Tobacco Cutworm (Spodoptera Litura) Larvae Silenced in the NADPH-Cytochrome P450 Reductase Gene Show Increased Susceptibility to Phoxim
Abstract
1. Introduction
2. Results
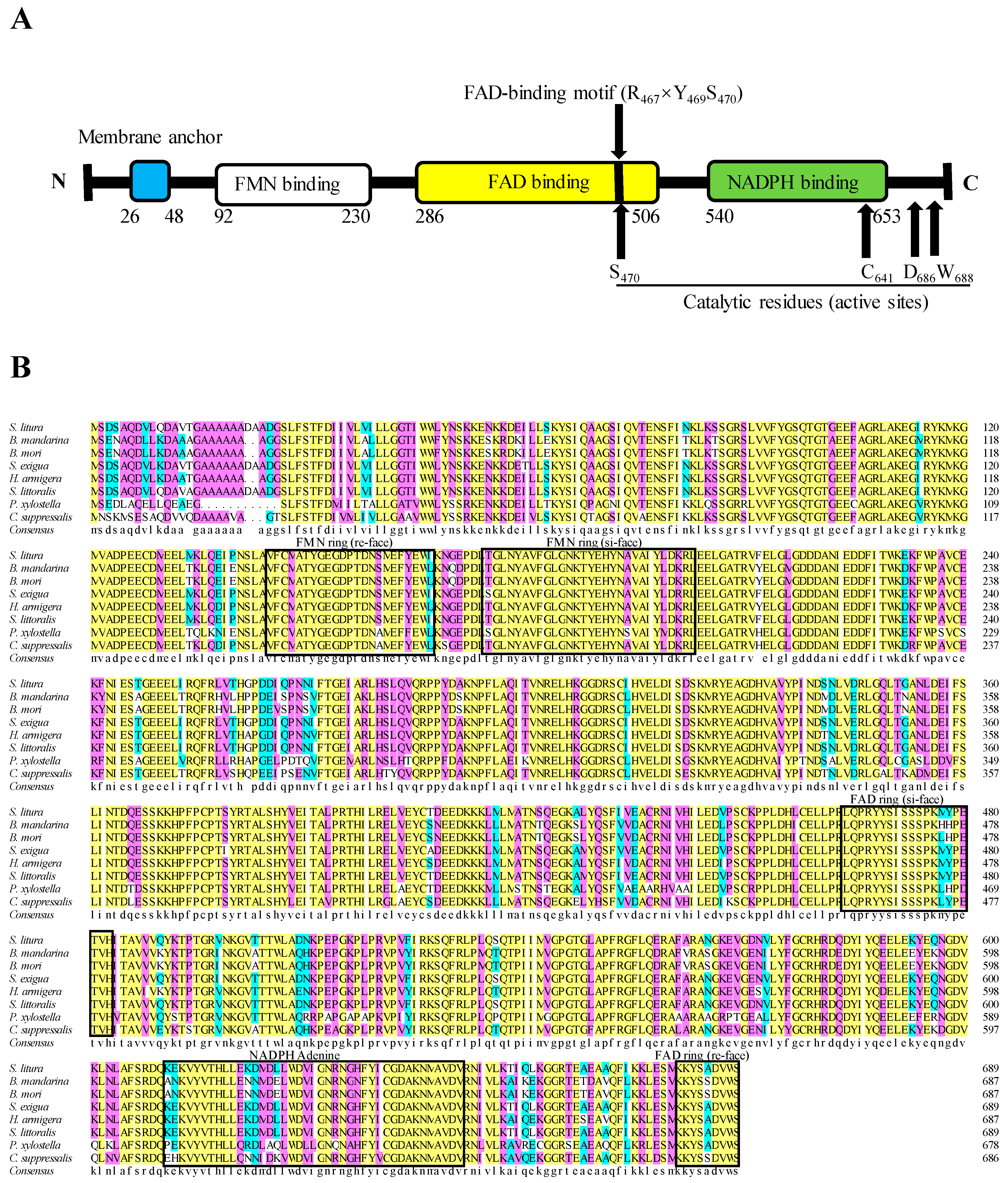
2.1. Cloning and Sequence Analysis of SlCPR
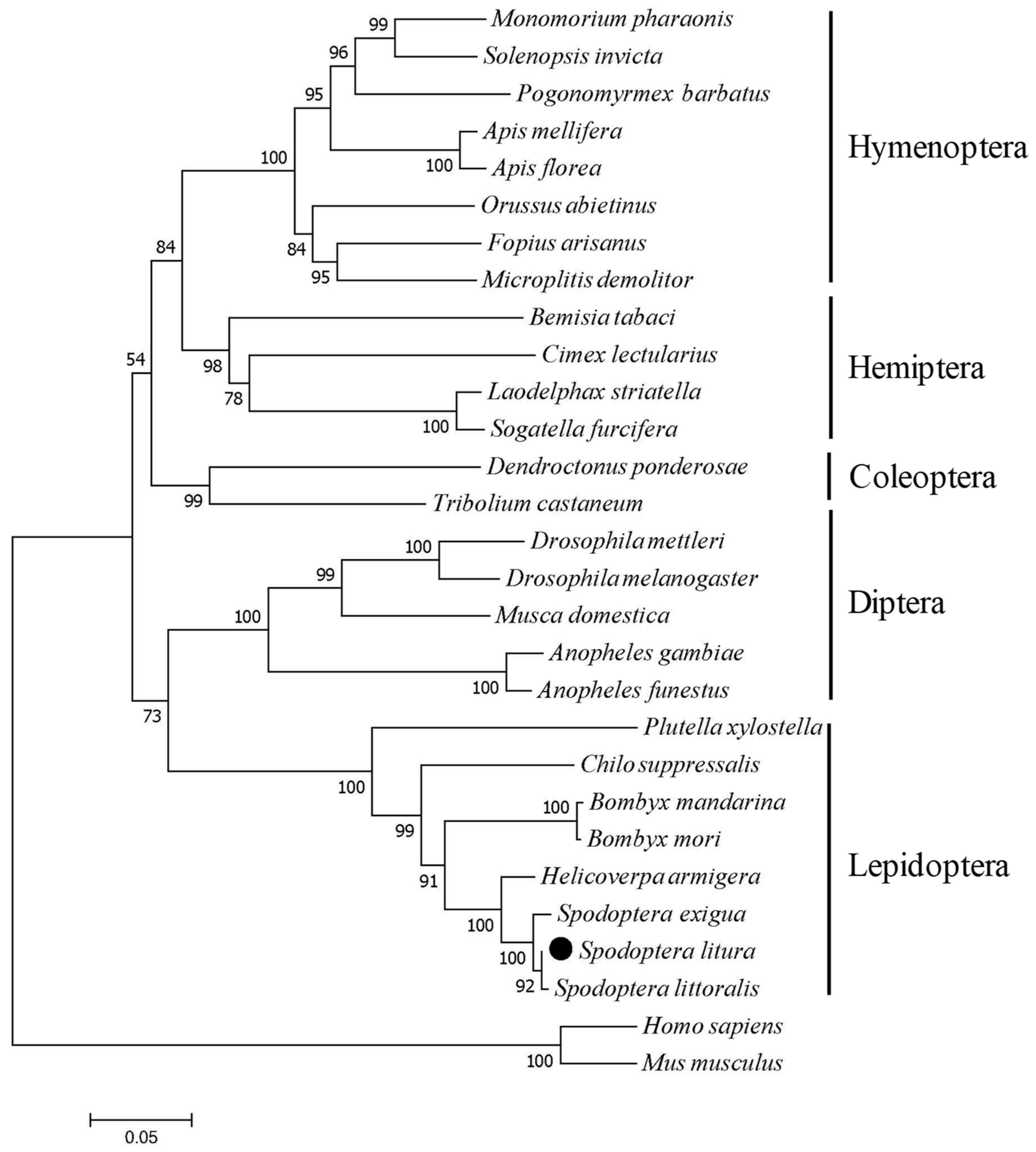
2.2. Phylogenetic Relation Between SlCPR and Other CPRs
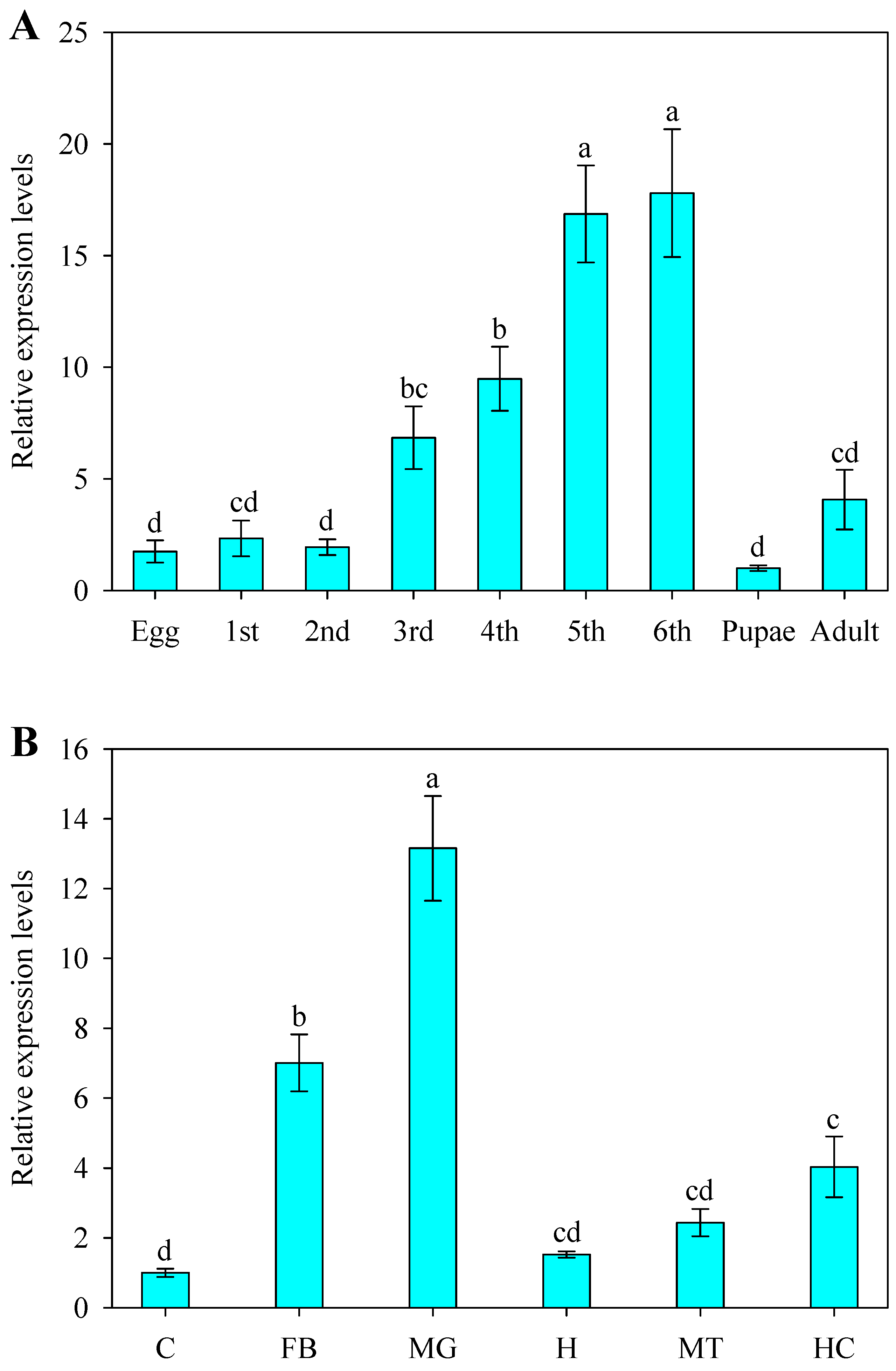
2.3. Developmental and Spatial Expression Patterns of SlCPR
2.4. Expression Response of SlCPR in Larvae Exposed to Phoxim
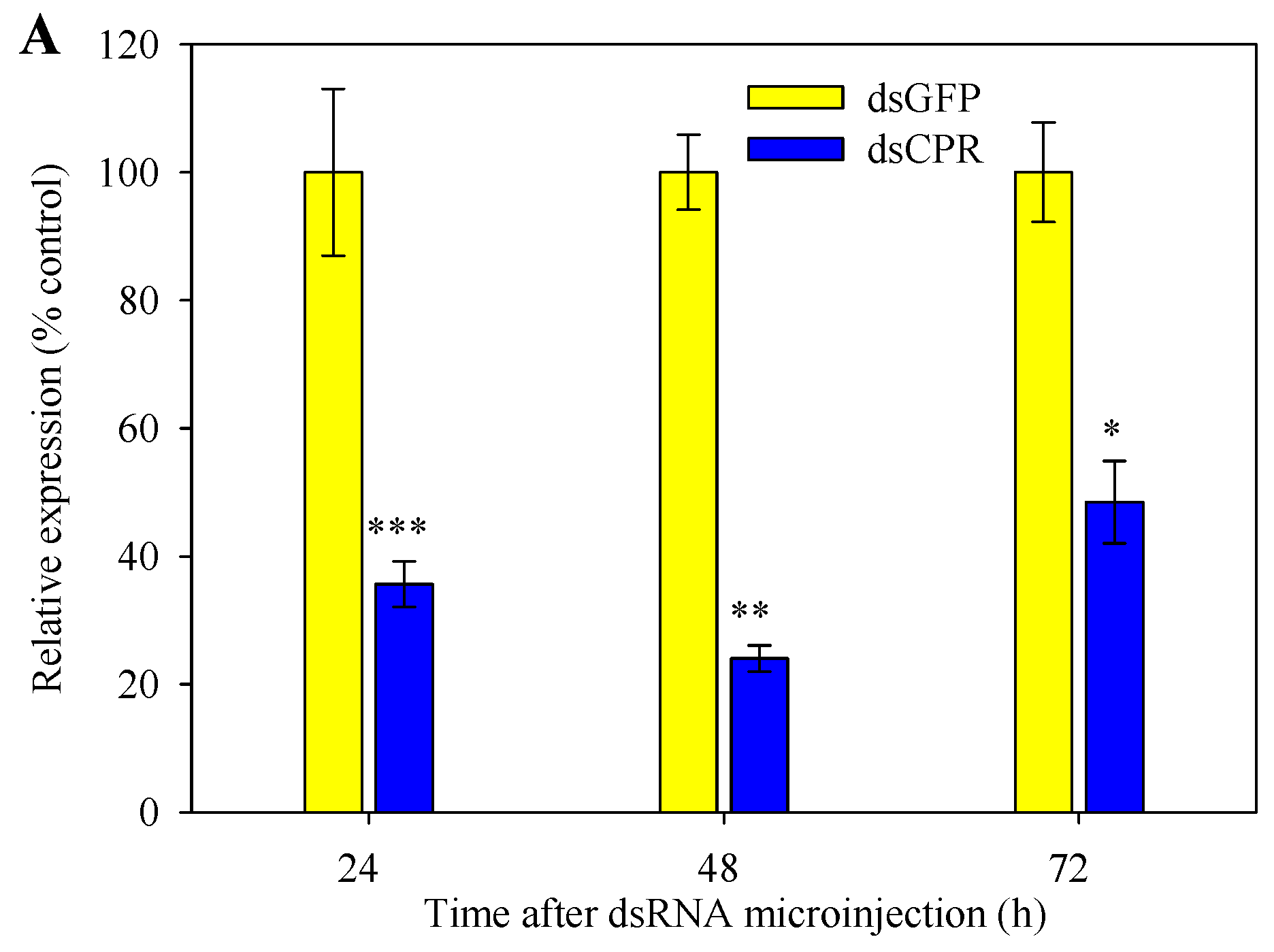
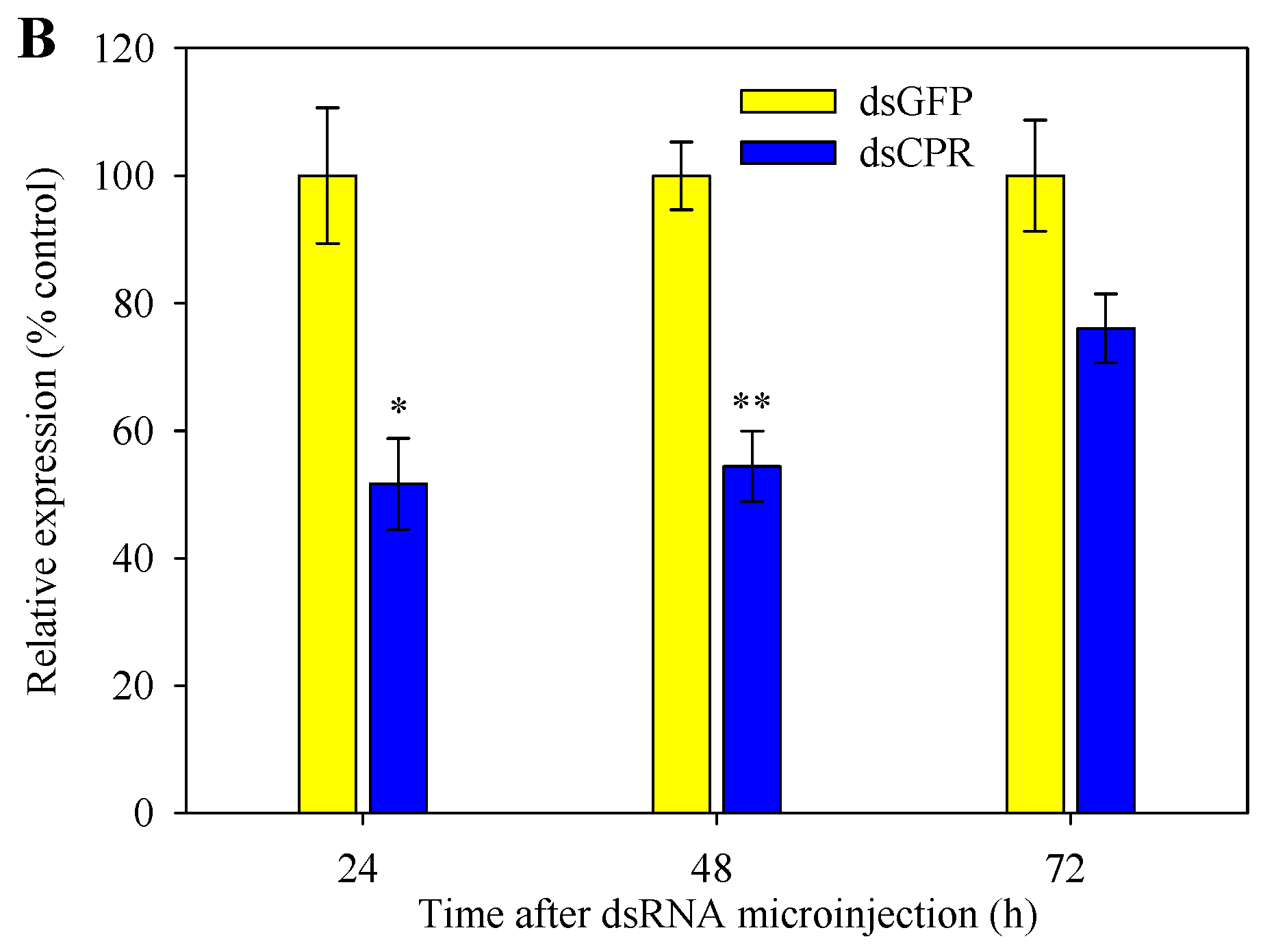
2.5. Silencing of SlCPR by RNAi
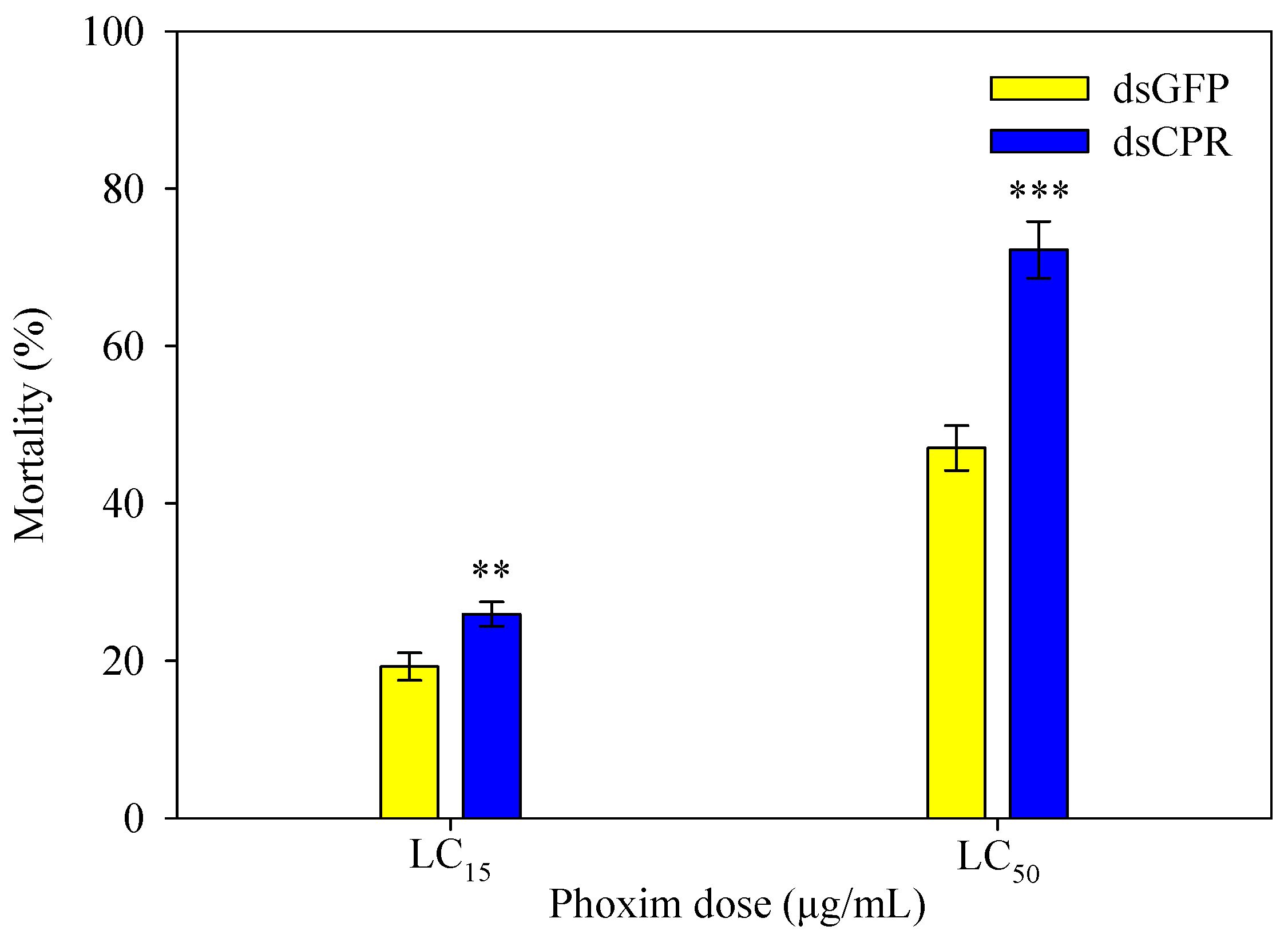
2.6. SlCPR-Silenced Larvae Show Increased Susceptibility to Phoxim
3. Discussion
4. Material and Methods
4.1. Insects
4.2. RNA Extraction and cDNA Synthesis
4.3. Cloning of SlCPR
4.4. Bioinformatic Analyses
4.5. SlCPR Expression Analysis
4.6. Analysis of SlCPR Expression in Larvae Exposed to Phoxim
4.7. Silencing of SlCPR by RNAi
4.8. Bioassays with Phoxim after RNAi
4.9. Data Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Ahmad, M.; Sayyed, A.H.; Saleem, M.A.; Ahmad, M. Evidence for field evolved resistance to newer insecticides in Spodoptera litura (Lepidoptera: Noctuidae) from Pakistan. Crop Prot. 2008, 27, 1367–1372. [Google Scholar] [CrossRef]
- Huang, S.; Han, Z. Mechanisms for multiple resistances in field populations of common cutworm, Spodoptera litura (Fabricius) in China. Pestic. Biochem. Physiol. 2007, 87, 14–22. [Google Scholar] [CrossRef]
- Ahmad, M.; Ahmad, M.; Saleem, M.A.; Aslam, M.; Sayyed, A.H. Resistance to selected organochlorin, organophosphate, carbamate and pyrethroid, in Spodoptera litura (Lepidoptera: Noctuidae) from Pakistan. J. Econ. Entomol. 2008, 101, 1667–1675. [Google Scholar]
