Gene Expression Profiling Reveals that PXR Activation Inhibits Hepatic PPARα Activity and Decreases FGF21 Secretion in Male C57Bl6/J Mice
Abstract
1. Introduction
2. Results
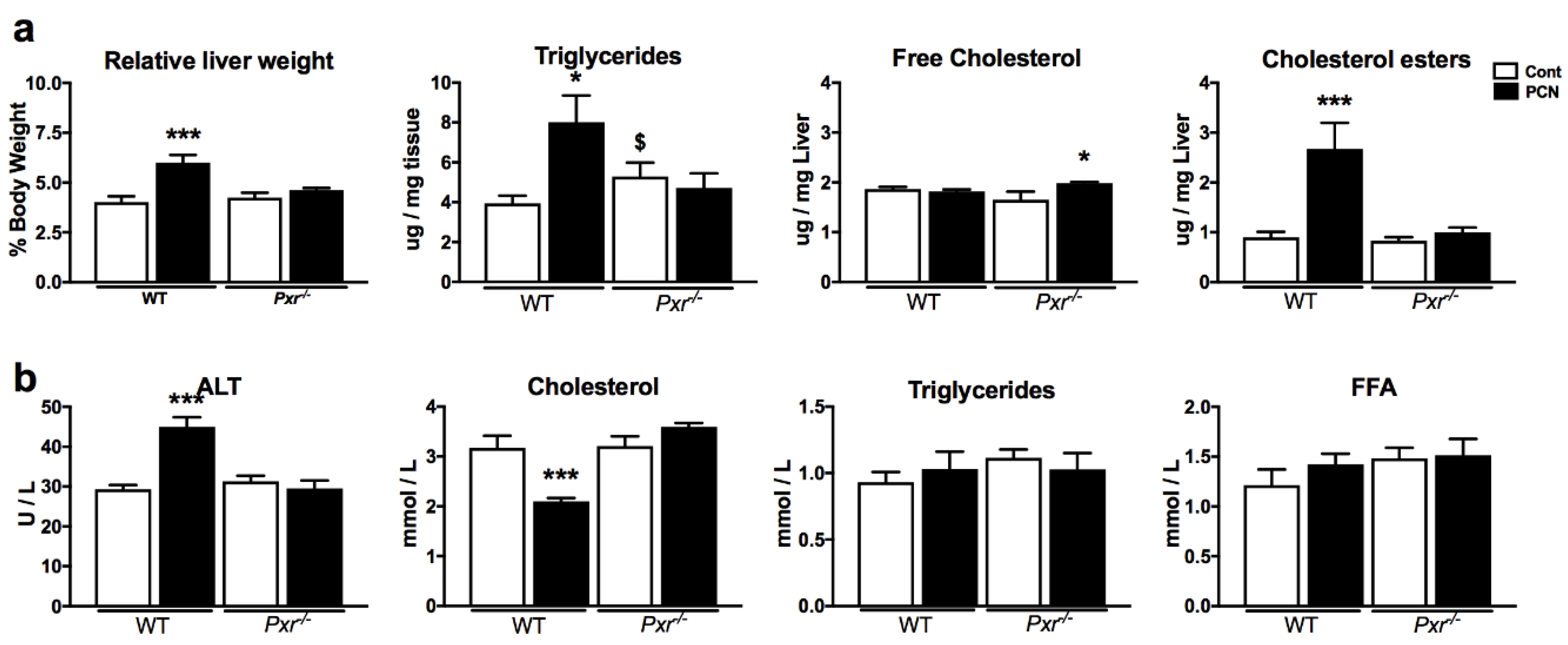
2.1. Effect of PXR Activation on Physiological Parameters and Liver Lipids
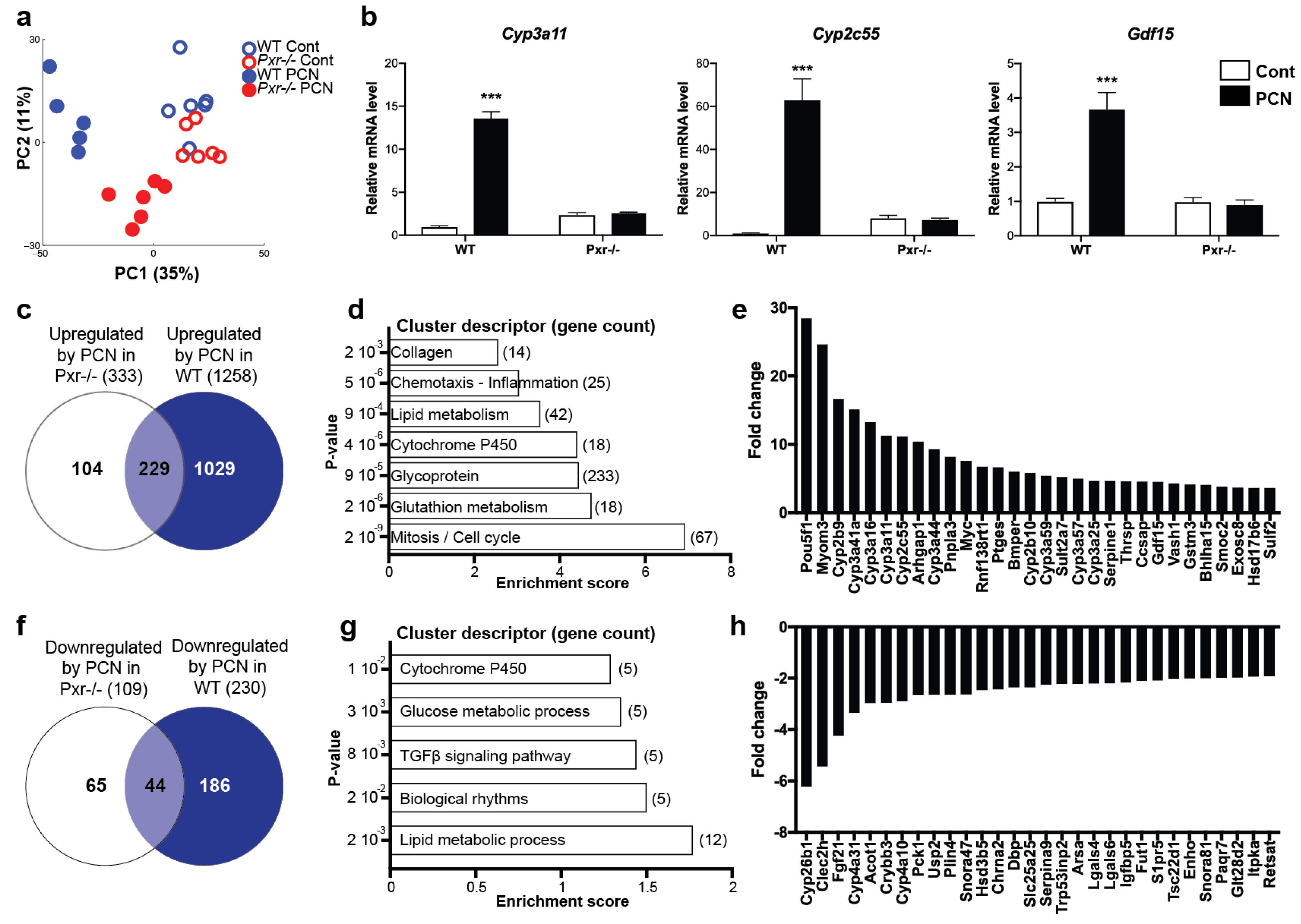
2.2. Effects of PXR Activation on the Hepatic Transcriptome
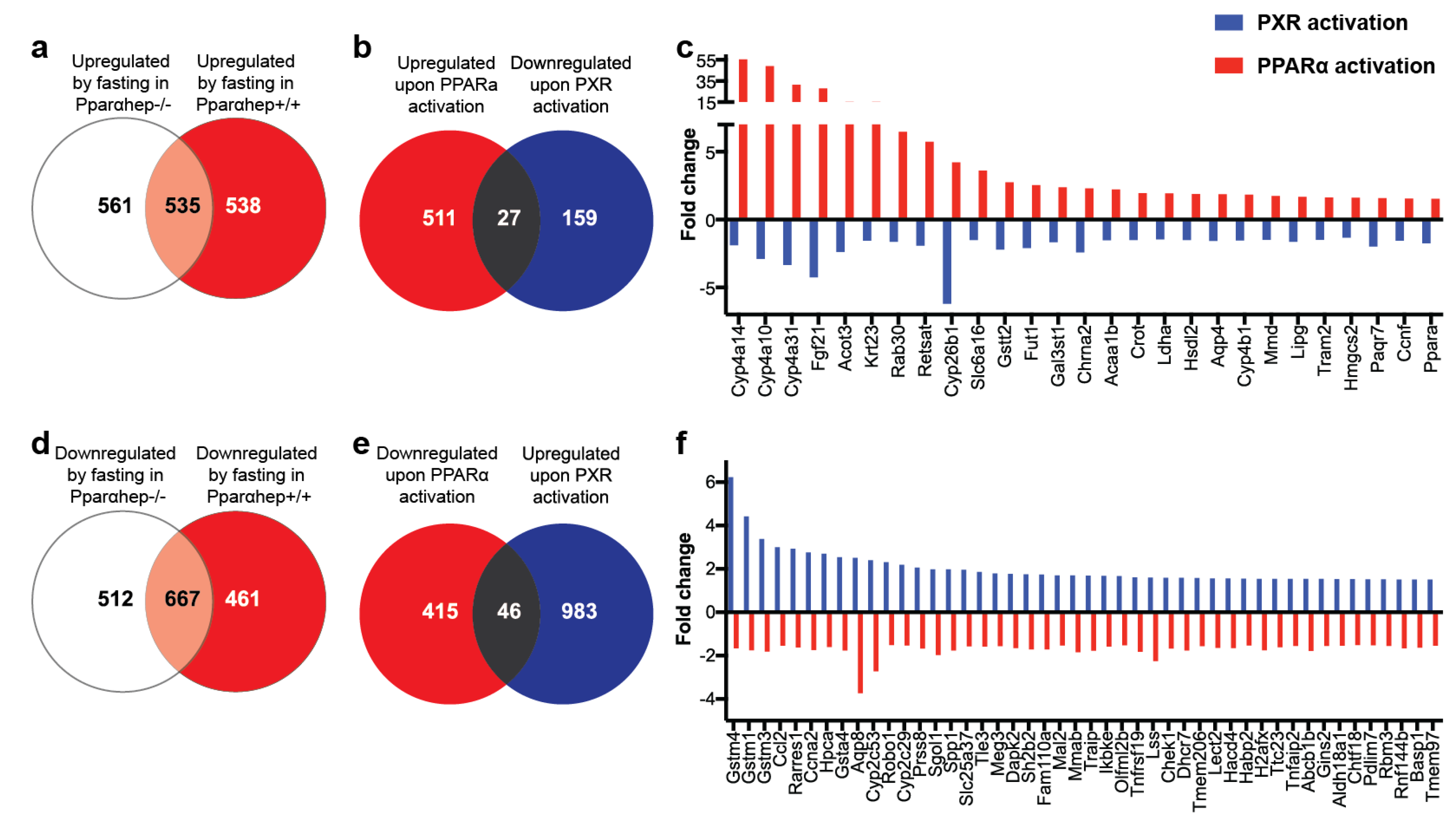
2.3. Comparison of PXR and PPARα-Dependent Transcriptome
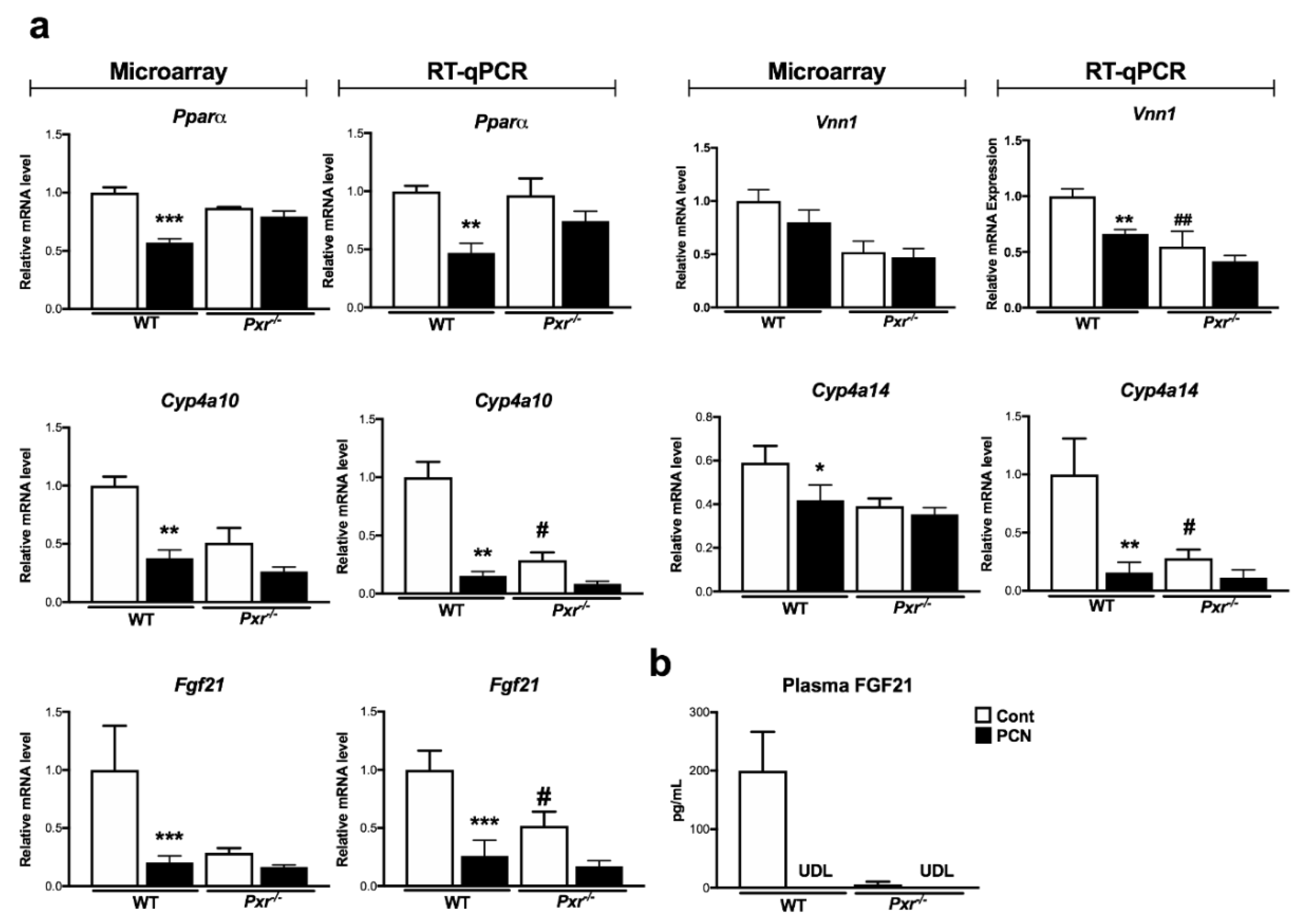
2.4. Regulation of FGF21
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. Blood and Tissue Samples
4.3. Gene Expression
4.4. Plasma Analysis
4.5. Liver Neutral Lipid Analysis
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ACAA | acetyl-Coenzyme A acyltransferase |
| Acot | acyl-coA thioeserase |
| AhR | aryl H receptor |
| CAR | constitutive androstane receptor |
| Cpt | carnitine palmitoyltransferase |
| CYP | cytochrome P450 |
| DR4 | direct repeat 4 |
| FASN | fatty acid synthase |
| FDR | false discovery rate |
| FGF21 | fibroblast growth factor 21 |
| G6Pase | glucose-6-phosphatase |
| GDF15 | growth/differentiation factor 15 |
| GCK | glucokinase |
| GO | gene ontology |
| GLUT2 | glucose transporter 2 |
| HMGCS | 3-hydroxy-3-methylglutaryl-CoenzymeA synthase |
| hPPARα | Human PPARα |
| hPXR | human PXR |
| Krt23 | keratin 23 |
| PXR | pregnane X receptor |
| PXRE | pregnane X receptor response elements |
| PCA | principal component analysis |
| PCN | pregnenolone 16α-carbonitrile |
| PEPCK | phosphoenopyruvate carboxykinase |
| PNPLA3 | patatin-like phospholipase domain containing 3 |
| PPARα | peroxisome proliferator-activated receptor α |
| Rab30 | ras-related protein rab-30 |
| RT-qPCR | quantitative reverse transcription PCR |
