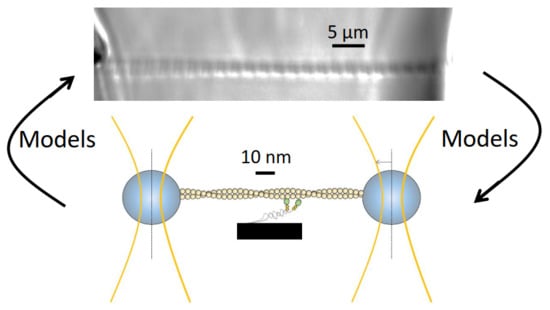
Do Actomyosin Single-Molecule Mechanics Data Predict Mechanics of Contracting Muscle?
Abstract
1. Introduction
2. Mechanical Measurements on Isolated Molecular Motors—Principles and Basic Findings
3. Challenges in Single-Molecule Mechanics
3.1. Directionality of Motor Induced Force and Displacement
3.2. Surface Immobilization and Surface-Motor Interactions
3.3. Solution Compositions Differing from that In Vivo
3.4. More on Lack of Accessory Proteins in Single-Molecule Mechanics
3.5. Time Resolution
4. Mechanical Experiments on Muscle Cells and Myofibrils—Concepts, Main Findings and Challenges
5. Key Cross-Bridge Characteristics from Single Molecules to Muscle
5.1. General
5.2. Cross-Bridge Stiffness
5.3. A Simple Model
5.4. The Model and Single Molecules
5.5. The Model and Muscle Mechanics
5.6. Section Summary
6. Top-Down and Bottom-Up Models
6.1. Challenges with Top-Down Models
6.2. Bottom-Up Models to the Rescue?
6.3. Bottom-Up Models and Non-Linear Cross-Bridge Elasticity
6.4. Jumping Cross-Bridges, Mechanosensing and Two Heads
6.5. Roles of Accessory Proteins and Non-Uniform Sarcomere Behavior in Muscle Contraction in Health and Disease
7. Conclusions and Perspectives
- Are there any sub-strokes in the power-stroke and is force-generation a Kramers or an Eyring process? Furthermore, related, the maximum power-stroke distance for which a single step is possible is about 9 nm (assuming an Eyring process)? Thus, a 9 nm working stroke with cross-bridge stiffness of 2 pN/nm corresponds to a maximum isometric cross-bridge force of less than 20 pN and maximum power of 81 pNnm ≈ 20 kBT, less than the free energy of ATP turnover under physiological conditions.
- What are the characteristics and magnitude of the cross-bridge elasticity in muscle cells? Is the stiffness about 2 pN/nm or higher at positive strain and is the elasticity non-linear as proposed by single-molecule studies [67]? Furthermore, what is the characteristic of the myofilament elasticity in muscle cells?
- What is the role of the two myosin heads? Is one of the heads just guiding the other to its binding site as suggested by single-molecule studies or can the ordered geometry of the myofilament lattice cause the heads to work in sequence?
- Can the high maximum power-output of muscle be explained based on bottom-up defined models without assuming cooperative effects (e.g., between two heads), altered kinetics in an ordered ensemble (e.g., jumping between sites) or effects of mechanosensing in the thick filaments?
- Can force-dependent rates obtained in optical tweezers studies predict the contractile behavior of muscle, for instance if the cross-bridge elasticity in muscle cells is non-linear as in [67]? Further model studies along these lines are of interest but also single-molecule studies to elucidate the force-dependence of rate constants in full length myosin.
- What is the main cross-bridge dependent mechanism of the force-enhancement during stretch? Are the two heads important or is there jumping between neighboring sites, possibly related to stretch induced changes in actin with strongly enhanced affinity at neighboring sites. For further insight, it would be of interest to supplement the muscle mechanics studies with single-molecule experiments. To the best of our knowledge such studies have not yet been performed.
- Are the modulatory roles of accessory proteins of increasing importance in diseases, e.g., as suggested for possible roles of MyBPC or titin in controlling the sequestered myosin head motif and also for mutations in RLC and other regulatory proteins such as troponin and tropomyosin. Further, are emergent phenomena enhanced due to increased sarcomere non-uniformities and mechanical instabilities in sarcomere myopathies?
Author Contributions
Funding
Conflicts of Interest
References
- Hudson, L.; Harford, J.J.; Denny, R.C.; Squire, J.M. Myosin head configuration in relaxed fish muscle: Resting state myosin heads must swing axially by up to 150 A or turn upside down to reach rigor. J. Mol. Biol. 1997, 273, 440–455. [Google Scholar] [CrossRef] [PubMed]
- Ebashi, S.; Endo, M. Calcium ion and muscle contraction. Prog. Biophys. Mol. Biol. 1968, 18, 123–183. [Google Scholar] [CrossRef]
- Spudich, J.A.; Watt, S. The regulation of rabbit skeletal muscle contraction. I. Biochemical studies of the interaction of the tropomyosin-troponin complex with actin and the proteolytic fragments of myosin. J. Biol. Chem. 1971, 246, 4866–4871. [Google Scholar] [PubMed]
- Offer, G.; Moos, C.; Starr, R. A new protein of the thick filaments of vertebrate skeletal myofibrils. Extractions, purification and characterization. J. Mol. Biol. 1973, 74, 653–676. [Google Scholar] [CrossRef]
- Previs, M.J.; Beck Previs, S.; Gulick, J.; Robbins, J.; Warshaw, D.M. Molecular mechanics of cardiac myosin-binding protein C in native thick filaments. Science 2012, 337, 1215–1218. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Ramirez-Mitchell, R.; Palter, D. Titin is an extraordinarily long, flexible, and slender myofibrillar protein. Proc. Natl. Acad. Sci. USA 1984, 81, 3685–3689. [Google Scholar] [CrossRef] [PubMed]
- Kellermayer, M.S.; Smith, S.B.; Granzier, H.L.; Bustamante, C. Folding-unfolding transitions in single titin molecules characterized with laser tweezers. Science 1997, 276, 1112–1116. [Google Scholar] [CrossRef] [PubMed]
- Linke, W.A.; Kruger, M. The giant protein titin as an integrator of myocyte signaling pathways. Physiology (Bethesda) 2010, 25, 186–198. [Google Scholar] [CrossRef] [PubMed]
- Raskin, A.; Lange, S.; Banares, K.; Lyon, R.C.; Zieseniss, A.; Lee, L.K.; Yamazaki, K.G.; Granzier, H.L.; Gregorio, C.C.; McCulloch, A.D.; et al. A novel mechanism involving four-and-a-half LIM domain protein-1 and extracellular signal-regulated kinase-2 regulates titin phosphorylation and mechanics. J. Biol. Chem. 2012, 287, 29273–29284. [Google Scholar] [CrossRef] [PubMed]
- Millman, B.M. The filament lattice of striated muscle. Physiol. Rev. 1998, 78, 359–391. [Google Scholar] [CrossRef] [PubMed]
- Luther, P.K.; Squire, J.M. The intriguing dual lattices of the Myosin filaments in vertebrate striated muscles: Evolution and advantage. Biology 2014, 3, 846–865. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Andersson-Lendahl, M.; Sejersen, T.; Arner, A. Knockdown of desmin in zebrafish larvae affects interfilament spacing and mechanical properties of skeletal muscle. J. Gen. Physiol. 2013, 141, 335–345. [Google Scholar] [CrossRef] [PubMed]
- Lieber, R.L.; Friden, J. Functional and clinical significance of skeletal muscle architecture. Muscle Nerve 2000, 23, 1647–1666. [Google Scholar] [CrossRef]
- Steinmetz, P.R.; Kraus, J.E.; Larroux, C.; Hammel, J.U.; Amon-Hassenzahl, A.; Houliston, E.; Worheide, G.; Nickel, M.; Degnan, B.M.; Technau, U. Independent evolution of striated muscles in cnidarians and bilaterians. Nature 2012, 487, 231–234. [Google Scholar] [CrossRef] [PubMed]
- Bennet-Clark, H.C. The energetics of the jump of the locust Schistocerca gregaria. J. Exp. Biol. 1975, 63, 53–83. [Google Scholar] [PubMed]
- Malik, F.I.; Hartman, J.J.; Elias, K.A.; Morgan, B.P.; Rodriguez, H.; Brejc, K.; Anderson, R.L.; Sueoka, S.H.; Lee, K.H.; Finer, J.T.; et al. Cardiac myosin activation: A potential therapeutic approach for systolic heart failure. Science 2011, 331, 1439–1443. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Ajtai, K.; Burghardt, T.P. Analytical comparison of natural and pharmaceutical ventricular myosin activators. Biochemistry 2014, 53, 5298–5306. [Google Scholar] [CrossRef] [PubMed]
- Aksel, T.; Choe Yu, E.; Sutton, S.; Ruppel, K.M.; Spudich, J.A. Ensemble force changes that result from human cardiac myosin mutations and a small-molecule effector. Cell Rep. 2015, 11, 910–920. [Google Scholar] [CrossRef] [PubMed]
- Rohde, J.A.; Thomas, D.D.; Muretta, J.M. Heart failure drug changes the mechanoenzymology of the cardiac myosin powerstroke. Proc. Natl. Acad. Sci. USA 2017, 114, E1796–E1804. [Google Scholar] [CrossRef] [PubMed]
