Regulation of Organic Anion Transporting Polypeptides (OATP) 1B1- and OATP1B3-Mediated Transport: An Updated Review in the Context of OATP-Mediated Drug-Drug Interactions
Abstract
1. Introduction
2. Scope
3. Substrate Transport Specificity and Transport Mechanism of OATP1B1 and OATP1B3
4. Altered Hepatic Disposition of OATP1B1/1B3 Substrates Due to Genetic Variation and Drug-Drug Interactions
5. Altered Expression of OATP1B1 and 1B3 in Pathological Conditions
6. Transcriptional Regulation of OATP1B1 and OATP1B3
7. Post-Translational Regulation of OATP1B1 and OATP1B3
7.1. Glycosylation
7.2. Phosphorylation
7.3. Ubiquitination
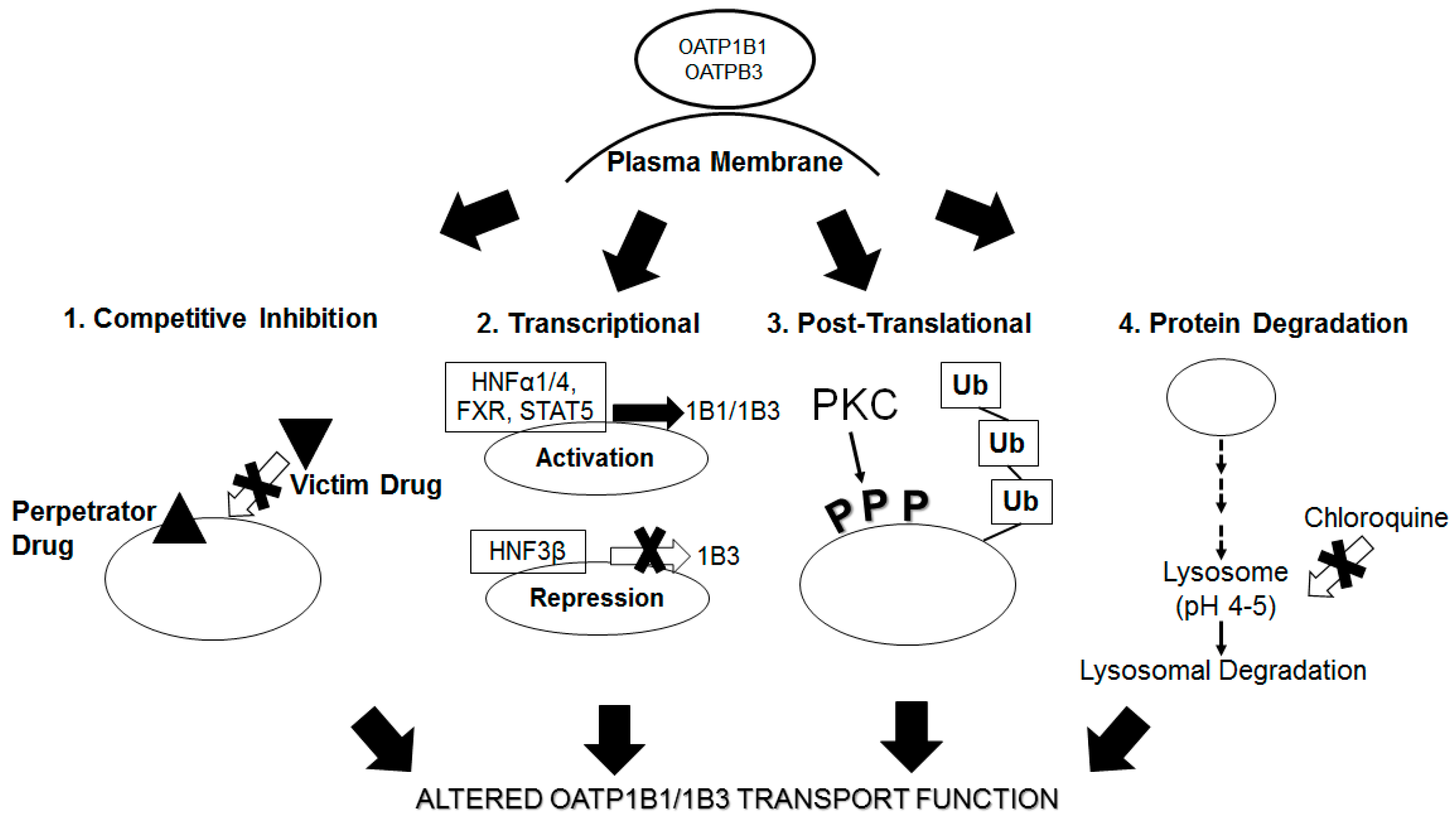
8. Regulation of OATP1B1/1B3 Transport Function by Drugs Perturbing Protein Degradation
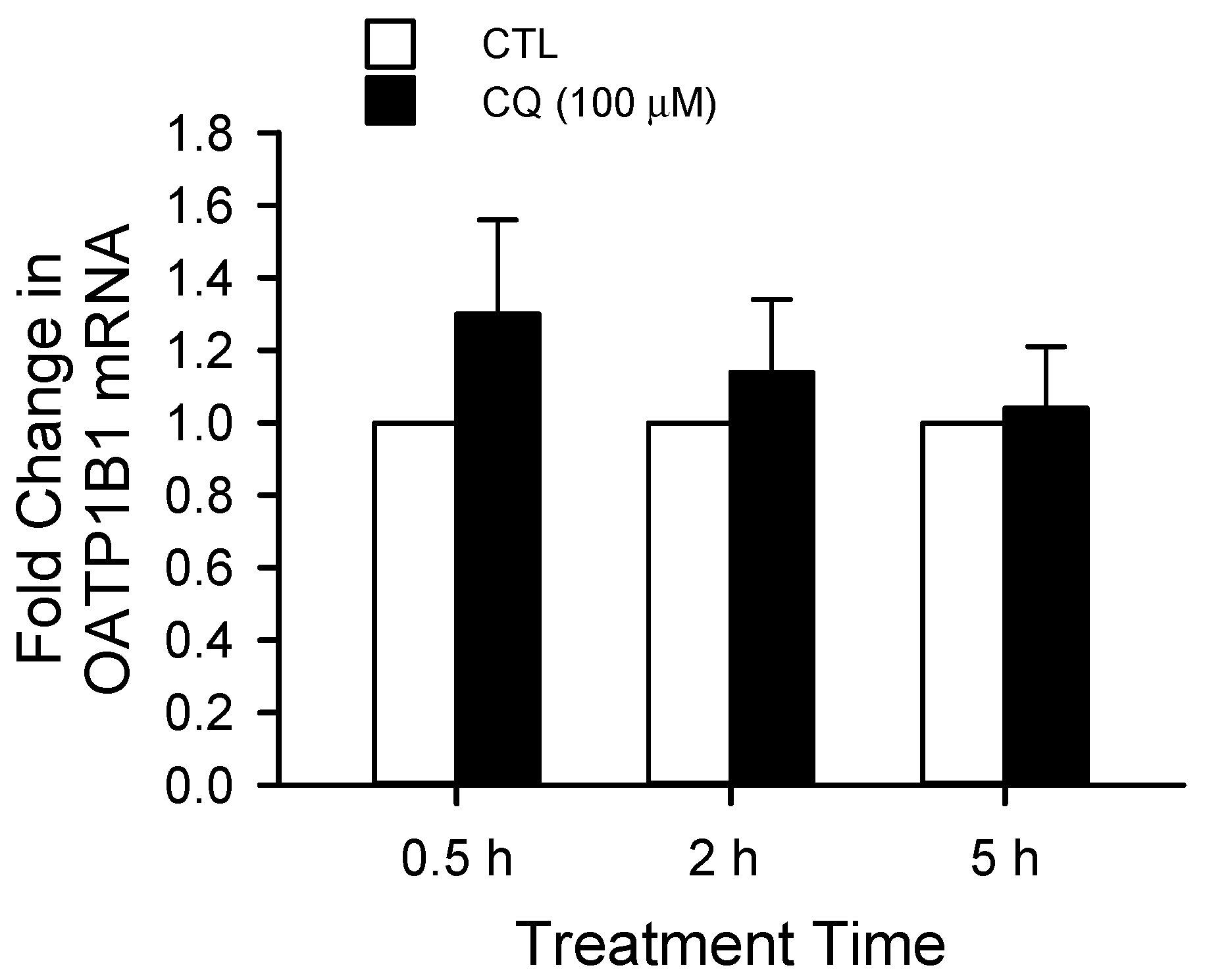
8.1. Regulation of OATP1B1 and OATP1B3 Transport Function by Lysosome Inhibition
8.2. Regulation of OATP1B1 and OATP1B3 Transport Function by Proteasome Inhibitors
9. Discussion and Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Hagenbuch, B.; Meier, P.J. Organic anion transporting polypeptides of the OATP/SLC21 family: Phylogenetic classification as OATP/SLCO superfamily, new nomenclature and molecular/functional properties. Pflugers Arch. 2004, 447, 653–665. [Google Scholar] [CrossRef] [PubMed]
- Kullak-Ublick, G.A.; Beuers, U.; Meier, P.J.; Domdey, H.; Paumgartner, G. Assignment of the human organic anion transporting polypeptide (OATP) gene to chromosome 12p12 by fluorescence in situ hybridization. J. Hepatol. 1996, 25, 985–987. [Google Scholar] [CrossRef]
- König, J.; Cui, Y.; Nies, A.T.; Keppler, D. Localization and genomic organization of a new hepatocellular organic anion transporting polypeptide. J. Biol. Chem. 2000, 275, 23161–23168. [Google Scholar] [CrossRef] [PubMed]
- Konig, J.; Cui, Y.; Nies, A.T.; Keppler, D. A novel human organic anion transporting polypeptide localized to the basolateral hepatocyte membrane. Am. J. Physiol. Gastrointest. Liver Physiol. 2000, 278, G156–G164. [Google Scholar] [CrossRef] [PubMed]
- Ho, R.H.; Tirona, R.G.; Leake, B.F.; Glaeser, H.; Lee, W.; Lemke, C.J.; Wang, Y.; Kim, R.B. Drug and bile acid transporters in rosuvastatin hepatic uptake: Function, expression, and pharmacogenetics. Gastroenterology 2006, 130, 1793–1806. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, T.; Tamai, I. Genetic polymorphisms of OATP transporters and their impact on intestinal absorption and hepatic disposition of drugs. Drug Metab. Pharmacokinet. 2012, 27, 106–121. [Google Scholar] [CrossRef] [PubMed]
- Kellick, K. Organic ion transporters and statin drug interactions. Curr. Atheroscler. Rep. 2017, 19, 65. [Google Scholar] [CrossRef] [PubMed]
- Tweedie, D.; Polli, J.W.; Berglund, E.G.; Huang, S.M.; Zhang, L.; Poirier, A.; Chu, X.; Feng, B.; International Transporter, C. Transporter studies in drug development: Experience to date and follow-up on decision trees from the International Transporter Consortium. Clin. Pharmacol. Ther. 2013, 94, 113–125. [Google Scholar] [CrossRef] [PubMed]
- European Medicines Agency (EMA). Guideline on the Investigation of Drug Interactions; EMA: London, UK, 2012. [Google Scholar]
- Food and Drug Administration (US FDA). In Vitro Metabolism and Transporter Mediated Drug-Drug Interaction Studies Guidance for Industry; Office of Communications, Division of Drug Information Center for Drug Evaluation and Research, Food and Drug Administration: Silver Spring, MD, USA, 2017.
