Atomic Force Microscopy Based Tip-Enhanced Raman Spectroscopy in Biology
Abstract
1. Introduction
2. Tip Enhanced Raman Spectroscopy (TERS) Experiment
2.1. Optical Geometries
2.2. Feedback Systems
2.2.1. Atomic Force Microscopy Based Tip Enhanced Raman Spectrum (AFM-TERS)
2.2.2. Scanning Tunneling Microscopy Based Tip Enhanced Raman Spectrum (STM-TERS)
2.2.3. Shear Force Microscopy Based Tip Enhanced Raman Spectrum (SFM-TERS)
2.3. Tip Preparation
2.3.1. Chemical Vapor Deposition
2.3.2. Mirror Reaction
2.4. Gap Mode
3. Applications of AFM-TERS in Biology
3.1. Nucleic Acids
3.2. Proteins
3.3. Pathogens
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Dieluweit, S.; Csiszr, A.; Rubner, W.; Fleischhauer, J.; Houben, S.; Merkel, R. Mechanical properties of bare and protein-coated giant unilamellar phospholipid vesicles. A comparative study of micropipet aspiration and atomic force microscopy. Langmuir 2010, 26, 11041–11049. [Google Scholar] [CrossRef] [PubMed]
- Santos, E.B.; Morris, J.K.; Glynos, E.; Sboros, V.; Koutsos, V. Nanomechanical properties of phospholipid microbubbles. Langmuir 2012, 28, 5753–5760. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sirghi, L. Atomic Force Microscopy Indentation of Living Cells; Formatex: Badajoz, Spain, 2010; pp. 433–440. [Google Scholar]
- Franz, V.; Loi, S.; Mller, H.; Bamberg, E.; Butt, H.J. Tip penetration through lipid bilayers in atomic force microscopy. Colloids Surf. B Biointerfaces 2002, 23, 191–200. [Google Scholar] [CrossRef]
- Gilbert, W. DNA sequencing and gene structure. Biosci. Rep. 1981, 1, 353–375. [Google Scholar] [CrossRef] [PubMed]
- Sanger, F.; Nicklen, S.; Coulson, A.R. DNA sequencing with chain-terminating inhibitors 1977. Biotechnology 1992, 24, 104–108. [Google Scholar] [PubMed]
- Gilbert, W. A new method for sequencing DNA. Proc. Natl. Acad. Sci. USA 1977, 74, 5463–5467. [Google Scholar]
- Lindsay, S.M.; Nagahara, L.A.; Thundat, T.; Oden, P. Sequence, packing and nanometer scale structure in STM images of nucleic acids under water. J. Biomol. Struct. Dyn. 1989, 7, 289–299. [Google Scholar] [CrossRef] [PubMed]
- Gutteridge, A.; Thornton, J.M. Understanding nature’s catalytic toolkit. Trends Biochem. Sci. 2005, 30, 622–629. [Google Scholar] [CrossRef] [PubMed]
- Copland, J.A.; Sheffield-Moore, M.; Kodzic-Zivanovic, N.; Gentry, S.; Lamprou, G.; Tzortzatou-Stathopoulou, F.; Zoumpourlis, V.; Urban, R.J.; Vlahopoulos, S.A. Sex steroid receptors in skeletal differentiation and epithelial neoplasia: Is tissue-specific intervention possible. Bioessays 2009, 31, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Samarin, S.; Nusrat, A. Regulation of epithelial apical junctional complex by Rho family GTPases. Front. Biosci. 2009, 14, 1129–1142. [Google Scholar] [CrossRef]
- Gonen, T.; Cheng, Y.F.; Sliz, P.; Hiroaki, Y.; Fujiyoshi, Y.; Harrison, S.C.; Walz, T. Lipid-protein interactions in double-layered two-dimensional AQP0 crystals. Nature 2005, 438, 633–638. [Google Scholar] [CrossRef] [PubMed]
