Aeginetia indica Decoction Inhibits Hepatitis C Virus Life Cycle
Abstract
:1. Introduction
2. Results
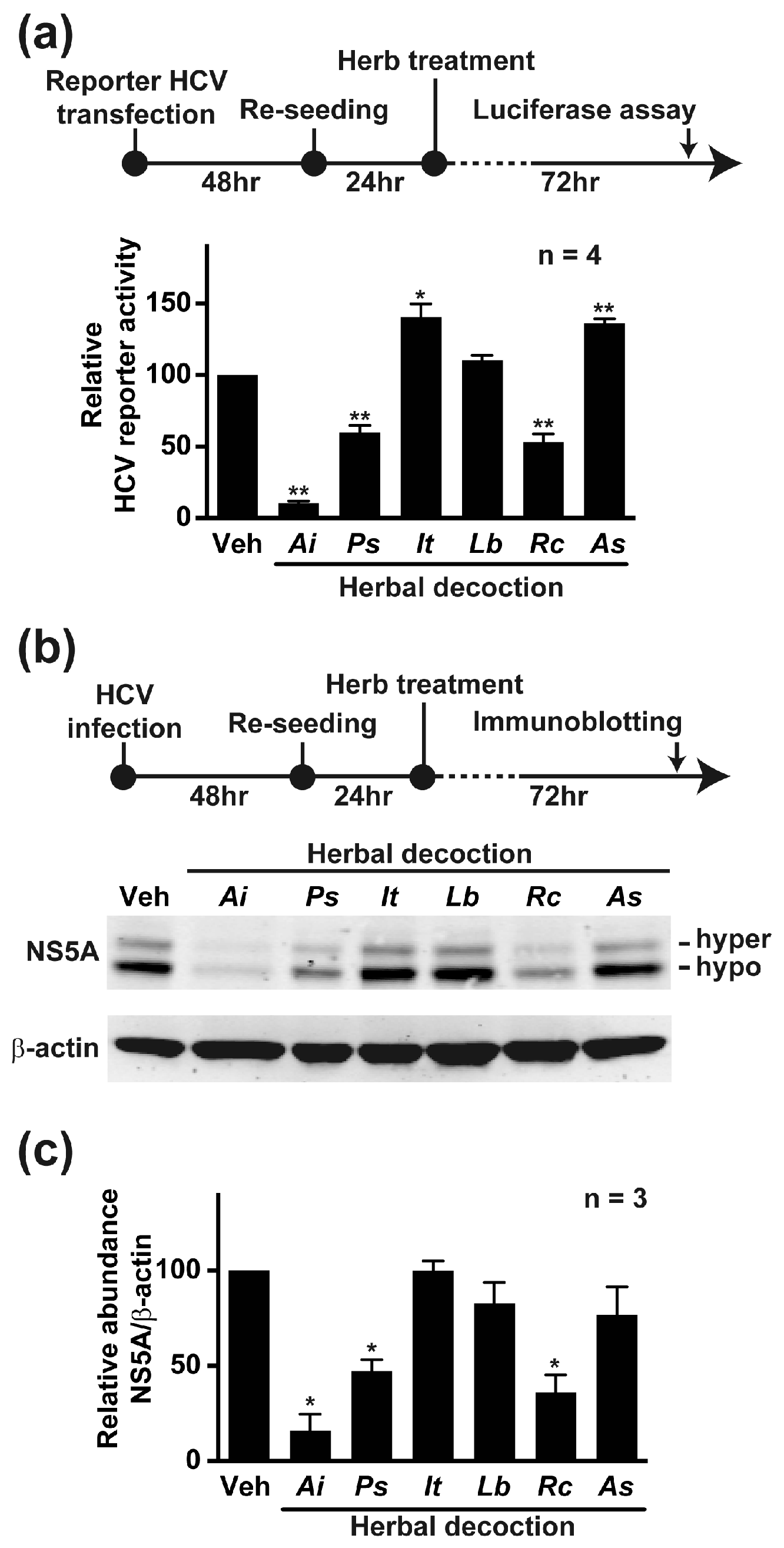
2.1. Aeginetia indica Decoction Inhibited HCV Reporter Activity
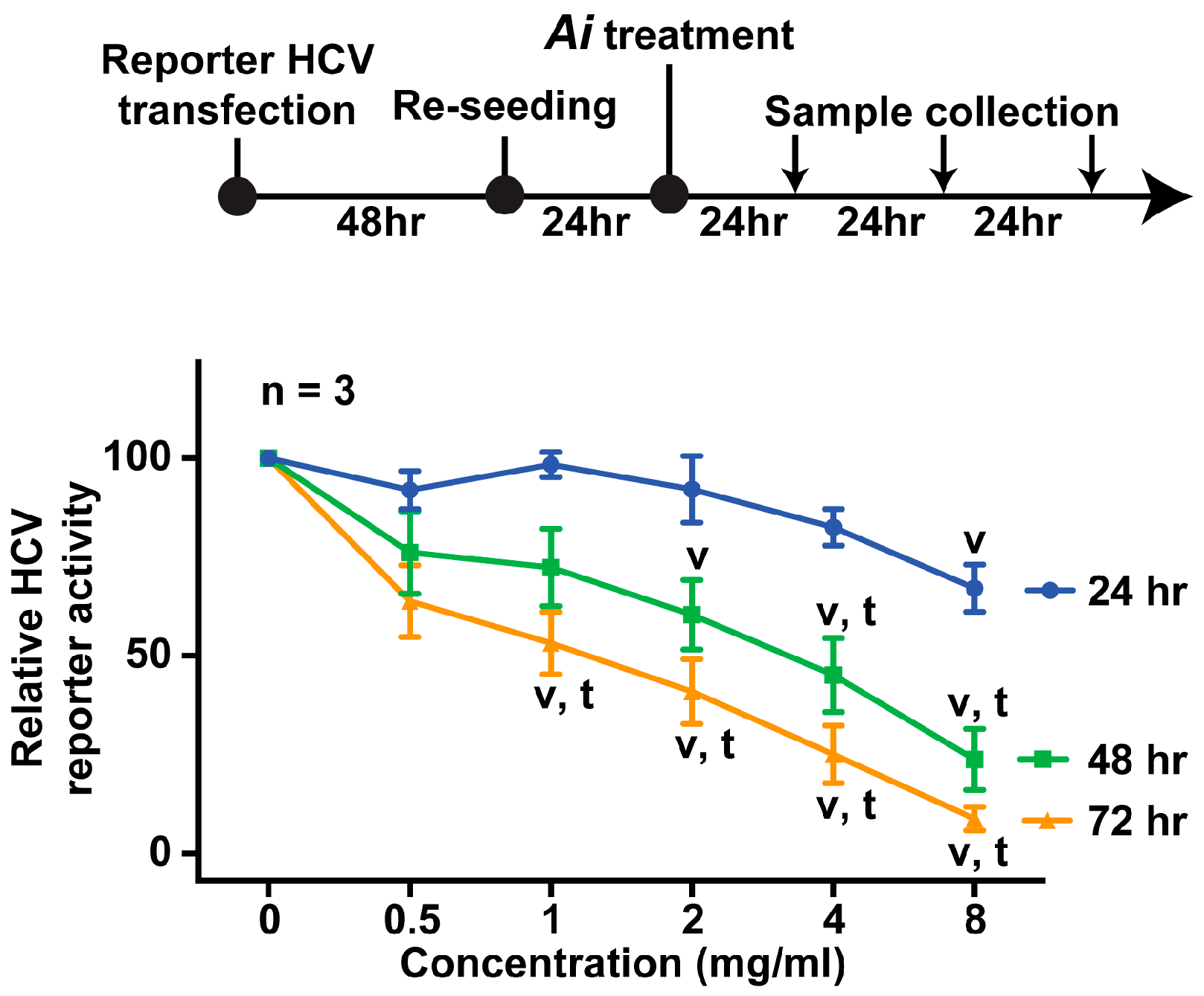
2.2. Aeginetia indica Decoction Dose- and Time-Dependently Inhibited HCV Reporter Activity
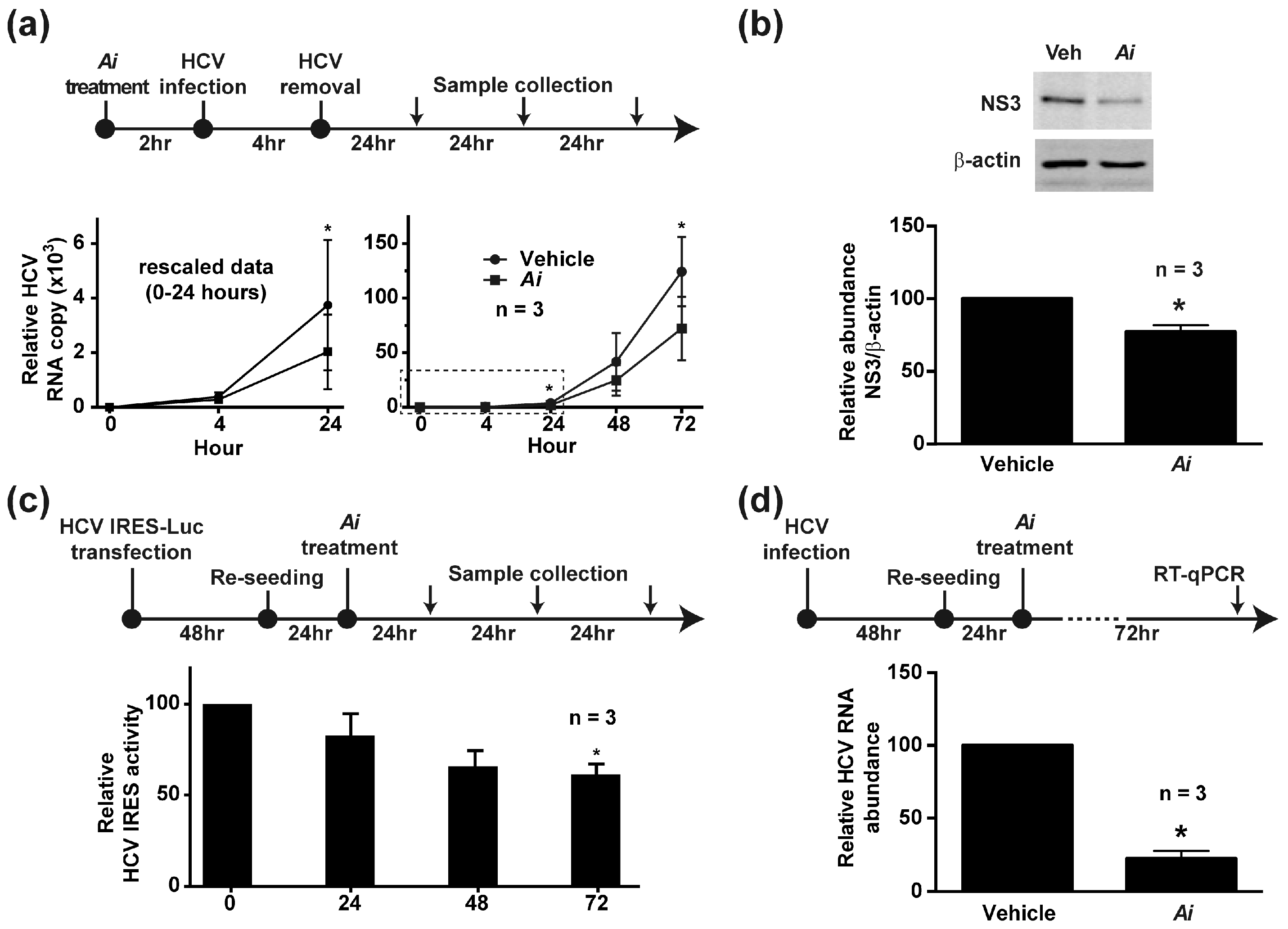
2.3. Pretreating the Cells with Aeginetia indica Decoction Reduced HCV Infection
2.4. Aeginetia indica Decoction Inhibited HCV IRES-Mediated Protein Translation
2.5. Aeginetia indica Decoction Reduced HCV RNA in the Infected Cells
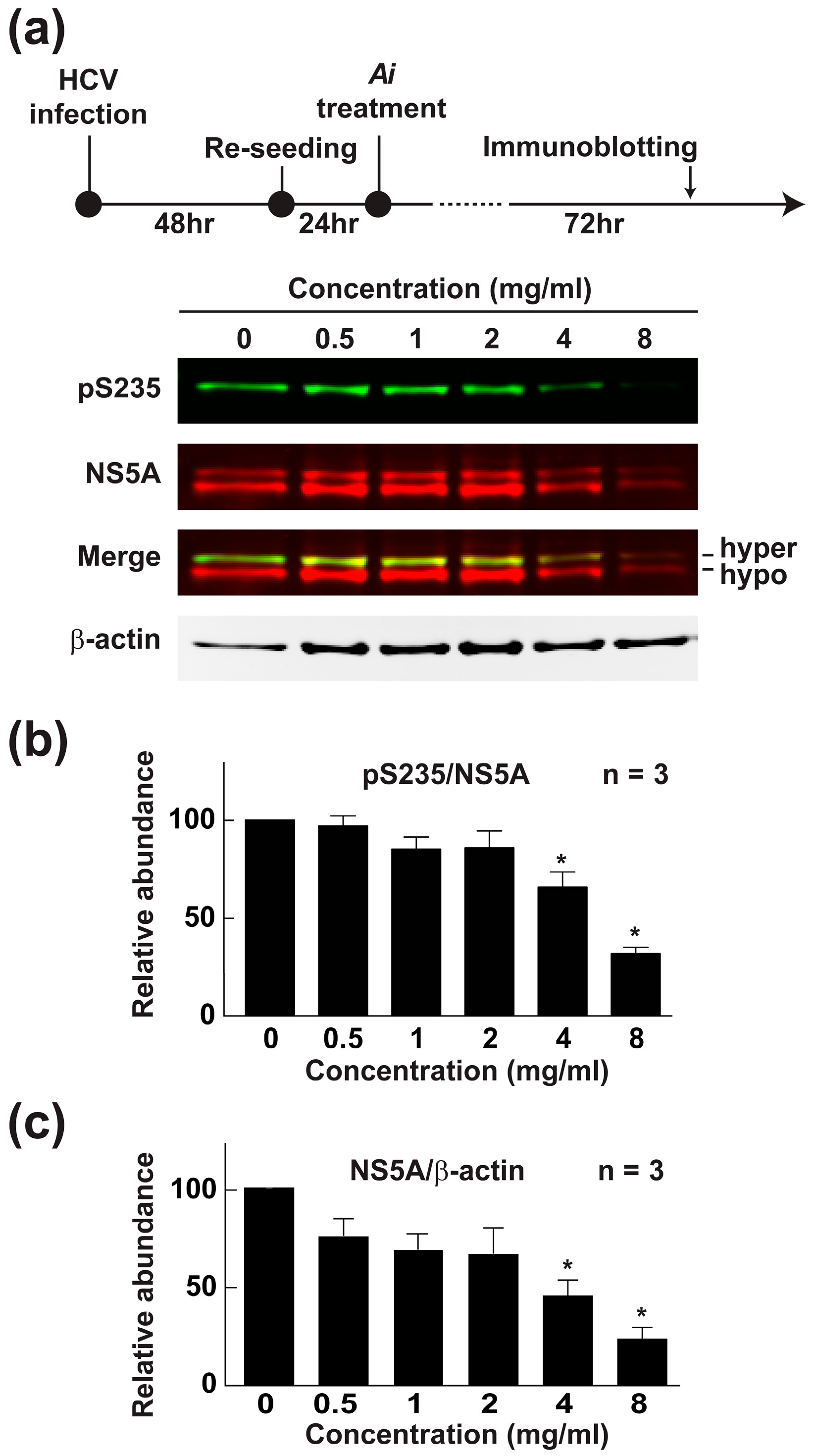
2.6. Aeginetia indica Decoction Reduced NS5A and NS5A Phosphorylation at Serine 235
3. Discussion
4. Materials and Methods
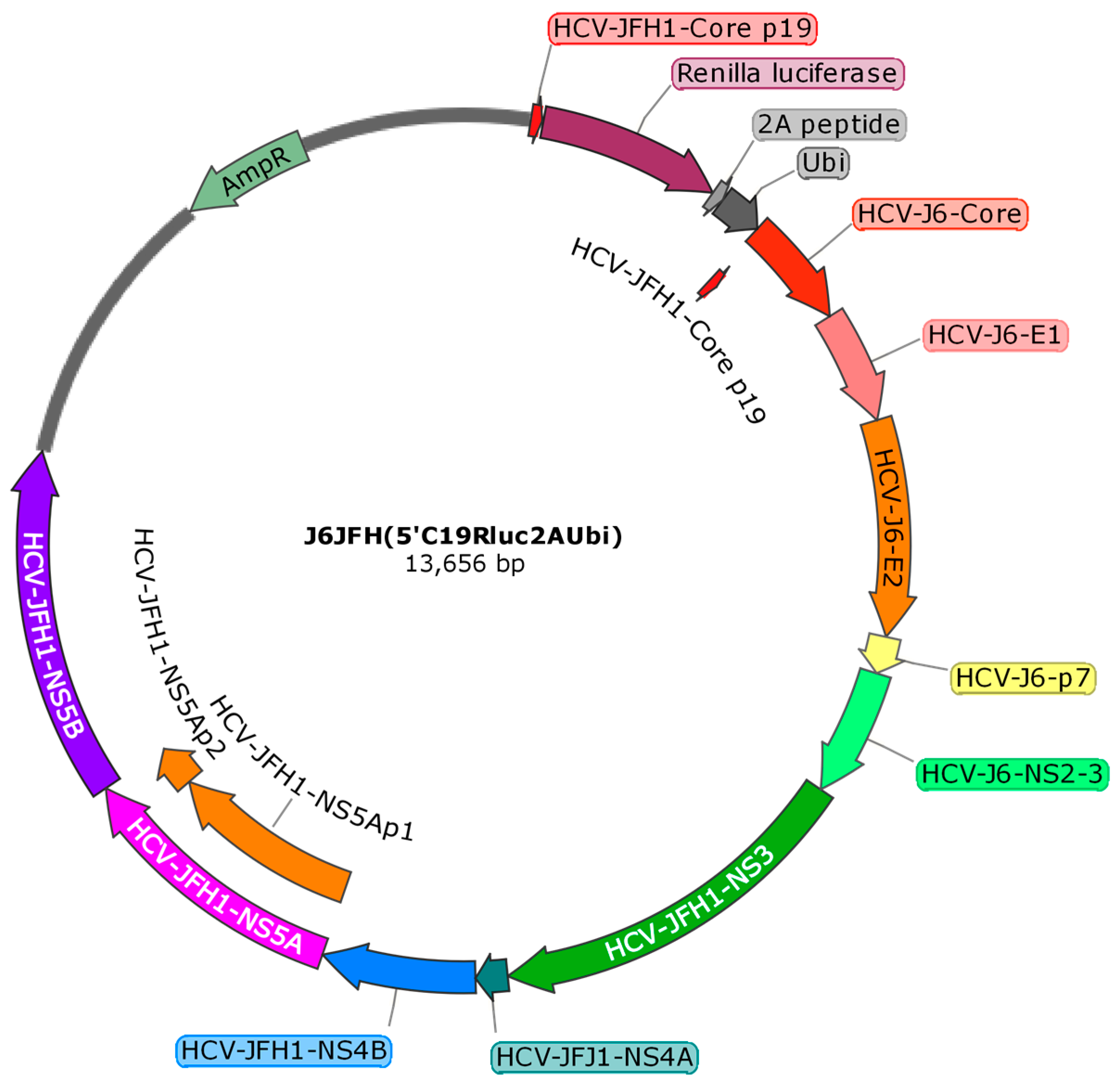
4.1. Cells and HCV Constructs
4.2. Herbal Decoction
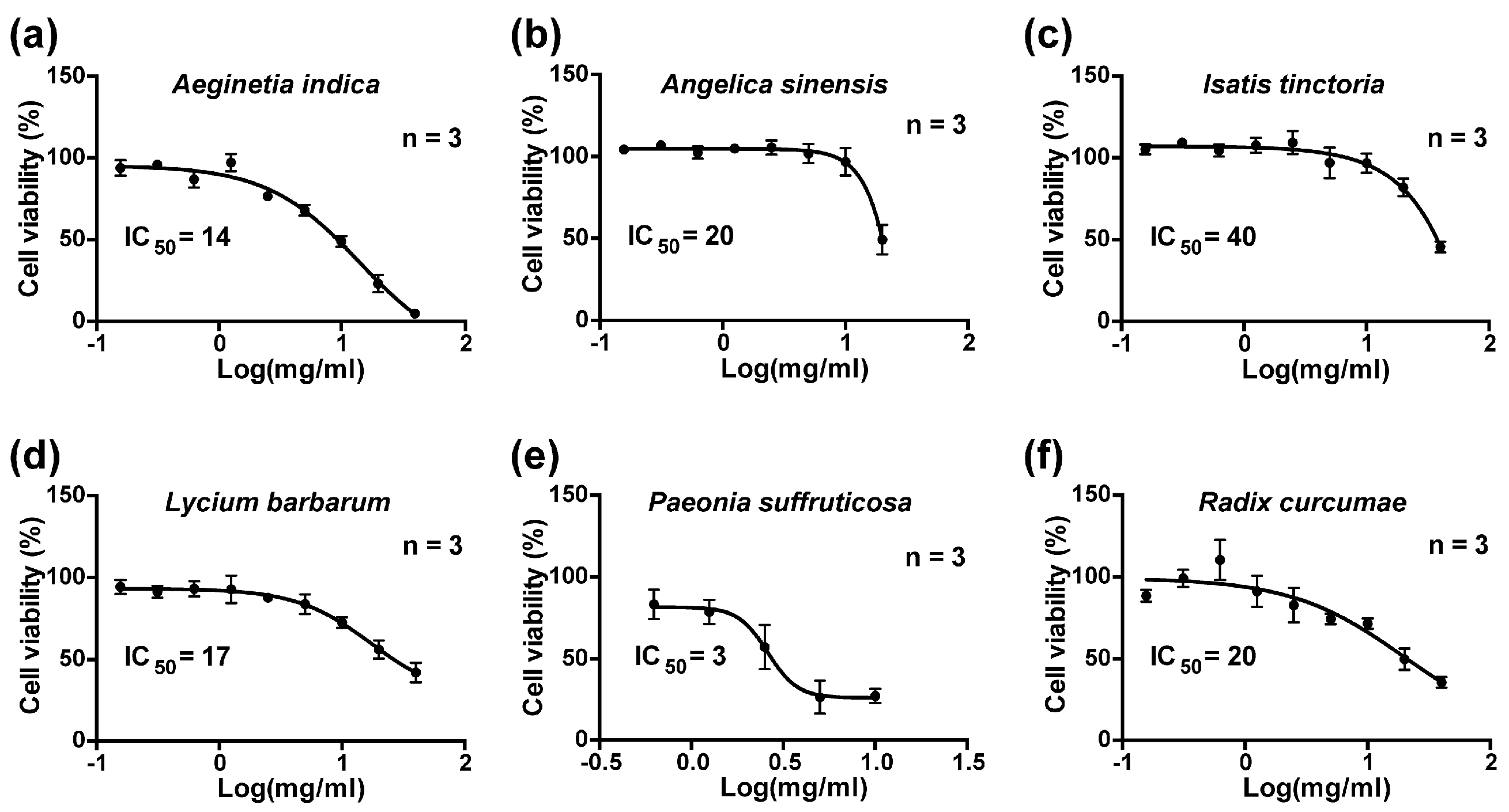
4.3. Cell Viability Assay
4.4. HCV RNA Preparation
4.5. HCV Reporter Activity Assay
4.6. Infectious HCV Virion Production
4.7. Immunoblotting
4.8. HCV IRES-Mediated Translation Activity Assay
4.9. HCV RNA Measurement
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Scheel, T.K.; Rice, C.M. Understanding the hepatitis C virus life cycle paves the way for highly effective therapies. Nat. Med. 2013, 19, 837–849. [Google Scholar] [CrossRef] [PubMed]