- Vaidyanathan, J.; Yoshida, K.; Arya, V.; Zhang, L. Comparing various in vitro prediction criteria to assess the potential of a new molecular entity to inhibit organic anion transporting polypeptide 1B1. J. Clin. Pharmacol. 2016, 56 (Suppl. S7), S59–S72. [Google Scholar] [CrossRef] [PubMed]
- Hirano, M.; Maeda, K.; Shitara, Y.; Sugiyama, Y. Contribution of OATP2 (OATP1B1) and OATP8 (OATP1B3) to the hepatic uptake of pitavastatin in humans. J. Pharmacol. Exp. Ther. 2004, 311, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, S.; Maeda, K.; Wang, Y.; Sugiyama, Y. Involvement of multiple transporters in the hepatobiliary transport of rosuvastatin. Drug Metab. Dispos. 2008, 36, 2014–2023. [Google Scholar] [CrossRef] [PubMed]
- Vavricka, S.R.; Van Montfoort, J.; Ha, H.R.; Meier, P.J.; Fattinger, K. Interactions of rifamycin SV and rifampicin with organic anion uptake systems of human liver. Hepatology 2002, 36, 164–172. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; König, J.; Leier, I.; Buchholz, U.; Keppler, D. Hepatic uptake of bilirubin and its conjugates by the human organic anion transporter SLC21A6. J. Biol. Chem. 2001, 276, 9626–9630. [Google Scholar] [CrossRef] [PubMed]
- Treiber, A.; Schneiter, R.; Hausler, S.; Stieger, B. Bosentan is a substrate of human OATP1B1 and OATP1B3: Inhibition of hepatic uptake as the common mechanism of its interactions with cyclosporin A, rifampicin, and sildenafil. Drug Metab. Dispos. 2007, 35, 1400–1407. [Google Scholar] [CrossRef] [PubMed]
- Yamashiro, W.; Maeda, K.; Hirouchi, M.; Adachi, Y.; Hu, Z.; Sugiyama, Y. Involvement of transporters in the hepatic uptake and biliary excretion of valsartan, a selective antagonist of the angiotensin II AT1-receptor, in humans. Drug Metab. Dispos. 2006, 34, 1247–1254. [Google Scholar] [CrossRef] [PubMed]
- Yamada, A.; Maeda, K.; Kamiyama, E.; Sugiyama, D.; Kondo, T.; Shiroyanagi, Y.; Nakazawa, H.; Okano, T.; Adachi, M.; Schuetz, J.D.; et al. Multiple human isoforms of drug transporters contribute to the hepatic and renal transport of olmesartan, a selective antagonist of the angiotensin II AT1-receptor. Drug Metab. Dispos. 2007, 35, 2166–2176. [Google Scholar] [CrossRef] [PubMed]
- Niemi, M. Role of OATP transporters in the disposition of drugs. Pharmacogenomics 2007, 8, 787–802. [Google Scholar] [CrossRef] [PubMed]
- Kalliokoski, A.; Niemi, M. Impact of OATP transporters on pharmacokinetics. Br. J. Pharmacol. 2009, 158, 693–705. [Google Scholar] [CrossRef] [PubMed]
- Ishiguro, N.; Maeda, K.; Kishimoto, W.; Saito, A.; Harada, A.; Ebner, T.; Roth, W.; Igarashi, T.; Sugiyama, Y. Predominant contribution of OATP1B3 to the hepatic uptake of telmisartan, an angiotensin II receptor antagonist, in humans. Drug Metab. Dispos. 2006, 34, 1109–1115. [Google Scholar] [CrossRef] [PubMed]
- Kullak-Ublick, G.A.; Ismair, M.G.; Stieger, B.; Landmann, L.; Huber, R.; Pizzagalli, F.; Fattinger, K.; Meier, P.J.; Hagenbuch, B. Organic anion-transporting polypeptide B (OATP-B) and its functional comparison with three other OATPs of human liver. Gastroenterology 2001, 120, 525–533. [Google Scholar] [CrossRef] [PubMed]
- Letschert, K.; Faulstich, H.; Keller, D.; Keppler, D. Molecular characterization and inhibition of amanitin uptake into human hepatocytes. Toxicol. Sci. 2006, 91, 140–149. [Google Scholar] [CrossRef] [PubMed]
- Gozalpour, E.; Greupink, R.; Wortelboer, H.M.; Bilos, A.; Schreurs, M.; Russel, F.G.; Koenderink, J.B. Interaction of digitalis-like compounds with liver uptake transporters NTCP, OATP1B1, and OATP1B3. Mol. Pharm. 2014, 11, 1844–1855. [Google Scholar] [CrossRef] [PubMed]
- Ismair, M.G.; Stieger, B.; Cattori, V.; Hagenbuch, B.; Fried, M.; Meier, P.J.; Kullak-Ublick, G.A. Hepatic uptake of cholecystokinin octapeptide by organic anion-transporting polypeptides OATP4 and OATP8 of rat and human liver. Gastroenterology 2001, 121, 1185–1190. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kullak-Ublick, G.A.; Hagenbuch, B.; Stieger, B.; Schteingart, C.D.; Hofmann, A.F.; Wolkoff, A.W.; Meier, P.J. Molecular and functional characterization of an organic anion transporting polypeptide cloned from human liver. Gastroenterology 1995, 109, 1274–1282. [Google Scholar] [CrossRef]
- König, J. Uptake transporters of the human OATP family: Molecular characteristics, substrates, their role in drug-drug interactions, and functional consequences of polymorphisms. Handb. Exp. Pharmacol. 2011, 1–28. [Google Scholar] [CrossRef]
- Stieger, B.; Hagenbuch, B. Organic anion-transporting polypeptides. Curr. Top. Membr. 2014, 73, 205–232. [Google Scholar] [PubMed]
- DeGorter, M.K.; Ho, R.H.; Leake, B.F.; Tirona, R.G.; Kim, R.B. Interaction of three regiospecific amino acid residues is required for OATP1B1 gain of OATP1B3 substrate specificity. Mol. Pharm. 2012, 9, 986–995. [Google Scholar] [CrossRef] [PubMed]
- Gui, C.; Hagenbuch, B. Amino acid residues in transmembrane domain 10 of organic anion transporting polypeptide 1B3 are critical for cholecystokinin octapeptide transport. Biochemistry 2008, 47, 9090–9097. [Google Scholar] [CrossRef] [PubMed]
- Gui, C.; Hagenbuch, B. Role of transmembrane domain 10 for the function of organic anion transporting polypeptide 1B1. Protein Sci. 2009, 18, 2298–2306. [Google Scholar] [CrossRef] [PubMed]
- Miyagawa, M.; Maeda, K.; Aoyama, A.; Sugiyama, Y. The eighth and ninth transmembrane domains in organic anion transporting polypeptide 1B1 affect the transport kinetics of estrone-3-sulfate and estradiol-17beta-D-glucuronide. J. Pharmacol. Exp. Ther. 2009, 329, 551–557. [Google Scholar] [CrossRef] [PubMed]
- Satlin, L.M.; Amin, V.; Wolkoff, A.W. Organic anion transporting polypeptide mediates organic anion/HCO3- exchange. J. Biol. Chem. 1997, 272, 26340–26345. [Google Scholar] [CrossRef] [PubMed]
- Leuthold, S.; Hagenbuch, B.; Mohebbi, N.; Wagner, C.A.; Meier, P.J.; Stieger, B. Mechanisms of pH-gradient driven transport mediated by organic anion polypeptide transporters. Am. J. Physiol. Cell Physiol. 2009, 296, C570–C582. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Lee, T.K.; Meier, P.J.; Ballatori, N. Identification of glutathione as a driving force and leukotriene C4 as a substrate for oatp1, the hepatic sinusoidal organic solute transporter. J. Biol. Chem. 1998, 273, 16184–16191. [Google Scholar] [CrossRef] [PubMed]
- Briz, O.; Romero, M.R.; Martinez-Becerra, P.; Macias, R.I.; Perez, M.J.; Jimenez, F.; San Martin, F.G.; Marin, J.J. OATP8/1B3-mediated cotransport of bile acids and glutathione: An export pathway for organic anions from hepatocytes? J. Biol. Chem. 2006, 281, 30326–30335. [Google Scholar] [CrossRef] [PubMed]