- Standley, D.M.; Kinjo, A.R.; Kinoshita, K.; Nakamura, H. Protein structure databases with new web services for structural biology and biomedical research. Brief. Bioinform. 2008, 9, 276–285. [Google Scholar] [CrossRef] [PubMed]
- Walian, P.; Cross, T.A.; Jap, B.K. Structural genomics of membrane proteins. Genome Biol. 2004, 5, 215. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sleator, R.D.; Walsh, P. An overview of in silico protein function prediction. Arch. Microbiol. 2010, 192, 151–155. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Parajo, M.F. Optical antennas focus in on biology. Nat. Photonics 2008, 2, 201–203. [Google Scholar] [CrossRef]
- Huang, B.; Bates, M.; Zhuang, X.W. Super-Resolution Fluorescence Microscopy. Annu. Rev. Biochem. 2009, 78, 993–1016. [Google Scholar] [CrossRef] [PubMed]
- Schmid, T.; Yeo, B.-S.; Leong, G.; Stadler, J.; Zenobi, R. Performing Tip-Enhanced Raman Spectroscopy in Liquids. J. Raman Spectrosc. 2009, 40, 1392–1399. [Google Scholar] [CrossRef]
- Yeo, B.S.; Stadler, J.; Schmid, T.; Zenobi, R.; Zhang, W.H. Tip Enhanced Raman Spectroscopy—Its Status, Challenges and Future Directions. Chem. Phys. Lett. 2009, 472, 1–13. [Google Scholar] [CrossRef]
- Nakata, A.; Nomoto, T.; Toyota, T.; Fujinami, M. Tip-enhanced Raman Spectroscopy of Lipid Bilayers in Water with an Alumina- and Silver-coated Tungsten Tip. Anal. Sci. 2013, 29, 865–869. [Google Scholar] [CrossRef] [PubMed]
- Kasas, S.; Thomson, N.H.; Smith, B.L.; Hansma, H.G.; Zhu, X.; Guthold, M.; Bustamante, C.; Kool, E.T.; Kashlev, M.; Hansma, P.K. Escherichia coli RNA polymerase activity observed using atomic force microscopy. Biochemistry 1997, 36, 461–468. [Google Scholar] [CrossRef] [PubMed]
- Muhammad, S.K.; Noura, S.D.; Ghulam, M.; Darayas, P.; Bakhrom, B.; John, D.W. Electrophysiology of Epithelial Sodium Channel (ENaC) Embedded in Supported Lipid Bilayer Using a Single Nanopore Chip. Langmuir 2017, 33, 13680–13688. [Google Scholar]
- Lipiec, E.; Sekine, R.; Bielecki, J.; Kwiatek, W.M.; Wood, B.R. Molecular Characterization of DNA Double Strand Breaks with Tip-Enhanced Raman Scattering. Angew. Chem. Int. Ed. 2014, 53, 169–172. [Google Scholar] [CrossRef] [PubMed]
- Deckert-Gaudig, T.; Bailo, E.; Deckert, V. Tip-enhanced Raman scattering (TERS) of oxidised glutathione on an ultraflat gold nanoplate. Phys. Chem. Chem. Phys. 2009, 11, 7360–7362. [Google Scholar] [CrossRef] [PubMed]
- Kurouski, D.; Deckert-Gaudig, T.; Deckert, V.; Lednev, I.K. Surface Characterization of Insulin Protofilaments and Fibril Polymorphs Using Tip-Enhanced Raman Spectroscopy (TERS). Biophys. J. 2014, 106, 263–271. [Google Scholar] [CrossRef] [PubMed]
- Wood, B.R.; Bailo, E.; Khiavi, M.A.; Tilley, L.; Deed, S.; Deckert-Gaudig, T.; McNaughton, D.; Deckert, V. Tip-Enhanced Raman Scattering (TERS) from Hemozoin Crystals within a Sectioned Erythrocyte. Nano Lett. 2011, 11, 1868–1873. [Google Scholar] [CrossRef] [PubMed]
- Bohme, R.; Richter, M.; Cialla, D.; Rosch, P.; Deckert, V.; Popp, J. Towards a specific characterisation of components on a cell surface—Combined TERS-investigations of lipids and human cells. J. Raman Spectrosc. 2009, 40, 1452–1457. [Google Scholar] [CrossRef]