- Feyereisen, R. Arthropod CYPomes illustrate the tempo and mode in P450 evolution. Biochim. Biophys. Acta Proteins Proteomics 2011, 1814, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Feyereisen, R. Insect P450 inhibitors and insecticides: Challenges and opportunities. Pest Manag. Sci. 2015, 71, 793–800. [Google Scholar] [CrossRef]
- Wang, R.L.; Staehelin, C.; Xia, Q.Q.; Su, Y.J.; Zeng, R.S. Identification and characterization of CYP9A40 from the tobacco cutworm moth (Spodoptera litura), a cytochrome P450 gene induced by plant allelochemicals and insecticides. Int. J. Mol. Sci. 2015, 16, 22606–22620. [Google Scholar] [CrossRef]
- Wang, R.L.; Zhu-Salzman, K.; Baerson, S.R.; Xin, X.W.; Li, J.; Su, Y.J.; Zeng, R.S. Identification of a novel cytochrome P450 CYP321B1 gene from tobacco cutworm (Spodoptera litura) and RNA interference to evaluate its role in commonly used insecticides. Insect Sci. 2017, 24, 235–247. [Google Scholar] [CrossRef]
- Li, F.; Ni, M.; Zhang, H.; Wang, B.; Xu, K.; Tian, J.; Hu, J.; Shen, W.; Li, B. Expression profile analysis of silkworm P450 family genes after phoxim induction. Pestic. Biochem. Physiol. 2015, 122, 103–109. [Google Scholar] [CrossRef]
- Porter, T.D.; Kasper, C.B. NADPH-cytochrome P-450 oxidoreductase: Flavin mononucleotide and flavin adenine dinucleotide domains evolved from different flavoproteins. Biochem. 1986, 25, 1682–1687. [Google Scholar] [CrossRef]
- Waskell, L.; Kim, J.J.P. Electron transfer partners of cytochrome P450. In Cytochrome P450: Structure, Mechanism, and Biochemistry, 4th ed.; Springer: Cham, Germany, 2015; pp. 33–68. [Google Scholar]
