GmFAD3A, A ω-3 Fatty Acid Desaturase Gene, Enhances Cold Tolerance and Seed Germination Rate under Low Temperature in Rice
Abstract
1. Introduction
2. Results
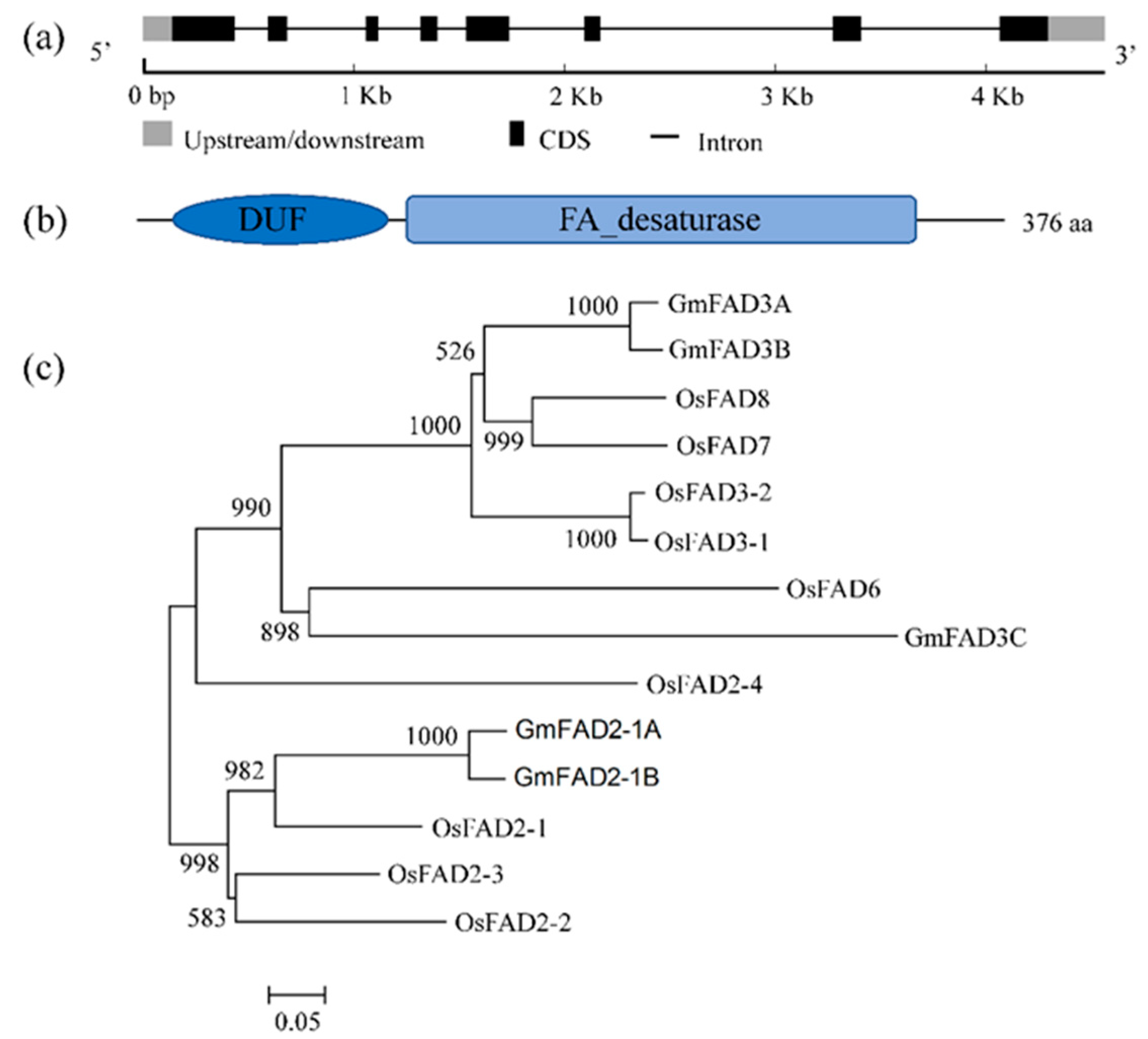
2.1. Structural Characteristics and Cladogram of GmFAD3A
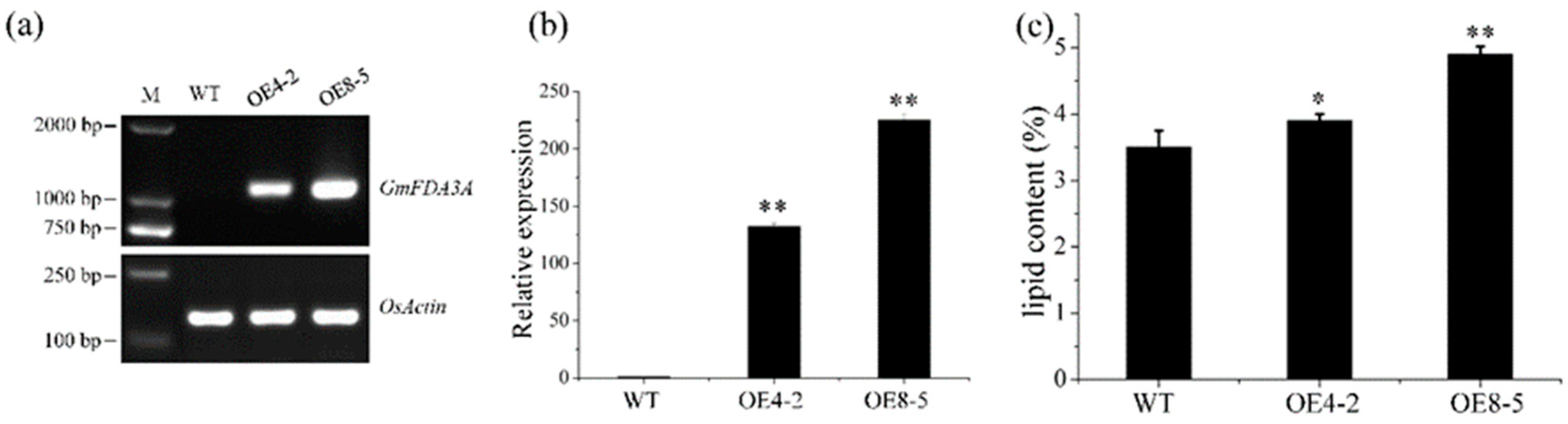
2.2. Ectopic Expression of GmFAD3A in Rice
2.3. Overexpression of GmFAD3A Increased Lipid Content
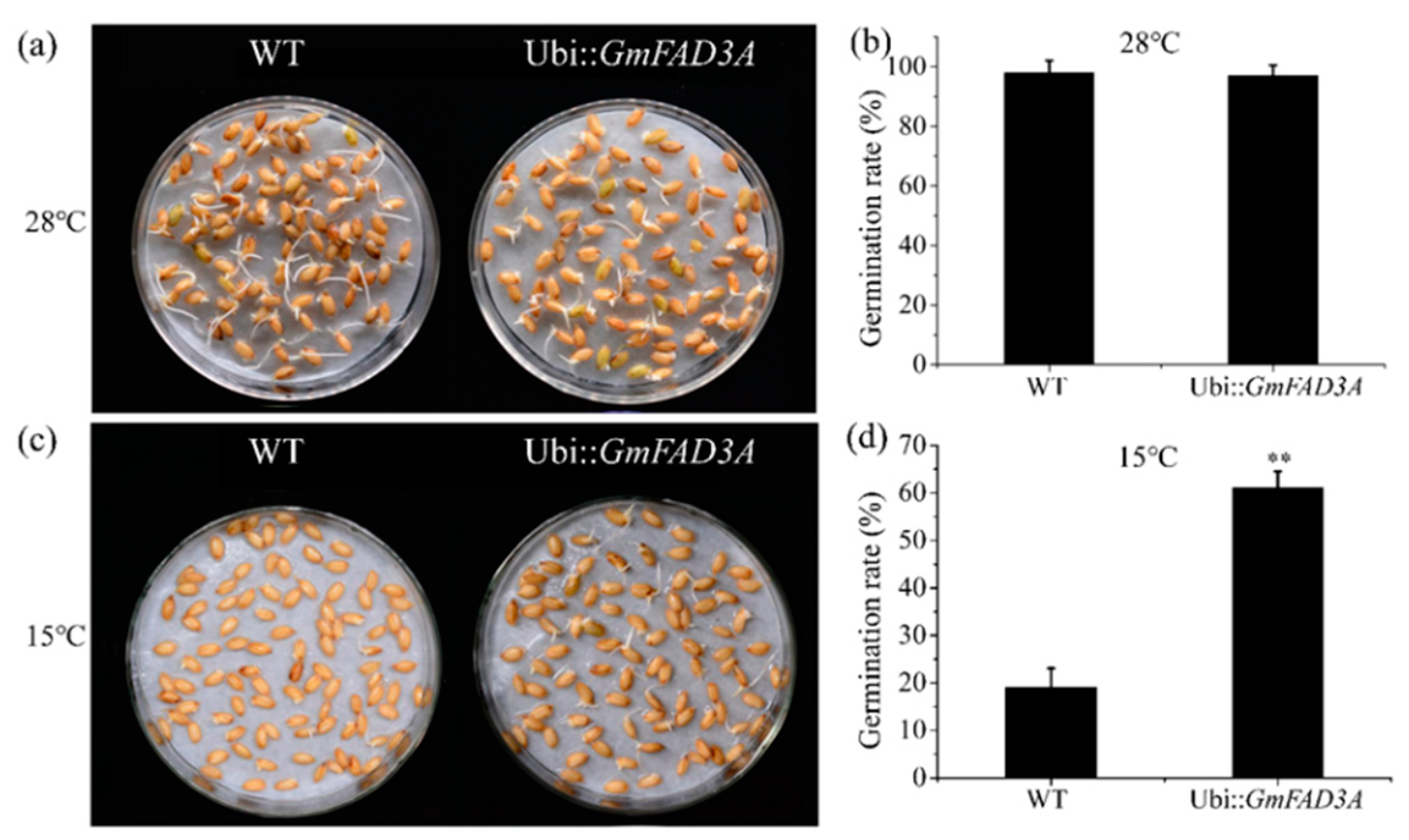
2.4. GmFAD3A Raised Seed Germination Rate at Low Temperature in Rice.
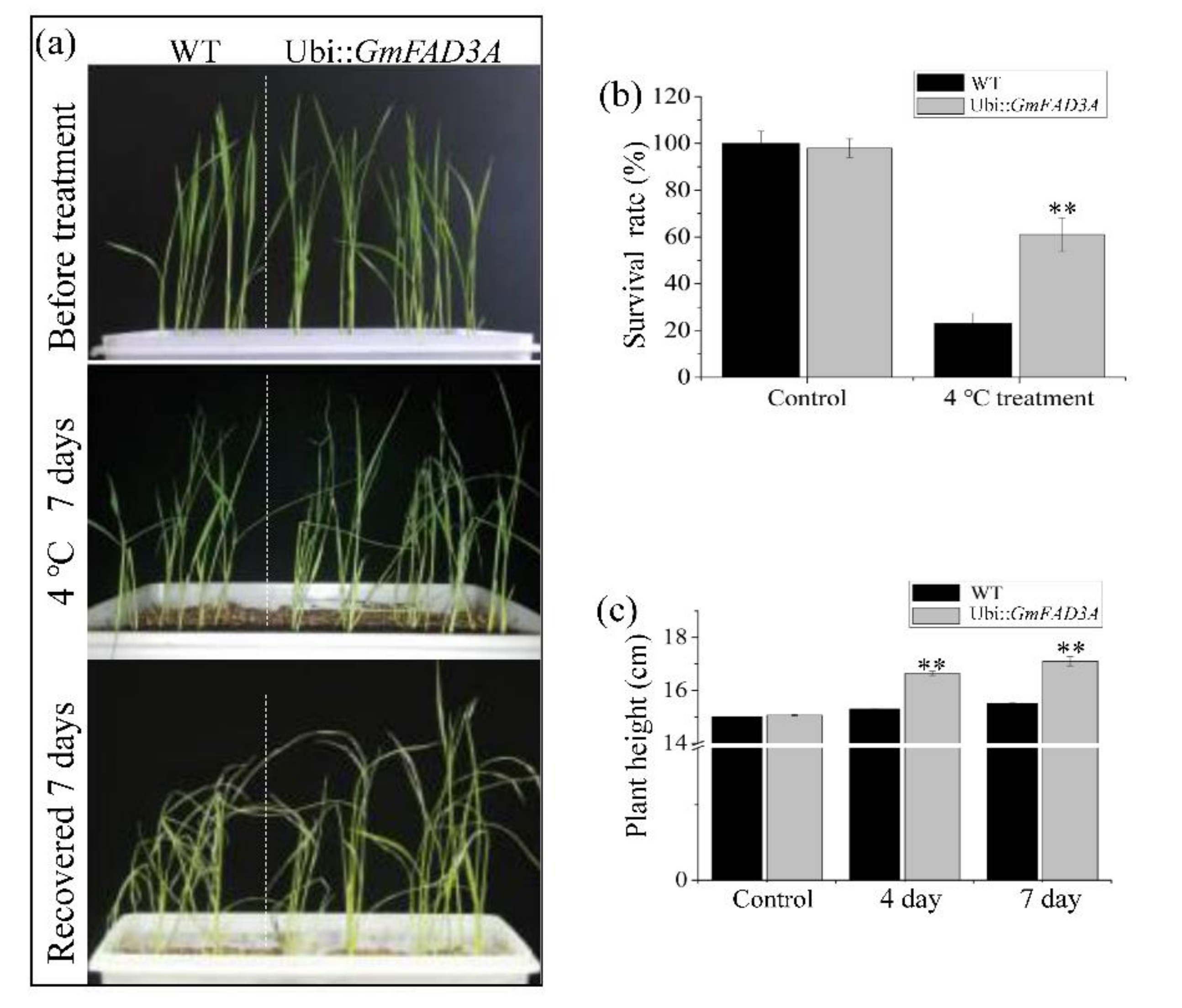
2.5. GmFAD3A Enhanced Cold Tolerance in Rice Seedlings
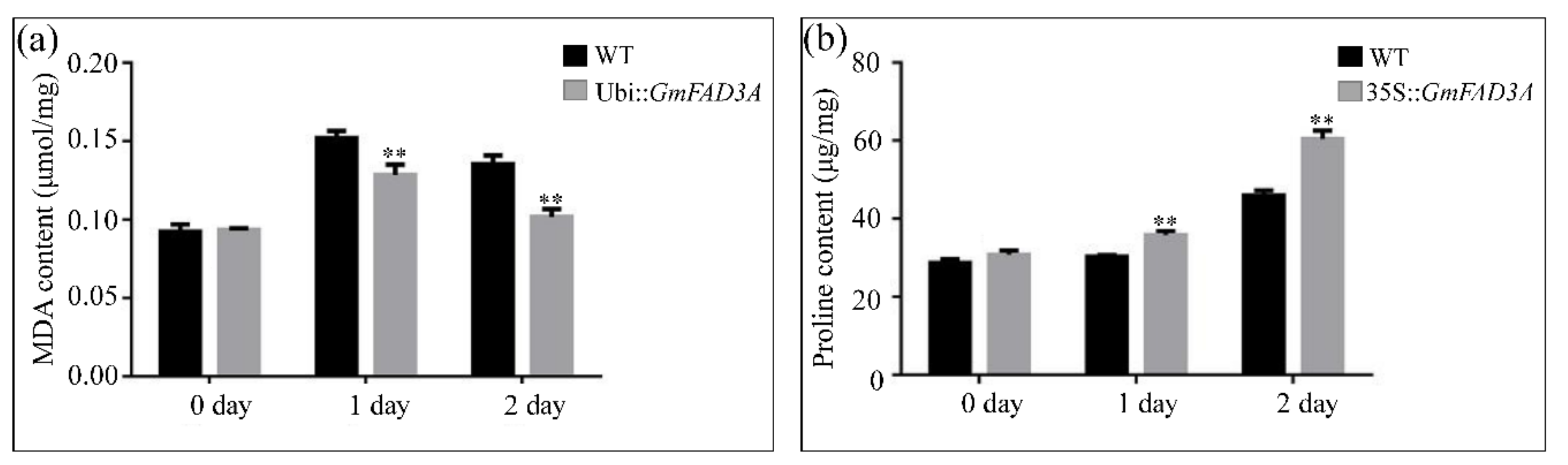
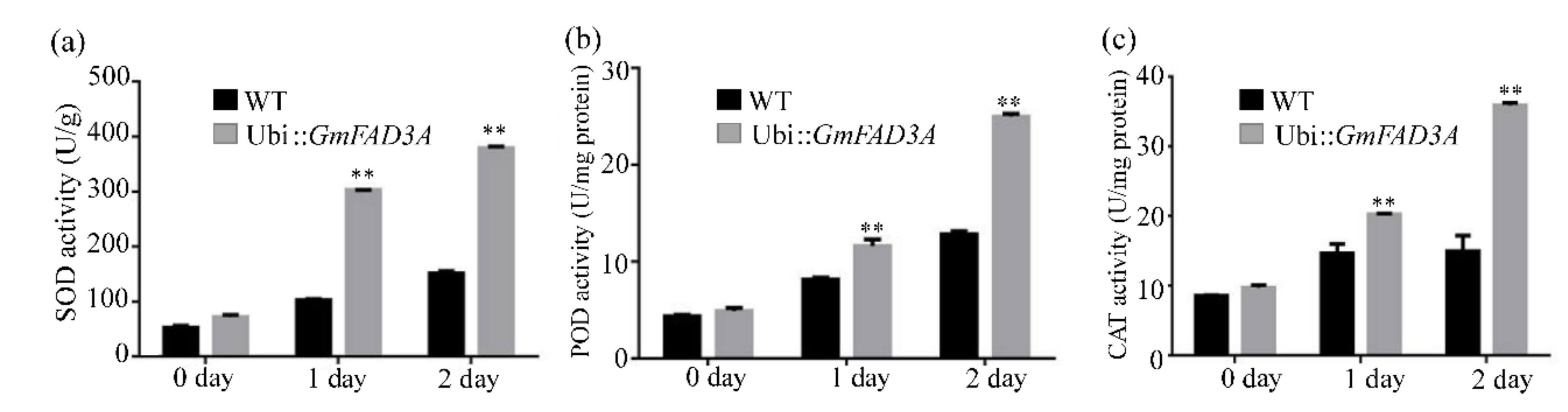
2.6. Determination of MDA, Proline Contents, and Antioxidant Enzymatic Activities
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Structure and Sequence Analysis of GmFAD3A
4.3. Plasmid Construction and Rice Transformation
4.4. RNA Isolation and Quantitative RT-PCR Analyses
4.5. Analysis of Lipid Content
4.6. Seed Germination Assays
4.7. Cold Stress Tolerance Experiment
4.8. Physiological and Biochemical Measurements
4.9. Enzyme Activity Assay
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| PUFA | polyunsaturated fatty acid |
| TAG | triacylglycerol |
| LA | linoleic acid |
| ALA | α-linolenic acid |
| MDA | malondialdehyde |
| SOD | superoxide dismutase |
| POD | peroxidase |
| CAT | hydroperoxidase |