- Swenson, A.M.; Tang, W.; Blair, C.A.; Fetrow, C.M.; Unrath, W.C.; Previs, M.J.; Campbell, K.S.; Yengo, C.M. Omecamtiv Mecarbil Enhances the Duty Ratio of Human β-Cardiac Myosin Resulting in Increased Calcium Sensitivity and Slowed Force Development in Cardiac Muscle. J. Biol. Chem. 2017, 292, 3768–3778. [Google Scholar] [CrossRef] [PubMed]
- Kampourakis, T.; Zhang, X.; Sun, Y.B.; Irving, M. Omecamtiv mercabil and blebbistatin modulate cardiac contractility by perturbing the regulatory state of the myosin filament. J. Physiol. 2018, 596, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Straight, A.F.; Cheung, A.; Limouze, J.; Chen, I.; Westwood, N.J.; Sellers, J.R.; Mitchison, T.J. Dissecting temporal and spatial control of cytokinesis with a myosin II Inhibitor. Science 2003, 299, 1743–1747. [Google Scholar] [CrossRef] [PubMed]
- Kovacs, M.; Toth, J.; Hetenyi, C.; Malnasi-Csizmadia, A.; Sellers, J.R. Mechanism of blebbistatin inhibition of myosin II. J. Biol. Chem. 2004, 279, 35557–35563. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Blair, C.A.; Walton, S.D.; Malnasi-Csizmadia, A.; Campbell, K.S.; Yengo, C.M. Modulating β-Cardiac Myosin Function at the Molecular and Tissue Levels. Front. Physiol. 2016, 7, 659. [Google Scholar] [CrossRef] [PubMed]
- Varkuti, B.H.; Kepiro, M.; Horvath, I.A.; Vegner, L.; Rati, S.; Zsigmond, A.; Hegyi, G.; Lenkei, Z.; Varga, M.; Malnasi-Csizmadia, A. A highly soluble, non-phototoxic, non-fluorescent blebbistatin derivative. Sci. Rep. 2016, 6, 26141. [Google Scholar] [CrossRef] [PubMed]
- Young, E.J.; Blouin, A.M.; Briggs, S.B.; Sillivan, S.E.; Lin, L.; Cameron, M.D.; Rumbaugh, G.; Miller, C.A. Nonmuscle myosin IIB as a therapeutic target for the prevention of relapse to methamphetamine use. Mol. Psychiatry 2016, 21, 615–623. [Google Scholar] [CrossRef] [PubMed]
- Young, E.J.; Briggs, S.B.; Rumbaugh, G.; Miller, C.A. Nonmuscle myosin II inhibition disrupts methamphetamine-associated memory in females and adolescents. Neurobiol. Learn. Mem. 2017, 139, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Green, E.M.; Wakimoto, H.; Anderson, R.L.; Evanchik, M.J.; Gorham, J.M.; Harrison, B.C.; Henze, M.; Kawas, R.; Oslob, J.D.; Rodriguez, H.M.; et al. A small-molecule inhibitor of sarcomere contractility suppresses hypertrophic cardiomyopathy in mice. Science 2016, 351, 617–621. [Google Scholar] [CrossRef] [PubMed]
- Spudich, J.A. Hypertrophic and dilated cardiomyopathy: Four decades of basic research on muscle lead to potential therapeutic approaches to these devastating genetic diseases. Biophys. J. 2014, 106, 1236–1249. [Google Scholar] [CrossRef] [PubMed]
- Trivedi, D.V.; Adhikari, A.S.; Sarkar, S.S.; Ruppel, K.M.; Spudich, J.A. Hypertrophic cardiomyopathy and the myosin mesa: Viewing an old disease in a new light. Biophys. Rev. 2018, 10, 27–48. [Google Scholar] [CrossRef] [PubMed]
- Spudich, J.A.; Aksel, T.; Bartholomew, S.R.; Nag, S.; Kawana, M.; Yu, E.C.; Sarkar, S.S.; Sung, J.; Sommese, R.F.; Sutton, S.; et al. Effects of hypertrophic and dilated cardiomyopathy mutations on power output by human β-cardiac myosin. J. Exp. Biol. 2016, 219 Pt 2, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Marston, S.B. How do mutations in contractile proteins cause the primary familial cardiomyopathies? J. Cardiovasc. Transl. Res. 2011, 4, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Memo, M.; Marston, S. Skeletal muscle myopathy mutations at the actin tropomyosin interface that cause gain- or loss-of-function. J. Muscle Res. Cell Motil. 2013, 34, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Teekakirikul, P.; Padera, R.F.; Seidman, J.G.; Seidman, C.E. Hypertrophic cardiomyopathy: Translating cellular cross talk into therapeutics. J. Cell Biol. 2012, 199, 417–421. [Google Scholar] [CrossRef] [PubMed]
- Månsson, A. Hypothesis and theory: Mechanical instabilities and non-uniformities in hereditary sarcomere myopathies. Front. Physiol. 2014, 5, 350. [Google Scholar] [PubMed]
- Marston, S.; Memo, M.; Messer, A.; Papadaki, M.; Nowak, K.; McNamara, E.; Ong, R.; El-Mezgueldi, M.; Li, X.; Lehman, W. Mutations in repeating structural motifs of tropomyosin cause gain of function in skeletal muscle myopathy patients. Hum. Mol. Genet. 2013, 22, 4978–4987. [Google Scholar] [CrossRef] [PubMed]
- Bai, F.; Wang, L.; Kawai, M. A study of tropomyosin’s role in cardiac function and disease using thin-filament reconstituted myocardium. J. Muscle Res. Cell Motil. 2013, 34, 295–310. [Google Scholar] [CrossRef] [PubMed]
- Nag, S.; Trivedi, D.V.; Sarkar, S.S.; Adhikari, A.S.; Sunitha, M.S.; Sutton, S.; Ruppel, K.M.; Spudich, J.A. The myosin mesa and the basis of hypercontractility caused by hypertrophic cardiomyopathy mutations. Nat. Struct. Mol. Biol. 2017, 24, 525–533. [Google Scholar] [CrossRef] [PubMed]
- Houdusse, A.; Sweeney, H.L. How Myosin Generates Force on Actin Filaments. Trends Biochem. Sci. 2016, 41, 989–997. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.; Naber, N.; Pate, E.; Cooke, R. The myosin inhibitor blebbistatin stabilizes the super-relaxed state in skeletal muscle. Biophys. J. 2014, 107, 1637–1646. [Google Scholar] [CrossRef] [PubMed]
- Hooijman, P.; Stewart, M.A.; Cooke, R. A new state of cardiac myosin with very slow ATP turnover: A potential cardioprotective mechanism in the heart. Biophys. J. 2011, 100, 1969–1976. [Google Scholar] [CrossRef] [PubMed]
- Margossian, S.S.; Lowey, S. Preparation of myosin and its subfragments from rabbit skeletal muscle. Methods Enzymol. 1982, 85 Pt B, 55–71. [Google Scholar] [PubMed]
- Kron, S.J.; Toyoshima, Y.Y.; Uyeda, T.Q.; Spudich, J.A. Assays for actin sliding movement over myosin-coated surfaces. Methods Enzymol. 1991, 196, 399–416. [Google Scholar] [PubMed]
- Tsiavaliaris, G.; Fujita-Becker, S.; Batra, R.; Levitsky, D.I.; Kull, F.J.; Geeves, M.A.; Manstein, D.J. Mutations in the relay loop region result in dominant-negative inhibition of myosin II function in Dictyostelium. EMBO Rep. 2002, 3, 1099–1105. [Google Scholar] [CrossRef] [PubMed]
- Anson, M.; Geeves, M.A.; Kurzawa, S.E.; Manstein, D.J. Myosin motors with artificial lever arms. EMBO J. 1996, 15, 6069–6074. [Google Scholar] [PubMed]
- Sung, J.; Mortensen, K.I.; Spudich, J.A.; Flyvbjerg, H. How to Measure Load-Dependent Kinetics of Individual Motor Molecules Without a Force-Clamp. Methods Enzymol. 2017, 582, 1–29. [Google Scholar] [PubMed]
- Sommese, R.F.; Sung, J.; Nag, S.; Sutton, S.; Deacon, J.C.; Choe, E.; Leinwand, L.A.; Ruppel, K.; Spudich, J.A. Molecular consequences of the R453C hypertrophic cardiomyopathy mutation on human β-cardiac myosin motor function. Proc. Natl. Acad. Sci. USA 2013, 110, 12607–12612. [Google Scholar] [CrossRef] [PubMed]
- Rock, R.S.; Rief, M.; Mehta, A.D.; Spudich, J.A. In vitro assays of processive myosin motors. Methods 2000, 22, 373–381. [Google Scholar] [CrossRef] [PubMed]
- Sheetz, M.P.; Spudich, J.A. Movement of myosin-coated structures on actin cables. Cell Motil. 1983, 3, 485–489. [Google Scholar] [CrossRef] [PubMed]
- Spudich, J.A.; Kron, S.J.; Sheetz, M.P. Movement of myosin-coated beads on oriented filaments reconstituted from purified actin. Nature 1985, 315, 584–586. [Google Scholar] [CrossRef] [PubMed]
- Kron, S.J.; Spudich, J.A. Fluorescent Actin-Filaments Move on Myosin Fixed to a Glass-Surface. Proc. Natl. Acad. Sci. USA 1986, 83, 6272–6276. [Google Scholar] [CrossRef] [PubMed]
- Kishino, A.; Yanagida, T. Force measurements by micromanipulation of a single actin filament by glass needles. Nature 1988, 334, 74–76. [Google Scholar] [CrossRef] [PubMed]
- Ashkin, A.; Dziedzic, J.M.; Yamane, T. Optical trapping and manipulation of single cells using infrared laser beams. Nature 1987, 330, 769–771. [Google Scholar] [CrossRef] [PubMed]
- Ashkin, A. Applications of laser radiation pressure. Science 1980, 210, 1081–1088. [Google Scholar] [CrossRef] [PubMed]
- Svoboda, K.; Schmidt, C.F.; Schnapp, B.J.; Block, S.M. Direct observation of kinesin stepping by optical trapping interferometry. Nature 1993, 365, 721–727. [Google Scholar] [CrossRef] [PubMed]
- Block, S.M.; Goldstein, L.S.; Schnapp, B.J. Bead movement by single kinesin molecules studied with optical tweezers. Nature 1990, 348, 348–352. [Google Scholar] [CrossRef] [PubMed]
