Genome-Wide Identification of MicroRNAs in Response to Cadmium Stress in Oilseed Rape (Brassica napus L.) Using High-Throughput Sequencing
Abstract
1. Introduction
2. Results
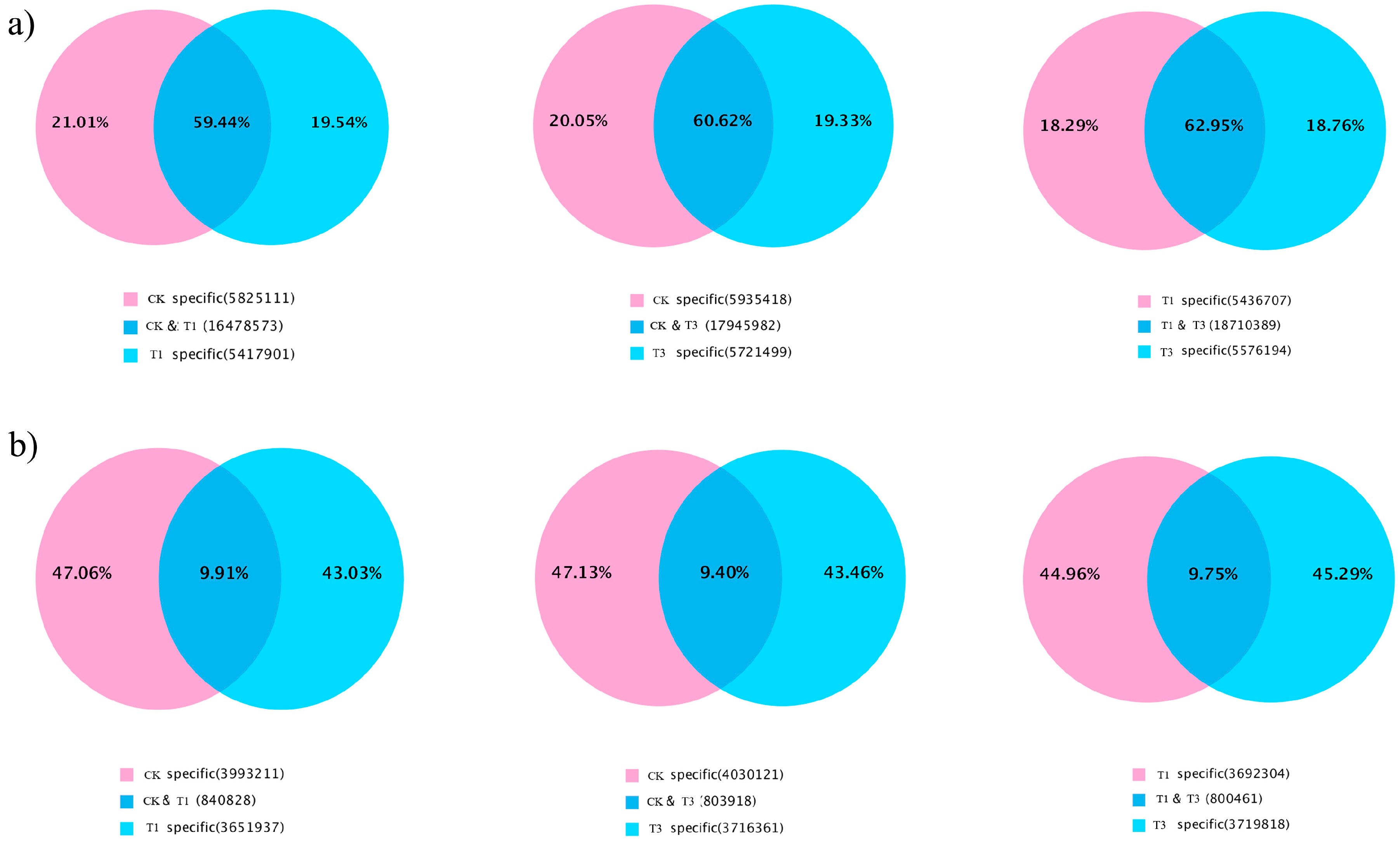
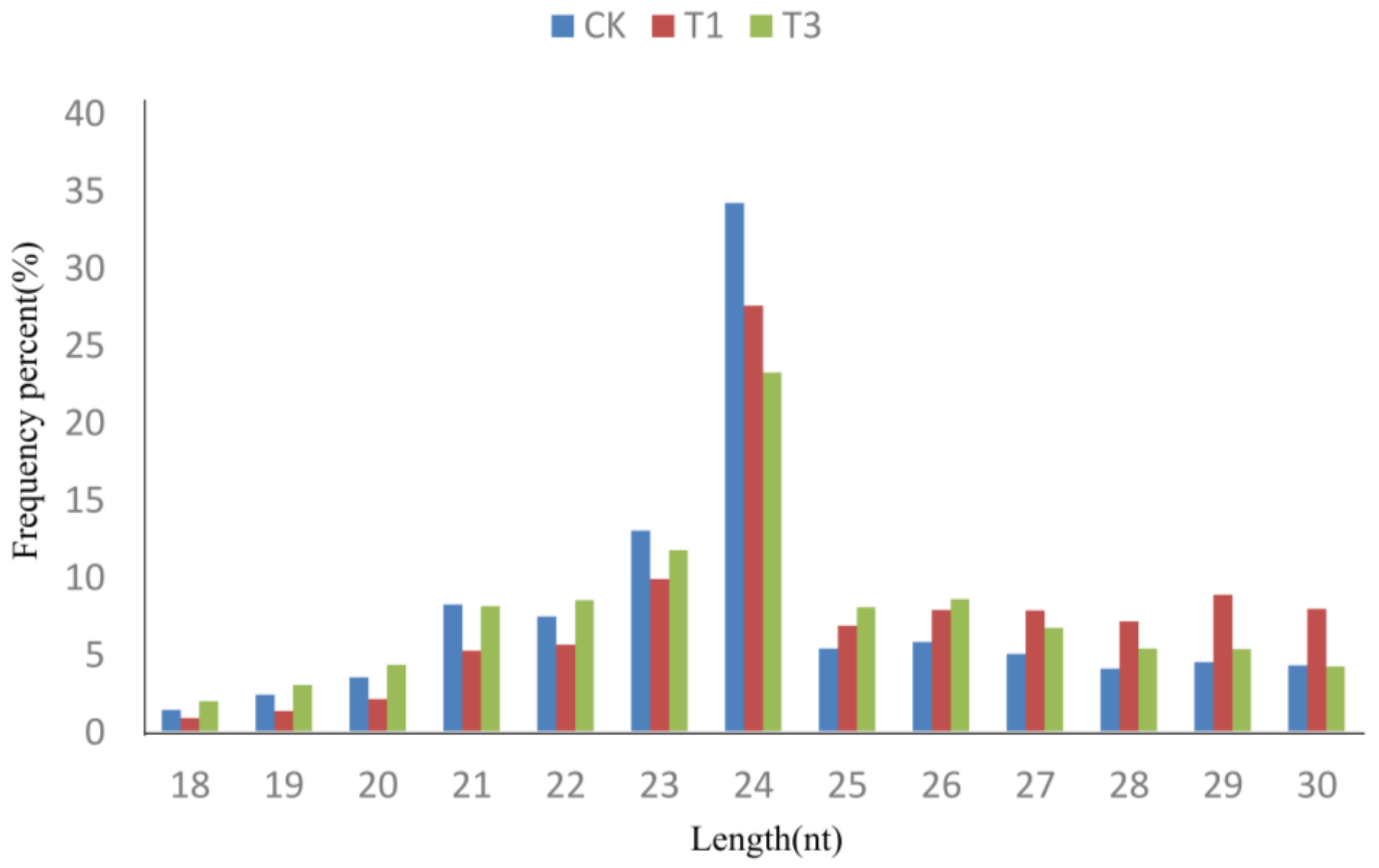
2.1. High-Throughput Sequencing of Small RNAs
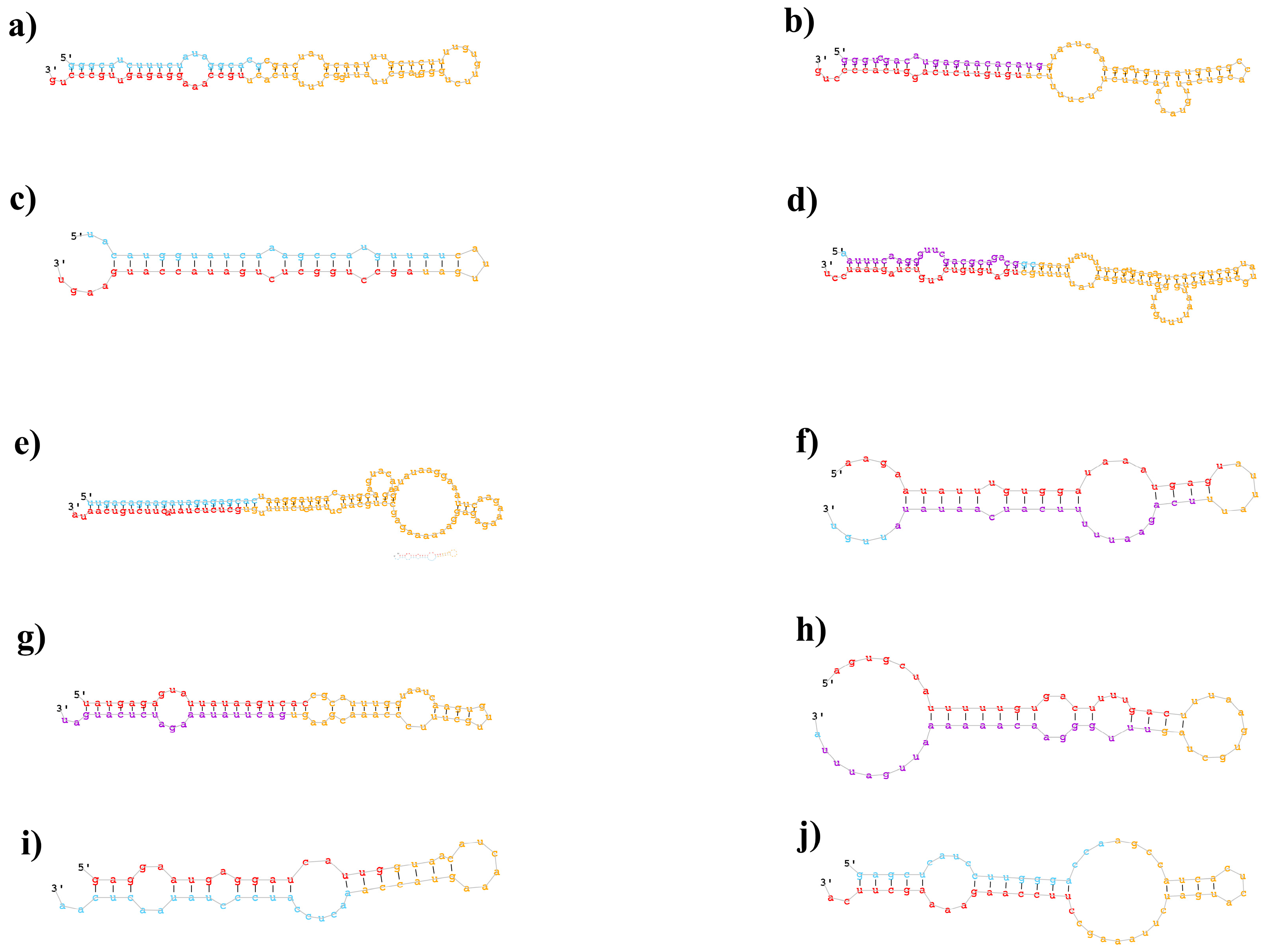
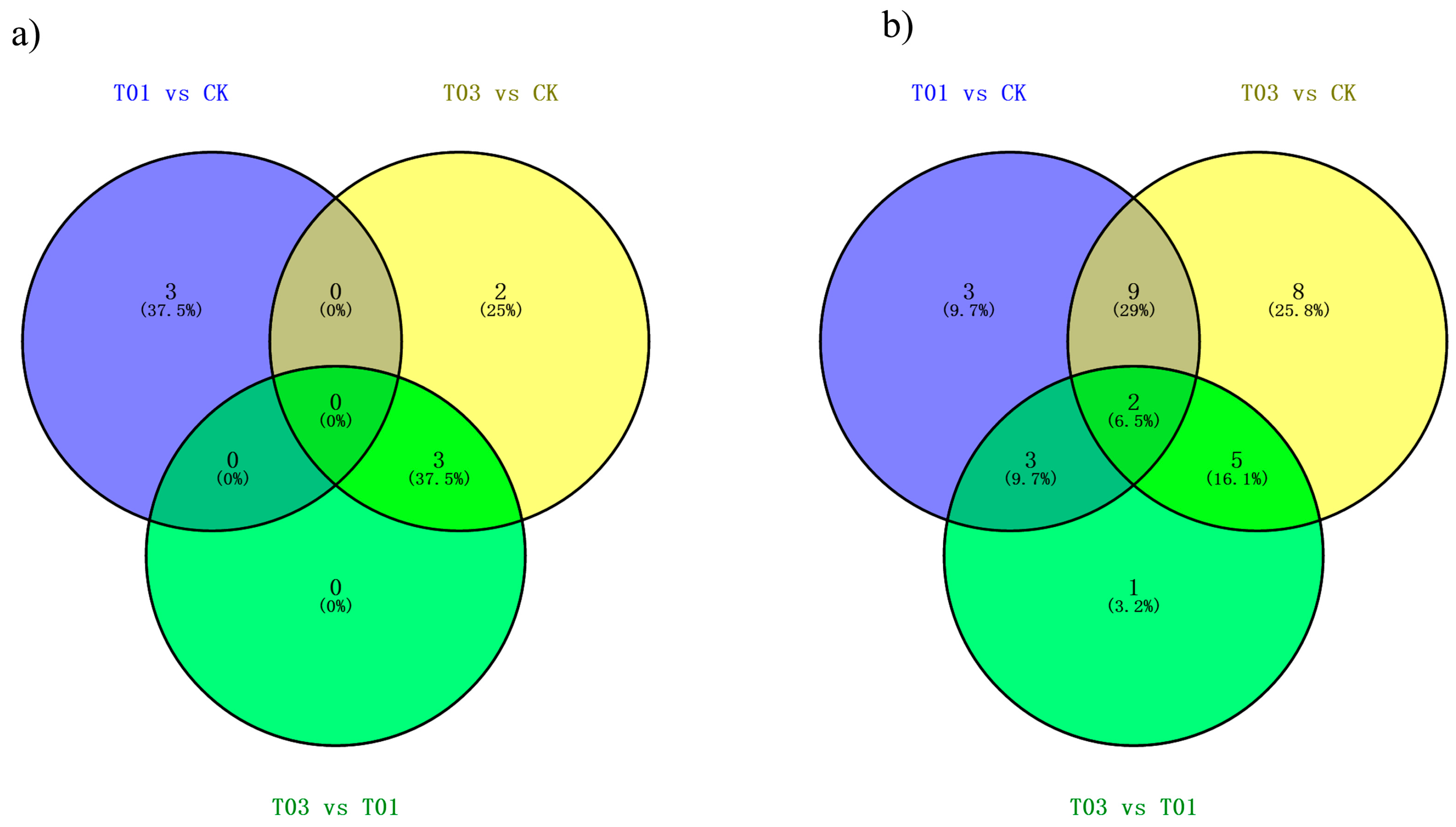
2.2. Identification of Known and Novel miRNAs in B. napus
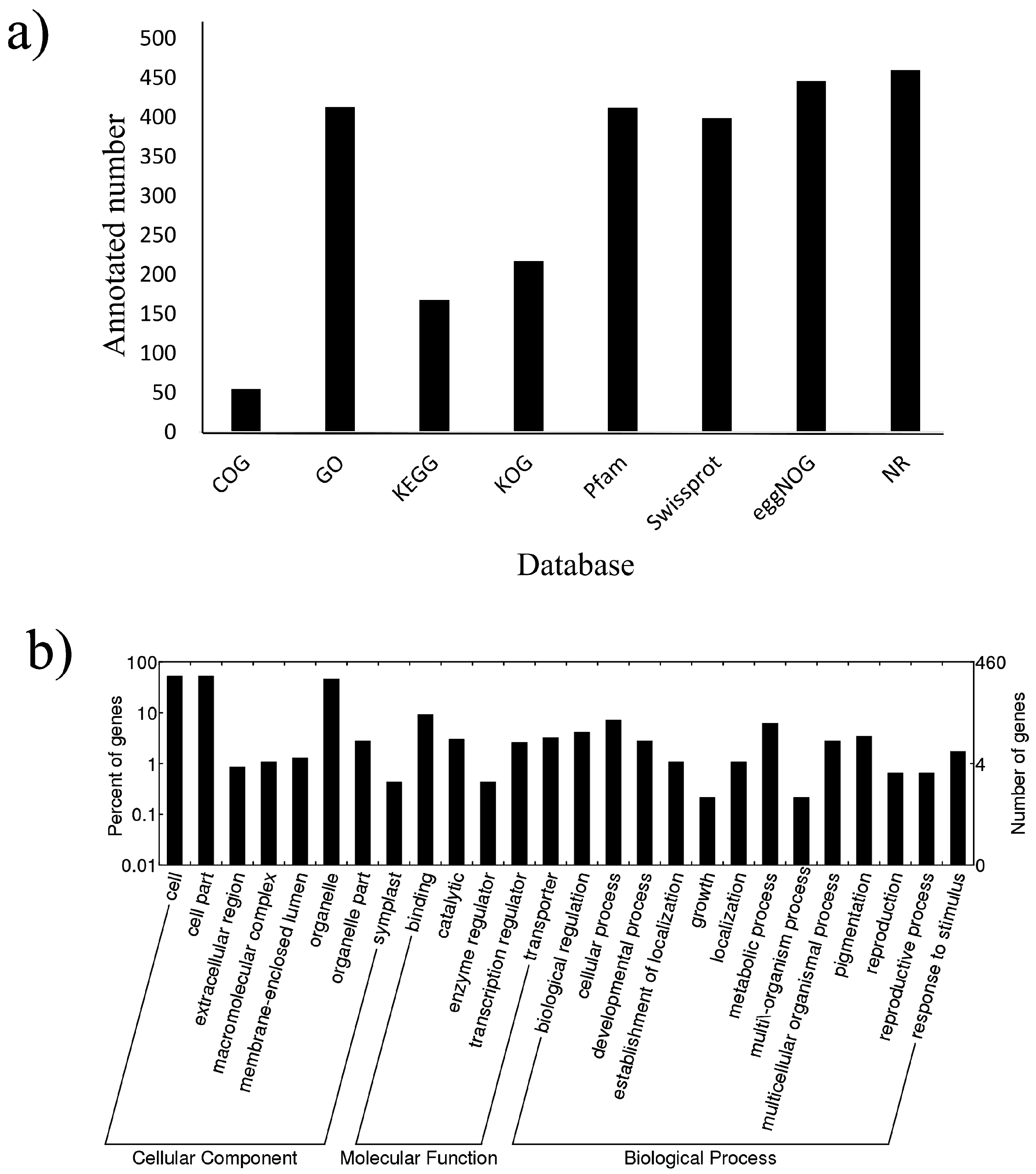
2.3. Target Prediction and Functional Analysis of miRNAs
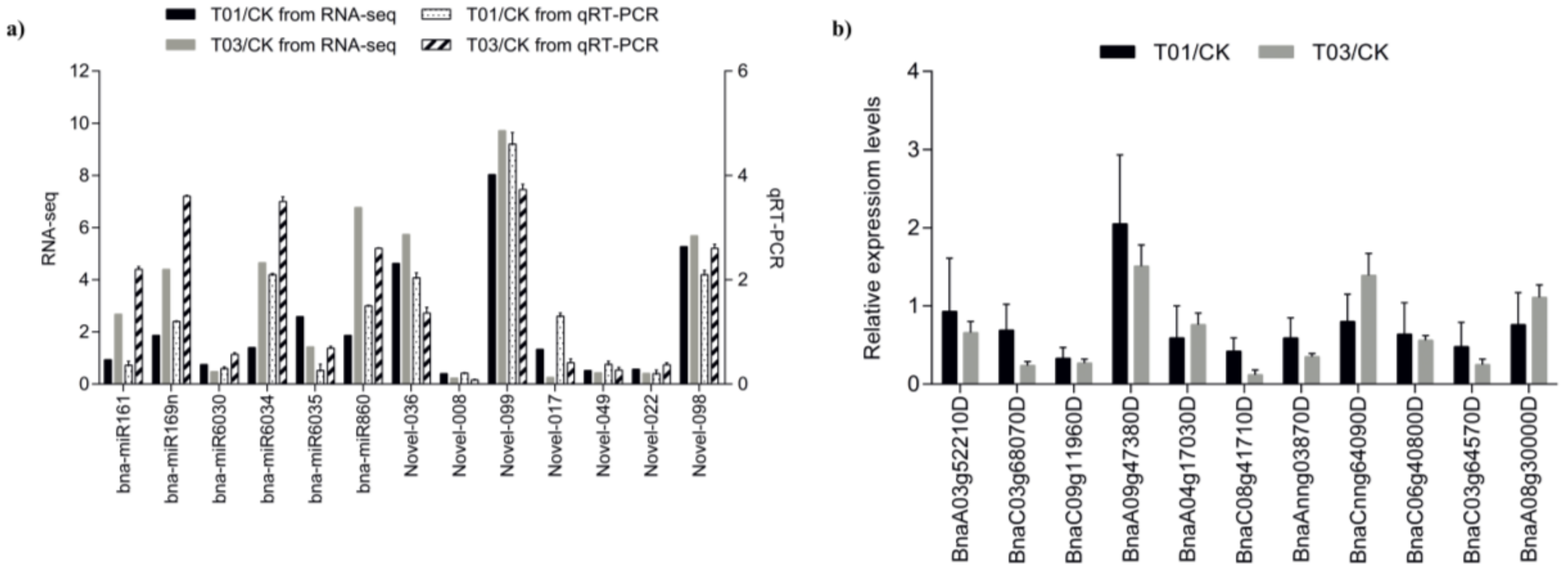
2.4. qRT-PCR Validation
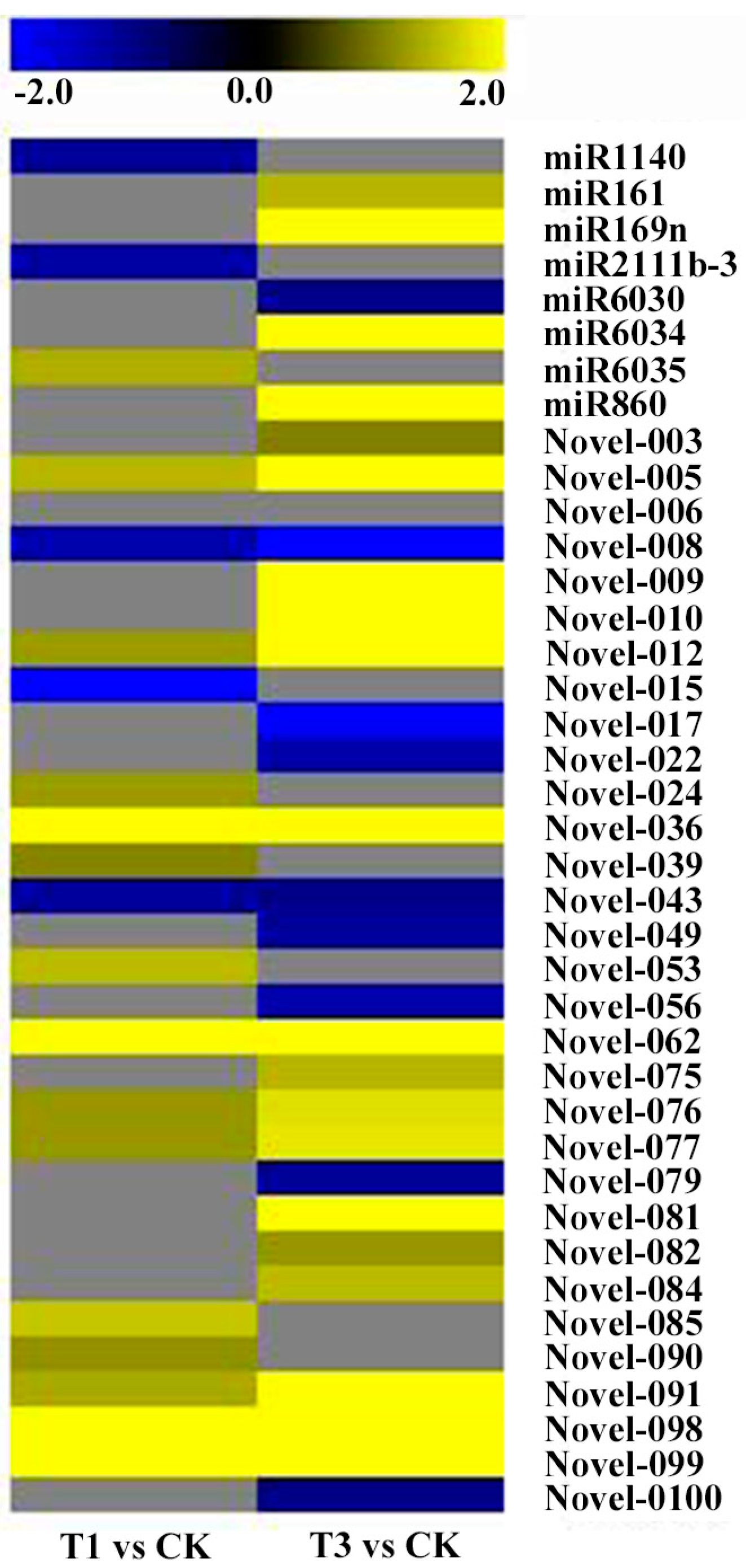
2.5. miRNA Expression in Response to Cd Stress
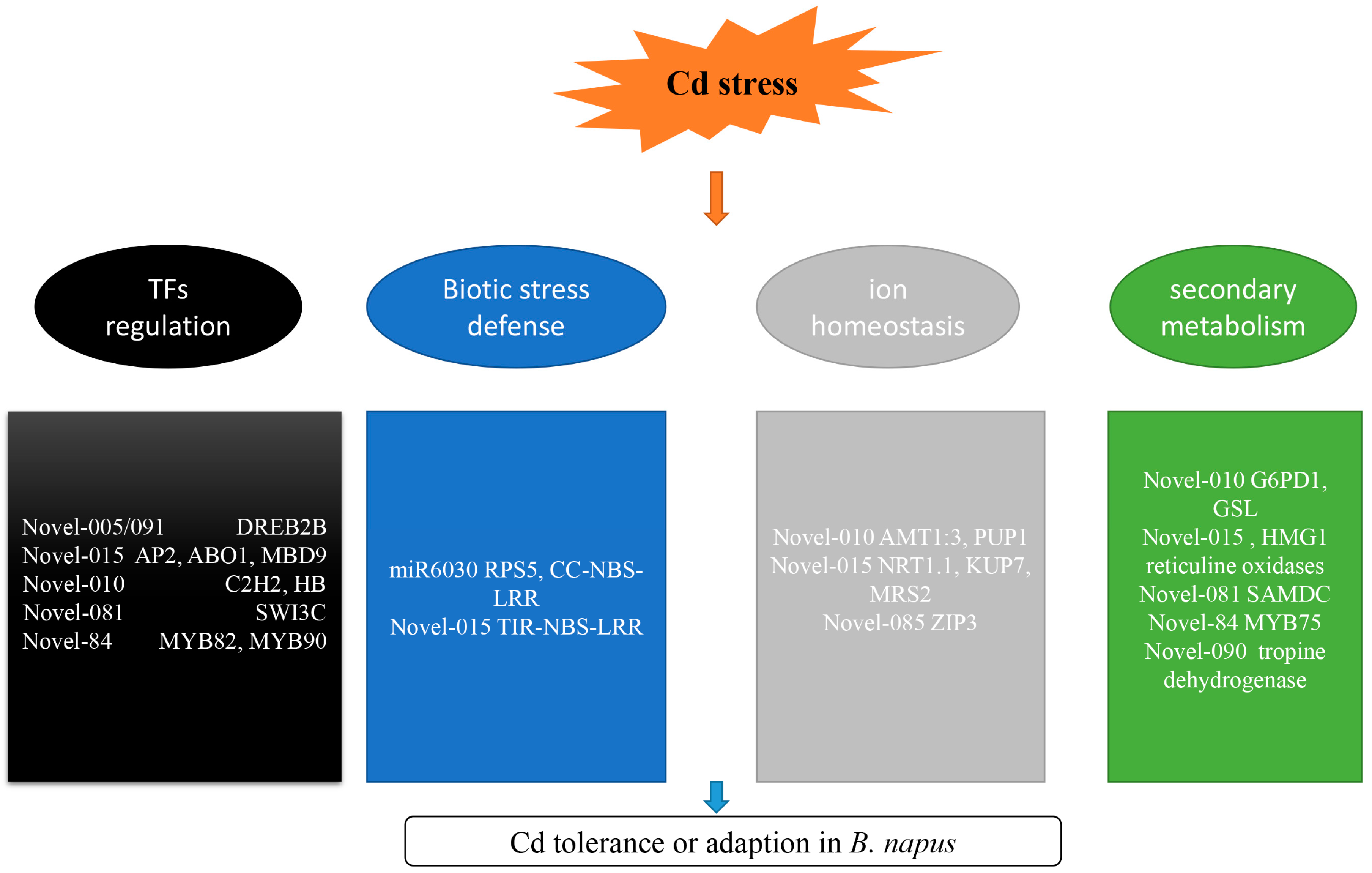
3. Discussion
3.1. miRNAs Involved in TF Regulation
3.2. miRNAs Involved in Biotic Stress Responses
3.3. miRNAs Involved in Ion Transport
3.4. miRNAs Involved in Secondary Metabolism
4. Materials and Methods
4.1. Plant Culture and Treatment
4.2. Construction and Sequencing of Small RNA Libraries
4.3. Analysis of Small RNA Sequencing Data
4.4. Differential Expression Analysis of miRNAs under Cd Stress
4.5. Prediction of miRNA Targets
4.6. Validation of Mature miRNAs and Corresponding Target Genes
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Xu, Y.C.; Chu, L.L.; Jin, Q.J.; Wang, Y.J.; Chen, X.; Zhao, H.; Xue, Z.Y. Transcriptome-Wide Identification of miRNAs and Their Targets from Typha angustifolia by RNA-Seq and Their Response to Cadmium Stress. PLoS ONE 2015, 10, e0125462. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Han, F.X.X.; Banin, A.; Su, Y.; Monts, D.L.; Plodinec, M.J.; Kingery, W.L.; Triplett, G.E. Industrial age anthropogenic inputs of heavy metals into the pedosphere. Naturwissenschaften 2002, 89, 497–504. [Google Scholar] [CrossRef] [PubMed]
- Nawrot, T.; Plusquin, M.; Hogervorst, J.; Roels, H.A.; Celis, H.; Thijs, L.; Vangronsveld, J.; Van Hecke, E.; Staessen, J.A. Environmental exposure to cadmium and risk of cancer: A prospective population-based study. Lancet Oncol. 2006, 7, 119–126. [Google Scholar] [CrossRef]
- Shamsi, I.H.; Wei, K.; Zhang, G.P.; Jilani, G.H.; Hassan, M.J. Interactive effects of cadmium and aluminum on growth and antioxidative enzymes in soybean. Biol. Plant. 2008, 52, 165–169. [Google Scholar] [CrossRef]
- Patra, M.; Bhowmik, N.; Bandopadhyay, B.; Sharma, A. Comparison of mercury, lead and arsenic with respect to genotoxic effects on plant systems and the development of genetic tolerance. Environ. Exp. Bot. 2004, 52, 199–223. [Google Scholar] [CrossRef]
- DalCorso, G.; Farinati, S.; Furini, A. Regulatory networks of cadmium stress in plants. Plant Signal. Behav. 2010, 5, 663–667. [Google Scholar] [CrossRef] [PubMed]
- Montero-Palmero, M.B.; Martin-Barranco, A.; Escobar, C.; Hernandez, L.E. Early transcriptional responses to mercury: A role for ethylene in mercury-induced stress. New Phytol. 2014, 201, 116–130. [Google Scholar] [CrossRef] [PubMed]
- Martinka, M.; Vaculik, M.; Lux, A. Plant Cell Responses to Cadmium and Zinc. Plant Cell Monogr. 2014, 22, 209–246. [Google Scholar]
- Patra, M.; Sharma, A. Mercury toxicity in plants. Bot. Rev. 2000, 66, 379–422. [Google Scholar] [CrossRef]
- Hassan, M.J.; Shao, G.S.; Zhang, G.P. Influence of cadmium toxicity on growth and antioxidant enzyme activity in rice cultivars with different grain cadmium accumulation. J. Plant Nutr. 2005, 28, 1259–1270. [Google Scholar] [CrossRef]
- Overmyer, K.; Brosche, M.; Kangasjarvi, J. Reactive oxygen species and hormonal control of cell death. Trends Plant Sci. 2003, 8, 335–342. [Google Scholar] [CrossRef]
