRNAs demonstrated increased uORF translation with an accompanying decrease in acORF translation. OGD induced translation from non-AUG codons, leading to an extended variability of protein isoforms, as demonstrated for Adm, Bcl211, Fam178b, Ptms, and Ppp1r2 mRNAs. The strongest increase in translation of uORFs was observed for Eif1a and Junb mRNAs with over 10-fold increase in ribosome density in uORFs during OGD. | [57] |
| 24 h in DMEM lacking glucose (0% w/v added) | Mus musculus (Ser51Ala eIF2α mutant), mouse embryonic fibroblasts (MEF) cell line | Upon low glucose, protein folding is inefficient, and PERK activated. This was demonstrated to result in an increased eIF2α phosphorylation. Lack of ternary complex (TC) was shown to inhibit translation (including of pre-pro-insulin) and lead to an increase of transcription of ER stress response genes. | [9] |
| Conditions of Stress | Organism | Main Observations and Conclusions or Models | Reference |
|---|---|---|---|
| 15 min in SC lacking (0% w/v) glucose at 30 °C | S. cerevisiae BY4741 and EY0690 (W303 MATa trp1-1 leu2-3 ura3-1 his3-11 can1-100) | Differential re-localization of mRNAs into P-bodies and SGs was observed. mRNAs of proteins responsible for fermentation of glucose to ethanol, such as PDC1 and involved in glycolysis and gluconeogenesis, such as PGK1 localized in P-bodies, whereas mRNAs transcriptionally activated by Hsf1p via heat-specific elements (such as HSP30 and HSP26) remained diffusely localized in the cytoplasm and highly translated. mRNA like GLC3 (required for glycogen biosynthesis) and HXK1 (Hexokinase) that are activated via stress-regulated elements independent of Hsf1p localized to both, P-bodies and SGs and were translationally repressed during glucose starvation. | [40,56] |
| 10 min in SC lacking (0% w/v) glucose at 30 °C | S. cerevisiae Yrp1724 (MATa leu2-3,112 trp1 ura3-52 his4-539 cup1::LEU2/PGK1pG/MFA2pG DHH1-GFP (NEO)) | P-bodies were formed and found to contain various proteins implicated in mRNA degradation. In glucose starved cells, P-bodies increased in brightness and number when stained for Dcp2p (decapping enzyme), Dhh1p (activator of decapping enzyme). An increase in the concentration of decapping activators like Lsm1p, Pat1p, Edc3p, Dcp1p, and Xrn1p exonuclease, was also observed within P-bodies upon glucose starvation stress. | [61,62] |
| 30 min in YP lacking glucose (0% w/v added) at 30 °C | S. cerevisiae (BY4741) | Ksp1p -dependent phosphorylation of eIF4G was found to promote degradation of mRNAs encoding glycolytic proteins (PGK1, TDH3, ENO2 and CDC19) by inducing Dhh1p recruitment. Ksp1p also was shown to cause rapid degradation of ribosomal protein (RP) mRNAs (e.g., RPL37A and RPS8B) during glucose starvation conditions. | [63,58] |
| 30 min in SC lacking (0% w/v) glucose at 30 °C | S. cerevisiae (BY4741) | Accumulation of eIF4E, eIF4G1, eIF4G2 and Pab1p was observed in cytoplasmic granules after 30 min, but not 10 min of starvation. | [42,64] |
| 15 min in SC lacking (0% w/v) glucose at 30 °C | S. cerevisiae yRP2065 (BY4741) | DEAD-box protein Ded1p (mammalian ortholog: DDX3) was found to accumulate in SGs. Ded1p was also shown to assemble translationally repressed mRNPs that accumulated in SGs independent of Ded1p ATPase activity. It was demonstrated that the Ded1p ATPase was required for the release of mRNPs from SGs. | [63] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Janapala, Y.; Preiss, T.; Shirokikh, N.E. Control of Translation at the Initiation Phase During Glucose Starvation in Yeast. Int. J. Mol. Sci. 2019, 20, 4043. https://doi.org/10.3390/ijms20164043
Janapala Y, Preiss T, Shirokikh NE. Control of Translation at the Initiation Phase During Glucose Starvation in Yeast. International Journal of Molecular Sciences. 2019; 20(16):4043. https://doi.org/10.3390/ijms20164043
Chicago/Turabian StyleJanapala, Yoshika, Thomas Preiss, and Nikolay E. Shirokikh. 2019. "Control of Translation at the Initiation Phase During Glucose Starvation in Yeast" International Journal of Molecular Sciences 20, no. 16: 4043. https://doi.org/10.3390/ijms20164043
APA StyleJanapala, Y., Preiss, T., & Shirokikh, N. E. (2019). Control of Translation at the Initiation Phase During Glucose Starvation in Yeast. International Journal of Molecular Sciences, 20(16), 4043. https://doi.org/10.3390/ijms20164043






