Epigenetic Biomarkers in Colorectal Cancer Patients Receiving Adjuvant or Neoadjuvant Therapy: A Systematic Review of Epidemiological Studies
Abstract
1. Introduction
2. Literature Search
2.1. Search Strategy and Selection Criteria
2.2. Selection Criteria and Data Collection
3. Results
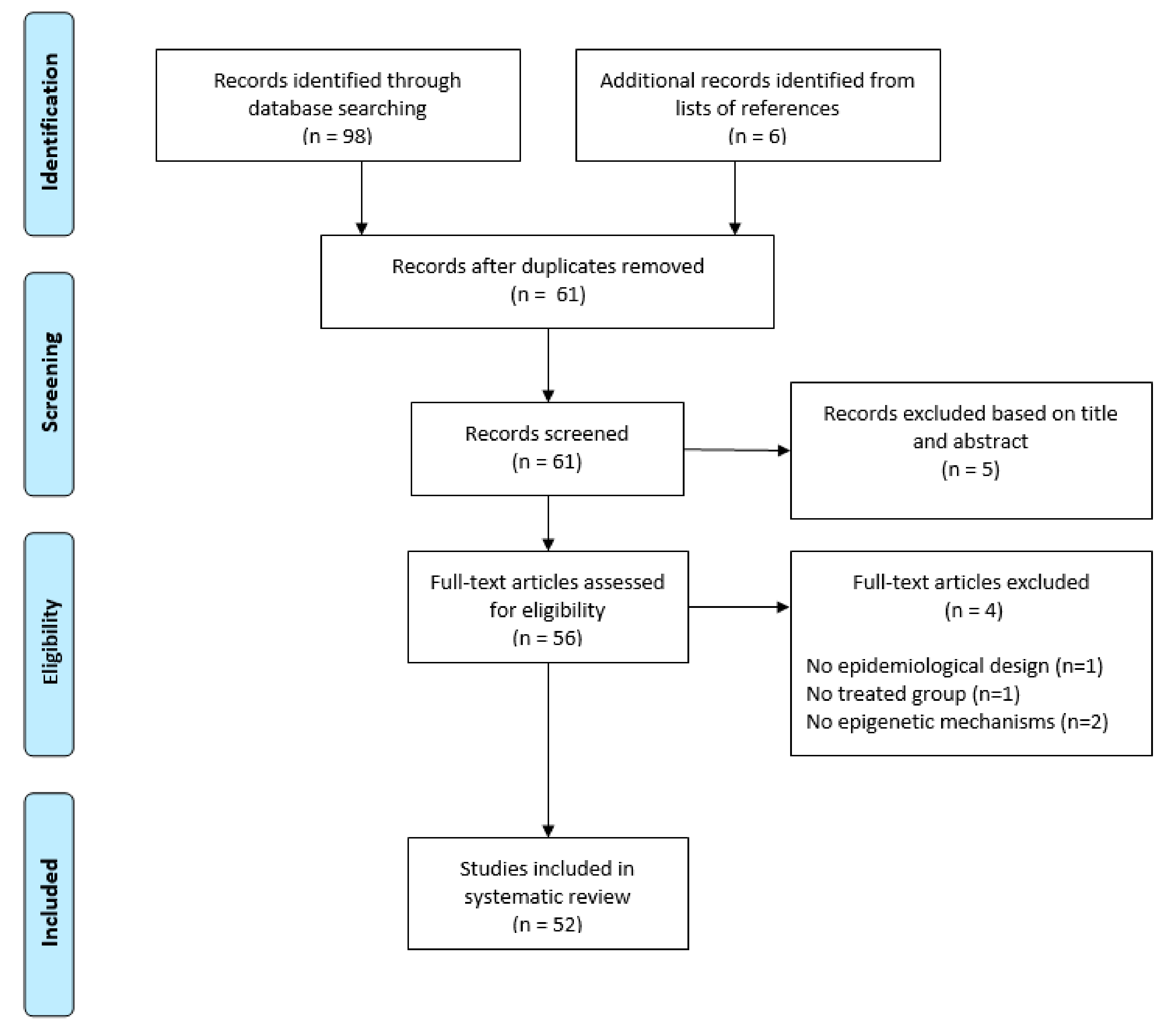
3.1. Study Characteristics
3.2. DNA Methylation in Patients Receiving Adjuvant Therapy
3.2.1. Gene-Specific Methylation
3.2.2. CpG Island Phenotype
3.2.3. Long Interspersed Nuclear Element-1 Methylation
3.3. DNA Methylation in Patients Receiving Neoadjuvant Therapy
3.3.1. Gene-Specific Methylation
3.3.2. CpG Island Methylator Phenotype
3.3.3. Global and Genome-Wide Methylation
3.4. MicroRNA in Patients Receiving Adjuvant Therapy
3.5. MicroRNA in Patients Receiving Neoadjuvant Therapy
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Globocan. Globocan 2018: Estimated Cancer Incidence, Mortality and Prevalence Worldwide in 2018. Available online: http://globocan.iarc.fr/Default.aspx (accessed on 1 May 2019).
- Abubakar, I.I.; Tillmann, T.; Banerjee, A. Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990–2013: A systematic analysis for the Global Burden of Disease Study 2013. Lancet 2015, 385, 117–171. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Jubb, A.M.; Bell, S.M.; Quirke, P. Methylation and colorectal cancer. J. Pathol. 2001, 195, 111–134. [Google Scholar] [CrossRef] [PubMed]
- Jubb, A.M.; Quirke, P.; Oates, A.J. DNA methylation, a biomarker for colorectal cancer: Implications for screening and pathological utility. Ann. N. Y. Acad. Sci. 2003, 983, 251–267. [Google Scholar] [CrossRef] [PubMed]
- Lao, V.V.; Grady, W.M. Epigenetics and colorectal cancer. Nat. Rev. Gastroenterol. Hepatol. 2011, 8, 686–700. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.H.; Petko, Z.; Dzieciatkowski, S.; Lin, L.; Ghiassi, M.; Stain, S.; Chapman, W.C.; Washington, M.K.; Willis, J.; Markowitz, S.D.; et al. CpG island methylation of genes accumulates during the adenoma progression step of the multistep pathogenesis of colorectal cancer. Genes Chromosom. Cancer 2006, 45, 781–789. [Google Scholar] [CrossRef] [PubMed]
- Hammoud, S.S.; Cairns, B.R.; Jones, D.A. Epigenetic regulation of colon cancer and intestinal stem cells. Curr. Opin. Cell Biol. 2013, 25, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Auclair, G.; Weber, M. Mechanisms of DNA methylation and demethylation in mammals. Biochimie 2012, 94, 2202–2211. [Google Scholar] [CrossRef]
- Worthley, D.L.; Whitehall, V.L.; Buttenshaw, R.L.; Irahara, N.; Greco, S.A.; Ramsnes, I.; Mallitt, K.A.; Le Leu, R.K.; Winter, J.; Hu, Y.; et al. DNA methylation within the normal colorectal mucosa is associated with pathway-specific predisposition to cancer. Oncogene 2010, 29, 1653–1662. [Google Scholar] [CrossRef]
- Guo, H.; Ingolia, N.T.; Weissman, J.S.; Bartel, D.P. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 2010, 466, 835–840. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Sayed, D.; Abdellatif, M. MicroRNAs in development and disease. Physiol. Rev. 2011, 91, 827–887. [Google Scholar] [CrossRef] [PubMed]
- Barchitta, M.; Maugeri, A.; Quattrocchi, A.; Agrifoglio, O.; Agodi, A. The Role of miRNAs as Biomarkers for Pregnancy Outcomes: A Comprehensive Review. Int. J. Genom. 2017, 2017, 8067972. [Google Scholar] [CrossRef] [PubMed]
- Rapado-González, Ó.; Álvarez-Castro, A.; López-López, R.; Iglesias-Canle, J.; Suárez-Cunqueiro, M.M.; Muinelo-Romay, L. Circulating microRNAs as Promising Biomarkers in Colorectal Cancer. Cancers 2019, 11, 898. [Google Scholar] [CrossRef] [PubMed]
- Fearon, E.R.; Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 1990, 61, 759–767. [Google Scholar] [CrossRef]
- Lengauer, C.; Kinzler, K.W.; Vogelstein, B. Genetic instabilities in human cancers. Nature 1998, 396, 643–649. [Google Scholar] [CrossRef] [PubMed]
- Kinzler, K.W.; Vogelstein, B. Lessons from hereditary colorectal cancer. Cell 1996, 87, 159–170. [Google Scholar] [CrossRef]
- Feinberg, A.P.; Tycko, B. The history of cancer epigenetics. Nat. Rev. Cancer 2004, 4, 143–153. [Google Scholar] [CrossRef]
- Sawan, C.; Herceg, Z. Histone modifications and cancer. Adv. Genet. 2010, 70, 57–85. [Google Scholar] [CrossRef]
- Van Engeland, M.; Derks, S.; Smits, K.M.; Meijer, G.A.; Herman, J.G. Colorectal cancer epigenetics: Complex simplicity. J. Clin. Oncol. 2011, 29, 1382–1391. [Google Scholar] [CrossRef] [PubMed]
- Kuipers, E.J.; Grady, W.M.; Lieberman, D.; Seufferlein, T.; Sung, J.J.; Boelens, P.G.; van de Velde, C.J.; Watanabe, T. Colorectal cancer. Nat. Rev. Dis. Prim. 2015, 1, 15065. [Google Scholar] [CrossRef] [PubMed]
- Labianca, R.; Nordlinger, B.; Beretta, G.D.; Mosconi, S.; Mandalà, M.; Cervantes, A.; Arnold, D.; Group, E.G.W. Early colon cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2013, 24, vi64–vi72. [Google Scholar] [CrossRef] [PubMed]
- Van Cutsem, E.; Cervantes, A.; Nordlinger, B.; Arnold, D.; Group, E.G.W. Metastatic colorectal cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2014, 25, iii1–iii9. [Google Scholar] [CrossRef] [PubMed]
- Juul, T.; Ahlberg, M.; Biondo, S.; Emmertsen, K.J.; Espin, E.; Jimenez, L.M.; Matzel, K.E.; Palmer, G.; Sauermann, A.; Trenti, L.; et al. International validation of the low anterior resection syndrome score. Ann. Surg. 2014, 259, 728–734. [Google Scholar] [CrossRef] [PubMed]
- Bregendahl, S.; Emmertsen, K.J.; Lindegaard, J.C.; Laurberg, S. Urinary and sexual dysfunction in women after resection with and without preoperative radiotherapy for rectal cancer: A population-based cross-sectional study. Colorectal Dis. 2015, 17, 26–37. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, A.; Ziegler, L.; Martland, M.; Davidson, S.; Efficace, F.; Sebag-Montefiore, D.; Velikova, G. Systematic Review of Radiation Therapy Toxicity Reporting in Randomized Controlled Trials of Rectal Cancer: A Comparison of Patient-Reported Outcomes and Clinician Toxicity Reporting. Int. J. Radiat. Oncol. Biol. Phys. 2015, 92, 555–567. [Google Scholar] [CrossRef]
- Williamson, J.S.; Harris, D.A.; Beynon, J.; Jenkins, G.J. Review of the development of DNA methylation as a marker of response to neoadjuvant therapy and outcomes in rectal cancer. Clin. Epigenet. 2015, 7, 70. [Google Scholar] [CrossRef]
- Dayde, D.; Tanaka, I.; Jain, R.; Tai, M.C.; Taguchi, A. Predictive and Prognostic Molecular Biomarkers for Response to Neoadjuvant Chemoradiation in Rectal Cancer. Int. J. Mol. Sci. 2017, 18, 573. [Google Scholar] [CrossRef]