- Edreva, A. Generation and scavenging of reactive oxygen species in chloroplasts: A submolecular approach. Agric. Ecosyst. Environ. 2005, 106, 119–133. [Google Scholar] [CrossRef]
- Kim, D.Y.; Bovet, L.; Maeshima, M.; Martinoia, E.; Lee, Y. The ABC transporter AtPDR8 is a cadmium extrusion pump conferring heavy metal resistance. Plant J. 2007, 50, 207–218. [Google Scholar] [CrossRef] [PubMed]
- Ueno, D.; Milner, M.J.; Yamaji, N.; Yokosho, K.; Koyama, E.; Zambrano, M.C.; Kaskie, M.; Ebbs, S.; Kochian, L.V.; Ma, J.F. Elevated expression of TcHMA3 plays a key role in the extreme Cd tolerance in a Cd-hyperaccumulating ecotype of Thlaspi caerulescens. Plant J. 2011, 66, 852–862. [Google Scholar] [CrossRef] [PubMed]
- Shukla, D.; Huda, K.M.K.; Banu, M.S.A.; Gill, S.S.; Tuteja, R.; Tuteja, N. OsACA6, a P-type 2B Ca2+ ATPase functions in cadmium stress tolerance in tobacco by reducing the oxidative stress load. Planta 2014, 240, 809–824. [Google Scholar] [CrossRef] [PubMed]
- Jonak, C.; Nakagami, H.; Hirt, H. Heavy metal stress. Activation of distinct mitogen-activated protein kinase pathways by copper and cadmium. Plant Physiol. 2004, 136, 3276–3283. [Google Scholar] [CrossRef] [PubMed]
- Maksymiec, W. Signaling responses in plants to heavy metal stress. Acta Physiol. Plant. 2007, 29, 177–187. [Google Scholar] [CrossRef]
- DalCorso, G.; Farinati, S.; Maistri, S.; Furini, A. How plants cope with cadmium: Staking all on metabolism and gene expression. J. Integr. Plant Biol. 2008, 50, 1268–1280. [Google Scholar] [CrossRef] [PubMed]
- Romero-Puertas, M.C.; Corpas, F.J.; Rodriguez-Serrano, M.; Gomez, M.; del Rio, L.A.; Sandalio, L.M. Differential expression and regulation of antioxidative enzymes by cadmium in pea plants. J. Plant Physiol. 2007, 164, 1346–1357. [Google Scholar] [CrossRef] [PubMed]
- De Mortel, J.E.V.; Schat, H.; Moerland, P.D.; Van Themaat, E.V.L.; Van der Ent, S.; Blankestijn, H.; Ghandilyan, A.; Tsiatsiani, S.; Aarts, M.G.M. Expression differences for genes involved in lignin, glutathione and sulphate metabolism in response to cadmium in Arabidopsis thaliana and the related Zn/Cd-hyperaccumulator Thlaspi caerulescens. Plant Cell Environ. 2008, 31, 301–324. [Google Scholar] [CrossRef] [PubMed]
- Fusco, N.; Micheletto, L.; Dal Corso, G.; Borgato, L.; Furini, A. Identification of cadmium-regulated genes by cDNA-AFLP in the heavy metal accumulator Brassica juncea L. J. Exp. Bot. 2005, 56, 3017–3027. [Google Scholar] [CrossRef] [PubMed]
- Meng, H.B.; Hua, S.J.; Shamsi, I.H.; Jilani, G.; Li, Y.L.; Jiang, L.X. Cadmium-induced stress on the seed germination and seedling growth of Brassica napus L.; and its alleviation through exogenous plant growth regulators. Plant Growth Regul. 2009, 58, 47–59. [Google Scholar] [CrossRef]
- Weber, M.; Trampczynska, A.; Clemens, S. Comparative transcriptome analysis of toxic metal responses in Arabidopsis thaliana and the Cd2+-hypertolerant facultative metallophyte Arabidopsis halleri. Plant Cell Environ. 2006, 29, 950–963. [Google Scholar] [CrossRef] [PubMed]
- Tamas, L.; Dudikova, J.; Durcekova, K.; Halugkova, L.; Huttova, J.; Mistrik, I.; Olle, M. Alterations of the gene expression, lipid peroxidation, proline and thiol content along the barley root exposed to cadmium. J. Plant Physiol. 2008, 165, 1193–1203. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Zhang, Y.X.; Han, L.; Guan, Z.Q.; Chai, T.Y. A novel WRKY transcriptional factor from Thlaspi caerulescens negatively regulates the osmotic stress tolerance of transgenic tobacco. Plant Cell Rep. 2008, 27, 795–803. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Koizumi, N.; Sano, H. Screening of cadmium-responsive genes in Arabidopsis thaliana. Plant Cell Environ. 2001, 24, 1177–1188. [Google Scholar] [CrossRef]
