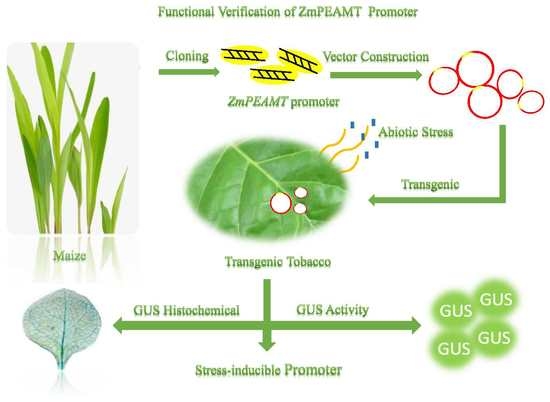
Cloning and Functional Analysis of Phosphoethanolamine Methyltransferase Promoter from Maize (Zea mays L.)
Abstract
:1. Introduction
2. Results
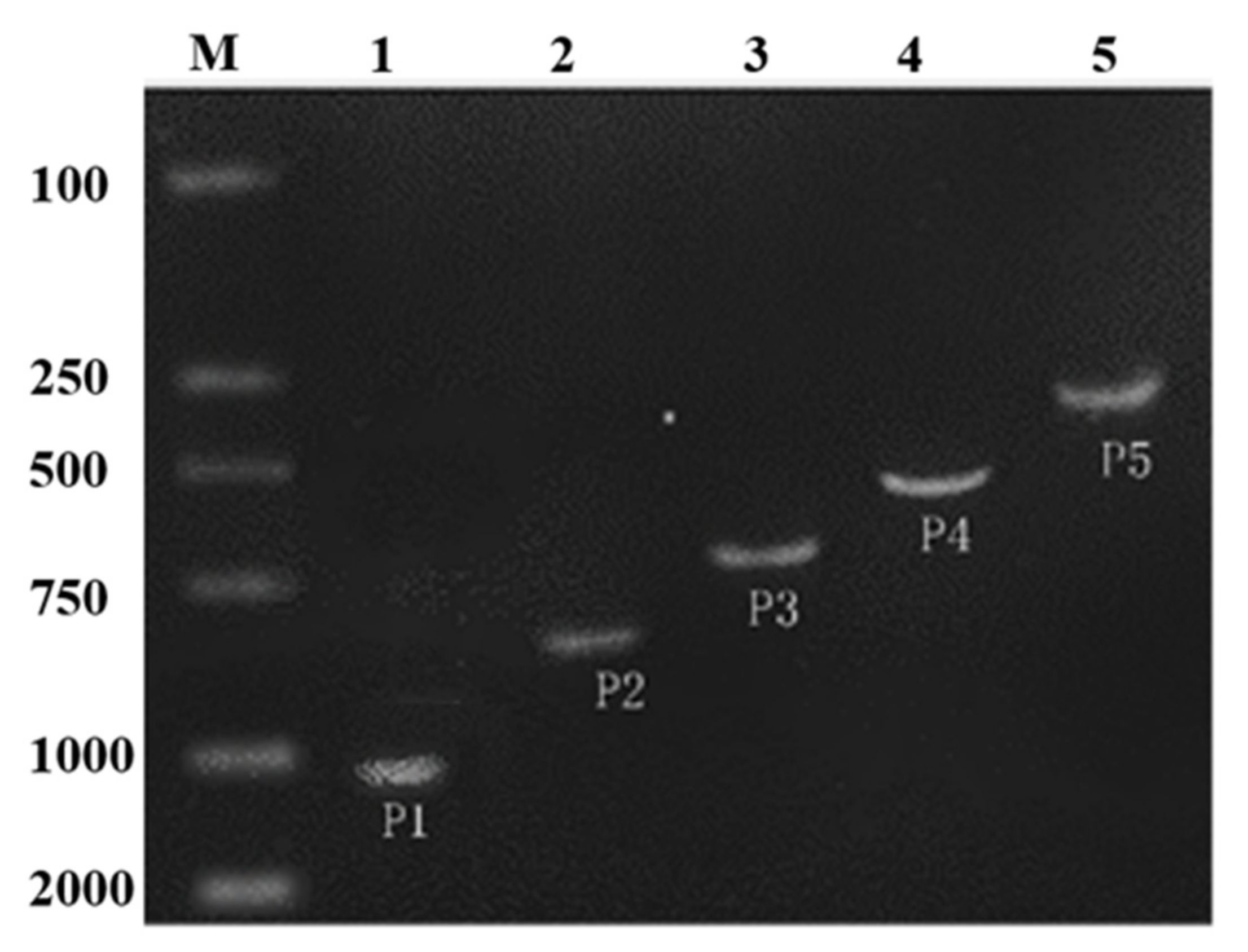
2.1. Cloning of ZmPEAMT-P and In Silico Analysis
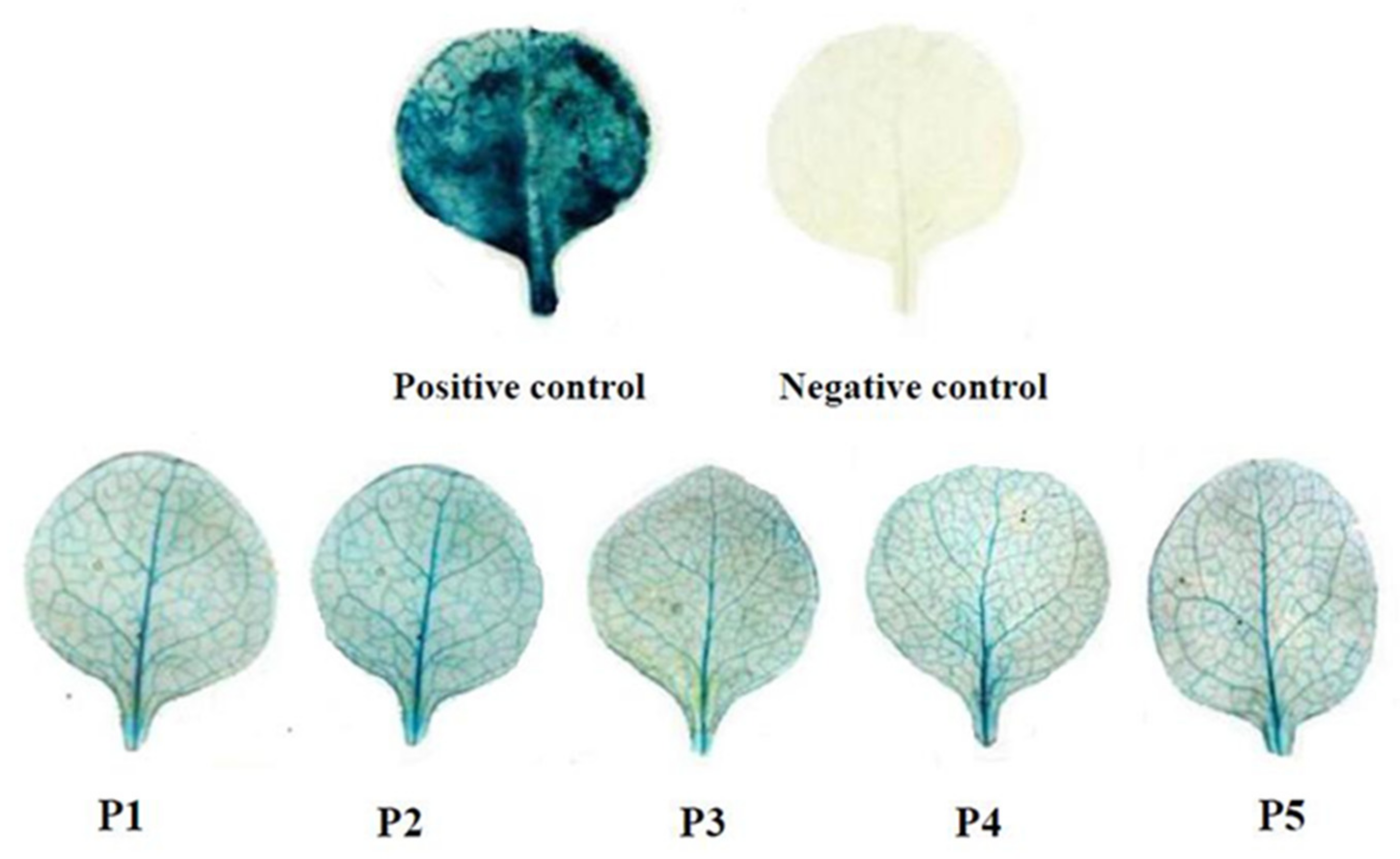
2.2. Promoter Deletion Constructs and Histochemical GUS Assay
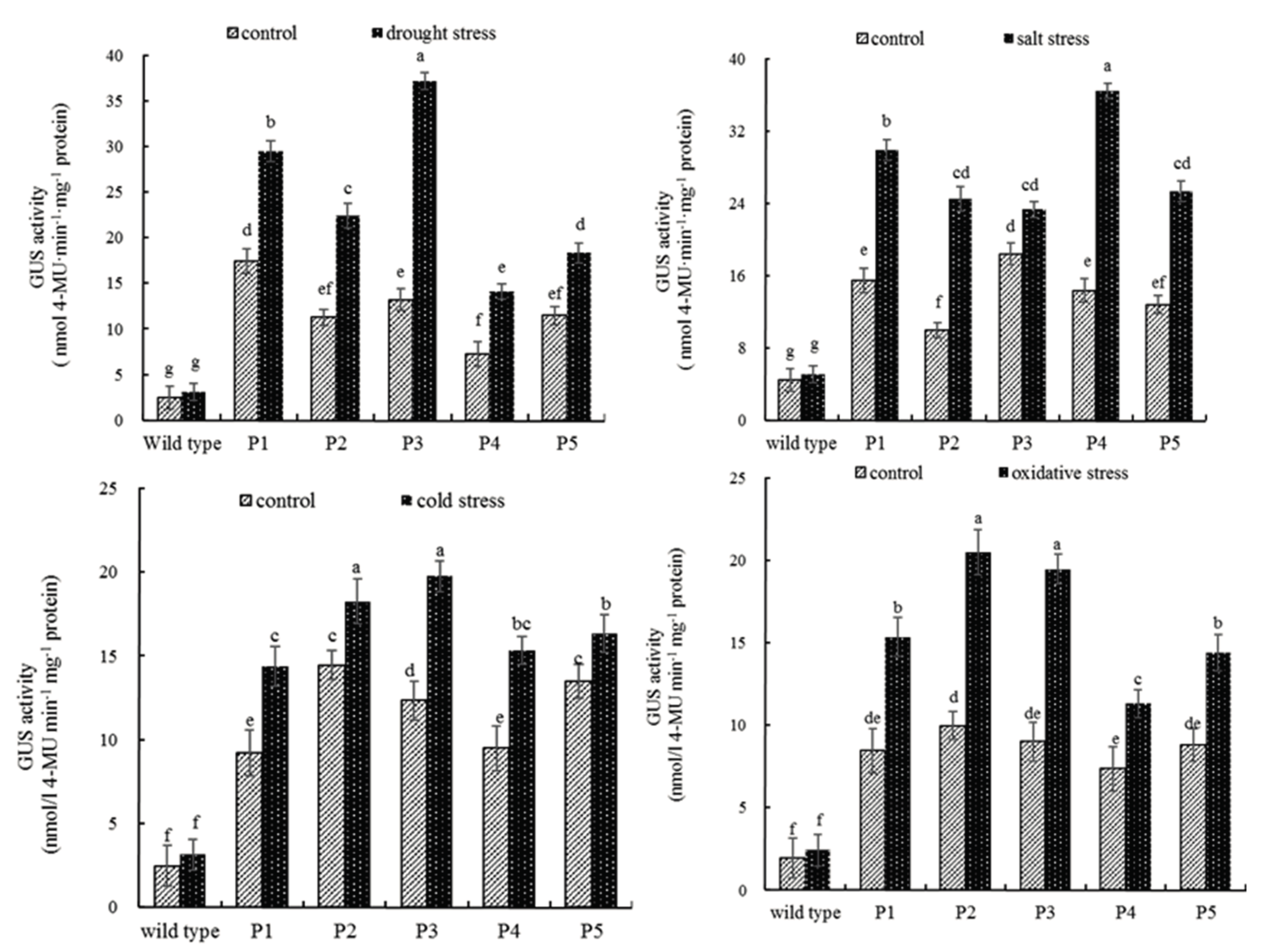
2.3. Quantitative MUG Assays of PEAMT Gene Promoter under Abiotic Stress Conditions
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Isolation of 5′-Upstream Cis-Acting Elements and In Silico Analysis
4.3. Construction of ZmPEAMT Promoter Expression Vector and Tobacco Transformation
4.4. GUS Histochemical
4.5. Quantitative Assay under Different Stress Conditions
4.6. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| PEAMT | Phosphoethanolamine N-Methyltransferase |
| GUS | Escherichia Coli β-Glucuronidase gene |
| P-Etn | Phosphoethanolamine |
| P-Cho | Phosphocholine |
| P-MMEtn | Phosphomonomethylethanolamine |
| Wt | Wild Type |
| AtADH1 | Alcohol Dehydrogenase |
| CTAB | Cetyltrimethylammonium Bromide |
References
- Scholz, A.; Stahl, J.; De, B.V.; Müller, V.; Averhoff, B. Osmotic stress response in acinetobacter baylyi: Identification of a glycine-betaine biosynthesis pathway and regulation of osmoadaptive choline uptake and glycine-betaine synthesis through a choline-responsive BetI repressor. Env. Microbiol. Rep. 2016, 8, 316–322. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.L.; Xie, J.H.; Ma, X.Q.; Li, D. Molecular cloning of Phosphoethanolamine N-methyltransferase (PEAMT) gene and its promoter from the halophyte Suaeda liaotungensis and their response to salt stress. Acta Physiol. Plant 2016, 38, 39. [Google Scholar] [CrossRef]
- BeGora, M.D.; Macleod, M.J.R.; McCarry, B.E.; Summers, P.S.; Weretilnyk, E.A. Identification of phosphomethylethanolamine N-methyltransferase from Arabidopsis and its role in choline and phospholipid metabolism. J. Biol. Chem. 2010, 285, 29147–29155. [Google Scholar] [CrossRef] [PubMed]