- Moher, D.; Shamseer, L.; Clarke, M.; Ghersi, D.; Liberati, A.; Petticrew, M.; Shekelle, P.; Stewart, L.A.; Group, P.P. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst. Rev. 2015, 4, 1. [Google Scholar] [CrossRef]
- Nagasaka, T.; Sharp, G.B.; Notohara, K.; Kambara, T.; Sasamoto, H.; Isozaki, H.; MacPhee, D.G.; Jass, J.R.; Tanaka, N.; Matsubara, N. Hypermethylation of O6-methylguanine-DNA methyltransferase promoter may predict nonrecurrence after chemotherapy in colorectal cancer cases. Clin. Cancer Res. 2003, 9, 5306–5312. [Google Scholar] [PubMed]
- Chen, S.P.; Chiu, S.C.; Wu, C.C.; Lin, S.Z.; Kang, J.C.; Chen, Y.L.; Lin, P.C.; Pang, C.Y.; Harn, H.J. The association of methylation in the promoter of APC and MGMT and the prognosis of Taiwanese CRC patients. Genet. Test. Mol. Biomark. 2009, 13, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Miyoshi, Y.; Nagase, H.; Ando, H.; Horii, A.; Ichii, S.; Nakatsuru, S.; Aoki, T.; Miki, Y.; Mori, T.; Nakamura, Y. Somatic mutations of the APC gene in colorectal tumors: Mutation cluster region in the APC gene. Hum. Mol. Genet. 1992, 1, 229–233. [Google Scholar] [PubMed]
- Ide, T.; Kitajima, Y.; Ohtaka, K.; Mitsuno, M.; Nakafusa, Y.; Miyazaki, K. Expression of the hMLH1 gene is a possible predictor for the clinical response to 5-fluorouracil after a surgical resection in colorectal cancer. Oncol. Rep. 2008, 19, 1571–1576. [Google Scholar] [PubMed]
- Sinicrope, F.A.; Shi, Q.; Smyrk, T.C.; Thibodeau, S.N.; Dienstmann, R.; Guinney, J.; Bot, B.M.; Tejpar, S.; Delorenzi, M.; Goldberg, R.M.; et al. Molecular markers identify subtypes of stage III colon cancer associated with patient outcomes. Gastroenterology 2015, 148, 88–99. [Google Scholar] [CrossRef]
- Perez-Carbonell, L.; Balaguer, F.; Toiyama, Y.; Egoavil, C.; Rojas, E.; Guarinos, C.; Andreu, M.; Llor, X.; Castells, A.; Jover, R.; et al. IGFBP3 methylation is a novel diagnostic and predictive biomarker in colorectal cancer. PLoS ONE 2014, 9, e104285. [Google Scholar] [CrossRef]
- Heitzer, E.; Artl, M.; Filipits, M.; Resel, M.; Graf, R.; Weißenbacher, B.; Lax, S.; Gnant, M.; Wrba, F.; Greil, R.; et al. Differential survival trends of stage II colorectal cancer patients relate to promoter methylation status of PCDH10, SPARC, and UCHL1. Mod. Pathol. 2014, 27, 906–915. [Google Scholar] [CrossRef]
- Jeanes, A.; Gottardi, C.J.; Yap, A.S. Cadherins and cancer: How does cadherin dysfunction promote tumor progression? Oncogene 2008, 27, 6920–6929. [Google Scholar] [CrossRef]
- Ying, J.; Li, H.; Seng, T.J.; Langford, C.; Srivastava, G.; Tsao, S.W.; Putti, T.; Murray, P.; Chan, A.T.; Tao, Q. Functional epigenetics identifies a protocadherin PCDH10 as a candidate tumor suppressor for nasopharyngeal, esophageal and multiple other carcinomas with frequent methylation. Oncogene 2006, 25, 1070–1080. [Google Scholar] [CrossRef]
- Li, L.; Tao, Q.; Jin, H.; van Hasselt, A.; Poon, F.F.; Wang, X.; Zeng, M.S.; Jia, W.H.; Zeng, Y.X.; Chan, A.T.; et al. The tumor suppressor UCHL1 forms a complex with p53/MDM2/ARF to promote p53 signaling and is frequently silenced in nasopharyngeal carcinoma. Clin. Cancer Res. 2010, 16, 2949–2958. [Google Scholar] [CrossRef]
- Kim, J.H.; Jung, E.J.; Lee, H.S.; Kim, M.A.; Kim, W.H. Comparative analysis of DNA methylation between primary and metastatic gastric carcinoma. Oncol. Rep. 2009, 21, 1251–1259. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.Y.; Kuo, C.C.; Wu, C.C.; Hsiao, C.W.; Hu, J.M.; Hsu, C.H.; Chou, Y.C.; Shih, Y.L.; Lin, Y.W. NKX6.1 hypermethylation predicts the outcome of stage II colorectal cancer patients undergoing chemotherapy. Genes Chromosom. Cancer 2018, 57, 268–277. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.C.; Huang, R.L.; Wang, H.C.; Liao, Y.P.; Yu, M.H.; Lai, H.C. High methylation rate of LMX1A, NKX6-1, PAX1, PTPRR, SOX1, and ZNF582 genes in cervical adenocarcinoma. Int. J. Gynecol. Cancer 2014, 24, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Taylor, K.H.; Pena-Hernandez, K.E.; Davis, J.W.; Arthur, G.L.; Duff, D.J.; Shi, H.; Rahmatpanah, F.B.; Sjahputera, O.; Caldwell, C.W. Large-scale CpG methylation analysis identifies novel candidate genes and reveals methylation hotspots in acute lymphoblastic leukemia. Cancer Res. 2007, 67, 2617–2625. [Google Scholar] [CrossRef] [PubMed]
- Asada, K.; Nakajima, T.; Shimazu, T.; Yamamichi, N.; Maekita, T.; Yokoi, C.; Oda, I.; Ando, T.; Yoshida, T.; Nanjo, S.; et al. Demonstration of the usefulness of epigenetic cancer risk prediction by a multicentre prospective cohort study. Gut 2015, 64, 388–396. [Google Scholar] [CrossRef] [PubMed]
- Pfütze, K.; Benner, A.; Hoffmeister, M.; Jansen, L.; Yang, R.; Bläker, H.; Herpel, E.; Ulrich, A.; Ulrich, C.M.; Chang-Claude, J.; et al. Methylation status at HYAL2 predicts overall and progression-free survival of colon cancer patients under 5-FU chemotherapy. Genomics 2015, 106, 348–354. [Google Scholar] [CrossRef] [PubMed]
- Van Rijnsoever, M.; Elsaleh, H.; Joseph, D.; McCaul, K.; Iacopetta, B. CpG island methylator phenotype is an independent predictor of survival benefit from 5-fluorouracil in stage III colorectal cancer. Clin. Cancer Res. 2003, 9, 2898–2903. [Google Scholar] [PubMed]
- Min, B.H.; Bae, J.M.; Lee, E.J.; Yu, H.S.; Kim, Y.H.; Chang, D.K.; Kim, H.C.; Park, C.K.; Lee, S.H.; Kim, K.M.; et al. The CpG island methylator phenotype may confer a survival benefit in patients with stage II or III colorectal carcinomas receiving fluoropyrimidine-based adjuvant chemotherapy. BMC Cancer 2011, 11, 344. [Google Scholar] [CrossRef]
- Shiovitz, S.; Bertagnolli, M.M.; Renfro, L.A.; Nam, E.; Foster, N.R.; Dzieciatkowski, S.; Luo, Y.; Lao, V.V.; Monnat, R.J.; Emond, M.J.; et al. CpG island methylator phenotype is associated with response to adjuvant irinotecan-based therapy for stage III colon cancer. Gastroenterology 2014, 147, 637–645. [Google Scholar] [CrossRef]
- Cohen, S.A.; Wu, C.; Yu, M.; Gourgioti, G.; Wirtz, R.; Raptou, G.; Gkakou, C.; Kotoula, V.; Pentheroudakis, G.; Papaxoinis, G.; et al. Evaluation of CpG Island Methylator Phenotype as a Biomarker in Colorectal Cancer Treated with Adjuvant Oxaliplatin. Clin. Colorectal Cancer 2016, 15, 164–169. [Google Scholar] [CrossRef][Green Version]
- Jover, R.; Nguyen, T.P.; Pérez-Carbonell, L.; Zapater, P.; Payá, A.; Alenda, C.; Rojas, E.; Cubiella, J.; Balaguer, F.; Morillas, J.D.; et al. 5-Fluorouracil adjuvant chemotherapy does not increase survival in patients with CpG island methylator phenotype colorectal cancer. Gastroenterology 2011, 140, 1174–1181. [Google Scholar] [CrossRef] [PubMed]