- Finer, J.T.; Simmons, R.M.; Spudich, J.A. Single myosin molecule mechanics: Piconewton forces and nanometre steps. Nature 1994, 368, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Molloy, J.E.; Burns, J.E.; Kendrick-Jones, J.; Tregear, R.T.; White, D.C. Movement and force produced by a single myosin head. Nature 1995, 378, 209–212. [Google Scholar] [CrossRef] [PubMed]
- Veigel, C.; Coluccio, L.M.; Jontes, J.D.; Sparrow, J.C.; Milligan, R.A.; Molloy, J.E. The motor protein myosin-I produces its working stroke in two steps. Nature 1999, 398, 530–533. [Google Scholar] [CrossRef] [PubMed]
- Veigel, C.; Wang, F.; Bartoo, M.L.; Sellers, J.R.; Molloy, J.E. The gated gait of the processive molecular motor, myosin V. Nat. Cell Biol. 2002, 4, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Veigel, C.; Molloy, J.E.; Schmitz, S.; Kendrick-Jones, J. Load-dependent kinetics of force production by smooth muscle myosin measured with optical tweezers. Nat. Cell Biol. 2003, 5, 980–986. [Google Scholar] [CrossRef] [PubMed]
- Steffen, W.; Smith, D.; Simmons, R.; Sleep, J. Mapping the actin filament with myosin. Proc. Natl. Acad. Sci. USA 2001, 98, 14949–14954. [Google Scholar] [CrossRef] [PubMed]
- Capitanio, M.; Canepari, M.; Cacciafesta, P.; Lombardi, V.; Cicchi, R.; Maffei, M.; Pavone, F.S.; Bottinelli, R. Two independent mechanical events in the interaction cycle of skeletal muscle myosin with actin. Proc. Natl. Acad. Sci. USA 2006, 103, 87–92. [Google Scholar] [CrossRef] [PubMed]
- Capitanio, M.; Canepari, M.; Maffei, M.; Beneventi, D.; Monico, C.; Vanzi, F.; Bottinelli, R.; Pavone, F.S. Ultrafast force-clamp spectroscopy of single molecules reveals load dependence of myosin working stroke. Nat. Methods 2012, 9, 1013–1019. [Google Scholar] [CrossRef] [PubMed]
- Sung, J.; Nag, S.; Mortensen, K.I.; Vestergaard, C.L.; Sutton, S.; Ruppel, K.; Flyvbjerg, H.; Spudich, J.A. Harmonic force spectroscopy measures load-dependent kinetics of individual human β-cardiac myosin molecules. Nat. Commun. 2015, 6, 7931. [Google Scholar] [CrossRef] [PubMed]
- Ishijima, A.; Kojima, H.; Funatsu, T.; Tokunaga, M.; Higuchi, H.; Tanaka, H.; Yanagida, T. Simultaneous observation of individual ATPase and mechanical events by a single myosin molecule during interaction with actin. Cell 1998, 92, 161–171. [Google Scholar] [CrossRef]
- Kaya, M.; Higuchi, H. Nonlinear elasticity and an 8-nm working stroke of single myosin molecules in myofilaments. Science 2010, 329, 686–689. [Google Scholar] [CrossRef] [PubMed]
- Abbondanzieri, E.A.; Greenleaf, W.J.; Shaevitz, J.W.; Landick, R.; Block, S.M. Direct observation of base-pair stepping by RNA polymerase. Nature 2005, 438, 460–465. [Google Scholar] [CrossRef] [PubMed]
- Fazal, F.M.; Meng, C.A.; Murakami, K.; Kornberg, R.D.; Block, S.M. Real-time observation of the initiation of RNA polymerase II transcription. Nature 2015, 525, 274–277. [Google Scholar] [CrossRef] [PubMed]
- Wruck, F.; Katranidis, A.; Nierhaus, K.H.; Buldt, G.; Hegner, M. Translation and folding of single proteins in real time. Proc. Natl. Acad. Sci. USA 2017, 114, E4399–E4407. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.L.; Bax, N.A.; Buckley, C.D.; Weis, W.I.; Dunn, A.R. Vinculin forms a directionally asymmetric catch bond with F-actin. Science 2017, 357, 703–706. [Google Scholar] [CrossRef] [PubMed]
- Capitanio, M.; Pavone, F.S. Interrogating biology with force: Single molecule high-resolution measurements with optical tweezers. Biophys. J. 2013, 105, 1293–1303. [Google Scholar] [CrossRef] [PubMed]
- Perkins, T.T. Angstrom-precision optical traps and applications. Ann. Rev. Biophys. 2014, 43, 279–302. [Google Scholar] [CrossRef] [PubMed]
- Rayment, I.; Holden, H.M.; Whittaker, M.; Yohn, C.B.; Lorenz, M.; Holmes, K.C.; Milligan, R.A. Structure of the actin-myosin complex and its implications for muscle contraction. Science 1993, 261, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Rayment, I.; Rypniewski, W.R.; Schmidt-Base, K.; Smith, R.; Tomchick, D.R.; Benning, M.M.; Winkelmann, D.A.; Wesenberg, G.; Holden, H.M. Three-dimensional structure of myosin subfragment-1: A molecular motor. Science 1993, 261, 50–58. [Google Scholar] [CrossRef] [PubMed]
- Jontes, J.D.; Wilson-Kubalek, E.M.; Milligan, R.A. A 32 degree tail swing in brush border myosin I on ADP release. Nature 1995, 378, 751–753. [Google Scholar] [CrossRef] [PubMed]
- Whittaker, M.; Wilson-Kubalek, E.M.; Smith, J.E.; Faust, L.; Milligan, R.A.; Sweeney, H.L. A 35-A movement of smooth muscle myosin on ADP release. Nature 1995, 378, 748–751. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, M.J.; Shuman, H.; Ostap, E.M. Inherent force-dependent properties of β-cardiac myosin contribute to the force-velocity relationship of cardiac muscle. Biophys. J. 2014, 107, L41–L44. [Google Scholar] [CrossRef] [PubMed]
- Albet-Torres, N.; Bloemink, M.J.; Barman, T.; Candau, R.; Frölander, K.; Geeves, M.A.; Golker, K.; Herrmann, C.; Lionne, C.; Piperio, C.; et al. Drug effect unveils inter-head cooperativity and strain-dependent ADP release in fast skeletal actomyosin. J. Biol. Chem. 2009, 284, 22926–22937. [Google Scholar] [CrossRef] [PubMed]
- Llinas, P.; Isabet, T.; Song, L.; Ropars, V.; Zong, B.; Benisty, H.; Sirigu, S.; Morris, C.; Kikuti, C.; Safer, D.; et al. How actin initiates the motor activity of Myosin. Dev. Cell 2015, 33, 401–412. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Dudko, O.K. A transformation for the mechanical fingerprints of complex biomolecular interactions. Proc. Natl. Acad. Sci. USA 2013, 110, 16432–16437. [Google Scholar] [CrossRef] [PubMed]
- Sellers, J.R.; Kachar, B. Polarity and velocity of sliding filaments: Control of direction by actin and of speed by myosin. Science 1990, 249, 406–408. [Google Scholar] [CrossRef] [PubMed]
- Yamada, A.; Ishii, N.; Takahashi, K. Direction and speed of actin filaments moving along thick filaments isolated from molluscan smooth muscle. J. Biochem. 1990, 108, 341–343. [Google Scholar] [CrossRef] [PubMed]
- Yamada, A.; Yoshio, M.; Nakayama, H. Bi-directional movement of actin filaments along long bipolar tracks of oriented rabbit skeletal muscle myosin molecules. FEBS Lett. 1997, 409, 380–384. [Google Scholar] [CrossRef]
- Tanaka, H.; Ishijima, A.; Honda, M.; Saito, K.; Yanagida, T. Orientation dependence of displacements by a single one-headed myosrelative to the actin filament. Biophys. J. 1998, 75, 1886–1894. [Google Scholar] [CrossRef]
- Molloy, J.E.; Burns, J.E.; Sparrow, J.C.; Tregear, R.T.; Kendrick-Jones, J.; White, D.C. Single-molecule mechanics of heavy meromyosin and S1 interacting with rabbit or Drosophila actins using optical tweezers. Biophys. J. 1995, 68 (Suppl. 4), 298S–305S. [Google Scholar] [PubMed]
- Mehta, A.D.; Finer, J.T.; Spudich, J.A. Detection of single-molecule interactions using correlated thermal diffusion. Proc. Natl. Acad. Sci. USA 1997, 94, 7927–7931. [Google Scholar] [CrossRef] [PubMed]
- Ishijima, A.; Harada, Y.; Kojima, H.; Funatsu, T.; Higuchi, H.; Yanagida, T. Single-molecule analysis of the actomyosin motor using nano-manipulation. Biochem. Biophys. Res. Commun. 1994, 199, 1057–1063. [Google Scholar] [CrossRef] [PubMed]
- Guo, B.; Guilford, W.H. Mechanics of actomyosin bonds in different nucleotide states are tuned to muscle contraction. Proc. Natl. Acad. Sci. USA 2006, 103, 9844–9849. [Google Scholar] [CrossRef] [PubMed]
- Nishizaka, T.; Seo, R.; Tadakuma, H.; Kinosita, K., Jr.; Ishiwata, S. Characterization of single actomyosin rigor bonds: Load dependence of lifetime and mechanical properties. Biophys. J. 2000, 79, 962–974. [Google Scholar] [CrossRef]
- Toyoshima, Y.Y.; Kron, S.J.; McNally, E.M.; Niebling, K.R.; Toyoshima, C.; Spudich, J.A. Myosin subfragment-1 is sufficient to move actin filaments in vitro. Nature 1987, 328, 536–539. [Google Scholar] [CrossRef] [PubMed]
- Guo, B.; Guilford, W.H. The tail of myosin reduces actin filament velocity in the in vitro motility assay. Cell Motil. Cytoskel. 2004, 59, 264–272. [Google Scholar] [CrossRef] [PubMed]
- Toyoshima, Y.Y. How are myosin fragments bound to nitrocellulose film? Adv. Exp. Med. Biol. 1993, 332, 259–265. [Google Scholar] [PubMed]
