2A-DUB/Mysm1 Regulates Epidermal Development in Part by Suppressing p53-Mediated Programs
Abstract
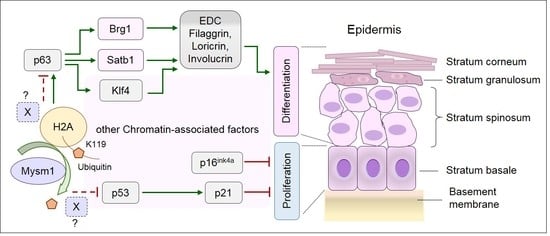
1. Introduction
2. Results
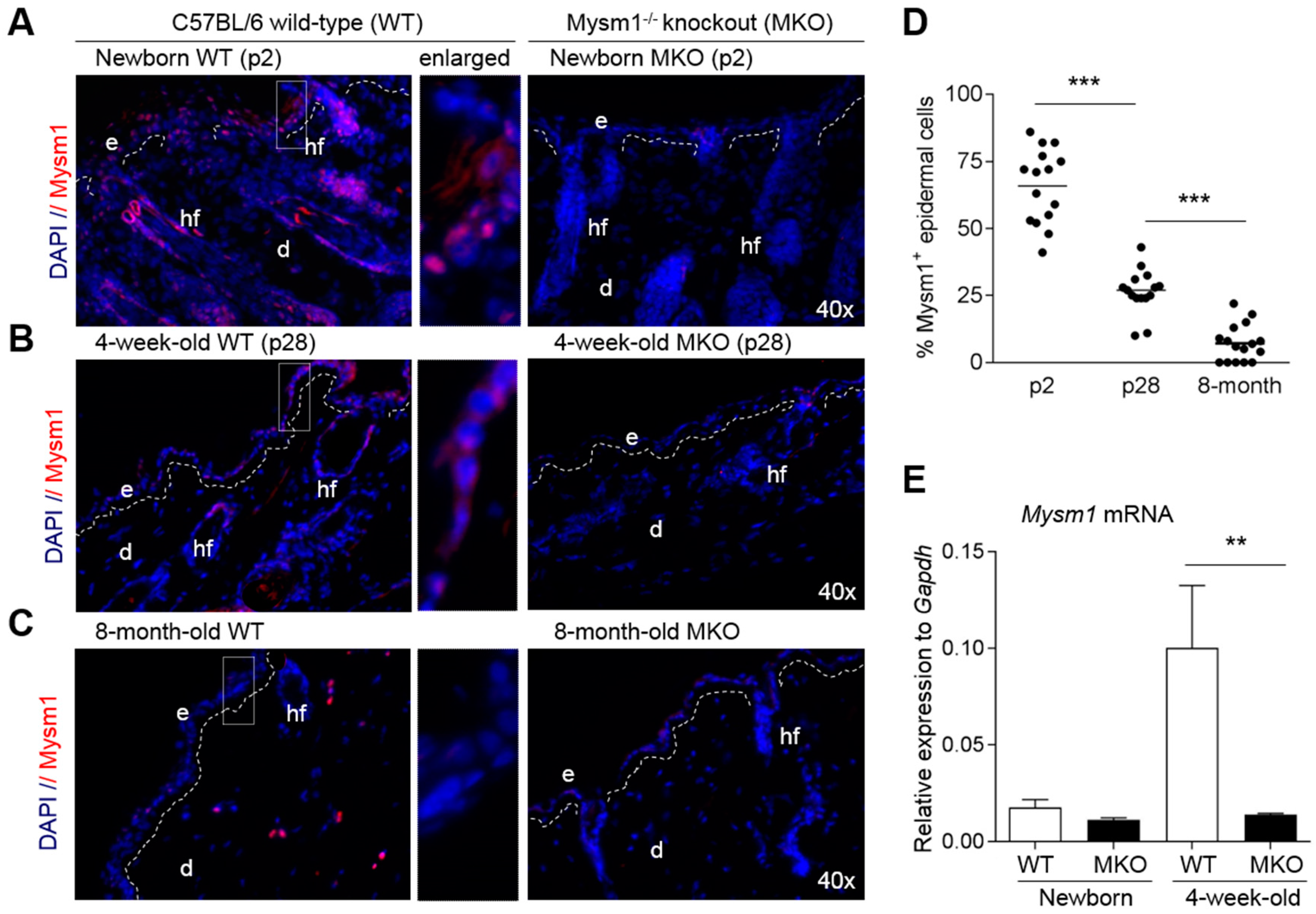
2.1. Mysm1 Expression in Murine Skin Declines with Age
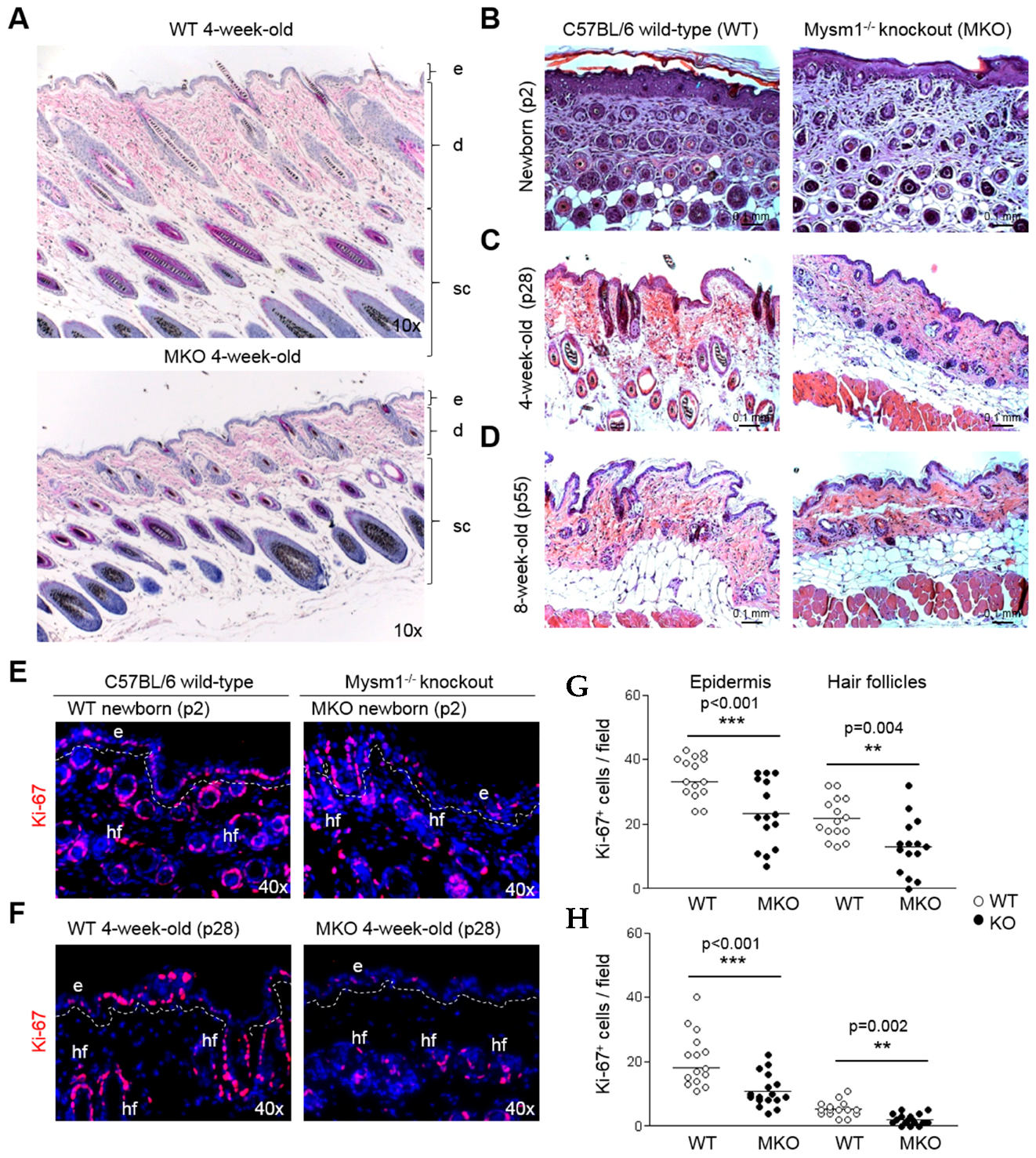
2.2. Loss of Mysm1 Causes Skin Atrophy and Reduced Skin Cellularity in Mysm1-Deficient Mice
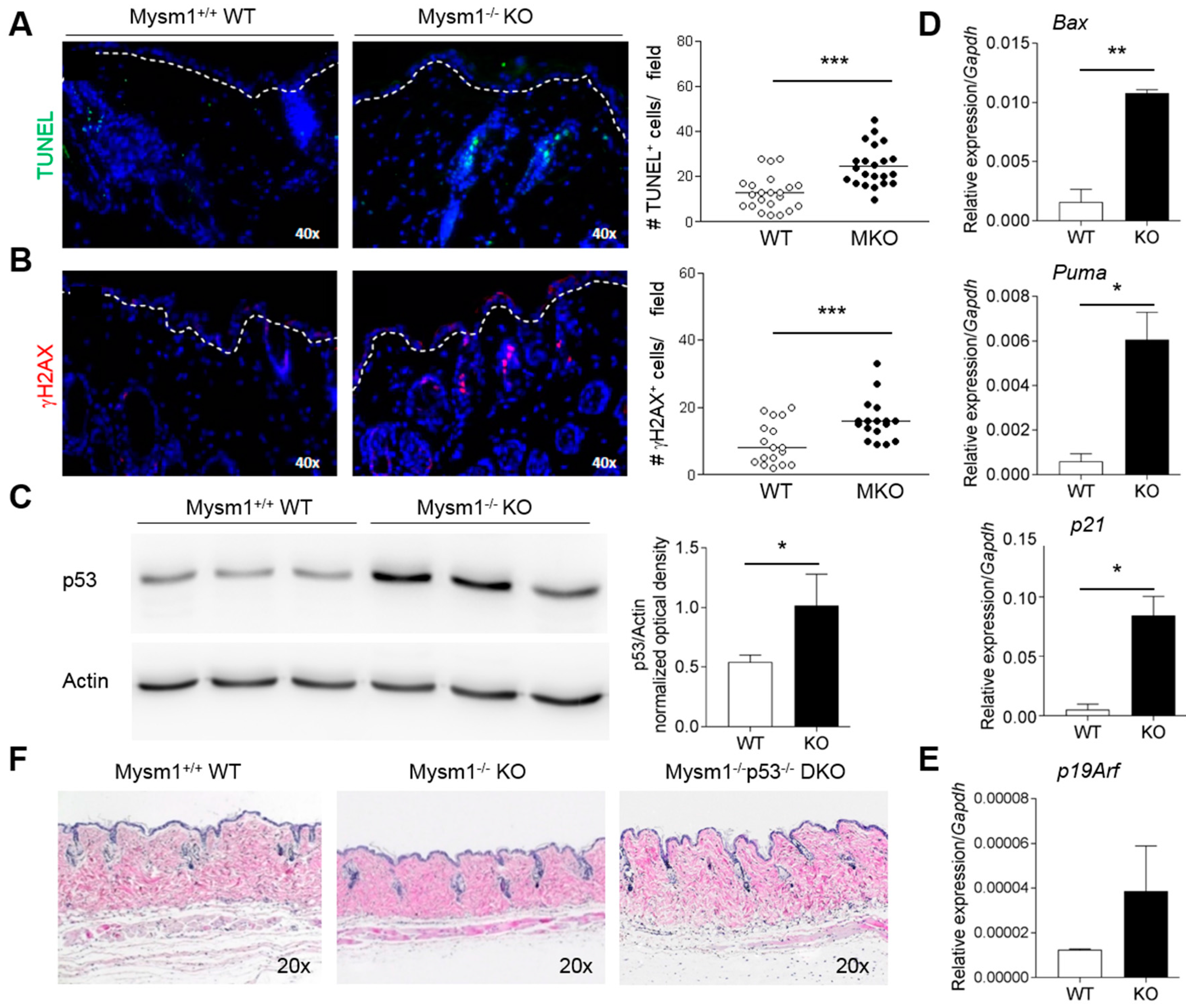
2.3. Increased p53-Dependent Apoptosis in Mysm1−/− Skin and Rescue of Skin Atrophy in Mysm1−/−p53−/− Mice
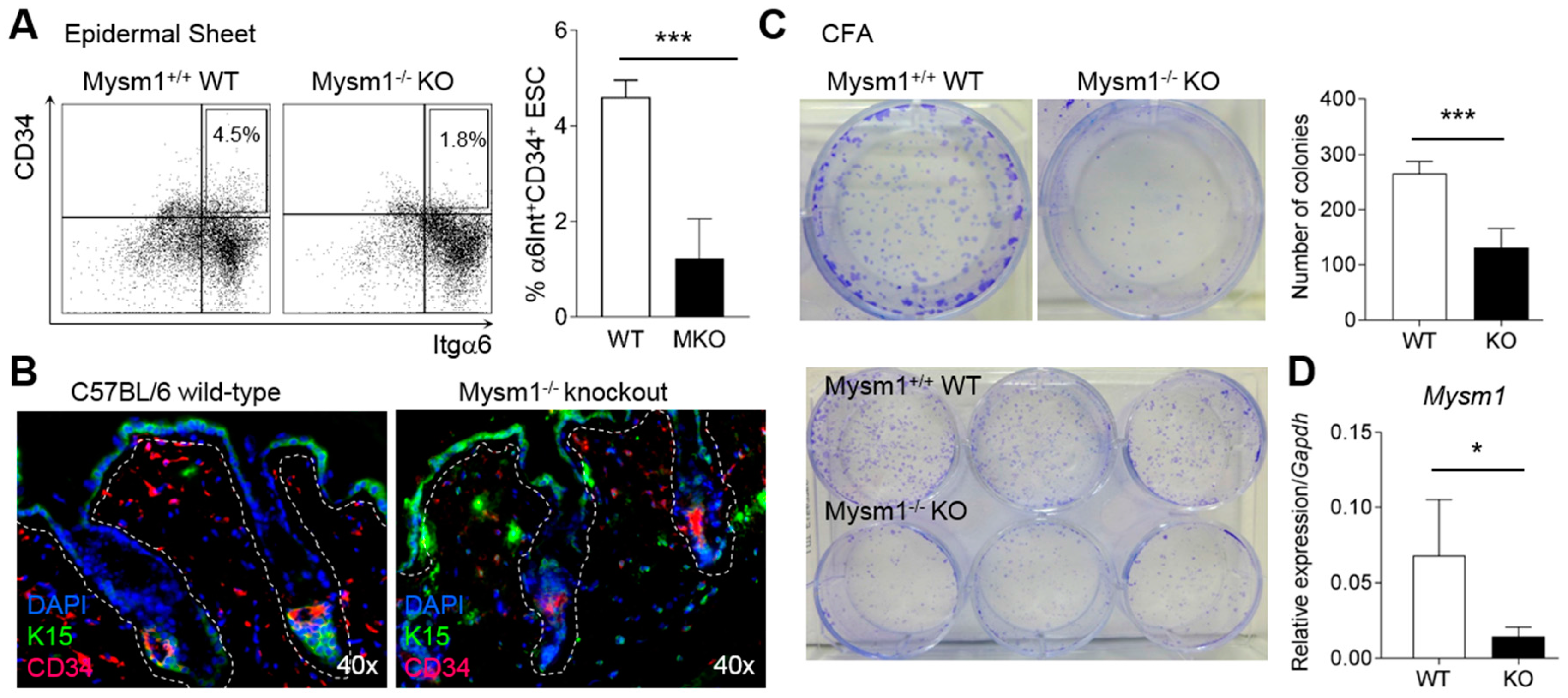
2.4. Mysm1-Deficiency Affects Epidermal Stem Cells
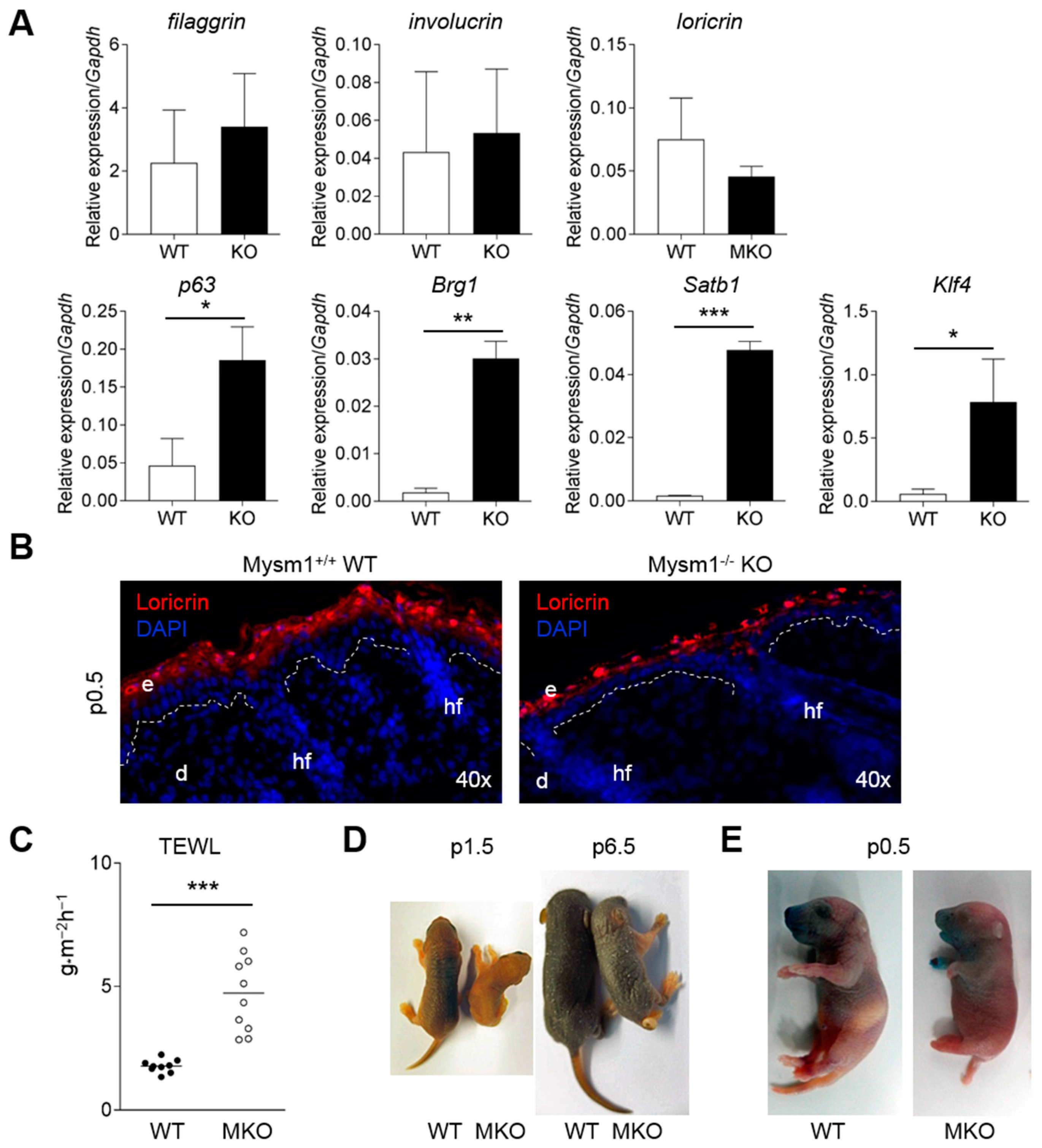
2.5. Defective Epidermal Barrier Formation in Mysm1-Deficient Mice
3. Discussion
4. Materials and Methods
4.1. Mouse Models
4.2. Histology and Immunofluorescent Analyses
4.3. qPCR
4.4. Western Blot
4.5. Colony Formation Assay (CFA)
4.6. Transepidermal Water Loss (TEWL)
4.7. Statistical Analyses
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Levine, A.J.; Berger, S.L. The interplay between epigenetic changes and the p53 protein in stem cells. Genes Dev. 2017, 31, 1195–1201. [Google Scholar] [CrossRef] [PubMed]
- Avgustinova, A.; Benitah, S.A. Epigenetic control of adult stem cell function. Nat. Rev. Mol. Cell Biol. 2016, 17, 643–658. [Google Scholar] [CrossRef] [PubMed]
