LHCSR Expression under HSP70/RBCS2 Promoter as a Strategy to Increase Productivity in Microalgae
Abstract
:1. Introduction
2. Results
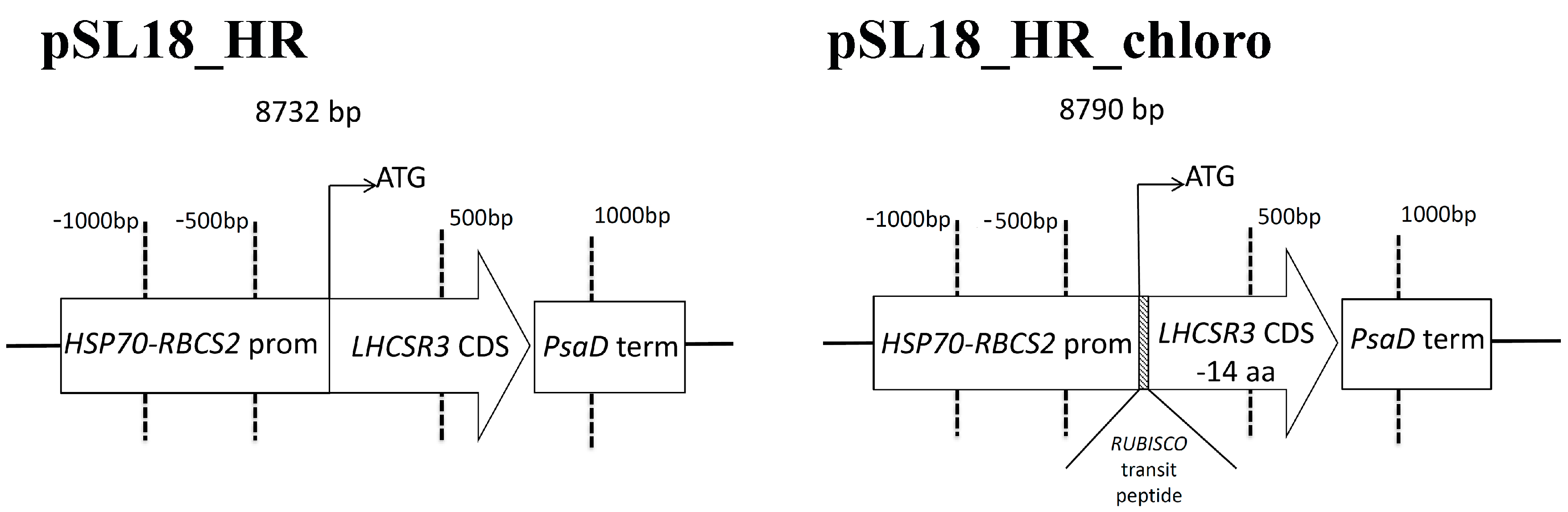
2.1. Complementation of npq4 lhcsr1 Strain
2.2. Pigment Composition of Transformed Lines
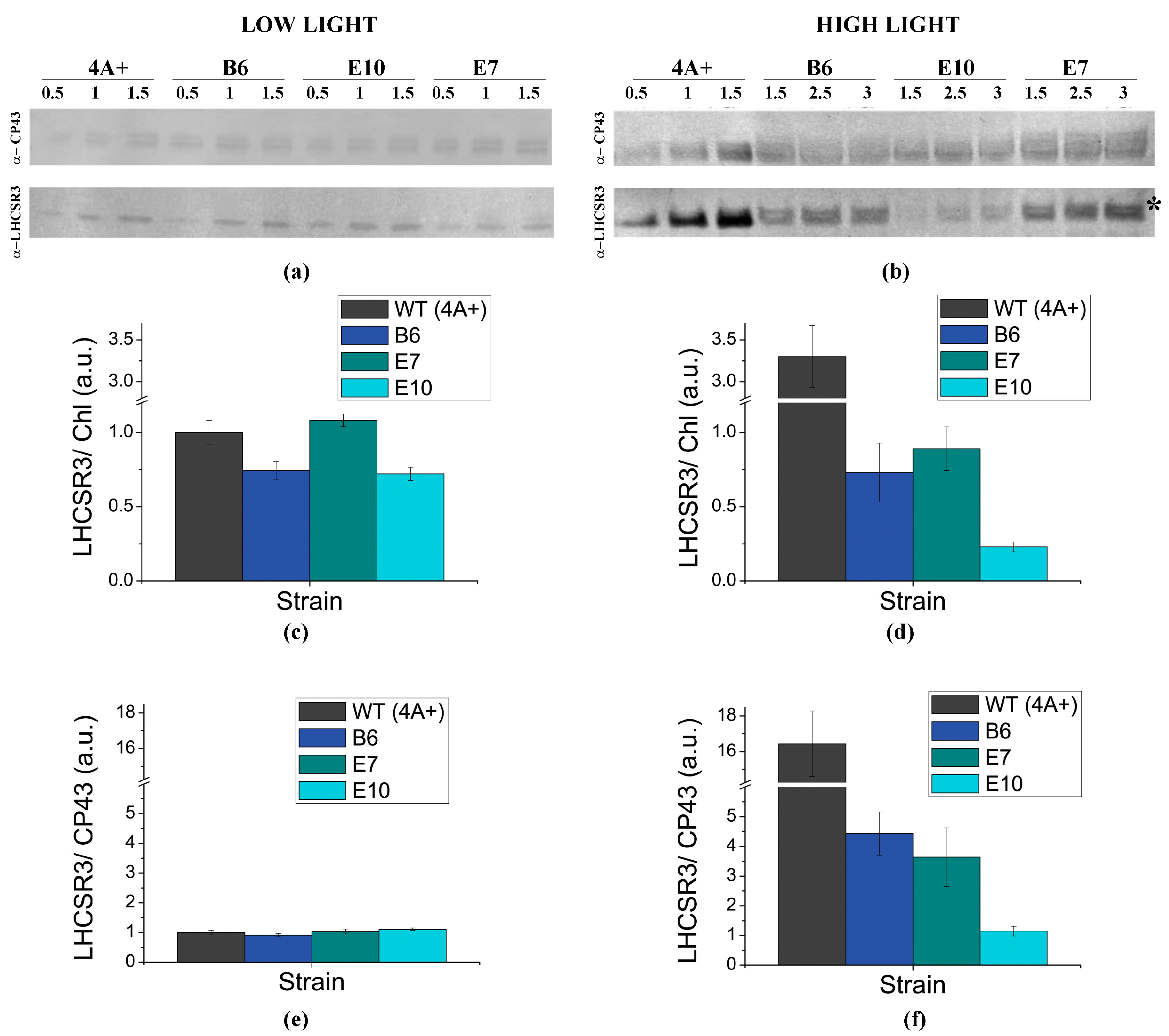
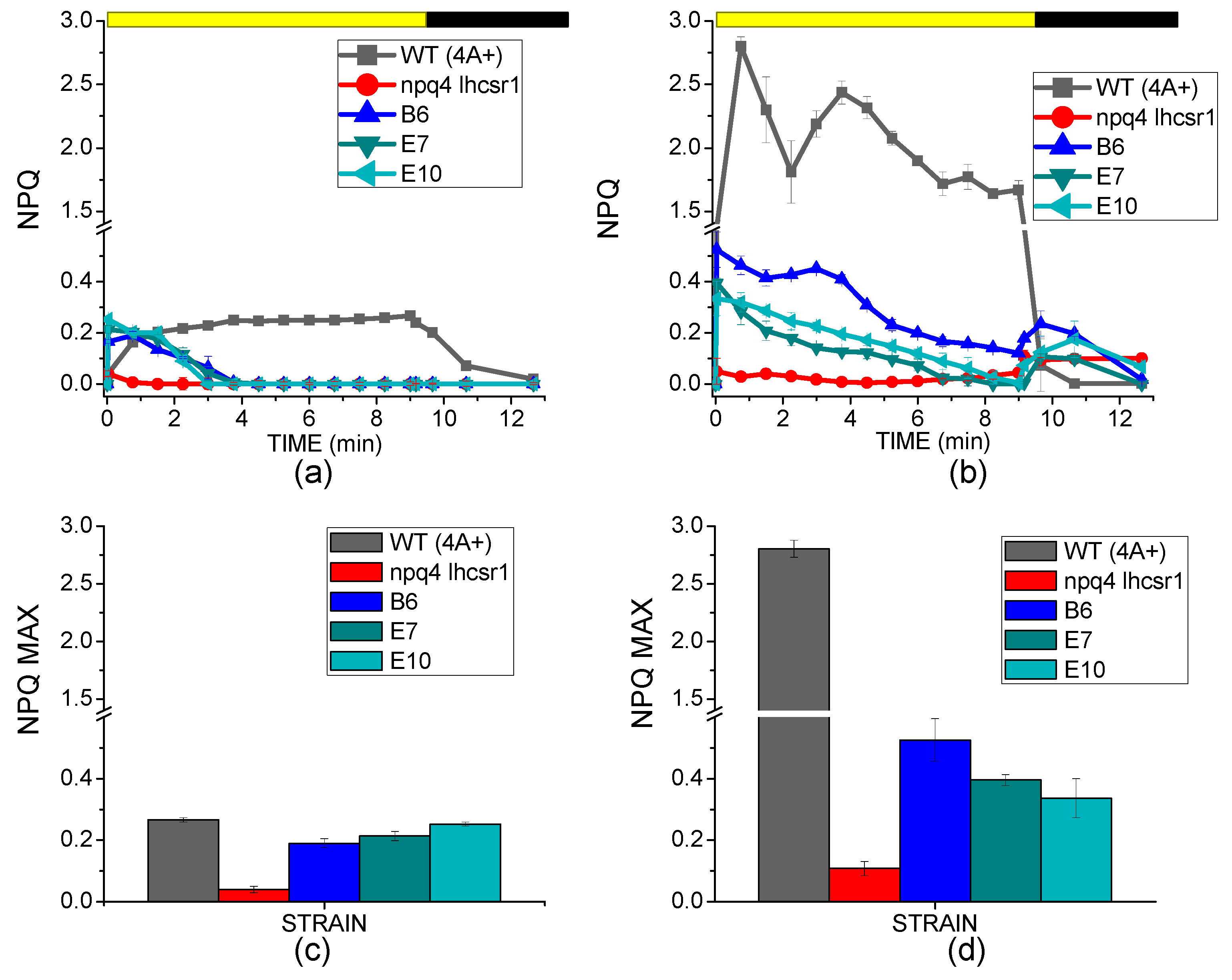
2.3. NPQ Induction and LHCSR3 Accumulation
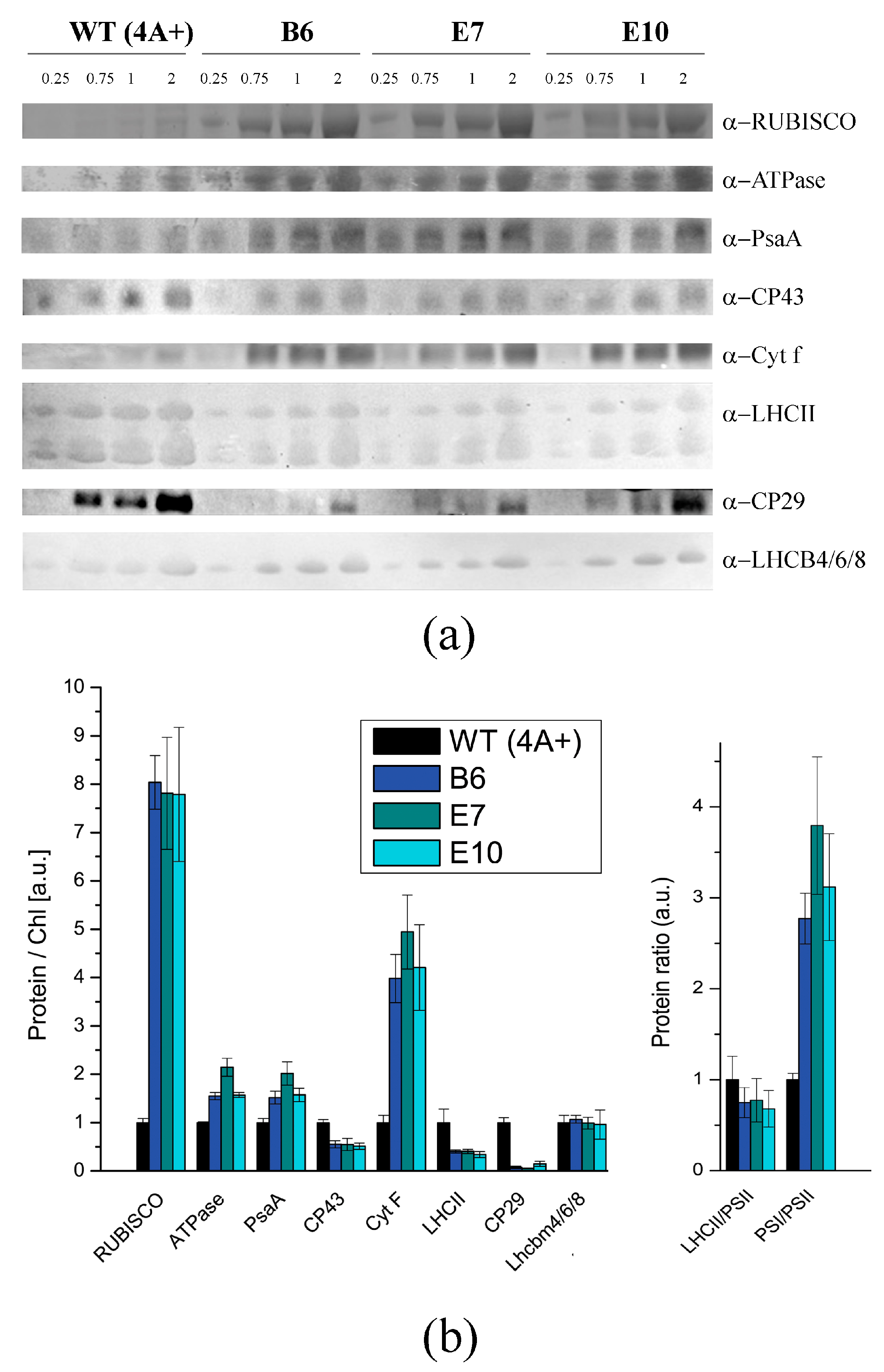
2.4. Composition of Photosynthetic Subunits
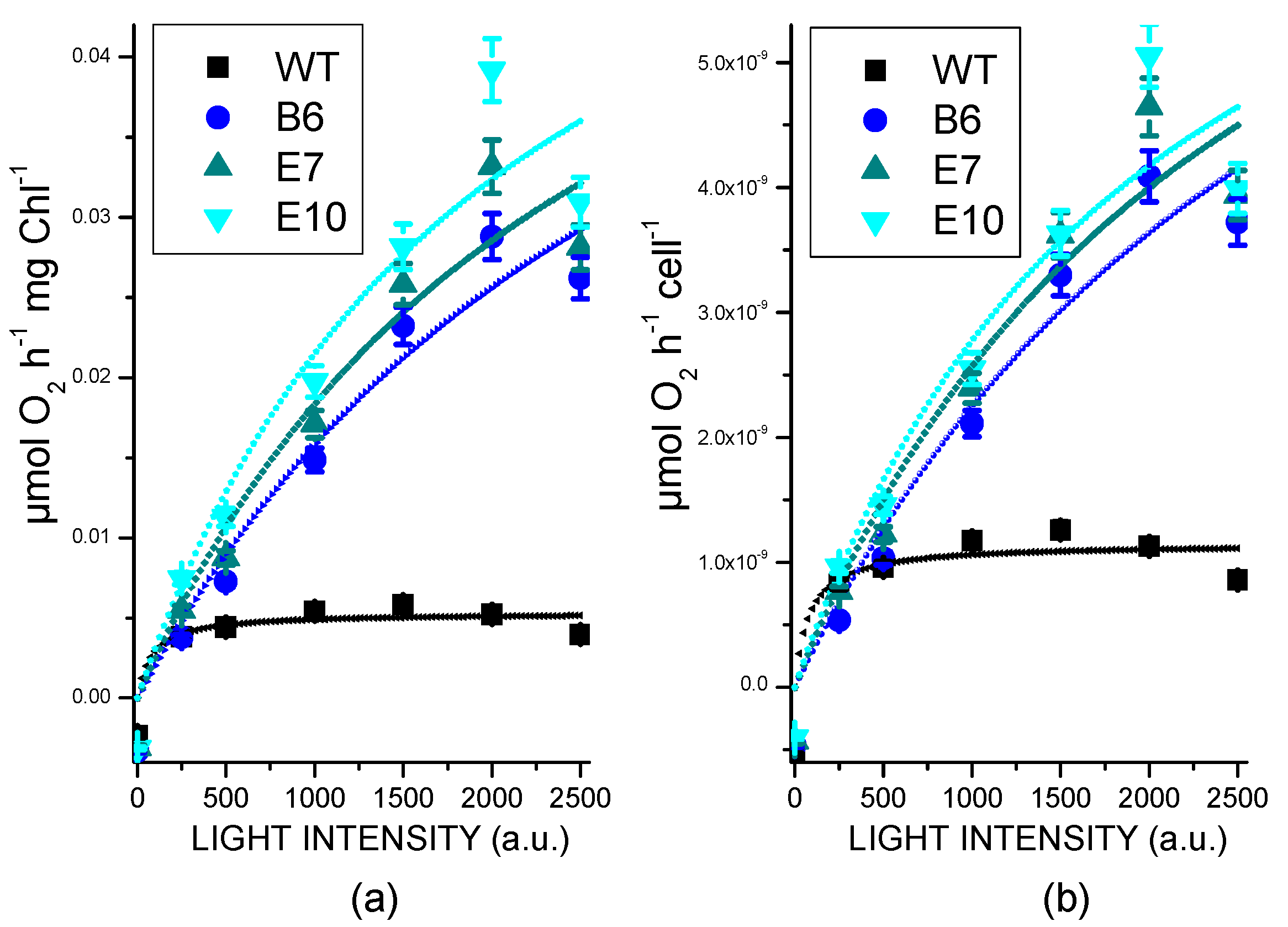
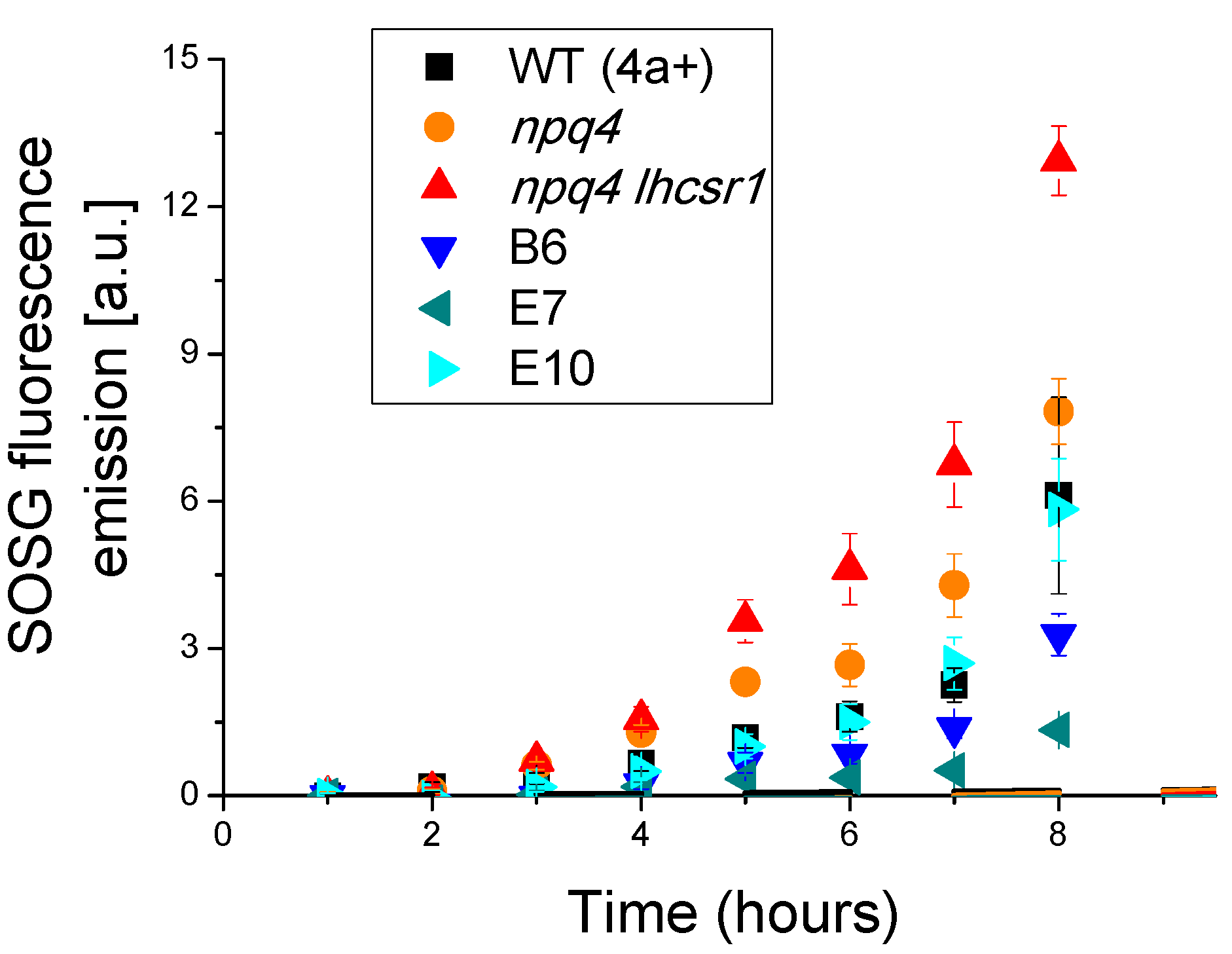
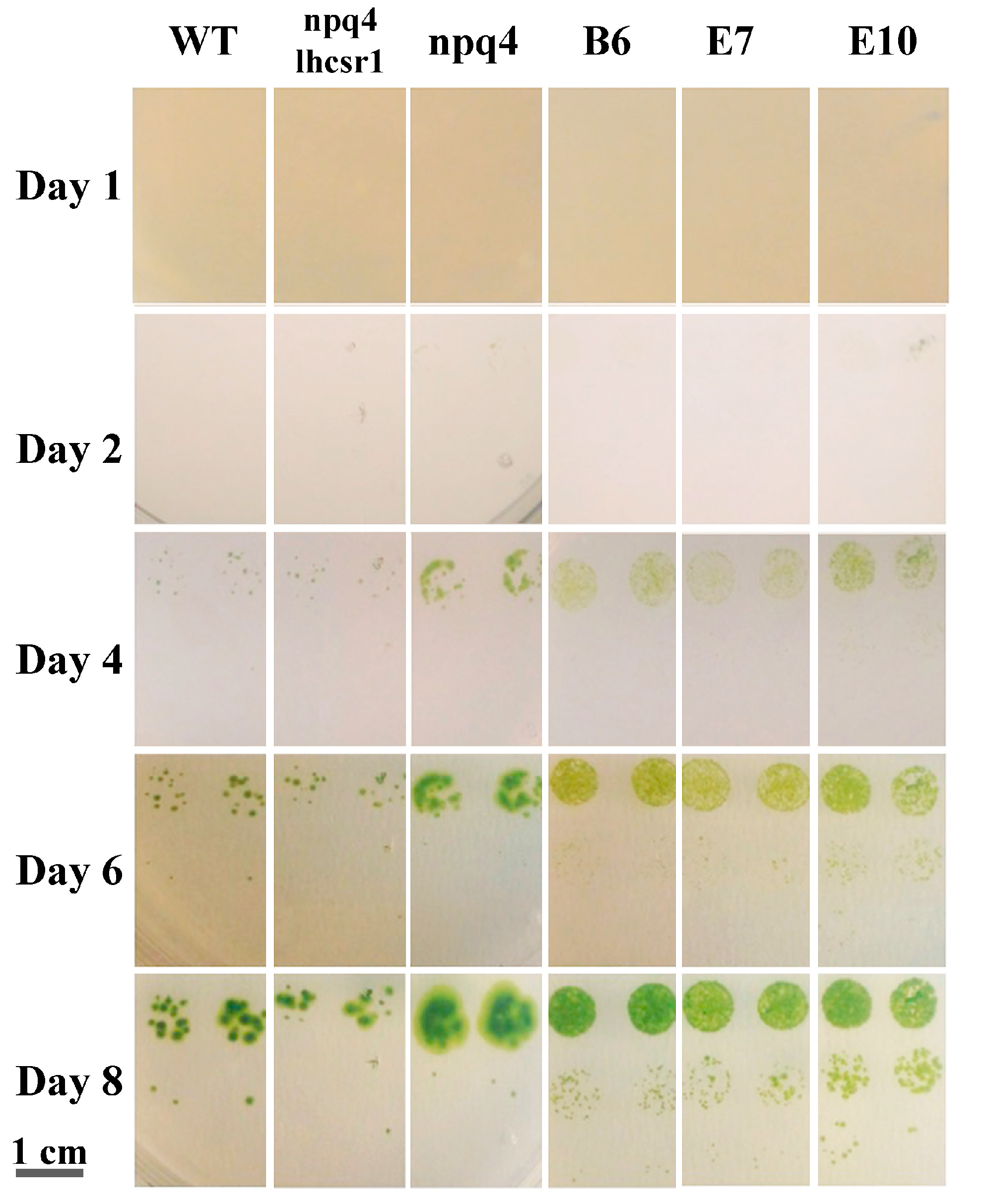
2.5. Light Dependent Oxygen Evolution vs. Photoprotection
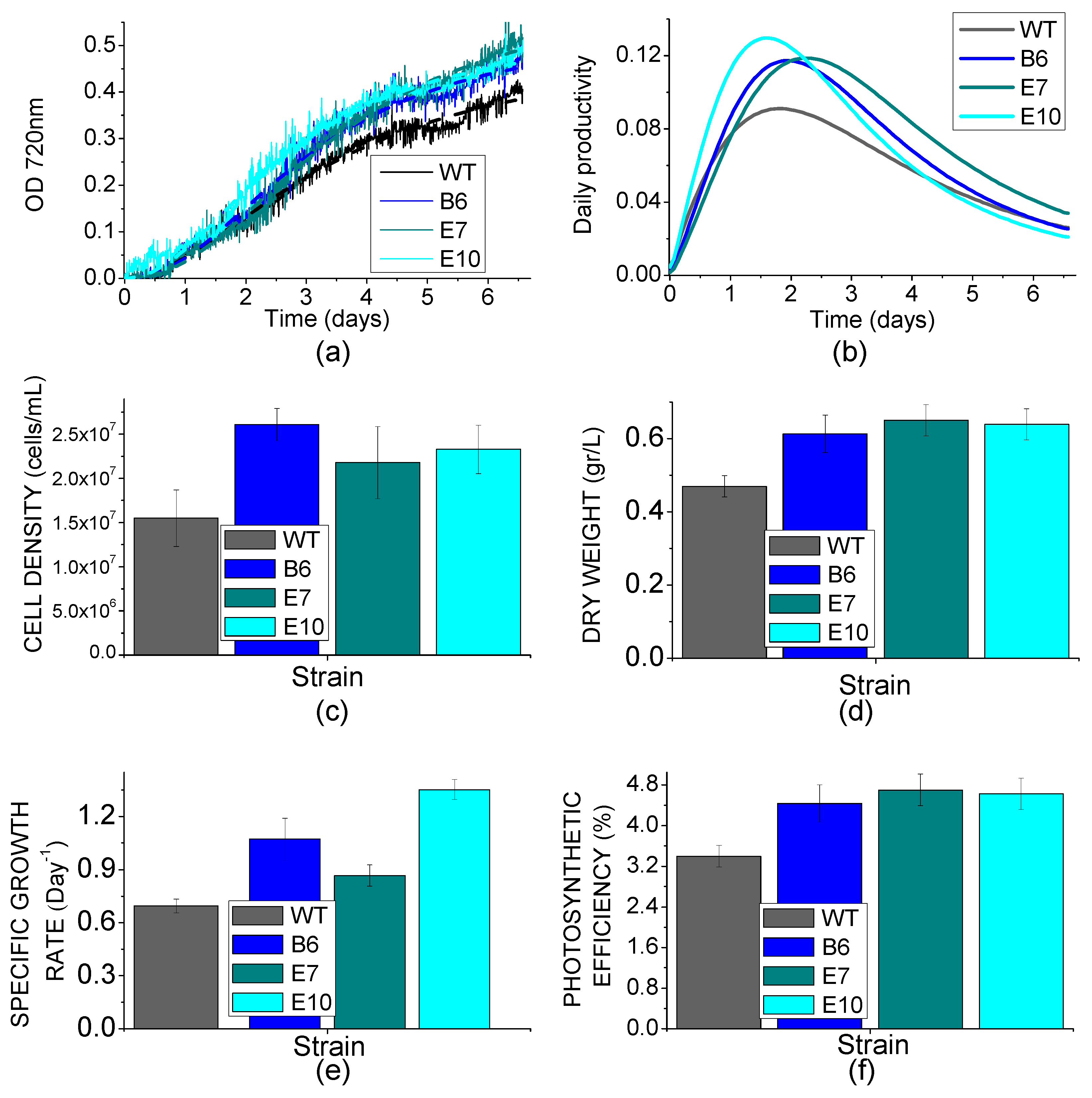
2.6. Biomass Productivity
3. Discussion
4. Materials and Methods
4.1. Chlamydomonas Reinhardtii Transformation
4.2. Chlamydomonas Reinhardtii Cultivation
4.3. Chlorophyll Fluorescence and Oxygen Evolution Measurement
4.4. Pigment Analysis
4.5. SDS-PAGE Analysis, Immunoblot Assays and Western Blotting Quantifications
4.6. Singlet Oxygen Production
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| NPQ | Non Photochemical Quenching |
| PSI/II | Photosystem I, II |
| LHC | Light Harvesting Complex |
| LHCSR | Light Harvesting Complex Stress Related |
| LHCII | Light Harvesting Complex II |
| HSP | Heat Shock Protein |
| RBCS2 | Ribulose-1,5-bisphosphate Carboxylase/Oxygenase Small Subunit |
| Chl | Chlorophyll |
| Car | Carotenoids |
References
- Ben-Shem, A.; Frolow, F.; Nelson, N. Crystal structure of plant photosystem I. Nature 2003, 426, 630–635. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Su, X.; Cao, P.; Liu, X.; Chang, W.; Li, M.; Zhang, X.; Liu, Z. Structure of spinach photosystem II-LHCII supercomplex at 3.2 Å resolution. Nature 2016, 534, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Hope, A.B. Electron transfers amongst cytochrome f, plastocyanin and photosystem I: Kinetics and mechanisms. Biochim. Biophys. Acta 2000, 1456, 5–26. [Google Scholar] [CrossRef]
- Caffarri, S.; Tibiletti, T.; Jennings, R.C.; Santabarbara, S. A comparison between plant photosystem I and photosystem II architecture and functioning. Curr. Protein Pept. Sci. 2014, 15, 296–331. [Google Scholar] [CrossRef] [PubMed]