- Han, S.W.; Lee, H.J.; Bae, J.M.; Cho, N.Y.; Lee, K.H.; Kim, T.Y.; Oh, D.Y.; Im, S.A.; Bang, Y.J.; Jeong, S.Y.; et al. Methylation and microsatellite status and recurrence following adjuvant FOLFOX in colorectal cancer. Int. J. Cancer 2013, 132, 2209–2216. [Google Scholar] [CrossRef] [PubMed]
- Bae, J.M.; Kim, J.H.; Kwak, Y.; Lee, D.W.; Cha, Y.; Wen, X.; Lee, T.H.; Cho, N.Y.; Jeong, S.Y.; Park, K.J.; et al. Distinct clinical outcomes of two CIMP-positive colorectal cancer subtypes based on a revised CIMP classification system. Br. J. Cancer 2017, 116, 1012–1020. [Google Scholar] [CrossRef] [PubMed]
- Babatz, T.D.; Burns, K.H. Functional impact of the human mobilome. Curr. Opin. Genet. Dev. 2013, 23, 264–270. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Fabris, S.; Ronchetti, D.; Agnelli, L.; Baldini, L.; Morabito, F.; Bicciato, S.; Basso, D.; Todoerti, K.; Lombardi, L.; Lambertenghi-Deliliers, G.; et al. Transcriptional features of multiple myeloma patients with chromosome 1q gain. Leukemia 2007, 21, 1113–1116. [Google Scholar] [CrossRef] [PubMed]
- Carreira, P.E.; Richardson, S.R.; Faulkner, G.J. L1 retrotransposons, cancer stem cells and oncogenesis. FEBS J. 2014, 281, 63–73. [Google Scholar] [CrossRef]
- Rodić, N.; Burns, K.H. Long interspersed element-1 (LINE-1): Passenger or driver in human neoplasms? PLoS Genet. 2013, 9, e1003402. [Google Scholar] [CrossRef] [PubMed]
- Barchitta, M.; Quattrocchi, A.; Maugeri, A.; Canto, C.; La Rosa, N.; Cantarella, M.A.; Spampinato, G.; Scalisi, A.; Agodi, A. LINE-1 hypermethylation in white blood cell DNA is associated with high-grade cervical intraepithelial neoplasia. BMC Cancer 2017, 17, 601. [Google Scholar] [CrossRef]
- Maugeri, A.; Barchitta, M.; Mazzone, M.G.; Giuliano, F.; Basile, G.; Agodi, A. Resveratrol modulates SIRT1 and DNMT functions and restores LINE-1 methylation levels in ARPE-19 cells under oxidative stress and inflammation. Int. J. Mol. Sci. 2018, 19, 2118. [Google Scholar] [CrossRef]
- Maugeri, A.; Mazzone, M.G.; Giuliano, F.; Vinciguerra, M.; Basile, G.; Barchitta, M.; Agodi, A. Curcumin Modulates DNA Methyltransferase Functions in a Cellular Model of Diabetic Retinopathy. Oxid. Med. Cell Longev. 2018, 2018, 5407482. [Google Scholar] [CrossRef]
- Bollati, V.; Schwartz, J.; Wright, R.; Litonjua, A.; Tarantini, L.; Suh, H.; Sparrow, D.; Vokonas, P.; Baccarelli, A. Decline in genomic DNA methylation through aging in a cohort of elderly subjects. Mech. Ageing Dev. 2009, 130, 234–239. [Google Scholar] [CrossRef] [PubMed]
- Bollati, V.; Galimberti, D.; Pergoli, L.; Dalla Valle, E.; Barretta, F.; Cortini, F.; Scarpini, E.; Bertazzi, P.A.; Baccarelli, A. DNA methylation in repetitive elements and Alzheimer disease. Brain Behav. Immun. 2011, 25, 1078–1083. [Google Scholar] [CrossRef] [PubMed]
- Barchitta, M.; Quattrocchi, A.; Maugeri, A.; Vinciguerra, M.; Agodi, A. LINE-1 hypomethylation in blood and tissue samples as an epigenetic marker for cancer risk: A systematic review and meta-analysis. PLoS ONE 2014, 9, e109478. [Google Scholar] [CrossRef] [PubMed]
- Agodi, A.; Barchitta, M.; Quattrocchi, A.; Maugeri, A.; Canto, C.; Marchese, A.E.; Vinciguerra, M. Low fruit consumption and folate deficiency are associated with LINE-1 hypomethylation in women of a cancer-free population. Genes Nutr. 2015, 10, 480. [Google Scholar] [CrossRef] [PubMed]
- Barchitta, M.; Maugeri, A.; Quattrocchi, A.; Barone, G.; Mazzoleni, P.; Catalfo, A.; De Guidi, G.; Iemmolo, M.G.; Crimi, N.; Agodi, A. Mediterranean Diet and Particulate Matter Exposure Are Associated With LINE-1 Methylation: Results From a Cross-Sectional Study in Women. Front. Genet. 2018, 9, 514. [Google Scholar] [CrossRef]
- Baccarelli, A.; Wright, R.O.; Bollati, V.; Tarantini, L.; Litonjua, A.A.; Suh, H.H.; Zanobetti, A.; Sparrow, D.; Vokonas, P.S.; Schwartz, J. Rapid DNA methylation changes after exposure to traffic particles. Am. J. Respir. Crit. Care Med. 2009, 179, 572–578. [Google Scholar] [CrossRef]
- De Prins, S.; Koppen, G.; Jacobs, G.; Dons, E.; Van de Mieroop, E.; Nelen, V.; Fierens, F.; Int Panis, L.; De Boever, P.; Cox, B.; et al. Influence of ambient air pollution on global DNA methylation in healthy adults: A seasonal follow-up. Environ. Int. 2013, 59, 418–424. [Google Scholar] [CrossRef]
- Schulz, W.A. L1 retrotransposons in human cancers. J. Biomed. Biotechnol. 2006, 2006, 83672. [Google Scholar] [CrossRef]
- Slotkin, R.K.; Martienssen, R. Transposable elements and the epigenetic regulation of the genome. Nat. Rev. Genet. 2007, 8, 272–285. [Google Scholar] [CrossRef]
- Kawakami, K.; Matsunoki, A.; Kaneko, M.; Saito, K.; Watanabe, G.; Minamoto, T. Long interspersed nuclear element-1 hypomethylation is a potential biomarker for the prediction of response to oral fluoropyrimidines in microsatellite stable and CpG island methylator phenotype-negative colorectal cancer. Cancer Sci. 2011, 102, 166–174. [Google Scholar] [CrossRef]
- Ogino, S.; Kawasaki, T.; Nosho, K.; Ohnishi, M.; Suemoto, Y.; Kirkner, G.J.; Fuchs, C.S. LINE-1 hypomethylation is inversely associated with microsatellite instability and CpG island methylator phenotype in colorectal cancer. Int. J. Cancer 2008, 122, 2767–2773. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Wen, X.; Song, Y.S.; Rhee, Y.Y.; Lee, T.H.; Cho, N.Y.; Han, S.W.; Kim, T.Y.; Kang, G.H. Associations and prognostic implications of Eastern Cooperative Oncology Group performance status and tumoral LINE-1 methylation status in stage III colon cancer patients. Clin. Epigenet. 2016, 8, 36. [Google Scholar] [CrossRef] [PubMed]
- Lou, Y.T.; Chen, C.W.; Fan, Y.C.; Chang, W.C.; Lu, C.Y.; Wu, I.C.; Hsu, W.H.; Huang, C.W.; Wang, J.Y. LINE-1 Methylation Status Correlates Significantly to Post-Therapeutic Recurrence in Stage III Colon Cancer Patients Receiving FOLFOX-4 Adjuvant Chemotherapy. PLoS ONE 2014, 10, e0123973. [Google Scholar] [CrossRef] [PubMed]
- De Maat, M.F.; van de Velde, C.J.; Benard, A.; Putter, H.; Morreau, H.; van Krieken, J.H.; Meershoek Klein-Kranenbarg, E.; de Graaf, E.J.; Tollenaar, R.A.; Hoon, D.S. Identification of a quantitative MINT locus methylation profile predicting local regional recurrence of rectal cancer. Clin. Cancer Res. 2010, 16, 2811–2818. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.P.; Tänzer, M.; Balluff, B.; Burgermeister, E.; Kretzschmar, A.K.; Hughes, D.J.; Tetzner, R.; Lofton-Day, C.; Rosenberg, R.; Reinacher-Schick, A.C.; et al. TFAP2E-DKK4 and chemoresistance in colorectal cancer. N. Engl. J. Med. 2012, 366, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Molinari, C.; Casadio, V.; Foca, F.; Zingaretti, C.; Giannini, M.; Avanzolini, A.; Lucci, E.; Saragoni, L.; Passardi, A.; Amadori, D.; et al. Gene methylation in rectal cancer: Predictive marker of response to chemoradiotherapy? J. Cell. Physiol. 2013, 228, 2343–2349. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Sun, Y.; Zhu, M.; Wang, Z.; Zhang, H.; Xin, Y.; Jiang, G.; Guo, X.; Zhang, Z.; Liu, Y. The role of plasma cell-free DNA detection in predicting preoperative chemoradiotherapy response in rectal cancer patients. Oncol. Rep. 2014, 31, 1466–1472. [Google Scholar] [CrossRef] [PubMed]