- Blanpain, C.; Fuchs, E. Epidermal stem cells of the skin. Annu. Rev. Cell Dev. Biol. 2006, 22, 339–373. [Google Scholar] [CrossRef] [PubMed]
- Shen, Q.; Jin, H.; Wang, X. Epidermal stem cells and their epigenetic regulation. Int. J. Mol. Sci. 2013, 14, 17861–17880. [Google Scholar] [CrossRef] [PubMed]
- Botchkarev, V.A. Integration of the transcription factor-regulated and epigenetic mechanisms in the control of keratinocyte differentiation. J. Investig. Dermatol. Symp. Proc. 2015, 17, 30–32. [Google Scholar] [CrossRef] [PubMed]
- Abhishek, S.; Palamadai Krishnan, S. Epidermal differentiation complex: A review on its epigenetic regulation and potential drug targets. Cell J. 2016, 18, 1–6. [Google Scholar] [PubMed]
- Frye, M.; Benitah, S.A. Chromatin regulators in mammalian epidermis. Semin. Cell Dev. Biol. 2012, 23, 897–905. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhang, H.; Duan, E. Epidermal development in mammals: Key regulators, signals from beneath, and stem cells. Int. J. Mol. Sci. 2013, 14, 10869–10895. [Google Scholar] [CrossRef] [PubMed]
- Koster, M.I.; Kim, S.; Mills, A.A.; DeMayo, F.L.; Roop, D.R. p63 is the molecular switch for initiation of an epithelial stratification programm. Genes Dev. 2004, 18, 126–131. [Google Scholar] [CrossRef] [PubMed]
