Effect of Hybridization on Somatic Mutations and Genomic Rearrangements in Plants
Abstract
1. Introduction
2. Hybridization Affects Somatic Mutations in Plants
Theories Behind Altered Mutation Rate Behavior in Hybrids—Trends Emerging from the Literature
3. Hybridization Alters Genome Size
Transposition Events Play an Important Role in Bringing Genome Size Changes and Cause Genomic Restructuring in Hybrids
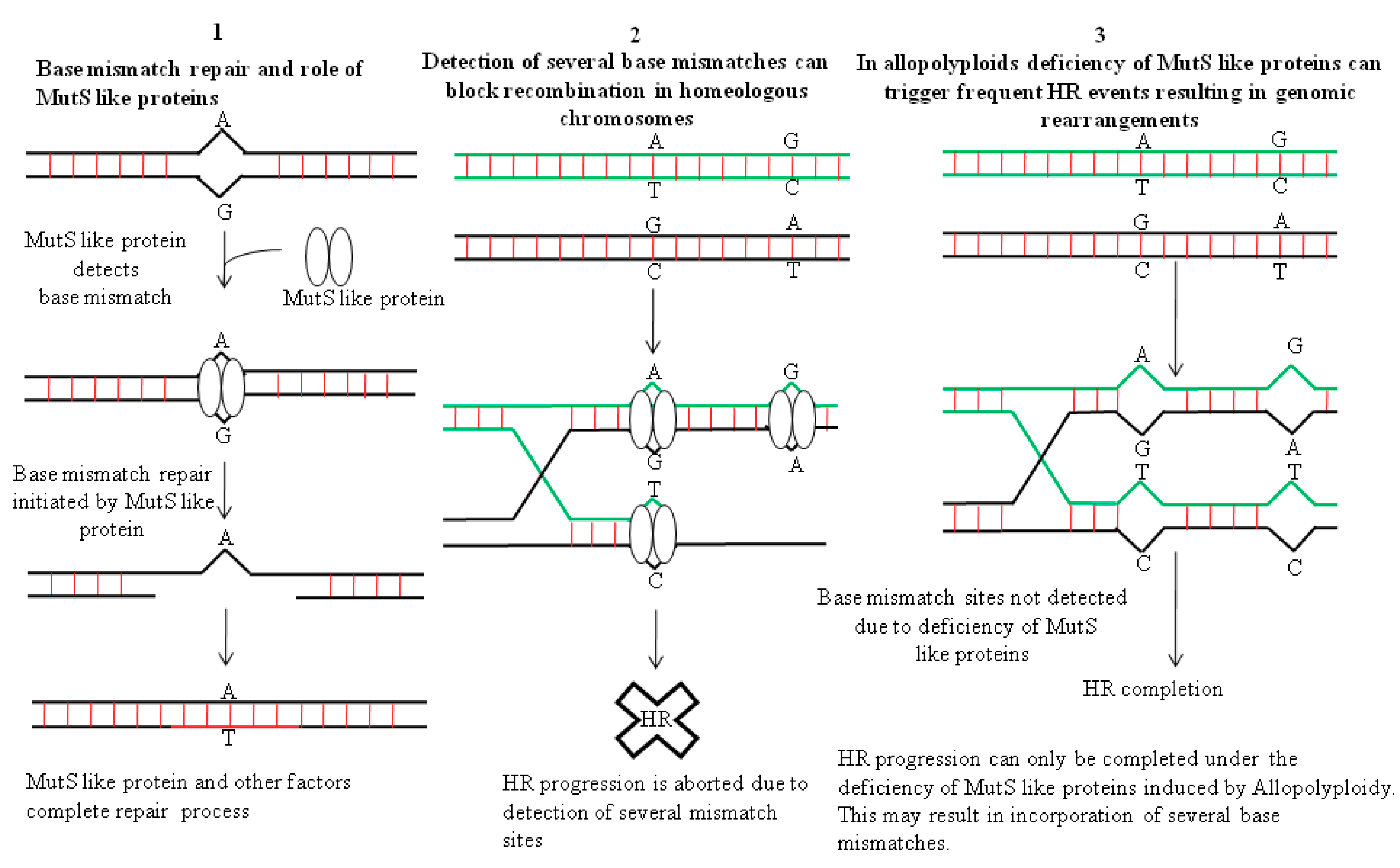
4. Genomic Restructuring in Hybrids: The Role Played by Recombination Events in Facilitating Changes in the Architecture of the Genome
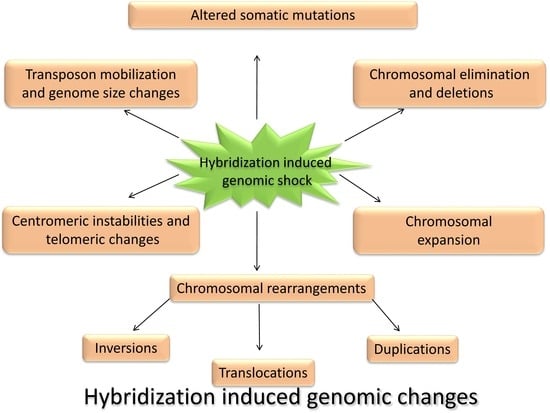
5. Hybridization Triggers Changes in Chromosome Architecture
5.1. Chromosomal Elimination
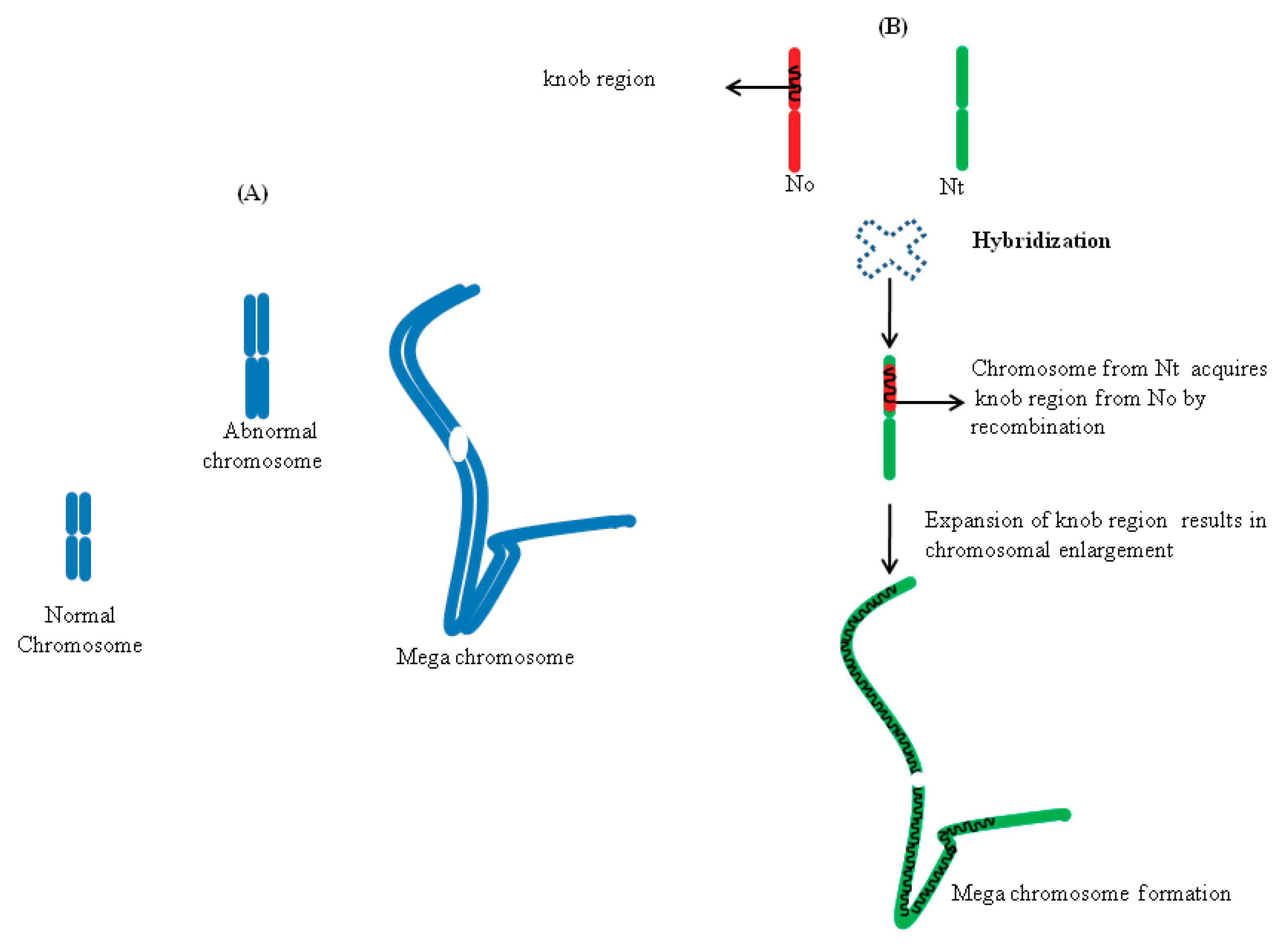
5.2. Chromosome Expansion in Hybrids
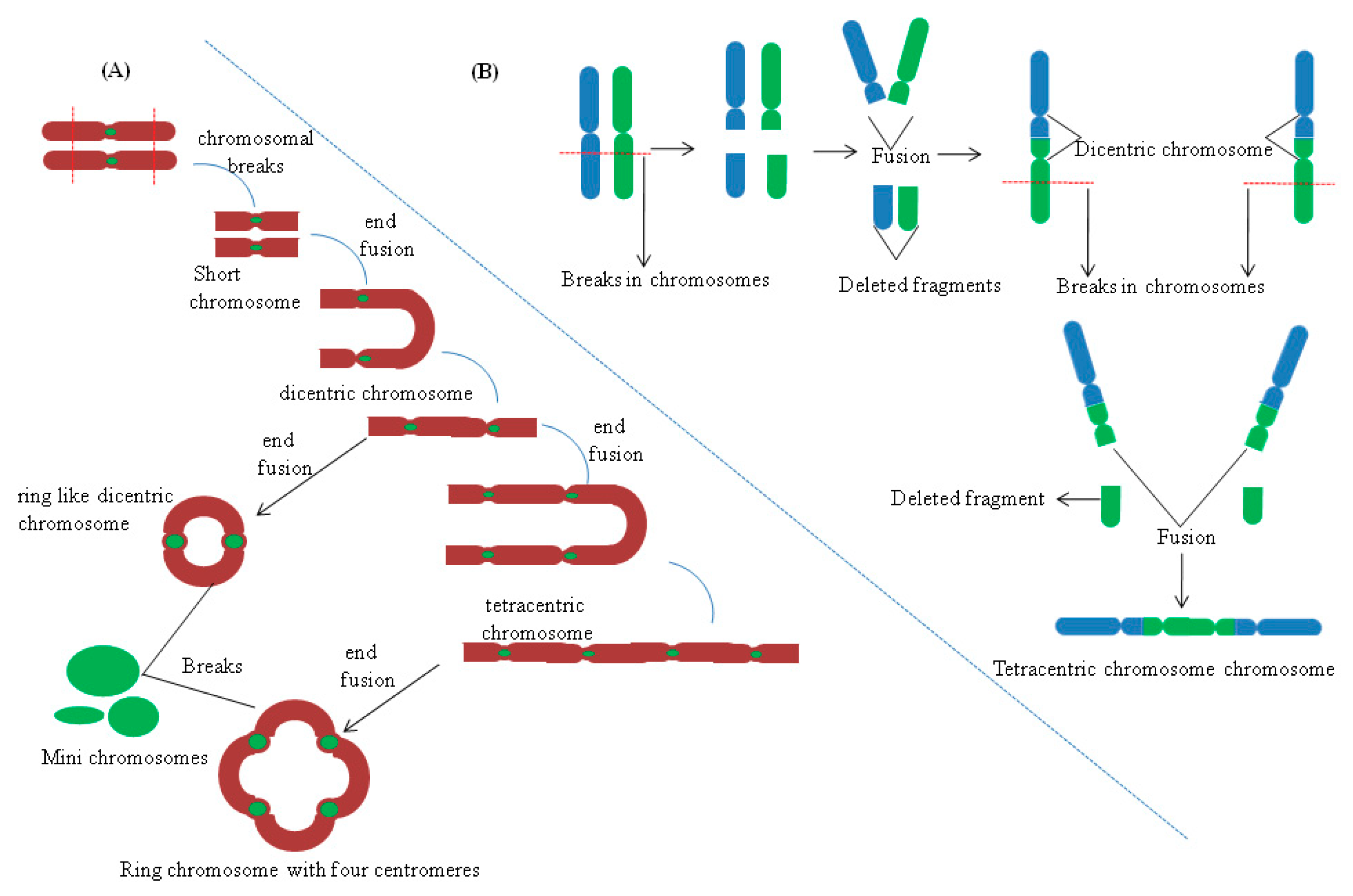
5.3. Chromosomal Rearrangements
6. Effects of Hybridization on Chromatin Structure and Centromere Organization
7. Conclusions and Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Darwin, C.R. The Effects of Cross and Self Fertilization in the Vegetable Kingdom; John Murray: UK, London, 1876. [Google Scholar]
- Duvick, D.N. Heterosis: Feeding people and protecting natural resources. In Genetics and Exploitation of Heterosis in Crops; Coors, J.G., Pandey, S., Eds.; Crop Science Society of America: Madison, WI, USA, 1999; pp. 19–29. [Google Scholar]
- Khush, G.S. Green revolution: The way forward. Nat. Rev. Genet. 2001, 2, 815–822. [Google Scholar] [CrossRef] [PubMed]
- Mallet, J. Hybrid speciation. Nature 2007, 446, 279–283. [Google Scholar] [CrossRef] [PubMed]
- McClintock, B. The significance of responses of the genome to challenge. Science 1984, 226, 792–801. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.J. Molecular mechanisms of polyploidy and hybrid vigor. Trends Plant Sci. 2010, 15, 57–71. [Google Scholar] [CrossRef] [PubMed]
- Del Pozo, J.C.; Ramirez-Parra, E. Whole genome duplications in plants: An overview from Arabidopsis. J. Exp. Bot. 2015, 66, 6991–7003. [Google Scholar] [CrossRef] [PubMed]
- Shaked, H.; Kashkush, K.; Ozkan, H.; Feldman, M.; Levy, A.A. Sequence elimination and cytosine methylation are rapid and reproducible responses of the genome to wide hybridization and allopolyploidy in wheat. Plant Cell 2001, 13, 1749–1759. [Google Scholar] [CrossRef] [PubMed]
- Vitte, C.; Bennetzen, J.L. Analysis of retrotransposon structural diversity uncovers properties and propensities in angiosperm genome evolution. Proc. Natl. Acad. Sci. USA 2006, 103, 17638–17643. [Google Scholar] [CrossRef] [PubMed]
