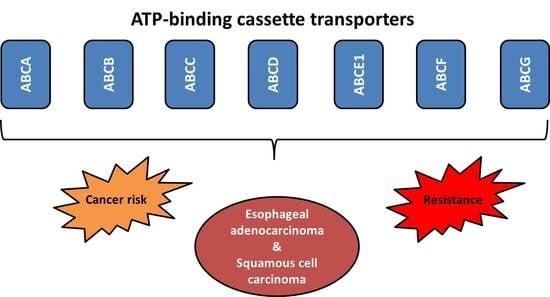
ABC Transporters and Their Role in the Neoadjuvant Treatment of Esophageal Cancer
Abstract
1. Introduction
2. Search Results and Discussion
2.1. Genetic Variation
2.1.1. Single Nucleotide Polymorphisms
2.1.2. Somatic Aberrations
2.1.3. Structural Variations in ABC Transporters Specifically Studied in Esophageal Cancer
2.2. Epigenetic Dysregulation
2.2.1. DNA Methylation
2.2.2. Histone Modifications
2.2.3. MicroRNAs
2.3. Phenotype Changes
2.3.1. Transcript Expression
2.3.2. Protein Expression
2.4. Signaling Pathways
2.5. ABC Transporters-Induced Radiotherapy Resistance
3. Conclusions and Future Prospects
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- SEER Cancer Stat Facts: Esophageal Cancer. Available online: https://seer.cancer.gov/statfacts/html/esoph.html (accessed on 26 January 2018).
- Castro, C.; Peleteiro, B.; Lunet, N. Modifiable factors and esophageal cancer: A systematic review of published meta-analyses. J. Gastroenterol. 2018, 53, 37–51. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.; Dhingra, S. Pathology of esophageal cancer and Barrett’s esophagus. Ann. Cardiothorac. Surg. 2017, 6, 99–109. [Google Scholar] [CrossRef] [PubMed]
- Minsky, B.D.; Pajak, T.F.; Ginsberg, R.J.; Pisansky, T.M.; Martenson, J.; Komaki, R.; Okawara, G.; Rosenthal, S.A.; Kelsen, D.P. INT 0123 (Radiation Therapy Oncology Group 94-05) phase III trial of combined-modality therapy for esophageal cancer: High-dose versus standard-dose radiation therapy. J. Clin. Oncol. 2002, 20, 1167–1174. [Google Scholar] [CrossRef] [PubMed]
- Cooper, J.S.; Guo, M.D.; Herskovic, A.; Macdonald, J.S.; Martenson, J.A., Jr.; Al-Sarraf, M.; Byhardt, R.; Russell, A.H.; Beitler, J.J.; Spencer, S.; et al. Chemoradiotherapy of locally advanced esophageal cancer: Long-term follow-up of a prospective randomized trial (RTOG 85-01). Radiation Therapy Oncology Group. JAMA 1999, 281, 1623–1627. [Google Scholar] [CrossRef] [PubMed]
- Matzenauer, M.; Vrana, D.; Vlachova, Z.; Aujesky, R.; Vrba, R.; Neoral, C.; Melichar, B. Stereotactic radiotherapy in the treatment of local recurrences of esophageal cancer. Oncol. Lett. 2017, 13, 1807–1810. [Google Scholar] [CrossRef] [PubMed]
- Gottesman, M.M.; Fojo, T.; Bates, S.E. Multidrug resistance in cancer: Role of ATP-dependent transporters. Nat. Rev. Cancer 2002, 2, 48–58. [Google Scholar] [CrossRef] [PubMed]
- Hlavata, I.; Mohelnikova-Duchonova, B.; Vaclavikova, R.; Liska, V.; Pitule, P.; Novak, P.; Bruha, J.; Vycital, O.; Holubec, L.; Treska, V.; et al. The role of ABC transporters in progression and clinical outcome of colorectal cancer. Mutagenesis 2012, 27, 187–196. [Google Scholar] [CrossRef] [PubMed]
- Hlavac, V.; Brynychova, V.; Vaclavikova, R.; Ehrlichova, M.; Vrana, D.; Pecha, V.; Kozevnikovova, R.; Trnkova, M.; Gatek, J.; Kopperova, D.; et al. The expression profile of ATP-binding cassette transporter genes in breast carcinoma. Pharmacogenomics 2013, 14, 515–529. [Google Scholar] [CrossRef] [PubMed]
- Mohelnikova-Duchonova, B.; Brynychova, V.; Oliverius, M.; Honsova, E.; Kala, Z.; Muckova, K.; Soucek, P. Differences in transcript levels of ABC transporters between pancreatic adenocarcinoma and nonneoplastic tissues. Pancreas 2013, 42, 707–716. [Google Scholar] [CrossRef] [PubMed]
- Elsnerova, K.; Mohelnikova-Duchonova, B.; Cerovska, E.; Ehrlichova, M.; Gut, I.; Rob, L.; Skapa, P.; Hruda, M.; Bartakova, A.; Bouda, J.; et al. Gene expression of membrane transporters: Importance for prognosis and progression of ovarian carcinoma. Oncol. Rep. 2016, 35, 2159–2170. [Google Scholar] [CrossRef] [PubMed]
- Melichar, B. Laboratory medicine and medical oncology: The tale of two Cinderellas. Clin. Chem. Lab. Med. 2013, 51, 99–112. [Google Scholar] [CrossRef] [PubMed]
- Saier, M.H., Jr.; Tran, C.V.; Barabote, R.D. TCDB: The Transporter Classification Database for membrane transport protein analyses and information. Nucleic Acids Res. 2006, 34, D181–D186. [Google Scholar] [CrossRef] [PubMed]
- Narumiya, K.; Metzger, R.; Bollschweiler, E.; Alakus, H.; Brabender, J.; Drebber, U.; Holscher, A.H.; Warnecke-Eberz, U. Impact of ABCB1 C3435T polymorphism on lymph node regression in multimodality treatment of locally advanced esophageal cancer. Pharmacogenomics 2011, 12, 205–214. [Google Scholar] [CrossRef] [PubMed]
- SNPedia. Available online: https://www.snpedia.com/index.php/SNPedia (accessed on 26 January 2018).
- Komoto, C.; Nakamura, T.; Sakaeda, T.; Kroetz, D.L.; Yamada, T.; Omatsu, H.; Koyama, T.; Okamura, N.; Miki, I.; Tamura, T.; et al. MDR1 haplotype frequencies in Japanese and Caucasian, and in Japanese patients with colorectal cancer and esophageal cancer. Drug Metab. Pharmacokinet. 2006, 21, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Gu, J.; Wu, T.T.; Swisher, S.G.; Liao, Z.; Correa, A.M.; Liu, J.; Etzel, C.J.; Amos, C.I.; Huang, M.; et al. Genetic variations in radiation and chemotherapy drug action pathways predict clinical outcomes in esophageal cancer. J. Clin. Oncol. 2006, 24, 3789–3798. [Google Scholar] [CrossRef] [PubMed]
- Gusella, M.; Giacopuzzi, S.; Bertolaso, L.; Zanoni, A.; Pezzolo, E.; Modena, Y.; Menon, D.; Paganin, P.; Weindelmayer, J.; Crepaldi, G.; et al. Genetic prediction of long-term survival after neoadjuvant chemoradiation in locally advanced esophageal cancer. Pharmacogenomics J. 2017, 17, 252–257. [Google Scholar] [CrossRef] [PubMed]
- Okuno, T.; Tamura, T.; Yamamori, M.; Chayahara, N.; Yamada, T.; Miki, I.; Okamura, N.; Kadowaki, Y.; Shirasaka, D.; Aoyama, N.; et al. Favorable genetic polymorphisms predictive of clinical outcome of chemoradiotherapy for stage II/III esophageal squamous cell carcinoma in Japanese. Am. J. Clin. Oncol. 2007, 30, 252–257. [Google Scholar] [CrossRef] [PubMed]
- Miura, M.; Motoyama, S.; Hinai, Y.; Niioka, T.; Endo, M.; Hayakari, M.; Ogawa, J. Influence of CYP2C19 and ABCB1 polymorphisms on plasma concentrations of lansoprazole enantiomers after enteral administration. Xenobiotica 2010, 40, 630–636. [Google Scholar] [CrossRef] [PubMed]
- Findlay, J.M.; Middleton, M.R.; Tomlinson, I. A systematic review and meta-analysis of somatic and germline DNA sequence biomarkers of esophageal cancer survival, therapy response and stage. Ann. Oncol. 2015, 26, 624–644. [Google Scholar] [CrossRef] [PubMed]
- Zou, N.; Yang, L.; Chen, L.; Li, T.; Jin, T.; Peng, H.; Zhang, S.; Wang, D.; Li, R.; Liu, C.; et al. Heterozygote of TAP1 Codon637 decreases susceptibility to HPV infection but increases susceptibility to esophageal cancer among the Kazakh populations. J. Exp. Clin. Cancer Res. 2015, 34, 70. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Lu, H.; Wang, J.; Chen, Z.; Wang, Y.; Ji, R.; Li, Q.; Zhao, Y.; Zhang, H.; Zhou, Y. Association between TAP1 gene polymorphism and esophageal cancer in a Han Gansu population. Int. J. Clin. Exp. Med. 2016, 9, 12097–12102. [Google Scholar]
- Cao, B.; Tian, X.; Li, Y.; Jiang, P.; Ning, T.; Xing, H.; Zhao, Y.; Zhang, C.; Shi, X.; Chen, D.; et al. LMP7/TAP2 gene polymorphisms and HPV infection in esophageal carcinoma patients from a high incidence area in China. Carcinogenesis 2005, 26, 1280–1284. [Google Scholar] [CrossRef] [PubMed]
- Rumiato, E.; Boldrin, E.; Malacrida, S.; Battaglia, G.; Bocus, P.; Castoro, C.; Cagol, M.; Chiarion-Sileni, V.; Ruol, A.; Amadori, A.; et al. A germline predictive signature of response to platinum chemotherapy in esophageal cancer. Transl. Res. 2016, 171, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Shi, N.; Lu, H.; Zhang, J.; Ma, Y.; Qiao, Y.; Mao, Y.; Jia, K.; Han, L.; Liu, F.; et al. ABCC4 copy number variation is associated with susceptibility to esophageal squamous cell carcinoma. Carcinogenesis 2014, 35, 1941–1950. [Google Scholar] [CrossRef] [PubMed]
- COSMIC COSMIC, the Catalogue of Somatic Mutations in Cancer. Available online: http://cancer.sanger.ac.uk/cosmic (accessed on 26 January 2018).