- Tikhonov, A.N. The cytochrome b6f complex at the crossroad of photosynthetic electron transport pathways. Plant Physiol. Biochem. 2014, 81, 163–183. [Google Scholar] [CrossRef] [PubMed]
- Erickson, E.; Wakao, S.; Niyogi, K.K. Light stress and photoprotection in Chlamydomonas reinhardtii. Plant J. 2015, 82, 449–465. [Google Scholar] [CrossRef] [PubMed]
- Niyogi, K.K. PHOTOPROTECTION REVISITED: Genetic and Molecular Approaches. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1999, 50, 333–359. [Google Scholar] [CrossRef] [PubMed]
- DemmigAdams, B.; Adams, W.W.; Barker, D.H.; Logan, B.A.; Bowling, D.R.; Verhoeven, A.S. Using chlorophyll fluorescence to assess the fraction of absorbed light allocated to thermal dissipation of excess excitation. Physiol. Plant. 1996, 98, 253–264. [Google Scholar] [CrossRef]
- Belgio, E.; Kapitonova, E.; Chmeliov, J.; Duffy, C.D.; Ungerer, P.; Valkunas, L.; Ruban, A.V. Economic photoprotection in photosystem II that retains a complete light-harvesting system with slow energy traps. Nat. Commun. 2014, 5, 4433. [Google Scholar] [CrossRef] [PubMed]
- Patila, S.; Pandit, R.; Lali, A. Responses of algae to high light exposure: Prerequisite for species selection for outdoor cultivation. J. Algal Biomass Utln. 2017, 8, 75–83. [Google Scholar]
- Brestic, M.; Zivcak, M.; Kunderlikova, K.; Allakhverdiev, S. High temperature specifically affects the photoprotective responses of chlorophyll b-deficient wheat mutant lines. Photosynth. Res. 2016, 130, 251–266. [Google Scholar] [CrossRef] [PubMed]
- Kalaji, H.; Jajoo, A.; Oukarroum, A.; Brestic, M.; Zivcak, M.; Samborska, I.; Cetner, M.; Lukasik, I.; Goltsev, V.; Ladle, R. Chlorophyll a fluorescence as a tool to monitor physiological status of plants under abiotic stress conditions. Acta Physiol. Plant. 2016, 38. [Google Scholar] [CrossRef]
- Campbell, D.; Hurry, V.; Clarke, A.K.; Gustafsson, P.; Oquist, G. Chlorophyll fluorescence analysis of cyanobacterial photosynthesis and acclimation. Microbiol. Mol. Biol. Rev. 1998, 62, 667–683. [Google Scholar] [PubMed]
- Li, X.P.; Björkman, O.; Shih, C.; Grossman, A.R.; Rosenquist, M.; Jansson, S.; Niyogi, K.K. A pigment-binding protein essential for regulation of photosynthetic light harvesting. Nature 2000, 403, 391–395. [Google Scholar] [CrossRef] [PubMed]