- Bartenschlager, R.; Lohmann, V.; Penin, F. The molecular and structural basis of advanced antiviral therapy for hepatitis C virus infection. Nat. Rev. Microbiol. 2013, 11, 482–496. [Google Scholar] [CrossRef] [PubMed]
- Pawlotsky, J.M. The end of the hepatitis c burden: Really? Hepatology 2016, 64, 1404–1407. [Google Scholar] [CrossRef] [PubMed]
- McCauley, J.A.; Rudd, M.T. Hepatitis C virus NS3/4a protease inhibitors. Curr. Opin. Pharmacol. 2016, 30, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Ross-Thriepland, D.; Harris, M. Hepatitis C virus NS5A: Enigmatic but still promiscuous 10 years on! J. Gen. Virol. 2015, 96, 727–738. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.; O’Boyle, D.R., II; Roberts, S. HCV NS5A replication complex inhibitors. Curr. Opin. Pharmacol. 2016, 30, 151–157. [Google Scholar] [CrossRef] [PubMed]
- Keating, G.M. Ledipasvir/sofosbuvir: A review of its use in chronic hepatitis C. Drugs 2015, 75, 675–685. [Google Scholar] [CrossRef] [PubMed]
- Masaki, T.; Matsunaga, S.; Takahashi, H.; Nakashima, K.; Kimura, Y.; Ito, M.; Matsuda, M.; Murayama, A.; Kato, T.; Hirano, H.; et al. Involvement of hepatitis C virus NS5A hyperphosphorylation mediated by casein kinase i-alpha in infectious virus production. J. Virol. 2014, 88, 7541–7555. [Google Scholar] [CrossRef] [PubMed]
- Fridell, R.A.; Valera, L.; Qiu, D.; Kirk, M.J.; Wang, C.; Gao, M. Intragenic complementation of hepatitis C virus NS5A RNA replication-defective alleles. J. Virol. 2013, 87, 2320–2329. [Google Scholar] [CrossRef] [PubMed]