| ROS | reactive oxygen species |
References
- Wang, X.; Zhou, W.; Lu, Z.; Ouyang, Y.; Yao, J. A lipid transfer protein, OsLTPL36, is essential for seed development and seed quality in rice. Plant Sci. 2015, 239, 200–208. [Google Scholar] [CrossRef] [PubMed]
- Endo, T.; Chiba, B.; Wagatsuma, K.; Saeki, K.; Ando, T.; Shomura, A.; Mizubayashi, T.; Ueda, T.; Yamamoto, T.; Nishio, T. Detection of QTLs for cold tolerance of rice cultivar ‘Kuchum’ and effect of QTL pyramiding. Theor. Appl. Genet. 2016, 129, 631–640. [Google Scholar] [CrossRef] [PubMed]
- Eremina, M.; Rozhon, W.; Poppenberger, B. Hormonal control of cold stress responses in plants. Cell Mol. Life Sci. 2016, 73, 797–810. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Xu, C.; Zhu, Y.; Zhang, L.; Chen, T.; Zhou, F.; Chen, H.; Lin, Y. The calcium-dependent kinase OsCPK24 functions in cold stress responses in rice. J. Integr. Plant Biol. 2018, 60, 173–188. [Google Scholar] [CrossRef] [PubMed]
- Beney, L.; Gervais, P. Influence of the fluidity of the membrane on the response of microorganisms to environmental stresses. Appl. Microbiol. Biotechnol. 2001, 57, 34–42. [Google Scholar] [PubMed]
- Dyer, J.M.; Mullen, R.T. Engineering plant oils as high-value industrial feedstocks for biorefining: The need for underpinning cell biology research. Physiol. Plant 2008, 132, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Wang, M.J.; Zhao, S.T.; Hu, J.J.; Lu, M.Z. Changes in freezing tolerance in hybrid poplar caused by up- and down-regulation of PtFAD2 gene expression. Transgenic Res. 2010, 19, 647–654. [Google Scholar] [CrossRef] [PubMed]