- Baake, E.; Gabriel, W. Biological evolution through mutation, selection and drift: An introductory review. Annu. Rev. Comp. Phys. 1999, 7, 203–264. [Google Scholar]
- Kovalchuk, I.; Kovalchuk, O.; Hohn, B. Genome-wide variation of the somatic mutation frequency in transgenic plants. EMBO J. 2000, 19, 4431–4438. [Google Scholar] [CrossRef] [PubMed]
- Ossowski, S.; Schneeberger, K.; Lucas-Lledo, J.I.; Warthmann, N.; Clark, R.M.; Shaw, R.G.; Weigel, D.; Lynch, M. The rate and molecular spectrum of spontaneous mutations in Arabidopsis thaliana. Science 2010, 327, 92–94. [Google Scholar] [CrossRef] [PubMed]
- Bashir, T.; Sailer, C.; Gerber, F.; Loganathan, N.; Bhoopalan, H.; Eichenberger, C.; Grossniklaus, U.; Baskar, R. Hybridization alters spontaneous mutation rates in a parent-of-origin-dependent fashion in Arabidopsis. Plant Physiol. 2014, 165, 424–437. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Wang, L.; Wang, L.; Wang, Z.; Lu, Z.; Tian, D.; Yang, S.; Hurst, L.D. Mutation rate analysis via parent-progeny sequencing of the perennial peach. I. A low rate in woody perennials and a higher mutagenicity in hybrids. Proc. Biol. Sci. 2016, 283. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Wang, L.; Huang, J.; Zhang, X.; Yuan, Y.; Chen, J.Q.; Hurst, L.D.; Tian, D. Parent-progeny sequencing indicates higher mutation rates in heterozygotes. Nature 2015, 523, 463–467. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhang, Y.; Qin, C.; Tian, D.; Yang, S.; Hurst, L.D. Mutation rate analysis via parent-progeny sequencing of the perennial peach. II. No evidence for recombination-associated mutation. Proc. Biol. Sci. 2016, 283. [Google Scholar] [CrossRef] [PubMed]
- Kostoff, D. On the increase of mutation frequency following interspecific hybridization in Nicotiana. Curr. Sci. 1935, 3, 302–304. [Google Scholar]
- Napoli, C.; Lemieux, C.; Jorgensen, R. Introduction of a Chimeric Chalcone Synthase Gene into Petunia Results in Reversible Co-Suppression of Homologous Genes in trans. Plant Cell 1990, 2, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Gerats, A.G.; Huits, H.; Vrijlandt, E.; Marana, C.; Souer, E.; Beld, M. Molecular characterization of a nonautonomous transposable element (dTph1) of petunia. Plant Cell 1990, 2, 1121–1128. [Google Scholar] [CrossRef] [PubMed]
- Belgovsky, M.L. A comparison of the frequency of induced mutations in D. simulans and in its hybrid with D. melanogaster. Genetica 1937, 19, 370–386. [Google Scholar] [CrossRef]