- Ding, Y.; Chen, Z.; Zhu, C. Microarray-based analysis of cadmium-responsive microRNAs in rice (Oryza sativa). J. Exp. Bot. 2011, 62, 3563–3573. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.F.; Mao, D.H.; Xu, L.W.; Li, D.Y.; Song, S.H.; Chen, C.Y. Integrated analysis of miRNA and mRNA expression profiles in response to Cd exposure in rice seedlings. BMC Genom. 2014, 15. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Wang, Y.; Zhai, L.L.; Xu, Y.Y.; Wang, L.J.; Zhu, X.W.; Gong, Y.Q.; Yu, R.G.; Limera, C.; Liu, L.W. Genome-wide identification and characterization of cadmium-responsive microRNAs and their target genes in radish (Raphanus sativus L.) roots. J. Exp. Bot. 2013, 64, 4271–4287. [Google Scholar] [CrossRef] [PubMed]
- Ali, E.; Maodzeka, A.; Hussain, N.; Shamsi, I.H.; Jiang, L.X. The alleviation of cadmium toxicity in oilseed rape (Brassica napus) by the application of salicylic acid. Plant Growth Regul. 2015, 75, 641–655. [Google Scholar] [CrossRef]
- Ali, B.; Qian, P.; Jin, R.; Ali, S.; Khan, M.; Aziz, R.; Tian, T.; Zhou, W. Physiological and ultra-structural changes in Brassica napus seedlings induced by cadmium stress. Biol. Plant. 2014, 58, 131–138. [Google Scholar] [CrossRef]
- Ali, B.; Wang, B.; Ali, S.; Ghani, M.A.; Hayat, M.T.; Yang, C.; Xu, L.; Zhou, W.J. 5-Aminolevulinic Acid Ameliorates the Growth, Photosynthetic Gas Exchange Capacity, and Ultrastructural Changes Under Cadmium Stress in Brassica napus L. J. Plant Growth Regul. 2013, 32, 604–614. [Google Scholar] [CrossRef]
- Ali, B.; Song, W.J.; Hu, W.Z.; Luo, X.N.; Gill, R.A.; Wang, J.; Zhou, W.J. Hydrogen sulfide alleviates lead-induced photosynthetic and ultrastructural changes in oilseed rape. Ecotoxicol. Environ. Saf. 2014, 102, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Jian, H.J.; Wang, J.; Wang, T.Y.; Wei, L.J.; Li, J.; Liu, L.Z. Identification of Rapeseed MicroRNAs Involved in Early Stage Seed Germination under Salt and Drought Stresses. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Jian, H.J.; Wang, T.Y.; Wei, L.J.; Li, J.N.; Li, C.; Liu, L.Z. Identification of microRNAs Actively Involved in Fatty Acid Biosynthesis in Developing Brassica napus Seeds Using High-Throughput Sequencing. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Hao, M.; Wang, W.; Mei, D.; Wells, R.; Liu, J.; Wang, H.; Sang, S.; Tang, M.; Zhou, R.; et al. Integrative RNA- and miRNA-Profile Analysis Reveals a Likely Role of BR and Auxin Signaling in Branch Angle Regulation of B. napus. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.Y.; Xu, Y.P.; Zhao, L.; Li, S.S.; Cai, X.Z. Tight regulation of the interaction between Brassica napus and Sclerotinia sclerotiorum at the microRNA level. Plant Mol. Biol. 2016, 92, 39–55. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.L.; Huang, S.Q.; Guo, K.; Xiang, A.L.; Zhu, Y.Y.; Nie, L.; Yang, Z.M. Computational identification of novel microRNAs and targets in Brassica napus. FEBS Lett. 2007, 581, 1464–1474. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.Q.; Xiang, A.L.; Che, L.L.; Chen, S.; Li, H.; Song, J.B.; Yang, Z.M. A set of miRNAs from Brassica napus in response to sulphate deficiency and cadmium stress. Plant Biotechnol. J. 2010, 8, 887–899. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.S.; Song, J.B.; Yang, Z.M. Genome-wide identification of Brassica napus microRNAs and their targets in response to cadmium. J. Exp. Bot. 2012, 63, 4597–4613. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A. and Griffiths-Jones, S. Mirbase: Integrating miRNAs annotation and deep-sequencing data. Nucleic. Acids Res. 2011, 39, 152–157. [Google Scholar] [CrossRef] [PubMed]
