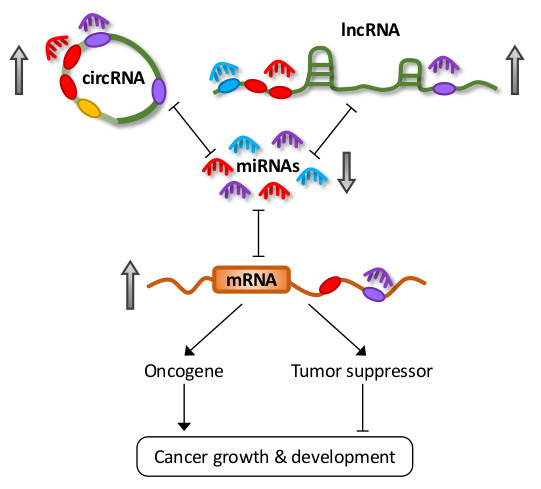
Noncoding RNA:RNA Regulatory Networks in Cancer
Abstract
1. Introduction
2. Competing Endogenous RNA (ceRNA) Networks and Regulation
3. The Links between Long Noncoding RNAs and microRNAs
3.1. Cytoplasmic lncRNAs
3.1.1. H19
3.1.2. Growth Arrest-Specific 5 (GAS5)
3.1.3. LincRNA, Regulator of Reprogramming (Linc-ROR)
3.1.4. Noncoding RNA Activated by DNA Damage (NORAD)
3.2. Nuclear LncRNAs
3.2.1. X-Inactive Transcript (XIST)
3.2.2. Nuclear Enriched Abundant Transcript 1 (NEAT1)
3.2.3. Metastasis Associated Lung Adenocarcinoma Transcript 1 (MALAT1)
3.2.4. Plasmacytoma Variant Translocation 1 (PVT1)
3.3. Antisense RNAs
3.3.1. Hox Transcript Antisense RNA (HOTAIR)
3.3.2. HOXD Antisense Growth-Associated lncRNA (HOXD-AS1)
3.4. Pseudogenes
3.4.1. Tumor Suppressive Pseudogenes
3.4.2. Oncogenic Pseudogenes
3.5. Circular RNAs
3.5.1. Cerebellar Degeneration-Related Protein 1 Antisense RNA (CDR1as)
3.5.2. Circ-ITCH (Itchy E3 Ubiquitin Protein Ligase)
3.5.3. CircHIPK3 (Homeodomain Interacting Protein Kinase 3)
3.5.4. CircPVT1
3.5.5. Other Newly Identified circRNAs
4. The Impact of Cellular Localization on miRNA-Mediated Gene Regulation
5. The Diagnostic and Prognostic Potential of ceRNA Interactions
6. Closing Remarks
Acknowledgments
Conflicts of Interest
Abbreviations
| ADAR2 | adenosine deaminase, RNA specific B1 |
| AGO2 | argonaute 2, RISC catalytic component |
| AKT | AKT serine/threonine kinase 1/protein kinase B |
| ANXA1 | annexin A1 |
| AR | androgen receptor |
| ARHI | DIRAS family GTPase 3 |
| ATG7 | autophagy related 7 |
| AVPR2 | arginine vasopressin receptor 2 |
| AXL | AXL receptor tyrosine kinase |
| BCL2 | B-cell lymphoma 2, apoptosis regulator |
| BECN1 | beclin 1 |
| BMF | Bcl2 modifying factor |
| BRAF | B-Raf proto-oncogene, serine/threonine kinase |
| BRAFP1 | B-Raf proto-oncogene, serine/threonine kinase pseudogene 1 |
| BUB1 | BUB1 mitotic checkpoint serine/threonine kinase |
| CCND1 | cyclin D1 |
| CCNE1 | cyclin E1 |
| CCNJ | cyclin J |
| CDC42 | cell division cycle 42 |
| CDK3 | cyclin dependent kinase 3 |
| CDK6 | cyclin dependent kinase 6 |
| ceRNA | competing endogenous RNA |
| c-Kit | KIT proto-oncogene receptor tyrosine kinase |
| c-Met | MET proto-oncogene, receptor tyrosine kinase |
| CRC | colorectal carcinoma |
| CRISPR | clustered regularly interspaced short palindromic repeats |
| CTNNA1 | catenin alpha 1 |
| CTNNA1P | catenin alpha 1 pseudogene |
| CXCR4 | C-X-C motif chemokine receptor 4 |
| CYP4Z1 | cytochrome P450 family 4 subfamily Z member 1 |
| CYP4Z2P | cytochrome P450 family 4 subfamily Z member 2, pseudogene |
| CYTH3 | cytohesin 3 |
| DNMT3B | DNA methyltransferase 3 beta |
| DRCE | drug-response related ceRNA |
| DVL2 | disheveled segment polarity protein 2 |
| EGFR | epidermal growth factor receptor |
| EGR1 | early growth response 1 |
| EMT | epithelial mesenchymal transition |
| EPHA4 | EPH receptor A4 |
| EZH2 | enhancer of zeste 2 polycomb repressive complex 2 subunit |
| FAK | focal adhesion kinase |
| FANCC | Fanconi anemia complementation group C |
| FASN | fatty acid synthase |
| FOXO3 | forkhead box O3 |
| FOXO3P | forkhead box O3 pseudogene |
| FTH1 | ferritin heavy chain 1 |
| FZD4 | frizzled class receptor 4 |
| GIT2 | GIT ArfGAP 2 |
| HCC | hepatocellular carcinoma |
| HER2 | erb-b2 receptor tyrosine kinase 2 |
| HIF-1α | hypoxia-inducible factor 1-alpha |
| HMGA1P6/7 | high mobility group AT-hook 1 pseudogene 6/7 |
| HMGA2 | high mobility group AT-hook 2 |
| HMGB1 | high mobility group box 1 |
| hTERT | human telomerase reverse transcriptase |
| IGF1R | insulin like growth factor 1 receptor |
| IGF2 | insulin like growth factor 2 |
| INST6 | integrator complex subunit 6 |
| INTS6P1 | integrator complex subunit 6 pseudogene 1 |
| KRAS | KRAS proto-oncogene, GTPase |
| LIN28 | lin-28 homolog |
| lncRNA | long noncoding RNA |
| LSCC | laryngeal squamous cell carcinoma |
| MCL1 | MCL1, BCL2 family apoptosis regulator |
| MDR1 | ATP binding cassette subfamily B member 1 |
| miRNA | microRNA |
| MKI67 | marker of proliferation Ki-67 |
| MMP2 | matrix metallopeptidase 2 |
| MRP1 | ATP binding cassette subfamily C member 1 |
| MYCN | MYCN proto-oncogene, bHLH transcription factor |
| NF-κB | nuclear factor kappa B |
| NSCLC | non-small cell lung cancer |
| OCT4/POU5F1 | POU class 5 homeobox 1 |
| OSCC | oral squamous cell carcinoma |
| PDK1 | pyruvate dehydrogenase kinase 1 |
| PI3K | phosphatidylinositol-4,5-bisphosphate 3-kinase |
| PIK3CD | phosphatidylinositol-4,5-Bisphosphate 3-Kinase Catalytic Subunit Delta |
| PLXNC1 | plexin C1 |
| PPIB | peptidylprolyl isomerase B |
| PRC2 | polycomb repressive complex 2 |
| PTEN | phosphatase and tensin homolog |
| PTENP1 | phosphatase and tensin homolog pseudogene 1 |
| RELA | RELA proto-oncogene, NF-κB subunit |
| RHOA | ras homolog family member A |
| RISC | RNA-induced silencing complex |
| ROCK2 | Rho associated coiled-coil containing protein kinase 2 |
| RSU1P2 | Ras suppressor protein 1 pseudogene 2 |
| SMAD7 | SMAD family member 7 |
| SOCS6 | suppressor of cytokine signaling 6 |
| SOX9 | SRY (sex determining region Y)-box 9 |
| STAT3 | signal transducer and activator of transcription 3 |
| TEAD | TEA domain transcription factor |
| TIMP2/3 | tissue inhibitor of metalloproteinases 2/3 |
| TNRC6 | trinucleotide repeat containing 6A |
| TP53 | tumor protein p53 |
| TUSC2 | tumor suppressor 2, mitochondrial calcium regulator |
| TWIST1 | twist family bHLH transcription factor 1 |
| UCA1 | urothelial cancer associated 1 |
| VEGFA | vascular endothelial growth factor A |
| VEGFR2 | kinase insert domain receptor |
| VIM | vimentin |
| YAP1 | Yes associated protein 1 |
| YY1 | Yin Yang 1 transcription factor |
| ZEB1/2 | zinc finger E-box binding homeobox 1/2 |
References
- Carninci, P.; Kasukawa, T.; Katayama, S.; Gough, J.; Frith, M.C.; Maeda, N.; Oyama, R.; Ravasi, T.; Lenhard, B.; Wells, C.; et al. The transcriptional landscape of the mammalian genome. Science 2005, 309, 1559–1563. [Google Scholar] [PubMed]
- Djebali, S.; Davis, C.A.; Merkel, A.; Dobin, A.; Lassmann, T.; Mortazavi, A.; Tanzer, A.; Lagarde, J.; Lin, W.; Schlesinger, F.; et al. Landscape of transcription in human cells. Nature 2012, 489, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P.; Chen, C.Z. Micromanagers of gene expression: The potentially widespread influence of metazoan microRNAs. Nat. Rev. Genet. 2004, 5, 396–400. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Rinn, J.L. Modular regulatory principles of large non-coding RNAs. Nature 2012, 482, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Cech, T.R.; Steitz, J.A. The noncoding RNA revolution-trashing old rules to forge new ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef] [PubMed]