- Los, D.A.; Murata, N. Membrane fluidity and its roles in the perception of environmental signals. Biochim Biophys. Acta 2004, 1666, 142–157. [Google Scholar] [CrossRef] [PubMed]
- Ishizaki-Nishizawa, O.; Fujii, T.; Azuma, M.; Sekiguchi, K.; Murata, N.; Ohtani, T.; Toguri, T. Low-temperature resistance of higher plants is significantly enhanced by a nonspecific cyanobacterial desaturase. Nat. Biotechnol 1996, 14, 1003–1006. [Google Scholar] [CrossRef]
- Pham, A.T.; Shannon, J.G.; Bilyeu, K.D. Combinations of mutant FAD2 and FAD3 genes to produce high oleic acid and low linolenic acid soybean oil. Theor. Appl. Genet. 2012, 125, 503–515. [Google Scholar] [CrossRef]
- Schmutz, J.; Cannon, S.B.; Schlueter, J.; Ma, J.; Mitros, T.; Nelson, W.; Hyten, D.L.; Song, Q.; Thelen, J.J.; Cheng, J.; et al. Genome sequence of the palaeopolyploid soybean. Nature 2010, 463, 178–183. [Google Scholar] [CrossRef]
- Rajwade, A.V.; Kadoo, N.Y.; Borikar, S.P.; Harsulkar, A.M.; Ghorpade, P.B.; Gupta, V.S. Differential transcriptional activity of SAD, FAD2 and FAD3 desaturase genes in developing seeds of linseed contributes to varietal variation in alpha-linolenic acid content. Phytochemistry 2014, 98, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Silva, L.C.C.; Bueno, R.D.; da