- Griffiths-Jones, S.; Saini, H.K.; Van Dongen, S.; Enright, A.J. Mirbase: Tools for miRNA genomics. Nucleic. Acids Res. 2008, 36, 154–158. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Zhao, P.X. Psrnatarget: A plant small RNA target analysis server. Nucleic. Acids Res. 2011, 39, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Carbon, S.; Ireland, A.; Mungall, C.J.; Shu, S.; Marshall, B.; Lewis, S.G.O. Hub ami, and group web presence working. amigo: Online access to ontology and annotation data. Bioinformatics 2009, 25, 288–289. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Sun, L.; Yang, X.E.; Liu, J.X. Transcriptomic Analysis of Cadmium Stress Response in the Heavy Metal Hyperaccumulator Sedum alfredii Hance. PLoS ONE 2013, 8, e64643. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.F.; Zhu, C. The role of microRNAs in copper and cadmium homeostasis. Biochem. Biophys. Res. Commun. 2009, 386, 6–10. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.B.; Foley, R.C.; Onate-Sanchez, L. Transcription factors in plant defense and stress responses. Curr. Opin. Plant Biol. 2002, 5, 430–436. [Google Scholar] [CrossRef]
- Hirai, M.Y.; Sugiyama, K.; Sawada, Y.; Tohge, T.; Obayashi, T.; Suzuki, A.; Araki, R.; Sakurai, N.; Suzuki, H.; Aoki, K.; et al. Omics-based identification of Arabidopsis Myb transcription factors regulating aliphatic glucosinolate biosynthesis. Proc. Natl. Acad. Sci. USA 2007, 104, 6478–6483. [Google Scholar] [CrossRef] [PubMed]
- Karasov, T.L.; Kniskern, J.M.; Gao, L.P.; DeYoung, B.J.; Ding, J.; Dubiella, U.; Lastra, R.O.; Nallu, S.; Roux, F.; Innes, R.W.; et al. The long-term maintenance of a resistance polymorphism through diffuse interactions. Nature 2014, 512, 436–440. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Millan, A.F.; Ellis, D.R.; Grusak, M.A. Identification and characterization of several new members of the ZIP family of metal ion transporters in Medicago truncatula. Plant Mol. Biol. 2004, 54, 583–596. [Google Scholar] [CrossRef] [PubMed]
- Kramer, U.; Talke, I.N.; Hanikenne, M. Transition metal transport. FEBS Lett. 2007, 581, 2263–2272. [Google Scholar] [CrossRef] [PubMed]
- Deckert, J. Cadmium toxicity in plants: Is there any analogy to its carcinogenic effect in mammalian cells? Biometals 2005, 18, 475–481. [Google Scholar] [CrossRef] [PubMed]
- Nevo, Y.; Nelson, N. The NRAMP family of metal-ion transporters. BBA Mol. Cell Res. 2006, 1763, 609–620. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.Y.; Bovet, L.; Kushnir, S.; Noh, E.W.; Martinoia, E.; Lee, Y. AtATM3 is involved in heavy metal resistance in Arabidopsis. Plant Physiol. 2006, 140, 922–932. [Google Scholar] [CrossRef] [PubMed]
- Pusztahelyi, T.; Holb, I.J.; Pocsi, I. Secondary metabolites in fungus-plant interactions. Front. Plant Sci. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Lorenc-Kukula, K.; Zuk, M.; Kulma, A.; Czemplik, M.; Kostyn, K.; Skala, J.; Starzycki, M.; Szopa, J. Engineering Flax with the GT Family 1 Solanum sogarandinum Glycosyltransferase SsGT1 Confers Increased Resistance to Fusarium Infection. J. Agric. Food Chem. 2009, 57, 6698–6705. [Google Scholar] [CrossRef] [PubMed]
- Shin, D.H.; Cho, M.; Choi, M.G.; Das, P.K.; Lee, S.K.; Choi, S.B.; Park, Y.I. Identification of genes that may regulate the expression of the transcription factor production of anthocyanin pigment 1 (PAP1)/MYB75 involved in Arabidopsis anthocyanin biosynthesis. Plant Cell Rep. 2015, 34, 805–815. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Shi, M.Z.; Xie, D.Y. Regulation of anthocyanin biosynthesis in Arabidopsis thaliana red pap1-D cells metabolically programmed by auxins. Planta 2014, 239, 765–781. [Google Scholar] [CrossRef] [PubMed]