- Peers, G.; Truong, T.B.; Ostendorf, E.; Busch, A.; Elrad, D.; Grossman, A.R.; Hippler, M.; Niyogi, K.K. An ancient light-harvesting protein is critical for the regulation of algal photosynthesis. Nature 2009, 462, 518–521. [Google Scholar] [CrossRef] [PubMed]
- Demmig-Adams, B.; Cohu, C.M.; Muller, O.; Adams, W.W. Modulation of photosynthetic energy conversion efficiency in nature: From seconds to seasons. Photosynth. Res. 2012, 113, 75–88. [Google Scholar] [CrossRef] [PubMed]
- Demmig-Adams, B.; Adams, W.W. Photoprotection in an ecological context: The remarkable complexity of thermal energy dissipation. New Phytol. 2006, 172, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Lepetit, B.; Sturm, S.; Rogato, A.; Gruber, A.; Sachse, M.; Falciatore, A.; Kroth, P.; Lavaud, J. High light acclimation in the secondary plastids containing diatom Phaeodactylum tricornutum is triggered by the redox state of the plastoquinone pool. Plant Physiol. 2012. [Google Scholar] [CrossRef] [PubMed]
- Wilson, A.; Ajlani, G.; Verbavatz, J.M.; Vass, I.; Kerfeld, C.A.; Kirilovsky, D. A soluble carotenoid protein involved in phycobilisome-related energy dissipation in cyanobacteria. Plant Cell 2006, 18, 992–1007. [Google Scholar] [CrossRef] [PubMed]
- Berteotti, S.; Ballottari, M.; Bassi, R. Increased biomass productivity in green algae by tuning non-photochemical quenching. Sci. Rep. 2016, 6, 21339. [Google Scholar] [CrossRef] [PubMed]
- Kirst, H.; Gabilly, S.T.; Niyogi, K.K.; Lemaux, P.G.; Melis, A. Photosynthetic antenna engineering to improve crop yields. Planta 2017, 245, 1009–1020. [Google Scholar] [CrossRef] [PubMed]
- Chisti, Y. Biodiesel from microalgae. Biotechnol. Adv. 2007, 25, 294–306. [Google Scholar] [CrossRef] [PubMed]
- Akbari, F.; Eskandani, M.; Khosroushahi, A.Y. The potential of transgenic green microalgae; a robust photobioreactor to produce recombinant therapeutic proteins. World J. Microbiol. Biotechnol. 2014, 30, 2783–2796. [Google Scholar] [CrossRef] [PubMed]