- Iwane, A.H.; Kitamura, K.; Tokunaga, M.; Yanagida, T. Myosin subfragment-1 is fully equipped with factors essential for motor function. Biochem. Biophys. Res. Commun. 1997, 230, 76–80. [Google Scholar] [CrossRef] [PubMed]
- Albet-Torres, N.; O’Mahony, J.; Charlton, C.; Balaz, M.; Lisboa, P.; Aastrup, T.; Månsson, A.; Nicholls, I.A. Mode of heavy meromyosin adsorption and motor function correlated with surface hydrophobicity and charge. Langmuir 2007, 23, 11147–11156. [Google Scholar] [CrossRef] [PubMed]
- Persson, M.; Albet-Torres, N.; Sundberg, M.; Ionov, L.; Diez, S.; Höök, F.; Månsson, A.; Balaz, M. Heavy Meromyosin Molecules Extend more than 50 nm above Adsorbing Electronegative Surfaces. Langmuir 2010, 26, 9927–9936. [Google Scholar] [CrossRef] [PubMed]
- Balaz, M.; Sundberg, M.; Persson, M.; Kvassman, J.; Månsson, A. Effects of Surface Adsorption on Catalytic Activity of Heavy Meromyosin Studied using Fluorescent ATP Analogue. Biochemistry 2007, 46, 7233–7251. [Google Scholar] [CrossRef] [PubMed]
- Jaber, J.A.; Chase, P.B.; Schlenoff, J.B. Actomyosin-driven motility on patterned polyelectrolyte mono- and multilayers. Nano Lett. 2003, 3, 1505–1509. [Google Scholar] [CrossRef]
- Albet-Torres, N.; Gunnarsson, A.; Persson, M.; Balaz, M.; Höök, F.; Månsson, A. Molecular motors on lipid bilayers and silicon dioxide: Different driving forces for adsorption. Soft Matter 2010, 6, 3211–3219. [Google Scholar] [CrossRef]
- Van Zalinge, H.; Ramsey, L.C.; Aveyard, J.; Persson, M.; Månsson, A.; Nicolau, D.V. Surface-Controlled Properties of Myosin Studied by Electric Field Modulation. Langmuir 2015, 31, 8354–8361. [Google Scholar] [CrossRef] [PubMed]
- Van Zalinge, H.; Aveyard, J.; Hajne, J.; Persson, M.; Månsson, A.; Nicolau, D.V. Actin filament motility induced variation of resonance frequency and rigidity of polymer surfaces studied by quartz crystal microbalance. Langmuir 2012, 28, 15033–15037. [Google Scholar] [CrossRef] [PubMed]
- Hanson, K.L.; Fulga, F.; Dobroiu, S.; Solana, G.; Kaspar, O.; Tokarova, V.; Nicolau, D.V. Polymer surface properties control the function of heavy meromyosin in dynamic nanodevices. Biosens. Bioelectron. 2017, 93, 305–314. [Google Scholar] [CrossRef] [PubMed]
- Dupuis, D.E.; Guilford, W.H.; Wu, J.; Warshaw, D.M. Actin filament mechanics in the laser trap. J. Muscle Res. Cell Motil. 1997, 18, 17–30. [Google Scholar] [CrossRef] [PubMed]
- Tyska, M.J.; Dupuis, D.E.; Guilford, W.H.; Patlak, J.B.; Waller, G.S.; Trybus, K.M.; Warshaw, D.M.; Lowey, S. Two heads of myosin are better than one for generating force and motion. Proc. Natl. Acad. Sci. USA 1999, 96, 4402–4407. [Google Scholar] [CrossRef] [PubMed]
- Takagi, Y.; Homsher, E.E.; Goldman, Y.E.; Shuman, H. Force generation in single conventional actomyosin complexes under high dynamic load. Biophys. J. 2006, 90, 1295–1307. [Google Scholar] [CrossRef] [PubMed]
- Kaya, M.; Higuchi, H. Stiffness, working stroke, and force of single-myosin molecules in skeletal muscle: Elucidation of these mechanical properties via nonlinear elasticity evaluation. Cell. Mol. Life Sci. 2013, 70, 4275–4292. [Google Scholar] [CrossRef] [PubMed]
- Hoppe, T.; Minton, A.P. An equilibrium model for the combined effect of macromolecular crowding and surface adsorption on the formation of linear protein fibrils. Biophys. J. 2015, 108, 957–966. [Google Scholar] [CrossRef] [PubMed]
- Ge, J.; Bouriyaphone, S.D.; Serebrennikova, T.A.; Astashkin, A.V.; Nesmelov, Y.E. Macromolecular Crowding Modulates Actomyosin Kinetics. Biophys. J. 2016, 111, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Nyitrai, M.; Rossi, R.; Adamek, N.; Pellegrino, M.A.; Bottinelli, R.; Geeves, M.A. What limits the velocity of fast-skeletal muscle contraction in mammals? J. Mol. Biol. 2006, 355, 432–442. [Google Scholar] [CrossRef] [PubMed]
- Ranatunga, K.W. Force and power generating mechanism(s) in active muscle as revealed from temperature perturbation studies. J Physiol. 2010, 588 Pt 19, 3657–3670. [Google Scholar] [CrossRef] [PubMed]
- Brenner, B.; Schoenberg, M.; Chalovich, J.M.; Greene, L.E.; Eisenberg, E. Evidence for cross-bridge attachment in relaxed muscle at low ionic strength. Proc. Natl. Acad. Sci. USA 1982, 79, 7288–7291. [Google Scholar] [CrossRef] [PubMed]
- Homsher, E.; Wang, F.; Sellers, J.R. Factors affecting movement of F-actin filaments propelled by skeletal muscle heavy meromyosin. Am. J. Physiol. 1992, 262 Pt 1, C714–C723. [Google Scholar] [CrossRef] [PubMed]
- Vikhorev, P.G.; Vikhoreva, N.N.; Månsson, A. Bending flexibility of actin filaments during motor-induced sliding. Biophys. J. 2008, 95, 5809–5819. [Google Scholar] [CrossRef] [PubMed]
- Hook, P.; Larsson, L. Actomyosin interactions in a novel single muscle fiber in vitro motility assay. J. Muscle Res. Cell Motil. 2000, 21, 357–365. [Google Scholar] [CrossRef] [PubMed]
- Homsher, E.; Nili, M.; Chen, I.Y.; Tobacman, L.S. Regulatory proteins alter nucleotide binding to acto-myosin of sliding filaments in motility assays. Biophys. J. 2003, 85, 1046–1052. [Google Scholar] [CrossRef]
- Ranatunga, K.W. Temperature Effects on Force and Actin(-)Myosin Interaction in Muscle: A Look Back on Some Experimental Findings. Int. J. Mol. Sci. 2018, 19, 1538. [Google Scholar] [CrossRef] [PubMed]
- Homsher, E.; Lee, D.M.; Morris, C.; Pavlov, D.; Tobacman, L.S. Regulation of force and unloaded sliding speed in single thin filaments: Effects of regulatory proteins and calcium. J. Physiol. 2000, 524 Pt 1, 233–243. [Google Scholar] [CrossRef] [PubMed]
- Kad, N.M.; Kim, S.; Warshaw, D.M.; VanBuren, P.; Baker, J.E. Single-myosin crossbridge interactions with actin filaments regulated by troponin-tropomyosin. Proc. Natl. Acad. Sci. USA 2005, 102, 16990–16995. [Google Scholar] [CrossRef] [PubMed]
- Fusi, L.; Brunello, E.; Yan, Z.; Irving, M. Thick filament mechano-sensing is a calcium-independent regulatory mechanism in skeletal muscle. Nat. Commun. 2016, 7, 13281. [Google Scholar] [CrossRef] [PubMed]
- Linari, M.; Brunello, E.; Reconditi, M.; Fusi, L.; Caremani, M.; Narayanan, T.; Piazzesi, G.; Lombardi, V.; Irving, M. Force generation by skeletal muscle is controlled by mechanosensing in myosin filaments. Nature 2015, 528, 276–279. [Google Scholar] [CrossRef] [PubMed]
- McNamara, J.W.; Li, A.; Smith, N.J.; Lal, S.; Graham, R.M.; Kooiker, K.B.; van Dijk, S.J.; Remedios, C.G.D.; Harris, S.P.; Cooke, R. Ablation of cardiac myosin binding protein-C disrupts the super-relaxed state of myosin in murine cardiomyocytes. J. Mol. Cell. Cardiol. 2016, 94, 65–71. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; White, H.D.; Offer, G.W.; Yu, L.C. Stabilization of helical order in the thick filaments by blebbistatin: Further evidence of coexisting multiple conformations of myosin. Biophys. J. 2009, 96, 3673–3681. [Google Scholar] [CrossRef] [PubMed]
- Kawana, M.; Sarkar, S.S.; Sutton, S.; Ruppel, K.M.; Spudich, J.A. Biophysical properties of human β-cardiac myosin with converter mutations that cause hypertrophic cardiomyopathy. Sci. Adv. 2017, 3, e1601959. [Google Scholar] [CrossRef] [PubMed]
- Gangadharan, B.; Sunitha, M.S.; Mukherjee, S.; Chowdhury, R.R.; Haque, F.; Sekar, N.; Sowdhamini, R.; Spudich, J.A.; Mercer, J.A. Molecular mechanisms and structural features of cardiomyopathy-causing troponin T mutants in the tropomyosin overlap region. Proc. Natl. Acad. Sci. USA 2017, 114, 11115–11120. [Google Scholar] [CrossRef] [PubMed]
- Homburger, J.R.; Green, E.M.; Caleshu, C.; Sunitha, M.S.; Taylor, R.E.; Ruppel, K.M.; Metpally, R.P.; Colan, S.D.; Michels, M.; Day, S.M.; et al. Multidimensional structure-function relationships in human β-cardiac myosin from population-scale genetic variation. Proc. Natl. Acad. Sci. USA 2016, 113, 6701–6706. [Google Scholar] [CrossRef] [PubMed]
- Adhikari, A.S.; Kooiker, K.B.; Sarkar, S.S.; Liu, C.; Bernstein, D.; Spudich, J.A.; Ruppel, K.M. Early-Onset Hypertrophic Cardiomyopathy Mutations Significantly Increase the Velocity, Force, and Actin-Activated ATPase Activity of Human β-Cardiac Myosin. Cell Rep. 2016, 17, 2857–2864. [Google Scholar] [CrossRef] [PubMed]