- Kumar, N.; Rae, A.; Roy, D. Accurate measurement of enhancement factor in tip-enhanced Raman spectroscopy through elimination of far-field artefacts. Appl. Phys. Lett. 2014, 104, 123106. [Google Scholar] [CrossRef]
- Lerman, G.M.; Levy, U. Effect of radial polarization and apodization on spot size under tight focusing conditions. Opt. Express 2008, 16, 4567–4581. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wu, X.; Wang, R.; Zhang, M. Detection of Carbon Nanotubes Using Tip-Enhanced Raman Spectroscopy. In Electronic Properties of Carbon Nanotubes; Marulanda, J.M., Ed.; InTech: Vukova, Croatia, 2011. [Google Scholar]
- Hayazawa, N.; Inouye, Y.; Sekkat, Z.; Kawata, S. Metallized tip amplification of near-field Raman scattering. Opt. Commun. 2000, 183, 333–336. [Google Scholar] [CrossRef]
- Hayazawa, N. Near-field Raman scattering enhanced by ametallized tip. Chem. Phys. Lett. 2001, 335, 369–374. [Google Scholar] [CrossRef]
- Hayazawa, N. Near-field Raman imaging of organic mole-cules by an apertureless metallic probe scanning optical microscope. J. Chem. Phys. 2002, 117, 1296–1301. [Google Scholar] [CrossRef]
- Mehtani, J.D. Nano-Raman spectroscopy with side-illumination optics. J. Raman Spectrosc. 2005, 36, 1068–1075. [Google Scholar] [CrossRef]
- Saito, Y.; Motohashi, M.; Hayazawa, N.; Iyoki, M.; Kawata, S. Nanoscale characterization of strained silicon by tip-enhanced Raman spectroscope in reflection mode. Appl. Phys. Lett. 2006, 88, 143109. [Google Scholar] [CrossRef]
- Stadler, J.; Schmid, T.; Zenobi, R. Nanoscale chemical imaging using top-illumination tip-enhanced Raman spectroscopy. Nano Lett. 2010, 10, 4514–4520. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, R.D. Compact metal probes: A solution for atomic force microscopy based tip-enhanced Raman spectroscopy. Rev. Sci. Instrum. 2012, 83, 123708. [Google Scholar] [CrossRef] [PubMed]
- Naumenko, D.; Snitka, V.; Serviene, E.; Bruzaite, I.; Snopok, B. In vivo characterization of protein uptake by yeast cell envelope: Single cell AFM imaging and μ-tipenhanced Raman scattering study. Analyst 2013, 138, 5371–5383. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Saito, Y.; Ichimura, T.; Kawata, S.; Verma, P. Far-field free tapping-mode tip-enhanced Raman microscopy. Appl. Phys. Lett. 2013, 102, 123110. [Google Scholar] [CrossRef]
- Anderson, M.S. Locally enhanced Raman spectroscopy with an atomic force microscope. Appl. Phys. Lett. 2000, 76, 3130–3132. [Google Scholar] [CrossRef]
- Rickman, R.H.; Dunstan, P.R. Enhancement of lattice defect signatures in graphene and ultrathin graphite using tip-enhanced Raman spectroscopy. J. Raman Spectrosc. 2014, 45, 15–21. [Google Scholar] [CrossRef]
- Ghislandi, M.; Hoffmann, G.G.; Tkalya, E.; Xue, L.J.; De With, G. Tip-Enhanced Raman Spectroscopy and Mapping of Graphene Sheets. Appl. Spectrosc. Rev. 2012, 47, 371–381. [Google Scholar] [CrossRef]