- Chong, W.M.; Hsu, S.C.; Kao, W.T.; Lo, C.W.; Lee, K.Y.; Shao, J.S.; Chen, Y.H.; Chang, J.; Chen, S.S.; Yu, M.J. Phosphoproteomics identified an NS5A phosphorylation site involved in hepatitis C virus replication. J. Biol. Chem. 2016, 291, 3918–3931. [Google Scholar] [CrossRef] [PubMed]
- Eyre, N.S.; Hampton-Smith, R.J.; Aloia, A.L.; Eddes, J.S.; Simpson, K.J.; Hoffmann, P.; Beard, M.R. Phosphorylation of NS5A serine-235 is essential to hepatitis C virus RNA replication and normal replication compartment formation. Virology 2016, 491, 27–44. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.C.; Lo, C.W.; Pan, T.C.; Lee, K.Y.; Yu, M.J. Serine 235 is the primary ns5a hyper-phosphorylation site responsible for hcv replication. J. Virol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.Y.; Chen, Y.H.; Hsu, S.C.; Yu, M.J. Phosphorylation of serine 235 of the hepatitis C virus non-structural protein NS5A by multiple kinases. PLoS ONE 2016, 11, e0166763. [Google Scholar] [CrossRef] [PubMed]
- Patil, V.M.; Masand, N.; Gupta, S.P. HCV inhibitors: Role of compounds from botanical sources. Curr. Top. Med. Chem. 2016, 16, 1402–1409. [Google Scholar] [CrossRef] [PubMed]
- Calland, N.; Dubuisson, J.; Rouille, Y.; Seron, K. Hepatitis C virus and natural compounds: A new antiviral approach? Viruses 2012, 4, 2197–2217. [Google Scholar] [CrossRef] [PubMed]