- Qi, T.C.; Song, S.S.; Ren, Q.C.; Wu, D.W.; Huang, H.; Chen, Y.; Fan, M.; Peng, W.; Ren, C.M.; Xie, D.X. The Jasmonate-ZIM-Domain Proteins Interact with the WD-Repeat/bHLH/MYB Complexes to Regulate Jasmonate-Mediated Anthocyanin Accumulation and Trichome Initiation in Arabidopsis thaliana. Plant Cell 2011, 23, 1795–1814. [Google Scholar] [CrossRef] [PubMed]
- Leivar, P.; Antolin-Llovera, M.; Ferrero, S.; Closa, M.; Arro, M.; Ferrer, A.; Boronat, A.; Campos, N. Multilevel control of Arabidopsis 3-hydroxy-3-methylglutaryl coenzyme A reductase by protein phosphatase 2A. Plant Cell 2011, 23, 1494–1511. [Google Scholar] [CrossRef] [PubMed]
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.; Tang, H.; Wang, X.; Chiquet, J.; Belcram, H.; Tong, C.; Samans, B.; et al. Plant genetics. early allopolyploid evolution in the post-neolithic Brassica napus oilseed genome. Science 2014, 345, 950–953. [Google Scholar] [CrossRef] [PubMed]
- Li, R.Q.; Yu, C.; Li, Y.R.; Lam, T.W.; Yiu, S.M.; Kristiansen, K.; Wang, J. SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 2009, 25, 1966–1967. [Google Scholar] [CrossRef] [PubMed]
- Meyers, B.C.; Axtell, M.J.; Bartel, B.; Bartel, D.P.; Baulcombe, D.; Bowman, J.L.; Cao, X.; Carrington, J.C.; Chen, X.M.; Green, P.J.; et al. Criteria for Annotation of Plant MicroRNAs. Plant Cell 2008, 20, 3186–3190. [Google Scholar] [CrossRef] [PubMed]

| Samples | Raw_Reads | Low_Quality | containing’n’reads | Length < 18 | Length > 30 | Clean_Reads |
|---|---|---|---|---|---|---|
| CK | 16,892,251 | 0 | 1164 | 575,989 | 2,514,501 | 13,800,597 |
| T1 | 20,019,156 | 0 | 1413 | 799,829 | 5,296,926 | 13,920,988 |
| T3 | 19,891,839 | 0 | 1417 | 1,122,546 | 2,965,574 | 15,802,302 |
| miRNA | Length | CK | T1 | T3 | Mature Sequence |
|---|---|---|---|---|---|
| miR1140 | 21 | 99.84 | 44.49 | 88.45 | ACAGCCTAAACCAATCGGAGC |
| miR156a | 21 | 25,154.03 | 16,041.64 | 13,758.44 | TGACAGAAGAGAGTGAGCACA |
| miR156b | 21 | 17,575.97 | 17,616.44 | 24,364.75 | TTGACAGAAGATAGAGAGCAC |
| miR156d | 20 | 25,634.20 | 16,450.91 | 14,216.79 | TGACAGAAGAGAGTGAGCAC |
| miR159 | 21 | 283,188.49 | 271,657.99 | 306,215.83 | TTTGGATTGAAGGGAGCTCTA |
| miR160a | 21 | 2976.08 | 1975.18 | 3031.52 | TGCCTGGCTCCCTGTATGCCA |
| miR161 | 21 | 57.05 | 53.38 | 152.78 | TCAATGCACTGAAAGTGACTA |
| miR164a | 21 | 2362.80 | 2411.14 | 2621.42 | TGGAGAAGCAGGGCACGTGCA |
| miR164b | 21 | 969.84 | 898.62 | 1045.35 | TGGAGAAGCAGGGCACGTGCG |
| miR166a | 21 | 100,777.77 | 105,636.37 | 92,513.67 | TCGGACCAGGCTTCATTCCCC |
| miR166f | 21 | 22,025.82 | 21,807.02 | 13,613.70 | TCGGACCAGGCTTCATCCCCC |
| miR167a | 22 | 13,753.66 | 10,694.43 | 9504.66 | TGAAGCTGCCAGCATGATCTAA |
| miR167c | 21 | 13,810.71 | 10,694.43 | 9617.24 | TGAAGCTGCCAGCATGATCTA |
| miR167d | 20 | 5357.89 | 3905.87 | 4615.63 | TGAAGCTGCCAGCATGATCT |
| miR168a | 21 | 13,929.56 | 14,404.56 | 14,787.71 | TCGCTTGGTGCAGGTCGGGAA |
| miR169a | 21 | 19.02 | 17.79 | 40.21 | CAGCCAAGGATGACTTGCCGA |
| miR169g | 22 | 9.51 | 0.00 | 0.00 | TAGCCAAGGATGACTTGCCTGC |
| miR169m | 21 | 19.02 | 8.90 | 24.12 | TGAGCCAAAGATGACTTGCCG |
| miR169n | 21 | 23.77 | 44.49 | 104.54 | CAGCCAAGGATGACTTGCCGG |
| miR171a | 21 | 456.40 | 631.70 | 321.65 | TTGAGCCGTGCCAATATCACG |
| miR171f | 21 | 1150.50 | 845.23 | 1085.56 | TGATTGAGCCGCGCCAATATC |
| miR171g | 22 | 1150.50 | 845.23 | 1085.56 | TGATTGAGCCGCGCCAATATCT |