- Neugebauer, U.; Rosch, P.; Schmitt, M.; Popp, J.; Julien, C.; Rasmussen, A.; Budich, C.; Deckert, V. On the way to nanometer-sized information of the bacterial surface by tip-enhanced Raman spectroscopy. Chemphyschem 2006, 7, 1428–1430. [Google Scholar] [CrossRef] [PubMed]
- Bailo, E.; Deckert, V. Tip-enhanced Raman spectroscopy of single RNA strands: Towards a novel direct-sequencing method. Angew. Chem. Int. Ed. 2008, 47, 1658–1661. [Google Scholar] [CrossRef] [PubMed]
- Michaels, A.M.; Jiang, J.; Brus, L. Ag nanocrystal junctions as the site for surface-enhanced Raman scattering of single Rhodamine 6G molecules. J. Phys. Chem. B 2000, 104, 11965–11971. [Google Scholar] [CrossRef]
- Williams, C.; Roy, D. Fabrication of gold tips suitable for tip-enhanced Raman spectroscopy. J. Vac. Sci. Technol. B 2008, 26, 1761–1764. [Google Scholar] [CrossRef]
- Taguchi, A.; Hayazawa, N.; Furusawa, K.; Ishitobi, H.; Kawata, S. Deep-UV tip-enhanced Raman scattering. J. Raman Spectrosc. 2009, 40, 1324–1330. [Google Scholar] [CrossRef]
- Meng, L.Y.; Huang, T.X.; Wang, X.; Chen, S.; Yang, Z.L.; Ren, B. Gold-coated AFM tips for tip-enhanced Raman spectroscopy: Theoretical calculation and experimental demonstration. Opt. Express 2015, 23, 13804–13813. [Google Scholar] [CrossRef] [PubMed]
- Stockle, R.M.; Suh, Y.D.; Deckert, V.; Zenobi, R. Nanoscale chemical analysis by tip-enhanced Raman spectroscopy. Chem. Phys. Lett. 2000, 318, 131–136. [Google Scholar] [CrossRef]
- Xu, H.; Bjerneld, E.J.; Kall, M.; Borjesson, L. Spectroscopy of Single Hemoglobin Molecules by Surface Enhanced Raman Scattering. Phys. Rev. Lett. 1999, 83, 4357–4360. [Google Scholar] [CrossRef]
- Xu, H.; Aizpurua, J.; Kall, M.; Apell, P. Electromagnetic Contributions to Single-Molecule Sensitivity in Surface-Enhanced Raman Scattering. Phys. Rev. E 2000, 62, 4318–4324. [Google Scholar] [CrossRef]
- Klingsporn, J.M.; Jiang, N.; Pozzi, E.A.; Sonntag, M.D.; Chulhai, D.; Seideman, T.; Jensen, L.; Hersam, M.C.; Van Duyne, R.P. Intramolecular Insight into Adsorbate-Substrate Interactions via Low-Temperature, Ultrahigh-Vacuum Tip-Enhanced Raman Spectrocopy. J. Am. Chem. Soc. 2014, 136, 3881–3887. [Google Scholar] [CrossRef] [PubMed]
- Touzalin, T.; Dauphin, A.L.; Joiret, S.; Lucas, I.T.; Maisonhaute, E. Tip-Enhanced Raman Spectroscopy Imaging of Opaque Samples in Organic Liquid. Phys. Chem. Chem. Phys. 2016, 18, 15510–15513. [Google Scholar] [CrossRef] [PubMed]
- Kurouski, D.; Mattei, M.; Van Duyne, R.P. Probing Redox Reactions at the Nanoscale with Electrochemical Tip-Enhanced Raman Spectroscopy. Nano Lett. 2015, 15, 7956–7962. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Z.C.; Huang, S.C.; Wu, D.Y.; Meng, L.Y.; Li, M.H.; Huang, T.X.; Zhong, J.H.; Wang, X.; Yang, Z.L.; Ren, B. Electrochemical Tip-Enhanced Raman Spectroscopy. J. Am. Chem. Soc. 2015, 137, 11928–11931. [Google Scholar] [CrossRef] [PubMed]
- Sabanes, N.M.; Driessen, L.M.A.; Domke, K.F. Versatile Side-Illumination Geometry for Tip-Enhanced Raman Spectroscopy at Solid/Liquid Interfaces. Anal. Chem. 2016, 88, 7108–7114. [Google Scholar] [CrossRef] [PubMed]