| RXR | retinoid X receptor |
| SPOT14 | thyroid hormone-responsive spot 14 |
| TGFβ | transforming growth factor-ß |
| Thrsp | thyroid hormone-responsive spot 14 |
| WT | wild-type |
| ZT | Zeitgeber time |
References
- Bookout, A.L.; Jeong, Y.; Downes, M.; Yu, R.T.; Evans, R.M.; Mangelsdorf, D.J. Anatomical profiling of nuclear receptor expression reveals a hierarchical transcriptional network. Cell 2006, 126, 789–799. [Google Scholar] [CrossRef] [PubMed]
- Kliewer, S.A.; Moore, J.T.; Wade, L.; Staudinger, J.L.; Watson, M.A.; Jones, S.A.; McKee, D.D.; Oliver, B.B.; Willson, T.M.; Zetterström, R.H.; et al. An orphan nuclear receptor activated by pregnanes defines a novel steroid signaling pathway. Cell 1998, 92, 73–82. [Google Scholar] [CrossRef]
- Hernandez, J.P.; Mota, L.C.; Baldwin, W.S. Activation of CAR and PXR by Dietary, Environmental and Occupational Chemicals Alters Drug Metabolism, Intermediary Metabolism, and Cell Proliferation. Curr. Pharm. Pers. Med. 2009, 7, 81–105. [Google Scholar] [CrossRef] [PubMed]
- Guengerich, F.P. Cytochrome P-450 3A4: Regulation and role in drug metabolism. Annu. Rev. Pharm. Toxicol. 1999, 39, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Drozdzik, M.; Busch, D.; Lapczuk, J.; Müller, J.; Ostrowski, M.; Kurzawski, M.; Oswald, S. Protein Abundance of Clinically Relevant Drug-Metabolizing Enzymes in the Human Liver and Intestine: A Comparative Analysis in Paired Tissue Specimens. Clin. Pharmacol. Ther. 2018, 104, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Petrie, I.D.; Levy, R.H.; Ragueneau-Majlessi, I. Mechanisms and Clinical Significance of Pharmacokinetic-based Drug-drug Interactions with Drugs Approved by the U.S. Food and Drug Administration in 2017. Drug Metab. Dispos. 2019, 47, 135–144. [Google Scholar] [CrossRef]