- Lum, K.K.; Kim, J.; Lei, X.G. Dual potential of microalgae as a sustainable biofuel feedstock and animal feed. J. Anim. Sci. Biotechnol. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Chisti, Y. Biodiesel from microalgae beats bioethanol. Trends Biotechnol. 2008, 26, 126–131. [Google Scholar] [CrossRef] [PubMed]
- Roth, M.S.; Cokus, S.J.; Gallaher, S.D.; Walter, A.; Lopez, D.; Erickson, E.; Endelman, B.; Westcott, D.; Larabell, C.A.; Merchant, S.S.; et al. Chromosome-level genome assembly and transcriptome of the green alga Chromochloris zofingiensis illuminates astaxanthin production. Proc. Natl. Acad. Sci. USA 2017, 114, E4296–E4305. [Google Scholar] [CrossRef] [PubMed]
- Stephens, E.; Ross, I.; Mussgnug, J.; Wagner, L.; Borowitzka, M.; Posten, C.; Kruse, O.; Hankamer, B. Future prospects of microalgal biofuel production systems. Trends Plant Sci. 2010, 15, 554–564. [Google Scholar] [CrossRef] [PubMed]
- Ballottari, M.; Truong, T.B.; De Re, E.; Erickson, E.; Stella, G.R.; Fleming, G.R.; Bassi, R.; Niyogi, K.K. Identification of pH-sensing Sites in the Light Harvesting Complex Stress-related 3 Protein Essential for Triggering Non-photochemical Quenching in Chlamydomonas reinhardtii. J. Biol. Chem. 2016, 291, 7334–7346. [Google Scholar] [CrossRef] [PubMed]
- Liguori, N.; Roy, L.M.; Opacic, M.; Durand, G.; Croce, R. Regulation of light harvesting in the green alga Chlamydomonas reinhardtii: The C-terminus of LHCSR is the knob of a dimmer switch. J. Am. Chem. Soc. 2013, 135, 18339–18342. [Google Scholar] [CrossRef] [PubMed]
- Pinnola, A.; Cazzaniga, S.; Alboresi, A.; Nevo, R.; Levin-Zaidman, S.; Reich, Z.; Bassi, R. Light-Harvesting Complex Stress-Related Proteins Catalyze Excess Energy Dissipation in Both Photosystems of Physcomitrella patens. Plant Cell 2015, 27, 3213–3227. [Google Scholar] [CrossRef] [PubMed]
- Dinc, E.; Tian, L.; Roy, L.M.; Roth, R.; Goodenough, U.; Croce, R. LHCSR1 induces a fast and reversible pH-dependent fluorescence quenching in LHCII in Chlamydomonas reinhardtii cells. Proc. Natl. Acad. Sci. USA 2016, 113, 7673–7678. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, S.; Tokutsu, R.; Minagawa, J. Transcriptional regulation of the stress-responsive light harvesting complex genes in Chlamydomonas reinhardtii. Plant Cell Physiol. 2014, 55, 1304–1310. [Google Scholar] [CrossRef] [PubMed]