- Vermilion, J.L.; Ballou, D.P.; Massey, V.; Coon, M.J. Separate roles for FMN and FAD in catalysis by liver microsomal NADPH-cytochrome P-450 reductase. J. Biol. Chem. 1981, 256, 266–277. [Google Scholar]
- Zhang, Y.; Wang, Y.; Wang, L.; Yao, J.; Guo, H.; Fang, J. Knockdown of NADPH-cytochrome P450 reductase results in reduced resistance to buprofezin in the small brown planthopper, Laodelphax striatellus (fallén). Pestic. Biochem. Physiol. 2016, 127, 21–27. [Google Scholar] [CrossRef]
- Cheng, T.; Wu, J.; Wu, Y.; Chilukuri, R.V.; Huang, L.; Yamamoto, K.; Feng, L.; Li, W.; Chen, Z.; Guo, H.; et al. Genomic adaptation to polyphagy and insecticides in a major East Asian noctuid pest. Nat. Ecol. Evol. 2017, 1, 1747–1756. [Google Scholar] [CrossRef] [PubMed]
- Pottier, M.A.; Bozzolan, F.; Chertemps, T.; Jacquin-Joly, E.; Lalouette, L.; Siaussat, D.; Maïbèche-Coisne, M. Cytochrome P450s and cytochrome P450 reductase in the olfactory organ of the cotton leafworm Spodoptera littoralis. Insect Mol. Biol. 2012, 21, 568–580. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F.; Sams, S.; Moural, T.; Haynes, K.F.; Potter, M.F.; Palli, S.R. RNA interference of NADPH-cytochrome P450 reductase results in reduced insecticide resistance in the bed bug, Cimex lectularius. PLoS ONE 2012, 7, e31037. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Tang, T.; Feng, X.; Qiu, L. Cloning and characterisation of NADPH-dependent cytochrome P450 reductase gene in the cotton bollworm, Helicoverpa armigera. Pest Manag. Sci. 2014, 70, 130–139. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Liang, Q.M.; Zhou, W.W.; Jiang, Y.D.; Zhu, Q.Z.; Yu, H.; Zhang, C.X.; Gurr, G.M.; Zhu, Z.R. RNA interference of NADPH-cytochrome P450 reductase of the rice brown planthopper, Nilaparvata lugens, increases susceptibility to insecticides. Pest Manag. Sci. 2015, 71, 32–39. [Google Scholar] [CrossRef]
- Huang, Y.; Lu, X.P.; Wang, L.L.; Wei, D.; Feng, Z.J.; Zhang, Q.; Xiao, L.F.; Dou, W.; Wang, J.J. Functional characterization of NADPH-cytochrome P450 reductase from Bactrocera dorsalis: Possible involvement in susceptibility to malathion. Sci. Rep. 2015, 5, 18394. [Google Scholar] [CrossRef]
- Zhao, C.; Feng, X.; Tang, T.; Qiu, L.; Swale, D. Isolation and expression analysis of CYP9A11 and cytochrome P450 reductase gene in the beet armyworm (Lepidoptera: Noctuidae). J. Insect Sci. 2015, 15, 122. [Google Scholar] [CrossRef][Green Version]
- Zhang, Y.X.; Li, S.G.; Rao, X.J.; Liu, S. Molecular characterization of a NADPH–cytochrome P450 reductase gene from the rice leaffolder, Cnaphalocrocis medinalis (Lepidoptera: Pyralidae). Appl. Entomol. Zool. 2018, 53, 19–27. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, J.; Liu, J.; Li, Y.; Liu, X.; Wu, H.; Ma, E.; Zhang, J. Knockdown of NADPH-cytochrome P450 reductase increases the susceptibility to carbaryl in the migratory locust, Locusta migratoria. Chemosphere 2017, 188, 517–524. [Google Scholar] [CrossRef]
- Jing, T.X.; Tan, Y.; Ding, B.Y.; Dou, W.; Wei, D.D.; Wang, J.J. NADPH-cytochrome P450 reductase mediates the resistance of Aphis (Toxoptera) citricidus (Kirkaldy) to abamectin. Front. Physiol. 2018, 9, 986. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zhang, Y. Identification and characterization of NADPH-dependent cytochrome P450 reductase gene and cytochrome b5 gene from Plutella xylostella: Possible involvement in resistance to beta-cypermethrin. Gene 2015, 558, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Terenius, O.; Papanicolaou, A.; Garbutt, J.S.; Eleftherianos, I.; Huvenne, H.; Kanginakudru, S.; Albrechtsen, M.; An, C.; Aymeric, J.L.; Barthel, A.; et al. RNA interference in Lepidoptera: An overview of successful and unsuccessful studies and implications for experimental design. J. Insect Physiol. 2011, 57, 231–245. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.L.; He, Y.N.; Staehelin, C.; Liu, S.W.; Su, Y.J.; Zhang, J.E. Identification of two cytochrome monooxygenase P450 genes, CYP321A7 and CYP321A9, from the tobacco cutworm moth (Spodoptera litura) and their expression in response to plant allelochemicals. Int. J. Mol. Sci. 2017, 18, 2278. [Google Scholar] [CrossRef] [PubMed]
- Ingelman, M.; Bianchi, V.; Eklund, H. The three-dimensional structure of flavodoxin reductase from Escherichia coli at 1.7 Å resolution. J. Mol. Biol. 1997, 268, 147–157. [Google Scholar] [CrossRef]
- Hubbard, P.A.; Shen, A.L.; Paschke, R.; Kasper, C.B.; Kim, J.J.P. NADPH-cytochrome P450 oxidoreductase structural basis for hydride and electron transfer. J. Biol. Chem. 2001, 276, 29163–29170. [Google Scholar] [CrossRef] [PubMed]
- Lamb, D.C.; Warrilow, A.G.S.; Venkateswarlu, K.; Kelly, D.E.; Kelly, S.L. Activities and kinetic mechanisms of native and soluble NADPH-cytochrome P450 reductase. Biochem. Biophys. Res. Commun. 2001, 286, 48–54. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Liang, Q.M.; Huang, Y.J.; Yuan, X.; Zhou, W.W.; Qiao, F.; Cheng, J.; Gurr, G.M.; Zhu, Z.R. Cloning, functional characterization, and expression profiles of NADPH-cytochrome P450 reductase gene from the Asiatic rice striped stem borer, Chilo suppressalis (Lepidoptera: Pyralidae). Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2013, 166, 225–231. [Google Scholar] [CrossRef]
- Flück, C.E.; Mullis, P.E.; Pandey, A.V. Modeling of human P450 oxidoreductase structure by in silico mutagenesis and MD simulation. Mol. Cell. Endocrinol. 2009, 313, 17–22. [Google Scholar] [CrossRef]
- Liu, D.; Zhou, X.; Li, M.; Zhu, S.; Qiu, X. Characterization of NADPH-cytochrome P450 reductase gene from the cotton bollworm, Helicoverpa armigera. Gene 2014, 545, 262–270. [Google Scholar] [CrossRef]
- Wang, K.; Peng, X.; Zuo, Y.; Li, Y.; Chen, M. Molecular cloning, expression pattern and polymorphisms of NADPH-cytochrome P450 reductase in the bird cherry-oat aphid Rhopalosiphum padi (L.). PLoS ONE 2016, 11, e0154633. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.J.; Li, G.H.; Pang, Y. A simple artificial diet for mass rearing of some noctuid species. Entomol. Knowl. 2000, 37, 8–10. [Google Scholar]
- Miller, J.S.; Stanley, D.W. Lipopolysaccharide evokes microaggregation reactions in hemocytes isolated from tobacco hornworms, Manduca sexta. Comp. Biochem. Physiol. Part A: Mol. Integr. Physiol. 2004, 137, 285–295. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhou, J.; Zhang, G.; Zhou, Q. Molecular characterization of cytochrome P450 CYP6B47 cDNAs and 5′-flanking sequence from Spodoptera litura (Lepidoptera: Noctuidae): Its response to lead stress. J. Insect Physiol. 2012, 58, 726–736. [Google Scholar] [CrossRef] [PubMed]