- Spudich, J.A. The myosin mesa and a possible unifying hypothesis for the molecular basis of human hypertrophic cardiomyopathy. Biochem. Soc. Trans. 2015, 43, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Kraft, T.; Witjas-Paalberends, E.R.; Boontje, N.M.; Tripathi, S.; Brandis, A.; Montag, J.; Hodgkinson, J.L.; Francino, A.; Navarro-Lopez, F.; Brenner, B.; et al. Familial hypertrophic cardiomyopathy: Functional effects of myosin mutation R723G in cardiomyocytes. J. Mol. Cell. Cardiol. 2013, 57, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, M.J.; Kazmierczak, K.; Szczesna-Cordary, D.; Moore, J.R. Cardiomyopathy-linked myosin regulatory light chain mutations disrupt myosin strain-dependent biochemistry. Proc. Natl. Acad. Sci. USA 2010, 107, 17403–17408. [Google Scholar] [CrossRef] [PubMed]
- Yuan, C.C.; Kazmierczak, K.; Liang, J.; Zhou, Z.; Yadav, S.; Gomes, A.V.; Irving, T.C.; Szczesna-Cordary, D. Sarcomeric perturbations of myosin motors lead to dilated cardiomyopathy in genetically modified MYL2 mice. Proc. Natl. Acad. Sci. USA 2018, 115, E2338–E2347. [Google Scholar] [CrossRef] [PubMed]
- Szczesna-Cordary, D.; de Tombe, P.P. Myosin light chain phosphorylation, novel targets to repair a broken heart? Cardiovasc. Res. 2016, 111, 5–7. [Google Scholar] [CrossRef] [PubMed]
- Stewart, M.; Franks-Skiba, K.; Cooke, R. Myosin regulatory light chain phosphorylation inhibits shortening velocities of skeletal muscle fibers in the presence of the myosin inhibitor blebbistatin. J. Muscle Res. Cell Motil. 2009, 30, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Kamm, K.E.; Stull, J.T. Signaling to myosin regulatory light chain in sarcomeres. J. Biol. Chem. 2011, 286, 9941–9947. [Google Scholar] [CrossRef] [PubMed]
- Woody, M.S.; Capitanio, M.; Ostap, E.M.; Goldman, Y.E. Electro-optic deflectors deliver advantages over acousto-optical deflectors in a high resolution, ultra-fast force-clamp optical trap. Opt. Express 2018, 26, 11181–11193. [Google Scholar] [CrossRef] [PubMed]
- Guilford, W.H.; Dupuis, D.E.; Kennedy, G.; Wu, J.; Patlak, J.B.; Warshaw, D.M. Smooth muscle and skeletal muscle myosins produce similar unitary forces and displacements in the laser trap. Biophys. J. 1997, 72, 1006–1021. [Google Scholar] [CrossRef]
- Smith, D.A.; Steffen, W.; Simmons, R.M.; Sleep, J. Hidden-markov methods for the analysis of single-molecule actomyosin displacement data: The variance-hidden-markov method. Biophys. J. 2001, 81, 2795–2816. [Google Scholar] [CrossRef]
- Shalabi, N.; Cornachione, A.; de Souza Leite, F.; Vengallatore, S.; Rassier, D.E. Residual force enhancement is regulated by titin in skeletal and cardiac myofibrils. J. Physiol. 2017, 595, 2085–2098. [Google Scholar] [CrossRef] [PubMed]
- Rassier, D.E.; Pavlov, I. Force produced by isolated sarcomeres and half-sarcomeres after an imposed stretch. Am. J. Physiol. Cell Physiol. 2012, 302, C240–C248. [Google Scholar] [CrossRef] [PubMed]
- Piroddi, N.; Belus, A.; Scellini, B.; Tesi, C.; Giunti, G.; Cerbai, E.; Mugelli, A.; Poggesi, C. Tension generation and relaxation in single myofibrils from human atrial and ventricular myocardium. Pflugers Arch. 2007, 454, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Stehle, R.; Solzin, J.; Iorga, B.; Poggesi, C. Insights into the kinetics of Ca(2+)-regulated contraction and relaxation from myofibril studies. Pflugers Arch. 2009, 458, 337–357. [Google Scholar] [CrossRef] [PubMed]
- Huxley, A.F.; Simmons, R.M. Proposed mechanism of force generation in striated muscle. Nature 1971, 233, 533–538. [Google Scholar] [CrossRef] [PubMed]
- Ford, L.E.; Huxley, A.F.; Simmons, R.M. Tension responses to sudden length change in stimulated frog muscle fibres near slack length. J. Physiol. 1977, 269, 441–515. [Google Scholar] [CrossRef] [PubMed]
- Edman, K.A.P. Double-hyperbolic force-velocity relation in frog muscle fibres. J. Physiol. (Lond.) 1988, 404, 301–321. [Google Scholar] [CrossRef]
- Edman, K.A.P.; Flitney, F.W. Laser diffraction studies of sarcomere dynamics during ‘isometric’ relaxation in isolated muscle fibres of the frog. J. Physiol. (Lond.) 1982, 329, 1–20. [Google Scholar] [CrossRef]
- Edman, K.A.P.; Mulieri, L.A.; Scubon-Mulieri, B. Non-hyperbolic force-velocity relationship in single muscle fibres. Acta Physiol. Scand. 1976, 98, 143–156. [Google Scholar] [CrossRef] [PubMed]
- Ford, L.E.; Huxley, A.F.; Simmons, R.M. Tension transients during steady shortening of frog muscle fibres. J. Physiol. (Lond.) 1985, 361, 131–150. [Google Scholar] [CrossRef]
- Ford, L.E.; Huxley, A.F.; Simmons, R.M. The relation between stiffness and filament overlap in stimulated frog muscle fibres. J. Physiol. (Lond.) 1981, 311, 219–249. [Google Scholar] [CrossRef]
- Fenn, W.O. A quantitative comparison between the energy liberated and the work performed by the isolated sartorius muscle of the frog. J. Physiol. 1923, 58, 175–203. [Google Scholar] [CrossRef] [PubMed]
- Hill, A.V. The heat of shortening and the dynamic constants of muscle. Proc. R. Soc. B 1938, 126, 136–195. [Google Scholar] [CrossRef]
- Månsson, A.; Edman, K.A.P. Effects of amrinone on twitch, tetanus and force-velocity relationship in frog skeletal muscle. Acta Physiol. Scand. 1984, 120, 473–475. [Google Scholar] [CrossRef] [PubMed]
- Lombardi, V.; Piazzesi, G. The contractile response during steady lengthening of stimulated frog muscle fibres. J. Physiol. (Lond.) 1990, 431, 141–171. [Google Scholar] [CrossRef]
- Piazzesi, G.; Francini, F.; Linari, M.; Lombardi, V. Tension transients during steady lengthening of tetanized muscle fibres of the frog. J. Physiol. (Lond.) 1992, 445, 659–711. [Google Scholar] [CrossRef]
- Curtin, N.A.; Davies, R.E. Very high tension with very little ATP breakdown by active skeletal muscle. J. Mechanochem. Cell Motil. 1975, 3, 147–154. [Google Scholar] [PubMed]
- Brunello, E.; Reconditi, M.; Elangovan, R.; Linari, M.; Sun, Y.B.; Narayanan, T.; Panine, P.; Piazzesi, G.; Irving, M.; Lombardi, V. Skeletal muscle resists stretch by rapid binding of the second motor domain of myosin to actin. Proc. Natl. Acad. Sci. USA 2007, 104, 20114–20119. [Google Scholar] [CrossRef] [PubMed]
- Piazzesi, G.; Reconditi, M.; Linari, M.; Lucii, L.; Bianco, P.; Brunello, E.; Decostre, V.; Stewart, A.; Gore, D.B.; Irving, T.C.; et al. Skeletal muscle performance determined by modulation of number of Myosin motors rather than motor force or stroke size. Cell 2007, 131, 784–795. [Google Scholar] [CrossRef] [PubMed]
- Caremani, M.; Melli, L.; Dolfi, M.; Lombardi, V.; Linari, M. The working stroke of the myosin II motor in muscle is not tightly coupled to release of orthophosphate from its active site. J. Physiol. 2013, 591 Pt 20, 5187–5205. [Google Scholar] [CrossRef] [PubMed]
- Roots, H.; Pinniger, G.J.; Offer, G.W.; Ranatunga, K.W. Mechanism of force enhancement during and after lengthening of active muscle: A temperature dependence study. J. Muscle Res. Cell Motil. 2012, 33, 313–325. [Google Scholar] [CrossRef] [PubMed]
- Nocella, M.; Bagni, M.A.; Cecchi, G.; Colombini, B. Mechanism of force enhancement during stretching of skeletal muscle fibres investigated by high time-resolved stiffness measurements. J. Muscle Res. Cell Motil. 2013, 34, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Colombini, B.; Nocella, M.; Benelli, G.; Cecchi, G.; Bagni, M.A. Crossbridge properties during force enhancement by slow stretching in single intact frog muscle fibres. J. Physiol. 2007, 585 Pt 2, 607–615. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, P.J.; Ashley, C.C.; Bagni, M.A.; Maeda, Y.; Cecchi, G. Cross-bridge attachment and stiffness during isotonic shortening of intact single muscle fibers. Biophys. J. 1993, 64, 1150–1160. [Google Scholar] [CrossRef]
- Huxley, A.F. Muscle structure and theories of contraction. Prog. Biophys. Biophys. Chem. 1957, 7, 255–318. [Google Scholar] [PubMed]
- Eisenberg, E.; Hill, T.L. A cross-bridge model of muscle contraction. Prog. Biophys. Mol. Biol. 1978, 33, 55–82. [Google Scholar] [CrossRef]
- Eisenberg, E.; Hill, T.L.; Chen, Y. Cross-bridge model of muscle contraction. Quantitative analysis. Biophys. J. 1980, 29, 195–227. [Google Scholar] [CrossRef]