- Petroutsos, D.; Busch, A.; Janssen, I.; Trompelt, K.; Bergner, S.V.; Weinl, S.; Holtkamp, M.; Karst, U.; Kudla, J.; Hippler, M. The chloroplast calcium sensor CAS is required for photoacclimation in Chlamydomonas reinhardtii. Plant Cell 2011, 23, 2950–2963. [Google Scholar] [CrossRef] [PubMed]
- Cantrell, M.; Peers, G. A mutant of Chlamydomonas without LHCSR maintains high rates of photosynthesis, but has reduced cell division rates in sinusoidal light conditions. PLoS ONE 2017, 12, e0179395. [Google Scholar] [CrossRef] [PubMed]
- Schroda, M.; Blöcker, D.; Beck, C.F. The HSP70A promoter as a tool for the improved expression of transgenes in Chlamydomonas. Plant J. 2000, 21, 121–131. [Google Scholar] [CrossRef] [PubMed]
- Bonente, G.; Ballottari, M.; Truong, T.B.; Morosinotto, T.; Ahn, T.K.; Fleming, G.R.; Niyogi, K.K.; Bassi, R. Analysis of LhcSR3, a Protein Essential for Feedback De-Excitation in the Green Alga Chlamydomonas reinhardtii. PLoS Biol. 2011, 9. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Lee, D.S.; Choi, I.S.; Ahn, S.J.; Kim, Y.H.; Bae, H.J. Arabidopsis thaliana Rubisco small subunit transit peptide increases the accumulation of Thermotoga maritima endoglucanase Cel5A in chloroplasts of transgenic tobacco plants. Transgenic Res. 2010, 19, 489–497. [Google Scholar] [CrossRef] [PubMed]
- Bonente, G.; Pippa, S.; Castellano, S.; Bassi, R.; Ballottari, M. Acclimation of Chlamydomonas reinhardtii to Different Growth Irradiances. J. Biol. Chem. 2012, 287, 5833–5847. [Google Scholar] [CrossRef] [PubMed]
- Petroutsos, D.; Tokutsu, R.; Maruyama, S.; Flori, S.; Greiner, A.; Magneschi, L.; Cusant, L.; Kottke, T.; Mittag, M.; Hegemann, P.; et al. A blue-light photoreceptor mediates the feedback regulation of photosynthesis. Nature 2016, 537, 563–566. [Google Scholar] [CrossRef] [PubMed]
- Bergner, S.V.; Scholz, M.; Trompelt, K.; Barth, J.; Gäbelein, P.; Steinbeck, J.; Xue, H.; Clowez, S.; Fucile, G.; Goldschmidt-Clermont, M.; et al. STATE TRANSITION7-Dependent Phosphorylation Is Modulated by Changing Environmental Conditions, and Its Absence Triggers Remodeling of Photosynthetic Protein Complexes. Plant Physiol. 2015, 168, 615–634. [Google Scholar] [CrossRef] [PubMed]