- Hirashima, T.; Toyoshima, M.; Moriyama, T.; Nakamura, Y.; Sato, N. Characterization of phosphoethanolamine-N-methyltransferases in green algae. Biochem. Bioph. Res. Commun. 2017, 488, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Bolognese, C.P.; Mcgraw, P. The isolation and characterization in yeast of a gene for Arabidopsis S-adenosylmethionine: Phospho-ethanolamine N-methyltransferase. Plant Physiol. 2000, 124, 1800–1813. [Google Scholar] [CrossRef] [PubMed]
- Nuccio, M.L.; Ziemak, M.J.; Henry, S.A.; Weretilnyk, E.A.; Hanson, A.D. cDNA cloning of phosphoethanolamine N-methyltransferase from Spinach by complementation in Schizosac charomyces pombe and characterization of the recombinant enzyme. J. Biol. Chem. 2000, 275, 14095–14101. [Google Scholar] [CrossRef] [PubMed]
- Charron, J.B.F.; Breton, G.; Danyluk, J.; Muzac, I.; Ibrahim, R.K.; Sarhan, F. Molecular and biochemical characterization of a cold-regulated phosphoethanolamine N-methyltransferase from wheat. Plant Physiol. 2002, 129, 363–373. [Google Scholar] [CrossRef] [PubMed]
- Tabuchi, T.; Kawaguchi, Y.; Azuma, T.; Nanmori, T.; Yasuda, T. Similar regulation patterns of choline monooxygenase, phosphoethanolamine N-methyltransferase and S-adenosyl-l-methionine synthetase in leaves of the halophyte Atriplex nummularia L. Plant Cell Physiol. 2005, 46, 505–513. [Google Scholar] [CrossRef] [PubMed]
- Tabuchi, T.; Okada, T.; Takashima, Y.; Azuma, T.; Nanmori, T.; Yasuda, T. Transcriptional response of glycine betaine-related genes to salt stress and light in leaf beet. Plant Biotechnol. J. 2006, 23, 317–320. [Google Scholar] [CrossRef]
- Wu, S.; Yu, Z.; Wang, F.; Li, W.; Ye, C.; Li, J.; Tang, J.H.; Ding, J.Q.; Zhao, J.R.; Wang, B. Cloning, characterization, and transformation of the phosphoethanolamine N-methyltransferase gene (ZmPEAMT) in maize (Zea mays L.). Mol. Biotechnol. 2007, 36, 102–112. [Google Scholar] [CrossRef] [PubMed]
- Jost, R.; Berkowitz, O.; Shaw, J.; Masle, J. Biochemical characterization of two wheat phosphoethanolamine N-methyltransferase isoforms with different sensitivities to inhibition by phosphatidic acid. J. Biol. Chem. 2009, 284, 31962–31971. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Murzello, C.; Sun, Y.; Kim, M.S.; Xie, X.; Jeter, R.M.; Zak, J.C.; Dowd, S.C.; Paré, P.W. Choline and osmotic-stress tolerance induced in Arabidopsis by the soil microbe Bacillus subtilis (GB03). Mol. Plant Microbe Interact. 