- Jo, P.; Jung, K.; Grade, M.; Conradi, L.C.; Wolff, H.A.; Kitz, J.; Becker, H.; Rüschoff, J.; Hartmann, A.; Beissbarth, T.; et al. CpG island methylator phenotype infers a poor disease-free survival in locally advanced rectal cancer. Surgery 2012, 151, 564–570. [Google Scholar] [CrossRef]
- Kohonen-Corish, M.R.; Tseung, J.; Chan, C.; Currey, N.; Dent, O.F.; Clarke, S.; Bokey, L.; Chapuis, P.H. KRAS mutations and CDKN2A promoter methylation show an interactive adverse effect on survival and predict recurrence of rectal cancer. Int. J. Cancer 2014, 134, 2820–2828. [Google Scholar] [CrossRef]
- Tsang, J.S.; Vencken, S.; Sharaf, O.; Leen, E.; Kay, E.W.; McNamara, D.A.; Deasy, J.; Mulligan, E.D. Global DNA methylation is altered by neoadjuvant chemoradiotherapy in rectal cancer and may predict response to treatment—A pilot study. Eur. J. Surg. Oncol. 2014, 40, 1459–1466. [Google Scholar] [CrossRef]
- Gaedcke, J.; Leha, A.; Claus, R.; Weichenhan, D.; Jung, K.; Kitz, J.; Grade, M.; Wolff, H.A.; Jo, P.; Doyen, J.; et al. Identification of a DNA methylation signature to predict disease-free survival in locally advanced rectal cancer. Oncotarget 2014, 5, 8123–8135. [Google Scholar] [CrossRef] [PubMed]
- Schetter, A.J.; Leung, S.Y.; Sohn, J.J.; Zanetti, K.A.; Bowman, E.D.; Yanaihara, N.; Yuen, S.T.; Chan, T.L.; Kwong, D.L.; Au, G.K.; et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA 2008, 299, 425–436. [Google Scholar] [CrossRef] [PubMed]
- Oue, N.; Anami, K.; Schetter, A.J.; Moehler, M.; Okayama, H.; Khan, M.A.; Bowman, E.D.; Mueller, A.; Schad, A.; Shimomura, M.; et al. High miR-21 expression from FFPE tissues is associated with poor survival and response to adjuvant chemotherapy in colon cancer. Int. J. Cancer 2014, 134, 1926–1934. [Google Scholar] [CrossRef] [PubMed]
- Coebergh van den Braak, R.R.J.; Sieuwerts, A.M.; Lalmahomed, Z.S.; Smid, M.; Wilting, S.M.; Bril, S.I.; Xiang, S.; van der Vlugt-Daane, M.; de Weerd, V.; van Galen, A.; et al. Confirmation of a metastasis-specific microRNA signature in primary colon cancer. Sci. Rep. 2018, 8, 5242. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Zhang, P.; Wang, F.; Zhang, H.; Yang, J.; Peng, J.; Liu, W.; Qin, H. miR-150 as a potential biomarker associated with prognosis and therapeutic outcome in colorectal cancer. Gut 2012, 61, 1447–1453. [Google Scholar] [CrossRef] [PubMed]
- Perez-Carbonell, L.; Sinicrope, F.A.; Alberts, S.R.; Oberg, A.L.; Balaguer, F.; Castells, A.; Boland, C.R.; Goel, A. MiR-320e is a novel prognostic biomarker in colorectal cancer. Br. J. Cancer 2015, 113, 83–90. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.X.; Song, W.; Chen, Z.H.; Wei, J.H.; Liao, Y.J.; Lei, J.; Hu, M.; Chen, G.Z.; Liao, B.; Lu, J.; et al. Prognostic and predictive value of a microRNA signature in stage II colon cancer: A microRNA expression analysis. Lancet Oncol. 2013, 14, 1295–1306. [Google Scholar] [CrossRef]
- To, K.K.; Leung, W.W.; Ng, S.S. Exploiting a novel miR-519c-HuR-ABCG2 regulatory pathway to overcome chemoresistance in colorectal cancer. Exp. Cell Res. 2015, 338, 222–231. [Google Scholar] [CrossRef]
- Diaz, T.; Tejero, R.; Moreno, I.; Ferrer, G.; Cordeiro, A.; Artells, R.; Navarro, A.; Hernandez, R.; Tapia, G.; Monzo, M. Role of miR-200 family members in survival of colorectal cancer patients treated with fluoropyrimidines. J. Surg. Oncol. 2014, 109, 676–683. [Google Scholar] [CrossRef]
- Li, S.; Gao, J.; Gu, J.; Yuan, J.; Hua, D.; Shen, L. MicroRNA-215 inhibits relapse of colorectal cancer patients following radical surgery. Med. Oncol. 2013, 30, 549. [Google Scholar] [CrossRef]
- Dou, R.; Deng, Y.; Huang, L.; Fu, S.; Tan, S.; Wang, L.; Lian, L.; Fang, L.; Fan, X.; Jin, G.; et al. Multi-microarray identifies lower AQP9 expression in adjuvant chemotherapy nonresponders with stage III colorectal cancer. Cancer Lett. 2013, 336, 106–113. [Google Scholar] [CrossRef] [PubMed]
- Conev, N.V.; Donev, I.S.; Konsoulova-Kirova, A.A.; Chervenkov, T.G.; Kashlov, J.K.; Ivanov, K.D. Serum expression levels of miR-17, miR-21, and miR-92 as potential biomarkers for recurrence after adjuvant chemotherapy in colon cancer patients. Biosci. Trends 2015, 9, 393–401. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Eng, C.; Shen, J.; Lu, Y.; Takata, Y.; Mehdizadeh, A.; Chang, G.J.; Rodriguez-Bigas, M.A.; Li, Y.; Chang, P.; et al. Serum exosomal miR-4772-3p is a predictor of tumor recurrence in stage II and III colon cancer. Oncotarget 2016, 7, 76250–76260. [Google Scholar] [CrossRef] [PubMed]
- Gaedcke, J.; Grade, M.; Camps, J.; Søkilde, R.; Kaczkowski, B.; Schetter, A.J.; Difilippantonio, M.J.; Harris, C.C.; Ghadimi, B.M.; Møller, S.; et al. The rectal cancer microRNAome—Microrna expression in rectal cancer and matched normal mucosa. Clin. Cancer Res. 2012, 18, 4919–4930. [Google Scholar] [CrossRef] [PubMed]
- Drebber, U.; Lay, M.; Wedemeyer, I.; Vallböhmer, D.; Bollschweiler, E.; Brabender, J.; Mönig, S.P.; Hölscher, A.H.; Dienes, H.P.; Odenthal, M. Altered levels of the onco-microRNA 21 and the tumor-supressor microRNAs 143 and 145 in advanced rectal cancer indicate successful neoadjuvant chemoradiotherapy. Int. J. Oncol. 2011, 39, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Svoboda, M.; Sana, J.; Fabian, P.; Kocakova, I.; Gombosova, J.; Nekvindova, J.; Radova, L.; Vyzula, R.; Slaby, O. MicroRNA expression profile associated with response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer patients. Radiat. Oncol. 2012, 7, 195. [Google Scholar] [CrossRef] [PubMed]
- Kheirelseid, E.A.; Miller, N.; Chang, K.H.; Curran, C.; Hennessey, E.; Sheehan, M.; Newell, J.; Lemetre, C.; Balls, G.; Kerin, M.J. miRNA expressions in rectal cancer as predictors of response to neoadjuvant chemoradiation therapy. Int. J. Colorectal Dis. 2013, 28, 247–260. [Google Scholar] [CrossRef]
- Della Vittoria Scarpati, G.; Falcetta, F.; Carlomagno, C.; Ubezio, P.; Marchini, S.; De Stefano, A.; Singh, V.K.; D’Incalci, M.; De Placido, S.; Pepe, S. A specific miRNA signature correlates with complete pathological response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Int. J. Radiat. Oncol. Biol. Phys. 2012, 83, 1113–1119. [Google Scholar] [CrossRef]