| miR172a | 21 | 123.61 | 124.56 | 88.45 | AGAATCTTGATGATGCTGCAT |
| miR172b | 21 | 199.67 | 373.68 | 225.15 | GGAATCTTGATGATGCTGCAT |
| miR172d | 21 | 23.77 | 26.69 | 16.08 | AGAATCTTGATGATGCTGCAG |
| miR2111a-3p | 21 | 38.03 | 44.49 | 16.08 | GTCCTCGGGATGCGGATTACC |
| miR2111a-5p | 21 | 0.00 | 0.00 | 16.08 | TAATCTGCATCCTGAGGTTTA |
| miR2111b-3p | 21 | 85.57 | 35.59 | 64.33 | ATCCTCGGGATACAGATTACC |
| miR2111c | 21 | 4.75 | 0.00 | 0.00 | TAATCTGCATCCTGGGGTTTA |
| miR390a | 21 | 2700.34 | 1788.34 | 1648.44 | AAGCTCAGGAGGGATAGCGCC |
| miR393 | 21 | 52.30 | 44.49 | 80.41 | TCCAAAGGGATCGCATTGATC |
| miR394a | 20 | 3342.14 | 3016.15 | 3337.09 | TTGGCATTCTGTCCACCTCC |
| miR395a | 21 | 16,382.69 | 21,735.84 | 27,243.49 | CTGAAGTGTTTGGGGGAACTC |
| miR395d | 21 | 6608.22 | 6317.01 | 5588.61 | CTGAAGTGTTTGGGGGGACTC |
| miR397a | 22 | 4.75 | 17.79 | 8.04 | TCATTGAGTGCAGCGTTGATGT |
| miR399a | 21 | 0.00 | 8.90 | 0.00 | TGCCAAAGGAGATTTGCCCGG |
| miR403 | 21 | 26,927.32 | 24,342.72 | 26,310.71 | TTAGATTCACGCACAAACTCG |
| miR6029 | 21 | 256.72 | 302.50 | 217.11 | TGGGGTTGTGATTTCAGGCTT |
| miR6030 | 22 | 1093.45 | 818.54 | 522.68 | TCCACCCATACCATACAGACCC |
| miR6031 | 24 | 1179.02 | 1574.80 | 868.45 | AAGAGGTTCGGAGCGGTTTGAAGC |
| miR6034 | 21 | 19.02 | 26.69 | 88.45 | TCTGATGTATATAGCTTTGGG |
| miR6035 | 21 | 61.80 | 160.15 | 88.45 | TGGAGTAGAAAATGCAGTCGT |
| miR824 | 21 | 1911.16 | 1823.92 | 1286.59 | TAGACCATTTGTGAGAAGGGA |
| miR860 | 21 | 4.75 | 8.90 | 32.16 | TCAATACATTGGACTACATAT |
| miRNA | Length | CK | T1 | T3 | Hairpin Energy (kcal moL−1) | Mature Sequence |
|---|---|---|---|---|---|---|
| Novel-001 | 21 | 128.4 | 160.1 | 72.4 | −80.5 | auaacuugguuuugcuccuac |
| Novel-002 | 21 | 12,845.6 | 12,011.2 | 14,924.4 | −67.3 | cggcucugauaccaauugaug |
| Novel-003 | 23 | 1625.9 | 2117.5 | 3264.7 | −78.5 | ugcucacggcucuuucugucagu |
| Novel-004 | 21 | 228.2 | 213.5 | 193.0 | −73.8 | ugccaaaggagaguugcccug |
| Novel-005 | 20 | 85.6 | 231.3 | 386.0 | −61.2 | uguguucucaggucaccccu |
| Novel-006 | 21 | 366.1 | 284.7 | 587.0 | −51.2 | uuggagcaucgagugaagagc |
| Novel-007 | 21 | 4278.7 | 3772.4 | 3393.4 | −67.3 | ucgauaaaccucugcauccag |
| Novel-008 | 24 | 2386.6 | 952.0 | 554.8 | −72.9 | gaugacgguaucucuccuacguag |
| Novel-009 | 22 | 23.8 | 17.8 | 217.1 | −119.2 | cagaagauguaugcuaaauugg |
| Novel-010 | 18 | 0.0 | 17.8 | 128.7 | −49.4 | gaggaaugaggaucauug |
| Novel-011 | 24 | 19,216.1 | 30,481.8 | 22,209.7 | −69.5 | agagauuuuuguuacuguuaacug |
| Novel-012 | 25 | 118.9 | 275.8 | 635.3 | −57.7 | gucaauugauggguaguaguucauu |
| Novel-013 | 24 | 2752.6 | 2811.5 | 1889.7 | −60.5 | agccuggcucugauaccaugaagu |
| Novel-014 | 21 | 294.8 | 266.9 | 241.2 | −68.3 | gacuuauaauaaucucaugaa |
| Novel-015 | 18 | 57.0 | 0.0 | 48.2 | −69.1 | cuuccaagaaaagcuuca |
| Novel-016 | 20 | 789.2 | 765.2 | 675.5 | −75.9 | ugaaugucuuucucuucauc |
| Novel-017 | 21 | 1141.0 | 1512.5 | 297.5 | −76.1 | uuguggaaccgugugaauacc |
| Novel-018 | 24 | 57.0 | 53.4 | 72.4 | −55.4 | ugaugugucaugucuagaaauccu |
| Novel-019 | 21 | 95.1 | 53.4 | 64.3 | −95.6 | uccccaguuuggauuguuugc |
| Novel-020 | 21 | 9555.8 | 7028.8 | 8202.0 | −88 | uaugugugcucacucucuauc |
| Novel-021 | 24 | 42.8 | 80.1 | 72.4 | −56.1 | acaggugguggaacaaauaugagu |
| Novel-022 | 21 | 713.1 | 409.3 | 289.5 | −72.1 | cgauauugguacgguugaauc |
| Novel-023 | 21 | 746.4 | 596.1 | 1101.6 | −76.4 | gauccucugaacacuucauug |
| Novel-024 | 24 | 142.6 | 329.2 | 176.9 | −135.7 | auuuuuagaacuucaauggguaga |
| Novel-025 | 21 | 171.1 | 160.1 | 257.3 | −62.4 | aucaugcgaucucuucggauu |
| Novel-026 | 21 | 26,375.8 | 21,495.6 | 24,670.3 | −82.4 | gcucucuagucuucugucauc |
| Novel-027 | 21 | 351.8 | 249.1 | 482.5 | −75.1 | guucccugaaacgcuucauug |