- Chenna, R.; Sugawara, H.; Koike, T.; Lopez, R.; Gibson, T.J.; Higgins, D.G.; Thompson, J.D. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 2003, 31, 3497–3500. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Yuan, M.; Gao, X.; Kang, T.; Zhan, S.; Wan, H.; Li, J. Identification and validation of reference genes for gene expression analysis using quantitative PCR in Spodoptera litura (Lepidoptera: Noctuidae). PLoS ONE. 2013, 8, e68059. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]

| Order | Species | Accession Number | Identity (%) |
|---|---|---|---|
| Hymenoptera | Monomorium pharaonis | XP_012541364 | 62.8 |
| Solenopsis invicta | XP_011157063 | 62.6 | |
| Pogonomyrmex barbatus | XP_011643152 | 61.5 | |
| Apis mellifera | XP_001119949 | 62.1 | |
| Apis florea | NP_001351669 | 62.6 | |
| Orussus abietinus | XP_012275162 | 62.6 | |
| Fopius arisanus | XP_011306347 | 63.4 | |
| Microplitis demolitor | XP_008548684 | 62.3 | |
| Hemiptera | Bemisia tabaci | AGT15701 | 61.9 |
| Cimex lectularius | AFD50507 | 62.7 | |
| Laodelphax striatella | AID55422 | 63.8 | |
| Sogatella furcifera | AHM93009 | 64.4 | |
| Coleoptera | Dendroctonus ponderosae | AFI45002 | 64.9 |
| Tribolium castaneum | XP_971174 | 67.6 | |
| Diptera | Drosophila mettleri | AAB48964 | 62.8 |
| Drosophila melanogaster | NP_477158 | 66.6 | |
| Musca domestica | AAA29295 | 68.4 | |
| Anopheles gambiae | AAO24765 | 66.6 | |
| Anopheles funestus | EF152578 | 67.5 | |
| Lepidoptera | Plutella xylostella | NP_001292469 | 79.4 |
| Chilo suppressalis | AGM20565 | 85.4 | |
| Bombyx mandarina | ABJ97709 | 87.0 | |
| Bombyx mori | NP_001104834 | 87.0 | |