- Hakkola, J.; Rysä, J.; Hukkanen, J. Regulation of hepatic energy metabolism by the nuclear receptor PXR. Biochim. Biophys. Acta 2016, 1859, 1072–1082. [Google Scholar] [CrossRef]
- Kodama, S.; Koike, C.; Negishi, M.; Yamamoto, Y. Nuclear receptors CAR and PXR cross talk with FOXO1 to regulate genes that encode drug-metabolizing and gluconeogenic enzymes. Mol. Cell. Biol. 2004, 24, 7931–7940. [Google Scholar] [CrossRef]
- Rysä, J.; Buler, M.; Savolainen, M.J.; Ruskoaho, H.; Hakkola, J.; Hukkanen, J. Pregnane X receptor agonists impair postprandial glucose tolerance. Clin. Pharmacol. Ther. 2013, 93, 556–563. [Google Scholar] [CrossRef]
- Bitter, A.; Rümmele, P.; Klein, K.; Kandel, B.A.; Rieger, J.K.; Nüssler, A.K.; Zanger, U.M.; Trauner, M.; Schwab, M.; Burk, O. Pregnane X receptor activation and silencing promote steatosis of human hepatic cells by distinct lipogenic mechanisms. Arch. Toxicol. 2015, 89, 2089–2103. [Google Scholar] [CrossRef]
- Meng, Z.; Gwag, T.; Sui, Y.; Park, S.-H.; Zhou, X.; Zhou, C. The atypical antipsychotic quetiapine induces hyperlipidemia by activating intestinal PXR signaling. JCI Insight 2019, 4, 1. [Google Scholar] [CrossRef]
- Gwag, T.; Meng, Z.; Sui, Y.; Helsley, R.N.; Park, S.-H.; Wang, S.; Greenberg, R.N.; Zhou, C. Non-nucleoside reverse transcriptase inhibitor efavirenz activates PXR to induce hypercholesterolemia and hepatic steatosis. J. Hepatol. 2019, 70, 930–940. [Google Scholar] [CrossRef]
- Kliewer, S.A.; Mangelsdorf, D.J. A Dozen Years of Discovery: Insights into the Physiology and Pharmacology of FGF21. Cell Metab. 2019, 29, 246–253. [Google Scholar] [CrossRef] [PubMed]