- Adams, B.D.; Anastasiadou, E.; Esteller, M.; He, L.; Slack, F.J. The Inescapable Influence of Noncoding RNAs in Cancer. Cancer Res. 2015, 75, 5206–5210. [Google Scholar] [CrossRef] [PubMed]
- Kwok, Z.H.; Tay, Y. Long noncoding RNAs: Lincs between human health and disease. Biochem. Soc. Trans. 2017, 45, 805–812. [Google Scholar] [CrossRef] [PubMed]
- Anastasiadou, E.; Jacob, L.S.; Slack, F.J. Non-coding RNA networks in cancer. Nat. Rev. Cancer 2018, 18, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Bartel, D.P. Metazoan MicroRNAs. Cell 2018, 173, 20–51. [Google Scholar] [CrossRef] [PubMed]
- Bracken, C.P.; Scott, H.S.; Goodall, G.J. A network-biology perspective of microRNA function and dysfunction in cancer. Nat. Rev. Genet. 2016, 17, 719–732. [Google Scholar] [CrossRef] [PubMed]
- Svoronos, A.A.; Engelman, D.M.; Slack, F.J. OncomiR or Tumor Suppressor? The Duplicity of MicroRNAs in Cancer. Cancer Res. 2016, 76, 3666–3670. [Google Scholar] [CrossRef] [PubMed]
- Quinn, J.J.; Chang, H.Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 2016, 17, 47–62. [Google Scholar] [CrossRef] [PubMed]
- Ward, M.; McEwan, C.; Mills, J.D.; Janitz, M. Conservation and tissue-specific transcription patterns of long noncoding RNAs. J. Hum. Transcr. 2015, 1, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Xiao, Y.; Huang, F.; Deng, W.; Zhao, H.; Shi, X.; Wang, S.; Yu, X.; Zhang, L.; Han, Z.; et al. Spatiotemporal-specific lncRNAs in the brain, colon, liver and lung of macaque during development. Mol. Biosyst. 2015, 11, 3253–3263. [Google Scholar] [CrossRef] [PubMed]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The Rosetta Stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Tay, Y.; Rinn, J.; Pandolfi, P.P. The multilayered complexity of ceRNA crosstalk and competition. Nature 2014, 505, 344–352. [Google Scholar] [CrossRef] [PubMed]
- Poliseno, L.; Marranci, A.; Pandolfi, P.P. Pseudogenes in Human Cancer. Front. Med. (Lausanne) 2015, 2, 68. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Croce, C.M. The role of MicroRNAs in human cancer. Signal Transduct. Target. Ther. 2016, 1, 15004. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, L.S.; Hansen, T.B.; Veno, M.T.; Kjems, J. Circular RNAs in cancer: Opportunities and challenges in the field. Oncogene 2018, 37, 555–565. [Google Scholar] [CrossRef] [PubMed]
- De Rie, D.; Abugessaisa, I.; Alam, T.; Arner, E.; Arner, P.; Ashoor, H.; Astrom, G.; Babina, M.; Bertin, N.; Burroughs, A.M.; et al. An integrated expression atlas of miRNAs and their promoters in human and mouse. Nat. Biotechnol. 2017, 35, 872–878. [Google Scholar] [CrossRef] [PubMed]
- Ulitsky, I.; Bartel, D.P. LincRNAs: Genomics, evolution, and mechanisms. Cell 2013, 154, 26–46. [Google Scholar] [CrossRef] [PubMed]
- Brown, B.D.; Gentner, B.; Cantore, A.; Colleoni, S.; Amendola, M.; Zingale, A.; Baccarini, A.; Lazzari, G.; Galli, C.; Naldini, L. Endogenous microRNA can be broadly exploited to regulate transgene expression according to tissue, lineage and differentiation state. Nat. Biotechnol. 2007, 25, 1457–1467. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.S.; Neilson, J.R.; Sharp, P.A. MicroRNA sponges: Competitive inhibitors of small RNAs in mammalian cells. Nat. Methods 2007, 4, 721–726. [Google Scholar] [CrossRef] [PubMed]
- Brown, B.D.; Naldini, L. Exploiting and antagonizing microRNA regulation for therapeutic and experimental applications. Nat. Rev. Genet. 2009, 10, 578–585. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.S.; Sharp, P.A. MicroRNA sponges: Progress and possibilities. RNA 2010, 16, 2043–2050. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; Garcia, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef] [PubMed]
- Seitz, H. Redefining microRNA targets. Curr. Biol. 2009, 19, 870–873. [Google Scholar] [CrossRef] [PubMed]
- Poliseno, L.; Salmena, L.; Zhang, J.; Carver, B.; Haveman, W.J.; Pandolfi, P.P. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 2010, 465, 1033–1038. [Google Scholar] [CrossRef] [PubMed]
- Tay, Y.; Kats, L.; Salmena, L.; Weiss, D.; Tan, S.M.; Ala, U.; Karreth, F.; Poliseno, L.; Provero, P.; Di Cunto, F.; et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 2011, 147, 344–357. [Google Scholar] [CrossRef] [PubMed]
- Karreth, F.A.; Tay, Y.; Perna, D.; Ala, U.; Tan, S.M.; Rust, A.G.; DeNicola, G.; Webster, K.A.; Weiss, D.; Perez-Mancera, P.A.; et al. In vivo identification of tumor-suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell 2011, 147, 382–395. [Google Scholar] [CrossRef] [PubMed]
- Sumazin, P.; Yang, X.; Chiu, H.S.; Chung, W.J.; Iyer, A.; Llobet-Navas, D.; Rajbhandari, P.; Bansal, M.; Guarnieri, P.; Silva, J.; et al. An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Cell 2011, 147, 370–381. [Google Scholar] [CrossRef] [PubMed]
- Karreth, F.A.; Reschke, M.; Ruocco, A.; Ng, C.; Chapuy, B.; Leopold, V.; Sjoberg, M.; Keane, T.M.; Verma, A.; Ala, U.; et al. The BRAF pseudogene functions as a competitive endogenous RNA and induces lymphoma in vivo. Cell 2015, 161, 319–332. [Google Scholar] [CrossRef] [PubMed]
- Smillie, C.L.; Sirey, T.; Ponting, C.P. Complexities of post-transcriptional regulation and the modeling of ceRNA crosstalk. Crit. Rev. Biochem. Mol. Biol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Ala, U.; Karreth, F.A.; Bosia, C.; Pagnani, A.; Taulli, R.; Leopold, V.; Tay, Y.; Provero, P.; Zecchina, R.; Pandolfi, P.P. Integrated transcriptional and competitive endogenous RNA networks are cross-regulated in permissive molecular environments. Proc. Natl. Acad. Sci. USA 2013, 110, 7154–7159. [Google Scholar] [CrossRef] [PubMed]
- Bosson, A.D.; Zamudio, J.R.; Sharp, P.A. Endogenous miRNA and target concentrations determine susceptibility to potential ceRNA competition. Mol. Cell 2014, 56, 347–359. [Google Scholar] [CrossRef] [PubMed]
- Denzler, R.; Agarwal, V.; Stefano, J.; Bartel, D.P.; Stoffel, M. Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol. Cell 2014, 54, 766–776. [Google Scholar] [CrossRef] [PubMed]
