ФC31 Integrase-Mediated Isolation and Characterization of Novel Safe Harbors for Transgene Expression in the Pig Genome
Abstract
:1. Introduction
2. Results
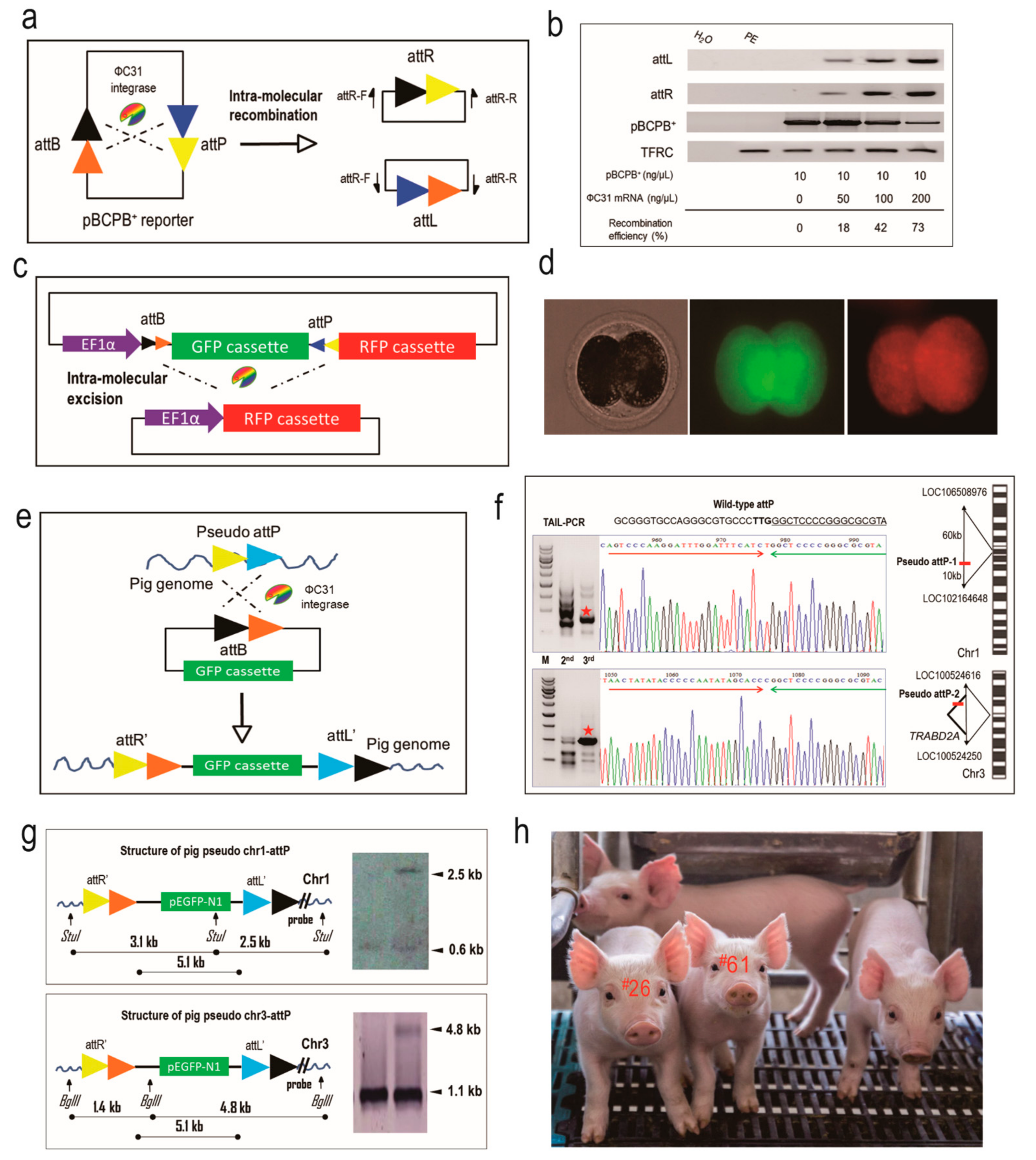
2.1. The ФC31 Integrase is Functional in Catalyzing Site-Specific Recombination in Porcine Embryos
2.2. The Integration of a Reporter Donor into Pseudo attP Sites Using ФC31 Integrase by Zygote Microinjection
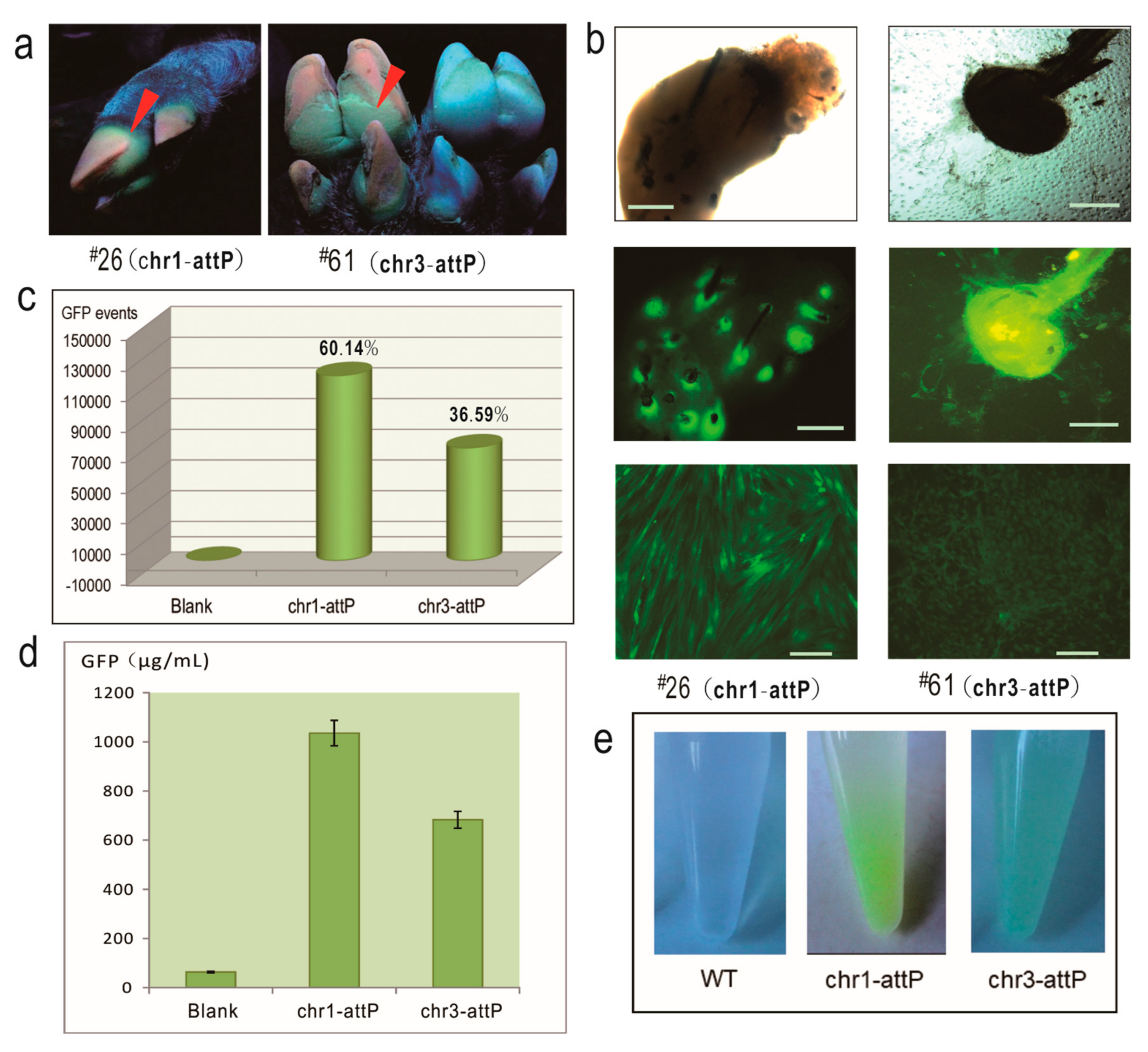
2.3. The Two Pseudo attP Sites Are Safe Harbors for Transgene Expression
3. Discussion
4. Materials and Methods
4.1. Animal Ethics
4.2. Plasmids and Strains
4.3. Nucleic Acid Manipulation
4.4. Pig Embryo Engineering and Microinjection
4.5. End Point Real-Time and TAIL-PCR
4.6. Cell Culture
4.7. FACS and ELISA
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Tiyaboonchai, A.; Mac, H.; Shamsedeen, R.; Mills, J.A.; Kishore, S.; French, D.L.; Gadue, P. Utilization of the AAVS1 safe harbor locus for hematopoietic specific transgene expression and gene knockdown in human ES cells. Stem Cell Res. 2014, 12, 630–637. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yang, Y.; Bu, L.; Guo, X.; Tang, C.; Song, J.; Fan, N.; Zhao, B.; Ouyang, Z.; Liu, Z.; et al. Rosa26-targeted swine models for stable gene over-expression and Cre-mediated lineage tracing. Cell Res. 2014, 24, 501–504. [Google Scholar] [CrossRef] [PubMed]
- Ruan, J.; Li, H.; Xu, K.; Wu, T.; Wei, J.; Zhou, R.; Liu, Z.; Mu, Y.; Yang, S.; Ouyang, H.; et al. Highly efficient CRISPR/Cas9-mediated transgene knockin at the H11 locus in pigs. Sci. Rep. 2015, 5, 14253. [Google Scholar] [CrossRef] [PubMed]
- Park, K.-E.; Park, C.-H.; Powell, A.; Martin, J.; Donovan, D.M.; Telugu, B.P. Targeted Gene Knockin in Porcine Somatic Cells Using CRISPR/Cas Ribonucleoproteins. Int. J. Mol. Sci. 2016, 17, 810. [Google Scholar] [CrossRef] [PubMed]