- Thompson, J.N., Jr.; Woodruff, R.C. Increased mutation in crosses between geographically separated strains of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 1980, 77, 1059–1062. [Google Scholar] [CrossRef] [PubMed]
- Sturtevant, A.H. High Mutation Frequency Induced by Hybridization. Proc. Natl. Acad. Sci. USA 1939, 25, 308–310. [Google Scholar] [CrossRef] [PubMed]
- Simmons, M.J.; Johnson, N.A.; Fahey, T.M.; Nellett, S.M.; Raymond, J.D. High mutability in male hybrids of Drosophila melanogaster. Genetics 1980, 96, 479–480. [Google Scholar] [PubMed]
- Amos, W. Heterozygosity and mutation rate: Evidence for an interaction and its implications: The potential for meiotic gene conversions to influence both mutation rate and distribution. Bioessays 2010, 32, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Comai, L.; Madlung, A.; Josefsson, C.; Tyagi, A. Do the different parental ‘heteromes’ cause genomic shock in newly formed allopolyploids? Philos. Trans. R. Soc. Lond. B Biol. Sci. 2003, 358, 1149–1155. [Google Scholar] [CrossRef] [PubMed]
- Demerec, M. Changes in the Rate of Mutability of the Mutable Miniature Gene of Drosophila Virilis. Proc. Natl. Acad. Sci. USA 1929, 15, 870–876. [Google Scholar] [CrossRef] [PubMed]
- Amos, W. Even small SNP clusters are non-randomly distributed: Is this evidence of mutational non-independence? Proc. Biol. Sci. 2010, 277, 1443–1449. [Google Scholar] [CrossRef] [PubMed]
- Woodruff, R.C.; Thompson, J.N., Jr.; Lyman, R.F. Intraspecific hybridisation and the release of mutator activity. Nature 1979, 278, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Bennetzen, J.L.; Ma, J.; Devos, K.M. Mechanisms of recent genome size variation in flowering plants. Ann. Bot. 2005, 95, 127–132. [Google Scholar] [CrossRef] [PubMed]
- Rogers, S.O.; Bendich, A.J. Heritability and Variability in Ribosomal RNA Genes of Vicia faba. Genetics 1987, 117, 285–295. [Google Scholar] [PubMed]
- Petrov, D.A.; Schutzman, J.L.; Hartl, D.L.; Lozovskaya, E.R. Diverse transposable elements are mobilized in hybrid dysgenesis in Drosophila virilis. Proc. Natl. Acad. Sci. USA 1995, 92, 8050–8054. [Google Scholar] [CrossRef] [PubMed]
- Labrador, M.; Farre, M.; Utzet, F.; Fontdevila, A. Interspecific hybridization increases transposition rates of Osvaldo. Mol. Biol. Evol. 1999, 16, 931–937. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, R.J.; O’Neill, M.J.; Graves, J.A. Undermethylation associated with retroelement activation and chromosome remodelling in an interspecific mammalian hybrid. Nature 1998, 393, 68–72. [Google Scholar] [CrossRef] [PubMed]
