Plasma Membrane-Localized Calcium Pumps and Copines Coordinately Regulate Pollen Germination and Fertility in Arabidopsis
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. Plant Materials
3.2. Plant Growth Conditions
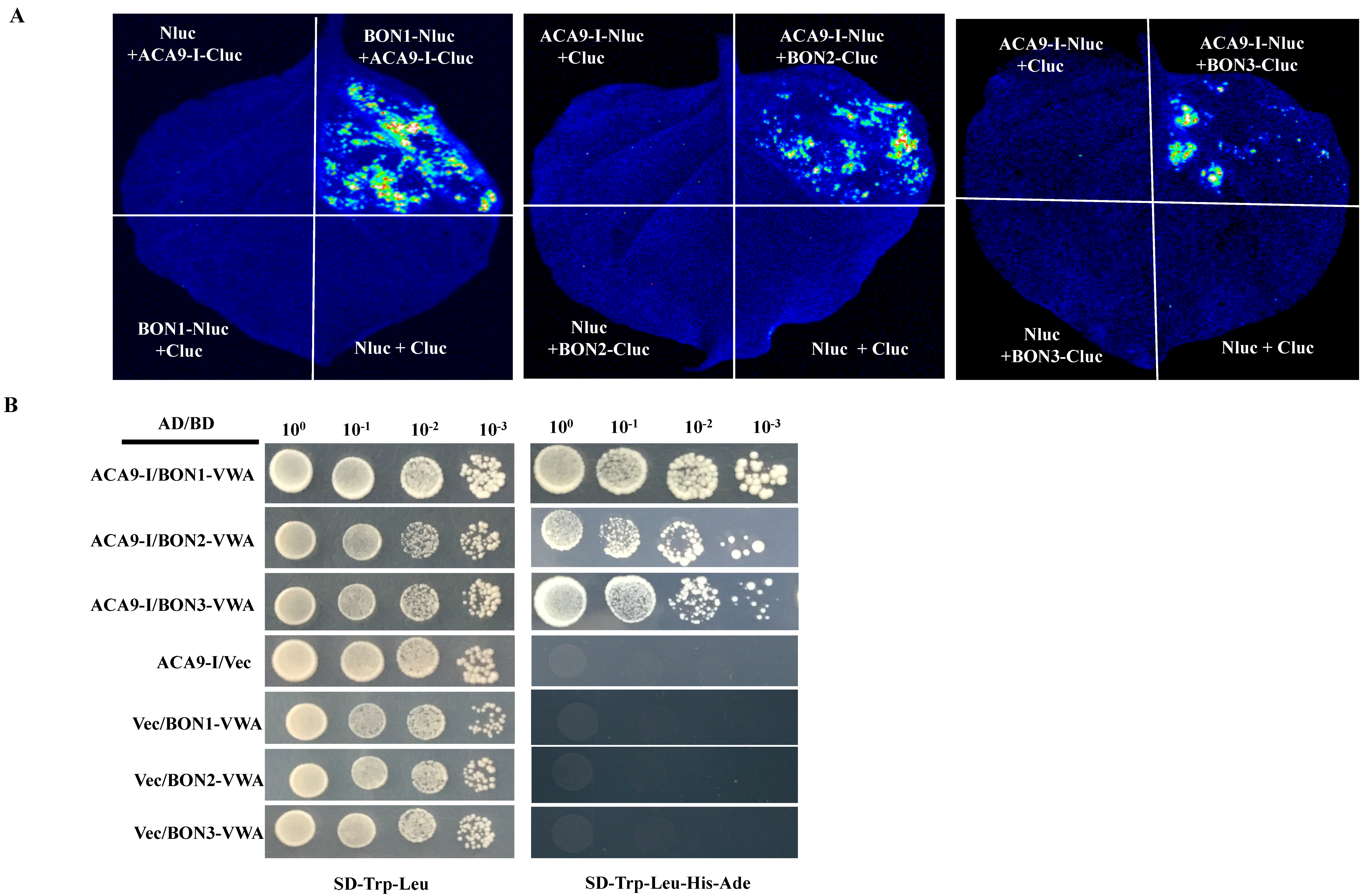
3.3. Split-Luc Assay
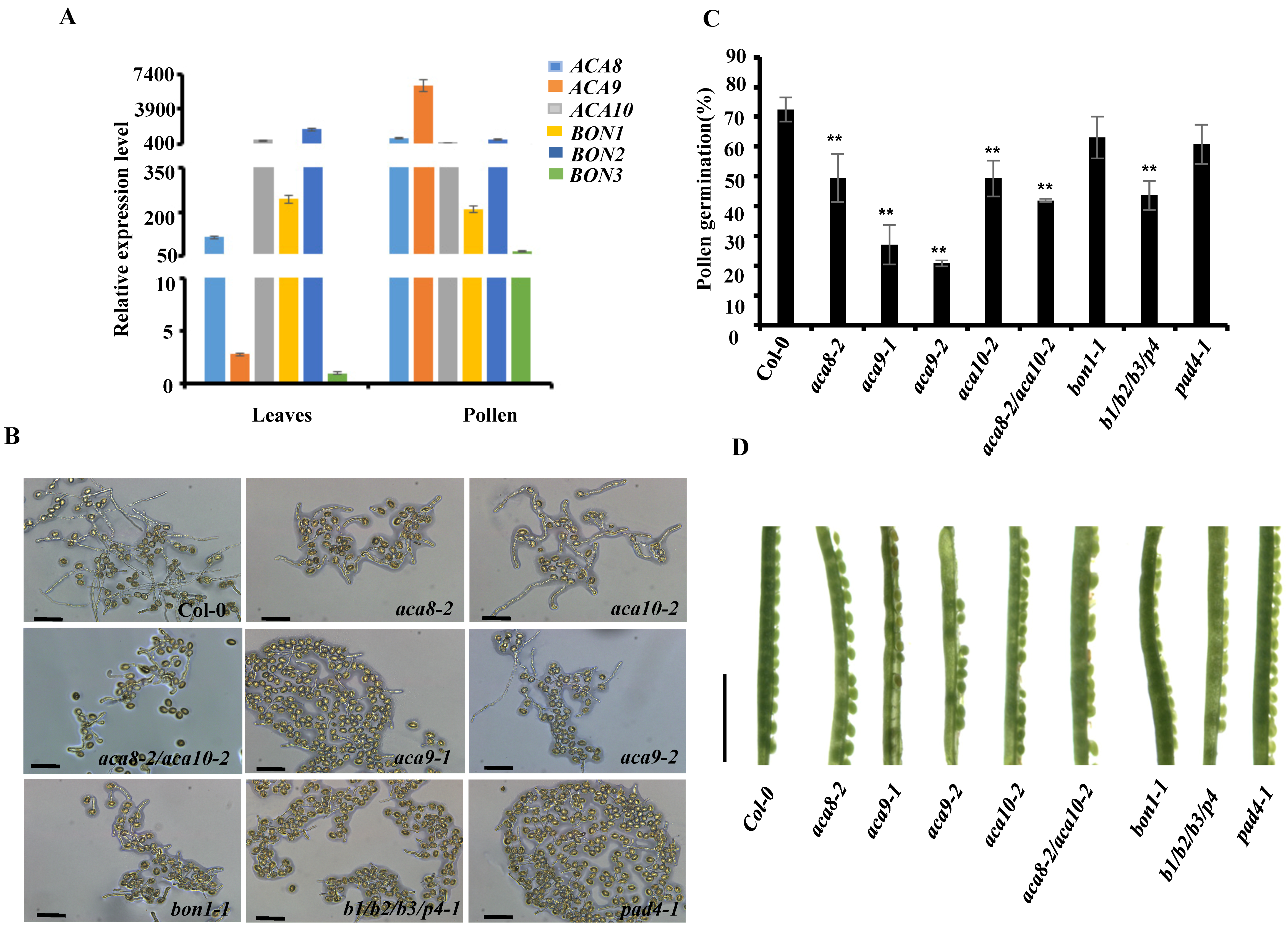
3.4. Pollen Germination Assay
3.5. Yeast-Two-Hybrid Assay
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- White, P.J.; Broadley, M.R. Calcium in plants. Ann. Bot. 2003, 92, 487–511. [Google Scholar] [CrossRef] [PubMed]
- Dodd, A.N.; Kudla, J.; Sanders, D. The language of calcium signaling. Annu. Rev. Plant Biol. 2010, 61, 593–620. [Google Scholar] [CrossRef] [PubMed]
- Kudla, J.; Batistic, O.; Hashimoto, K. Calcium signals: the lead currency of plant information processing. Plant Cell 2010, 22, 541–563. [Google Scholar] [CrossRef] [PubMed]
- Bonza, M.C.; De Michelis, M.I. The plant Ca2+—ATPase repertoire: biochemical features and physiological functions. Plant Biol. (Stuttg. Ger.) 2011, 13, 421–430. [Google Scholar] [CrossRef] [PubMed]
- Steinhorst, L.; Kudla, J. Calcium—A central regulator of pollen germination and tube growth. Biochim. Biophys. Acta 2013, 1833, 1573–1581. [Google Scholar] [CrossRef] [PubMed]
- Geisler, M.; Axelsen, K.B.; Harper, J.F.; Palmgren, M.G. Molecular aspects of higher plant P-type Ca(2+)-ATPases. Biochim. Biophys. Acta 2000, 1465, 52–78. [Google Scholar] [CrossRef]
- Sze, H.; Liang, F.; Hwang, I.; Curran, A.C.; Harper, J.F. Diversity and regulation of plant Ca2+ pumps: Insights from expression in yeast. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2000, 51, 433–462. [Google Scholar] [CrossRef] [PubMed]
- Boursiac, Y.; Harper, J.F. The origin and function of calmodulin regulated Ca2+ pumps in plants. J. Bioenerg. Biomembr. 2007, 39, 409–414. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Yan, J.; Du, X.; Hua, J. Overlapping and differential roles of plasma membrane calcium ATPase ACAs in Arabidopsis growth and environmental responses. J. Exp. Bot. 2018, 69, 2693–2703. [Google Scholar] [CrossRef] [PubMed]