2010, 23, 1097–1104. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Chen, C.; Chen, Z. Expression profiles of the arabidopsis wrky gene superfamily during plant defense response. Plant Mol. Biol. 2003, 51, 21–37. [Google Scholar]
- Sakuma, Y.; Maruyama, K.; Osakabe, Y.; Qin, F.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozakia, K. Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 2006, 18, 1292–1309. [Google Scholar] [CrossRef] [PubMed]
- Ohmetakagi, M.; Suzuki, K.; Shinshi, H. Regulation of ethylene-induced transcription of defense genes. Plant Cell Physiol. 2000, 41, 1187–1192. [Google Scholar] [CrossRef]
- Sharma, R.; Rawat, V.; Suresh, C.G. Genome-wide identification and tissue-specific expression analysis of nucleotide binding site-leucine rich repeat gene family in Cicer arietinum (kabuli chickpea). Genomics Data. 2017, 14, 24–31. [Google Scholar] [CrossRef] [PubMed]
- BhatnagarMathur, P.; Vadez, V.; Sharma, K.K. Transgenic approaches for abiotic stress tolerance in plants: Retrospect and prospects. Plant Cell Rep. 2008, 27, 411–424. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, V.; Patel, M.K.; Chaturvedi, A.K.; Mishra, A.; Jha, B. Functional Characterization of the Tau class glutathione-s-transferases gene (SbGSTU) promoter of Salicornia brachiata under salinity and osmotic stress. PLoS ONE 2016, 11, e0148494. [Google Scholar] [CrossRef] [PubMed]
- Ye, C.; Wu, S.; Yang, Q.; Ma, C.; Yang, G.; Wang, B. Cloning, sequencing and salt induced expression of peamt and badh in oilseed rape (brassica napus). DNA Seq. 2005, 16, 364–371. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Garcia, C.M.; Finer, J.J. Identification and validation of promoters and cis-acting regulatory elements. Plant Sci. 2014, 217, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.B.; Luo, L.; He, L.L.; Ni, J.; Xu, Z.F. A promoter analysis of mother of ft and rft11 (JcMFT1), a seed-preferential gene from the biofuel plant Jatropha curcas. J. Plant Res. 2014, 127, 513–524. [Google Scholar] [CrossRef] [PubMed]