- Hong, L.; Han, Y.; Zhang, H.; Li, M.; Gong, T.; Sun, L.; Wu, K.; Zhao, Q.; Fan, D. The prognostic and chemotherapeutic value of miR-296 in esophageal squamous cell carcinoma. Ann. Surg. 2010, 251, 1056–1063. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Hong, L. Prediction value of miR-483 and miR-214 in prognosis and multidrug resistance of esophageal squamous cell carcinoma. Genet. Test. Mol. Biomarkers 2013, 17, 470–474. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.J.; Chen, K.N.; Xu, G.W.; Xing, H.P.; Shi, X.T. Congenital expression of mdr-1 gene in tissues of carcinoma and its relation with pathomorphology and prognosis. World J. Gastroenterol. 1999, 5, 53–56. [Google Scholar] [CrossRef] [PubMed]
- Langer, R.; Specht, K.; Becker, K.; Ewald, P.; Ott, K.; Lordick, F.; Siewert, J.R.; Hofler, H. Comparison of pretherapeutic and posttherapeutic expression levels of chemotherapy-associated genes in adenocarcinomas of the esophagus treated by 5-fluorouracil- and cisplatin-based neoadjuvant chemotherapy. Am. J. Clin. Pathol. 2007, 128, 191–197. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Langer, R.; Ott, K.; Feith, M.; Lordick, F.; Specht, K.; Becker, K.; Hofler, H. High pretherapeutic thymidylate synthetase and MRP-1 protein levels are associated with nonresponse to neoadjuvant chemotherapy in oesophageal adenocarcinoma patients. J. Surg. Oncol. 2010, 102, 503–508. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Ling, Y.; Xu, Y.; Lu, M.Z.; Liu, Y.P.; Zhang, C.S. Elevated expression of MDR1 associated with Line-1 hypomethylation in esophageal squamous cell carcinoma. Int. J. Clin. Exp. Pathol. 2015, 8, 14392–14400. [Google Scholar] [PubMed]
- Zhang, G.; Gao, R.; Wang, J.; Fu, J.; Zhang, M.; Jin, X. Various doses of fractioned irradiation modulates multidrug resistance 1 expression differently through hypoxia-inducible factor 1α in esophageal cancer cells. Dis. Esophagus 2011, 24, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Langer, R.; Specht, K.; Becker, K.; Ewald, P.; Bekesch, M.; Sarbia, M.; Busch, R.; Feith, M.; Stein, H.J.; Siewert, J.R.; et al. Association of pretherapeutic expression of chemotherapy-related genes with response to neoadjuvant chemotherapy in Barrett carcinoma. Clin. Cancer Res. 2005, 11, 7462–7469. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, M.; Makino, T.; Masuzawa, T.; Kurokawa, Y.; Miyata, H.; Takiguchi, S.; Nakajima, K.; Fujiwara, Y.; Matsuura, N.; Mori, M.; et al. Role of multidrug resistance protein 2 (MRP2) in chemoresistance and clinical outcome in oesophageal squamous cell carcinoma. Br. J. Cancer 2011, 104, 707–713. [Google Scholar] [CrossRef] [PubMed]
- Tsunoda, S.; Okumura, T.; Ito, T.; Kondo, K.; Ortiz, C.; Tanaka, E.; Watanabe, G.; Itami, A.; Sakai, Y.; Shimada, Y. ABCG2 expression is an independent unfavorable prognostic factor in esophageal squamous cell carcinoma. Oncology 2006, 71, 251–258. [Google Scholar] [CrossRef] [PubMed]
- Harpole, D.H., Jr.; Moore, M.B.; Herndon, J.E., 2nd; Aloia, T.; D’Amico, T.A.; Sporn, T.; Parr, A.; Linoila, I.; Allegra, C. The prognostic value of molecular marker analysis in patients treated with trimodality therapy for esophageal cancer. Clin. Cancer Res. 2001, 7, 562–569. [Google Scholar] [PubMed]
- Liu, Q.; Hao, C.; Su, P.; Shi, J. Down-regulation of HLA class I antigen-processing machinery components in esophageal squamous cell carcinomas: Association with disease progression. Scand. J. Gastroenterol. 2009, 44, 960–969. [Google Scholar] [CrossRef] [PubMed]
- Shimada, Y.; Imamura, M.; Watanabe, G.; Uchida, S.; Harada, H.; Makino, T.; Kano, M. Prognostic factors of oesophageal squamous cell carcinoma from the perspective of molecular biology. Br. J. Cancer 1999, 80, 1281–1288. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhu, S.M.; Xu, X.L.; Zhao, A.N.; Hu, J.L. Expression levels of HER2 and MRP1 are not prognostic factors of long-term survival in 829 patients with esophageal squamous cell carcinoma. Oncol. Lett. 2016, 11, 745–752. [Google Scholar] [CrossRef] [PubMed]
- Gan, S.Y.; Zhong, X.Y.; Xie, S.M.; Li, S.M.; Peng, H.; Luo, F. Expression and significance of tumor drug resistance related proteins and beta-catenin in esophageal squamous cell carcinoma. Chin. J. Cancer 2010, 29, 300–305. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Xu, J.; Guo, J.; Jin, Y.; Wang, X.; Zhang, Q.; Liu, L. Circulating autoantibody to ABCC3 may be a potential biomarker for esophageal squamous cell carcinoma. Clin. Transl. Oncol. 2013, 15, 398–402. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Gong, X.; Zhou, H.; Xiong, F.; Wang, S. Depleting ABCE1 expression induces apoptosis and inhibits the ability of proliferation and migration of human esophageal carcinoma cells. Int. J. Clin. Exp. Pathol. 2014, 7, 584–592. [Google Scholar] [PubMed]
- Hang, D.; Dong, H.C.; Ning, T.; Dong, B.; Hou, D.L.; Xu, W.G. Prognostic value of the stem cell markers CD133 and ABCG2 expression in esophageal squamous cell carcinoma. Dis. Esophagus 2012, 25, 638–644. [Google Scholar] [CrossRef] [PubMed]
- Simon, R.M.; Paik, S.; Hayes, D.F. Use of archived specimens in evaluation of prognostic and predictive biomarkers. J. Natl. Cancer Inst. 2009, 101, 1446–1452. [Google Scholar] [CrossRef] [PubMed]
- Lin, E.W.; Karakasheva, T.A.; Hicks, P.D.; Bass, A.J.; Rustgi, A.K. The tumor microenvironment in esophageal cancer. Oncogene 2016, 35, 5337–5349. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Ji, Y.; Chen, L.; Li, M.; Wu, F.; Hu, J.; Jiang, J.; Cui, X.; Chen, Y.; Pang, L.; et al. Genetic variability in LMP2 and LMP7 is associated with the risk of esophageal squamous cell carcinoma in the Kazakh population but is not associated with HPV infection. PLoS ONE 2017, 12, e0186319. [Google Scholar] [CrossRef] [PubMed]
- Petrick, J.L.; Wyss, A.B.; Butler, A.M.; Cummings, C.; Sun, X.; Poole, C.; Smith, J.S.; Olshan, A.F. Prevalence of human papillomavirus among oesophageal squamous cell carcinoma cases: Systematic review and meta-analysis. Br. J. Cancer 2014, 110, 2369–2377. [Google Scholar] [CrossRef] [PubMed]
- Gharahkhani, P.; Fitzgerald, R.C.; Vaughan, T.L.; Palles, C.; Gockel, I.; Tomlinson, I.; Buas, M.F.; May, A.; Gerges, C.; Anders, M.; et al. Genome-wide association studies in oesophageal adenocarcinoma and Barrett’s oesophagus: A large-scale meta-analysis. Lancet Oncol. 2016, 17, 1363–1373. [Google Scholar] [CrossRef]
- Pasello, G.; Agata, S.; Bonaldi, L.; Corradin, A.; Montagna, M.; Zamarchi, R.; Parenti, A.; Cagol, M.; Zaninotto, G.; Ruol, A.; et al. DNA copy number alterations correlate with survival of esophageal adenocarcinoma patients. Mod. Pathol. 2009, 22, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Obara, K.; Ghazizadeh, M.; Shimizu, H.; Arai, R.; Tenjin, T.; Suzuki, S.; Moriyama, Y.; Kawanami, O. Comparative genomic hybridization study of genetic changes associated with vindesine resistance in esophageal carcinoma. Int. J. Oncol. 2002, 20, 255–260. [Google Scholar] [CrossRef] [PubMed]
- Saito, T.; Hikita, M.; Kohno, K.; Tanimura, H.; Miyahara, M.; Kobayashi, M. Enhanced expression of the multidrug resistance gene in vindesine-resistant human esophageal cancer cells. Oncology 1994, 51, 440–445. [Google Scholar] [CrossRef] [PubMed]
- Albrecht, B.; Hausmann, M.; Zitzelsberger, H.; Stein, H.; Siewert, J.R.; Hopt, U.; Langer, R.; Hofler, H.; Werner, M.; Walch, A. Array-based comparative genomic hybridization for the detection of DNA sequence copy number changes in Barrett’s adenocarcinoma. J. Pathol. 2004, 203, 780–788. [Google Scholar] [CrossRef] [PubMed]