- Mahagita, C.; Grassl, S.M.; Piyachaturawat, P.; Ballatori, N. Human organic anion transporter 1B1 and 1B3 function as bidirectional carriers and do not mediate GSH-bile acid cotransport. Am. J. Physiol. Gastrointest. Liver Physiol. 2007, 293, G271–G278. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Becerra, P.; Briz, O.; Romero, M.R.; Macias, R.I.; Perez, M.J.; Sancho-Mateo, C.; Lostao, M.P.; Fernandez-Abalos, J.M.; Marin, J.J. Further characterization of the electrogenicity and pH sensitivity of the human organic anion-transporting polypeptides OATP1B1 and OATP1B3. Mol. Pharmacol. 2011, 79, 596–607. [Google Scholar] [CrossRef] [PubMed]
- Tirona, R.G.; Leake, B.F.; Merino, G.; Kim, R.B. Polymorphisms in OATP-C. Identification of multiple allelic variants associated with altered transport activity among European and African-Americans. J. Biol. Chem. 2001, 276, 35669–35675. [Google Scholar] [CrossRef] [PubMed]
- SEARCH Collaborative Group. SLCO1B1 variants and statin-induced myopathy—A genomewide study. N. Engl. J. Med. 2008, 359, 789–799. [Google Scholar]
- Grapci, A.D.; Dimovski, A.J.; Kapedanovska, A.; Vavlukis, M.; Eftimov, A.; Geshkovska, N.M.; Labachevski, N.; Jakjovski, K.; Gorani, D.; Kedev, S.; et al. Frequencies of single-nucleotide polymorphisms and haplotypes of the SLCO1B1 gene in selected populations of the western balkans. Balkan J. Med. Genet. 2015, 18, 5–21. [Google Scholar] [CrossRef] [PubMed]
- Pasanen, M.K.; Fredrikson, H.; Neuvonen, P.J.; Niemi, M. Different effects of SLCO1B1 polymorphism on the pharmacokinetics of atorvastatin and rosuvastatin. Clin. Pharmacol. Ther. 2007, 82, 726–733. [Google Scholar] [CrossRef] [PubMed]
- Niemi, M.; Pasanen, M.K.; Neuvonen, P.J. Organic anion transporting polypeptide 1B1: A genetically polymorphic transporter of major importance for hepatic drug uptake. Pharmacol. Rev. 2011, 63, 157–181. [Google Scholar] [CrossRef] [PubMed]
- Niemi, M.; Kivisto, K.T.; Hofmann, U.; Schwab, M.; Eichelbaum, M.; Fromm, M.F. Fexofenadine pharmacokinetics are associated with a polymorphism of the SLCO1B1 gene (encoding OATP1B1). Br. J. Clin. Pharmacol. 2005, 59, 602–604. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, Y.; Zhao, P.; Huang, S. Predicting drug-drug interactions: An FDA perspective. AAPS J. 2009, 11, 300–306. [Google Scholar] [CrossRef] [PubMed]
- Kohlrausch, F.B.; de Cassia Estrela, R.; Barroso, P.F.; Suarez-Kurtz, G. The impact of SLCO1B1 polymorphisms on the plasma concentration of lopinavir and ritonavir in HIV-infected men. Br. J. Clin. Pharmacol. 2010, 69, 95–98. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; He, Y.; Han, C.; Liu, Z.; Li, Q.; Fan, L.; Tan, Z.; Zhang, W.; Yu, B.; Wang, D.; et al. Effect of SLCO1B1 genetic polymorphism on the pharmacokinetics of nateglinide. Br. J. Clin. Pharmacol. 2006, 62, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Niemi, M.; Pasanen, M.K.; Neuvonen, P.J. SLCO1B1 polymorphism and sex affect the pharmacokinetics of pravastatin but not fluvastatin. Clin. Pharmacol. Ther. 2006, 80, 356–366. [Google Scholar] [CrossRef] [PubMed]
- Kalliokoski, A.; Neuvonen, M.; Neuvonen, P.J.; Niemi, M. The effect of SLCO1B1 polymorphism on repaglinide pharmacokinetics persists over a wide dose range. Br. J. Clin. Pharmacol. 2008, 66, 818–825. [Google Scholar] [CrossRef] [PubMed]
- Hamada, A.; Sissung, T.; Price, D.K.; Danesi, R.; Chau, C.H.; Sharifi, N.; Venzon, D.; Maeda, K.; Nagao, K.; Sparreboom, A.; et al. Effect of SLCO1B3 haplotype on testosterone transport and clinical outcome in caucasian patients with androgen-independent prostatic cancer. Clin. Cancer Res. 2008, 14, 3312–3318. [Google Scholar] [CrossRef] [PubMed]
- Picard, N.; Yee, S.W.; Woillard, J.B.; Lebranchu, Y.; Le Meur, Y.; Giacomini, K.M.; Marquet, P. The role of organic anion-transporting polypeptides and their common genetic variants in mycophenolic acid pharmacokinetics. Clin. Pharmacol. Ther. 2010, 87, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Van de Steeg, E.; Stranecky, V.; Hartmannova, H.; Noskova, L.; Hrebicek, M.; Wagenaar, E.; van Esch, A.; de Waart, D.R.; Oude Elferink, R.P.; Kenworthy, K.E.; et al. Complete OATP1B1 and OATP1B3 deficiency causes human Rotor syndrome by interrupting conjugated bilirubin reuptake into the liver. J. Clin. Investig. 2012, 122, 519–528. [Google Scholar] [CrossRef] [PubMed]
- De Graaf, W.; Hausler, S.; Heger, M.; van Ginhoven, T.M.; van Cappellen, G.; Bennink, R.J.; Kullak-Ublick, G.A.; Hesselmann, R.; van Gulik, T.M.; Stieger, B. Transporters involved in the hepatic uptake of (99m)Tc-mebrofenin and indocyanine green. J. Hepatol. 2011, 54, 738–745. [Google Scholar] [CrossRef] [PubMed]
- Tirona, R.G.; Leake, B.F.; Wolkoff, A.W.; Kim, R.B. Human organic anion transporting polypeptide-C (SLC21A6) is a major determinant of rifampin-mediated pregnane X receptor activation. J. Pharmacol. Exp. Ther. 2003, 304, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Shitara, Y.; Hirano, M.; Sato, H.; Sugiyama, Y. Gemfibrozil and its glucuronide inhibit the organic anion transporting polypeptide 2 (OATP2/OATP1B1:SLC21A6)-mediated hepatic uptake and CYP2C8-mediated metabolism of cerivastatin: Analysis of the mechanism of the clinically relevant drug-drug interaction between cerivastatin and gemfibrozil. J. Pharmacol. Exp. Ther. 2004, 311, 228–236. [Google Scholar] [PubMed]
- Shitara, Y.; Itoh, T.; Sato, H.; Li, A.P.; Sugiyama, Y. Inhibition of transporter-mediated hepatic uptake as a mechanism for drug-drug interaction between cerivastatin and cyclosporin A. J. Pharmacol. Exp. Ther. 2003, 304, 610–616. [Google Scholar] [CrossRef] [PubMed]
- Kiser, J.J.; Gerber, J.G.; Predhomme, J.A.; Wolfe, P.; Flynn, D.M.; Hoody, D.W. Drug/Drug interaction between lopinavir/ritonavir and rosuvastatin in healthy volunteers. J. Acquir. Immune Defic. Syndr. 2008, 47, 570–578. [Google Scholar] [CrossRef] [PubMed]
- Chauvin, B.; Drouot, S.; Barrail-Tran, A.; Taburet, A.M. Drug-drug interactions between HMG-CoA reductase inhibitors (statins) and antiviral protease inhibitors. Clin. Pharmacokinet. 2013, 52, 815–831. [Google Scholar] [CrossRef] [PubMed]
- Olsson, A.G.; McTaggart, F.; Raza, A. Rosuvastatin: A highly effective new HMG-CoA reductase inhibitor. Cardiovasc. Drug Rev. 2002, 20, 303–328. [Google Scholar] [CrossRef] [PubMed]
- Schneck, D.W.; Birmingham, B.K.; Zalikowski, J.A.; Mitchell, P.D.; Wang, Y.; Martin, P.D.; Lasseter, K.C.; Brown, C.D.; Windass, A.S.; Raza, A. The effect of gemfibrozil on the pharmacokinetics of rosuvastatin. Clin. Pharmacol. Ther. 2004, 75, 455–463. [Google Scholar] [CrossRef] [PubMed]
- Shitara, Y.; Takeuchi, K.; Nagamatsu, Y.; Wada, S.; Sugiyama, Y.; Horie, T. Long-lasting Inhibitory Effects of Cyclosporin A, but Not Tacrolimus, on OATP1B1- and OATP1B3-mediated Uptake. Drug Metab. Pharmacokinet. 2012, 27, 368–378. [Google Scholar] [CrossRef] [PubMed]