- Park, K.-E.; Powell, A.; Sandmaier, S.E.S.; Kim, C.-M.; Mileham, A.; Donovan, D.M.; Telugu, B.P. Targeted gene knock-in by CRISPR/Cas ribonucleoproteins in porcine zygotes. Sci. Rep. 2017, 7, 42458. [Google Scholar] [CrossRef] [PubMed]
- Ahmadi, M.; Mahboudi, F.; Akbari Eidgahi, M.R.; Nasr, R.; Nematpour, F.; Ahmadi, S.; Ebadat, S.; Aghaeepoor, M.; Davami, F. Evaluating the efficiency of phiC31 integrase-mediated monoclonal antibody expression in CHO cells. Biotechnol. Prog. 2016, 32, 1570–1576. [Google Scholar] [CrossRef] [PubMed]
- Groth, A.C.; Olivares, E.C.; Thyagarajan, B.; Calos, M.P. A phage integrase directs efficient site-specific integration in human cells. Proc. Natl. Acad. Sci. USA 2000, 97, 5995–6000. [Google Scholar] [CrossRef] [PubMed]
- Olivares, E.C.; Hollis, R.P.; Chalberg, T.W.; Meuse, L.; Kay, M.A.; Calos, M.P. Site-specific genomic integration produces therapeutic Factor IX levels in mice. Nat. Biotechnol. 2002, 20, 1124–1128. [Google Scholar] [CrossRef] [PubMed]
- Chalberg, T.W.; Genise, H.L.; Vollrath, D.; Calos, M.P. φC31 Integrase Confers Genomic Integration and Long-Term Transgene Expression in Rat Retina. Investig. Opthalmol. Vis. Sci. 2005, 46, 2140. [Google Scholar] [CrossRef] [PubMed]
- Yonemura, N.; Tamura, T.; Uchino, K.; Kobayashi, I.; Tatematsu, K.; Iizuka, T.; Tsubota, T.; Sezutsu, H.; Muthulakshmi, M.; Nagaraju, J.; et al. phiC31-integrase-mediated, site-specific integration of transgenes in the silkworm, Bombyx mori (Lepidoptera: Bombycidae). Appl. Entomol. Zool. 2013, 48, 265–273. [Google Scholar] [CrossRef]