| Helicoverpa armigera | ADK25060 | 95.6 | |
| Spodoptera exigua | ADX95746 | 98.4 | |
| Spodoptera littoralis | AFP20584 | 99.6 | |
| Spodoptera litura | MH638288 | 100 | |
| Rodentia | Mus musculus | NM_008898 | 53.5 |
| Primates | Homo sapiens | NP_000932 | 56.2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ji, H.-Y.; Staehelin, C.; Jiang, Y.-P.; Liu, S.-W.; Ma, Z.-H.; Su, Y.-J.; Zhang, J.-E.; Wang, R.-L. Tobacco Cutworm (Spodoptera Litura) Larvae Silenced in the NADPH-Cytochrome P450 Reductase Gene Show Increased Susceptibility to Phoxim. Int. J. Mol. Sci. 2019, 20, 3839. https://doi.org/10.3390/ijms20153839
Ji H-Y, Staehelin C, Jiang Y-P, Liu S-W, Ma Z-H, Su Y-J, Zhang J-E, Wang R-L. Tobacco Cutworm (Spodoptera Litura) Larvae Silenced in the NADPH-Cytochrome P450 Reductase Gene Show Increased Susceptibility to Phoxim. International Journal of Molecular Sciences. 2019; 20(15):3839. https://doi.org/10.3390/ijms20153839
Chicago/Turabian StyleJi, Hong-Yi, Christian Staehelin, Yan-Ping Jiang, Shi-Wei Liu, Zhi-Hui Ma, Yi-Juan Su, Jia-En Zhang, and Rui-Long Wang. 2019. "Tobacco Cutworm (Spodoptera Litura) Larvae Silenced in the NADPH-Cytochrome P450 Reductase Gene Show Increased Susceptibility to Phoxim" International Journal of Molecular Sciences 20, no. 15: 3839. https://doi.org/10.3390/ijms20153839
APA StyleJi, H.-Y., Staehelin, C., Jiang, Y.-P., Liu, S.-W., Ma, Z.-H., Su, Y.-J., Zhang, J.-E., & Wang, R.-L. (2019). Tobacco Cutworm (Spodoptera Litura) Larvae Silenced in the NADPH-Cytochrome P450 Reductase Gene Show Increased Susceptibility to Phoxim. International Journal of Molecular Sciences, 20(15), 3839. https://doi.org/10.3390/ijms20153839







