Identification of Circulating miRNAs Differentially Regulated by Opioid Treatment
Abstract
1. Introduction
2. Results
2.1. Overview of Differentially Regulated miRNAs
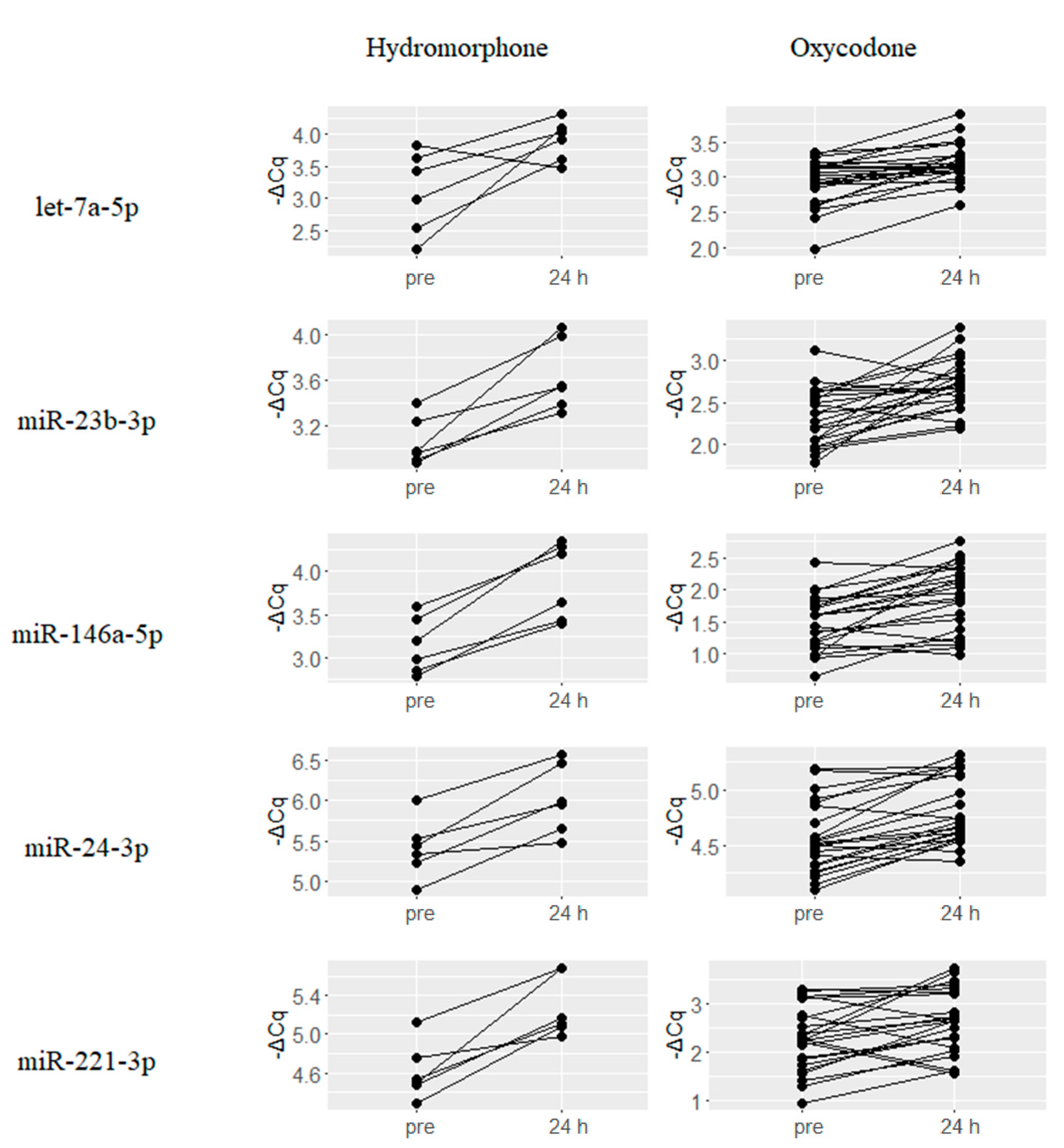
2.2. Upregulated miRNAs upon Hydromorphone and