- Baack, E.J.; Whitney, K.D.; Rieseberg, L.H. Hybridization and genome size evolution: Timing and magnitude of nuclear DNA content increases in Helianthus homoploid hybrid species. New Phytol. 2005, 167, 623–630. [Google Scholar] [CrossRef] [PubMed]
- Burke, J.M.; Lai, Z.; Salmaso, M.; Nakazato, T.; Tang, S.; Heesacker, A.; Knapp, S.J.; Rieseberg, L.H. Comparative mapping and rapid karyotypic evolution in the genus helianthus. Genetics 2004, 167, 449–457. [Google Scholar] [CrossRef] [PubMed]
- Ungerer, M.C.; Strakosh, S.C.; Zhen, Y. Genome expansion in three hybrid sunflower species is associated with retrotransposon proliferation. Curr. Biol. 2006, 16, 872–873. [Google Scholar] [CrossRef] [PubMed]
- Caceres, M.E.; De Pace, C.; Mugnozza, G.T.S.; Kotsonis, P.; Ceccarelli, M.; Cionini, P.G. Genome size variations within Dasypyrum villosum: Correlations with chromosomal traits, environmental factors and plant phenotypic characteristics and behaviour in reproduction. Theor. Appl. Genet. 1998, 96, 559–567. [Google Scholar] [CrossRef]
- Thomas, C.A., Jr. The genetic organization of chromosomes. Ann. Rev. Genet. 1971, 5, 237–256. [Google Scholar] [CrossRef] [PubMed]
- Grotkopp, E.; Rejmanek, M.; Sanderson, M.J.; Rost, T.L. Evolution of genome size in pines (Pinus) and its life-history correlates: Supertree analyses. Evolution 2004, 58, 1705–1729. [Google Scholar] [CrossRef] [PubMed]
- Vinogradov, A.E. Selfish DNA is maladaptive: Evidence from the plant Red List. Trends Genet. 2003, 19, 609–614. [Google Scholar] [CrossRef] [PubMed]
- Leitch, I.; Bennett, M. Genome downsizing in polyploid plants. Biol. J. Linn. Soc. 2004, 82, 651–663. [Google Scholar] [CrossRef]
- Petrov, D.A. Evolution of genome size: New approaches to an old problem. Trends Genet. 2001, 17, 23–28. [Google Scholar] [CrossRef]
- Bennetzen, J.L. Mechanisms and rates of genome expansion and contraction in flowering plants. Genetica 2002, 115, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Gregory, T.R. Insertion-deletion biases and the evolution of genome size. Gene 2004, 324, 15–34. [Google Scholar] [CrossRef] [PubMed]
- Petrov, D.A. Mutational equilibrium model of genome size evolution. Theor. Popul. Biol. 2002, 61, 531–544. [Google Scholar] [CrossRef] [PubMed]
- Feschotte, C.; Pritham, E.J. DNA transposons and the evolution of eukaryotic genomes. Ann. Rev Genet. 2007, 41, 331–368. [Google Scholar] [CrossRef] [PubMed]
- Bennetzen, J.L.; Wang, H. The contributions of transposable elements to the structure, function, and evolution of plant genomes. Ann. Rev. Plant Biol. 2014, 65, 505–530. [Google Scholar] [CrossRef] [PubMed]