- Allorent, G.; Tokutsu, R.; Roach, T.; Peers, G.; Cardol, P.; Girard-Bascou, J.; Seigneurin-Berny, D.; Petroutsos, D.; Kuntz, M.; Breyton, C.; et al. A dual strategy to cope with high light in Chlamydomonas reinhardtii. Plant Cell 2013, 25, 545–557. [Google Scholar] [CrossRef] [PubMed]
- Girolomoni, L.; Ferrante, P.; Berteotti, S.; Giuliano, G.; Bassi, R.; Ballottari, M. The function of LHCBM4/6/8 antenna proteins in Chlamydomonas reinhardtii. J. Exp. Bot. 2017, 68, 627–641. [Google Scholar] [PubMed]
- Flors, C.; Fryer, M.J.; Waring, J.; Reeder, B.; Bechtold, U.; Mullineaux, P.M.; Nonell, S.; Wilson, M.T.; Baker, N.R. Imaging the production of singlet oxygen in vivo using a new fluorescent sensor, Singlet Oxygen Sensor Green. J. Exp. Bot. 2006, 57, 1725–1734. [Google Scholar] [CrossRef] [PubMed]
- Betterle, N.; Ballottari, M.; Hienerwadel, R.; Dall’Osto, L.; Bassi, R. Dynamics of zeaxanthin binding to the photosystem II monomeric antenna protein Lhcb6 (CP24) and modulation of its photoprotection properties. Arch. Biochem. Biophys. 2010, 504, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Giordano, M.; Norici, A.; Forssen, M.; Eriksson, M.; Raven, J. An anaplerotic role for mitochondrial carbonic anhydrase in Chlamydomonas reinhardtii. Plant Physiol. 2003, 132, 2126–2134. [Google Scholar] [CrossRef] [PubMed]
- Kliphuis, A.; Klok, A.; Martens, D.; Lamers, P.; Janssen, M.; Wijffels, R. Metabolic modeling of Chlamydomonas reinhardtii: Energy requirements for photoautotrophic growth and maintenance. J. Appl. Phycol. 2012, 24, 253–266. [Google Scholar] [CrossRef] [PubMed]
- Langner, U.; Jakob, T.; Stehfest, K.; Wilhelm, C. An energy balance from absorbed photons to new biomass for Chlamydomonas reinhardtii and Chlamydomonas acidophila under neutral and extremely acidic growth conditions. Plant Cell Environ. 2009, 32, 250–258. [Google Scholar] [CrossRef] [PubMed]
- Vitova, M.; Bisova, K.; Hlavova, M.; Kawano, S.; Zachleder, V.; Cizkova, M. Chlamydomonas reinhardtii: Duration of its cell cycle and phases at growth rates affected by temperature. Planta 2011, 234, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Carbonera, D.; Gerotto, C.; Posocco, B.; Giacometti, G.M.; Morosinotto, T. NPQ activation reduces chlorophyll triplet state formation in the moss Physcomitrella patens. Biochim. Biophys. Acta 2012, 1817, 1608–1615. [Google Scholar] [CrossRef] [PubMed]