- Denzler, R.; McGeary, S.E.; Title, A.C.; Agarwal, V.; Bartel, D.P.; Stoffel, M. Impact of MicroRNA Levels, Target-Site Complementarity, and Cooperativity on Competing Endogenous RNA-Regulated Gene Expression. Mol. Cell 2016, 64, 565–579. [Google Scholar] [CrossRef] [PubMed]
- Powers, J.T.; Tsanov, K.M.; Pearson, D.S.; Roels, F.; Spina, C.S.; Ebright, R.; Seligson, M.; de Soysa, Y.; Cahan, P.; Theissen, J.; et al. Multiple mechanisms disrupt the let-7 microRNA family in neuroblastoma. Nature 2016, 535, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Gilot, D.; Migault, M.; Bachelot, L.; Journe, F.; Rogiers, A.; Donnou-Fournet, E.; Mogha, A.; Mouchet, N.; Pinel-Marie, M.L.; Mari, B.; et al. A non-coding function of TYRP1 mRNA promotes melanoma growth. Nat. Cell Biol. 2017, 19, 1348–1357. [Google Scholar] [CrossRef] [PubMed]
- Brannan, C.I.; Dees, E.C.; Ingram, R.S.; Tilghman, S.M. The product of the H19 gene may function as an RNA. Mol. Cell Biol. 1990, 10, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Bartolomei, M.S.; Zemel, S.; Tilghman, S.M. Parental imprinting of the mouse H19 gene. Nature 1991, 351, 153–155. [Google Scholar] [CrossRef] [PubMed]
- Barlow, D.P.; Stoger, R.; Herrmann, B.G.; Saito, K.; Schweifer, N. The mouse insulin-like growth factor type-2 receptor is imprinted and closely linked to the Tme locus. Nature 1991, 349, 84–87. [Google Scholar] [CrossRef] [PubMed]
- Borsani, G.; Tonlorenzi, R.; Simmler, M.C.; Dandolo, L.; Arnaud, D.; Capra, V.; Grompe, M.; Pizzuti, A.; Muzny, D.; Lawrence, C.; et al. Characterization of a murine gene expressed from the inactive X chromosome. Nature 1991, 351, 325–329. [Google Scholar] [CrossRef] [PubMed]
- Brown, C.J.; Ballabio, A.; Rupert, J.L.; Lafreniere, R.G.; Grompe, M.; Tonlorenzi, R.; Willard, H.F. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature 1991, 349, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Brockdorff, N.; Ashworth, A.; Kay, G.F.; Cooper, P.; Smith, S.; McCabe, V.M.; Norris, D.P.; Penny, G.D.; Patel, D.; Rastan, S. Conservation of position and exclusive expression of mouse Xist from the inactive X chromosome. Nature 1991, 351, 329–331. [Google Scholar] [CrossRef] [PubMed]
- Kopp, F.; Mendell, J.T. Functional Classification and Experimental Dissection of Long Noncoding RNAs. Cell 2018, 172, 393–407. [Google Scholar] [CrossRef] [PubMed]
- Rachmilewitz, J.; Goshen, R.; Ariel, I.; Schneider, T.; de Groot, N.; Hochberg, A. Parental imprinting of the human H19 gene. FEBS Lett. 1992, 309, 25–28. [Google Scholar] [CrossRef]
- Park, I.Y.; Sohn, B.H.; Choo, J.H.; Joe, C.O.; Seong, J.K.; Lee, Y.I.; Chung, J.H. Deregulation of DNA methyltransferases and loss of parental methylation at the insulin-like growth factor II (Igf2)/H19 loci in p53 knockout mice prior to tumor development. J. Cell. Biochem. 2005, 94, 585–596. [Google Scholar] [CrossRef] [PubMed]
- Shoshani, O.; Massalha, H.; Shani, N.; Kagan, S.; Ravid, O.; Madar, S.; Trakhtenbrot, L.; Leshkowitz, D.; Rechavi, G.; Zipori, D. Polyploidization of murine mesenchymal cells is associated with suppression of the long noncoding RNA H19 and reduced tumorigenicity. Cancer Res. 2012, 72, 6403–6413. [Google Scholar] [CrossRef] [PubMed]
- Raveh, E.; Matouk, I.J.; Gilon, M.; Hochberg, A. The H19 Long non-coding RNA in cancer initiation, progression and metastasis—A proposed unifying theory. Mol. Cancer 2015, 14, 184. [Google Scholar] [CrossRef] [PubMed]
- Lv, M.; Zhong, Z.; Huang, M.; Tian, Q.; Jiang, R.; Chen, J. lncRNA H19 regulates epithelial-mesenchymal transition and metastasis of bladder cancer by miR-29b-3p as competing endogenous RNA. Biochim. Biophys. Acta 2017, 1864, 1887–1899. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Ye, X.L.; Xu, J.; Cao, M.G.; Fang, Z.Y.; Li, L.Y.; Guan, G.H.; Liu, Q.; Qian, Y.H.; Xie, D. The lncRNA H19 mediates breast cancer cell plasticity during EMT and MET plasticity by differentially sponging miR-200b/c and let-7b. Sci. Signal. 2017, 10. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Nong, K.; Zhu, H.; Wang, W.; Huang, X.; Yuan, Z.; Ai, K. H19 promotes pancreatic cancer metastasis by derepressing let-7’s suppression on its target HMGA2-mediated EMT. Tumour Biol. 2014, 35, 9163–9169. [Google Scholar] [CrossRef] [PubMed]
- Kou, N.; Liu, S.; Li, X.; Li, W.; Zhong, W.; Gui, L.; Chai, S.; Ren, X.; Na, R.; Zeng, T.; et al. H19 facilitates tongue squamous cell carcinoma migration and invasion via sponging miR-let-7. Oncol. Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.C.; Fu, W.M.; Wong, C.W.; Wang, Y.; Wang, W.M.; Hu, G.X.; Zhang, L.; Xiao, L.J.; Wan, D.C.; Zhang, J.F.; et al. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget 2015, 6, 22513–22525. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Li, X.; Li, X.; Li, X.; Chen, Z. LncRNA H19 promotes epithelial-mesenchymal transition (EMT) by targeting miR-484 in human lung cancer cells. J. Cell. Biochem. 2018, 119, 4447–4457. [Google Scholar] [CrossRef] [PubMed]
- Kallen, A.N.; Zhou, X.B.; Xu, J.; Qiao, C.; Ma, J.; Yan, L.; Lu, L.; Liu, C.; Yi, J.S.; Zhang, H.; et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol. Cell 2013, 52, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Peng, F.; Li, T.T.; Wang, K.L.; Xiao, G.Q.; Wang, J.H.; Zhao, H.D.; Kang, Z.J.; Fan, W.J.; Zhu, L.L.; Li, M.; et al. H19/let-7/LIN28 reciprocal negative regulatory circuit promotes breast cancer stem cell maintenance. Cell Death Dis. 2017, 8, e2569. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Li, Z. Long non-coding RNA growth arrest-specific transcript 5 in tumor biology. Oncol. Lett. 2015, 10, 1953–1958. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.F.; Ye, Y.; Zhao, S.J. LncRNA Gas5 acts as a ceRNA to regulate PTEN expression by sponging miR-222-3p in papillary thyroid carcinoma. Oncotarget 2018, 9, 3519–3530. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Gu, J.; Lu, H. The GAS5/miR-222 Axis Regulates Proliferation of Gastric Cancer Cells through the PTEN/Akt/mTOR Pathway. Dig. Dis. Sci. 2017, 62, 3426–3437. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Song, W.Q.; Sun, P.; Jin, L.; Dai, H.Y. LncRNA-GAS5 induces PTEN expression through inhibiting miR-103 in endometrial cancer cells. J. Biomed. Sci. 2015, 22, 100. [Google Scholar] [CrossRef] [PubMed]