- Koster, M.I.; Dai, D.; Marinari, B.; Sano, Y.; Costanzo, A.; Karin, M.; Roop, D.R. p63 induces key target genes required for epidermal morphogenesis. Proc. Natl. Acad. Sci. USA 2007, 104, 3255–3260. [Google Scholar] [CrossRef] [PubMed]
- Ezhkova, E.; Pasolli, H.A.; Parker, J.S.; Stokes, N.; Su, I.H.; Hannon, G.; Tarakhovsky, A.; Fuchs, E. Ezh2 orchestrates gene expression for the stepweise differentiation of tissue-specific stem cells. Cell 2009, 136, 1122–1135. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Adhikary, G.; Balasubramanian, S.; Gopalakrishnan, R.; McCormick, T.; Dimri, G.P.; Eckert, R.L.; Rorke, E.A. Expression of Bmi-1 in epidermis enhandes cell survival by altering cell cycle regulatory protein expression and inhibiting apoptosis. J. Investig. Dermatol. 2008, 128, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Vissers, J.H.; Nicassio, F.; van Lohuizen, M.; Di Fiore, P.P.; Citterio, E. The many faces of ubiquitinated histone H2A: Insights from the DUBs. Cell Div. 2008, 3, 8. [Google Scholar] [CrossRef] [PubMed]
- Zhu, P.; Zhou, W.; Wang, J.; Puc, J.; Ohgi, K.A.; Erdjument-Bromage, H.; Tempst, P.; Glass, C.K.; Rosenfeld, M.G. A histone H2A deubiquitinase complex coordinating histone acetylation and H1 dissociation in transcriptional regulation. Mol. Cell 2007, 7, 609–621. [Google Scholar] [CrossRef] [PubMed]