- Peng, X.; Liu, H.; Wang, D.; Shen, S. Genome-wide identification of the Jatropha curcas MYB family and functional analysis of the abiotic stress responsive gene JcMYB2. BMC Genom. 2016, 17, 251. [Google Scholar] [CrossRef] [PubMed]
- Roy, S. Function of MYB domain transcription factors in abiotic stress and epigenetic control of stress response in plant genome. Plant Signal. Behav. 2016, 11, e1117723. [Google Scholar] [CrossRef] [PubMed]
- Bruxelles, G.L.D.; Peacock, W.J.; Dennis, E.S.; Dolferus, R. Abscisic Acid Induces the Alcohol Dehydrogenase Gene in Arabidopsis. Plant Physiol. 1996, 111, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Guiltinan, M.J.; Marcotte, W.R., Jr.; Quatrano, R.S. A plant leucine zipper protein that recognizes an abscisic acid response element. Science 1990, 250, 267. [Google Scholar] [CrossRef] [PubMed]
- Narusaka, Y.; Nakashima, K.; Shinwari, Z.K.; Sakuma, Y.; Furihata, T.; Abe, H.; Narusaka, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J. 2003, 34, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Bang, S.W.; Jeong, J.S.; Jung, H.; Redillas, M.C.F.R.; Kim, H.I.; Lee, L.H.; Kim, Y.S.; Kim, J.K. Analysis of the APX, PGD1 and R1G1B constitutive gene promoters in various organs over three homozygous generations of transgenic rice plants. Planta 2012, 235, 1397–1408. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.; Roychoudhury, A. WRKY proteins: Signaling and regulation of expression during abiotic stress responses. Sci. World J. 2015, 2015, 807560. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Yoshida, T.; Yamaguchi-Shinozaki, K. Pivotal role of the AREB/ABF-SnRK2 pathway in ABRE-mediated transcription in response to osmotic stress in plants. Physiol. Plantarum. 2013, 147, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Kakali, M.; Roy, C.A.; Bhaskar, G.; Sudhiranjan, G.; Sengupta, D. An abre-binding factor, osbz8, is highly expressed in salt tolerant cultivars than in salt sensitive cultivars of indica rice. BMC Plant Biol. 2006, 6, 1–14. [Google Scholar]
- Chinnusamy, V.; Schumaker, K.; Zhu, J.K. Molecular genetic perspectives on cross-talk and specificity in abiotic stress signalling in plants. J. Exp. Bot. 2004, 55, 225–236. [Google Scholar] [CrossRef] [PubMed]