- Simonson, S.G.; Raza, A.; Martin, P.D.; Mitchell, P.D.; Jarcho, J.A.; Brown, C.D.; Windass, A.S.; Schneck, D.W. Rosuvastatin pharmacokinetics in heart transplant recipients administered an antirejection regimen including cyclosporine. Clin. Pharmacol. Ther. 2004, 76, 167–177. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, K.; Maeda, K.; Sugiyama, Y. Transporter-mediated drug-drug interactions involving OATP substrates: Predictions based on in vitro inhibition studies. Clin. Pharmacol. Ther. 2012, 91, 1053–1064. [Google Scholar] [CrossRef] [PubMed]
- Chu, X.; Cai, X.; Cui, D.; Tang, C.; Ghosal, A.; Chan, G.; Green, M.D.; Kuo, Y.; Liang, Y.; Maciolek, C.M.; et al. In vitro assessment of drug-drug interaction potential of boceprevir associated with drug metabolizing enzymes and transporters. Drug Metab. Dispos. 2013, 41, 668–681. [Google Scholar] [CrossRef] [PubMed]
- Hulskotte, E.G.; Feng, H.P.; Xuan, F.; Gupta, S.; van Zutven, M.G.; O’Mara, E.; Wagner, J.A.; Butterton, J.R. Pharmacokinetic evaluation of the interaction between hepatitis C virus protease inhibitor boceprevir and 3-hydroxy-3-methylglutaryl coenzyme A reductase inhibitors atorvastatin and pravastatin. Antimicrob. Agents Chemother. 2013, 57, 2582–2588. [Google Scholar] [CrossRef] [PubMed]
- Sane, R.S.; Steinmann, G.G.; Huang, Q.; Li, Y.; Podila, L.; Mease, K.; Olson, S.; Taub, M.E.; Stern, J.O.; Nehmiz, G.; et al. Mechanisms underlying benign and reversible unconjugated hyperbilirubinemia observed with faldaprevir administration in hepatitis C virus patients. J. Pharmacol. Exp. Ther. 2014, 351, 403–412. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.; Marzin, K.; Koenen, R.; Kammerer, K.P.; Strelkowa, N.; Elgadi, M.; Quinson, A.M.; Haertter, S. Effect of steady-state faldaprevir on pharmacokinetics of atorvastatin or rosuvastatin in healthy volunteers: A prospective open-label, fixed-sequence crossover study. J. Clin. Pharmacol. 2017, 57, 1305–1314. [Google Scholar] [CrossRef] [PubMed]
- Ayalasomayajula, S.; Han, Y.; Langenickel, T.; Malcolm, K.; Zhou, W.; Hanna, I.; Alexander, N.; Natrillo, A.; Goswami, B.; Hinder, M.; et al. In vitro and clinical evaluation of OATP-mediated drug interaction potential of sacubitril/valsartan (LCZ696). J. Clin. Pharm. Ther. 2016, 41, 424–431. [Google Scholar] [CrossRef] [PubMed]
- Hanna, I.; Alexander, N.; Crouthamel, M.H.; Davis, J.; Natrillo, A.; Tran, P.; Vapurcuyan, A.; Zhu, B. Transport properties of valsartan, sacubitril and its active metabolite (LBQ657) as determinants of disposition. Xenobiotica 2018, 48, 300–313. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Ji, T.; Einolf, H.; Ayalasomayajula, S.; Lin, T.H.; Hanna, I.; Heimbach, T.; Breen, C.; Jarugula, V.; He, H. Evaluation of drug-drug interaction potential between sacubitril/valsartan (LCZ696) and statins using a physiologically based pharmacokinetic model. J. Pharm. Sci. 2017, 106, 1439–1451. [Google Scholar] [CrossRef] [PubMed]
- Janssen Pharmaceuticals, Inc. OLYSIO (Simeprevir) Capsules: Package Insert; Janssen Pharmaceuticals, Inc.: Titusville, NJ, USA, 2013. [Google Scholar]
- Ouwerkerk-Mahadevan, S.; Snoeys, J.; Peeters, M.; Beumont-Mauviel, M.; Simion, A. Drug-drug interactions with the NS3/4A protease inhibitor simeprevir. Clin. Pharmacokinet. 2016, 55, 197–208. [Google Scholar] [CrossRef] [PubMed]
- Kunze, A.; Huwyler, J.; Camenisch, G.; Gutmann, H. Interaction of the antiviral drug telaprevir with renal and hepatic drug transporters. Biochem. Pharmacol. 2012, 84, 1096–1102. [Google Scholar] [CrossRef] [PubMed]
- Shin, K.H.; Kim, T.E.; Kim, S.E.; Lee, M.G.; Song, I.S.; Yoon, S.H.; Cho, J.Y.; Jang, I.J.; Shin, S.G.; Yu, K.S. The effect of the newly developed angiotensin receptor II antagonist fimasartan on the pharmacokinetics of atorvastatin in relation to OATP1B1 in healthy male volunteers. J. Cardiovasc. Pharmacol. 2011, 58, 492–499. [Google Scholar] [CrossRef] [PubMed]
- Hirano, M.; Maeda, K.; Shitara, Y.; Sugiyama, Y. Drug-drug interaction between pitavastatin and various drugs via OATP1B1. Drug Metab. Dispos. 2006, 34, 1229–1236. [Google Scholar] [CrossRef] [PubMed]
- Karlgren, M.; Vildhede, A.; Norinder, U.; Wisniewski, J.R.; Kimoto, E.; Lai, Y.; Haglund, U.; Artursson, P. Classification of inhibitors of hepatic organic anion transporting polypeptides (OATPs): Influence of protein expression on drug-drug interactions. J. Med. Chem. 2012, 55, 4740–4763. [Google Scholar] [CrossRef] [PubMed]
- Pham, P.A.; la Porte, C.J.; Lee, L.S.; van Heeswijk, R.; Sabo, J.P.; Elgadi, M.M.; Piliero, P.J.; Barditch-Crovo, P.; Fuchs, E.; Flexner, C.; et al. Differential effects of tipranavir plus ritonavir on atorvastatin or rosuvastatin pharmacokinetics in healthy volunteers. Antimicrob. Agents Chemother. 2009, 53, 4385–4392. [Google Scholar] [CrossRef] [PubMed]
- Pahwa, S.; Alam, K.; Crowe, A.; Farasyn, T.; Neuhoff, S.; Hatley, O.; Ding, K.; Yue, W. Pretreatment with rifampicin and tyrosine kinase inhibitor dasatinib potentiates the inhibitory effects toward OATP1B1- and OATP1B3-mediated transport. J. Pharm. Sci. 2017, 106, 2123–2135. [Google Scholar] [CrossRef] [PubMed]
- Prueksaritanont, T.; Chu, X.; Evers, R.; Klopfer, S.O.; Caro, L.; Kothare, P.A.; Dempsey, C.; Rasmussen, S.; Houle, R.; Chan, G.; et al. Pitavastatin is a more sensitive and selective organic anion-transporting polypeptide 1B clinical probe than rosuvastatin. Br. J. Clin. Pharmacol. 2014, 78, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhang, W.; Huang, W.; Tan, Z.; Wang, Y.; Huang, X.; Zhou, H. Effect of a single-dose rifampin on the pharmacokinetics of pitavastatin in healthy volunteers. Eur. J. Clin. Pharmacol. 2013, 69, 1933–1938. [Google Scholar] [CrossRef] [PubMed]
- Furihata, T.; Matsumoto, S.; Fu, Z.; Tsubota, A.; Sun, Y.; Matsumoto, S.; Kobayashi, K.; Chiba, K. Different interaction profiles of direct-acting anti-hepatitis C virus agents with human organic anion transporting polypeptides. Antimicrob. Agents Chemother. 2014, 58, 4555–4564. [Google Scholar] [CrossRef] [PubMed]
- Garimella, T.; Tao, X.; Sims, K.; Chang, Y.T.; Rana, J.; Myers, E.; Wind-Rotolo, M.; Bhatnagar, R.; Eley, T.; LaCreta, F.; et al. Effects of a fixed-dose co-formulation of daclatasvir, asunaprevir, and beclabuvir on the pharmacokinetics of a cocktail of cytochrome P450 and drug transporter substrates in healthy subjects. Drugs R&D 2017, 18, 55–65. [Google Scholar]
- Annaert, P.; Ye, Z.W.; Stieger, B.; Augustijns, P. Interaction of HIV protease inhibitors with OATP1B1, 1B3, and 2B1. Xenobiotica 2010, 40, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Aquilante, C.L.; Kiser, J.J.; Anderson, P.L.; Christians, U.; Kosmiski, L.A.; Daily, E.B.; Hoffman, K.L.; Hopley, C.W.; Predhomme, J.A.; Schniedewind, B.; et al. Influence of SLCO1B1 polymorphisms on the drug-drug interaction between darunavir/ritonavir and pravastatin. J. Clin. Pharmacol. 2012, 52, 1725–1738. [Google Scholar] [CrossRef] [PubMed]