- Van Dekken, H.; Vissers, K.; Tilanus, H.W.; Kuo, W.L.; Tanke, H.J.; Rosenberg, C.; Ijszenga, M.; Szuhai, K. Genomic array and expression analysis of frequent high-level amplifications in adenocarcinomas of the gastro-esophageal junction. Cancer Genet. Cytogenet. 2006, 166, 157–162. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Guo, L.; Peiffer, D.A.; Zhou, L.; Chan, O.T.; Bibikova, M.; Wickham-Garcia, E.; Lu, S.H.; Zhan, Q.; Wang-Rodriguez, J.; et al. Genomic profiling of 766 cancer-related genes in archived esophageal normal and carcinoma tissues. Int. J. Cancer 2008, 122, 2249–2254. [Google Scholar] [CrossRef] [PubMed]
- Garcia, E.; Hayden, A.; Birts, C.; Britton, E.; Cowie, A.; Pickard, K.; Mellone, M.; Choh, C.; Derouet, M.; Duriez, P.; et al. Authentication and characterisation of a new oesophageal adenocarcinoma cell line: MFD-1. Sci. Rep. 2016, 6, 32417. [Google Scholar] [CrossRef] [PubMed]
- Li, J.S.; Ying, J.M.; Wang, X.W.; Wang, Z.H.; Tao, Q.; Li, L.L. Promoter methylation of tumor suppressor genes in esophageal squamous cell carcinoma. Chin. J. Cancer 2013, 32, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Kaz, A.M.; Grady, W.M. Epigenetic biomarkers in esophageal cancer. Cancer Lett. 2014, 342, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Ahrens, T.D.; Werner, M.; Lassmann, S. Epigenetics in esophageal cancers. Cell Tissue Res. 2014, 356, 643–655. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Li, J.; Li, Q. CDKN2A methylation in esophageal cancer: A meta-analysis. Oncotarget 2017, 8, 50071–50083. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Ren, J.; Brock, M.V.; Herman, J.G.; Carraway, H.E. Promoter methylation of HIN-1 in the progression to esophageal squamous cancer. Epigenetics 2008, 3, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Lima, S.C.; Hernandez-Vargas, H.; Simao, T.; Durand, G.; Kruel, C.D.; Le Calvez-Kelm, F.; Ribeiro Pinto, L.F.; Herceg, Z. Identification of a DNA methylome signature of esophageal squamous cell carcinoma and potential epigenetic biomarkers. Epigenetics 2011, 6, 1217–1227. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hibi, K.; Taguchi, M.; Nakayama, H.; Takase, T.; Kasai, Y.; Ito, K.; Akiyama, S.; Nakao, A. Molecular detection of p16 promoter methylation in the serum of patients with esophageal squamous cell carcinoma. Clin. Cancer Res. 2001, 7, 3135–3138. [Google Scholar] [PubMed]
- Li, B.; Wang, B.; Niu, L.J.; Jiang, L.; Qiu, C.C. Hypermethylation of multiple tumor-related genes associated with DNMT3b up-regulation served as a biomarker for early diagnosis of esophageal squamous cell carcinoma. Epigenetics 2011, 6, 307–316. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Toh, Y.; Egashira, A.; Yamamoto, M. Epigenetic alterations and their clinical implications in esophageal squamous cell carcinoma. Gen. Thorac. Cardiovasc. Surg. 2013, 61, 262–269. [Google Scholar] [CrossRef] [PubMed]
- Zare, M.; Jazii, F.R.; Alivand, M.R.; Nasseri, N.K.; Malekzadeh, R.; Yazdanbod, M. Qualitative analysis of Adenomatous Polyposis Coli promoter: Hypermethylation, engagement and effects on survival of patients with esophageal cancer in a high risk region of the world, a potential molecular marker. BMC Cancer 2009, 9, 24. [Google Scholar] [CrossRef] [PubMed]
- Ling, Z.Q.; Li, P.; Ge, M.H.; Zhao, X.; Hu, F.J.; Fang, X.H.; Dong, Z.M.; Mao, W.M. Hypermethylation-modulated down-regulation of CDH1 expression contributes to the progression of esophageal cancer. Int. J. Mol. Med. 2011, 27, 625–635. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Lee, B.B.; Han, J.; Cho, E.Y.; Shim, Y.M.; Park, J.; Kim, D.H. CpG island hypermethylation of E-cadherin (CDH1) and integrin alpha4 is associated with recurrence of early stage esophageal squamous cell carcinoma. Int. J. Cancer 2008, 123, 2073–2079. [Google Scholar] [CrossRef] [PubMed]
- Cui, L.; Xu, L.Y.; Shen, Z.Y.; Tao, Q.; Gao, S.Y.; Lv, Z.; Du, Z.P.; Fang, W.K.; Li, E.M. NGALR is overexpressed and regulated by hypomethylation in esophageal squamous cell carcinoma. Clin. Cancer Res. 2008, 14, 7674–7681. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Yin, B.L.; He, B.; Chen, C.; Zhao, M.; Zhang, W.; Xia, Z.K.; Pan, Y.; Tang, J.; Zhou, X.; et al. Overexpression of DNA damage-induced 45 alpha gene contributes to esophageal squamous cell cancer by promoter hypomethylation. J. Exp. Clin. Cancer Res. 2012, 31, 11. [Google Scholar] [CrossRef] [PubMed]
- Iwagami, S.; Baba, Y.; Watanabe, M.; Shigaki, H.; Miyake, K.; Ishimoto, T.; Iwatsuki, M.; Sakamaki, K.; Ohashi, Y.; Baba, H. LINE-1 hypomethylation is associated with a poor prognosis among patients with curatively resected esophageal squamous cell carcinoma. Ann. Surg. 2013, 257, 449–455. [Google Scholar] [CrossRef] [PubMed]
- Baba, Y.; Watanabe, M.; Murata, A.; Shigaki, H.; Miyake, K.; Ishimoto, T.; Iwatsuki, M.; Iwagami, S.; Yoshida, N.; Oki, E.; et al. LINE-1 hypomethylation, DNA copy number alterations, and CDK6 amplification in esophageal squamous cell carcinoma. Clin. Cancer Res. 2014, 20, 1114–1124. [Google Scholar] [CrossRef] [PubMed]
- Kawano, H.; Saeki, H.; Kitao, H.; Tsuda, Y.; Otsu, H.; Ando, K.; Ito, S.; Egashira, A.; Oki, E.; Morita, M.; et al. Chromosomal instability associated with global DNA hypomethylation is associated with the initiation and progression of esophageal squamous cell carcinoma. Ann. Surg. Oncol. 2014, 21 (Suppl. S4), S696–S702. [Google Scholar] [CrossRef] [PubMed]
- Kelly, T.K.; Miranda, T.B.; Liang, G.; Berman, B.P.; Lin, J.C.; Tanay, A.; Jones, P.A. H2A.Z maintenance during mitosis reveals nucleosome shifting on mitotically silenced genes. Mol. Cell 2010, 39, 901–911. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.N.; Tan, J.; Chen, Y.X.; Fang, J.Y. Genetic variants in the histone methylation and acetylation pathway and their risks in eight types of cancer. J. Dig. Dis. 2018. [Google Scholar] [CrossRef] [PubMed]
- Herceg, Z.; Ushijima, T. Introduction: Epigenetics and cancer. Adv. Genet. 2010, 70, 1–23. [Google Scholar] [PubMed]
- Dawson, M.A.; Kouzarides, T. Cancer epigenetics: From mechanism to therapy. Cell 2012, 150, 12–27. [Google Scholar] [CrossRef] [PubMed]
- Vaissiere, T.; Sawan, C.; Herceg, Z. Epigenetic interplay between histone modifications and DNA methylation in gene silencing. Mutat. Res. 2008, 659, 40–48. [Google Scholar] [CrossRef] [PubMed]
- Toh, Y.; Yamamoto, M.; Endo, K.; Ikeda, Y.; Baba, H.; Kohnoe, S.; Yonemasu, H.; Hachitanda, Y.; Okamura, T.; Sugimachi, K. Histone H4 acetylation and histone deacetylase 1 expression in esophageal squamous cell carcinoma. Oncol. Rep. 2003, 10, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Toh, Y.; Ohga, T.; Endo, K.; Adachi, E.; Kusumoto, H.; Haraguchi, M.; Okamura, T.; Nicolson, G.L. Expression of the metastasis-associated MTA1 protein and its relationship to deacetylation of the histone H4 in esophageal squamous cell carcinomas. Int. J. Cancer 2004, 110, 362–367. [Google Scholar] [CrossRef] [PubMed]
- Tzao, C.; Tung, H.J.; Jin, J.S.; Sun, G.H.; Hsu, H.S.; Chen, B.H.; Yu, C.P.; Lee, S.C. Prognostic significance of global histone modifications in resected squamous cell carcinoma of the esophagus. Mod. Pathol. 2009, 22, 252–260. [Google Scholar] [CrossRef] [PubMed]
- I, H.; Ko, E.; Kim, Y.; Cho, E.Y.; Han, J.; Park, J.; Kim, K.; Kim, D.H.; Shim, Y.M. Association of global levels of histone modifications with recurrence-free survival in stage IIB and III esophageal squamous cell carcinomas. Cancer Epidemiol. Biomarkers Prev. 2010, 19, 566–573. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Chen, Z.; Zhang, L.; Zhou, F.; Shi, S.; Feng, X.; Li, B.; Meng, X.; Ma, X.; Luo, M.; et al. Distinctive microRNA profiles relating to patient survival in esophageal squamous cell carcinoma. Cancer Res. 2008, 68, 26–33. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Gu, J.; Wang, K.K.; Zhang, W.; Xing, J.; Chen, Z.; Ajani, J.A.; Wu, X. MicroRNA expression signatures in Barrett’s esophagus and esophageal adenocarcinoma. Clin. Cancer Res. 2009, 15, 5744–5752. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.G.; Qin, X.G.; Zhao, B.S.; Qi, B.; Yao, W.J.; Wang, T.Y.; Li, H.C.; Wu, X.N. Differential expression of miRNAs in esophageal cancer tissue. Oncol. Lett. 2013, 5, 1639–1642. [Google Scholar] [CrossRef] [PubMed]