- Campayo, M.; Navarro, A.; Benítez, J.C.; Santasusagna, S.; Ferrer, C.; Monzó, M.; Cirera, L. miR-21, miR-99b and miR-375 combination as predictive response signature for preoperative chemoradiotherapy in rectal cancer. PLoS ONE 2018, 13, e0206542. [Google Scholar] [CrossRef]
- Lopes-Ramos, C.M.; Habr-Gama, A.; Quevedo, B.e.S.; Felício, N.M.; Bettoni, F.; Koyama, F.C.; Asprino, P.F.; Galante, P.A.; Gama-Rodrigues, J.; Camargo, A.A.; et al. Overexpression of miR-21-5p as a predictive marker for complete tumor regression to neoadjuvant chemoradiotherapy in rectal cancer patients. BMC Med. Genom. 2014, 7, 68. [Google Scholar] [CrossRef]
- Caramés, C.; Cristobal, I.; Moreno, V.; Marín, J.P.; González-Alonso, P.; Torrejón, B.; Minguez, P.; Leon, A.; Martín, J.I.; Hernández, R.; et al. MicroRNA-31 Emerges as a Predictive Biomarker of Pathological Response and Outcome in Locally Advanced Rectal Cancer. Int. J. Mol. Sci. 2016, 17, 878. [Google Scholar] [CrossRef] [PubMed]
- D’Angelo, E.; Zanon, C.; Sensi, F.; Digito, M.; Rugge, M.; Fassan, M.; Scarpa, M.; Pucciarelli, S.; Nitti, D.; Agostini, M. miR-194 as predictive biomarker of responsiveness to neoadjuvant chemoradiotherapy in patients with locally advanced rectal adenocarcinoma. J. Clin. Pathol. 2018, 71, 344–350. [Google Scholar] [CrossRef] [PubMed]
- Du, B.; Wang, X.; Wu, D.; Wang, T.; Yang, X.; Wang, J.; Shi, X.; Chen, L.; Zhang, W. MicroRNA expression profiles identify biomarkers for predicting the response to chemoradiotherapy in rectal cancer. Mol. Med. Rep. 2018, 18, 1909–1916. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Liu, L.; Zhou, N.; Shen, J.; Sun, Q.; Zhu, Y.; Chen, M. miR-519b-3p promotes responsiveness to preoperative chemoradiotherapy in rectal cancer patients by targeting ARID4B. Gene 2018, 655, 84–90. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Li, N.; Wang, X.; Ren, H.; Wang, W.; Wang, S.; Song, Y.; Liu, Y.; Li, Y.; Zhou, X.; et al. Circulating serum microRNA-345 correlates with unfavorable pathological response to preoperative chemoradiotherapy in locally advanced rectal cancer. Oncotarget 2016, 7, 64233–64243. [Google Scholar] [CrossRef] [PubMed]
- Menéndez, P.; Padilla, D.; Villarejo, P.; Palomino, T.; Nieto, P.; Menéndez, J.M.; Rodríguez-Montes, J.A. Prognostic implications of serum microRNA-21 in colorectal cancer. J. Surg. Oncol. 2013, 108, 369–373. [Google Scholar] [CrossRef]
- Hiyoshi, Y.; Akiyoshi, T.; Inoue, R.; Murofushi, K.; Yamamoto, N.; Fukunaga, Y.; Ueno, M.; Baba, H.; Mori, S.; Yamaguchi, T. Serum miR-143 levels predict the pathological response to neoadjuvant chemoradiotherapy in patients with locally advanced rectal cancer. Oncotarget 2017, 8, 79201–79211. [Google Scholar] [CrossRef]
- Toyota, M.; Ahuja, N.; Ohe-Toyota, M.; Herman, J.G.; Baylin, S.B.; Issa, J.P. CpG island methylator phenotype in colorectal cancer. Proc. Natl. Acad. Sci. USA 1999, 96, 8681–8686. [Google Scholar] [CrossRef]
- Berg, M.; Hagland, H.R.; Søreide, K. Comparison of CpG island methylator phenotype (CIMP) frequency in colon cancer using different probe- and gene-specific scoring alternatives on recommended multi-gene panels. PLoS ONE 2014, 9, e86657. [Google Scholar] [CrossRef]
- Sottoriva, A.; Kang, H.; Ma, Z.; Graham, T.A.; Salomon, M.P.; Zhao, J.; Marjoram, P.; Siegmund, K.; Press, M.F.; Shibata, D.; et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 2015, 47, 209–216. [Google Scholar] [CrossRef]
| First Author and Year | Tumor | Overall Population (Treated Patients) | Treatment | Samples | Markers | Methods | Response to Treatment | Other Findings |
|---|---|---|---|---|---|---|---|---|
| Nagasaka, T., 2003 [32] | CRC | 90 (50) | 5-FU | Tumor and normal tissues | MGMT | MS-PCR | In the unmethylated group, patients receiving adjuvant chemotherapy had greater risk of recurrence than those who did not receive chemotherapy | Patients with unmethylated MGMT were more likely to experience recurrence within 36 months than those with MGMT methylation |
| Chen, S.P., 2009 [33] | CRC | 117 (61) | 5-FU | Tumor tissues | APC and MGMT | MS-PCR | No interaction between adjuvant chemotherapy and methylation markers was evident | Patients with APC methylation had lower risk of death or CRC-related deaths. No significant association MGMT methylation and prognosis was evident |
| Ide, T., 2008 [35] | CRC | 94 (35) | 5-FU | Tumor and normal tissues | MLH1 | MS-PCR | Among patients receiving adjuvant chemotherapy, those with low hMLH1 mRNA expression levels had longer disease-free survival | MLH1 mRNA expression levels were significantly lower in CRC tissues with MLH1 methylation compared with unmethylated hMLH1 tissues |
| Sinicrope, F.A., 2015 [36] | Colon cancer | 2720 (2720) | mFOLFOX6 alone or with cetuximab | Tumor tissues | MLH1 | MS-PCR | Not examined | Two subtypes of colon cancer deficient in mismatch repair were identified, based on mismatch repair status and detection of BRAFV600E or mutations in KRAS. However, time of disease-free survival was similar between patients deficient in mismatch repair and those without BRAFV600E or KRAS mutations |
| Perez-Carbonell, L., 2014 [37] | CRC | 425 (425) | 5-FU | Tumor tissues | SEPT9, TWIST1, IGFBP3, GAS7, ALX4 and miR137 | Pyrosequencing | Patients with IGFBP3 hypermethylation did not benefit from adjuvant chemotherapy | Methylation levels of all genes analyzed were significantly higher in tumor tissues than in normal mucosa. IGFBP3 hypomethylation was an independent risk factor for poor disease-free survival |
| Heitzer, E., 2014 [38] | CRC | 143 (71) | 5-FU and leucovorin | Tumor tissues | PCDH10, SPARC, and UCHL1, | Real-time PCR | Patients receiving adjuvant therapy with no methylation of a combined set of three genes had longer overall survival than those with hypermethylation. In the surveillance group, unmethylated genes were associated with shorter disease-free survival and overall survival. | NA |
| Chang, S.Y., 2018 [43] | CRC | 151 (151) | NA | Tumor and normal tissues | NKX6.1 | MS-PCR | Patients receiving adjuvant chemotherapy with NKX6.1 methylation had shorter 5-year overall survival and disease-free survival than patients with no NKX6.1 methylation | NKX6.1 methylation was significantly higher in tumor tissues than in adjacent normal tissues |