- Hartschuh, A.; Qian, H.; Meixner, A.J.; Anderson, N.; Novotny, L. Nanoscale Optical Imaging of Single-Walled Carbon Nanotubes. J. Lumin. 2006, 119–120, 204–208. [Google Scholar] [CrossRef]
- Hartschuh, A.; Anderson, N.; Novotny, L. Near-Field Raman Spectroscopy using a Sharp Metal Tip. J. Microsc. 2003, 210, 234–240. [Google Scholar] [CrossRef] [PubMed]
- Anger, P.; Bharadwaj, P.; Novotny, L. Enhancement and Quenching of Single-Molecule Fluorescence. Phys. Rev. Lett. 2006, 96, 113002. [Google Scholar] [CrossRef] [PubMed]
- Neacsu, C.C.; Berweger, S.; Raschke, M.B. Tip-enhanced Raman imaging and nanospectroscopy: Sensitivity, symmetry, and selection rules. Nanobiotechnology 2007, 3, 172–196. [Google Scholar] [CrossRef][Green Version]
- Lloyd, J.S.; Williams, A.; Rickman, R.H.; McCowen, A.; Dunstan, P.R. Reproducible electrochemical etching of silver probes with a radius of curvature of 20 nm for tip-enhanced Raman applications. Appl. Phys. Lett. 2011, 99, 143108. [Google Scholar] [CrossRef]
- Yang, Z.; Aizpurua, J.; Xu, H. Electromagnetic field enhancement in TERS configurations. J. Raman Spectrosc. 2009, 40, 1343–1348. [Google Scholar] [CrossRef]
- Festy, F.; Demming, A.; Richards, D. Resonant excitation of tip plasmons for tipenhanced Raman SNOM. Ultramicroscopy 2004, 100, 437–441. [Google Scholar] [CrossRef] [PubMed]
- Yeo, B.S.; Zhang, W.; Vannier, C.; Zenobi, R. Enhancement of Raman signals with silver-coated tips. Appl. Spectrosc. 2006, 60, 1142–1147. [Google Scholar] [CrossRef] [PubMed]
- Kharintsev, S.S.; Hoffmann, G.G.; Dorozhkin, P.S.; de With, G.; Loos, J. Atomic force and shear force based tip-enhanced Raman spectroscopy and imaging. Nanotechnology 2007, 18, 315502. [Google Scholar] [CrossRef]
- Johnson, P.B.; Christy, R.W. Optical constants of the noble metals. Phys. Rev. B 1972, 6, 4370–4379. [Google Scholar] [CrossRef]
- Novotny, L.; Bian, R.; Xie, X. Theory of nanometric optical tweezers. Phys. Rev. Lett. 1997, 79, 645–648. [Google Scholar] [CrossRef]
- Mcmahon, M.D.; Lopez, R.; Meyer, H.M.; Feldman, L.C.; Haglund, R.F. Rapid tarnishing of silver nanoparticles in ambient laboratory air. Appl. Phys. B 2005, 80, 915–921. [Google Scholar] [CrossRef]
- Stockle, R.M.; Deckert, V.; Fokas, C.; Zenobi, R. Controlled formation of isolated silver islands for surface-enhanced Raman scattering. Appl. Spectrosc. 2000, 54, 1577–1583. [Google Scholar] [CrossRef]
- Nieman, L.T.; Krampert, G.M.; Martinez, R.E. An apertureless near-field scanning optical microscope and its application to surface-enhanced Raman spectroscopy and multiphoton fluorescence imaging. Rev. Sci. Instrum. 2001, 72, 1691–1699. [Google Scholar] [CrossRef]
- Saito, Y.; Wang, J.J.; Smith, D.A.; Batchelder, D.N. A simple chemical method for the preparation of silver surfaces for efficient SERS. Langmuir 2002, 18, 2959–2961. [Google Scholar] [CrossRef]
- Wang, J.J.; Saito, Y.; Batchelder, D.N.; Kirkham, J.; Robinson, C.; Smith, D.A. Controllable method for the preparation of metallized probes for efficient scanning near-field optical Raman microscopy. Appl. Phys. Lett. 2005, 86, 263111. [Google Scholar] [CrossRef]