- Mao, X.; Jia, D.; Li, A.; Zhang, H.; Tian, S.; Zhang, X.; Jing, R. Transgenic expression of TaMYB2A confers enhanced tolerance to multiple abiotic stresses in Arabidopsis. Funct. Integr. Genomic. 2011, 11, 445–457. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Ren, W.; Zhi, D.; Wang, L.; Xia, G. Arabidopsis dreb1a/cbf3 bestowed transgenic tall fescue increased tolerance to drought stress. Plant Cell Rep. 2007, 26, 1521–1528. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Wang, L.; Nawaz, M.A.; Niu, M.; Sun, J.; Xie, J.; Kong, Q.; Huang, Y.; Cheng, F.; Bie, Z. Ectopic expression of pumpkin nac transcription factor cmnac1 improves multiple abiotic stress tolerance in arabidopsis. Front. Plant Sci. 2017, 8, 2052. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Zhao, L.; Zhao, S.; Wang, J.; Shi, H. Biochemical and transcriptomic analyses of drought stress responses of LY1306 tobacco strain. Sci. Rep. 2017, 7, 17442. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wang, Y.; Du, Y.; Zhao, Z.; Zhu, X.; Jiang, X.; Shu, Z.; Yin, Y.; Li, X. Overexpression of camellia sinensis h1 histone gene confers abiotic stress tolerance in transgenic tobacco. Plant Cell Rep. 2014, 33, 1829–1841. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, L.; Zhang, T.; Yang, X.; Li, D. Functional characterization of ks-type dehydrin zmdhn13 and its related conserved domains under oxidative stress. Sci. Rep. 2017, 7, 7361. [Google Scholar] [CrossRef] [PubMed]
- Siegel, C.S.; Stevenson, F.O.; Zimmer, E.A. Evaluation and comparison of fta card and ctab dna extraction methods for non-agricultural taxa1. Appl. Plant Sci. 2017, 5, 1600109. [Google Scholar] [CrossRef] [PubMed]
- Higo, K.; Iwamoto, M.; Higo, H. Place: A database of plant cis-acting regulatory DNA elements. Nucleic Acids Res. 1998, 26, 358–359. [Google Scholar] [CrossRef] [PubMed]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van, D.; Rouze, P.; Rombauts, S. Plantcare, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Higo, K.; Ugawa, Y.; Iwamoto, M.; Korenaga, T. Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res. 1999, 27, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Battraw, M.J.; Hall, T.C. Histochemical analysis of CaMV35S promoter-β-glucuronidase gene expression in transgenic rice plants. Plant Mol. Biol. 1990, 5, 527–538. [Google Scholar] [CrossRef]