- Ho, J.C.; Chen, C.M.; Li, Z.Q.; Row, L.C. Phenylpropanoid glycosides from the parasitic plant, Aeginetia indica. J. Chin. Chem. Soc. 2004, 51, 1073–1076. [Google Scholar] [CrossRef]
- Liu, Y.H.; Li, M.L.; Hsu, M.Y.; Pang, Y.Y.; Chen, I.L.; Chen, C.K.; Tang, S.W.; Lin, H.Y.; Lin, J.Y. Effects of a chinese herbal medicine, Guan-Jen-Huang (Aeginetia indica linn.), on renal cancer cell growth and metastasis. Evid.-Based Complement. Altern. Med. 2012, 2012, 935860. [Google Scholar]
- Wu, S.J.; Ng, L.T.; Lin, C.C. Antioxidant activities of some common ingredients of traditional chinese medicine, angelica sinensis, lycium barbarum and poria cocos. Phytother. Res. 2004, 18, 1008–1012. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.M.; Tsai, F.J.; Tsai, Y. Inhibitory effect of angelica sinensis extract in the presence of 2-hydroxypropyl-β-cyclodextrin. Carbohydr. Polym. 2014, 114, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.W.; Chen, Y.M.; Hsu, Y.J.; Huang, C.C.; Wu, Y.T.; Hsu, M.C. Protective effects of the roots of angelica sinensis on strenuous exercise-induced sports anemia in rats. J. Ethnopharmacol. 2016, 193, 169–178. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.S.; Wu, S.J.; Tsai, Y.H.; Lin, Y.H.; Chao, J.C. Hot water extracted lycium barbarum and rehmannia glutinosa inhibit liver inflammation and fibrosis in rats. Am. J. Chin. Med. 2011, 39, 1173–1191. [Google Scholar] [CrossRef] [PubMed]