- Régnier, M.; Polizzi, A.; Lippi, Y.; Fouché, E.; Michel, G.; Lukowicz, C.; Smati, S.; Marrot, A.; Lasserre, F.; Naylies, C.; et al. Insights into the role of hepatocyte PPARα activity in response to fasting. Mol. Cell. Endocrinol. 2017, 471, 75–88. [Google Scholar] [CrossRef]
- Montagner, A.; Polizzi, A.; Fouché, E.; Ducheix, S.; Lippi, Y.; Lasserre, F.; Barquissau, V.; Régnier, M.; Lukowicz, C.; Benhamed, F.; et al. Liver PPARα is crucial for whole-body fatty acid homeostasis and is protective against NAFLD. Gut 2016, 65, 1202–1214. [Google Scholar] [CrossRef] [PubMed]
- BonDurant, L.D.; Potthoff, M.J. Fibroblast Growth Factor 21: A Versatile Regulator of Metabolic Homeostasis. Annu. Rev. Nutr. 2018, 38, 173–196. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, I.; Hager, G.L. Transcriptional and Chromatin Regulation during Fasting–The Genomic Era. Trends Endocrinol. Metab. 2015, 26, 699–710. [Google Scholar] [CrossRef]
- Konno, Y.; Negishi, M.; Kodama, S. The roles of nuclear receptors CAR and PXR in hepatic energy metabolism. Drug Metab. Pharmacokinet. 2008, 23, 8–13. [Google Scholar] [CrossRef] [PubMed]
- Montagner, A.; Korecka, A.; Polizzi, A.; Lippi, Y.; Blum, Y.; Canlet, C.; Tremblay-Franco, M.; Gautier-Stein, A.; Burcelin, R.; Yen, Y.-C.; et al. Hepatic circadian clock oscillators and nuclear receptors integrate microbiome-derived signals. Sci. Rep. 2016, 6, 20127. [Google Scholar] [CrossRef]
- Li, C.Y.; Cheng, S.L.; Bammler, T.K.; Cui, J.Y. Editor’s Highlight: Neonatal Activation of the Xenobiotic-Sensors PXR and CAR Results in Acute and Persistent Down-regulation of PPARα-Signaling in Mouse Liver. Toxicol. Sci. 2016, 153, 282–302. [Google Scholar] [CrossRef]
- Cui, J.Y.; Klaassen, C.D. RNA-Seq reveals common and unique PXR- and CAR-target gene signatures in the mouse liver transcriptome. Biochim. Biophys. Acta 2016, 1859, 1198–1217. [Google Scholar] [CrossRef]