| Order | Name | Sequence | Number | Function |
|---|---|---|---|---|
| 1 | A-box | CCGTCC | 3 | cis-acting regulatory element |
| 2 | AAGAA-motif | GAAAGAA | 2 | |
| 3 | ABRE | GGACACGTGGC | 6 | cis-acting element involved in the abscisic acid responsiveness |
| ACGTGGC | ||||
| ACACGTGGC | ||||
| CACGTG | ||||
| GCAACGTGTC | ||||
| CACGTG | ||||
| 4 | ACE | AAAACGTTTA | 4 | cis-acting element involved in light responsiveness |
| ACTACGTTGG | ||||
| 5 | ARE | TGGTTT | 1 | cis-acting regulatory element essential for the anaerobic induction |
| 6 | AT1-motif | AT1-motif | 1 | part of a light responsive module |
| 7 | ATC-motif | TGCTATCCG | 3 | part of a conserved DNA module involved in light responsiveness |
| 8 | ATCT-motif | AATCTAATCT | 2 | part of a conserved DNA module involved in light responsiveness |
| AATCTGATCG | ||||
| 9 | AuxRE | TGTCTCAATAAG | 1 | part of an auxin-responsive element |
| 10 | Box 4 | ATTAAT | 2 | part of a conserved DNA module involved in light responsiveness |
| 11 | CAAT-box | gGCAAT | 16 | common cis-acting element in promoter and enhancer regions |
| CAAT | ||||
| CAAAT | ||||
| 12 | CAT-box | GCCACT | 1 | cis-acting regulatory element related to meristem expression |
| 13 | CCGTCC-box | CCGTCC | 3 | cis-acting regulatory element related to meristem specific activation |
| 14 | CGTCA-motif | CGTCA | 1 | cis-acting regulatory element involved in the MeJA-responsiveness |
| 15 | G-Box | CACGTG | 2 | cis-acting regulatory element involved in light responsiveness |
| 16 | G-box | tgACACGTGGCA | 4 | cis-acting regulatory element involved in light responsiveness |
| CACGTG | ||||
| ACACGTGGC | ||||
| 17 | GCN4_motif | CAAGCCA | 1 | cis-regulatory element involved in endosperm expression |
| 18 | HSE | AAAAAATTTC | 2 | cis-acting element involved in heat stress responsiveness |
| AGAAAATTCG | ||||
| 19 | LTR | CCGAAA | 5 | cis-acting element involved in low-temperature responsiveness |
| 20 | MBS | CGGTCA | 1 | MYB Binding Site |