Matta, L.B.; Pereira, P.H.S.; Mayrink, D.B.; Piovesan, N.D.; Sediyama, C.S.; Fontes, E.P.B.; Cardinal, A.J.; Dal-Bianco, M. Characterization of a new GmFAD3A allele in Brazilian CS303TNKCA soybean cultivar. Theor. Appl. Genet. 2018, 131, 1099–1110. [Google Scholar] [CrossRef] [PubMed]
- Shimada, T.; Wakita, Y.; Otani, M.; Iba, K. Modification of Fatty Acid Composition in Rice Plants by Transformation with a Tobacco Microsomal omega;-3 Fatty Acid Desaturase Gene (NtFAD3). Plant Biotechnol. 2000, 17, 43–48. [Google Scholar] [CrossRef]
- Anai, T.; Koga, M.; Tanaka, H.; Kinoshita, T.; Rahman, S.M.; Takagi, Y. Improvement of rice (Oryza sativa L.) seed oil quality through introduction of a soybean microsomal omega-3 fatty acid desaturase gene. Plant Cell Rep. 2003, 21, 988–992. [Google Scholar] [CrossRef] [PubMed]
- Ying, J.Z.; Shan, J.X.; Gao, J.P.; Zhu, M.Z.; Shi, M.; Lin, H.X. Identification of quantitative trait Loci for lipid metabolism in rice seeds. Mol. Plant 2012, 5, 865–875. [Google Scholar] [CrossRef] [PubMed]
- Varshney, R.K.; Bansal, K.C.; Aggarwal, P.K.; Datta, S.K.; Craufurd, P.Q. Agricultural biotechnology for crop improvement in a variable climate: Hope or hype? Trends Plant Sci. 2011, 16, 363–371. [Google Scholar] [CrossRef]
- Wang, J.; Wang, J.; Wang, X.; Li, R.; Chen, B. Proteomic response of hybrid wild rice to cold stress at the seedling stage. PLoS ONE 2018, 13, e0198675. [Google Scholar] [CrossRef]