- Nijnik, A.; Clare, S.; Hale, C.; Raisen, C.; McIntyre, R.E.; Yusa, K.; Everitt, A.R.; Mottram, L.; Podrini, C.; Lucas, M.; et al. The critical role of histone H2A-deubiquitinase Mysm1 in hematopoiesis and lymphocyte differentiation. Blood 2012, 119, 1370–1379. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Nandakumar, V.; Jiang, X.X.; Jones, L.; Yang, A.G.; Huang, X.F.; Chen, S.Y. The control of hematopoietic stem cell maintenance, self-renewal, and differentiation by Mysm1-mediated epigenetic regulation. Blood 2013, 122, 2812–2822. [Google Scholar] [CrossRef] [PubMed]
- Gatzka, M.; Tasdogan, A.; Hainzl, A.; Allies, G.; Maity, P.; Wilms, C.; Wlaschek, M.; Scharffetter-Kochanek, K. Interplay of H2A deubiquitinase 2A-DUB/Mysm1 and the p19(ARF)/p53 axis in hematopoiesis, early T-cell development and tissue differentiation. Cell Death Differ. 2015, 22, 1451–1462. [Google Scholar] [CrossRef] [PubMed]
- Belle, J.I.; Langlais, D.; Petrov, J.C.; Pardo, M.; Jones, R.G.; Gros, P.; Nijnik, A. p53 mediates loss of hematopoietic stem cell function and lymphopenia in Mysm1 deficiency. Blood 2015, 125, 2344–2348. [Google Scholar] [CrossRef] [PubMed]
- Belle, J.I.; Petrov, J.C.; Langlais, D.; Robert, F.; Cencic, R.; Shen, S.; Pelletier, J.; Gros, P.; Nijnik, A. Repression of p53-target gene Bbc3/PUMA by MYSM1 is essential for the survival of hematopoietic multipotent progenitors and contributes to stem cell maintenance. Cell Death Differ. 2016, 23, 759–775. [Google Scholar] [CrossRef] [PubMed]