- Kromdijk, J.; Głowacka, K.; Leonelli, L.; Gabilly, S.T.; Iwai, M.; Niyogi, K.K.; Long, S.P. Improving photosynthesis and crop productivity by accelerating recovery from photoprotection. Science 2016, 354, 857–861. [Google Scholar] [CrossRef] [PubMed]
- Chaux, F.; Johnson, X.; Auroy, P.; Beyly-Adriano, A.; Te, I.; Cuiné, S.; Peltier, G. PGRL1 and LHCSR3 Compensate for Each Other in Controlling Photosynthesis and Avoiding Photosystem I Photoinhibition during High Light Acclimation of Chlamydomonas Cells. Mol. Plant 2017, 10, 216–218. [Google Scholar] [CrossRef] [PubMed]
- Xue, H.; Tokutsu, R.; Bergner, S.V.; Scholz, M.; Minagawa, J.; Hippler, M. PHOTOSYSTEM II SUBUNIT R is required for efficient binding of LIGHT-HARVESTING COMPLEX STRESS-RELATED PROTEIN3 to photosystem II-light-harvesting supercomplexes in Chlamydomonas reinhardtii. Plant Physiol. 2015, 167, 1566–1578. [Google Scholar] [CrossRef] [PubMed]
- Semchonok, D.A.; Sathish Yadav, K.N.; Xu, P.; Drop, B.; Croce, R.; Boekema, E.J. Interaction between the photoprotective protein LHCSR3 and C2S2 Photosystem II supercomplex in Chlamydomonas reinhardtii. Biochim. Biophys. Acta 2017, 1858, 379–385. [Google Scholar] [CrossRef] [PubMed]
- Pinnola, A.; Staleva-Musto, H.; Capaldi, S.; Ballottari, M.; Bassi, R.; Polívka, T. Electron transfer between carotenoid and chlorophyll contributes to quenching in the LHCSR1 protein from Physcomitrella patens. Biochim. Biophys. Acta 2016, 1857, 1870–1878. [Google Scholar] [CrossRef] [PubMed]
- Bonente, G.; Formighieri, C.; Mantelli, M.; Catalanotti, C.; Giuliano, G.; Morosinotto, T.; Bassi, R. Mutagenesis and phenotypic selection as a strategy toward domestication of Chlamydomonas reinhardtii strains for improved performance in photobioreactors. Photosynth. Res. 2011, 108. [Google Scholar] [CrossRef] [PubMed]
- León, R.; Couso, I.; Fernández, E. Metabolic engineering of ketocarotenoids biosynthesis in the unicelullar microalga Chlamydomonas reinhardtii. J. Biotechnol. 2007, 130, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Kindle, K.L. High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc. Natl. Acad. Sci. USA 1990, 87, 1228–1232. [Google Scholar] [CrossRef] [PubMed]