- Damavandi, N.; Raigani, M.; Joudaki, A.; Davami, F.; Zeinali, S. Rapid characterization of the CHO platform cell line and identification of pseudo attP sites for PhiC31 integrase. Protein Expr. Purif. 2017, 140, 60–64. [Google Scholar] [CrossRef] [PubMed]
- Bi, Y.; Liu, X.; Zhang, L.; Shao, C.; Ma, Z.; Hua, Z.; Zhang, L.; Li, L.; Hua, W.; Xiao, H.; et al. Pseudo attP sites in favor of transgene integration and expression in cultured porcine cells identified by streptomyces phage phiC31 integrase. BMC Mol. Biol. 2013, 14, 20. [Google Scholar] [CrossRef] [PubMed]
- Lister, J.A. Transgene excision in zebrafish using the phiC31 integrase. Genesis 2010, 48, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Wang, Y.; Tong, Q.; Liu, X.; Su, F.; Quan, F.; Guo, Z.; Zhang, Y. A Site-Specific Recombinase-Based Method to Produce Antibiotic Selectable Marker Free Transgenic Cattle. PLoS ONE 2013, 8, e62457. [Google Scholar] [CrossRef] [PubMed]
- Ou, H.-L.; Huang, Y.; Qu, L.-J.; Xu, M.; Yan, J.-B.; Ren, Z.-R.; Huang, S.-Z.; Zeng, Y.-T. A ϕC31 integrase-mediated integration hotspot in favor of transgene expression exists in the bovine genome. FEBS J. 2009, 276, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Southern, E.M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J. Mol. Biol. 1975, 98, 503–517. [Google Scholar] [CrossRef]
- Abeydeera, L.R.; Wang, W.H.; Cantley, T.C.; Rieke, A.; Murphy, C.N.; Prather, R.S.; Day, B.N. Development and viability of pig oocytes matured in a protein-free medium containing epidermal growth factor. Theriogenology 2000, 54, 787–797. [Google Scholar] [CrossRef]
- Liu, Y.-G.; Mitsukawa, N.; Oosumi, T.; Whittier, R.F. Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J. 1995, 8, 457–463. [Google Scholar] [CrossRef] [PubMed]
| Surrogate | Route | mRNA/DNA Dosage (ng/μL) | Injected/Transferred (%) | Liveborn Founders (%) * | Integrated (%) ** | Pseudo attP Site |