- Jia, L.; Li, W.; Li, J.; Li, Y.; Song, H.; Luan, Y.; Qi, H.; Ma, L.; Lu, X.; Yang, Y. Lycium barbarum polysaccharide attenuates high-fat diet-induced hepatic steatosis by up-regulating SIRT1 expression and deacetylase activity. Sci. Rep. 2016, 6, 36209. [Google Scholar] [CrossRef] [PubMed]
- Ha do, T.; Phuong, T.T.; Oh, J.; Bae, K.; Thuan, N.D.; Na, M. Palbinone from paeonia suffruticosa protects hepatic cells via up-regulation of heme oxygenase-1. Phytother. Res. 2014, 28, 308–311. [Google Scholar] [PubMed]
- Zhou, Y.; Xie, M.; Song, Y.; Wang, W.; Zhao, H.; Tian, Y.; Wang, Y.; Bai, S.; Zhao, Y.; Chen, X.; et al. Two traditional chinese medicines curcumae radix and curcumae rhizoma: An ethnopharmacology, phytochemistry, and pharmacology review. Evid.-Based Complement. Altern. Med. 2016, 2016, 4973128. [Google Scholar] [CrossRef] [PubMed]
- Kumagai, T.; Aratsu, Y.; Sugawara, R.; Sasaki, T.; Miyairi, S.; Nagata, K. Indirubin, a component of Ban-Lan-Gen, activates CYP3A4 gene transcription through the human pregnane X receptor. Drug Metab. Pharmacokinet. 2016, 31, 139–145. [Google Scholar] [CrossRef] [PubMed]
- Galani Tietcheu, B.R.; Sass, G.; Njayou, N.F.; Mkounga, P.; Tiegs, G.; Moundipa, P.F. Anti-hepatitis C virus activity of crude extract and fractions of entada africana in genotype 1b replicon systems. Am. J. Chin. Med. 2014, 42, 853–868. [Google Scholar] [CrossRef] [PubMed]