- Wen, Q.; Liu, Y.; Lyu, H.; Xu, X.; Wu, Q.; Liu, N.; Yin, Q.; Li, J.; Sheng, X. Long Noncoding RNA GAS5, Which Acts as a Tumor Suppressor via microRNA 21, Regulates Cisplatin Resistance Expression in Cervical Cancer. Int. J. Gynecol. Cancer 2017, 27, 1096–1108. [Google Scholar] [CrossRef] [PubMed]
- Cao, L.; Chen, J.; Ou, B.; Liu, C.; Zou, Y.; Chen, Q. GAS5 knockdown reduces the chemo-sensitivity of non-small cell lung cancer (NSCLC) cell to cisplatin (DDP) through regulating miR-21/PTEN axis. Biomed. Pharmacother. 2017, 93, 570–579. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhao, W.; Lu, Z.; Zhang, W.; Yang, X. Long non-coding RNA GAS5 promotes proliferation, migration and invasion by regulation of miR-301a in esophageal cancer. Oncol. Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Loewer, S.; Cabili, M.N.; Guttman, M.; Loh, Y.H.; Thomas, K.; Park, I.H.; Garber, M.; Curran, M.; Onder, T.; Agarwal, S.; et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat. Genet. 2010, 42, 1113–1117. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xu, Z.; Jiang, J.; Xu, C.; Kang, J.; Xiao, L.; Wu, M.; Xiong, J.; Guo, X.; Liu, H. Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev. Cell 2013, 25, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Fu, Z.; Li, G.; Li, Z.; Wang, Y.; Zhao, Y.; Zheng, S.; Ye, H.; Luo, Y.; Zhao, X.; Wei, L.; et al. Endogenous miRNA Sponge LincRNA-ROR promotes proliferation, invasion and stem cell-like phenotype of pancreatic cancer cells. Cell Death Discov. 2017, 3, 17004. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yu, X.; Zhang, Z.; Pang, L.; Xu, J.; Jiang, J.; Liang, W.; Chai, Y.; Hou, J.; Li, F. Linc-ROR promotes esophageal squamous cell carcinoma progression through the derepression of SOX9. J. Exp. Clin. Cancer Res. 2017, 36, 182. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Kopp, F.; Chang, T.C.; Sataluri, A.; Chen, B.; Sivakumar, S.; Yu, H.; Xie, Y.; Mendell, J.T. Noncoding RNA NORAD Regulates Genomic Stability by Sequestering PUMILIO Proteins. Cell 2016, 164, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wang, X.; Wen, C.; Huo, Z.; Wang, W.; Zhan, Q.; Cheng, D.; Chen, H.; Deng, X.; Peng, C.; et al. Long noncoding RNA NORAD, a novel competing endogenous RNA, enhances the hypoxia-induced epithelial-mesenchymal transition to promote metastasis in pancreatic cancer. Mol. Cancer 2017, 16, 169. [Google Scholar] [CrossRef] [PubMed]
- Ng, K.; Pullirsch, D.; Leeb, M.; Wutz, A. Xist and the order of silencing. EMBO Rep. 2007, 8, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, L.K.; Yang, Y.T.; Ma, X.; Han, B.; Wang, Z.S.; Zhao, Q.Y.; Wu, L.Q.; Qu, Z.Q. MicroRNA-92b promotes hepatocellular carcinoma progression by targeting Smad7 and is mediated by long non-coding RNA XIST. Cell Death Dis. 2016, 7, e2203. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.; Chen, B.; Wang, X.; Wu, K.; Sun, Y. Long non-coding RNA XIST regulates PTEN expression by sponging miR-181a and promotes hepatocellular carcinoma progression. BMC Cancer 2017, 17, 248. [Google Scholar] [CrossRef] [PubMed]
- Mo, Y.; Lu, Y.; Wang, P.; Huang, S.; He, L.; Li, D.; Li, F.; Huang, J.; Lin, X.; Li, X.; et al. Long non-coding RNA XIST promotes cell growth by regulating miR-139-5p/PDK1/AKT axis in hepatocellular carcinoma. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.L.; Ju, H.Q.; Lu, Y.X.; Chen, L.Z.; Zeng, Z.L.; Zhang, D.S.; Luo, H.Y.; Wang, F.; Qiu, M.Z.; Wang, D.S.; et al. Long non-coding RNA XIST regulates gastric cancer progression by acting as a molecular sponge of miR-101 to modulate EZH2 expression. J. Exp. Clin. Cancer Res. 2016, 35, 142. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Wang, L.; Li, Y.; Chen, M.; He, W.; Qi, L. The Long Non-Coding RNA XIST Interacted with MiR-124 to Modulate Bladder Cancer Growth, Invasion and Migration by Targeting Androgen Receptor (AR). Cell. Physiol. Biochem. 2017, 43, 405–418. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Liu, Y.; Lu, Y.; Yang, B.; Tang, L. LncRNA XIST Promotes Pancreatic Cancer Proliferation through miR-133a/EGFR. J. Cell. Biochem. 2017, 118, 3349–3358. [Google Scholar] [CrossRef] [PubMed]
- Brockdorff, N. Polycomb complexes in X chromosome inactivation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2017, 372. [Google Scholar] [CrossRef] [PubMed]
- Sunwoo, H.; Dinger, M.E.; Wilusz, J.E.; Amaral, P.P.; Mattick, J.S.; Spector, D.L. MEN epsilon/beta nuclear-retained non-coding RNAs are up-regulated upon muscle differentiation and are essential components of paraspeckles. Genome Res. 2009, 19, 347–359. [Google Scholar] [CrossRef] [PubMed]
- Clemson, C.M.; Hutchinson, J.N.; Sara, S.A.; Ensminger, A.W.; Fox, A.H.; Chess, A.; Lawrence, J.B. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol. Cell 2009, 33, 717–726. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Wu, T.; Zhou, H.; Jin, Q.; He, G.; Yu, H.; Xuan, L.; Wang, X.; Tian, L.; Sun, Y.; et al. Long noncoding RNA NEAT1 promotes laryngeal squamous cell cancer through regulating miR-107/CDK6 pathway. J. Exp. Clin. Cancer Res. 2016, 35, 22. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Xiao, Z.; Du, X.; Huang, L.; Du, G. Silencing of the long non-coding RNA NEAT1 suppresses glioma stem-like properties through modulation of the miR-107/CDK6 pathway. Oncol. Rep. 2017, 37, 555–562. [Google Scholar] [CrossRef] [PubMed]
- Tan, H.Y.; Wang, C.; Liu, G.; Zhou, X. Long noncoding RNA NEAT1-modualted miR-506 regulates gastric cancer development through targeting STAT3. J. Cell. Biochem. 2018. [Google Scholar] [CrossRef] [PubMed]
- Gong, W.; Zheng, J.; Liu, X.; Ma, J.; Liu, Y.; Xue, Y. Knockdown of NEAT1 restrained the malignant progression of glioma stem cells by activating microRNA let-7e. Oncotarget 2016, 7, 62208–62223. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Li, T.; Wei, G.; Liu, L.; Chen, Q.; Xu, L.; Zhang, K.; Zeng, D.; Liao, R. The long non-coding RNA NEAT1 regulates epithelial to mesenchymal transition and radioresistance in through miR-204/ZEB1 axis in nasopharyngeal carcinoma. Tumour Biol. 2016, 37, 11733–11741. [Google Scholar] [CrossRef] [PubMed]
- Han, D.; Wang, J.; Cheng, G. LncRNA NEAT1 enhances the radio-resistance of cervical cancer via miR-193b-3p/CCND1 axis. Oncotarget 2018, 9, 2395–2409. [Google Scholar] [CrossRef] [PubMed]
- Hutchinson, J.N.; Ensminger, A.W.; Clemson, C.M.; Lynch, C.R.; Lawrence, J.B.; Chess, A. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genom. 2007, 8, 39. [Google Scholar] [CrossRef] [PubMed]