|---|---|---|---|---|---|---|
| 886 | PNI | 200/10 | 32/28 (88) | 9 (32) | 2 (22) | 0 |
| 1023 | PNI | 200/10 | 33/25 (76) | 8 (32) | 2 (25) | 1 |
| 654 | PNI | 200/10 | 28/20 (71) | 8 (40) | 3 (38) | 1 |
| 982 | ICI | 200/10 | 29/24 (83) | 7 (29) | 0 (0) | N/A |
| 10,665 | ICI | 200/10 | 30/23 (77) | return | N/A | N/A |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bi, Y.; Hua, Z.; Ren, H.; Zhang, L.; Xiao, H.; Liu, X.; Hua, W.; Mei, S.; Molenaar, A.; Laible, G.; et al. ФC31 Integrase-Mediated Isolation and Characterization of Novel Safe Harbors for Transgene Expression in the Pig Genome. Int. J. Mol. Sci. 2018, 19, 149. https://doi.org/10.3390/ijms19010149
Bi Y, Hua Z, Ren H, Zhang L, Xiao H, Liu X, Hua W, Mei S, Molenaar A, Laible G, et al. ФC31 Integrase-Mediated Isolation and Characterization of Novel Safe Harbors for Transgene Expression in the Pig Genome. International Journal of Molecular Sciences. 2018; 19(1):149. https://doi.org/10.3390/ijms19010149
Chicago/Turabian StyleBi, Yanzhen, Zaidong Hua, Hongyan Ren, Liping Zhang, Hongwei Xiao, Ximei Liu, Wenjun Hua, Shuqi Mei, Adrian Molenaar, Götz Laible, and et al. 2018. "ФC31 Integrase-Mediated Isolation and Characterization of Novel Safe Harbors for Transgene Expression in the Pig Genome" International Journal of Molecular Sciences 19, no. 1: 149. https://doi.org/10.3390/ijms19010149
APA StyleBi, Y., Hua, Z., Ren, H., Zhang, L., Xiao, H., Liu, X., Hua, W., Mei, S., Molenaar, A., Laible, G., & Zheng, X. (2018). ФC31 Integrase-Mediated Isolation and Characterization of Novel Safe Harbors for Transgene Expression in the Pig Genome. International Journal of Molecular Sciences, 19(1), 149. https://doi.org/10.3390/ijms19010149