- Sakai, N.S.; Samia-Aly, E.; Barbera, M.; Fitzgerald, R.C. A review of the current understanding and clinical utility of miRNAs in esophageal cancer. Semin. Cancer Biol. 2013, 23, 512–521. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhang, S.Y.; Gao, Y.M.; Liu, Y.F.; Liu, Y.B.; Zhao, Z.G.; Yang, K. MicroRNAs as oncogenes or tumour suppressors in oesophageal cancer: Potential biomarkers and therapeutic targets. Cell Prolif. 2014, 47, 277–286. [Google Scholar] [CrossRef] [PubMed]
- Bobryshev, Y.V.; Orekhov, A.N.; Chistiakov, D.A. MicroRNAs in Esophageal Adenocarcinoma: Functional Significance and Potential for the Development of New Molecular Disease Markers. Curr. Pharm. Des. 2015, 21, 3402–3416. [Google Scholar] [CrossRef] [PubMed]
- Hemmatzadeh, M.; Mohammadi, H.; Karimi, M.; Musavishenas, M.H.; Baradaran, B. Differential role of microRNAs in the pathogenesis and treatment of Esophageal cancer. Biomed. Pharmacother. 2016, 82, 509–519. [Google Scholar] [CrossRef] [PubMed]
- Vrana, D.; Matzenauer, M.; Aujesky, R.; Vrba, R.; Neoral, C.; Melichar, B.; Souček, P. Potential predictive role of microRNAs in the neoadjuvant treatment of esophageal cancer. Anticancer Res. 2017, 37, 403–412. [Google Scholar] [CrossRef] [PubMed]
- Imanaka, Y.; Tsuchiya, S.; Sato, F.; Shimada, Y.; Shimizu, K.; Tsujimoto, G. MicroRNA-141 confers resistance to cisplatin-induced apoptosis by targeting YAP1 in human esophageal squamous cell carcinoma. J. Hum. Genet. 2011, 56, 270–276. [Google Scholar] [CrossRef] [PubMed]
- Hummel, R.; Watson, D.I.; Smith, C.; Kist, J.; Michael, M.Z.; Haier, J.; Hussey, D.J. miR-148a improves response to chemotherapy in sensitive and resistant oesophageal adenocarcinoma and squamous cell carcinoma cells. J. Gastrointest. Surg. 2011, 15, 429–438. [Google Scholar] [CrossRef] [PubMed]
- Hummel, R.; Sie, C.; Watson, D.I.; Wang, T.; Ansar, A.; Michael, M.Z.; Van der Hoek, M.; Haier, J.; Hussey, D.J. MicroRNA signatures in chemotherapy resistant esophageal cancer cell lines. World J. Gastroenterol. 2014, 20, 14904–14912. [Google Scholar] [CrossRef] [PubMed]
- Hamano, R.; Miyata, H.; Yamasaki, M.; Kurokawa, Y.; Hara, J.; Moon, J.H.; Nakajima, K.; Takiguchi, S.; Fujiwara, Y.; Mori, M.; et al. Overexpression of miR-200c induces chemoresistance in esophageal cancers mediated through activation of the Akt signaling pathway. Clin. Cancer Res. 2011, 17, 3029–3038. [Google Scholar] [CrossRef] [PubMed]
- Sugimura, K.; Miyata, H.; Tanaka, K.; Hamano, R.; Takahashi, T.; Kurokawa, Y.; Yamasaki, M.; Nakajima, K.; Takiguchi, S.; Mori, M.; et al. Let-7 expression is a significant determinant of response to chemotherapy through the regulation of IL-6/STAT3 pathway in esophageal squamous cell carcinoma. Clin. Cancer Res. 2012, 18, 5144–5153. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, Y.; Herbst, A.; Kalinski, T.; Qin, J.; Wang, X.; Jiang, Z.; Benedix, F.; Franke, S.; Wartman, T.; et al. miR-221 Mediates Chemoresistance of Esophageal Adenocarcinoma by Direct Targeting of DKK2 Expression. Ann. Surg. 2016, 264, 804–814. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Li, M.; Han, Y.; Hong, L.; Gong, T.; Sun, L.; Zheng, X. Down-regulation of miR-27a might reverse multidrug resistance of esophageal squamous cell carcinoma. Dig. Dis. Sci. 2010, 55, 2545–2551. [Google Scholar] [CrossRef] [PubMed]
- Phatak, P.; Byrnes, K.A.; Mansour, D.; Liu, L.; Cao, S.; Li, R.; Rao, J.N.; Turner, D.J.; Wang, J.Y.; Donahue, J.M. Overexpression of miR-214-3p in esophageal squamous cancer cells enhances sensitivity to cisplatin by targeting survivin directly and indirectly through CUG-BP1. Oncogene 2016, 35, 2087–2097. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Yang, Y.; Zhao, J.; Zhao, S. BAG3-mediated miRNA let-7g and let-7i inhibit proliferation and enhance apoptosis of human esophageal carcinoma cells by targeting the drug transporter ABCC10. Cancer Lett. 2016, 371, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Odenthal, M.; Bollschweiler, E.; Grimminger, P.P.; Schroder, W.; Brabender, J.; Drebber, U.; Holscher, A.H.; Metzger, R.; Vallbohmer, D. MicroRNA profiling in locally advanced esophageal cancer indicates a high potential of miR-192 in prediction of multimodality therapy response. Int. J. Cancer 2013, 133, 2454–2463. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.Y.; Chen, Q.J.; Xu, K.; Ren, H.T.; Bao, X.; Ma, Y.N.; Wei, Y.; Ma, H.B. Involvement of microRNA-141-3p in 5-fluorouracil and oxaliplatin chemo-resistance in esophageal cancer cells via regulation of PTEN. Mol. Cell. Biochem. 2016, 422, 161–170. [Google Scholar] [CrossRef] [PubMed]
- Schwarzenbach, H.; Nishida, N.; Calin, G.A.; Pantel, K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat. Rev. Clin. Oncol. 2014, 11, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Wang, C.; Chen, X.; Yang, C.; Li, K.; Wang, J.; Dai, J.; Hu, Z.; Zhou, X.; Chen, L.; et al. Expression profile of microRNAs in serum: A fingerprint for esophageal squamous cell carcinoma. Clin. Chem. 2010, 56, 1871–1879. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, S.; Ichikawa, D.; Hirajima, S.; Kawaguchi, T.; Miyamae, M.; Okajima, W.; Ohashi, T.; Arita, T.; Konishi, H.; Shiozaki, A.; et al. Plasma microRNA profiles: Identification of miR-25 as a novel diagnostic and monitoring biomarker in oesophageal squamous cell carcinoma. Br. J. Cancer 2014, 111, 1614–1624. [Google Scholar] [CrossRef] [PubMed]
- Kurashige, J.; Kamohara, H.; Watanabe, M.; Tanaka, Y.; Kinoshita, K.; Saito, S.; Hiyoshi, Y.; Iwatsuki, M.; Baba, Y.; Baba, H. Serum microRNA-21 is a novel biomarker in patients with esophageal squamous cell carcinoma. J. Surg. Oncol. 2012, 106, 188–192. [Google Scholar] [CrossRef] [PubMed]
- Hirajima, S.; Komatsu, S.; Ichikawa, D.; Takeshita, H.; Konishi, H.; Shiozaki, A.; Morimura, R.; Tsujiura, M.; Nagata, H.; Kawaguchi, T.; et al. Clinical impact of circulating miR-18a in plasma of patients with oesophageal squamous cell carcinoma. Br. J. Cancer 2013, 108, 1822–1829. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P.; Saraya, A.; Gupta, P.; Sharma, R. Decreased levels of circulating and tissue miR-107 in human esophageal cancer. Biomarkers 2013, 18, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Liao, J.; Yang, M.; Shi, Y.; Peng, Y.; Wang, Y.; Pan, E.; Guo, W.; Pu, Y.; Yin, L. Circulating miR-155 expression in plasma: A potential biomarker for early diagnosis of esophageal cancer in humans. J. Toxicol. Environ. Health A 2012, 75, 1154–1162. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, N.; Hoshino, I.; Mori, M.; Akutsu, Y.; Hanari, N.; Yoneyama, Y.; Ikeda, N.; Isozaki, Y.; Maruyama, T.; Akanuma, N.; et al. Serum microRNA expression profile: miR-1246 as a novel diagnostic and prognostic biomarker for oesophageal squamous cell carcinoma. Br. J. Cancer 2013, 108, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.; Miyata, H.; Yamasaki, M.; Sugimura, K.; Takahashi, T.; Kurokawa, Y.; Nakajima, K.; Takiguchi, S.; Mori, M.; Doki, Y. Circulating miR-200c levels significantly predict response to chemotherapy and prognosis of patients undergoing neoadjuvant chemotherapy for esophageal cancer. Ann. Surg. Oncol. 2013, 20 (Suppl. S3), S607–S615. [Google Scholar] [CrossRef] [PubMed]
- miRTarBase. miRTarBase: The Experimentally Validated microRNA-Target Interactions Database. Available online: http://mirtarbase.mbc.nctu.edu.tw/php/index.php (accessed on 26 January 2018).