- Frei dit Frey, N.; Mbengue, M.; Kwaaitaal, M.; Nitsch, L.; Altenbach, D.; Haweker, H.; Lozano-Duran, R.; Njo, M.F.; Beeckman, T.; Huettel, B.; et al. Plasma membrane calcium ATPases are important components of receptor-mediated signaling in plant immune responses and development. Plant Physiol. 2012, 159, 798–809. [Google Scholar] [CrossRef] [PubMed]
- Costa, A.; Luoni, L.; Marrano, C.A.; Hashimoto, K.; Koster, P.; Giacometti, S.; De Michelis, M.I.; Kudla, J.; Bonza, M.C. Ca2+-dependent phosphoregulation of the plasma membrane Ca2+-ATPase ACA8 modulates stimulus-induced calcium signatures. J. Exp. Bot. 2017, 68, 3215–3230. [Google Scholar] [CrossRef] [PubMed]
- Giacometti, S.; Marrano, C.A.; Bonza, M.C.; Luoni, L.; Limonta, M.; De Michelis, M.I. Phosphorylation of serine residues in the N-terminus modulates the activity of ACA8, a plasma membrane Ca2+-ATPase of Arabidopsis thaliana. J. Exp. Bot. 2012, 63, 1215–1224. [Google Scholar] [CrossRef] [PubMed]
- Tidow, H.; Poulsen, L.R.; Andreeva, A.; Knudsen, M.; Hein, K.L.; Wiuf, C.; Palmgren, M.G.; Nissen, P. A bimodular mechanism of calcium control in eukaryotes. Nature 2012, 491, 468–472. [Google Scholar] [CrossRef] [PubMed]
- Bonza, M.C.; Morandini, P.; Luoni, L.; Geisler, M.; Palmgren, M.G.; De Michelis, M.I. At-ACA8 encodes a plasma membrane-localized calcium-ATPase of Arabidopsis with a calmodulin-binding domain at the N terminus. Plant Physiol. 2000, 123, 1495–1506. [Google Scholar] [CrossRef] [PubMed]
- Schiott, M.; Romanowsky, S.M.; Baekgaard, L.; Jakobsen, M.K.; Palmgren, M.G.; Harper, J.F. A plant plasma membrane Ca2+ pump is required for normal pollen tube growth and fertilization. Proc. Natl. Acad. Sci. USA 2004, 101, 9502–9507. [Google Scholar] [CrossRef] [PubMed]