- Liu, C.T.; Wang, W.; Mao, B.G.; Chu, C.C. Cold stress tolerance in rice: Physiological changes, molecular mechanism, and future prospects. Yi Chuan 2018, 40, 171–185. [Google Scholar]
- Man, L.; Xiang, D.; Wang, L.; Zhang, W.; Wang, X.; Qi, G. Stress-responsive gene RsICE1 from Raphanus sativus increases cold tolerance in rice. Protoplasma 2017, 254, 945–956. [Google Scholar] [CrossRef]
- Liu, W.X.; Liu, H.L.; Qu le, Q. Embryo-specific expression of soybean oleosin altered oil body morphogenesis and increased lipid content in transgenic rice seeds. Theor. Appl. Genet. 2013, 126, 2289–2297. [Google Scholar] [CrossRef]
- Wang, H.S.; Yu, C.; Tang, X.F.; Wang, L.Y.; Dong, X.C.; Meng, Q.W. Antisense-mediated depletion of tomato endoplasmic reticulum omega-3 fatty acid desaturase enhances thermal tolerance. J. Integr. Plant Biol. 2010, 52, 568–577. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.L.; Yin, Z.J.; Xiao, L.; Xu, Y.N.; Qu le, Q. Identification and evaluation of omega-3 fatty acid desaturase genes for hyperfortifying alpha-linolenic acid in transgenic rice seed. J. Exp. Bot. 2012, 63, 3279–3287. [Google Scholar] [CrossRef]
- Nishida, I.; Murata, N. CHILLING SENSITIVITY IN PLANTS AND CYANOBACTERIA: The Crucial Contribution of Membrane Lipids. Annu. Rev. Plant Biol. 1996, 47, 541–568. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.P.; Shi, J.L.; Li, Z.; Ming, F. Isolation of OsFAD2, OsFAD6 and FAD family members response to abiotic stresses in Oryza sativa L. Yi chuan= Hereditas 2010, 32, 839–847. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Li, X.; Yuan, W.; Chen, G.; Kilian, A.; Li, J.; Xu, C.; Li, X.; Zhou, D.X.; Wang, S.; et al. Development of enhancer trap lines for functional analysis of the rice genome. Plant J. 2003, 35, 418–427. [Google Scholar] [CrossRef] [PubMed]
- Berry, S. Lipid Analysis: Isolation, Separation, Identification and Structural Analysis of Lipids. Nutr. Bulletin 2010, 29, 72–73. [Google Scholar] [CrossRef]
- Cruz-Hernandez, C.; Deng, Z.; Zhou, J.; Hill, A.R.; Yurawecz, M.P.; Delmonte, P.; Mossoba, M.M.; Dugan, M.E.; Kramer, J.K. Methods for analysis of conjugated linoleic acids and trans-18:1 isomers in dairy fats by using a combination of gas chromatography, silver-ion thin-layer chromatography/gas chromatography, and silver-ion liquid chromatography. J. AOAC Int. 2004, 87, 545–562. [Google Scholar]
- Byun, M.Y.; Cui, L.H.; Lee, J.; Park, H.; Lee, A.; Kim, W.T.; Lee, H. Identification of Rice Genes Associated with Enhanced Cold Tolerance by Comparative Transcriptome Analysis with Two Transgenic Rice Plants Overexpressing DaCBF4 or DaCBF7, Isolated From Antarctic Flowering Plant Deschampsia antarctica. Front. Plant Sci. 2018, 9, 601. [Google Scholar] [CrossRef]