- Haenisch, S.; Werk, A.N.; Cascorbi, I. MicroRNAs and their relevance to ABC transporters. Br. J. Clin. Pharmacol. 2014, 77, 587–596. [Google Scholar] [CrossRef] [PubMed]
- Caliskan, M.; Guler, H.; Bozok Cetintas, V. Current updates on microRNAs as regulators of chemoresistance. Biomed. Pharmacother. 2017, 95, 1000–1012. [Google Scholar] [CrossRef] [PubMed]
- Shang, Y.; Zhang, Z.; Liu, Z.; Feng, B.; Ren, G.; Li, K.; Zhou, L.; Sun, Y.; Li, M.; Zhou, J.; et al. miR-508-5p regulates multidrug resistance of gastric cancer by targeting ABCB1 and ZNRD1. Oncogene 2014, 33, 3267–3276. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Yang, Z.; Xia, L.; Nie, Y.; Wu, K.; Shi, Y.; Fan, D. Methylation of miR-129-5p CpG island modulates multi-drug resistance in gastric cancer by targeting ABC transporters. Oncotarget 2014, 5, 11552–11563. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Zhang, M.; Chen, X.; Liu, Y.; Lou, G. MicroRNA-595 sensitizes ovarian cancer cells to cisplatin by targeting ABCB1. Oncotarget 2016, 7, 87091–87099. [Google Scholar] [CrossRef] [PubMed]
- Ikemura, K.; Yamamoto, M.; Miyazaki, S.; Mizutani, H.; Iwamoto, T.; Okuda, M. MicroRNA-145 post-transcriptionally regulates the expression and function of P-glycoprotein in intestinal epithelial cells. Mol. Pharmacol. 2013, 83, 399–405. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.C.; Renuse, S.; Pinto, S.; Kumar, P.; Yang, Y.; Chaerkady, R.; Godsey, B.; Mendell, J.T.; Halushka, M.K.; Civin, C.I.; et al. Identification of miR-145 targets through an integrated omics analysis. Mol. Biosyst. 2015, 11, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Ren, Y.; Tang, H.; Wang, W.; He, Q.; Sun, J.; Zhou, X.; Wang, A. Deregulation of the miR-222-ABCG2 regulatory module in tongue squamous cell carcinoma contributes to chemoresistance and enhanced migratory/invasive potential. Oncotarget 2015, 6, 44538–44550. [Google Scholar] [CrossRef] [PubMed]
- Pogribny, I.P.; Filkowski, J.N.; Tryndyak, V.P.; Golubov, A.; Shpyleva, S.I.; Kovalchuk, O. Alterations of microRNAs and their targets are associated with acquired resistance of MCF-7 breast cancer cells to cisplatin. Int. J. Cancer 2010, 127, 1785–1794. [Google Scholar] [CrossRef] [PubMed]
- Moscow, J.A.; Fairchild, C.R.; Madden, M.J.; Ransom, D.T.; Wieand, H.S.; O’Brien, E.E.; Poplack, D.G.; Cossman, J.; Myers, C.E.; Cowan, K.H. Expression of anionic glutathione-S-transferase and P-glycoprotein genes in human tissues and tumors. Cancer Res. 1989, 49, 1422–1428. [Google Scholar] [PubMed]
- Nakashima, A.; Okabayashi, T.; Nakatani, S.; Kataoka, M.; Johira, H.; Takeda, A.; Orita, K. [Analysis of MDR1 (multidrug resistance) gene expression by RT-PCR]. Gan Kagaku Ryoho 1993, 20, 831–833. [Google Scholar]
- Nooter, K.; Westerman, A.M.; Flens, M.J.; Zaman, G.J.; Scheper, R.J.; van Wingerden, K.E.; Burger, H.; Oostrum, R.; Boersma, T.; Sonneveld, P.; et al. Expression of the multidrug resistance-associated protein (MRP) gene in human cancers. Clin. Cancer Res. 1995, 1, 1301–1310. [Google Scholar] [PubMed]
- Di Nicolantonio, F.; Mercer, S.J.; Knight, L.A.; Gabriel, F.G.; Whitehouse, P.A.; Sharma, S.; Fernando, A.; Glaysher, S.; Di Palma, S.; Johnson, P.; et al. Cancer cell adaptation to chemotherapy. BMC Cancer 2005, 5, 78. [Google Scholar] [CrossRef] [PubMed]
- Nooter, K.; Kok, T.; Bosman, F.T.; van Wingerden, K.E.; Stoter, G. Expression of the multidrug resistance protein (MRP) in squamous cell carcinoma of the oesophagus and response to pre-operative chemotherapy. Eur. J. Cancer 1998, 34, 81–86. [Google Scholar] [CrossRef]
- Milano, F.; Guarriera, M.; Rygiel, A.M.; Krishnadath, K.K. Trastuzumab mediated T-cell response against HER-2/neu overexpressing esophageal adenocarcinoma depends on intact antigen processing machinery. PLoS ONE 2010, 5, e12424. [Google Scholar] [CrossRef] [PubMed]
- Leichman, L.; Lawrence, D.; Leichman, C.G.; Nava, H.; Nava, E.; Proulx, G.; Clark, K.; Khushalani, N.I.; Berdzik, J.; Greco, W.; et al. Expression of genes related to activity of oxaliplatin and 5-fluorouracil in endoscopic biopsies of primary esophageal cancer in patients receiving oxaliplatin, 5-flourouracil and radiation: Characterization and exploratory analysis with survival. J. Chemother. 2006, 18, 514–524. [Google Scholar] [CrossRef] [PubMed]
- Dvorak, K.; Watts, G.S.; Ramsey, L.; Holubec, H.; Payne, C.M.; Bernstein, C.; Jenkins, G.J.; Sampliner, R.E.; Prasad, A.; Garewal, H.S.; et al. Expression of bile acid transporting proteins in Barrett’s esophagus and esophageal adenocarcinoma. Am. J. Gastroenterol. 2009, 104, 302–309. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Xu, C.Q.; Zhu, S.T.; Wang, M.; Guo, S.L.; Sun, X.J.; Cheng, R.; Xing, J.; Wang, W.H.; Shao, L.L.; Zhang, S.T. Pathway analysis of differentially expressed genes in human esophageal squamous cell carcinoma. Eur. Rev. Med. Pharmacol. Sci. 2015, 19, 1652–1661. [Google Scholar] [PubMed]
- Xu, D.D.; Zhou, P.J.; Wang, Y.; Zhang, L.; Fu, W.Y.; Ruan, B.B.; Xu, H.P.; Hu, C.Z.; Tian, L.; Qin, J.H.; et al. Reciprocal activation between STAT3 and miR-181b regulates the proliferation of esophageal cancer stem-like cells via the CYLD pathway. Mol. Cancer 2016, 15, 40. [Google Scholar] [CrossRef] [PubMed]
- Kim, R.; Hirabayashi, N.; Nishiyama, M.; Jinushi, K.; Toge, T.; Okada, K. Factors contributing to adriamycin sensitivity in human xenograft tumors: The relationship between expression of the MDR1, GST-pi and topoisomerase II genes and tumor sensitivity to adriamycin. Anticancer Res. 1992, 12, 241–245. [Google Scholar] [PubMed]
- Hu, X.; Akutsu, Y.; Suganami, A.; Qin, W.; Hanari, N.; Murakam, K.; Kano, M.; Usui, A.; Suito, H.; Takahashi, M. Low-dose hyperthermia enhances the antitumor effects of chemotherapy in squamous cell carcinoma. Dis. Esophagus 2017, 30, 1–7. [Google Scholar] [CrossRef]
- Saito, T.; Hikita, M.; Kohno, K.; Sato, S.; Takano, H.; Kobayashi, M. Different sensitivities of human esophageal cancer cells to multiple anti-cancer agents and related mechanisms. Cancer 1992, 70, 2402–2409. [Google Scholar] [CrossRef]
- Oosthuizen, M.M.; Nel, M.J.; Greyling, D. Heat shock treated oesophageal cancer cells become thermosensitized against anticancer drugs. Anticancer Res. 2000, 20, 2697–2703. [Google Scholar] [PubMed]
- Murase, M.; Kodera, Y.; Kondo, K.; Sekiguchi, H.; Fujiwara, M.; Kasai, Y.; Akiyama, S.; Ito, K.; Takagi, H. Expression of MRP and mdr1 in human gastrointestinal cancer cell lines: A correlation with resistance against doxorubicin. J. Surg. Oncol. 1996, 61, 223–229. [Google Scholar] [CrossRef]
- Wen, J.; Zheng, B.; Hu, Y.; Zhang, X.; Yang, H.; Luo, K.J.; Zhang, X.; Li, Y.F.; Fu, J.H. Establishment and biological analysis of the EC109/CDDP multidrug-resistant esophageal squamous cell carcinoma cell line. Oncol. Rep. 2009, 22, 65–71. [Google Scholar] [PubMed]