- Gutschner, T.; Hammerle, M.; Eissmann, M.; Hsu, J.; Kim, Y.; Hung, G.; Revenko, A.; Arun, G.; Stentrup, M.; Gross, M.; et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013, 73, 1180–1189. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Peng, W.X.; Mo, Y.Y.; Luo, D. MALAT1-mediated tumorigenesis. Front. Biosci. 2017, 22, 66–80. [Google Scholar]
- Jie, Y.; Zhao, H.; Meng, W.-Y.; Wu, Q. LncRNA MALAT1 induces colon cancer development by regulating miR-129-5p/HMGB1 axis. J. Cell. Physiol. 2018. [Google Scholar] [CrossRef]
- Chou, J.; Wang, B.; Zheng, T.; Li, X.; Zheng, L.; Hu, J.; Zhang, Y.; Xing, Y.; Xi, T. MALAT1 induced migration and invasion of human breast cancer cells by competitively binding miR-1 with cdc42. Biochem. Biophys. Res. Commun. 2016, 472, 262–269. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.H.; Zhang, W.J.; Wu, X.C.; Zhang, M.D.; Weng, M.Z.; Zhou, D.; Wang, J.D.; Quan, Z.W. Long non-coding RNA Malat1 promotes gallbladder cancer development by acting as a molecular sponge to regulate miR-206. Oncotarget 2016, 7, 37857–37867. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Mei, Z.; Hu, H.B.; Zhang, X. The lncRNA MALAT1 contributes to non-small cell lung cancer development via modulating miR-124/STAT3 axis. J. Cell. Physiol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.M.; Hu, W.W. Long non-coding RNA MALAT1 promotes oral squamous cell carcinoma development via microRNA-125b/STAT3 axis. J. Cell. Physiol. 2018, 233, 3384–3396. [Google Scholar] [CrossRef] [PubMed]
- Fang, Z.; Chen, W.; Yuan, Z.; Liu, X.; Jiang, H. LncRNA-MALAT1 contributes to the cisplatin-resistance of lung cancer by upregulating MRP1 and MDR1 via STAT3 activation. Biomed. Pharmacother. 2018, 101, 536–542. [Google Scholar] [CrossRef] [PubMed]
- Kwok, Z.H.; Roche, V.; Chew, X.H.; Fadieieva, A.; Tay, Y. A non-canonical tumor suppressive role for the long non-coding RNA MALAT1 in colon and breast cancers. Int. J. Cancer 2018. [Google Scholar] [CrossRef] [PubMed]
- Tseng, Y.Y.; Moriarity, B.S.; Gong, W.; Akiyama, R.; Tiwari, A.; Kawakami, H.; Ronning, P.; Reuland, B.; Guenther, K.; Beadnell, T.C.; et al. PVT1 dependence in cancer with MYC copy-number increase. Nature 2014, 512, 82–86. [Google Scholar] [CrossRef] [PubMed]
- Lan, T.; Yan, X.; Li, Z.; Xu, X.; Mao, Q.; Ma, W.; Hong, Z.; Chen, X.; Yuan, Y. Long non-coding RNA PVT1 serves as a competing endogenous RNA for miR-186-5p to promote the tumorigenesis and metastasis of hepatocellular carcinoma. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.; Liu, H.W.; Chen, J.Q.; Wang, S.H.; Hao, L.Q.; Liu, M.; Wang, B. The long noncoding RNA PVT1 functions as a competing endogenous RNA by sponging miR-186 in gastric cancer. Biomed. Pharmacother. 2017, 88, 302–308. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Wang, P.; Xue, Y.; Qu, C.; Zheng, J.; Liu, X.; Ma, J.; Liu, Y. PVT1 affects growth of glioma microvascular endothelial cells by negatively regulating miR-186. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Chang, Z.; Cui, J.; Song, Y. Long noncoding RNA PVT1 promotes EMT via mediating microRNA-186 targeting of Twist1 in prostate cancer. Gene 2018, 654, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Han, C.; Zhang, Y.; Liu, F. LncRNA PVT1 regulate expression of HIF1alpha via functioning as ceRNA for miR199a5p in nonsmall cell lung cancer under hypoxia. Mol. Med. Rep. 2018, 17, 1105–1110. [Google Scholar] [PubMed]
- Zhou, Q.; Chen, F.; Zhao, J.; Li, B.; Liang, Y.; Pan, W.; Zhang, S.; Wang, X.; Zheng, D. Long non-coding RNA PVT1 promotes osteosarcoma development by acting as a molecular sponge to regulate miR-195. Oncotarget 2016, 7, 82620–82633. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Zhou, H.; Liu, P.; Yan, L.; Yao, W.; Chen, K.; Zeng, J.; Li, H.; Hu, J.; Xu, H.; et al. lncRNA PVT1 and its splicing variant function as competing endogenous RNA to regulate clear cell renal cell carcinoma progression. Oncotarget 2017, 8, 85353–85367. [Google Scholar] [CrossRef] [PubMed]
- Conte, F.; Fiscon, G.; Chiara, M.; Colombo, T.; Farina, L.; Paci, P. Role of the long non-coding RNA PVT1 in the dysregulation of the ceRNA-ceRNA network in human breast cancer. PLoS ONE 2017, 12, e0171661. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Zhang, G.; Liu, J. Long noncoding RNA PVT1 promotes cervical cancer progression through epigenetically silencing miR-200b. APMIS 2016, 124, 649–658. [Google Scholar] [CrossRef] [PubMed]
- Huppi, K.; Volfovsky, N.; Runfola, T.; Jones, T.L.; Mackiewicz, M.; Martin, S.E.; Mushinski, J.F.; Stephens, R.; Caplen, N.J. The identification of microRNAs in a genomically unstable region of human chromosome 8q24. Mol. Cancer Res. 2008, 6, 212–221. [Google Scholar] [CrossRef] [PubMed]
- Cui, M.; You, L.; Ren, X.; Zhao, W.; Liao, Q.; Zhao, Y. Long non-coding RNA PVT1 and cancer. Biochem. Biophys. Res. Commun. 2016, 471, 10–14. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.W.; Sun, M.; Xia, R.; Zhang, E.B.; Liu, X.H.; Zhang, Z.H.; Xu, T.P.; De, W.; Liu, B.R.; Wang, Z.X. LincHOTAIR epigenetically silences miR34a by binding to PRC2 to promote the epithelial-to-mesenchymal transition in human gastric cancer. Cell Death Dis. 2015, 6, e1802. [Google Scholar] [CrossRef] [PubMed]
- Li, C.H.; Xiao, Z.; Tong, J.H.; To, K.F.; Fang, X.; Cheng, A.S.; Chen, Y. EZH2 coupled with HOTAIR to silence MicroRNA-34a by the induction of heterochromatin formation in human pancreatic ductal adenocarcinoma. Int. J. Cancer 2017, 140, 120–129. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Du, P.; Yuan, W.; Du, Z.; Yu, M.; Yu, X.; Hu, T. Long non-coding RNA HOTAIR regulates cyclin J via inhibition of microRNA-205 expression in bladder cancer. Cell Death Dis. 2015, 6, e1907. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; An, Y.; Chen, X.; Sun, D.; Chen, T.; Peng, Y.; Zhu, F.; Jiang, Y.; He, X. Epigenetic inhibition of miR-663b by long non-coding RNA HOTAIR promotes pancreatic cancer cell proliferation via up-regulation of insulin-like growth factor 2. Oncotarget 2016, 7, 86857–86870. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhao, F.; Cui, D.; Jiang, R.; Chen, J.; Huang, Q.; Shi, J. HOXD-AS1/miR-130a sponge regulates glioma development by targeting E2F8. Int. J. Cancer 2018, 142, 2313–2322. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Huo, X.; Yang, X.R.; He, J.; Cheng, L.; Wang, N.; Deng, X.; Jin, H.; Wang, N.; Wang, C.; et al. STAT3-mediated upregulation of lncRNA HOXD-AS1 as a ceRNA facilitates liver cancer metastasis by regulating SOX4. Mol. Cancer 2017, 16, 136. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, W.; Wang, Y.; Wang, S. HOXD-AS1 promotes cell proliferation, migration and invasion through miR-608/FZD4 axis in ovarian cancer. Am. J. Cancer Res. 2018, 8, 170–182. [Google Scholar] [PubMed]