- Wang, X.; Chen, J.; Liu, C.; Luo, J.; Yan, X.; Aihua, A.; Cai, Y.; Xie, H.; Ding, X.; Peng, X. Over-expression of a protein disulfide isomerase gene from Methanothermobacter thermautotrophicus, enhances heat stress tolerance in rice. Gene 2019, 684, 124–130. [Google Scholar] [CrossRef]
- Landi, M. Commentary to: “Improving the thiobarbituric acid-reactive-substances assay for estimating lipid peroxidation in plant tissues containing anthocyanin and other interfering compounds” by Hodges et al., Planta (1999) 207:604–611. Planta 2017, 245, 1067. [Google Scholar] [CrossRef] [PubMed]
- Kaur, N.; Dhawan, M.; Sharma, I.; Pati, P.K. Interdependency of Reactive Oxygen Species generating and scavenging system in salt sensitive and salt tolerant cultivars of rice. BMC Plant Biol. 2016, 16, 131. [Google Scholar] [CrossRef] [PubMed]
| Fatty Acid | Content (mg/g) | ||
|---|---|---|---|
| WT | OE4-3 | OE8-5 | |
| C13:0 (internal standard) | 1.982 ± 0.004 | 1.848 ± 0.005 | 1.435 ± 0.004 |
| C16:0 (palmitic acid) | 1.414 ± 0.056 | 1.587 ± 0.087 | 1.566 ± 0.077 |
| C18:0 (stearic acid) | 0.263 ± 0.007 | 0.273 ± 0.009 | 0.275 ± 0.005 |
| 9cC18:1 (oleic acid) | 3.468 ± 0.014 | 3.256 ± 0.018 * | 3.186 ± 0.020 * |
| C18:2 (linoleic acid) | 3.842 ± 0.046 | 4.058 ± 0.016 ** | 4.669 ± 0.014 ** |
| C18:3n-3 (linolenic acid) | 0.127 ± 0.013 | 0.159 ± 0.015 * | 0.187 ± 0.013 ** |
| C20:0 (arachidic acid) | 0.058 ± 0.004 | 0.061 ± 0.005 | 0.062 ± 0.003 |