- Wang, T.H.; Wan, J.Y.; Gong, X.; Li, H.Z.; Cheng, Y. Tetrandrine enhances cytotoxicity of cisplatin in human drug-resistant esophageal squamous carcinoma cells by inhibition of multidrug resistance-associated protein 1. Oncol. Rep. 2012, 28, 1681–1686. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, R.; Ishiguro, H.; Kuwabara, Y.; Kimura, M.; Mitsui, A.; Mori, Y.; Mori, R.; Tomoda, K.; Katada, T.; Harada, K.; et al. Identification of candidate genes involved in the radiosensitivity of esophageal cancer cells by microarray analysis. Dis. Esophagus 2008, 21, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, N.; Kimura, H.; Faried, A.; Sakai, M.; Sano, A.; Inose, T.; Sohda, M.; Okada, K.; Nakajima, M.; Miyazaki, T.; et al. Quantitative analysis of cisplatin sensitivity of human esophageal squamous cancer cell lines using in-air micro-PIXE. Cancer Sci. 2010, 101, 1487–1492. [Google Scholar] [CrossRef] [PubMed]
- Li, X.R.; Yang, L.Z.; Huo, X.Q.; Wang, Y.; Yang, Q.H.; Zhang, Q.Q. Effects of silencing the ATP-binding cassette protein E1 gene by electroporation on the proliferation and migration of EC109 human esophageal cancer cells. Mol. Med. Rep. 2015, 12, 837–842. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Minegaki, T.; Takara, K.; Hamaguchi, R.; Tsujimoto, M.; Nishiguchi, K. Factors affecting the sensitivity of human-derived esophageal carcinoma cell lines to 5-fluorouracil and cisplatin. Oncol. Lett. 2013, 5, 427–434. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Dvorak, P.; Pesta, M.; Soucek, P. ABC gene expression profiles have clinical importance and possibly form a new hallmark of cancer. Tumour Biol. 2017, 39, 1010428317699800. [Google Scholar] [CrossRef] [PubMed]
- Robey-Cafferty, S.S.; Rutledge, M.L.; Bruner, J.M. Expression of a multidrug resistance gene in esophageal adenocarcinoma. Correlation with response to chemotherapy and comparison with gastric adenocarcinoma. Am. J. Clin. Pathol. 1990, 93, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Sur, M.; Taylor, L.; Cooper, K.; Sur, R.K. Lack of correlation of P-glycoprotein expression with response to MIC chemotherapy in oesophageal cancer. J. Clin. Pathol. 1997, 50, 534. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Darnton, S.J.; Jenner, K.; Steyn, R.S.; Ferry, D.R.; Matthews, H.R. Lack of correlation of P-glycoprotein expression with response to MIC chemotherapy in oesophageal cancer. J. Clin. Pathol. 1995, 48, 1064–1066. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Sumarpo, A.; Saiki, Y.; Chen, N.; Sunamura, M.; Horii, A. ABCB1 Is Upregulated in Acquisition of Taxane Resistance: Lessons from Esophageal Squamous Cell Carcinoma Cell Lines. Tohoku J. Exp. Med. 2016, 240, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, Q.; Jin, S.; Deng, W.; Li, S.; Tong, Q.; Chen, Y. Up-regulation of P-glycoprotein is involved in the increased paclitaxel resistance in human esophageal cancer radioresistant cells. Scand. J. Gastroenterol. 2012, 47, 802–808. [Google Scholar] [CrossRef] [PubMed]
- Takebayashi, Y.; Akiyama, S.; Natsugoe, S.; Hokita, S.; Niwa, K.; Kitazono, M.; Sumizawa, T.; Tani, A.; Furukawa, T.; Aikou, T. The expression of multidrug resistance protein in human gastrointestinal tract carcinomas. Cancer 1998, 82, 661–666. [Google Scholar] [CrossRef]
- Zhang, S.; Cao, W.; Yue, M.; Zheng, N.; Hu, T.; Yang, S.; Dong, Z.; Lu, S.; Mo, S. Caveolin-1 affects tumor drug resistance in esophageal squamous cell carcinoma by regulating expressions of P-gp and MRP1. Tumour Biol. 2016, 37, 9189–9196. [Google Scholar] [CrossRef] [PubMed]
- Bharthuar, A.; Saif Ur Rehman, S.; Black, J.D.; Levea, C.; Malhotra, U.; Mashtare, T.L.; Iyer, R. Breast cancer resistance protein (BCRP) and excision repair cross complement-1 (ERCC1) expression in esophageal cancers and response to cisplatin and irinotecan based chemotherapy. J. Gastrointest. Oncol. 2014, 5, 253–258. [Google Scholar] [PubMed]
- Huang, L.; Lu, Q.; Han, Y.; Li, Z.; Zhang, Z.; Li, X. ABCG2/V-ATPase was associated with the drug resistance and tumor metastasis of esophageal squamous cancer cells. Diagn. Pathol. 2012, 7, 180. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Mathur, A.; Zhang, Y.; Xi, S.; Atay, S.; Hong, J.A.; Datrice, N.; Upham, T.; Kemp, C.D.; Ripley, R.T.; et al. Mithramycin represses basal and cigarette smoke-induced expression of ABCG2 and inhibits stem cell signaling in lung and esophageal cancer cells. Cancer Res. 2012, 72, 4178–4192. [Google Scholar] [CrossRef] [PubMed]
- To, K.K.; Yu, L.; Liu, S.; Fu, J.; Cho, C.H. Constitutive AhR activation leads to concomitant ABCG2-mediated multidrug resistance in cisplatin-resistant esophageal carcinoma cells. Mol. Carcinog. 2012, 51, 449–464. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Zuo, L.F.; Guo, J.W. ABCG2 gene amplification and expression in esophageal cancer cells with acquired adriamycin resistance. Mol. Med. Rep. 2014, 9, 1299–1304. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Ju, Y.; Wang, J.; Zhou, R. Epigallocatechin-3-gallate promotes apoptosis and reversal of multidrug resistance in esophageal cancer cells. Pathol. Res. Pract. 2017, 213, 1242–1250. [Google Scholar] [CrossRef] [PubMed]
- Pisarev, A.V.; Skabkin, M.A.; Pisareva, V.P.; Skabkina, O.V.; Rakotondrafara, A.M.; Hentze, M.W.; Hellen, C.U.; Pestova, T.V. The role of ABCE1 in eukaryotic posttermination ribosomal recycling. Mol. Cell 2010, 37, 196–210. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Hu, Z. Y-box-binding protein 1 promotes tumor progression and inhibits cisplatin chemosensitivity in esophageal squamous cell carcinoma. Biomed. Pharmacother. 2016, 79, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Wang, Q.; Tian, Y. Reversal effect of ouabain on multidrug resistance in esophageal carcinoma EC109/CDDP cells by inhibiting the translocation of Wnt/beta-catenin into the nucleus. Tumour Biol. 2016, 37, 15937–15947. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yan, Y.; Ma, W.; Wu, S. Knockdown of frizzled-7 inhibits cell growth and metastasis and promotes chemosensitivity of esophageal squamous cell carcinoma cells by inhibiting Wnt signaling. Biochem. Biophys. Res. Commun. 2017, 490, 1112–1118. [Google Scholar] [CrossRef] [PubMed]
- Sims-Mourtada, J.; Izzo, J.G.; Ajani, J.; Chao, K.S. Sonic Hedgehog promotes multiple drug resistance by regulation of drug transport. Oncogene 2007, 26, 5674–5679. [Google Scholar] [CrossRef] [PubMed]
- Takakura, Y.; Hinoi, T.; Oue, N.; Sasada, T.; Kawaguchi, Y.; Okajima, M.; Akyol, A.; Fearon, E.R.; Yasui, W.; Ohdan, H. CDX2 regulates multidrug resistance 1 gene expression in malignant intestinal epithelium. Cancer Res. 2010, 70, 6767–6778. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Zhang, X.; Zhang, M.; Zhang, J.; Sheng, Y.; Sun, X.; Chen, Q.; Wang, L.X. Increased Nanog expression promotes tumor development and Cisplatin resistance in human esophageal cancer cells. Cell. Physiol. Biochem. 2012, 30, 943–952. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Xiang, X.; Yang, F.; Xiao, D.; Liu, K.; Chen, Z.; Zhang, R.; Feng, G. Functional evidence that the self-renewal gene NANOG regulates esophageal squamous cancer development. Biochem. Biophys. Res. Commun. 2017, 490, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.; Gao, Q.; Guo, L.; Zhang, C.; Jiang, W.; Li, H.; Wang, J.; Han, X.; Shi, Y.; Lu, S.H. Isolation and identification of cancer stem-like cells in esophageal carcinoma cell lines. Stem Cells Dev. 2009, 18, 465–473. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, P.; Chueca, E.; Arruebo, M.; Strunk, M.; Solanas, E.; Serrano, T.; Garcia-Gonzalez, M.A.; Lanas, A. CD24 Expression Is Increased in 5-Fluorouracil-Treated Esophageal Adenocarcinoma Cells. Front. Pharmacol. 2017, 8, 321. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Bao, Q.; Schwarz, B.; Zhao, L.; Mysliwietz, J.; Ellwart, J.; Renner, A.; Hirner, H.; Niess, H.; Camaj, P.; et al. Stem cell-like side populations in esophageal cancer: A source of chemotherapy resistance and metastases. Stem Cells Dev. 2014, 23, 180–192. [Google Scholar] [CrossRef] [PubMed]