| Novel-028 | 20 | 8804.6 | 9164.1 | 9794.1 | −80.6 | uuggacugaagggagcuccu |
| Novel-029 | 21 | 309.0 | 302.5 | 442.3 | −83.2 | uuuugcguuucaacucggucc |
| Novel-030 | 21 | 13,311.5 | 10854.6 | 12,029.6 | −70.1 | cuugcauaucuuaggagcuuu |
| Novel-031 | 21 | 5191.5 | 2953.9 | 2830.5 | −67.3 | guucaauaaagcugugggaag |
| Novel-032 | 21 | 931.8 | 889.7 | 772.0 | −80.4 | uuggacugaagggaacucccu |
| Novel-033 | 24 | 156.9 | 222.4 | 241.2 | −53.5 | agcuaugguuuauguggacucagu |
| Novel-034 | 22 | 27,345.7 | 25,695.1 | 24,799.0 | −72.4 | gcucacugcucuuucugucaga |
| Novel-035 | 23 | 1606.9 | 1085.5 | 852.4 | −74.3 | gcucauucucguucugucauaac |
| Novel-036 | 21 | 1882.6 | 8719.2 | 10,807.3 | −69.9 | uguguucucaggucaccccug |
| Novel-037 | 21 | 1331.2 | 1245.6 | 1769.1 | −88.6 | gcuuacucucucucugucacc |
| Novel-038 | 23 | 156.9 | 302.5 | 233.2 | −58.6 | uucggaccaggcuucauucccca |
| Novel-039 | 23 | 608.5 | 1254.5 | 876.5 | −80 | guagguagacgcacuguuucuca |
| Novel-040 | 24 | 47.5 | 71.2 | 48.2 | −85.1 | ugaguuaucauuggucuuguguca |
| Novel-041 | 21 | 936.6 | 498.2 | 619.2 | −71.8 | gacuuauaaugaucucaugaa |
| Novel-042 | 21 | 185.4 | 329.2 | 209.1 | −100.9 | uugguuuaacuuggauuuuga |
| Novel-043 | 22 | 855.7 | 373.7 | 410.1 | −70.6 | cguacagaguagucaagcauga |
| Novel-044 | 21 | 256.7 | 462.7 | 257.3 | −65.1 | uguuuuguggguuucuaccga |
| Novel-045 | 21 | 2838.2 | 1868.4 | 3015.4 | −95.5 | cccgccuugcaucaacugaau |
| Novel-046 | 21 | 23,542.4 | 23,648.7 | 19,234.5 | −83.2 | uuggacugaagggagcucccu |
| Novel-047 | 24 | 584.8 | 898.6 | 619.2 | −98.7 | auauuccgauaagaacuucacucu |
| Novel-048 | 22 | 261.5 | 240.2 | 345.8 | −79.4 | uuucaucuuagagaauguuguc |
| Novel-049 | 22 | 551.5 | 284.7 | 241.2 | −63.6 | gcucucuauacuucugucaaua |
| Novel-050 | 23 | 2828.7 | 2615.8 | 2098.7 | −53.2 | uuggucacgugacuugugcuuua |
| Novel-051 | 24 | 5434.0 | 9635.7 | 6577.7 | −57.3 | aaacagcgguauucugacggacau |
| Novel-052 | 24 | 180.7 | 204.6 | 128.7 | −49.4 | aacuucggcuaagagauaguucuu |
| Novel-053 | 24 | 142.6 | 391.5 | 80.4 | −49.7 | aaauagaucuuuuugaaacaaauu |
| Novel-054 | 24 | 271.0 | 258.0 | 329.7 | −60.1 | auacagacucauauggaccaugug |
| Novel-055 | 21 | 375.6 | 293.6 | 514.6 | −83.8 | gcuuacucucucucugucacc |
| Novel-056 | 24 | 118.9 | 124.6 | 48.2 | −51.2 | aaagagcgacuauaguauaacuau |
| Novel-057 | 23 | 565.7 | 783.0 | 562.9 | −67.3 | agacuguaucuuauauuaugcua |
| Novel-058 | 21 | 266.2 | 160.1 | 144.7 | −67.5 | uugacagaagagagcgagcac |
| Novel-059 | 23 | 988.9 | 1067.7 | 1479.6 | −88 | ugcucaccucucuuucugucagu |
| Novel-060 | 21 | 1835.1 | 1708.3 | 1688.6 | −85.5 | uuuggauugaagggagcuccu |
| Novel-061 | 24 | 2572.0 | 4279.5 | 3473.8 | −101.8 | aggauuucauuuuccgucggaaug |
| Novel-062 | 24 | 14.3 | 177.9 | 64.3 | −32.2 | aaacgacguuguuuuauagcucuc |
| Novel-063 | 24 | 1373.9 | 2224.3 | 1897.7 | −84.5 | aggauuucauuuuccgucggaaug |
| Novel-064 | 24 | 3123.5 | 5035.8 | 4318.1 | −114 | aggauuucauuuuccgucggaaug |
| Novel-065 | 24 | 137.9 | 71.2 | 112.6 | −26.9 | aucuuaauugauugacaacucagc |
| Novel-066 | 24 | 589.5 | 702.9 | 418.1 | −61.1 | gauuaacccggaguucuuagaaug |
| Novel-067 | 24 | 285.2 | 427.1 | 217.1 | −65.8 | aggaugugguucuuagcggaaguu |
| Novel-068 | 24 | 347.1 | 240.2 | 249.3 | −36.9 | auaugaucugcacuuuugaacugu |
| Novel-069 | 20 | 7644.6 | 8399.0 | 8346.7 | −77 | ucccaaauguagacaaagca |
| Novel-070 | 24 | 3670.2 | 4341.8 | 3304.9 | −59 | agaauccgucucuuaacuuuuaac |
| Novel-071 | 21 | 698.9 | 1307.9 | 916.7 | −76.8 | uaagcugccagcaugaucuug |
| Novel-072 | 24 | 156.9 | 195.7 | 112.6 | −87.5 | aacgauuuuuguuacugauaacug |
| Novel-073 | 21 | 114.1 | 213.5 | 112.6 | −63.2 | uaugagaguauuauaagucac |
| Novel-074 | 21 | 389.8 | 231.3 | 225.2 | −171.9 | uaagacgagaaauugacaucc |