| Pfütze, K., 2015 [47] | Colon cancer | 232 (98) | 5-FU | Tumor tissues | HYAL2 | MALDI-TOF mass spectrometry | In patients receiving adjuvant chemotherapy, low methylation levels were associated with better overall survival and progression-free survival. In patients no receiving adjuvant chemotherapy, on the contrary, HYAL2 hypomethylation seemed to be associated with worse overall survival | NA |
| First Author and Year | Tumor | Overall Population (Treated Patients) | Treatment | Samples | Markers | Methods | Response to Treatment | Other Findings |
|---|---|---|---|---|---|---|---|---|
| Van Rijnsoever, M., 2003 [48] | CRC | 206 (103) | 5-FU | Tumor tissues | CIMP (CDKN2A, MINT-2 and MDR1) | MS-PCR | CIMP+ patients receiving adjuvant therapy exhibited improved survival, while no long-term benefits were evident in CIMP- patients | CIMP+ was associated with worse prognosis in patients treated with surgery alone |
| Min, B.H., 2011 [49] | CRC | 245 (124) | 5-FU or capecitabine | Tumor tissues | CIMP (CACNA1G, IGF2, NEUROG1, RUNX3, and SOCS1) | MethyLight assay | CIMP+ patients receiving adjuvant therapy exhibited increased recurrence-free survival than patients treated with surgery alone, while no benefits from adjuvant therapy were evident in CIMP- or CIMP-low patients | CIMP-high was also associated with high frequencies of MGMT methylation and microsatellite instability |
| Shiovitz, S., 2014 [50] | Colon cancer | 615 (316/299) | 5-FU and leucovorin alone or with irinotecan | Tumor tissues | CIMP (CACNA1G, IGF2, NEUROG1, RUNX3, and SOCS1) | MethyLight assay | CIMP+ patients treated with the FOLFIRI protocol had better overall survival than those treated with 5-FU and leucovorin alone. However, benefits of this combination were not reported in patients with CIMP- tumors | CIMP+ had shorter overall survival than those with CIMP- tumors |
| Cohen, S.A., 2016 [51] | CRC | 292 (144/148) | mFOLFOX6 or XELOX | Tumor tissues | CIMP (CACNA1G, IGF2, NEUROG1, RUNX3, and SOCS1) | MethyLight Assay | Not examined | CIMP+ status did not affect overall survival, but it was associated with right-side tumor location and BRAF mutations |
| Jover, R., 2011 [52] | CRC | 302 (93) | 5-FU | Tumor tissues | CIMP (CACNA1G, SOCS1, RUNX3, NEUROG1, and MLH1) | Pyrosequencing | Adjuvant therapy improved disease-free survival in CIMP- but not in CIMP+ patients. In patients not receiving adjuvant therapy, CIMP status was the only predictor of disease-free survival | NA |
| Han, S.W., 2013 [53] | CRC | 322 (322) | FOLFOX | Tumor and normal tissues | CIMP (CACNA1G, CDKN2A, CRABP1, IGF2, MLH1, NEUROG1, RUNX3 and SOCS1) | MethyLight assay | Not examined | CIMP status was associated with microsatellite instability but not with disease-free survival. Methylation of NEUROG1 and CDKN2A increased the risk of recurrence |
| Bae, J.M., 2017 [54] | CRC | 1370 (531/365/49) | 5-FU and leucovorin, FOLFOX, or FOLFIRI | Tumor tissues | CIMP (CACNA1G, CDKN2A, CRABP1, IGF2, MLH1, NEUROG1, RUNX3 and SOCS1) | MethyLight assay | Among patients treated FOLFOX or XELOX protocols, CIMP+1 tumors reported lower cancer-specific survival and relapse-free survival than those with CIMP- and CIMP+2 tumors | Among patients treated with surgery alone, those with CIMP+1 tumors reported lower cancer-specific survival and relapse-free survival than those with CIMP- and CIMP+2 tumors |
| First Author and Year | Tumor | Overall Population (Treated Patients) | Treatment | Samples | Markers | Methods | Response to Treatment | Other Findings |
|---|---|---|---|---|---|---|---|---|
| Kawakami, K., 2011 [71] | CRC | 155 (94) | 5-FU | Tumor tissues | LINE-1 | MS-PCR and Methylight assay | LINE-1 hypomethylation conferred a survival benefit in patients receiving adjuvant therapy. Benefits from adjuvant chemotherapy were not evident in patients with LINE-1 hypermethylation | In patients treated with surgery alone, LINE-1 hypomethylation was associated with worse prognosis |
| Chen, D., 2016 [73] | CRC | 336 (NA) | FOLFOX-4 or mFOLFOX-6 | Tumor tissues | LINE-1 | Pyrosequencing | LINE-1 methylation was associated with clinicopathological features and recurrence-free survival in CRC patients receiving adjuvant therapy based on the FOLFOX protocol | NA |
| Lou, Y.T., 2015 [74] | Colon cancer | 129 (129) | FOLFOX-4 | Tumor tissues | LINE-1 | Pyrosequencing | LINE-1 methylation level was lower in patients receiving adjuvant therapy with post-therapeutic recurrence than in those without recurrence. LINE-1 hypomethylation as an independent risk factor of post-therapeutic recurrence | Patients with LINE-1 hypomethylation had reduced disease free survival in the whole cohort and in those with post-therapeutic recurrence after 6 months |
| First Author and Year | Tumor | Overall Population (Treated Patients) | Treatment | Samples | Markers | Methods | Response to Treatment | Other Findings |
|---|---|---|---|---|---|---|---|---|
| De Maat, M.F., 2010 [75] | Rectal cancer | 251 (251) | Chemoradiation | Tumor tissues | MINT loci | Absolute quantitative assessmen tof methylated alleles | Not examined | MINT3 hypermethylation and MINT17 hypomethylation reduced the risk of recurrence |
| Ebert, M.P., 2012 [76] | Rectal cancer | 220 (NA) | Chemoradiation | Tumor tissues | TFAP2E | MethyLight assay | TFAP2E hypermethylation was associated with non-response to neoadjuvant chemoradiation. The odds of adequate response were higher in patients with TFAP2E hypomethylation compared with the whole cohort | NA |
| Molinari, C., 2013 [77] | Rectal cancer | 74 (74) | Chemoradiation | Tumor tissues | 24 genes | Methylation-specific multiplex ligation-dependent probeamplification | In patients receiving neoadjuvant therapy, TIMP3 methylation status was significantly different across four tumor regression grade classes | Compared with adjacent normal tissues, tumor samples exhibited hypermethylation of ESR1, CDH13, RARB, IGSF4, and APC genes |
| Sun, W., 2013 [78] | Rectal cancer | 34 (34) | Chemoradiation | Serum | MGMT | MS-PCR | Serum MGMT hypermethylation was associated with improved response to treatment and higher regression compared with MGMT hypomethylation | NA |
| First Author and Year | Tumor | Overall Population (Treated Patients) | Treatment | Samples | Markers | Methods | Response to Treatment | Other Findings |
|---|---|---|---|---|---|---|---|---|
| Jo, P., 2012 [79] | Rectal cancer | 150 (150) | Chemoradiation | Tumor tissues | CIMP (CACNA1G, IGF2, NEURO1G, RUNX3, and SOCS1) | MS-PCR | No association with response to neoadjuvant chemoradiation was evident | No association of CIMP status with clinicopathological features, and KRAS or BRAF mutations was evident. CIMP+ patients exhibited worse 3-year and 5-year disease-free survival |