- Hazzouri, K.M.; Mohajer, A.; Dejak, S.I.; Otto, S.P.; Wright, S.I. Contrasting patterns of transposable-element insertion polymorphism and nucleotide diversity in autotetraploid and allotetraploid Arabidopsis species. Genetics 2008, 179, 581–592. [Google Scholar] [CrossRef] [PubMed]
- Hughes, A.L.; Friedman, R.; Ekollu, V.; Rose, J.R. Non-random association of transposable elements with duplicated genomic blocks in Arabidopsis thaliana. Mol. Phylogenet. Evol. 2003, 29, 410–416. [Google Scholar] [CrossRef]
- Kumar, A.; Bennetzen, J.L. Plant retrotransposons. Ann. Rev. Genet. 1999, 33, 479–532. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Santerre-Ayotte, S.; Boivin, E.B.; Jean, M.; Belzile, F. A novel reporter for intrachromosomal homoeologous recombination in Arabidopsis thaliana. Plant J. 2004, 40, 1007–1015. [Google Scholar] [CrossRef] [PubMed]
- Sung, P.; Klein, H. Mechanism of homologous recombination: Mediators and helicases take on regulatory functions. Nat. Rev. Mol. Cell Biol. 2006, 7, 739–750. [Google Scholar] [CrossRef] [PubMed]
- Renkawitz, J.; Lademann, C.A.; Jentsch, S. Mechanisms and principles of homology search during recombination. Nat. Rev. Mol. Cell Biol. 2014, 15, 369–383. [Google Scholar] [CrossRef] [PubMed]
- Luo, M.C.; Dubcovsky, J.; Dvorak, J. Recognition of homeology by the wheat Ph1 locus. Genetics 1996, 144, 1195–1203. [Google Scholar] [PubMed]
- Moore, G. To pair or not to pair: Chromosome pairing and evolution. Curr. Opin. Plant Biol. 1998, 1, 116–122. [Google Scholar] [CrossRef]
- Comai, L. Genetic and epigenetic interactions in allopolyploid plants. Plant Mol. Biol. 2000, 43, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Lilley, D.M.J. Holliday junction-resolving enzymes-structures and mechanisms. FEBS Lett. 2017, 591, 1073–1082. [Google Scholar] [CrossRef] [PubMed]
- Culligan, K.M.; Hays, J.B. Arabidopsis MutS homologs-AtMSH2, AtMSH3, AtMSH6, and a novel AtMSH7-form three distinct protein heterodimers with different specificities for mismatched DNA. Plant Cell 2000, 12, 991–1002. [Google Scholar] [PubMed]
- Fu, S.; Lv, Z.; Guo, X.; Zhang, X.; Han, F. Alteration of terminal heterochromatin and chromosome rearrangements in derivatives of wheat-rye hybrids. J. Genet. Genomics 2013, 40, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Beaulieu, J.; Jean, M.; Belzile, F. The allotetraploid Arabidopsis thaliana-Arabidopsis lyrata subsp. petraea as an alternative model system for the study of polyploidy in plants. Mol. Genet. Genomics 2009, 281, 421–435. [Google Scholar] [CrossRef] [PubMed]