- Shebley, M.; Liu, J.; Kavetskaia, O.; Sydor, J.; de Morais, S.M.; Fischer, V.; Nijsen, M.; Bow, D.A.J. Mechanisms and predictions of drug-drug interactions of the hepatitis C virus three direct-acting antiviral regimen: Paritaprevir/ritonavir, ombitasvir, and dasabuvir. Drug Metab. Dispos. 2017, 45, 755–764. [Google Scholar] [CrossRef] [PubMed]
- Menon, R.M.; Badri, P.S.; Wang, T.; Polepally, A.R.; Zha, J.; Khatri, A.; Wang, H.; Hu, B.; Coakley, E.P.; Podsadecki, T.J.; et al. Drug-drug interaction profile of the all-oral anti-hepatitis C virus regimen of paritaprevir/ritonavir, ombitasvir, and dasabuvir. J. Hepatol. 2015, 63, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Tamraz, B.; Fukushima, H.; Wolfe, A.R.; Kaspera, R.; Totah, R.A.; Floyd, J.S.; Ma, B.; Chu, C.; Marciante, K.D.; Heckbert, S.R.; et al. OATP1B1-related drug-drug and drug-gene interactions as potential risk factors for cerivastatin-induced rhabdomyolysis. Pharmacogenet. Genom. 2013, 23, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.J.; Yoshikado, T.; Ieiri, I.; Maeda, K.; Kimura, M.; Irie, S.; Kusuhara, H.; Sugiyama, Y. Clarification of the mechanism of clopidogrel-mediated drug-drug interaction in a clinical cassette small-dose study and its prediction based on in vitro information. Drug Metab. Dispos. 2016, 44, 1622–1632. [Google Scholar] [CrossRef] [PubMed]
- Nakagomi-Hagihara, R.; Nakai, D.; Tokui, T.; Abe, T.; Ikeda, T. Gemfibrozil and its glucuronide inhibit the hepatic uptake of pravastatin mediated by OATP1B1. Xenobiotica 2007, 37, 474–486. [Google Scholar] [CrossRef] [PubMed]
- Noe, J.; Portmann, R.; Brun, M.E.; Funk, C. Substrate-dependent drug-drug interactions between gemfibrozil, fluvastatin and other organic anion-transporting peptide (OATP) substrates on OATP1B1, OATP2B1, and OATP1B3. Drug Metab. Dispos. 2007, 35, 1308–1314. [Google Scholar] [CrossRef] [PubMed]
- Honkalammi, J.; Niemi, M.; Neuvonen, P.J.; Backman, J.T. Dose-dependent interaction between gemfibrozil and repaglinide in humans: Strong inhibition of CYP2C8 with subtherapeutic gemfibrozil doses. Drug Metab. Dispos. 2011, 39, 1977–1986. [Google Scholar] [CrossRef] [PubMed]
- Honkalammi, J.; Niemi, M.; Neuvonen, P.J.; Backman, J.T. Gemfibrozil is a strong inactivator of CYP2C8 in very small multiple doses. Clin. Pharmacol. Ther. 2012, 91, 846–855. [Google Scholar] [CrossRef] [PubMed]
- Yoshikado, T.; Maeda, K.; Furihata, S.; Terashima, H.; Nakayama, T.; Ishigame, K.; Tsunemoto, K.; Kusuhara, H.; Furihata, K.I.; Sugiyama, Y. A clinical cassette dosing study for evaluating the contribution of hepatic OATPs and CYP3A to drug-drug interactions. Pharm. Res. 2017, 34, 1570–1583. [Google Scholar] [CrossRef] [PubMed]
- Custodio, J.M.; Wang, H.; Hao, J.; Lepist, E.I.; Ray, A.S.; Andrews, J.; Ling, K.H.; Cheng, A.; Kearney, B.P.; Ramanathan, S. Pharmacokinetics of cobicistat boosted-elvitegravir administered in combination with rosuvastatin. J. Clin. Pharmacol. 2014, 54, 649–656. [Google Scholar] [CrossRef] [PubMed]
- Elsby, R.; Martin, P.; Surry, D.; Sharma, P.; Fenner, K. Solitary inhibition of the breast cancer resistance protein efflux transporter results in a clinically significant drug-drug interaction with rosuvastatin by causing up to a 2-fold increase in statin exposure. Drug Metab. Dispos. 2016, 44, 398–408. [Google Scholar] [CrossRef] [PubMed]
- Martin, P.; Gillen, M.; Ritter, J.; Mathews, D.; Brealey, C.; Surry, D.; Oliver, S.; Holmes, V.; Severin, P.; Elsby, R. Effects of fostamatinib on the pharmacokinetics of oral contraceptive, warfarin, and the statins rosuvastatin and simvastatin: Results from phase I clinical studies. Drugs R&D 2016, 16, 93–107. [Google Scholar]
- Ebner, T.; Ishiguro, N.; Taub, M.E. The use of transporter probe drug cocktails for the assessment of transporter-based drug-drug interactions in a clinical setting-proposal of a four component transporter cocktail. J. Pharm. Sci. 2015, 104, 3220–3228. [Google Scholar] [CrossRef] [PubMed]
- Stopfer, P.; Giessmann, T.; Hohl, K.; Sharma, A.; Ishiguro, N.; Taub, M.E.; Zimdahl-Gelling, H.; Gansser, D.; Wein, M.; Ebner, T.; et al. Pharmacokinetic evaluation of a drug transporter cocktail consisting of digoxin, furosemide, metformin, and rosuvastatin. Clin. Pharmacol. Ther. 2016, 100, 259–267. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.F.; Hristeva, N.; Chang, J.; Liang, X.; Li, R.; Frassetto, L.; Benet, L.Z. Rosuvastatin pharmacokinetics in Asian and white subjects wild type for both OATP1B1 and BCRP under control and inhibited conditions. J. Pharm. Sci. 2017, 106, 2751–2757. [Google Scholar] [CrossRef] [PubMed]
- Lai, Y.; Mandlekar, S.; Shen, H.; Holenarsipur, V.K.; Langish, R.; Rajanna, P.; Murugesan, S.; Gaud, N.; Selvam, S.; Date, O.; et al. Coproporphyrins in plasma and urine can be appropriate clinical biomarkers to recapitulate drug-drug interactions mediated by organic anion transporting polypeptide inhibition. J. Pharmacol. Exp. Ther. 2016, 358, 397–404. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Huo, X.; Wang, C.; Meng, Q.; Liu, Z.; Sun, P.; Cang, J.; Sun, H.; Liu, K. Organic anion-transporting polypeptide and efflux transporter-mediated hepatic uptake and biliary excretion of cilostazol and its metabolites in rats and humans. J. Pharm. Sci. 2017, 106, 2515–2523. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Mathijssen, R.H.; de Bruijn, P.; Baker, S.D.; Sparreboom, A. Inhibition of OATP1B1 by tyrosine kinase inhibitors: In vitro-in vivo correlations. Br. J. Cancer 2014, 110, 894–898. [Google Scholar] [CrossRef] [PubMed]
- Pressler, H.; Sissung, T.M.; Venzon, D.; Price, D.K.; Figg, W.D. Expression of OATP family members in hormone-related cancers: Potential markers of progression. PLoS ONE 2011, 6, e20372. [Google Scholar] [CrossRef] [PubMed]
- Vander Borght, S.; Libbrecht, L.; Blokzijl, H.; Faber, K.N.; Moshage, H.; Aerts, R.; Van Steenbergen, W.; Jansen, P.L.; Desmet, V.J.; Roskams, T.A. Diagnostic and pathogenetic implications of the expression of hepatic transporters in focal lesions occurring in normal liver. J. Pathol. 2005, 207, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Vavricka, S.R.; Jung, D.; Fried, M.; Grutzner, U.; Meier, P.J.; Kullak-Ublick, G.A. The human organic anion transporting polypeptide 8 (SLCO1B3) gene is transcriptionally repressed by hepatocyte nuclear factor 3beta in hepatocellular carcinoma. J. Hepatol. 2004, 40, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Tsuboyama, T.; Onishi, H.; Kim, T.; Akita, H.; Hori, M.; Tatsumi, M.; Nakamoto, A.; Nagano, H.; Matsuura, N.; Wakasa, K.; et al. Hepatocellular carcinoma: Hepatocyte-selective enhancement at gadoxetic acid-enhanced MR imaging—Correlation with expression of sinusoidal and canalicular transporters and bile accumulation. Radiology 2010, 255, 824–833. [Google Scholar] [CrossRef] [PubMed]