| Kohonen-Corish, M.R., 2013 [80] | Rectal cancer | 381 (18) | Chemoradiation | Tumor tissues | CIMP (CACNA1G, IGF2, NEURO1G, RUNX3, and SOCS1) and CDKN2A | MethyLight assay | Not examined | No association of CIMP+ status with overall survival was evident. Combination of CDKN2A methylation and KRAS mutations independently predicted the risk of recurrence, thereby reducing overall and cancer-specific survival |
| First Author and Year | Tumor | Overall Population (Treated Patients) | Treatment | Samples | Markers | Methods | Response to Treatment | Other Findings |
|---|---|---|---|---|---|---|---|---|
| Tsang, J.S., 2014 [81] | Rectal cancer | 53 (53) | Chemoradiation | Tumor tissues | Global methylation | Immunohistochemical staining and image analysis of staining | A significant reduction of global DNA methylation in the majority of patients. Global methylation of pre-treatment specimens was significantly correlated with tumor stage and regression | NA |
| Gaedcke, J., 2014 [82] | Rectal cancer | 185 (185) | Chemoradiation | Tumor tissues | Whole genome methylation | Whole genome methylation CpG island array | Not examined | In a discovery cohort, 20 highly discriminative DMRs were identified. In two additional validation cohorts, 10 DMRs that allowed the discrimination of patients with different prognosis were confirmed |
| First Author and Year | Tumor | Overall Population (Treated Patients) | Treatment | Samples | Markers | Methods | Response to Treatment | Other Findings |
|---|---|---|---|---|---|---|---|---|
| Schetter, A.J., 2008 [83] | Colon cancer | 197 (72) | 5-FU | Tumor and normal tissues | miR-20a, miR-203, miR-21, miR-106a, miR-181b | Microarray and Real-time PCR | miR-20a, miR-21, miR-106a, miR-181b, and miR-203 were enriched in tumor tissues. MiR-21 upregulation was associated with poor survival and poor therapeutic response in both cohorts | 37 miRNAs were differentially expressed between tumor and normal tissues |
| Oue, N., 2014 [84] | Colon cancer | 301 (84) | 5-FU | Tumor tissues | miR-21 | Real-time PCR | Patients with miR-21 upregulation did not benefit from the adjuvant chemotherapy | Upregulation of miR-21 was associated with poor survival independent of clinicopathological features |
| Coebergh van den Braak, R.R.J., 2018 [85] | Colon cancer | 232 (77) | NA | Tumor and normal tissues | let-7i, miR-10b and miR-30b | Real-time PCR | The 2-miRNA signature (let-7i and miR-10b) had no prognostic value in stage III patients receiving adjuvant therapy | The 2-miRNA signature (let-7i and miR-10b) predicted hepatic recurrence in stage I-II patients, while the combination with miR-30b also predicted distant metastasis |
| Ma, Y., 2011 [86] | CRC | 424 (NA) | NA | Tumor and normal tissues | miR-150 | Microarray and Real-time PCR | Low miR-150 expression levels were associated with reduced overall survival and worse response to treatment | MiR-150 was downregulated in tumors compared with adjacent normal tissues |
| Perez-Carbonell, L., 2015 [87] | CRC | Discovery cohort: 100 (NA) Validation cohort: 237 (167) | FOLFOX, 5-FU, or 5-FU and oxaliplatin | Tumor tissues | miR-320e | Microarray and Real-time PCR | Not examined | MiR-320e was upregulated in patients with recurrence compared with those without recurrence. MiR-320e upregulation was associated with poor overall and disease-free survival and overall survival |
| Zhang, J.X., 2013 [88] | CRC | Training set: 138 (NA) Internal set: 137 (NA) Validation set: 360 (NA) | 5-FU | Tumor tissues | miR-20a-5p, miR-21-5p, miR-103a-3p, miR-106a-5p, miR-143-5p, and miR-215 | Microarray and Real-time PCR | Based on a 6-miRNA-based classifier, patients with high risk of disease progression exhibited a better response to 5-FU based adjuvant chemotherapy | 35 miRNAs were differentially expressed between tumor and normal tissues. Based on a 6-miRNA-based classifier, patients were classified according to their risk of disease progression. Five-year disease-free survival was higher in low risk patients compare with those at high risk |
| To, K.K., 2015 [89] | Colon cancer | 26 (26) | NA | Tumor tissues | miR-519c | Real-time PCR | Most of CRC samples from non-responders exhibited miR-519c downregulation | MiR-519c levels were positively correlated with ABCG2 expression |
| Diaz, T., 2014 [90] | CRC | 127 (56/24) | 5-FU or XELOX | Tumor and normal tissues | miR-200 family, miR-141, miR-429 | Real-time PCR | Among patients receiving 5-FU based adjuvant therapy, upregulation of miR-200a, miR-200c, miR-141, or miR-429 was positively associated with overall and disease-free survival | Upregulation of miR-200a, miR-200c and miR-429 was associated with better survival |
| Li, S., 2013 [91] | CRC | 125 (91) | 5-FU | Tumor tissues | miR-215 | Real-time PCR | MiR-215 downregulation was associated with worse response to 5-FU adjuvant therapy | Patients with recurrence exhibited lower miR-215 expression level than those without recurrence |
| Dou, R., 2013 | CRC | 35 (35) | FOLFOX6 | Tumor tissues | 17 miRNAs | Microarray and Real-time PCR | 17 miRNAs were differentially expressed between responders and non-responders | Three genes (AQP9, SATB2, and WIF1) were downregulated by the differentially expressed miRNAs |
| Conev, N.V., 2015 [93] | Colon cancer | 37 (37) | 5-FU | Serum | miR-17, miR-21, miR-29a, and miR-92 | Real-time PCR | Not examined | MiR-17, miR-21 and miR-92 were upregulated in patients with recurrence |
| Liu, C., 2016 [94] | Colon cancer | 84 (66) | FOLFOX | Serum | miR-4772-3p | RNA-seq and Real-time PCR | Downregulation of miR-4772-3p was associated with an increased risk of recurrence. This evidence was confirmed in patients receiving adjuvant therapy | 145 miRNAs were differentially expressed between patients with or without recurrence |
| First Author and Year | Tumor | Overall Population (Treated Patients) | Treatment | Samples | Markers | Methods | Response to Treatment | Other Findings |
|---|---|---|---|---|---|---|---|---|
| Gaedcke, J., 2012 [95] | Rectal cancer | 57 (57) | Chemoradiation | Tumor tissues | miR-135b | Microarray | Not examined | MiR-492, miR-542-5p, miR-584, miR-483-5p, miR-144, miR-2110, miR-652, miR-375, miR-147b, miR-148a, miR-190, miR-26a/b, and miR-338-3p were differentially expressed between rectal and colon cancer tissues. MiR-135b expression correlated with tumor regression grade, disease-free survival and cancer-specific survival |