- Vasile, M.J.; Grigg, D.A.; Griffith, J.E.; Fitzgerald, E.A.; Russell, P.E. Scanning probe tips formed by focused ion beams. Rev. Sci. Instrum. 1991, 62, 2167–2171. [Google Scholar] [CrossRef]
- Sánchez, E.; Novotny, L.; Xie, X. Near-field fluorescence microscopy based on two photon excitation with metal tips. Phys. Rev. Lett. 1999, 82, 4014–4017. [Google Scholar] [CrossRef]
- Becker, M.; Sivakov, V.; Gösele, U.; Stelzner, T.; Andrä, G.; Reich, H.J.; Hoffmann, S.; Michler, J.; Christiansen, S.H. Nanowires enabling signal-enhanced nanoscale Raman spectroscopy. Small 2008, 4, 398–404. [Google Scholar] [CrossRef] [PubMed]
- Becker, M.; Sivakov, V.; Andrä, G.; Geiger, R.; Schreiber, J.; Hoffmann, S.; Michler, J.; Milenin, A.P.; Werner, P.; Christiansen, S.H. The SERS and TERS effects obtained by gold droplets on top of Si nanowires. Nano Lett. 2007, 7, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Li, Z.-Y.; Nogami, M.; Tanemura, M.; Huang, Z. The controlled fabrication of “Tip-On-Tip” TERS probes. RSC Adv. 2014, 4, 4718–4722. [Google Scholar] [CrossRef]
- Johnson, T.W.; Lapin, Z.J.; Beams, R.; Lindquist, N.C.; Rodrigo, S.G.; Novotny, L. Highly reproducible near-field optical imaging with Sub-20-nm resolution based on template-stripped gold pyramids. ACS Nano 2012, 6, 9168–9174. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Walke, P.; De Feyter, S.; Hiroshi Uji-i, H. Remote excitation-tip-enhanced Raman scattering microscopy using silver nanowire. Jpn. J. Appl. Phys. 2016, 55, 08NB03. [Google Scholar] [CrossRef]
- Futamata, M.; Maruyama, Y.; Ishikawa, M. Local electric field and scattering cross section of Ag nanoparticles under surface plasmon resonance by finite difference time domain method. J. Phys. Chem. B 2003, 107, 7607–7617. [Google Scholar] [CrossRef]
- Verma, P.; Yamada, K.; Watanabe, H.; Inouye, Y.; Kawata, S. Near-field Raman scattering investigation of tip effects on C60 molecules. Phys. Rev. B 2006, 73, 045416. [Google Scholar] [CrossRef]
- Treffer, R.; Böhme, R.; Deckert-Gaudig, T.; Lau, K.; Tiede, S.; Lin, X.; Deckert, V. Advances in TERS (tip-enhanced Raman scattering) for biochemical applications. Biochem. Soc. Trans. 2012, 40, 609–614. [Google Scholar] [CrossRef] [PubMed]
- Bailo, E.; Deckert, V. Tip-enhanced Raman scattering. Chem. Soc. Rev. 2008, 37, 921–930. [Google Scholar] [CrossRef] [PubMed]
- Deckert-Gaudig, T.; Bailo, E.; Deckert, V. Perspectives for spatially resolved molecular spectroscopy—Raman on the nanometer scale. J. Biophotonics 2008, 1, 377–389. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, A.; Deckert, V. Surface- and Tip-Enhanced Raman Scattering of DNA Components. J. Raman Spectrosc. 2006, 37, 311–317. [Google Scholar] [CrossRef]
- Ichimura, T.; Hayazawa, N.; Hashimoto, M.; Inouye, Y.; Kawata, S. Tip-Enhanced Coherent Anti-Stokes Raman Scattering for Vibrational Nanoimaging. Phys. Rev. Lett. 2004, 92, 220801. [Google Scholar] [CrossRef] [PubMed]