- Nagai, M.; Furihata, T.; Matsumoto, S.; Ishii, S.; Motohashi, S.; Yoshino, I.; Ugajin, M.; Miyajima, A.; Matsumoto, S.; Chiba, K. Identification of a new organic anion transporting polypeptide 1B3 mRNA isoform primarily expressed in human cancerous tissues and cells. Biochem. Biophys. Res. Commun. 2012, 418, 818–823. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Furihata, T.; Ishii, S.; Nagai, M.; Harada, M.; Shimozato, O.; Kamijo, T.; Motohashi, S.; Yoshino, I.; Kamiichi, A.; et al. Unique expression features of cancer-type organic anion transporting polypeptide 1B3 mRNA expression in human colon and lung cancers. Clin. Transl. Med. 2014, 3, 37. [Google Scholar] [CrossRef] [PubMed]
- Thakkar, N.; Kim, K.; Jang, E.R.; Han, S.; Kim, K.; Kim, D.; Merchant, N.; Lockhart, A.C.; Lee, W. A cancer-specific variant of the SLCO1B3 gene encodes a novel human organic anion transporting polypeptide 1B3 (OATP1B3) localized mainly in the cytoplasm of colon and pancreatic cancer cells. Mol. Pharm. 2013, 10, 406–416. [Google Scholar] [CrossRef] [PubMed]
- Chun, S.E.; Thakkar, N.; Oh, Y.; Park, J.E.; Han, S.; Ryoo, G.; Hahn, H.; Maeng, S.H.; Lim, Y.R.; Han, B.W.; et al. The N-terminal region of organic anion transporting polypeptide 1B3 (OATP1B3) plays an essential role in regulating its plasma membrane trafficking. Biochem. Pharmacol. 2017, 131, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Thomson, M.M.; Hines, R.N.; Schuetz, E.G.; Meibohm, B. Expression patterns of organic anion transporting polypeptides 1B1 and 1B3 protein in human pediatric liver. Drug Metab. Dispos. 2016, 44, 999–1004. [Google Scholar] [CrossRef] [PubMed]
- Mooij, M.G.; Schwarz, U.I.; de Koning, B.A.; Leeder, J.S.; Gaedigk, R.; Samsom, J.N.; Spaans, E.; van Goudoever, J.B.; Tibboel, D.; Kim, R.B.; et al. Ontogeny of human hepatic and intestinal transporter gene expression during childhood: Age matters. Drug Metab. Dispos. 2014, 42, 1268–1274. [Google Scholar] [CrossRef] [PubMed]
- Jung, D.; Hagenbuch, B.; Gresh, L.; Pontoglio, M.; Meier, P.J.; Kullak-Ublick, G.A. Characterization of the human OATP-C (SLC21A6) gene promoter and regulation of liver-specific OATP genes by hepatocyte nuclear factor 1alpha. J. Biol. Chem. 2001, 276, 37206–37214. [Google Scholar] [CrossRef] [PubMed]
- Kamiyama, Y.; Matsubara, T.; Yoshinari, K.; Nagata, K.; Kamimura, H.; Yamazoe, Y. Role of human hepatocyte nuclear factor 4alpha in the expression of drug-metabolizing enzymes and transporters in human hepatocytes assessed by use of small interfering RNA. Drug Metab. Pharmacokinet. 2007, 22, 287–298. [Google Scholar] [CrossRef] [PubMed]
- Meyer Zu Schwabedissen, H.E.; Bottcher, K.; Chaudhry, A.; Kroemer, H.K.; Schuetz, E.G.; Kim, R.B. Liver X receptor alpha and farnesoid X receptor are major transcriptional regulators of OATP1B1. Hepatology 2010, 52, 1797–1807. [Google Scholar] [CrossRef] [PubMed]
- Wood, M.; Ananthanarayanan, M.; Jones, B.; Wooton-Kee, R.; Hoffman, T.; Suchy, F.J.; Vore, M. Hormonal regulation of hepatic organic anion transporting polypeptides. Mol. Pharmacol. 2005, 68, 218–225. [Google Scholar] [CrossRef] [PubMed]
- Jigorel, E.; Le Vee, M.; Boursier-Neyret, C.; Parmentier, Y.; Fardel, O. Differential regulation of sinusoidal and canalicular hepatic drug transporter expression by xenobiotics activating drug-sensing receptors in primary human hepatocytes. Drug. Metab. Dispos. 2006, 34, 1756–1763. [Google Scholar] [CrossRef] [PubMed]
- Le Vee, M.; Jouan, E.; Stieger, B.; Fardel, O. Differential regulation of drug transporter expression by all-trans retinoic acid in hepatoma HepaRG cells and human hepatocytes. Eur. J. Pharm. Sci. 2013, 48, 767–774. [Google Scholar] [CrossRef] [PubMed]
- Ichihara, S.; Kikuchi, R.; Kusuhara, H.; Imai, S.; Maeda, K.; Sugiyama, Y. DNA methylation profiles of organic anion transporting polypeptide 1B3 in cancer cell lines. Pharm. Res. 2010, 27, 510–516. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Kim, K.; Thakkar, N.; Kim, D.; Lee, W. Role of hypoxia inducible factor-1alpha in the regulation of the cancer-specific variant of organic anion transporting polypeptide 1B3 (OATP1B3), in colon and pancreatic cancer. Biochem. Pharmacol. 2013, 86, 816–823. [Google Scholar] [CrossRef] [PubMed]
- Yao, J.; Hong, W.; Huang, J.; Zhan, K.; Huang, H.; Hong, M. N-Glycosylation dictates proper processing of organic anion transporting polypeptide 1B1. PLoS ONE 2012, 7, e52563. [Google Scholar] [CrossRef] [PubMed]
- Murray, M.; Zhou, F. Trafficking and other regulatory mechanisms for organic anion transporting polypeptides and organic anion transporters that modulate cellular drug and xenobiotic influx and that are dysregulated in disease. Br. J. Pharmacol. 2017, 174, 1908–1924. [Google Scholar] [CrossRef] [PubMed]
- Traub, L.M.; Kornfeld, S. The trans-Golgi network: A late secretory sorting station. Curr. Opin. Cell Biol. 1997, 9, 527–533. [Google Scholar] [CrossRef]
- Opat, A.S.; Houghton, F.; Gleeson, P.A. Steady-state localization of a medial-Golgi glycosyltransferase involves transit through the trans-Golgi network. Biochem. J. 2001, 358, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Clarke, J.D.; Novak, P.; Lake, A.D.; Hardwick, R.N.; Cherrington, N.J. Impaired N-linked glycosylation of uptake and efflux transporters in human non-alcoholic fatty liver disease. Liver Int. 2017, 37, 1074–1081. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, U.I.; Meyer zu Schwabedissen, H.E.; Tirona, R.G.; Suzuki, A.; Leake, B.F.; Mokrab, Y.; Mizuguchi, K.; Ho, R.H.; Kim, R.B. Identification of novel functional organic anion-transporting polypeptide 1B3 polymorphisms and assessment of substrate specificity. Pharmacogenet. Genom. 2011, 21, 103–114. [Google Scholar] [CrossRef] [PubMed]
- You, G.; Kuze, K.; Kohanski, R.A.; Amsler, K.; Henderson, S. Regulation of mOAT-mediated organic anion transport by okadaic acid and protein kinase C in LLC-PK(1) cells. J. Biol. Chem. 2000, 275, 10278–10284. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, K.; Terada, T.; Asaka, J.; Katsura, T.; Inui, K. Human organic anion transporter 3 gene is regulated constitutively and inducibly via a cAMP-response element. J. Pharmacol. Exp. Ther. 2006, 319, 317–322. [Google Scholar] [CrossRef] [PubMed]
- Kock, K.; Koenen, A.; Giese, B.; Fraunholz, M.; May, K.; Siegmund, W.; Hammer, E.; Volker, U.; Jedlitschky, G.; Kroemer, H.K.; et al. Rapid modulation of the organic anion transporting polypeptide 2B1 (OATP2B1, SLCO2B1) function by protein kinase C-mediated internalization. J. Biol. Chem. 2010, 285, 11336–11347. [Google Scholar] [CrossRef] [PubMed]
- Chai, J.; Cai, S.; Liu, X.; Lian, W.; Chen, S.; Zhang, L.; Feng, X.; Cheng, Y.; He, X.; He, Y.; et al. Canalicular membrane MRP2/ABCC2 internalization is determined by Ezrin Thr567 phosphorylation in human obstructive cholestasis. J. Hepatol. 2015, 63, 1440–1448. [Google Scholar] [CrossRef] [PubMed]