- Zhang, Z.; Harrison, P.M.; Liu, Y.; Gerstein, M. Millions of years of evolution preserved: A comprehensive catalog of the processed pseudogenes in the human genome. Genome Res. 2003, 13, 2541–2558. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Pinto, S.M.; Getnet, D.; Nirujogi, R.S.; Manda, S.S.; Chaerkady, R.; Madugundu, A.K.; Kelkar, D.S.; Isserlin, R.; Jain, S.; et al. A draft map of the human proteome. Nature 2014, 509, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Yao, W.; Gumireddy, K.; Li, A.; Wang, J.; Xiao, W.; Chen, K.; Xiao, H.; Li, H.; Tang, K.; et al. Pseudogene PTENP1 functions as a competing endogenous RNA to suppress clear-cell renal cell carcinoma progression. Mol. Cancer Ther. 2014, 13, 3086–3097. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Ren, W.; Zhang, L.; Li, S.; Kong, X.; Zhang, H.; Dong, J.; Cai, G.; Jin, C.; Zheng, D.; et al. PTENp1, a natural sponge of miR-21, mediates PTEN expression to inhibit the proliferation of oral squamous cell carcinoma. Mol. Carcinog. 2017, 56, 1322–1334. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Guo, Y.; Ma, Z.; Ma, G.; Xue, Q.; Li, F.; Liu, L. Long non-coding RNA PTENP1 functions as a ceRNA to modulate PTEN level by decoying miR-106b and miR-93 in gastric cancer. Oncotarget 2017, 8, 26079–26089. [Google Scholar] [CrossRef] [PubMed]
- Gong, T.; Zheng, S.; Huang, S.; Fu, S.; Zhang, X.; Pan, S.; Yang, T.; Sun, Y.; Wang, Y.; Hui, B.; et al. PTENP1 inhibits the growth of esophageal squamous cell carcinoma by regulating SOCS6 expression and correlates with disease prognosis. Mol. Carcinog. 2017, 56, 2610–2619. [Google Scholar] [CrossRef] [PubMed]
- Rutnam, Z.J.; Du, W.W.; Yang, W.; Yang, X.; Yang, B.B. The pseudogene TUSC2P promotes TUSC2 function by binding multiple microRNAs. Nat. Commun. 2014, 5, 2914. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Ishida, M.; Li, L.; Saito, A.; Kamiya, A.; Hamilton, J.P.; Fu, R.; Olaru, A.V.; An, F.; Popescu, I.; et al. Pseudogene INTS6P1 regulates its cognate gene INTS6 through competitive binding of miR-17-5p in hepatocellular carcinoma. Oncotarget 2015, 6, 5666–5677. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zhu, H.; Wu, X.; Xie, X.; Huang, G.; Xu, X.; Li, S.; Xing, C. Downregulated pseudogene CTNNAP1 promote tumor growth in human cancer by downregulating its cognate gene CTNNA1 expression. Oncotarget 2016, 7, 55518–55528. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Du, W.W.; Li, X.; Yee, A.J.; Yang, B.B. Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene 2016, 35, 3919–3931. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.J.; Kwok, Z.H.; Chew, X.H.; Zhang, B.; Liu, C.; Soong, T.W.; Yang, H.; Tay, Y. A FTH1 gene:pseudogene:microRNA network regulates tumorigenesis in prostate cancer. Nucleic Acids Res. 2018, 46, 1998–2011. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Guo, Z.Y.; Zhang, R.; Xin, B.; Chen, R.; Zhao, J.; Wang, T.; Wen, W.H.; Jia, L.T.; Yao, L.B.; et al. Pseudogene OCT4-pg4 functions as a natural micro RNA sponge to regulate OCT4 expression by competing for miR-145 in hepatocellular carcinoma. Carcinogenesis 2013, 34, 1773–1781. [Google Scholar] [CrossRef] [PubMed]
- Bai, M.; Yuan, M.; Liao, H.; Chen, J.; Xie, B.; Yan, D.; Xi, X.; Xu, X.; Zhang, Z.; Feng, Y. OCT4 pseudogene 5 upregulates OCT4 expression to promote proliferation by competing with miR-145 in endometrial carcinoma. Oncol. Rep. 2015, 33, 1745–1752. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Guo, X.; Que, S.; Yang, X.; Fan, H.; Liu, M.; Li, X.; Tang, H. LncRNA RSU1P2 contributes to tumorigenesis by acting as a ceRNA against let-7a in cervical cancer cells. Oncotarget 2017, 8, 43768–43781. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Li, X.; Gu, Y.; Lv, X.; Xi, T. The 3’UTR of the pseudogene CYP4Z2P promotes tumor angiogenesis in breast cancer by acting as a ceRNA for CYP4Z1. Breast Cancer Res. Treat. 2015, 150, 105–118. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Li, X.; Meng, X.; Chou, J.; Hu, J.; Zhang, F.; Zhang, Z.; Xing, Y.; Liu, Y.; Xi, T. Competing endogenous RNA networks of CYP4Z1 and pseudogene CYP4Z2P confer tamoxifen resistance in breast cancer. Mol. Cell. Endocrinol. 2016, 427, 133–142. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zheng, L.; Xin, Y.; Tan, Z.; Zhang, Y.; Meng, X.; Wang, Z.; Xi, T. The competing endogenous RNA network of CYP4Z1 and pseudogene CYP4Z2P exerts an anti-apoptotic function in breast cancer. FEBS Lett. 2017, 591, 991–1000. [Google Scholar] [CrossRef] [PubMed]
- De Martino, M.; Forzati, F.; Marfella, M.; Pellecchia, S.; Arra, C.; Terracciano, L.; Fusco, A.; Esposito, F. HMGA1P7-pseudogene regulates H19 and Igf2 expression by a competitive endogenous RNA mechanism. Sci. Rep. 2016, 6, 37622. [Google Scholar] [CrossRef] [PubMed]
- De Martino, M.; Palma, G.; Azzariti, A.; Arra, C.; Fusco, A.; Esposito, F. The HMGA1 Pseudogene 7 Induces miR-483 and miR-675 Upregulation by Activating Egr1 through a ceRNA Mechanism. Genes 2017, 8, 330. [Google Scholar] [CrossRef] [PubMed]
- Cocquerelle, C.; Mascrez, B.; Hetuin, D.; Bailleul, B. Mis-splicing yields circular RNA molecules. FASEB J. 1993, 7, 155–160. [Google Scholar] [CrossRef] [PubMed]
- Salzman, J.; Gawad, C.; Wang, P.L.; Lacayo, N.; Brown, P.O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 2012, 7, e30733. [Google Scholar] [CrossRef] [PubMed]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.U.; Agarwal, V.; Guo, H.; Bartel, D.P. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014, 15, 409. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Bao, C.; Guo, W.; Li, S.; Chen, J.; Chen, B.; Luo, Y.; Lyu, D.; Li, Y.; Shi, G.; et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 2016, 7, 11215. [Google Scholar] [CrossRef] [PubMed]
- Jeck, W.R.; Sharpless, N.E. Detecting and characterizing circular RNAs. Nat. Biotechnol. 2014, 32, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.L. The biogenesis and emerging roles of circular RNAs. Nat. Rev. Mol. Cell Biol. 2016, 17, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Kjems, J.; Damgaard, C.K. Circular RNA and miR-7 in cancer. Cancer Res. 2013, 73, 5609–5612. [Google Scholar] [CrossRef] [PubMed]