| Novel-075 | 24 | 247.2 | 480.4 | 659.4 | −60.1 | gucaacugauggugagagagaggu |
| Novel-076 | 21 | 332.8 | 756.3 | 1117.7 | −86.8 | uuguagaauuuugggaagggc |
| Novel-077 | 21 | 332.8 | 756.3 | 1166.0 | −84.3 | uuguagaauuuugggaagggc |
| Novel-078 | 21 | 318.5 | 444.9 | 450.3 | −104.3 | auuuucgauugguggauugug |
| Novel-079 | 22 | 556.2 | 320.3 | 249.3 | −73.6 | uuuugccuacuccucccauacc |
| Novel-080 | 20 | 8747.6 | 9173.0 | 9770.0 | −80.7 | uuggacugaagggagcuccu |
| Novel-081 | 21 | 38.0 | 80.1 | 160.8 | −55.2 | uggaugaugcuuggcucgaga |
| Novel-082 | 22 | 294.8 | 533.8 | 667.4 | −78.9 | ugguauugguaguaaugagugu |
| Novel-083 | 21 | 23,875.2 | 23,897.9 | 19,491.8 | −87 | uuggacugaagggagcucccu |
| Novel-084 | 22 | 76.1 | 106.8 | 209.1 | −70.1 | ucuugcuuaaauggguauucca |
| Novel-085 | 24 | 5196.2 | 15,258.7 | 5315.2 | −32.2 | agugcuauuuuugugacuuuugac |
| Novel-086 | 21 | 99,598.8 | 102,388.9 | 109,022.2 | −79.3 | uuuccaaauguagacaaagca |
| Novel-087 | 21 | 4350.0 | 4964.6 | 5693.1 | −59.8 | uuagaugaccaucaacaaaua |
| Novel-088 | 21 | 418.4 | 409.3 | 530.7 | −65.5 | ggacuguugucuggcucgagg |
| Novel-089 | 24 | 404.1 | 409.3 | 394.0 | −79.6 | caccgagguucguacggaggacgu |
| Novel-090 | 23 | 76.1 | 169.0 | 88.5 | −68.6 | gauacaugguuuguuagcugaua |
| Novel-091 | 20 | 42.8 | 106.8 | 201.0 | −53.3 | uguguucucaggucaccccu |
| Novel-092 | 21 | 8785.6 | 7687.2 | 6915.4 | −64.9 | ucgauaaaccucugcauccag |
| Novel-093 | 21 | 394.6 | 355.9 | 587.0 | −63.1 | uuugauuaggagcucauuguu |
| Novel-094 | 21 | 161.6 | 160.1 | 128.7 | −67.8 | ggacuguugucuggcucgagg |
| Novel-095 | 24 | 432.6 | 409.3 | 289.5 | −54.7 | aaaccgauucuauuagagcaugau |
| Novel-096 | 21 | 36,853.9 | 36,327.2 | 29,527.2 | −97.3 | uuggacugaagggagcucccu |
| Novel-097 | 22 | 451.6 | 524.9 | 377.9 | −75.12 | cuuaucucgaacgacuuucucg |
| Novel-098 | 24 | 52.3 | 275.8 | 297.5 | −88.1 | guuauuggaguuaauagagaaccg |
| Novel-099 | 21 | 841.5 | 6753.0 | 8177.9 | −65.4 | acagggaacaagcagagcaug |
| Novel-100 | 21 | 1359.7 | 783.0 | 675.5 | −66 | uucgcaggagagauagcgcca |
| Novel-101 | 24 | 1963.5 | 2188.7 | 2267.6 | −123.2 | ucuuaacuucggauaagagacggu |
| Novel-102 | 24 | 1621.2 | 1743.8 | 1616.3 | −74.7 | uauuauagagugaaccauuggagu |
| Novel-103 | 24 | 242.5 | 436.0 | 418.1 | −36.3 | aaugauaaggcgacuguaccagcu |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jian, H.; Yang, B.; Zhang, A.; Ma, J.; Ding, Y.; Chen, Z.; Li, J.; Xu, X.; Liu, L. Genome-Wide Identification of MicroRNAs in Response to Cadmium Stress in Oilseed Rape (Brassica napus L.) Using High-Throughput Sequencing. Int. J. Mol. Sci. 2018, 19, 1431. https://doi.org/10.3390/ijms19051431
Jian H, Yang B, Zhang A, Ma J, Ding Y, Chen Z, Li J, Xu X, Liu L. Genome-Wide Identification of MicroRNAs in Response to Cadmium Stress in Oilseed Rape (Brassica napus L.) Using High-Throughput Sequencing. International Journal of Molecular Sciences. 2018; 19(5):1431. https://doi.org/10.3390/ijms19051431
Chicago/Turabian StyleJian, Hongju, Bo Yang, Aoxiang Zhang, Jinqi Ma, Yiran Ding, Zhiyou Chen, Jiana Li, Xinfu Xu, and Liezhao Liu. 2018. "Genome-Wide Identification of MicroRNAs in Response to Cadmium Stress in Oilseed Rape (Brassica napus L.) Using High-Throughput Sequencing" International Journal of Molecular Sciences 19, no. 5: 1431. https://doi.org/10.3390/ijms19051431
APA StyleJian, H., Yang, B., Zhang, A., Ma, J., Ding, Y., Chen, Z., Li, J., Xu, X., & Liu, L. (2018). Genome-Wide Identification of MicroRNAs in Response to Cadmium Stress in Oilseed Rape (Brassica napus L.) Using High-Throughput Sequencing. International Journal of Molecular Sciences, 19(5), 1431. https://doi.org/10.3390/ijms19051431