- Nagahori, H.; Nakamura, K.; Sumida, K.; Ito, S.; Ohtsuki, S. Combining Genomics To Identify the Pathways of Post-Transcriptional Nongenotoxic Signaling and Energy Homeostasis in Livers of Rats Treated with the Pregnane X Receptor Agonist, Pregnenolone Carbonitrile. J. Proteome Res. 2017, 16, 3634–3645. [Google Scholar] [CrossRef]
- Kandel, B.A.; Thomas, M.; Winter, S.; Damm, G.; Seehofer, D.; Burk, O.; Schwab, M.; Zanger, U.M. Genomewide comparison of the inducible transcriptomes of nuclear receptors CAR, PXR and PPARα in primary human hepatocytes. Biochim. Biophys. Acta 2016, 1859, 1218–1227. [Google Scholar] [CrossRef]
- LaFave, L.T.; Augustin, L.B.; Mariash, C.N. S14: Insights from knockout mice. Endocrinology 2006, 147, 4044–4047. [Google Scholar] [CrossRef]
- Moreau, A.; Téruel, C.; Beylot, M.; Albalea, V.; Tamasi, V.; Umbdenstock, T.; Parmentier, Y.; Sa-Cunha, A.; Suc, B.; Fabre, J.-M.; et al. A novel pregnane X receptor and S14-mediated lipogenic pathway in human hepatocyte. Hepatology 2009, 49, 2068–2079. [Google Scholar] [CrossRef]
- Dai, G.; Liu, P.; Li, X.; Zhou, X.; He, S. Association between PNPLA3 rs738409 polymorphism and nonalcoholic fatty liver disease (NAFLD) susceptibility and severity: A meta-analysis. Medicine 2019, 98, e14324. [Google Scholar] [CrossRef]
- Chrysovergis, K.; Wang, X.; Kosak, J.; Lee, S.-H.; Kim, J.S.; Foley, J.F.; Travlos, G.; Singh, S.; Baek, S.J.; Eling, T.E. NAG-1/GDF-15 prevents obesity by increasing thermogenesis, lipolysis and oxidative metabolism. Int. J. Obes. (Lond) 2014, 38, 1555–1564. [Google Scholar] [CrossRef]
- Lee, J.; Choi, J.; Scafidi, S.; Wolfgang, M.J. Hepatic Fatty Acid Oxidation Restrains Systemic Catabolism during Starvation. Cell Rep. 2016, 16, 201–212. [Google Scholar] [CrossRef]