Oxycodone Treatment
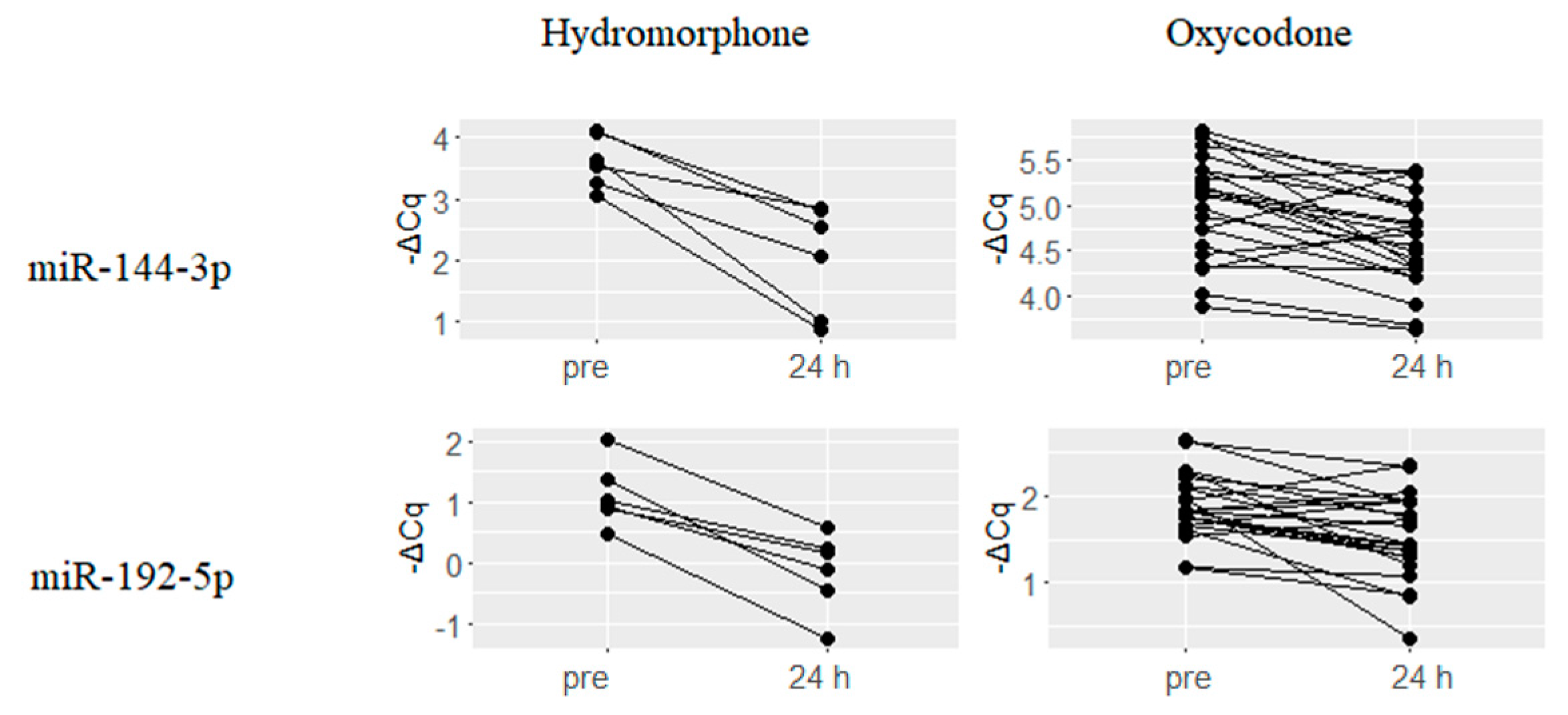
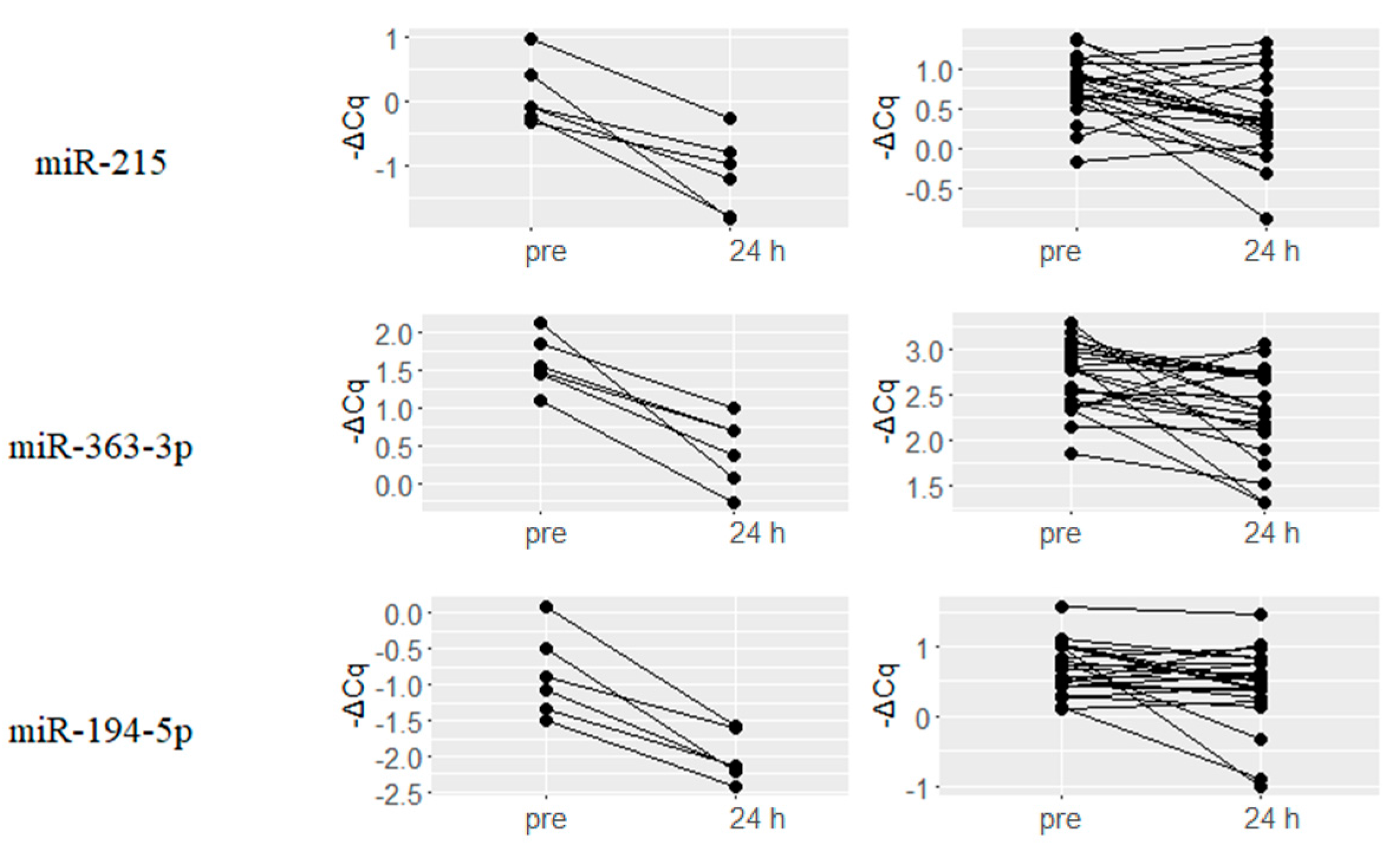
2.3. Downregulated miRNAs