| 21 | RY-element | CATGCATG | 1 | cis-acting regulatory element involved in seed-specific regulation |
| 22 | Skn-1_motif | GTCAT | 3 | cis-acting regulatory element required for endosperm expression |
| 23 | Sp1 | CC(G/A)CCC | 1 | light responsive element |
| 24 | TATA-box | TTTTA | 35 | core promoter element around −30 of transcription start |
| TATA | ||||
| TATAAA | ||||
| ATTATA | ||||
| TAATA | ||||
| TACAAAA | ||||
| TATAAAAT | ||||
| TATAAAA | ||||
| 25 | TCT-motif | TCTTAC | 1 | part of a light responsive element |
| 26 | TGACG-motif | TGACG | 1 | cis-acting regulatory element involved in the MeJA-responsiveness |
| 27 | Unnamed_1 | GCCACGTGGC | 4 | |
| CGTGG | ||||
| 28 | Unnamed_3 | CGTGG | 3 | |
| 29 | Unnamed_4 | CTCC | 6 | |
| 30 | box S | AGCCACC | 1 |
| Primer | Sequence (5′–3′) |
|---|---|
| F1 | AGCTCATCCATGCCATGTGTAATCC |
| F2 | TAGCACCGCCTACATACCTC |
| F3 | CGAACTGAGATACCTACAGC G |
| F4 | TCTTTCCTGCGTTATCCC |
| F5 | AATACGCAAACCGCCTCT |
| R1 | TCCACACAACATACGAGCC |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Niu, G.-L.; Gou, W.; Han, X.-L.; Qin, C.; Zhang, L.-X.; Abomohra, A.E.-F.; Ashraf, M. Cloning and Functional Analysis of Phosphoethanolamine Methyltransferase Promoter from Maize (Zea mays L.). Int. J. Mol. Sci. 2018, 19, 191. https://doi.org/10.3390/ijms19010191
Niu G-L, Gou W, Han X-L, Qin C, Zhang L-X, Abomohra AE-F, Ashraf M. Cloning and Functional Analysis of Phosphoethanolamine Methyltransferase Promoter from Maize (Zea mays L.). International Journal of Molecular Sciences. 2018; 19(1):191. https://doi.org/10.3390/ijms19010191
Chicago/Turabian StyleNiu, Gai-Li, Wei Gou, Xiang-Long Han, Cheng Qin, Li-Xin Zhang, Abd El-Fatah Abomohra, and Muhammad Ashraf. 2018. "Cloning and Functional Analysis of Phosphoethanolamine Methyltransferase Promoter from Maize (Zea mays L.)" International Journal of Molecular Sciences 19, no. 1: 191. https://doi.org/10.3390/ijms19010191
APA StyleNiu, G.-L., Gou, W., Han, X.-L., Qin, C., Zhang, L.-X., Abomohra, A. E.-F., & Ashraf, M. (2018). Cloning and Functional Analysis of Phosphoethanolamine Methyltransferase Promoter from Maize (Zea mays L.). International Journal of Molecular Sciences, 19(1), 191. https://doi.org/10.3390/ijms19010191