- Chambers, T.C.; Pohl, J.; Raynor, R.L.; Kuo, J.F. Identification of specific sites in human P-glycoprotein phosphorylated by protein kinase C. J. Biol. Chem. 1993, 268, 4592–4595. [Google Scholar] [PubMed]
- Bian, Y.; Song, C.; Cheng, K.; Dong, M.; Wang, F.; Huang, J.; Sun, D.; Wang, L.; Ye, M.; Zou, H. An enzyme assisted RP-RPLC approach for in-depth analysis of human liver phosphoproteome. J. Proteom. 2014, 96, 253–262. [Google Scholar] [CrossRef] [PubMed]
- Powell, J.; Farasyn, T.; Kock, K.; Meng, X.; Pahwa, S.; Brouwer, K.L.; Yue, W. Novel mechanism of impaired function of organic anion-transporting polypeptide 1B3 in human hepatocytes: Post-translational regulation of OATP1B3 by protein kinase C activation. Drug Metab. Dispos. 2014, 42, 1964–1970. [Google Scholar] [CrossRef] [PubMed]
- Hong, M.; Hong, W.; Ni, C.; Huang, J.; Zhou, C. Protein kinase C affects the internalization and recycling of organic anion transporting polypeptide 1B1. Biochim. Biophys. Acta 2015, 1848, 2022–2030. [Google Scholar] [CrossRef] [PubMed]
- Pickart, C.M.; Fushman, D. Polyubiquitin chains: Polymeric protein signals. Curr. Opin. Chem. Biol. 2004, 8, 610–616. [Google Scholar] [CrossRef] [PubMed]
- Ashida, H.; Kim, M.; Sasakawa, C. Exploitation of the host ubiquitin system by human bacterial pathogens. Nat. Rev. Microbiol. 2014, 12, 399–413. [Google Scholar] [CrossRef] [PubMed]
- Hershko, A.; Ciechanover, A. The ubiquitin system for protein degradation. Annu. Rev. Biochem. 1992, 61, 761–807. [Google Scholar] [CrossRef] [PubMed]
- Alam, K.; Farasyn, T.; Crowe, A.; Ding, K.; Yue, W. Treatment with proteasome inhibitor bortezomib decreases organic anion transporting polypeptide (OATP) 1B3-mediated transport in a substrate-dependent manner. PLoS ONE 2017, 12, e0186924. [Google Scholar] [CrossRef] [PubMed]
- Avci, D.; Lemberg, M.K. Clipping or extracting: Two ways to membrane protein degradation. Trends Cell Biol. 2015, 25, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Ohsumi, Y. Protein turnover. IUBMB Life 2006, 58, 363–369. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Roundtree, M.; Merikhi, A.; Lu, X.; Shentu, S.; Lesage, G. Degradation of the apical sodium-dependent bile acid transporter by the ubiquitin-proteasome pathway in cholangiocytes. J. Biol. Chem. 2004, 279, 44931–44937. [Google Scholar] [CrossRef] [PubMed]
- Jeffers, M.; Taylor, G.A.; Weidner, K.M.; Omura, S.; Vande Woude, G.F. Degradation of the Met tyrosine kinase receptor by the ubiquitin-proteasome pathway. Mol. Cell. Biol. 1997, 17, 799–808. [Google Scholar] [CrossRef] [PubMed]
- Kuhlkamp, T.; Keitel, V.; Helmer, A.; Haussinger, D.; Kubitz, R. Degradation of the sodium taurocholate cotransporting polypeptide (NTCP) by the ubiquitin-proteasome system. Biol. Chem. 2005, 386, 1065–1074. [Google Scholar] [CrossRef] [PubMed]
- Rezvani, K.; Teng, Y.; de Biasi, M. The ubiquitin-proteasome system regulates the stability of neuronal nicotinic acetylcholine receptors. J. Mol. Neurosci. 2010, 40, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Pons, V.; Serhan, N.; Gayral, S.; Malaval, C.; Nauze, M.; Malet, N.; Laffargue, M.; Gales, C.; Martinez, L.O. Role of the ubiquitin-proteasome system in the regulation of P2Y13 receptor expression: Impact on hepatic HDL uptake. Cell. Mol. Life Sci. 2014, 71, 1775–1788. [Google Scholar] [CrossRef] [PubMed]
- Cooper, G.M. The Cell: A Molecular Approach, 2nd ed.; Sinauer Associates: Sunderland, MA, USA, 2000. [Google Scholar]
- Sanofi-Aventis. Aralen® (Chloroquine Phosphate) Tablets Prescribing Information. 2013. Available online: http://www.accessdata.fda.gov/drugsatfda_docs/label/2013/006002s043lbl.pdf (accessed on 20 December 2014).
- Qin, H.; Shao, Q.; Igdoura, S.A.; Alaoui-Jamali, M.A.; Laird, D.W. Lysosomal and proteasomal degradation play distinct roles in the life cycle of Cx43 in gap junctional intercellular communication-deficient and -competent breast tumor cells. J. Biol. Chem. 2003, 278, 30005–30014. [Google Scholar] [CrossRef] [PubMed]
- Hsin, I.L.; Sheu, G.T.; Jan, M.S.; Sun, H.L.; Wu, T.C.; Chiu, L.Y.; Lue, K.H.; Ko, J.L. Inhibition of lysosome degradation on autophagosome formation and responses to GMI, an immunomodulatory protein from Ganoderma microsporum. Br. J. Pharmacol. 2012, 167, 1287–1300. [Google Scholar] [CrossRef] [PubMed]
- Rainsford, K.D.; Parke, A.L.; Clifford-Rashotte, M.; Kean, W.F. Therapy and pharmacological properties of hydroxychloroquine and chloroquine in treatment of systemic lupus erythematosus, rheumatoid arthritis and related diseases. Inflammopharmacology 2015, 23, 231–269. [Google Scholar] [CrossRef] [PubMed]
- Ben-Zvi, I.; Kivity, S.; Langevitz, P.; Shoenfeld, Y. Hydroxychloroquine: From malaria to autoimmunity. Clin. Rev. Allergy Immunol. 2012, 42, 145–153. [Google Scholar] [CrossRef] [PubMed]
- Bezerra, E.L.; Vilar, M.J.; da Trindade Neto, P.B.; Sato, E.I. Double-blind, randomized, controlled clinical trial of clofazimine compared with chloroquine in patients with systemic lupus erythematosus. Arthritis Rheum. 2005, 52, 3073–3078. [Google Scholar] [CrossRef] [PubMed]
- Meier, F.M.; Frerix, M.; Hermann, W.; Muller-Ladner, U. Current immunotherapy in rheumatoid arthritis. Immunotherapy 2013, 5, 955–974. [Google Scholar] [CrossRef] [PubMed]
- Avloclor Tablets-Summary of Product Characteristics. Electronic Medicines Compendium. Avloclor® Tablets-Summary of Product Characteristics. 2014. Available online: http://www.medicines.org.uk/emc/medicine/2272 (accessed on 6 January 2015).
- Inova Health Care Services. Study of the Efficacy of Chloroquine in the Treatment of Ductal Carcinoma in Situ (The PINC Trial). 2009. Available online: https://clinicaltrials.gov/ct2/show/study/NCT01023477 (accessed on 15 January 2015).
- Maastricht Radiation Oncology. Chloroquine as an Anti-Autophagic Radiosensitizing Drug in Stage I–III Small Cell Lung Cancer. 2012. Available online: https://clinicaltrials.gov/ct2/show/NCT01575782 (accessed on 15 January 2015).
- Maastricht Radiation Oncology. Chloroquine as an Anti-Autophagy Drug in Stage IV Small Cell Lung Cancer (SCLC) Patients (Chloroquine IV). 2009. Available online: https://clinicaltrials.gov/ct2/show/NCT00969306 (accessed on 15 January 2015).
- The Methodist Hospital System. Chloroquine With Taxane Chemotherapy for Advanced or Metastatic Breast Cancer Patients Who Have Failed an Anthracycline (CAT). 2011. Available online: https://clinicaltrials.gov/ct2/show/NCT01446016 (accessed on 15 January 2015).
- University of Cincinnati. Chloroquine in Combination With Carboplatin/Gemcitabine in Advanced Solid Tumors. 2014. Available online: https://clinicaltrials.gov/ct2/show/NCT02071537 (accessed on 15 January 2015).