- Ravi, M.; Chan, S.W. Haploid plants produced by centromere-mediated genome elimination. Nature 2010, 464, 615–618. [Google Scholar] [CrossRef] [PubMed]
- Tonosaki, K.; Osabe, K.; Kawanabe, T.; Fujimoto, R. The importance of reproductive barriers and the effect of allopolyploidization on crop breeding. Breed. Sci. 2016, 66, 333–349. [Google Scholar] [CrossRef] [PubMed]
- Ho, K.M.; Kasha, K.J. Genetic Control of Chromosome Elimination during Haploid Formation in Barley. Genetics 1975, 81, 263–275. [Google Scholar] [PubMed]
- Sanei, M.; Pickering, R.; Kumke, K.; Nasuda, S.; Houben, A. Loss of centromeric histone H3 (CENH3) from centromeres precedes uniparental chromosome elimination in interspecific barley hybrids. Proc. Natl. Acad. Sci. USA 2011, 108, 498–505. [Google Scholar] [CrossRef] [PubMed]
- Renny-Byfield, S.; Kovarik, A.; Chester, M.; Nichols, R.A.; Macas, J.; Novak, P.; Leitch, A.R. Independent, rapid and targeted loss of highly repetitive DNA in natural and synthetic allopolyploids of Nicotiana tabacum. PLoS ONE 2012, 7, e36963. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Li, M.; Chen, L.; Wang, Y.; Ren, Z.; Fu, S. New types of wheat chromosomal structural variations in derivatives of wheat-rye hybrids. PLoS ONE 2014, 9, e110282. [Google Scholar] [CrossRef] [PubMed]
- Han, F.; Fedak, G.; Guo, W.; Liu, B. Rapid and repeatable elimination of a parental genome-specific DNA repeat (pGc1R-1a) in newly synthesized wheat allopolyploids. Genetics 2005, 170, 1239–1245. [Google Scholar] [CrossRef] [PubMed]
- Tan, E.H.; Henry, I.M.; Ravi, M.; Bradnam, K.R.; Mandakova, T.; Marimuthu, M.P.; Korf, I.; Lysak, M.A.; Comai, L.; Chan, S.W. Catastrophic chromosomal restructuring during genome elimination in plants. Elife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- Bennett, M.D.; Finch, R.A.; Barclay, I.R. The time rate and mechanism of chromosome elimination in Hordeum hybrids. Chromosoma 1976, 54, 175–200. [Google Scholar] [CrossRef]
- Gupta, S.B. Duration of mitotic cycle and regulation of DNA replication in Nicotiana plumbaginifolia and a hybrid derivative of N. tabacum showing chromosome instability. Can. J. Genet. Cytol. 1969, 11, 133–142. [Google Scholar] [CrossRef]
- Finch, R.A. Tissue-specific elimination of alternative whole parental genomes in one barley hybrid. Chromosoma 1983, 88, 386–393. [Google Scholar] [CrossRef]