- Månsson, A. Actomyosin-ADP states, inter-head cooperativity and the force-velocity relation of skeletal muscle. Biophys. J. 2010, 98, 1237–1246. [Google Scholar] [CrossRef] [PubMed]
- Månsson, A. Actomyosin-based contraction: One mechanokinetic model from single molecules to muscle? J. Muscle Res. Cell Motil. 2016, 37, 181–194. [Google Scholar] [CrossRef] [PubMed]
- Duke, T.A. Molecular model of muscle contraction. Proc. Natl. Acad. Sci. USA 1999, 96, 2770–2775. [Google Scholar] [CrossRef] [PubMed]
- Offer, G.; Ranatunga, K.W. A cross-bridge cycle with two tension-generating steps simulates skeletal muscle mechanics. Biophys. J. 2013, 105, 928–940. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.A.; Mijailovich, S.M. Toward a unified theory of muscle contraction. II: Predictions with the mean-field approximation. Ann. Biomed. Eng. 2008, 36, 1353–1371. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.A.; Geeves, M.A.; Sleep, J.; Mijailovich, S.M. Towards a unified theory of muscle contraction. I: Foundations. Ann. Biomed. Eng. 2008, 36, 1624–1640. [Google Scholar] [CrossRef] [PubMed]
- Debold, E.P.; Patlak, J.B.; Warshaw, D.M. Slip sliding away: Load-dependence of velocity generated by skeletal muscle myosin molecules in the laser trap. Biophys. J. 2005, 89, L34–L36. [Google Scholar] [CrossRef] [PubMed]
- Debold, E.P.; Walcott, S.; Woodward, M.; Turner, M.A. Direct observation of phosphate inhibiting the force-generating capacity of a miniensemble of Myosin molecules. Biophys. J. 2013, 105, 2374–2384. [Google Scholar] [CrossRef] [PubMed]
- Minozzo, F.C.; Rassier, D.E. The effects of Ca2+ and MgADP on force development during and after muscle length changes. PLoS ONE 2013, 8, e68866. [Google Scholar] [CrossRef] [PubMed]
- Piazzesi, G.; Lucii, L.; Lombardi, V. The size and the speed of the working stroke of muscle myosin and its dependence on the force. J. Physiol. 2002, 545 Pt 1, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Roots, H.; Offer, G.W.; Ranatunga, K.W. Comparison of the tension responses to ramp shortening and lengthening in intact mammalian muscle fibres: Crossbridge and non-crossbridge contributions. J. Muscle Res.Cell Motil. 2007, 28, 123–139. [Google Scholar] [CrossRef] [PubMed]
- Minozzo, F.C.; Hilbert, L.; Rassier, D.E. Pre-power-stroke cross-bridges contribute to force transients during imposed shortening in isolated muscle fibers. PLoS ONE 2012, 7, e29356. [Google Scholar] [CrossRef] [PubMed]
- Edman, K.A. The force bearing capacity of frog muscle fibres during stretch: Its relation to sarcomere length and fibre width. J. Physiol. 1999, 519 Pt 2, 515–526. [Google Scholar] [CrossRef] [PubMed]
- Getz, E.B.; Cooke, R.; Lehman, S.L. Phase transition in force during ramp stretches of skeletal muscle. Biophys. J. 1998, 75, 2971–2983. [Google Scholar] [CrossRef]
- Rassier, D.E. Pre-power stroke cross bridges contribute to force during stretch of skeletal muscle myofibrils. Proc. Biol. Sci. R. Soc. 2008, 275, 2577–2586. [Google Scholar] [CrossRef] [PubMed]
- Minozzo, F.C.; Altman, D.; Rassier, D.E. MgADP activation contributes to force enhancement during fast stretch of isolated skeletal myofibrils. Biochem. Biophys. Res. Commun. 2015, 463, 1129–1134. [Google Scholar] [CrossRef] [PubMed]
- Edman, K.A. Residual force enhancement after stretch in striated muscle: A consequence of increased myofilament overlap? J. Physiol. 2012, 590 Pt 6, 1339–1345. [Google Scholar] [CrossRef] [PubMed]
- Cornachione, A.S.; Leite, F.; Bagni, M.A.; Rassier, D.E. The increase in non-cross-bridge forces after stretch of activated striated muscle is related to titin isoforms. Am. J. Physiol. Cell Physiol. 2016, 310, C19–C26. [Google Scholar] [CrossRef] [PubMed]
- Rassier, D.E. Residual force enhancement in skeletal muscles: One sarcomere after the other. J. Muscle Res. Cell Motil. 2012, 33, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Stehle, R. Force Responses and Sarcomere Dynamics of Cardiac Myofibrils Induced by Rapid Changes in [Pi]. Biophys. J. 2017, 112, 356–367. [Google Scholar] [CrossRef] [PubMed]
- Stehle, R.; Tesi, C. Kinetic coupling of phosphate release, force generation and rate-limiting steps in the cross-bridge cycle. J. Muscle Res. Cell Motil. 2017, 38, 275–289. [Google Scholar] [CrossRef] [PubMed]
- Kirschner, S.E.; Becker, E.; Antognozzi, M.; Kubis, H.P.; Francino, A.; Navarro-Lopez, F.; Bit-Avragim, N.; Perrot, A.; Mirrakhimov, M.M.; Osterziel, K.J.; et al. Hypertrophic cardiomyopathy-related β-myosin mutations cause highly variable calcium sensitivity with functional imbalances among individual muscle cells. Am. J. Physiol. Heart Circ. Physiol. 2005, 288, H1242–H1251. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, S.; Schultz, I.; Becker, E.; Montag, J.; Borchert, B.; Francino, A.; Navarro-Lopez, F.; Perrot, A.; Ozcelik, C.; Osterziel, K.J.; et al. Unequal allelic expression of wild-type and mutated β-myosin in familial hypertrophic cardiomyopathy. Basic Res. Cardiol. 2011, 106, 1041–1055. [Google Scholar] [CrossRef] [PubMed]
- Brenner, B.; Seebohm, B.; Tripathi, S.; Montag, J.; Kraft, T. Familial hypertrophic cardiomyopathy: Functional variance among individual cardiomyocytes as a trigger of FHC-phenotype development. Front. Physiol. 2014, 5, 392. [Google Scholar] [CrossRef] [PubMed]
- Montag, J.; Kowalski, K.; Makul, M.; Ernstberger, P.; Radocaj, A.; Beck, J.; Becker, E.; Tripathi, S.; Keyser, B.; Muhlfeld, C.; et al. Burst-Like Transcription of Mutant and Wildtype MYH7-Alleles as Possible Origin of Cell-to-Cell Contractile Imbalance in Hypertrophic Cardiomyopathy. Front. Physiol. 2018, 9, 359. [Google Scholar] [CrossRef] [PubMed]
- Montag, J.; Petersen, B.; Flogel, A.K.; Becker, E.; Lucas-Hahn, A.; Cost, G.J.; Muhlfeld, C.; Kraft, T.; Niemann, H.; Brenner, B. Successful knock-in of Hypertrophic Cardiomyopathy-mutation R723G into the MYH7 gene mimics HCM pathology in pigs. Sci. Rep. 2018, 8, 4786. [Google Scholar] [CrossRef] [PubMed]
- Montag, J.; Syring, M.; Rose, J.; Weber, A.L.; Ernstberger, P.; Mayer, A.K.; Becker, E.; Keyser, B.; Dos Remedios, C.; Perrot, A.; et al. Intrinsic MYH7 expression regulation contributes to tissue level allelic imbalance in hypertrophic cardiomyopathy. J. Muscle Res. Cell Motil. 2017, 38, 291–302. [Google Scholar] [CrossRef] [PubMed]
- Kraft, T.; Montag, J.; Radocaj, A.; Brenner, B. Hypertrophic Cardiomyopathy: Cell-to-Cell Imbalance in Gene Expression and Contraction Force as Trigger for Disease Phenotype Development. Circ. Res. 2016, 119, 992–995. [Google Scholar] [CrossRef] [PubMed]
- Offer, G.; Ranatunga, K.W. Crossbridge and filament compliance in muscle: Implications for tension generation and lever arm swing. J. Muscle Res. Cell Motil. 2010, 31, 245–265. [Google Scholar] [CrossRef] [PubMed]
- Barclay, C.J.; Woledge, R.C.; Curtin, N.A. Inferring crossbridge properties from skeletal muscle energetics. Prog. Biophys. Mol. Biol. 2010, 102, 53–71. [Google Scholar] [CrossRef] [PubMed]
- Huxley, H.E.; Stewart, A.; Sosa, H.; Irving, T. X-ray diffraction measurements of the extensibility of actin and myosin filaments in contracting muscle. Biophys. J. 1994, 67, 2411–2421. [Google Scholar] [CrossRef]
- Wakabayashi, K.; Sugimoto, Y.; Tanaka, H.; Ueno, Y.; Takezawa, Y.; Amemiya, Y. X-ray diffraction evidence for the extensibility of actin and myosin filaments during muscle contraction. Biophys. J. 1994, 67, 2422–2435. [Google Scholar] [CrossRef]
- Kojima, H.; Ishijima, A.; Yanagida, T. Direct measurement of stiffness of single actin filaments with and without tropomyosin by in vitro nanomanipulation. Proc. Natl. Acad. Sci. USA 1994, 91, 12962–12966. [Google Scholar] [CrossRef] [PubMed]
- Goldman, Y.E.; Huxley, A.F. Actin compliance: Are you pulling my chain? Biophys. J. 1994, 67, 2131–2133. [Google Scholar] [CrossRef]
- Higuchi, H.; Yanagida, T.; Goldman, Y.E. Compliance of thin filaments in skinned fibers of rabbit skeletal muscle. Biophys. J. 1995, 69, 1000–1010. [Google Scholar] [CrossRef]
- Linari, M.; Caremani, M.; Piperio, C.; Brandt, P.; Lombardi, V. Stiffness and fraction of Myosin motors responsible for active force in permeabilized muscle fibers from rabbit psoas. Biophys. J. 2007, 92, 2476–2490. [Google Scholar] [CrossRef] [PubMed]