- Liakath-Ali, K.; Vancollie, V.E.; Heath, E.; Smedley, D.P.; Estabel, J.; Sunter, D.; Ditommaso, T.; White, J.K.; Ramirez-Solis, R.; Smyth, I.; et al. Novel skin phenotypes revealed by a genome-wide mouse reverse genetic screen. Nat. Commun. 2014, 5, 3540. [Google Scholar] [CrossRef] [PubMed]
- DiTommaso, T.; Jones, L.K.; Cottle, D.L.; WTSI Mouse Genetics Program; Gerdin, A.K.; Vancollie, V.E.; Watt, F.M.; Ramirez-Solis, R.; Bradley, A.; Steel, K.P. Identification of genes important for cutaneous function revealed by a large scale reverse genetic screen in the mouse. PLoS Genet. 2014, 10, e1004705. [Google Scholar] [CrossRef] [PubMed]
- Wilms, C.; Kroeger, C.M.; Hainzl, A.V.; Banik, I.; Bruno, C.; Krikki, I.; Farsam, V.; Wlaschek, M.; Gatzka, M.V. MYSM1/2A-DUB is an epigenetic regulator in human melanoma and contributes to tumor cell growth. Oncotarget 2017, 8, 67287–67299. [Google Scholar] [CrossRef] [PubMed]
- Haffner-Luntzer, M.; Kovtun, A.; Fischer, V.; Prystaz, K.; Hainzl, A.; Kroeger, C.M.; Krikki, I.; Brinker, T.J.; Ignatius, A.; Gatzka, M. Loss of p53 compensates osteopenia in murine Mysm1-deficiency. FASEB J. 2017. [Google Scholar] [CrossRef] [PubMed]
- Panda, S.; Nilsson, J.A.; Gekara, N.O. Deubiquitinase MYSM1 regulates innate immunity through inactivation of TRAF3 and TRAF6 complexes. Immunity 2015, 43, 647–659. [Google Scholar] [CrossRef] [PubMed]