- Wagoner, J.; Negash, A.; Kane, O.J.; Martinez, L.E.; Nahmias, Y.; Bourne, N.; Owen, D.M.; Grove, J.; Brimacombe, C.; McKeating, J.A.; et al. Multiple effects of silymarin on the hepatitis C virus lifecycle. Hepatology 2010, 51, 1912–1921. [Google Scholar] [CrossRef] [PubMed]
- Gordon, A.; Hobbs, D.A.; Bowden, D.S.; Bailey, M.J.; Mitchell, J.; Francis, A.J.; Roberts, S.K. Effects of silybum marianum on serum hepatitis C virus RNA, alanine aminotransferase levels and well-being in patients with chronic hepatitis C. J. Gastroenterol. Hepatol. 2006, 21, 275–280. [Google Scholar] [CrossRef] [PubMed]
- Iwashina, T. Flavonoids from two parasitic and achlorophyllous plants, Aeginetia indica and Orobanche minor (orabanchaceae). Bull. Natl. Mus. Nat. Sci. Ser. B 2010, 36, 127–132. [Google Scholar]
- Chai, J.G.; Bando, T.; Kobashi, S.; Oka, M.; Nagasawa, H.; Nakai, S.; Maeda, K.; Himeno, K.; Sato, M.; Ohkubo, S. An extract of seeds from Aeginetia indica L., a parasitic plant, induces potent antigen-specific antitumor immunity in Meth A-bearing BALB/c mice. Cancer Immunol. Immunother. 1992, 35, 181–185. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.C.; Chen, W.C.; Wu, S.F.; Tseng, C.K.; Chiou, C.Y.; Chang, F.R.; Hsu, S.H.; Wu, Y.C. Anti-hepatitis C virus activity of acacia confusa extract via suppressing cyclooxygenase-2. Antivir. Res. 2011, 89, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Trujillo-Murillo, K.; Rincon-Sanchez, A.R.; Martinez-Rodriguez, H.; Bosques-Padilla, F.; Ramos-Jimenez, J.; Barrera-Saldana, H.A.; Rojkind, M.; Rivas-Estilla, A.M. Acetylsalicylic