- Lewalle, A.; Steffen, W.; Stevenson, O.; Ouyang, Z.; Sleep, J. Single-molecule measurement of the stiffness of the rigor myosin head. Biophys. J. 2008, 94, 2160–2169. [Google Scholar] [CrossRef] [PubMed]
- Seebohm, B.; Matinmehr, F.; Kohler, J.; Francino, A.; Navarro-Lopez, F.; Perrot, A.; Ozcelik, C.; McKenna, W.J.; Brenner, B.; Kraft, T. Cardiomyopathy mutations reveal variable region of myosin converter as major element of cross-bridge compliance. Biophys. J. 2009, 97, 806–824. [Google Scholar] [CrossRef] [PubMed]
- Sleep, J.A.; Hutton, R.L. Exchange between inorganic phosphate and adenosine 5′-triphosphate in the medium by actomyosin subfragment 1. Biochemistry 1980, 19, 1276–1283. [Google Scholar] [CrossRef] [PubMed]
- Hopkins, S.C.; Sabido-David, C.; van der Heide, U.A.; Ferguson, R.E.; Brandmeier, B.D.; Dale, R.E.; Kendrick-Jones, J.; Corrie, J.E.; Trentham, D.R.; Irving, M.; et al. Orientation changes of the myosin light chain domain during filament sliding in active and rigor muscle. J. Mol. Biol. 2002, 318, 1275–1291. [Google Scholar] [CrossRef]
- Fajer, P.G.; Fajer, E.A.; Brunsvold, N.J.; Thomas, D.D. Effects of AMPPNP on the orientation and rotational dynamics of spin-labeled muscle cross-bridges. Biophys. J. 1988, 53, 513–524. [Google Scholar] [CrossRef]
- Pate, E.; Cooke, R. Energetics of the actomyosin bond in the filament array of muscle fibers. Biophys. J. 1988, 53, 561–573. [Google Scholar] [CrossRef]
- Cooke, R.; Franks, K. All myosin heads form bonds with actin in rigor rabbit skeletal muscle. Biochemistry 1980, 19, 2265–2269. [Google Scholar] [CrossRef] [PubMed]
- Trinick, J.A.; Offer, G. Cross-linking of actin filaments by heavy meromyosin. J. Mol. Biol. 1979, 133, 549–556. [Google Scholar] [CrossRef]
- Chakrabarty, T.; Yengo, C.; Baldacchino, C.; Chen, L.Q.; Sweeney, H.L.; Selvin, P.R. Does the S2 rod of myosin II uncoil upon two-headed binding to actin? A leucine-zippered HMM study. Biochemistry 2003, 42, 12886–12892. [Google Scholar] [CrossRef] [PubMed]
- Planelles-Herrero, V.J.; Hartman, J.J.; Robert-Paganin, J.; Malik, F.I.; Houdusse, A. Mechanistic and structural basis for activation of cardiac myosin force production by omecamtiv mecarbil. Nat. Commun. 2017, 8, 190. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.A.; Usaj, M.; Rassier, D.E.; Månsson, A. Blebbistatin effects expose hidden secrets in the force-generating cycle of actin and myosin. Biophys. J. 2018, in press. [Google Scholar]
- Smith, D.A. A new mechanokinetic model for muscle contraction, where force and movement are triggered by phosphate release. J. Muscle Res. Cell Motil. 2014, 35, 295–306. [Google Scholar] [CrossRef] [PubMed]
- Månsson, A.; Rassier, D.; Tsiavaliaris, G. Poorly Understood Aspects of Striated Muscle Contraction. Biomed. Res. Int. 2015, 2015, 245154. [Google Scholar] [CrossRef] [PubMed]
- Howard, J. Mechanics of Motor Proteins and the Cytoskeleton; Sinauer Associates Inc.: Sunderland, MA, USA, 2001. [Google Scholar]
- Batters, C.; Veigel, C.; Homsher, E.; Sellers, J.R. To understand muscle you must take it apart. Front. Physiol. 2014, 5, 90. [Google Scholar] [CrossRef] [PubMed]
- Walker, M.L.; Burgess, S.A.; Sellers, J.R.; Wang, F.; Hammer, J.A., 3rd; Trinick, J.; Knight, P.J. Two-headed binding of a processive myosin to F-actin. Nature 2000, 405, 804–807. [Google Scholar] [CrossRef] [PubMed]
- Barclay, C.J. Estimation of cross-bridge stiffness from maximum thermodynamic efficiency. J. Muscle Res. Cell Motil. 1998, 19, 855–864. [Google Scholar] [CrossRef] [PubMed]
- He, Z.H.; Bottinelli, R.; Pellegrino, M.A.; Ferenczi, M.A.; Reggiani, C. ATP consumption and efficiency of human single muscle fibers with different myosin isoform composition. Biophys. J. 2000, 79, 945–961. [Google Scholar] [CrossRef]
- Sun, Y.B.; Hilber, K.; Irving, M. Effect of active shortening on the rate of ATP utilisation by rabbit psoas muscle fibres. J. Physiol. 2001, 531 Pt 3, 781–791. [Google Scholar] [CrossRef] [PubMed]
- Hill, T.L. Theoretical formalism for the sliding filament model of contraction of striated muscle. Part I. Prog. Biophys. Mol. Biol. 1974, 28, 267–340. [Google Scholar] [CrossRef]
- Kaya, M.; Tani, Y.; Washio, T.; Hisada, T.; Higuchi, H. Coordinated force generation of skeletal myosins in myofilaments through motor coupling. Nat. Commun. 2017, 8, 16036. [Google Scholar] [CrossRef] [PubMed]
- Persson, M.; Bengtsson, E.; ten Siethoff, L.; Månsson, A. Nonlinear cross-bridge elasticity and post-power-stroke events in fast skeletal muscle actomyosin. Biophys. J. 2013, 105, 1871–1881. [Google Scholar] [CrossRef] [PubMed]
- van der Heide, U.; Ketelaars, M.; Treijtel, B.W.; de Beer, E.L.; Blange, T. Strain dependence of the elastic properties of force-producing cross- bridges in rigor skeletal muscle. Biophys. J. 1997, 72 Pt 1, 814–821. [Google Scholar] [CrossRef]
- Seow, C.Y.; Shroff, S.G.; Ford, L.E. Detachment of low-force bridges contributes to the rapid tension transients of skinned rabbit skeletal muscle fibres. J. Physiol. 1997, 501 Pt 1, 149–164. [Google Scholar] [CrossRef] [PubMed]
- Piazzesi, G.; Linari, M.; Lombardi, V. The effect of hypertonicity on force generation in tetanized single fibres from frog skeletal muscle. J. Physiol. (Lond.) 1994, 476, 531–546. [Google Scholar] [CrossRef]
- Reconditi, M.; Linari, M.; Lucii, L.; Stewart, A.; Sun, Y.B.; Boesecke, P.; Narayanan, T.; Fischetti, R.F.; Irving, T.; Piazzesi, G.; et al. The myosin motor in muscle generates a smaller and slower working stroke at higher load. Nature 2004, 428, 578–581. [Google Scholar] [CrossRef] [PubMed]
- Nyitrai, M.; Geeves, M.A. Adenosine diphosphate and strain sensitivity in myosin motors. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2004, 359, 1867–1877. [Google Scholar] [PubMed]
- Piazzesi, G.; Dolfi, M.; Brunello, E.; Fusi, L.; Reconditi, M.; Bianco, P.; Linari, M.; Lombardi, V. The myofilament elasticity and its effect on kinetics of force generation by the myosin motor. Arch. Biochem. Biophys. 2014, 552–553, 108–116. [Google Scholar] [CrossRef] [PubMed]
- Offer, G.; Ranatunga, K.W. The endothermic ATP hydrolysis and crossbridge attachment steps drive the increase of force with temperature in isometric and shortening muscle. J. Physiol. 2015, 593, 1997–2016. [Google Scholar] [CrossRef] [PubMed]
- Reubold, T.F.; Eschenburg, S.; Becker, A.; Kull, F.J.; Manstein, D.J. A structural model for actin-induced nucleotide release in myosin. Nat. Struct. Biol. 2003, 10, 826–830. [Google Scholar] [CrossRef] [PubMed]
- Uyeda, T.Q.; Abramson, P.D.; Spudich, J.A. The neck region of the myosin motor domain acts as a lever arm to generate movement. Proc. Natl. Acad. Sci. USA 1996, 93, 4459–4464. [Google Scholar] [CrossRef] [PubMed]
- Houdusse, A.; Szent-Gyorgyi, A.G.; Cohen, C. Three conformational states of scallop myosin S1. Proc. Natl. Acad. Sci. USA 2000, 97, 11238–11243. [Google Scholar] [CrossRef] [PubMed]
- Billington, N.; Revill, D.J.; Burgess, S.A.; Chantler, P.D.; Knight, P.J. Flexibility within the heads of muscle myosin-2 molecules. J. Mol. Biol. 2014, 426, 894–907. [Google Scholar] [CrossRef] [PubMed]
- Uyeda, T.Q.; Kron, S.J.; Spudich, J.A. Myosin step size. Estimation from slow sliding movement of actin over low densities of heavy meromyosin. J. Mol. Biol. 1990, 214, 699–710. [Google Scholar] [CrossRef]
- Yanagida, T.; Ishijima, A.; Saito, K.; Harada, Y. Coupling between ATPase and force-generating attachment-detachment cycles of actomyosin in vitro. Adv. Exp. Med. Biol. 1993, 332, 339–347; discussion 347–349. [Google Scholar] [PubMed]
- Ujfalusi, Z.; Vera, C.D.; Mijailovich, S.M.; Svicevic, M.; Yu, E.C.; Kawana, M.; Ruppel, K.; Spudich, J.A.; Geeves, M.A.; Leinwand, L.A. Dilated cardiomyopathy myosin mutants have reduced force-generating capacity. J. Biol. Chem. 2018. [Google Scholar] [CrossRef] [PubMed]
- Walcott, S.; Warshaw, D.M.; Debold, E.P. Mechanical coupling between myosin molecules causes differences between ensemble and single-molecule measurements. Biophys. J. 2012, 103, 501–510. [Google Scholar] [CrossRef] [PubMed]