- Mullican, S.E.; Lin-Schmidt, X.; Chin, C.-N.; Chavez, J.A.; Furman, J.L.; Armstrong, A.A.; Beck, S.C.; South, V.J.; Dinh, T.Q.; Cash-Mason, T.D.; et al. GFRAL is the receptor for GDF15 and the ligand promotes weight loss in mice and nonhuman primates. Nat. Med. 2017, 23, 1150–1157. [Google Scholar] [CrossRef]
- Markan, K.R.; Naber, M.C.; Ameka, M.K.; Anderegg, M.D.; Mangelsdorf, D.J.; Kliewer, S.A.; Mohammadi, M.; Potthoff, M.J. Circulating FGF21 is liver derived and enhances glucose uptake during refeeding and overfeeding. Diabetes 2014, 63, 4057–4063. [Google Scholar] [CrossRef]
- Zhang, Y.; Xie, Y.; Berglund, E.D.; Coate, K.C.; He, T.T.; Katafuchi, T.; Xiao, G.; Potthoff, M.J.; Wei, W.; Wan, Y.; et al. The starvation hormone, fibroblast growth factor-21, extends lifespan in mice. Elife 2012, 1, e00065. [Google Scholar] [CrossRef]
- Owen, B.M.; Bookout, A.L.; Ding, X.; Lin, V.Y.; Atkin, S.D.; Gautron, L.; Kliewer, S.A.; Mangelsdorf, D.J. FGF21 contributes to neuroendocrine control of female reproduction. Nat. Med. 2013, 19, 1153–1156. [Google Scholar] [CrossRef]
- Iroz, A.; Montagner, A.; Benhamed, F.; Levavasseur, F.; Polizzi, A.; Anthony, E.; Régnier, M.; Fouché, E.; Lukowicz, C.; Cauzac, M.; et al. A Specific ChREBP and PPARα Cross-Talk Is Required for the Glucose-Mediated FGF21 Response. Cell Rep. 2017, 21, 403–416. [Google Scholar] [CrossRef]
- Ellero-Simatos, S.; Chakhtoura, G.; Barreau, C.; Langouët, S.; Benelli, C.; Penicaud, L.; Beaune, P.; de Waziers, I. Xenobiotic-metabolizing cytochromes p450 in human white adipose tissue: Expression and induction. Drug Metab. Dispos. 2010, 38, 679–686. [Google Scholar] [CrossRef]
- He, J.; Gao, J.; Xu, M.; Ren, S.; Stefanovic-Racic, M.; O’Doherty, R.M.; Xie, W. PXR ablation alleviates diet-induced and genetic obesity and insulin resistance in mice. Diabetes 2013, 62, 1876–1887. [Google Scholar] [CrossRef]