- Kim, N.S.; Armstrong, K.C.; Fedak, G.; Ho, K.; Park, N.I. A microsatellite sequence from the rice blast fungus (Magnaporthe grisea) distinguishes between the centromeres of Hordeum vulgare and H. bulbosum in hybrid plants. Genome 2002, 45, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Gernand, D.; Rutten, T.; Pickering, R.; Houben, A. Elimination of chromosomes in Hordeum vulgare x H. bulbosum crosses at mitosis and interphase involves micronucleus formation and progressive heterochromatinization. Cytogenet. Genome Res. 2006, 114, 169–174. [Google Scholar] [CrossRef] [PubMed]
- Ishii, T.; Ueda, T.; Tanaka, H.; Tsujimoto, H. Chromosome elimination by wide hybridization between Triticeae or oat plant and pearl millet: Pearl millet chromosome dynamics in hybrid embryo cells. Chromosome Res. 2010, 18, 821–831. [Google Scholar] [CrossRef] [PubMed]
- Linde-Laursen, I.; von Bothmer, R. Elimination and duplication of particular Hordeum vulgare chromosomes in aneuploid interspecific Hordeum hybrids. TAG. Theor. Appl. Genet. 1988, 76, 897–908. [Google Scholar] [CrossRef] [PubMed]
- Badaeva, E.D.; Dedkova, O.S.; Gay, G.; Pukhalskyi, V.A.; Zelenin, A.V.; Bernard, S.; Bernard, M. Chromosomal rearrangements in wheat: Their types and distribution. Genome 2007, 50, 907–926. [Google Scholar] [CrossRef] [PubMed]
- Raskina, O.; Barber, J.C.; Nevo, E.; Belyayev, A. Repetitive DNA and chromosomal rearrangements: Speciation-related events in plant genomes. Cytogenet. Genome Res. 2008, 120, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Alkhimova, A.G.; Heslop-Harrison, J.S.; Shchapova, A.I.; Vershinin, A.V. Rye chromosome variability in wheat-rye addition and substitution lines. Chromosome Res. 1999, 7, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Appels, R.; Gustafson, J.P.; May, C.E. Structural variation in the heterochromatin of rye chromosomes in triticales. Theor. Appl. Genet. 1982, 63, 235–244. [Google Scholar] [CrossRef] [PubMed]
- Bento, M.; Gustafson, P.; Viegas, W.; Silva, M. Genome merger: From sequence rearrangements in triticale to their elimination in wheat-rye addition lines. Theor. Appl. Genet. 2010, 121, 489–497. [Google Scholar] [CrossRef] [PubMed]
- Lapitan, N.L.; Sears, R.G.; Gill, B.S. Translocations and other karyotypic structural changes in wheat x rye hybrids regenerated from tissue culture. TAG. Theor. Appl. Genet. 1984, 68, 547–554. [Google Scholar] [CrossRef] [PubMed]
- Szakacs, E.; Molnar-Lang, M. Molecular cytogenetic evaluation of chromosome instability in Triticum aestivum-Secale cereale disomic addition lines. J. Appl. Genet. 2010, 51, 149–152. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.X.; Fu, S.L.; Ren, Z.L.; Zhou, J.P.; Yan, B.J.; Zhang, H.Q. Variations of tandem repeat, regulatory element, and promoter regions revealed by wheat-rye amphiploids. Genome 2008, 51, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, T. Screening of spontaneous translocations in cultivated emmer wheat. Wheat Inf. Serv. 1997, 85, 45–46. [Google Scholar]