- Cerovska, E.; Elsnerova, K.; Vaclavikova, R.; Soucek, P. The role of membrane transporters in ovarian cancer chemoresistance and prognosis. Expert Opin. Drug Metab. Toxicol. 2017, 13, 741–753. [Google Scholar] [CrossRef] [PubMed]
- Xiong, T.; Xu, G.; Huang, X.L.; Lu, K.Q.; Xie, W.Q.; Yin, K.; Tu, J. ATP-binding cassette transporter A1: A promising therapy target for prostate cancer. Mol. Clin. Oncol. 2018, 8, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Huang, D.; Yu, Y.; Zhang, J.; Fang, Q.; Xie, C. ABCG1 as a potential oncogene in lung cancer. Exp. Ther. Med. 2017, 13, 3189–3194. [Google Scholar] [CrossRef] [PubMed]
- Hlavac, V.; Soucek, P. Role of family D ATP-binding cassette transporters (ABCD) in cancer. Biochem. Soc. Trans. 2015, 43, 937–942. [Google Scholar] [CrossRef] [PubMed]
- Vogelstein, B.; Papadopoulos, N.; Velculescu, V.E.; Zhou, S.; Diaz, L.A., Jr.; Kinzler, K.W. Cancer genome landscapes. Science 2013, 339, 1546–1558. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Xiang, L.; Li, T.; Bai, Z. Cancer Hallmarks, Biomarkers and Breast Cancer Molecular Subtypes. J. Cancer 2016, 7, 1281–1294. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Chen, Y.; Huang, L.; Liu, T.; Huang, Y.; Zhao, J.; Wang, X.; Yang, K.; Ma, S.; Huang, L.; et al. Cetuximab enhanced the efficacy of chemotherapeutic agent in ABCB1/P-glycoprotein-overexpressing cancer cells. Oncotarget 2015, 6, 40850–40865. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Chen, Y.; Xu, M.; Chen, L.; Zhang, X.; To, K.K.; Zhao, H.; Wang, F.; Xia, Z.; Chen, X.; et al. Osimertinib (AZD9291) Enhanced the Efficacy of Chemotherapeutic Agents in ABCB1- and ABCG2-Overexpressing Cells In Vitro, In Vivo, and Ex Vivo. Mol. Cancer Ther. 2016, 15, 1845–1858. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Chen, Z.; To, K.K.W.; Fang, X.; Wang, F.; Cheng, B.; Fu, L. Effect of abemaciclib (LY2835219) on enhancement of chemotherapeutic agents in ABCB1 and ABCG2 overexpressing cells in vitro and in vivo. Biochem. Pharmacol. 2017, 124, 29–42. [Google Scholar] [CrossRef] [PubMed]
- Suntharalingam, M.; Winter, K.; Ilson, D.; Dicker, A.P.; Kachnic, L.; Konski, A.; Chakravarthy, A.B.; Anker, C.J.; Thakrar, H.; Horiba, N.; et al. Effect of the Addition of Cetuximab to Paclitaxel, Cisplatin, and Radiation Therapy for Patients with Esophageal Cancer: The NRG Oncology RTOG 0436 Phase 3 Randomized Clinical Trial. JAMA Oncol. 2017, 3, 1520–1528. [Google Scholar] [CrossRef] [PubMed]
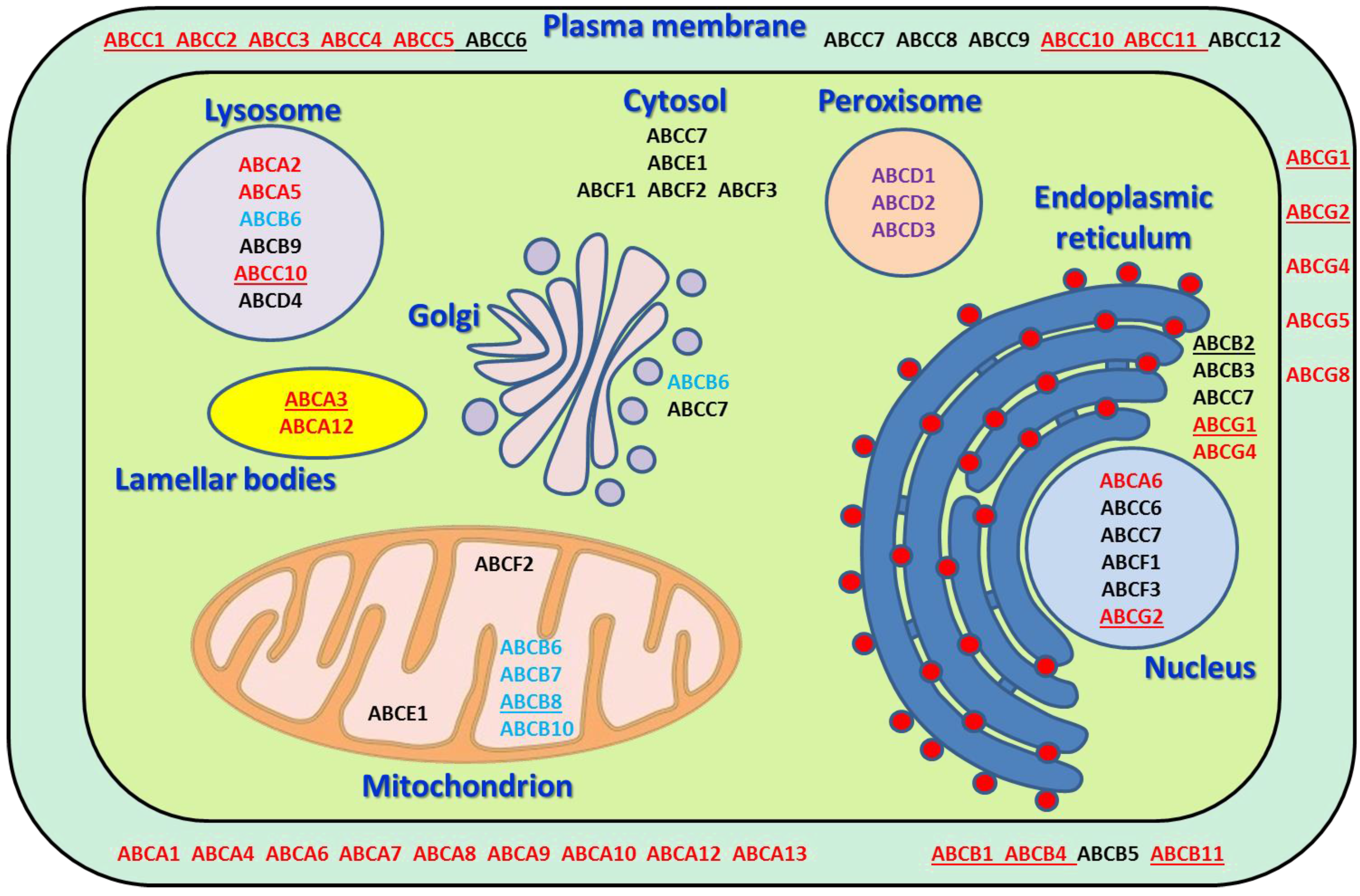

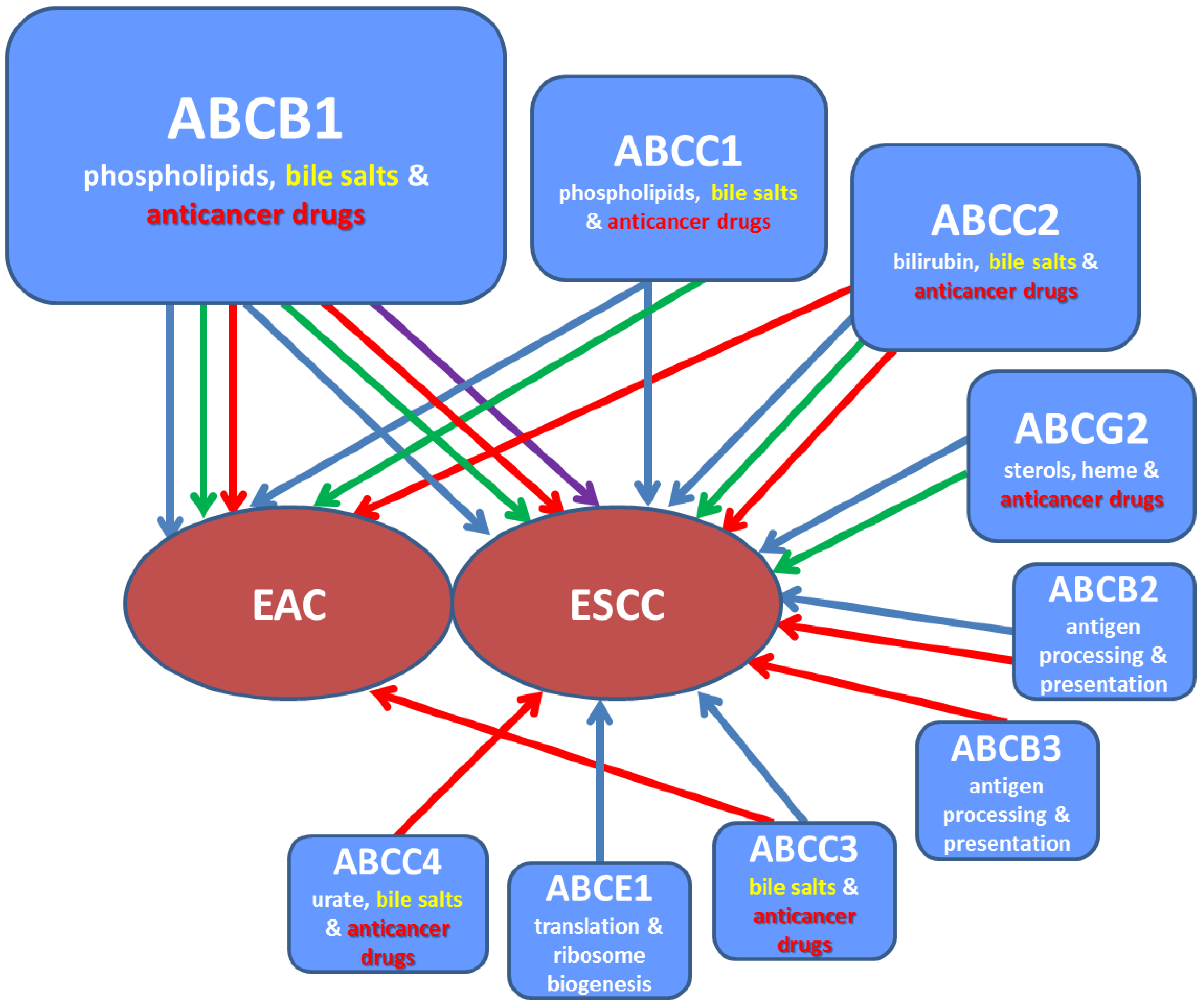

| Gene * | Chromosomal Location * | Gene (bp) * | Protein Name * | Protein (AA) * | Subcellular Localization * | Physiological Substrates/Function * | Examples of Translocated Drugs * | Pathology * |
|---|---|---|---|---|---|---|---|---|
| ABCA1 | 9q31.1 | 147,245 | ABC1 | 2261 | PM | cholesterol, phospholipids, sphingomyelin | statins, glyburide, fenofibrate | Tangier disease, hypercholesterolemia |
| ABCA2 | 9q34.3 | 21,748 | ABC2 | 2435 | lysosome | cholesterol, sterols | Pulmonary surfactant metabolism dysfunction | |
| ABCA3 | 16p13.3 | 64,869 | ABC3 | 1704 | lamellar bodies | phospholipids, sphingomyelin | imatinib, gleevec | Pulmonary surfactant metabolism dysfunction |
| ABCA4 | 1p22.1 | 128,315 | ABCR | 2273 | PM | retinoids, phospholipids | Stargardt disease, retinal dystrophy | |
| ABCA5 | 17q24.3 | 82,934 | ABCA5 | 1642 | lysosome | cholesterol | tacrolimus | Gingival fibromatosis with hypertrichosis |
| ABCA6 | 17q24.2-q24.3 | 63,904 | ABCA6 | 1617 | nucleus, PM | cholesterol, taurocholate | ||
| ABCA7 | 19p13.3 | 25,472 | ABCX | 2146 | PM | phospholipids | Alzheimer disease | |
| ABCA8 | 17q24.2 | 88,122 | ABCA8 | 1581 | PM | cholesterol | ||
| ABCA9 | 17q24.2 | 92,956 | ABCA9 | 1624 | PM | macrophage lipid homeostasis | ||
| ABCA10 | 17q24.3 | 97,633 | ABCA10 | 1543 | PM | macrophage lipid homeostasis | ||
| ABCA12 | 2q35 | 207,039 | ABCA12 | 2595 | PM, lamellar bodies | phospholipids | Ichthyosis | |
| ABCA13 | 7p12.3 | 476,050 | ABCA13 | 5058 | PM | unknown | ||
| ABCB1 | 7q21.12 | 209,691 | MDR1 | 1280 | PM | phospholipids, sphingolipids, bile salts | anthracyclines, nucleoside analogs, taxanes, vinca alkaloids, statins | |
| ABCB2 | 6p21.32 | 8770 | TAP1 | 808 | ER | antigen processing and presentation | lapatinib | |
| ABCB3 | 6p21.32 | 16,991 | TAP2 | 686 | ER | antigen processing and presentation | ||
| ABCB4 | 7q21.12 | 78,739 | MDR3 | 1286 | PM | phospholipids, bile salts | colchicine, anthracyclines, silodosin | Cholecystitis, familial intrahepatic cholestasis |
| ABCB5 | 7p21.1 | 161,832 | ABCB5 | 1257 | PM | glutathione level modulator | bilastine, dasabuvir, delafloxacin, naldemedine | |
| ABCB6 | 2q35 | 9225 | MTABC3 | 842 | mitochondrion, lysosome, Golgi | iron homeostasis | Dyschromatosis universalis hereditaria, familial pseudohyperkalemia, microphthalmia | |
| ABCB7 | Xq13.3 | 103,561 | ABC7 | 752 | mitochondrion | iron homeostasis | Spinocerebellar ataxia, sideroblastic anemia | |
| ABCB8 | 7q36.1 | 19,361 | MABC1 | 735 | mitochondrion | iron homeostasis | doxorubicin | |
| ABCB9 | 12q24.31 | 62,942 | TAPL | 766 | lysosome | cytosolic peptides | ||
| ABCB10 | 1q42.13 | 42,114 | MTABC2 | 738 | mitochondrion | iron homeostasis | ||
| ABCB11 | 2q31.1 | 112,089 | BSEP | 1321 | PM | phospholipids, bile salts | anthracyclines, nucleoside analogs, taxanes, vinca alkaloids, statins | Familial intrahepatic cholestasis |
| ABCC1 | 16p13.11 | 193,498 | MRP1 | 1531 | PM | phospholipids, bile salts, steroids, cobalamin | irinotecan, anthracyclines, taxanes, vinca alkaloids, statins | Pseudoxanthoma elasticum |
| ABCC2 | 10q24.2 | 69,979 | MRP2 | 1545 | PM | bilirubin, bile salts | irinotecan, anthracyclines, taxanes, vinca alkaloids, platinum derivatives, nucleoside analogs | Dubin-Johnson syndrome |
| ABCC3 | 17q21.33 | 57,476 | MRP3 | 1527 | PM | bile salts | anthracyclines, vinca alkaloids, platinum derivatives, nucleoside analogs | |
| ABCC4 | 13q32.1 | 281,618 | MRP4 | 1325 | PM | bile salts, uric acid | anthracyclines, cyclophosphamide, platinum derivatives, nucleoside analogs | |
| ABCC5 | 3q27.1 | 98,082 | MRP5 | 1437 | PM | bile salts, folate, glutamate | irinotecan, platinum derivatives, nucleoside analogs, atorvastatin, probenecid, rifampin | |
| ABCC6 | 16p13.11 | 74,746 | MRP6 | 1503 | PM, nucleus | putative biomineralization modulator | anthracyclines, platinum derivatives, taxanes, vinca alkaloids, etoposide, TKIs | Pseudoxanthoma elasticum, arterial calcifications |
| ABCC7 | 7q31.2 | 250,188 | CFTR | 1480 | all compartments | ATP-gated chloride channel | glyburide, ibuprofen, felodipine | Cystic fibrosis, hereditary pancreatitis |