- Piwecka, M.; Glazar, P.; Hernandez-Miranda, L.R.; Memczak, S.; Wolf, S.A.; Rybak-Wolf, A.; Filipchyk, A.; Klironomos, F.; Cerda Jara, C.A.; Fenske, P.; et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 2017, 357. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Gong, X.; Sun, L.; Zhou, Q.; Lu, B.; Zhu, L. The Circular RNA Cdr1as Act as an Oncogene in Hepatocellular Carcinoma through Targeting miR-7 Expression. PLoS ONE 2016, 11, e0158347. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Ji, M.; He, G.; Yang, L.; Niu, Z.; Jian, M.; Wei, Y.; Ren, L.; Xu, J. Silencing CDR1as inhibits colorectal cancer progression through regulating microRNA-7. Oncotargets Ther. 2017, 10, 2045–2056. [Google Scholar] [CrossRef] [PubMed]
- Su, C.; Han, Y.; Zhang, H.; Li, Y.; Yi, L.; Wang, X.; Zhou, S.; Yu, D.; Song, X.; Xiao, N.; et al. CiRS-7 targeting miR-7 modulates the progression of non-small cell lung cancer in a manner dependent on NF-κB signalling. J. Cell. Mol. Med. 2018. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.; Li, T.; Jiang, Y.; Pan, C.; Ding, Y.; Huang, Z.; Yu, H.; Kong, D. Overexpression of Circular RNA ciRS-7 Abrogates the Tumor Suppressive Effect of miR-7 on Gastric Cancer via PTEN/PI3K/AKT Signaling Pathway. J. Cell. Biochem. 2018, 119, 440–446. [Google Scholar] [CrossRef] [PubMed]
- Bernassola, F.; Karin, M.; Ciechanover, A.; Melino, G. The HECT family of E3 ubiquitin ligases: Multiple players in cancer development. Cancer Cell 2008, 14, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Zhang, L.; Li, W.; Deng, J.; Zheng, J.; An, M.; Lu, J.; Zhou, Y. Circular RNA ITCH has inhibitory effect on ESCC by suppressing the Wnt/beta-catenin pathway. Oncotarget 2015, 6, 6001–6013. [Google Scholar] [PubMed]
- Wan, L.; Zhang, L.; Fan, K.; Cheng, Z.X.; Sun, Q.C.; Wang, J.J. Circular RNA-ITCH Suppresses Lung Cancer Proliferation via Inhibiting the Wnt/beta-Catenin Pathway. Biomed. Res. Int. 2016, 2016, 1579490. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Yuan, W.; Yang, X.; Li, P.; Wang, J.; Han, J.; Tao, J.; Li, P.; Yang, H.; Lv, Q.; et al. Circular RNA circ-ITCH inhibits bladder cancer progression by sponging miR-17/miR-224 and regulating p21, PTEN expression. Mol. Cancer 2018, 17, 19. [Google Scholar] [CrossRef] [PubMed]
- Zeng, K.; Chen, X.; Xu, M.; Liu, X.; Hu, X.; Xu, T.; Sun, H.; Pan, Y.; He, B.; Wang, S. CircHIPK3 promotes colorectal cancer growth and metastasis by sponging miR-7. Cell Death Dis. 2018, 9, 417. [Google Scholar] [CrossRef] [PubMed]
- Panda, A.C.; Grammatikakis, I.; Kim, K.M.; De, S.; Martindale, J.L.; Munk, R.; Yang, X.; Abdelmohsen, K.; Gorospe, M. Identification of senescence-associated circular RNAs (SAC-RNAs) reveals senescence suppressor CircPVT1. Nucleic Acids Res. 2017, 45, 4021–4035. [Google Scholar] [CrossRef] [PubMed]
- Verduci, L.; Ferraiuolo, M.; Sacconi, A.; Ganci, F.; Vitale, J.; Colombo, T.; Paci, P.; Strano, S.; Macino, G.; Rajewsky, N.; et al. The oncogenic role of circPVT1 in head and neck squamous cell carcinoma is mediated through the mutant p53/YAP/TEAD transcription-competent complex. Genome Biol. 2017, 18, 237. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Z.; Huang, M.; Lv, M.; He, Y.; Duan, C.; Zhang, L.; Chen, J. Circular RNA MYLK as a competing endogenous RNA promotes bladder cancer progression through modulating VEGFA/VEGFR2 signaling pathway. Cancer Lett. 2017, 403, 305–317. [Google Scholar] [CrossRef] [PubMed]
- Han, D.; Li, J.; Wang, H.; Su, X.; Hou, J.; Gu, Y.; Qian, C.; Lin, Y.; Liu, X.; Huang, M.; et al. Circular RNA circMTO1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression. Hepatology 2017, 66, 1151–1164. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, K.Y.; Lin, Y.C.; Gupta, S.K.; Chang, N.; Yen, L.; Sun, H.S.; Tsai, S.J. Noncoding Effects of Circular RNA CCDC66 Promote Colon Cancer Growth and Metastasis. Cancer Res. 2017, 77, 2339–2350. [Google Scholar] [CrossRef] [PubMed]
- Leung, A.K.L. The Whereabouts of microRNA Actions: Cytoplasm and Beyond. Trends Cell Biol. 2015, 25, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G.; et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Yi, F.; Han, X.; Du, Q.; Liang, Z. MALAT-1 interacts with hnRNP C in cell cycle regulation. FEBS Lett. 2013, 587, 3175–3181. [Google Scholar] [CrossRef] [PubMed]
- Nishizawa, Y.; Konno, M.; Asai, A.; Koseki, J.; Kawamoto, K.; Miyoshi, N.; Takahashi, H.; Nishida, N.; Haraguchi, N.; Sakai, D.; et al. Hypoxia stimulates the cytoplasmic localization of oncogenic long noncoding RNA LINC00152 in colorectal cancer. Int. J. Oncol. 2018, 52, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Kino, T.; Hurt, D.E.; Ichijo, T.; Nader, N.; Chrousos, G.P. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci. Signal. 2010, 3. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Lei, C.; He, Q.; Pan, Z.; Xiao, D.; Tao, Y. Nuclear functions of mammalian MicroRNAs in gene regulation, immunity and cancer. Mol. Cancer 2018, 17, 64. [Google Scholar] [CrossRef] [PubMed]
- Portnoy, V.; Lin, S.H.; Li, K.H.; Burlingame, A.; Hu, Z.H.; Li, H.; Li, L.C. saRNA-guided Ago2 targets the RITA complex to promoters to stimulate transcription. Cell Res. 2016, 26, 320–335. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, J.C.; Younger, S.T.; Nguyen, N.B.; Hardy, D.B.; Monia, B.P.; Corey, D.R.; Janowski, B.A. Antisense transcripts are targets for activating small RNAs. Nat. Struct. Mol. Biol. 2008, 15, 842–848. [Google Scholar] [CrossRef] [PubMed]
- Leucci, E.; Patella, F.; Waage, J.; Holmstrom, K.; Lindow, M.; Porse, B.; Kauppinen, S.; Lund, A.H. microRNA-9 targets the long non-coding RNA MALAT1 for degradation in the nucleus. Sci. Rep. 2013, 3, 2535. [Google Scholar] [CrossRef] [PubMed]
- Li, C.Y.; Liang, G.Y.; Yao, W.Z.; Sui, J.; Shen, X.; Zhang, Y.Q.; Peng, H.; Hong, W.W.; Ye, Y.C.; Zhang, Z.Y.; et al. Integrated analysis of long non-coding RNA competing interactions reveals the potential role in progression of human gastric cancer. Int. J. Oncol. 2016, 48, 1965–1976. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Liu, B. Identification of potential prognostic ceRNA module biomarkers in patients with pancreatic adenocarcinoma. Oncotarget 2017, 8, 94493–94504. [Google Scholar] [CrossRef] [PubMed]