- Domke, K.F.; Zhang, D.; Pettinger, B. Tip-Enhanced Raman Spectra of Picomole Quantities of DNA Nucleobases at Au(111). J. Am. Chem. Soc. 2007, 129, 6708–6709. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Domke, K.F.; Pettinger, B. Tip-Enhanced Raman Spectroscopic Studies of the Hydrogen Bonding between Adenine and Thymine Adsorbed on Au(111). ChemPhysChem 2010, 11, 1662–1665. [Google Scholar] [CrossRef] [PubMed]
- Treffer, R.; Lin, X.; Bailo, E.; Deckert-Gaudig, T.; Deckert, V. Distinction of nucleobases—A tip-enhanced Raman approach. Beilstein J. Nanotechnol. 2011, 2, 628–637. [Google Scholar] [CrossRef] [PubMed]
- Najjar, S.; Talaga, D.; Schue, L.; Coffinier, Y.; Szunerits, S.; Boukherroub, R. Tip-enhanced Raman spectroscopy of combed double-stranded DNA bundles. J. Phys. Chem. C 2014, 118, 1174–1181. [Google Scholar] [CrossRef]
- Yeo, B.S.; Madler, S.; Schmid, T.; Zhang, W.H.; Zenobi, R. Tip-enhanced Raman spectroscopy can see more: The case of cytochrome C. J. Phys. Chem. C 2008, 112, 4867–4873. [Google Scholar] [CrossRef]
- Deckert-Gaudig, T.; Bailo, E.; Deckert, V. Tip-enhanced Raman scattering (TERS) and high-resolution bio nano-analysis—A comparison. Phys. Chem. Chem. Phys. 2010, 114, 7412–7420. [Google Scholar] [CrossRef]
- Moretti, M.; Zaccaria, R.P.; Descrovi, E.; Das, G.; Leoncini, M.; Liberale, C.; De Angelis, F.; Di Fabrizio, E. Reflection-mode TERS on insulin amyloid fibrils with top-visual AFM probes. Plasmonics 2013, 8, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Deckert-Gaudig, T.; Kämmer, E.; Deckert, V. Tracking of nanoscale structural variations on a single amyloid fibril with tip-enhanced Raman scattering. J. Biophotonics 2012, 5, 215–219. [Google Scholar] [CrossRef] [PubMed]
- Paulite, M.; Blum, C.; Schmid, T.; Opilik, L.; Eyer, K.; Walker, G.C.; Zenobi, R. Full spectroscopic tip-enhanced Raman imaging of single nanotapes formed from β-amyloid(1–40) peptide fragments. ACS Nano 2013, 7, 1911–1920. [Google Scholar] [CrossRef] [PubMed]
- Gullekson, C.; Lucas, L.; Hewitt, K.; Kreplak, L. Surface-sensitive Raman spectroscopy of collagen I fibrils. Biophys. J. 2011, 100, 1837–1845. [Google Scholar] [CrossRef] [PubMed]
- Neugebauer, U.; Schmid, U.; Baumann, K.; Ziebuhr, W.; Kozitskaya, S.; Deckert, V.; Schmitt, M.; Popp, J. Towards a detailed understanding of bacterial metabolism—Spectroscopic characterization of Staphylococcus epidermidis. ChemPhysChem 2007, 8, 124–137. [Google Scholar] [CrossRef] [PubMed]
- Cialla, D.; Deckert-Gaudig, T.; Budich, C.; Laue, M.; Moller, R.; Naumann, D.; Deckert, V.; Popp, J. Raman to the limit: Tip-enhanced Raman spectroscopic investigations of a single tobacco mosaic virus. J. Raman Spectrosc. 2009, 40, 240–243. [Google Scholar] [CrossRef]
- Rusciano, G.; Zito, G.; Isticato, R.; Sirec, T.; Ricca, E.; Bailo, E. Nanoscale Chemical Imaging of Bacillus subtilis Spores by Combining Tip-Enhanced Raman Scattering and Advanced Statistical Tools. ACS Nano 2014, 8, 12300–12309. [Google Scholar] [CrossRef] [PubMed]