| Drebber, U., 2011 [96] | Rectal cancer | 40 (40) | Chemoradiation | Tumor tissues | miR-21, miR-143 and miR-145 | Real-time PCR | Compared to pre-treatment tissues, miR-21 was downregulated while miR-143 and miR-145 were upregulated in post-treatment tissues. MiR-145 downregulation was associated with worse response to neoadjuvant therapy | MiR-21 was upregulated in tumor tissue than in adjacent normal mucosa |
| Svoboda, M., 2012 [97] | Rectal cancer | 20 (20) | Chemoradiation | Tumor tissues | miR-215, miR-190b, miR-29b-2, let-7e, miR-196b, miR-450a, miR-450b-5p and miR-99a | Microarray and Real-time PCR | Eight miRNAs were differentially expressed between responders and non-responders. MiR-215, miR-190b and miR-29b-2 were upregulated in non-responders, while let-7e, miR-196b, miR-450a, miR-450b-5p and miR-99a were upregulated in responders. This 8-miRNA signature allowed to correctly classify 90% of non-responders | NA |
| Kheirelseid, E.A., 2012 [98] | Rectal cancer | 12 (12) | Chemoradiation | Tumor tissues | miR-16, miR-590-5p, miR-153, miR-519c-3p, and miR-561 | Microarray and Real-time PCR | A 3-miRNA signature (miR-16, miR-590-5p and miR-153) predicted incomplete response to neoadjuvant therapy. A 2-miRNA signature (miR-519c-3p and miR-561) predicted poor response to neoadjuvant therapy with an accuracy of 100% | NA |
| Della Vittoria Scarpati, G., 2012 [99] | Rectal cancer | 38 (38) | Chemoradiation | Tumor tissues | miR-1183, miR-483-5p, miR-622, miR-125a-3p, miR-1224-5p, miR-188-5p, miR-1471, miR-671-5p, miR-1909, miR-630, miR-765, miR-1274b, miR-720 | Microarray and Real-time PCR | 14 miRNAs were differentially expressed in patients with complete response. 11 miRNAs were upregulated (miR-1183, miR-483-5p, miR-622, miR-125a-3p, miR-1224-5p, miR-188-5p, miR-1471, miR-671-5p, miR-1909, miR-630, miR-765), while 2 miRNAs were downregulated (miR-1274b, miR-720). MiR-622 and miR-630 showed 100% sensitivity and specificity in discriminating patients with complete response | NA |
| Campayo, M., 2018 [100] | Rectal cancer | 96 (96) | Chemoradiation | Tumor tissues | 377 miRNAs including miR-21, miR-99b and miR-375 | Microarray and Real-time PCR | 8 miRNAs (let-7b, let-7e, miR-21, miR-99b, miR-183, miR-328, miR-375 and miR-483-5p) were differentially expressed across different tumor regression grades. The combination of miR-21, miR-99b and miR-375 allowed to discriminate patients with complete response to neoadjuvant therapy | MiR-21, miR-99b and miR-375 were associated with disease-free and overall survival. |
| Lopes-Ramos, C.M., 2014 [101] | Rectal cancer | Training cohort: 27 (27) Validation cohort: 16 (16) | Chemoradiation | Tumor tissues | miR-21-5p | RNA-seq and Real-time PCR | In the training set, four miRNAs were differentially expressed between complete and incomplete responders. In the validation set, miR-21-5p was upregulated in complete responders. MiR-21-5p expression level allowed the prediction of complete response to neoadjuvant therapy with a sensitivity of 78% and a specificity of 86% | NA |
| Caramés, C., 2016 [102] | Rectal cancer | 78 (78) | Chemoradiation | Tumor tissues | mir-31 | Real-time PCR | MiR-31 expression level were higher in patients with complete response than in those with poor response | MiR-31 upregulation was associated with poor pathological response and worse overall survival |
| D’Angelo, E., 2017 [103] | Rectal cancer | 38 (38) | Chemoradiation | Tumor tissues | miR-194 | Real-time PCR | MiR-194 was overexpressed in patients who responded to neoadjuvant chemoradiation | NA |
| Du, B., 2018 [104] | Rectal cancer | 38 (38) | Chemoradiation | Tumor tissues | 41 miRNAs | Microarray | 41 miRNAs were differentially expressed between complete and incomplete responders to neoadjuvant therapy. MiR‑548c‑5p/miR‑548d‑5p and miR‑663a regulated genes associate with rectal cancer, thereby modulating the complete response to neoadjuvant chemoradiation | NA |
| Luo, J., 2018 [105] | Rectal cancer | 55 (55) | Chemoradiation | Tumor tissues | miR-519b-3p | Real-time PCR | In patients receiving neoadjuvant therapy, miR-519b-3p expression was positively correlated with response to treatment. A functional analysis suggested that miR-519b-3p was directly involved in response to neoadjuvant chemoradiation in an ARID4B-dependent way | NA |
| Yu, J., 2016 [106] | Rectal cancer | 149 (149) | Chemoradiation | Tumor tissues and serum | miR-345 | Microarray and Real-time PCR | MiR-345 upregulation was associated with a worse response to treatment either in tissue or serum | MiR-345 downregulation in serum was associated with better recurrence-free survival |
| Menéndez, P., 2013 [107] | Rectal cancer | 28 (28) | Chemoradiation | Serum | miR-21 | Real-time PCR | Not examined | Serum miR-21 downregulation was associated with high risk of recurrence and death. MiR-21 expression was an independent predictor of overall survival |
| Hiyoshi, Y., 2017 [108] | Rectal cancer | 94 (94) | Chemoradiation | Serum | let-7b, miR-15b, miR-20a, miR-21, miR-29a, miR-92a, miR-122, miR-125b, miR-141, miR-143, miR-145, miR-155, miR-200c, miR-221, miR-345, miR-423, miR-425, miR-1246 | Real-time PCR | Serum miR-143 expression was higher in responders than in non-responders. Serum miR-143 expression was an independent predictor of pathological response | NA |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barchitta, M.; Maugeri, A.; Li Destri, G.; Basile, G.; Agodi, A. Epigenetic Biomarkers in Colorectal Cancer Patients Receiving Adjuvant or Neoadjuvant Therapy: A Systematic Review of Epidemiological Studies. Int. J. Mol. Sci. 2019, 20, 3842. https://doi.org/10.3390/ijms20153842
Barchitta M, Maugeri A, Li Destri G, Basile G, Agodi A. Epigenetic Biomarkers in Colorectal Cancer Patients Receiving Adjuvant or Neoadjuvant Therapy: A Systematic Review of Epidemiological Studies. International Journal of Molecular Sciences. 2019; 20(15):3842. https://doi.org/10.3390/ijms20153842
Chicago/Turabian StyleBarchitta, Martina, Andrea Maugeri, Giovanni Li Destri, Guido Basile, and Antonella Agodi. 2019. "Epigenetic Biomarkers in Colorectal Cancer Patients Receiving Adjuvant or Neoadjuvant Therapy: A Systematic Review of Epidemiological Studies" International Journal of Molecular Sciences 20, no. 15: 3842. https://doi.org/10.3390/ijms20153842
APA StyleBarchitta, M., Maugeri, A., Li Destri, G., Basile, G., & Agodi, A. (2019). Epigenetic Biomarkers in Colorectal Cancer Patients Receiving Adjuvant or Neoadjuvant Therapy: A Systematic Review of Epidemiological Studies. International Journal of Molecular Sciences, 20(15), 3842. https://doi.org/10.3390/ijms20153842