| polyunsaturated fatty acids (PUFAs) | 3.945 ± 0.026 | 4.271 ± 0.035 ** | 4.983 ± 0.054 ** |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, X.; Yu, C.; Liu, Y.; Yang, L.; Li, Y.; Yao, W.; Cai, Y.; Yan, X.; Li, S.; Cai, Y.; et al. GmFAD3A, A ω-3 Fatty Acid Desaturase Gene, Enhances Cold Tolerance and Seed Germination Rate under Low Temperature in Rice. Int. J. Mol. Sci. 2019, 20, 3796. https://doi.org/10.3390/ijms20153796
Wang X, Yu C, Liu Y, Yang L, Li Y, Yao W, Cai Y, Yan X, Li S, Cai Y, et al. GmFAD3A, A ω-3 Fatty Acid Desaturase Gene, Enhances Cold Tolerance and Seed Germination Rate under Low Temperature in Rice. International Journal of Molecular Sciences. 2019; 20(15):3796. https://doi.org/10.3390/ijms20153796
Chicago/Turabian StyleWang, Xin, Chao Yu, Yi Liu, Lu Yang, Yang Li, Wen Yao, Yicong Cai, Xin Yan, Shaobo Li, Yaohui Cai, and et al. 2019. "GmFAD3A, A ω-3 Fatty Acid Desaturase Gene, Enhances Cold Tolerance and Seed Germination Rate under Low Temperature in Rice" International Journal of Molecular Sciences 20, no. 15: 3796. https://doi.org/10.3390/ijms20153796
APA StyleWang, X., Yu, C., Liu, Y., Yang, L., Li, Y., Yao, W., Cai, Y., Yan, X., Li, S., Cai, Y., Li, S., & Peng, X. (2019). GmFAD3A, A ω-3 Fatty Acid Desaturase Gene, Enhances Cold Tolerance and Seed Germination Rate under Low Temperature in Rice. International Journal of Molecular Sciences, 20(15), 3796. https://doi.org/10.3390/ijms20153796