- Ferrante, P.; Ballottari, M.; Bonente, G.; Giuliano, G.; Bassi, R. LHCBM1 and LHCBM2/7 polypeptides, components of major LHCII complex, have distinct functional roles in photosynthetic antenna system of Chlamydomonas reinhardtii. J. Biol. Chem. 2012, 287, 16276–16288. [Google Scholar] [CrossRef] [PubMed]
- Schägger, H.; von Jagow, G. Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal. Biochem. 1987, 166, 368–379. [Google Scholar] [CrossRef]
| Chl/Cell | s.d. | Chl a/b | s.d. | Chl/Car | s.d. | ||
|---|---|---|---|---|---|---|---|
| low light | WT | 2.16 × 10−6 | 1.99 × 10−7 | 1.86 | 0.01 | 3.93 | 0.02 |
| B6 | 1.42 × 10−6 | 1.26 × 10−7 | 3.22 | 0.01 | 2.46 | 0.01 | |
| E7 | 1.40 × 10−6 | 2.69 × 10−7 | 3.24 | 0.03 | 2.50 | 0.02 | |
| E10 | 1.29 × 10−6 | 7.70 × 10−8 | 3.24 | 0.10 | 2.49 | 0.06 | |
| high light | WT | 7.29 × 10−7 | 1.29 × 10−8 | 1.49 | 0.04 | 2.48 | 0.20 |
| B6 | 3.87 × 10−7 | 7.29 × 10−9 | 2.43 | 0.07 | 1.39 | 0.06 | |
| E7 | 4.38 × 10−7 | 1.09 × 10−8 | 2.53 | 0.03 | 1.15 | 0.03 | |
| E10 | 4.17 × 10−7 | 9.63 × 10−9 | 2.59 | 0.05 | 1.07 | 0.03 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Perozeni, F.; Stella, G.R.; Ballottari, M. LHCSR Expression under HSP70/RBCS2 Promoter as a Strategy to Increase Productivity in Microalgae. Int. J. Mol. Sci. 2018, 19, 155. https://doi.org/10.3390/ijms19010155
Perozeni F, Stella GR, Ballottari M. LHCSR Expression under HSP70/RBCS2 Promoter as a Strategy to Increase Productivity in Microalgae. International Journal of Molecular Sciences. 2018; 19(1):155. https://doi.org/10.3390/ijms19010155
Chicago/Turabian StylePerozeni, Federico, Giulio Rocco Stella, and Matteo Ballottari. 2018. "LHCSR Expression under HSP70/RBCS2 Promoter as a Strategy to Increase Productivity in Microalgae" International Journal of Molecular Sciences 19, no. 1: 155. https://doi.org/10.3390/ijms19010155
APA StylePerozeni, F., Stella, G. R., & Ballottari, M. (2018). LHCSR Expression under HSP70/RBCS2 Promoter as a Strategy to Increase Productivity in Microalgae. International Journal of Molecular Sciences, 19(1), 155. https://doi.org/10.3390/ijms19010155