acid inhibits hepatitis C virus RNA and protein expression through cyclooxygenase 2 signaling pathways. Hepatology 2008, 47, 1462–1472. [Google Scholar] [CrossRef] [PubMed]
- Shibata, C.; Ohno, M.; Otsuka, M.; Kishikawa, T.; Goto, K.; Muroyama, R.; Kato, N.; Yoshikawa, T.; Takata, A.; Koike, K. The flavonoid apigenin inhibits hepatitis C virus replication by decreasing mature microrna122 levels. Virology 2014, 462–463, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Jopling, C.L.; Yi, M.; Lancaster, A.M.; Lemon, S.M.; Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific microrna. Science 2005, 309, 1577–1581. [Google Scholar] [CrossRef] [PubMed]
- Manvar, D.; Mishra, M.; Kumar, S.; Pandey, V.N. Identification and evaluation of anti hepatitis C virus phytochemicals from eclipta alba. J. Ethnopharmacol. 2012, 144, 545–554. [Google Scholar] [CrossRef] [PubMed]
- Um, M.Y.; Ahn, J.; Ha, T.Y. Hypolipidaemic effects of cyanidin 3-glucoside rich extract from black rice through regulating hepatic lipogenic enzyme activities. J. Sci. Food Agric. 2013, 93, 3126–3128. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Pene, V.; Krishnamurthy, S.; Cha, H.; Liang, T.J. Hepatitis c virus infection activates an innate pathway involving ikk-α in lipogenesis and viral assembly. Nat. Med. 2013, 19, 722–729. [Google Scholar] [CrossRef] [PubMed]