- Spruiell, K.; Jones, D.Z.; Cullen, J.M.; Awumey, E.M.; Gonzalez, F.J.; Gyamfi, M.A. Role of human pregnane X receptor in high fat diet-induced obesity in pre-menopausal female mice. Biochem. Pharm. 2014, 89, 399–412. [Google Scholar] [CrossRef]
- Ma, Y.; Liu, D. Activation of pregnane X receptor by pregnenolone 16 α-carbonitrile prevents high-fat diet-induced obesity in AKR/J mice. PLoS ONE 2012, 7, e38734. [Google Scholar] [CrossRef]
- Lu, Y.-F.; Jin, T.; Xu, Y.; Zhang, D.; Wu, Q.; Zhang, Y.-K.J.; Liu, J. Sex differences in the circadian variation of cytochrome p450 genes and corresponding nuclear receptors in mouse liver. Chronobiol. Int. 2013, 30, 1135–1143. [Google Scholar] [CrossRef]
- Spruiell, K.; Gyamfi, A.A.; Yeyeodu, S.T.; Richardson, R.M.; Gonzalez, F.J.; Gyamfi, M.A. Pregnane X Receptor-Humanized Mice Recapitulate Gender Differences in Ethanol Metabolism but Not Hepatotoxicity. J. Pharm. Exp. 2015, 354, 459–470. [Google Scholar] [CrossRef]
- Staudinger, J.L.; Goodwin, B.; Jones, S.A.; Hawkins-Brown, D.; MacKenzie, K.I.; LaTour, A.; Liu, Y.; Klaassen, C.D.; Brown, K.K.; Reinhard, J.; et al. The nuclear receptor PXR is a lithocholic acid sensor that protects against liver toxicity. Proc. Natl. Acad. Sci. USA 2001, 98, 3369–3374. [Google Scholar] [CrossRef]
- Edgar, R.; Domrachev, M.; Lash, A.E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002, 30, 207–210. [Google Scholar] [CrossRef]
- Bligh, E.G.; Dyer, W.J. A RAPID METHOD OF TOTAL LIPID EXTRACTION AND PURIFICATION. Can. J. Biochem. Physiol. 2011, 37, 911–917. [Google Scholar] [CrossRef]
- Cheng, L.; Lo, L.-Y.; Tang, N.L.S.; Wang, D.; Leung, K.-S. CrossNorm: A novel normalization strategy for microarray data in cancers. Sci. Rep. 2016, 6, 18898. [Google Scholar] [CrossRef]
- Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barretto, S.A.; Lasserre, F.; Fougerat, A.; Smith, L.; Fougeray, T.; Lukowicz, C.; Polizzi, A.; Smati, S.; Régnier, M.; Naylies, C.; et al. Gene Expression Profiling Reveals that PXR Activation Inhibits Hepatic PPARα Activity and Decreases FGF21 Secretion in Male C57Bl6/J Mice. Int. J. Mol. Sci. 2019, 20, 3767. https://doi.org/10.3390/ijms20153767
Barretto SA, Lasserre F, Fougerat A, Smith L, Fougeray T, Lukowicz C, Polizzi A, Smati S, Régnier M, Naylies C, et al. Gene Expression Profiling Reveals that PXR Activation Inhibits Hepatic PPARα Activity and Decreases FGF21 Secretion in Male C57Bl6/J Mice. International Journal of Molecular Sciences. 2019; 20(15):3767. https://doi.org/10.3390/ijms20153767
Chicago/Turabian StyleBarretto, Sharon Ann, Frédéric Lasserre, Anne Fougerat, Lorraine Smith, Tiffany Fougeray, Céline Lukowicz, Arnaud Polizzi, Sarra Smati, Marion Régnier, Claire Naylies, and et al. 2019. "Gene Expression Profiling Reveals that PXR Activation Inhibits Hepatic PPARα Activity and Decreases FGF21 Secretion in Male C57Bl6/J Mice" International Journal of Molecular Sciences 20, no. 15: 3767. https://doi.org/10.3390/ijms20153767
APA StyleBarretto, S. A., Lasserre, F., Fougerat, A., Smith, L., Fougeray, T., Lukowicz, C., Polizzi, A., Smati, S., Régnier, M., Naylies, C., Bétoulières, C., Lippi, Y., Guillou, H., Loiseau, N., Gamet-Payrastre, L., Mselli-Lakhal, L., & Ellero-Simatos, S. (2019). Gene Expression Profiling Reveals that PXR Activation Inhibits Hepatic PPARα Activity and Decreases FGF21 Secretion in Male C57Bl6/J Mice. International Journal of Molecular Sciences, 20(15), 3767. https://doi.org/10.3390/ijms20153767