- Barros, E.; Carvajal, C. Urinary Exosomes and Their Cargo: Potential Biomarkers for Mineralocorticoid Arterial Hypertension. Front. Endocrinol. 2017, 8, 230. [Google Scholar] [CrossRef] [PubMed]
- Nalejska, E.; Mączyńska, E.; Lewandowska, M.A. Prognostic and Predictive Biomarkers: Tools in Personalized Oncology. Mol. Oncol. Gene. 2014, 18, 273–284. [Google Scholar] [CrossRef] [PubMed]
- Aronson, J. Biomarkers and surrogate endpoints. Brit. J. Clin. Pharmacol. 2005, 59, 491–494. [Google Scholar] [CrossRef] [PubMed]
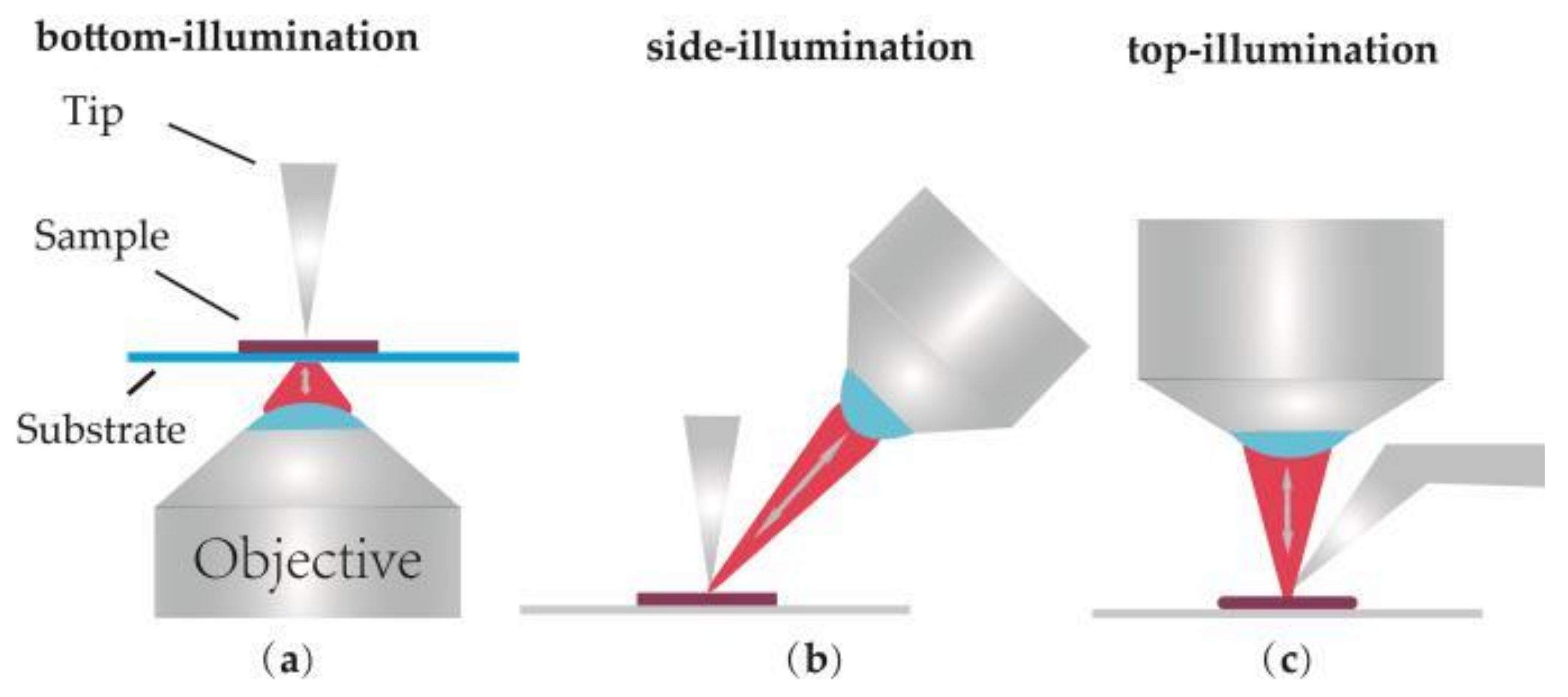
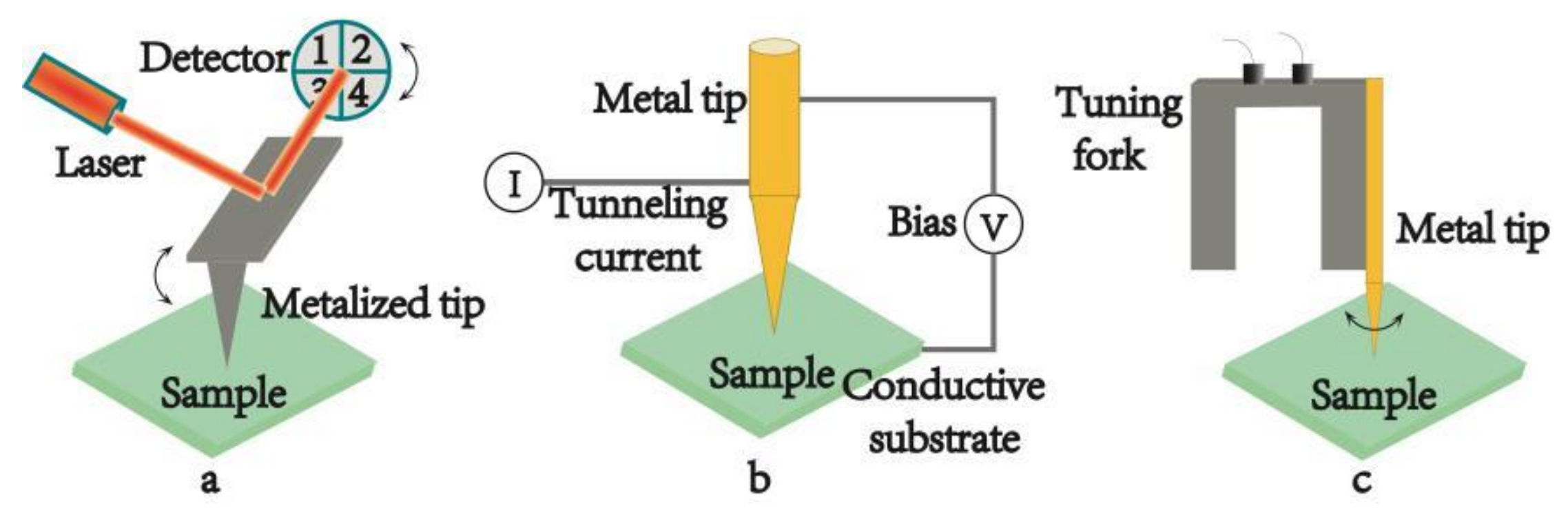
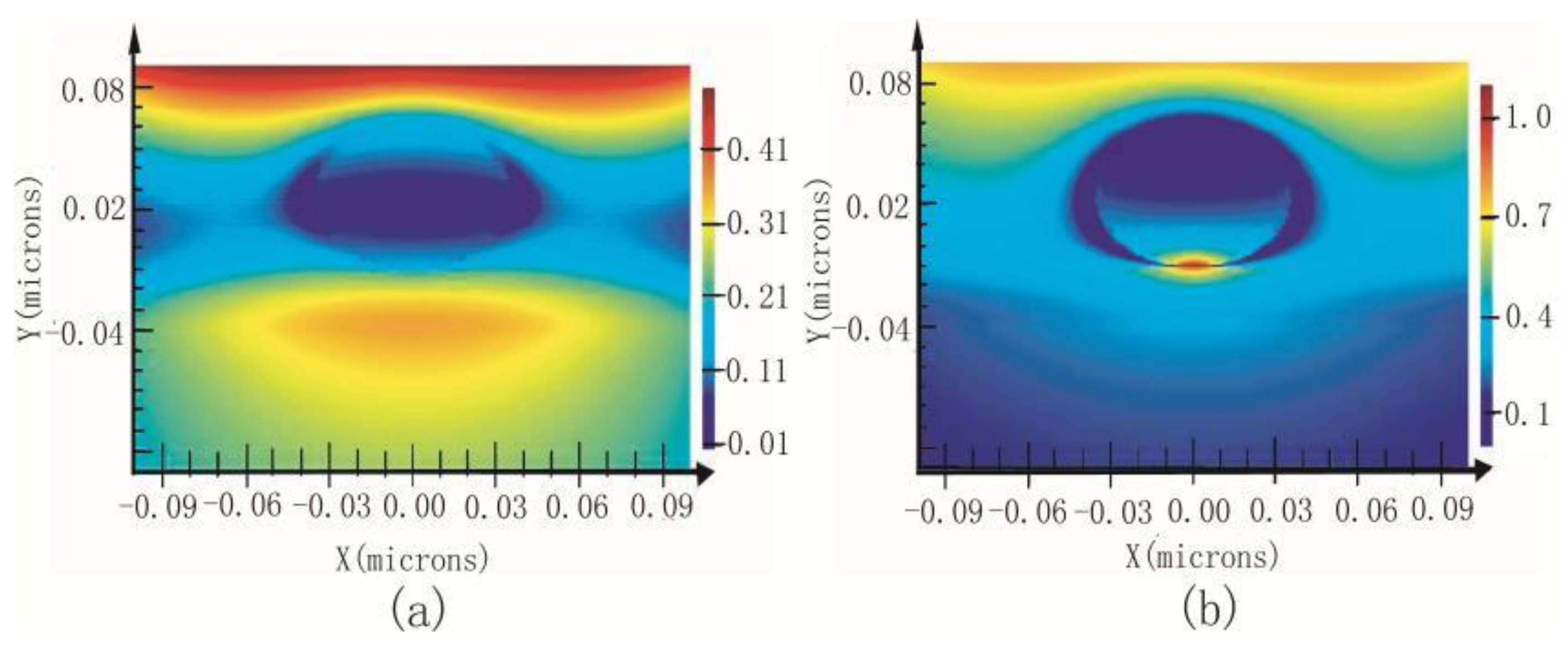
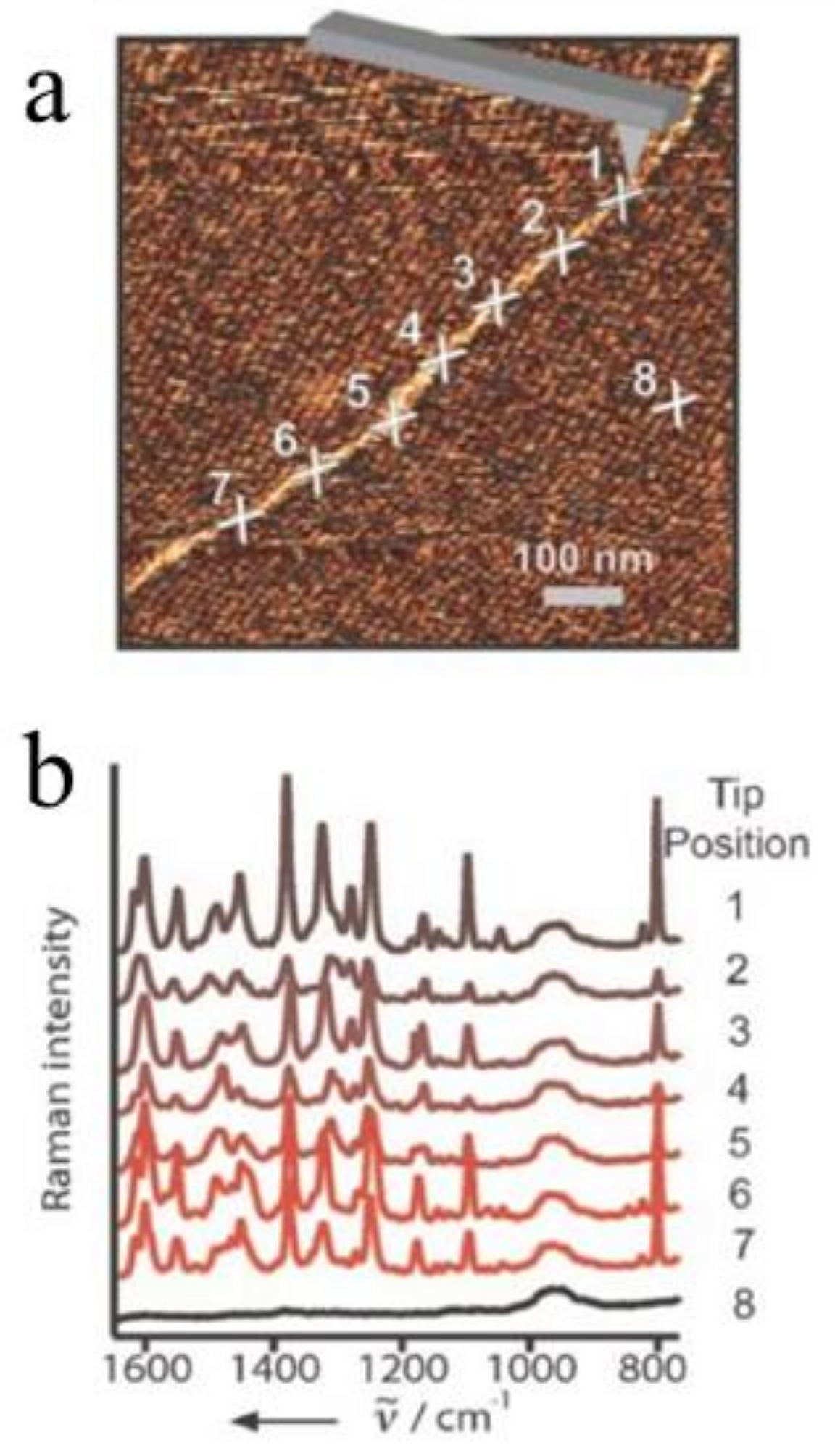
| Optical Geometries | Bottom-Illumination | Side-Illumination | Top-Illumination |
|---|---|---|---|
| Advantage | Efficient enhancement | Wide application | Wide application |
| Disadvantage | Only for transparent simple | Signal loss | top visual cantilevers should be used |
| Feedback System | AFM-TERS | STM-TERS | SFM-TERS |
|---|---|---|---|
| controller | Atomic force | tunneling current | Shear force |
| Tip material | silicon or silicon nitride tip covered with metal | metal | metal |
| optical geometries | reflection mode and transmission mode | reflection mode | reflection mode and transmission mode |
| substrate | all kinds of substrates [39,40,41,42,43,44,45,46] | conductive or semi-conductive substrates | all kinds of substrates |
| environment | Gas [39,40,41,42,43,44,45,46], liquid [49,50,51] | gas, liquid(special) [53,54,55,56], high-vacuum, low-temperature [52] | gas |
| Biomolecule | Nucleic Acid | Protein | Pathogen |
|---|---|---|---|
| Structure | A single- or double-stranded structure consisted of five nucleotides | Three-dimensional structure consisted of one or more long chains of amino acid residues | RNAs and Proteins |
| AFM-TERS test | direct-sequencing; DNA hybridization; DNA double strand breaks [19,44,82,83,84,85,86,87,88,89] | Conformal changes and amino acid distributions [25,90,91,92,93,94,95,96] | Peptides and polysaccharides [43,98,99] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gao, L.; Zhao, H.; Li, T.; Huo, P.; Chen, D.; Liu, B. Atomic Force Microscopy Based Tip-Enhanced Raman Spectroscopy in Biology. Int. J. Mol. Sci. 2018, 19, 1193. https://doi.org/10.3390/ijms19041193
Gao L, Zhao H, Li T, Huo P, Chen D, Liu B. Atomic Force Microscopy Based Tip-Enhanced Raman Spectroscopy in Biology. International Journal of Molecular Sciences. 2018; 19(4):1193. https://doi.org/10.3390/ijms19041193
Chicago/Turabian StyleGao, Lizhen, Huiling Zhao, Tianfeng Li, Peipei Huo, Dong Chen, and Bo Liu. 2018. "Atomic Force Microscopy Based Tip-Enhanced Raman Spectroscopy in Biology" International Journal of Molecular Sciences 19, no. 4: 1193. https://doi.org/10.3390/ijms19041193
APA StyleGao, L., Zhao, H., Li, T., Huo, P., Chen, D., & Liu, B. (2018). Atomic Force Microscopy Based Tip-Enhanced Raman Spectroscopy in Biology. International Journal of Molecular Sciences, 19(4), 1193. https://doi.org/10.3390/ijms19041193