- George, L.; Romanowsky, S.M.; Harper, J.F.; Sharrock, R.A. The ACA10 Ca2+-ATPase regulates adult vegetative development and inflorescence architecture in Arabidopsis. Plant Physiol. 2008, 146, 716–728. [Google Scholar] [CrossRef] [PubMed]
- Lucca, N.; Leon, G. Arabidopsis ACA7, encoding a putative auto-regulated Ca(2+)-ATPase, is required for normal pollen development. Plant Cell Rep. 2012, 31, 651–659. [Google Scholar] [CrossRef] [PubMed]
- Iwano, M.; Igarashi, M.; Tarutani, Y.; Kaothien-Nakayama, P.; Nakayama, H.; Moriyama, H.; Yakabe, R.; Entani, T.; Shimosato-Asano, H.; Ueki, M.; et al. A pollen coat-inducible autoinhibited Ca2+-ATPase expressed in stigmatic papilla cells is required for compatible pollination in the Brassicaceae. Plant Cell 2014, 26, 636–649. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.L.; Shi, Z.; Bao, Y.; Yan, J.; Yang, Z.; Yu, H.; Li, Y.; Gou, M.; Wang, S.; Zou, B.; et al. Calcium Pumps and Interacting BON1 Protein Modulate Calcium Signature, Stomatal Closure, and Plant Immunity. Plant Physiol. 2017, 175, 424–437. [Google Scholar] [CrossRef] [PubMed]
- Boursiac, Y.; Lee, S.M.; Romanowsky, S.; Blank, R.; Sladek, C.; Chung, W.S.; Harper, J.F. Disruption of the vacuolar calcium-ATPases in Arabidopsis results in the activation of a salicylic acid-dependent programmed cell death pathway. Plant Physiol. 2010, 154, 1158–1171. [Google Scholar] [CrossRef] [PubMed]
- Creutz, C.E.; Tomsig, J.L.; Snyder, S.L.; Gautier, M.C.; Skouri, F.; Beisson, J.; Cohen, J. The copines, a novel class of C2 domain-containing, calcium-dependent, phospholipid-binding proteins conserved from Paramecium to humans. J. Biol. Chem. 1998, 273, 1393–1402. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Yang, H.; Grisafi, P.; Sanchatjate, S.; Fink, G.R.; Sun, Q.; Hua, J. The BON/CPN gene family represses cell death and promotes cell growth in Arabidopsis. Plant J. 2006, 45, 166–179. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Hua, J. A Haplotype-Specific Resistance Gene Regulated by BONZAI1 Mediates Temperature-Dependent Growth Control in Arabidopsis. Plant Cell 2004, 16, 1060–1071. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Zou, Y.; Shang, Y.; Lin, H.; Wang, Y.; Cai, R.; Tang, X.; Zhou, J.-M. Firefly Luciferase Complementation Imaging Assay for Pro tein-Protein Interactions in Plants. Plant Physiol. 2008, 146, 368–376. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Lin, Y.; Heath, R.M.; Zhu, M.X.; Yang, Z. Control of pollen tube tip growth by a Rop GTPase-dependent pathway that leads to tip-localized calcium influx. Plant Cell 1999, 11, 1731–1742. [Google Scholar] [PubMed]
- Boavida, L.C.; McCormick, S. Temperature as a determinant factor for increased and reproducible in vitro pollen germination in Arabidopsis thaliana. Plant J. 2007, 52, 570–582. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Guo, J.; Yang, Z.; Yang, D.-L. Plasma Membrane-Localized Calcium Pumps and Copines Coordinately Regulate Pollen Germination and Fertility in Arabidopsis. Int. J. Mol. Sci. 2018, 19, 1774. https://doi.org/10.3390/ijms19061774
Li Y, Guo J, Yang Z, Yang D-L. Plasma Membrane-Localized Calcium Pumps and Copines Coordinately Regulate Pollen Germination and Fertility in Arabidopsis. International Journal of Molecular Sciences. 2018; 19(6):1774. https://doi.org/10.3390/ijms19061774
Chicago/Turabian StyleLi, Yun, Jinping Guo, Ziyuan Yang, and Dong-Lei Yang. 2018. "Plasma Membrane-Localized Calcium Pumps and Copines Coordinately Regulate Pollen Germination and Fertility in Arabidopsis" International Journal of Molecular Sciences 19, no. 6: 1774. https://doi.org/10.3390/ijms19061774
APA StyleLi, Y., Guo, J., Yang, Z., & Yang, D.-L. (2018). Plasma Membrane-Localized Calcium Pumps and Copines Coordinately Regulate Pollen Germination and Fertility in Arabidopsis. International Journal of Molecular Sciences, 19(6), 1774. https://doi.org/10.3390/ijms19061774