- Raj, D.; Brash, D.E.; Grossman, D. Keratinocyte apoptosis in epidermal development and disease. J. Investig. Dermatol. 2006, 126, 243–257. [Google Scholar] [CrossRef] [PubMed]
- Lacroix, M.; Caramel, J.; Goguet-Rubio, P.; Linares, L.K.; Estrach, S.; Hatchi, E.; Rodier, G.; Lledo, G.; de Bettignies, C.; Thépot, A.; et al. Transcription factor E4F1 is essential for epidermal stem cell maintenance and skin homeostasis. Proc. Natl. Acad. Sci. USA 2010, 107, 21076–21081. [Google Scholar] [CrossRef] [PubMed]
- D’Arcangelo, D.; Tinaburri, L.; Dellambra, E. The role of p16INK4a pathway in human epidermal stem cell self-renewal, aging, and cancer. Int. J. Mol. Sci. 2017, 18, 1591. [Google Scholar] [CrossRef] [PubMed]
- Bhaduri, A.; Ungewickell, A.; Boxer, L.D.; Lopez-Pajares, V.; Zarnegar, B.J.; Khavari, P.A. Network analysis identifies mitochondrial regulation of epidermal differentiation by MPZL3 and FXDR. Dev. Cell 2015, 35, 444–457. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.C.; Wang, D.Y.; Kao, M.H.; Chen, J.K. The growth-promoting effect of KGF on limbal epithelial cells is mediated by upregulation of DeltaNp63alpha through the p38 pathway. J. Cell Sci. 2009, 122, 4473–4480. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wagner, R.A.; Tabibiazar, R.; Liao, A.; Quertermous, T. Genome-wide expression dynamics during mouse embryonic development reveal similarities to Drosophila development. Dev. Biol. 2005, 288, 595–611. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mardaryev, A.N.; Liu, B.; Rapisarda, V.; Poterlowicz, K.; Malashchuk, I.; Rudolf, J.; Sharov, A.A.; Jahoda, C.A.; Fessing, M.Y.; Benitah, S.A.; et al. Cbx4 maintains the epithelial lineage identity and cell proliferation in the developing stratified epithelium. J. Cell Biol. 2016, 212, 77–89. [Google Scholar] [CrossRef] [PubMed]
- Sheehan, D.C.; Hrapchak, B.B. Theory and Practice of Histotechnology, 2nd ed.; Mosby: St. Louis, MO, USA, 1980; pp. 1–474. ISBN 13: 978-0801645730. [Google Scholar]
- Gatzka, M.; Prisco, M.; Baserga, R. Stabilization of the Ras oncoprotein by the insulin-like growth factor 1 receptor during anchorage-independent growth. Cancer Res. 2000, 60, 4222–4230. [Google Scholar] [PubMed]
- Gatzka, M.; Hainzl, A.; Peters, T.; Singh, K.; Tasdogan, A.; Wlaschek, M.; Scharffetter-Kochanek, K. Reduction of CD18 promotes expansion of inflammatory γδ T cells collaborating with CD4+ T cells in chronic murine psoriasiform dermatitis. J. Immunol. 2013, 191, 5477–5488. [Google Scholar] [CrossRef] [PubMed]