- Tscherne, D.M.; Jones, C.T.; Evans, M.J.; Lindenbach, B.D.; McKeating, J.A.; Rice, C.M. Time- and temperature-dependent activation of hepatitis C virus for low-ph-triggered entry. J. Virol. 2006, 80, 1734–1741. [Google Scholar] [CrossRef] [PubMed]
- Kalliampakou, K.I.; Kalamvoki, M.; Mavromara, P. Hepatitis C virus (HCV) NS5A protein downregulates HCV IRES-dependent translation. J. Gen. Virol. 2005, 86, 1015–1025. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, C.-W.; Lo, C.-W.; Tsai, C.-N.; Pan, T.-C.; Chen, P.-Y.; Yu, M.-J. Aeginetia indica Decoction Inhibits Hepatitis C Virus Life Cycle. Int. J. Mol. Sci. 2018, 19, 208. https://doi.org/10.3390/ijms19010208
Lin C-W, Lo C-W, Tsai C-N, Pan T-C, Chen P-Y, Yu M-J. Aeginetia indica Decoction Inhibits Hepatitis C Virus Life Cycle. International Journal of Molecular Sciences. 2018; 19(1):208. https://doi.org/10.3390/ijms19010208
Chicago/Turabian StyleLin, Cheng-Wei, Chieh-Wen Lo, Chia-Ni Tsai, Ting-Chun Pan, Pin-Yin Chen, and Ming-Jiun Yu. 2018. "Aeginetia indica Decoction Inhibits Hepatitis C Virus Life Cycle" International Journal of Molecular Sciences 19, no. 1: 208. https://doi.org/10.3390/ijms19010208
APA StyleLin, C.-W., Lo, C.-W., Tsai, C.-N., Pan, T.-C., Chen, P.-Y., & Yu, M.-J. (2018). Aeginetia indica Decoction Inhibits Hepatitis C Virus Life Cycle. International Journal of Molecular Sciences, 19(1), 208. https://doi.org/10.3390/ijms19010208