- Yao, K.; Wang, Q.; Jia, J.; Zhao, H. A competing endogenous RNA network identifies novel mRNA, miRNA and lncRNA markers for the prognosis of diabetic pancreatic cancer. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Huang, C.; Li, Q.; Wu, X. Construction and Comprehensive Analysis for Dysregulated Long Non-Coding RNA (lncRNA)-Associated Competing Endogenous RNA (ceRNA) Network in Gastric Cancer. Med. Sci. Monit. 2018, 24, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Wei, Y.; Zhu, Y.; Li, K.; Zhu, Y.; Zhao, Y.; Chang, Z.; Xu, Y. Identification of cancer-related potential biomarkers based on lncRNA-pseudogene-mRNA competitive networks. FEBS Lett. 2018, 592, 973–986. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, X.; Zhou, D.; Zhi, H.; Wang, P.; Gao, Y.; Guo, M.; Yue, M.; Wang, Y.; Shen, W.; et al. Inferences of individual drug responses across diverse cancer types using a novel competing endogenous RNA network. Mol. Oncol. 2018. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, M.R.; Oakes, B.L.; Sternberg, S.H.; East-Seletsky, A.; Kaplan, M.; Doudna, J.A. Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 2014, 516, 263–266. [Google Scholar] [CrossRef] [PubMed]
- Nelles, D.A.; Fang, M.Y.; O’Connell, M.R.; Xu, J.L.; Markmiller, S.J.; Doudna, J.A.; Yeo, G.W. Programmable RNA Tracking in Live Cells with CRISPR/Cas9. Cell 2016, 165, 488–496. [Google Scholar] [CrossRef] [PubMed]
- Abudayyeh, O.O.; Gootenberg, J.S.; Essletzbichler, P.; Han, S.; Joung, J.; Belanto, J.J.; Verdine, V.; Cox, D.B.T.; Kellner, M.J.; Regev, A.; et al. RNA targeting with CRISPR-Cas13. Nature 2017, 550, 280–284. [Google Scholar] [CrossRef] [PubMed]
- Cox, D.B.T.; Gootenberg, J.S.; Abudayyeh, O.O.; Franklin, B.; Kellner, M.J.; Joung, J.; Zhang, F. RNA editing with CRISPR-Cas13. Science 2017, 358, 1019–1027. [Google Scholar] [CrossRef] [PubMed]
| Noncoding RNA Species | Competing Endogenous RNAs | Shared miRNAs | Cancer Type | References | |
|---|---|---|---|---|---|
| LncRNA | H19 | DNMT3B | miR-29b-3p | Bladder | [54] |
| GIT2, CYTH3 | miR-200b, miR-200c, let-7b | Breast | [55] | ||
| HMGA2 | let-7 | Pancreas, tongue | [56,57] | ||
| VIM, ZEB1, ZEB2 | miR-138, miR-200a | Colon | [58] | ||
| ROCK2 | miR-484 | Lung | [59] | ||
| LIN28 | let-7 family | Breast | [60,61] | ||
| GAS5 | PTEN | miR-21, miR-222 | Thyroid, gastric, endometrial, cervical, lung | [63,64,65,66,67] | |
| CXCR4 | miR-301 | Esophageal | [68] | ||
| Linc-ROR | SOX9 | let-7 family, miR-93, miR-145, miR-320a, miR-320b | Pancreas | [71] | |
| miR-15b, miR-33a, miR-129, miR-145, and miR-206 | Esophageal | [72] | |||
| NORAD | RHOA | miR-125a-3p | Pancreas | [74] | |
| XIST | SMAD7 | miR-92b | Liver | [76] | |
| PTEN | miR-181a | Liver | [77] | ||
| PDK1 | miR-139-5p | Liver | [78] | ||
| EZH2 | miR-101 | Gastric | [79] | ||
| AR | miR-124 | Bladder | [80] | ||
| EGFR | miR-133a | Pancreas | [81] | ||
| NEAT1 | CDK6 | miR-107 | Laryngeal, glioma | [85,86] | |
| STAT3 | miR-506, let-7e | Gastric, glioma | [87,88] | ||
| ZEB1 | miR-204 | Nasopharyngeal | [89] | ||
| CCND1 | miR-193-3p | Cervical | [90] | ||
| MALAT1 | HMGB1 | miR-129-5p | Colon | [94] | |
| CDC42 | miR-1 | Breast | [95] | ||
| miR-206 | Gallbladder | [96] | |||
| STAT3 | miR-124, miR-125b | Lung, oral | [97,98] | ||
| PTEN | miR-17, miR-20a, miR-106b | Colon | [100] | ||
| PVT1 | YAP1, HIF1-α | miR-186-5p | Liver, gastric | [102,103] | |
| ATG7, BECN1 | miR-186 | Glioma | [104] | ||
| TWIST1 | miR-186 | Prostate | [105] | ||
| HIF1-α | miR-199a-5p | Lung | [106] | ||
| BCL2, CCND1, FASN | miR-195 | Osteosarcoma | [107] | ||
| miR-200 family | Renal, breast | [108,109] | |||
| HOTAIR | c-Met, SNAIL | miR-34a | Gastric, pancreas | [115,116] | |
| CCNJ | miR-205 | Bladder | [117] | ||
| IGF2 | miR-663b | Pancreas | [118] | ||
| HOXD-AS1 | E2F8, SOX4 | miR-130a | Glioma, liver | [119,120] | |
| FZD4 | miR-608 | Ovarian | [121] | ||
| Pseudogene | PTENP1 | PTEN | miR-19b, miR-20a, miR-21, miR-26a, miR-214, miR-93, miR-106b | Prostate, renal, oral, gastric | [31,124,125,126] |
| SOCS6 | miR-17-5p | Esophageal | [127] | ||
| TUSC2P | TUSC2, TIMP2, TIMP3 | miR-17, miR-93, miR-299-3p, miR-520a, miR-608, miR-661 | Breast, prostate | [128] | |
| INTS6P1 | INTS6 | miR-17-5p | Liver | [129] | |
| CTNNAP1 | CTNNA | miR-141 | Colon | [130] | |
| FOXO3P | FOXO3, circ-FOXO3 | miR-22, miR-136*, miR-138, miR-149*, miR-433, miR-762, miR-3614-5p, miR-3622b-5p | Breast | [131] | |
| FTH1P11, FTH1P16 | FTH1 | miR-19b, miR-181a, miR-210, miR-362, miR-616, miR-638 | Prostate | [132] | |
| BRAFP1 | BRAF | miR-30a, miR-182, miR-134, miR-543, miR-653, miR-876 | Melanoma | [35] | |
| KRASP1 | KRAS | let-7 family | Prostate | [31] | |
| OCT4-pg1, OCT4-pg3, OCT4-pg4, OCT4-pg5 | OCT4/POU5F1 | miR-145 | Liver, endometrial | [31,133,134] | |
| RSU1P2 | IGF1R, MYCN, EPHA4 | let-7a | Cervical | [135] | |
| CYP4Z2P | CYP4Z1, CDK3, hTERT | miR-125a, miR-197, miR-204, miR-1226 | Breast | [136,137,138] | |
| HMGA1P7 | HMGA1, H19, IGF2 | miR-15, miR-16, miR-214, miR-761 | Breast | [139,140] | |
| CircRNA | CDR1as | CCNE1, PIK3CD, EGFR, IGF1R, RELA | miR-7 | Liver, colon, lung, gastric | [144,149,151,152,153,154] |
| circ-ITCH | ITCH | miR-7, miR-17, miR-214 | Lung, esophageal | [156,157] | |
| p21, PTEN | miR-7, miR-224 | Bladder | [158] | ||
| circ-HIPK3 | FAK, IGF1R, EGFR, YY1 | miR-124 | Colon | [159] | |
| circ-PVT1 | IGF2BP1, KRAS, HMGA2 | let-7 | Breast, lung | [160] | |
| AURKA, MKI67, BUB1 | miR-497-5p | Head and neck | [161] | ||
| circ-MYLK | VEGFA, VEGFR2 | miR-29a | Bladder | [162] | |
| circ-MTO1 | p21 | miR-9 | Liver | [163] | |
| circ-CCDC66 | MYC | miR-93 | Colon | [164] | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chan, J.J.; Tay, Y. Noncoding RNA:RNA Regulatory Networks in Cancer. Int. J. Mol. Sci. 2018, 19, 1310. https://doi.org/10.3390/ijms19051310
Chan JJ, Tay Y. Noncoding RNA:RNA Regulatory Networks in Cancer. International Journal of Molecular Sciences. 2018; 19(5):1310. https://doi.org/10.3390/ijms19051310
Chicago/Turabian StyleChan, Jia Jia, and Yvonne Tay. 2018. "Noncoding RNA:RNA Regulatory Networks in Cancer" International Journal of Molecular Sciences 19, no. 5: 1310. https://doi.org/10.3390/ijms19051310
APA StyleChan, J. J., & Tay, Y. (2018). Noncoding RNA:RNA Regulatory Networks in Cancer. International Journal of Molecular Sciences, 19(5), 1310. https://doi.org/10.3390/ijms19051310