- Jensen, U.B.; Ghazizadeh, S.; Owens, D.M. Isolation and characterization of cutaneous epithelial stem cells. Methods Mol. Biol. 2013, 989, 61–69. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wilms, C.; Krikki, I.; Hainzl, A.; Kilo, S.; Alupei, M.; Makrantonaki, E.; Wagner, M.; Kroeger, C.M.; Brinker, T.J.; Gatzka, M. 2A-DUB/Mysm1 Regulates Epidermal Development in Part by Suppressing p53-Mediated Programs. Int. J. Mol. Sci. 2018, 19, 687. https://doi.org/10.3390/ijms19030687
Wilms C, Krikki I, Hainzl A, Kilo S, Alupei M, Makrantonaki E, Wagner M, Kroeger CM, Brinker TJ, Gatzka M. 2A-DUB/Mysm1 Regulates Epidermal Development in Part by Suppressing p53-Mediated Programs. International Journal of Molecular Sciences. 2018; 19(3):687. https://doi.org/10.3390/ijms19030687
Chicago/Turabian StyleWilms, Christina, Ioanna Krikki, Adelheid Hainzl, Sonja Kilo, Marius Alupei, Evgenia Makrantonaki, Maximilian Wagner, Carsten M. Kroeger, Titus Josef Brinker, and Martina Gatzka. 2018. "2A-DUB/Mysm1 Regulates Epidermal Development in Part by Suppressing p53-Mediated Programs" International Journal of Molecular Sciences 19, no. 3: 687. https://doi.org/10.3390/ijms19030687
APA StyleWilms, C., Krikki, I., Hainzl, A., Kilo, S., Alupei, M., Makrantonaki, E., Wagner, M., Kroeger, C. M., Brinker, T. J., & Gatzka, M. (2018). 2A-DUB/Mysm1 Regulates Epidermal Development in Part by Suppressing p53-Mediated Programs. International Journal of Molecular Sciences, 19(3), 687. https://doi.org/10.3390/ijms19030687