- Solomon, V.R.; Lee, H. Chloroquine and its analogs: A new promise of an old drug for effective and safe cancer therapies. Eur. J. Pharmacol. 2009, 625, 220–233. [Google Scholar] [CrossRef] [PubMed]
- Alam, K.; Pahwa, S.; Wang, X.; Zhang, P.; Ding, K.; Abuznait, A.H.; Li, L.; Yue, W. Downregulation of organic anion transporting polypeptide (OATP) 1B1 transport function by lysosomotropic drug chloroquine: Implication in oatp-mediated drug-drug interactions. Mol. Pharm. 2016, 13, 839–851. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Zhu, L.; Chan, T.; Lu, X.; Shen, W.; Madigan, M.C.; Gillies, M.C.; Zhou, F. Chloroquine and Hydroxychloroquine are novel inhibitors of human organic anion transporting polypeptide 1A2. J. Pharm. Sci. 2016, 105, 884–890. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Hershko, A.; Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 1998, 67, 425–479. [Google Scholar] [CrossRef] [PubMed]
- Brun, J.; Gray, D.A. Targeting the ubiquitin proteasome pathway for the treatment of septic shock in patients. Crit. Care 2009, 13, 311. [Google Scholar] [CrossRef] [PubMed]
- Attaix, D.; Ventadour, S.; Codran, A.; Bechet, D.; Taillandier, D.; Combaret, L. The ubiquitin-proteasome system and skeletal muscle wasting. Essays Biochem. 2005, 41, 173–186. [Google Scholar] [CrossRef] [PubMed]
- Paul, S. Dysfunction of the ubiquitin-proteasome system in multiple disease conditions: Therapeutic approaches. Bioessays 2008, 30, 1172–1184. [Google Scholar] [CrossRef] [PubMed]
- Shen, M.; Schmitt, S.; Buac, D.; Dou, Q.P. Targeting the ubiquitin-proteasome system for cancer therapy. Expert Opin. Ther. Targets 2013, 17, 1091–1108. [Google Scholar] [CrossRef] [PubMed]
- Radwan, M.; Wilkinson, D.J.; Hui, W.; Destrument, A.P.; Charlton, S.H.; Barter, M.J.; Gibson, B.; Coulombe, J.; Gray, D.A.; Rowan, A.D.; et al. Protection against murine osteoarthritis by inhibition of the 26S proteasome and lysine-48 linked ubiquitination. Ann. Rheum. Dis. 2015, 74, 1580–1587. [Google Scholar] [CrossRef] [PubMed]
- Allende-Vega, N.; Saville, M.K. Targeting the ubiquitin-proteasome system to activate wild-type p53 for cancer therapy. Semin. Cancer Biol. 2010, 20, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Cvek, B.; Dvorak, Z. The ubiquitin-proteasome system (UPS) and the mechanism of action of bortezomib. Curr. Pharm. Des. 2011, 17, 1483–1499. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Quinney, S.K.; Wang, Z.; Zhang, P.; Duke, J.; Desta, Z.; Elmendorf, J.S.; Flockhart, D.A.; Li, L. Identification and Mechanistic investigation of drug-drug interactions associated with myopathy: A translational approach. Clin. Pharmacol. Ther. 2015, 98, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Raschi, E.; Poluzzi, E.; Koci, A.; Salvo, F.; Pariente, A.; Biselli, M.; Moretti, U.; Moore, N.; De Ponti, F. Liver injury with novel oral anticoagulants: Assessing post-marketing reports in the US Food and Drug Administration adverse event reporting system. Br. J. Clin. Pharmacol. 2015, 80, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Baker, W.L.; Datta, R. Pitavastatin: A new 3-hydroxy-3-methylglutaryl coenzyme a reductase inhibitor for the treatment of hyperlipidemia. Adv. Ther. 2011, 28, 13–27. [Google Scholar] [CrossRef] [PubMed]
- Jacobsen, W.; Kirchner, G.; Hallensleben, K.; Mancinelli, L.; Deters, M.; Hackbarth, I.; Benet, L.Z.; Sewing, K.F.; Christians, U. Comparison of cytochrome P-450-dependent metabolism and drug interactions of the 3-hydroxy-3-methylglutaryl-CoA reductase inhibitors lovastatin and pravastatin in the liver. Drug Metab. Dispos. 1999, 27, 173–179. [Google Scholar] [PubMed]
- Gertz, M.; Cartwright, C.M.; Hobbs, M.J.; Kenworthy, K.E.; Rowland, M.; Houston, J.B.; Galetin, A. Cyclosporine inhibition of hepatic and intestinal CYP3A4, uptake and efflux transporters: Application of PBPK modeling in the assessment of drug-drug interaction potential. Pharm. Res. 2013, 30, 761–780. [Google Scholar] [CrossRef] [PubMed]
- Izumi, S.; Nozaki, Y.; Maeda, K.; Komori, T.; Takenaka, O.; Kusuhara, H.; Sugiyama, Y. Investigation of the impact of substrate selection on in vitro organic anion transporting polypeptide 1B1 inhibition profiles for the prediction of drug-drug interactions. Drug. Metab. Dispos. 2015, 43, 235–247. [Google Scholar] [CrossRef] [PubMed]
- Yoshikado, T.; Yoshida, K.; Kotani, N.; Nakada, T.; Asaumi, R.; Toshimoto, K.; Maeda, K.; Kusuhara, H.; Sugiyama, Y. Quantitative analyses of hepatic OATP-Mediated interactions between statins and inhibitors using PBPK modeling with a parameter optimization method. Clin. Pharmacol. Ther. 2016, 100, 513–523. [Google Scholar] [CrossRef] [PubMed]
- Shitara, Y.; Sugiyama, Y. Preincubation-dependent and long-lasting inhibition of organic anion transporting polypeptide (OATP) and its impact on drug-drug interactions. Pharmacol. Ther. 2017, 177, 67–80. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Li, S.; Patterson, C.; You, G. Lysine 48-linked polyubiquitination of organic anion transporter-1 is essential for its protein kinase C-regulated endocytosis. Mol. Pharmacol. 2013, 83, 217–224. [Google Scholar] [CrossRef] [PubMed]
| Substrate | Perpetrator Drugs | OATP1B1 Inhibition | OATP1B3 Inhibition | Reported AUCR |
|---|---|---|---|---|
| Atorvastatin | Boceprevir | [64] | [64] | 2.3 [65] |
| Faldaprevir | [66] | [66] | 9 [67] | |
| Sacubitril/Valsartan | [68,69] | [68,69] | 1.3 [70] | |
| Simprevir | [71] | [71] | 2.1 [72] | |
| Telaprevir | [73] | [73] | 7.8 [74] | |
| Tipranavir/Ritonavir | [75] | [76] | 9.4 [77] | |
| Pitavastatin | Rifampicin | [78] | [78] | 5.7–7.6 [79], 5.7 [80] |
| Pravastatin | Boceprevir | [64] | [64] | 1.6 [65] |
| Daclatasvir/Beclabuvir/Asunaprevir cocktail | [81] | [81] | 1.7 [82] | |
| Darunavir/Ritonavir | [75,83] | [76,83] | 2.1 [84] | |
| Paritaprevir/Ritonavir/Ombitasvir/Dasabuvir | [85] | [85] | 1.8 [86] | |
| Repaglinide | Clopidogrel | NA | [87] | 3.1 [88] |
| Gemfibrozil | [89] | [90] | 1.8 [91], 3.4 [92] | |
| Rifampicin | [78] | [78] | 1.9 [93] | |
| Rosuvastatin | Elvitegravir/Cobicistat/Emtricitabine/Tenovfovir | [94] | [94] | 1.4 [94] |
| Faldaprevir | [66] | [66] | 15 [67] | |
| Fostamatinib | [95] | NA | 2.0 [96] | |
| Furosemide/Digoxin/Metformin | [97] | [22,97] | 1.4 [98] | |
| Paritaprevir/Ritonavir/Ombitasvir/Dasabuvir | [85] | [85] | 1.6 [86] | |
| Rifampicin | [78] | [78] | 2 [99], 4.6–5.2 [79], 5 [100] | |
| Simprevir | [71] | [71] | 2.8 [72] | |
| Telmisartan | [101] | [101] | 1.3 [102] |
| Drugs | Number of Patients Taking Drug (N) | Number of Myopathy (M) | Risk | Relative Risk/p-Value |
|---|---|---|---|---|
| Metabolically stable statins † | 88,682 | 8149 | 9.2% | |
| Bortezomib and metabolically stable statins † | 311 | 25 | 8.0% | 0.87/0.58 |
| All statins § | 339,094 | 29,910 | 8.8% | |
| Bortezomib and all statins § | 1296 | 112 | 8.6% | 0.98/0.87 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alam, K.; Crowe, A.; Wang, X.; Zhang, P.; Ding, K.; Li, L.; Yue, W. Regulation of Organic Anion Transporting Polypeptides (OATP) 1B1- and OATP1B3-Mediated Transport: An Updated Review in the Context of OATP-Mediated Drug-Drug Interactions. Int. J. Mol. Sci. 2018, 19, 855. https://doi.org/10.3390/ijms19030855
Alam K, Crowe A, Wang X, Zhang P, Ding K, Li L, Yue W. Regulation of Organic Anion Transporting Polypeptides (OATP) 1B1- and OATP1B3-Mediated Transport: An Updated Review in the Context of OATP-Mediated Drug-Drug Interactions. International Journal of Molecular Sciences. 2018; 19(3):855. https://doi.org/10.3390/ijms19030855
Chicago/Turabian StyleAlam, Khondoker, Alexandra Crowe, Xueying Wang, Pengyue Zhang, Kai Ding, Lang Li, and Wei Yue. 2018. "Regulation of Organic Anion Transporting Polypeptides (OATP) 1B1- and OATP1B3-Mediated Transport: An Updated Review in the Context of OATP-Mediated Drug-Drug Interactions" International Journal of Molecular Sciences 19, no. 3: 855. https://doi.org/10.3390/ijms19030855
APA StyleAlam, K., Crowe, A., Wang, X., Zhang, P., Ding, K., Li, L., & Yue, W. (2018). Regulation of Organic Anion Transporting Polypeptides (OATP) 1B1- and OATP1B3-Mediated Transport: An Updated Review in the Context of OATP-Mediated Drug-Drug Interactions. International Journal of Molecular Sciences, 19(3), 855. https://doi.org/10.3390/ijms19030855