- Chester, M.; Gallagher, J.P.; Symonds, V.V.; Cruz da Silva, A.V.; Mavrodiev, E.V.; Leitch, A.R.; Soltis, P.S.; Soltis, D.E. Extensive chromosomal variation in a recently formed natural allopolyploid species, Tragopogon miscellus (Asteraceae). Proc. Natl. Acad. Sci. USA 2012, 109, 1176–1181. [Google Scholar] [CrossRef] [PubMed]
- Pontes, O.; Neves, N.; Silva, M.; Lewis, M.S.; Madlung, A.; Comai, L.; Viegas, W.; Pikaard, C.S. Chromosomal locus rearrangements are a rapid response to formation of the allotetraploid Arabidopsis suecica genome. Proc. Natl. Acad. Sci. USA 2004, 101, 18240–18245. [Google Scholar] [CrossRef] [PubMed]
- Metcalfe, C.J.; Bulazel, K.V.; Ferreri, G.C.; Schroeder-Reiter, E.; Wanner, G.; Rens, W.; Obergfell, C.; Eldridge, M.D.; O’Neill, R.J. Genomic instability within centromeres of interspecific marsupial hybrids. Genetics 2007, 177, 2507–2517. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.; Yang, M.; Fei, Y.; Tan, F.; Ren, Z.; Yan, B.; Zhang, H.; Tang, Z. Alterations and abnormal mitosis of wheat chromosomes induced by wheat-rye monosomic addition lines. PLoS ONE 2013, 8, e70483. [Google Scholar] [CrossRef] [PubMed]
- Salina, E.A.; Numerova, O.M.; Ozkan, H.; Feldman, M. Alterations in subtelomeric tandem repeats during early stages of allopolyploidy in wheat. Genome 2004, 47, 860–867. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bashir, T.; Chandra Mishra, R.; Hasan, M.M.; Mohanta, T.K.; Bae, H. Effect of Hybridization on Somatic Mutations and Genomic Rearrangements in Plants. Int. J. Mol. Sci. 2018, 19, 3758. https://doi.org/10.3390/ijms19123758
Bashir T, Chandra Mishra R, Hasan MM, Mohanta TK, Bae H. Effect of Hybridization on Somatic Mutations and Genomic Rearrangements in Plants. International Journal of Molecular Sciences. 2018; 19(12):3758. https://doi.org/10.3390/ijms19123758
Chicago/Turabian StyleBashir, Tufail, Ratnesh Chandra Mishra, Md. Mohidul Hasan, Tapan Kumar Mohanta, and Hanhong Bae. 2018. "Effect of Hybridization on Somatic Mutations and Genomic Rearrangements in Plants" International Journal of Molecular Sciences 19, no. 12: 3758. https://doi.org/10.3390/ijms19123758
APA StyleBashir, T., Chandra Mishra, R., Hasan, M. M., Mohanta, T. K., & Bae, H. (2018). Effect of Hybridization on Somatic Mutations and Genomic Rearrangements in Plants. International Journal of Molecular Sciences, 19(12), 3758. https://doi.org/10.3390/ijms19123758