| ABCC8 | 11p15.1 | 83,961 | SUR1 | 1581 | PM | ATP-sensitive potassium channel regulation | glyburide, tolbutamide, tolazamide. | Neonatal diabetes mellitus, hyperinsulinemic hypoglycemia, familial hyperinsulinism |
| ABCC9 | 12p12.1 | 145,141 | SUR2 | 1549 | PM | ATP-sensitive potassium channel regulation | glyburide, nicorandil | Dilated cardiomyopathy, hypertrichotic osteochondrodysplasia |
| ABCC10 | 6p21.1 | 24,615 | MRP7 | 1492 | PM, lysosome | bile salts, steroids, glutathione conjugates | anthracyclines, nucleoside analogs, taxanes, vinca alkaloids, etoposide | |
| ABCC11 | 16q12.1 | 80,659 | MRP8 | 1382 | PM | bile salts, steroids, cyclic nucleotides, glutathione conjugates, folate, taurocholate | estrogens, fluorouracil, methotrexate, probenecid, indomethacin | |
| ABCC12 | 16q12.1 | 73,046 | MRP9 | 1359 | PM | unknown | ||
| ABCD1 | Xq28 | 19,912 | ALD | 745 | peroxisome | very long chain fatty acids | ergocalciferol | Adrenoleukodystrophy, Addison’s disease |
| ABCD2 | 12q12 | 101,296 | ALDL1 | 740 | peroxisome | very long chain fatty acids | ||
| ABCD3 | 1p21.3 | 100,581 | PXMP1 | 659 | peroxisome | very long and long chain fatty acids, branched-chain fatty acids, bile acid intermediates | Bile acid synthesis defect | |
| ABCD4 | 14q24.3 | 17,788 | PXMP1L | 606 | lysosome | cobalamin | Methylmalonic aciduria and homocystinuria | |
| ABCE1 | 4q31.21 | 31,593 | RNASEL1 | 599 | mitochondrion, cytosol | translation and ribosome biogenesis | ||
| ABCF1 | 6p21.33 | 25,804 | ABC50 | 845 | nucleus, cytosol | translation initiation | ||
| ABCF2 | 7q36.1 | 19,545 | ABC28 | 623 | mitochondrion, cytosol | cell volume regulation | ||
| ABCF3 | 3q27.1 | 8430 | ABCF3 | 709 | nucleus, cytosol | putative translational regulation and apoptosis | ||
| ABCG1 | 21q22.3 | 104,699 | ABC8 | 678 | ER, PM | cholesterol, phospholipids, sterols | irinotecan, fluorouracil, leucovorin | Tangier disease, sitosterolemia |
| ABCG2 | 4q22.1 | 141,154 | BCRP | 655 | PM, nucleus | sterols, heme | anthracyclines, platinum derivatives, nucloside analogs, taxanes, vinca alkaloids, etoposide | Tangier disease, sitosterolemia |
| ABCG4 | 11q23.3 | 13,655 | WHITE2 | 646 | ER, PM | lipids, cholesterol, sterols | ||
| ABCG5 | 2p21 | 32,711 | sterolin 1 | 651 | PM | cholesterol, sterols | ezetimibe | Sitosterolemia |
| ABCG8 | 2p21 | 51,236 | sterolin 2 | 673 | PM | cholesterol, sterols | ezetimibe, atorvastatin | sitosterolemia, atherosclerosis |
| Level | Gene | Factor | Patients (N) | Association | Significance | Reference |
|---|---|---|---|---|---|---|
| Gene germline | ABCB1 | rs1045642-T allele | 210 ESCC and EAC NACRT-treated | longer overall survival after cisplatin | p = 0.030 | [17] |
| ABCB1 | rs1045642 | 31 ESCC | no effect on response and survival | NS | [19] | |
| ABCB1 | rs1045642-T allele | 116 ESCC and 146 EAC NACRT-treated | longer overall survival | p = 0.048 | [14] | |
| ABCB1 | rs1045642 | meta-analysis | putative biomarker of recurrence and survival | LOE IV # | [21] | |
| ABCB1 ** | rs2032582 ** | 62 ESCC and 62 EAC NACRT-treated | relapse-free and cancer-free survival | p < 0.001 ** | [18] | |
| TAP1 | rs1135216-G allele | 361 ESCC and 66 controls | higher risk of ESCC | 0.018 | [22] | |
| TAP1 | rs1135216-AG heterozygote | 200 ESCC and 100 controls | higher risk of ESCC | p = 0.035 | [23] | |
| TAP2 | rs1800454-AA homozygote | 265 ESCC and 357 controls | higher risk of ESCC | p = 0.023 | [24] | |
| ABCC2 * | rs17222723 * | 116 ESCC and EAC NACRT-treated | response | p = 0.002 * | [25] | |
| ABCC3 * | rs2277624 * | 116 ESCC and EAC NACRT-treated | response | p = 0.002 * | [25] | |
| Gene somatic | ABCC4 | presence of CNV | 1048 Chinese Han subjects | higher ESCC risk and poor overall survival | p = 0.001 | [26] |
| all ABCs | SNV | 2046 ESCC and 568 EAC | unknown | unknown | [27] | |
| Epigenetics-miRNA | ABCB1 | high miR-296 | 25 ESCC | poor survival | p < 0.05 | [28] |
| ABCB1 | high miR-483 and miR-214 | 104 ESCC | poor survival | p < 0.05 | [29] | |
| Transcript | ABCB1 | expression | 46 ESCC | no prognostic role | NS | [30] |
| ABCB1 | expression | 31 EAC NACRT-treated | no prognostic role | NS | [31] | |
| ABCB1 | high expression | 40 EAC NACRT-treated | no prognostic role | NS | [32] | |
| ABCB1 | expression | 310 ESCC | poor overall survival | p = 0.014 | [33] | |
| ABCB1 | expression | 54 unspecified EC derived cell lines | ↑ after high dose ↓ after low dose radiotherapy | p < 0.05 | [34] | |
| ABCC1 | high expression | 38 EAC NACRT-treated | longer overall survival and response | p = 0.017/p = 0.007 | [35] | |
| ABCC1 | low expression | 31 EAC NACRT-treated | response to NACRT | p = 0.041 | [31] | |
| ABCC1 | high expression | 40 EAC NACRT-treated | no prognostic role | NS | [32] | |
| ABCC2 | high expression | 42 ESCC NACRT-treated | poor response to NACT | p = 0.003 | [36] | |
| ABCG2 | high expression | 33 ESCC | poor survival | p = 0.017 | [37] | |
| Protein | ABCB1 | high expression | 118 EAC and ESCC NACRT-treated | poor cancer-free survival | p = 0.05 | [38] |
| ABCB2 | expression | 143 ESCC | correlates with tumor grade and metastasis | p < 0.05 | [39] | |
| ABCC1 | high expression | 40 EAC NACRT-treated | poor response to chemotherapy | p = 0.036 | [32] | |
| ABCC1 | expression | 116 ESCC | no prognostic role | NS | [40] | |
| ABCC1 | expression | 829 ESCC | no prognostic role | NS | [41] | |
| ABCC2 | high expression | 582 ESCC | correlates with tumor grade | p < 0.01 | [42] | |
| ABCC2 | expression | 81 ESCC | poor overall survival and response to NACRT | p = 0.027/p = 0.003 | [36] | |
| ABCC3 | IgA autoantibodies | 114 ESCC and 226 controls | diagnostic and predictive biomarker | p < 0.001 | [43] | |
| ABCE1 | expression | 112 ESCC | correlates with ESCC grade and stage | p < 0.001 | [44] | |
| ABCG2 | expression | 100 ESCC | poor survival | p = 0.009 | [37] | |
| ABCG2 | high expression | 110 ESCC | poor overall survival | p = 0.005 | [45] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vrana, D.; Hlavac, V.; Brynychova, V.; Vaclavikova, R.; Neoral, C.; Vrba, J.; Aujesky, R.; Matzenauer, M.; Melichar, B.; Soucek, P. ABC Transporters and Their Role in the Neoadjuvant Treatment of Esophageal Cancer. Int. J. Mol. Sci. 2018, 19, 868. https://doi.org/10.3390/ijms19030868
Vrana D, Hlavac V, Brynychova V, Vaclavikova R, Neoral C, Vrba J, Aujesky R, Matzenauer M, Melichar B, Soucek P. ABC Transporters and Their Role in the Neoadjuvant Treatment of Esophageal Cancer. International Journal of Molecular Sciences. 2018; 19(3):868. https://doi.org/10.3390/ijms19030868
Chicago/Turabian StyleVrana, David, Viktor Hlavac, Veronika Brynychova, Radka Vaclavikova, Cestmir Neoral, Jiri Vrba, Rene Aujesky, Marcel Matzenauer, Bohuslav Melichar, and Pavel Soucek. 2018. "ABC Transporters and Their Role in the Neoadjuvant Treatment of Esophageal Cancer" International Journal of Molecular Sciences 19, no. 3: 868. https://doi.org/10.3390/ijms19030868
APA StyleVrana, D., Hlavac, V., Brynychova, V., Vaclavikova, R., Neoral, C., Vrba, J., Aujesky, R., Matzenauer, M., Melichar, B., & Soucek, P. (2018). ABC Transporters and Their Role in the Neoadjuvant Treatment of Esophageal Cancer. International Journal of Molecular Sciences, 19(3), 868. https://doi.org/10.3390/ijms19030868