- Baker, J.E.; Brosseau, C.; Joel, P.B.; Warshaw, D.M. The biochemical kinetics underlying actin movement generated by one and many skeletal muscle Myosin molecules. Biophys. J. 2002, 82, 2134–2147. [Google Scholar] [CrossRef]
- Brizendine, R.K.; Alcala, D.B.; Carter, M.S.; Haldeman, B.D.; Facemyer, K.C.; Baker, J.E.; Cremo, C.R. Velocities of unloaded muscle filaments are not limited by drag forces imposed by myosin cross-bridges. Proc. Natl. Acad. Sci. USA 2015, 112, 11235–11240. [Google Scholar] [CrossRef] [PubMed]
- Mijailovich, S.M.; Nedic, D.; Svicevic, M.; Stojanovic, B.; Walklate, J.; Ujfalusi, Z.; Geeves, M.A. Modeling the Actin.myosin ATPase Cross-Bridge Cycle for Skeletal and Cardiac Muscle Myosin Isoforms. Biophys. J. 2017, 112, 984–996. [Google Scholar] [CrossRef] [PubMed]
- Piazzesi, G.; Lombardi, V. A cross-bridge model that is able to explain mechanical and energetic properties of shortening muscle. Biophys. J. 1995, 68, 1966–1979. [Google Scholar] [CrossRef]
- Wang, Y.; Ajtai, K.; Burghardt, T.P. The Qdot-labeled actin super-resolution motility assay measures low-duty cycle muscle myosin step size. Biochemistry 2013, 52, 1611–1621. [Google Scholar] [CrossRef] [PubMed]
- Pate, E.; Cooke, R. A model of crossbridge action: The effects of ATP, ADP and Pi. J. Muscle Res. Cell Motil. 1989, 10, 181–196. [Google Scholar] [CrossRef] [PubMed]
- Adamovic, I.; Mijailovich, S.M.; Karplus, M. The elastic properties of the structurally characterized myosin II S2 subdomain: A molecular dynamics and normal mode analysis. Biophys. J. 2008, 94, 3779–3789. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, H.; Goldman, Y.E. Sliding distance per ATP molecule hydrolyzed by myosin heads during isotonic shortening of skinned muscle fibers. Biophys. J. 1995, 69, 1491–1507. [Google Scholar] [CrossRef]
- Fusi, L.; Reconditi, M.; Linari, M.; Brunello, E.; Elangovan, R.; Lombardi, V.; Piazzesi, G. The mechanism of the resistance to stretch of isometrically contracting single muscle fibres. J. Physiol. 2010, 588 Pt 3, 495–510. [Google Scholar] [CrossRef] [PubMed]
- Fusi, L.; Brunello, E.; Reconditi, M.; Piazzesi, G.; Lombardi, V. The non-linear elasticity of the muscle sarcomere and the compliance of myosin motors. J. Physiol. 2014, 592 Pt 5, 1109–1118. [Google Scholar] [CrossRef] [PubMed]
- Månsson, A. Significant impact on muscle mechanics of small nonlinearities in myofilament elasticity. Biophys. J. 2010, 99, 1869–1875. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, K.; Tokunaga, M.; Iwane, A.H.; Yanagida, T. A single myosin head moves along an actin filament with regular steps of 5.3 nanometres. Nature 1999, 397, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Longyear, T.; Walcott, S.; Debold, E.P. The molecular basis of thin filament activation: From single molecule to muscle. Sci. Rep. 2017, 7, 1822. [Google Scholar] [CrossRef] [PubMed]
- Brenner, B.; Yu, L.C.; Greene, L.E.; Eisenberg, E.; Schoenberg, M. Ca2+-sensitive cross-bridge dissociation in the presence of magnesium pyrophosphate in skinned rabbit psoas fibers. Biophys. J. 1986, 50, 1101–1108. [Google Scholar] [CrossRef]
- Månsson, A. The tension response to stretch of intact skeletal muscle fibres of the frog at varied tonicity of the extracellular medium. J. Muscle Res. Cell Motil. 1994, 15, 145–157. [Google Scholar] [CrossRef] [PubMed]
- Cornachione, A.S.; Rassier, D.E. A non-cross-bridge, static tension is present in permeabilized skeletal muscle fibers after active force inhibition or actin extraction. Am. J. Physiol. Cell Physiol. 2012, 302, C566–C574. [Google Scholar] [CrossRef] [PubMed]
- Campbell, K.S.; Moss, R.L. History-dependent mechanical properties of permeabilized rat soleus muscle fibers. Biophys.J. 2002, 82, 929–943. [Google Scholar] [CrossRef]
- Labeit, D.; Watanabe, K.; Witt, C.; Fujita, H.; Wu, Y.; Lahmers, S.; Funck, T.; Labeit, S.; Granzier, H. Calcium-dependent molecular spring elements in the giant protein titin. Proc. Natl. Acad. Sci. USA 2003, 100, 13716–13721. [Google Scholar] [CrossRef] [PubMed]
- Edman, K.A.P.; Månsson, A.; Caputo, C. The biphasic force-velocity relationship in frog muscle fibres and its evaluation in terms of cross-bridge function. J. Physiol. (Lond.) 1997, 503 Pt 1, 141–156. [Google Scholar] [CrossRef]
- Marcucci, L.; Reggiani, C. Mechanosensing in Myosin Filament Solves a 60 Years Old Conflict in Skeletal Muscle Modeling between High Power Output and Slow Rise in Tension. Front. Physiol. 2016, 7, 427. [Google Scholar] [CrossRef] [PubMed]
- Huxley, A.F. Muscular contraction. J. Physiol. (Lond.) 1974, 243, 1–43. [Google Scholar] [CrossRef]
- Kad, N.M.; Rovner, A.S.; Fagnant, P.M.; Joel, P.B.; Kennedy, G.G.; Patlak, J.B.; Warshaw, D.M.; Trybus, K.M. A mutant heterodimeric myosin with one inactive head generates maximal displacement. J. Cell Biol. 2003, 162, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Knupp, C.; Offer, G.; Ranatunga, K.W.; Squire, J.M. Probing muscle myosin motor action: X-ray (m3 and m6) interference measurements report motor domain not lever arm movement. J. Mol. Biol. 2009, 390, 168–181. [Google Scholar] [CrossRef] [PubMed]
- Bagni, M.A.; Cecchi, G.; Griffiths, P.J.; Maeda, Y.; Rapp, G.; Ashley, C.C. Lattice spacing changes accompanying isometric tension development in intact single muscle fibers. Biophys. J. 1994, 67, 1965–1975. [Google Scholar] [CrossRef]
- Yu, L.C.; Brenner, B. Structures of actomyosin crossbridges in relaxed and rigor muscle fibers. Biophys. J. 1989, 55, 441–453. [Google Scholar] [CrossRef]
- Edman, K.A.; Reggiani, C. Redistribution of sarcomere length during isometric contraction of frog muscle fibres and its relation to tension creep. J. Physiol. (Lond.) 1984, 351, 169–198. [Google Scholar] [CrossRef]
- Edman, K.A.; Reggiani, C.; te Kronnie, G. Differences in maximum velocity of shortening along single muscle fibres of the frog. J. Physiol. (Lond.) 1985, 365, 147–163. [Google Scholar] [CrossRef]
- Shalabi, N.; Persson, M.; Månsson, A.; Vengallatore, S.; Rassier, D.E. Sarcomere Stiffness during Stretching and Shortening of Rigor Skeletal Myofibrils. Biophys. J. 2017, 113, 2768–2776. [Google Scholar] [CrossRef] [PubMed]
- Tajsharghi, H.; Oldfors, A. Myosinopathies: Pathology and mechanisms. Acta Neuropathol. 2013, 125, 3–18. [Google Scholar] [CrossRef] [PubMed]
- Jungbluth, H.; Treves, S.; Zorzato, F.; Sarkozy, A.; Ochala, J.; Sewry, C.; Phadke, R.; Gautel, M.; Muntoni, F. Congenital myopathies: Disorders of excitation-contraction coupling and muscle contraction. Nat. Rev. Neurol. 2018, 14, 151–167. [Google Scholar] [CrossRef] [PubMed]
- Jungbluth, H.; Ochala, J.; Treves, S.; Gautel, M. Current and future therapeutic approaches to the congenital myopathies. Semin. Cell Dev. Biol. 2017, 64, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Ochala, J.; Sun, Y.B. Novel myosin-based therapies for congenital cardiac and skeletal myopathies. J. Med. Genet. 2016, 53, 651–654. [Google Scholar] [CrossRef] [PubMed]
- Lindqvist, J.; Levy, Y.; Pati-Alam, A.; Hardeman, E.C.; Gregorevic, P.; Ochala, J. Modulating myosin restores muscle function in a mouse model of nemaline myopathy. Ann. Neurol. 2016, 79, 717–725. [Google Scholar] [CrossRef] [PubMed]

© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Månsson, A.; Ušaj, M.; Moretto, L.; Rassier, D.E. Do Actomyosin Single-Molecule Mechanics Data Predict Mechanics of Contracting Muscle? Int. J. Mol. Sci. 2018, 19, 1863. https://doi.org/10.3390/ijms19071863
Månsson A, Ušaj M, Moretto L, Rassier DE. Do Actomyosin Single-Molecule Mechanics Data Predict Mechanics of Contracting Muscle? International Journal of Molecular Sciences. 2018; 19(7):1863. https://doi.org/10.3390/ijms19071863
Chicago/Turabian StyleMånsson, Alf, Marko Ušaj, Luisa Moretto, and Dilson E. Rassier. 2018. "Do Actomyosin Single-Molecule Mechanics Data Predict Mechanics of Contracting Muscle?" International Journal of Molecular Sciences 19, no. 7: 1863. https://doi.org/10.3390/ijms19071863
APA StyleMånsson, A., Ušaj, M., Moretto, L., & Rassier, D. E. (2018). Do Actomyosin Single-Molecule Mechanics Data Predict Mechanics of Contracting Muscle? International Journal of Molecular Sciences, 19(7), 1863. https://doi.org/10.3390/ijms19071863