upon Hydromorphone and Oxycodone Treatment
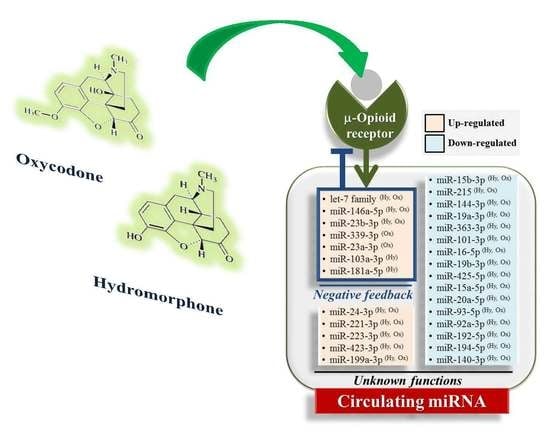
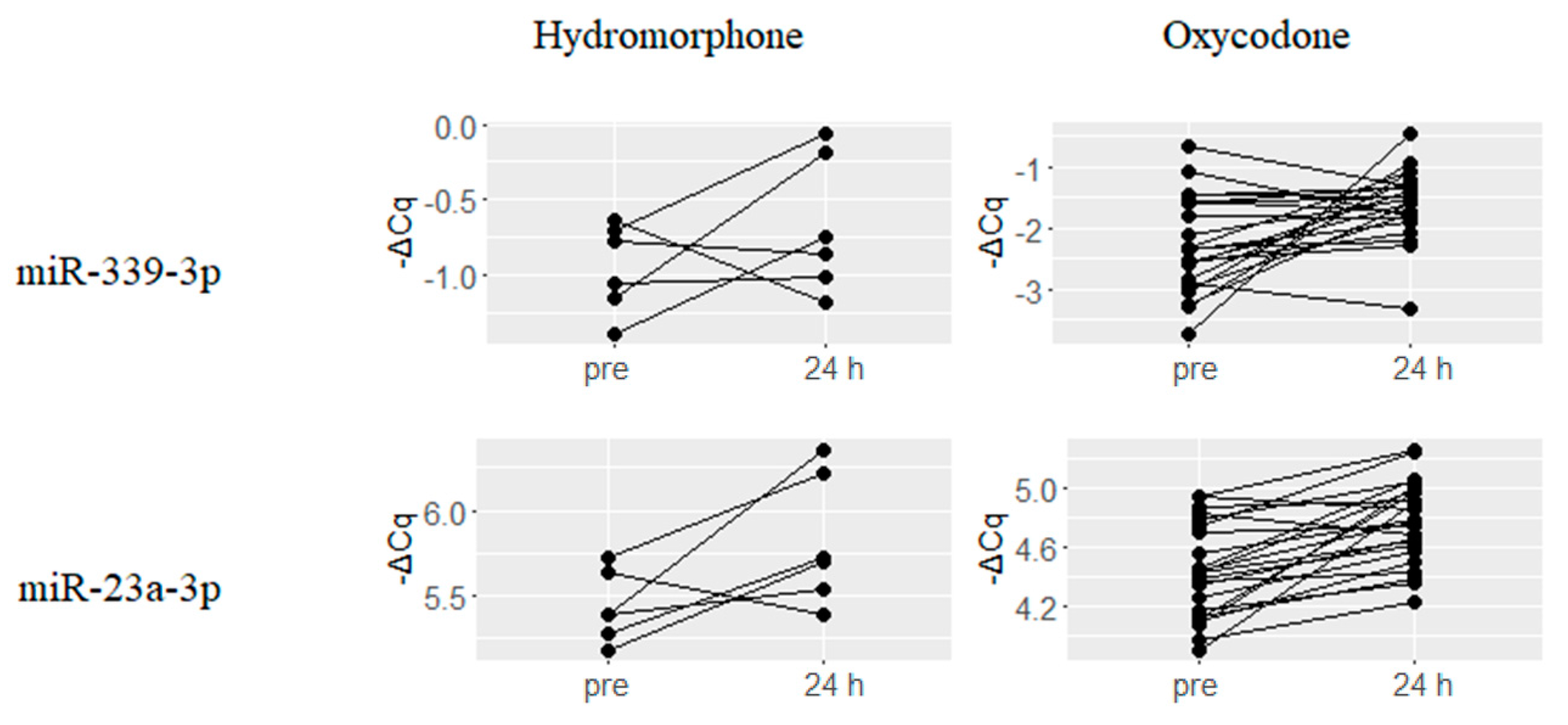
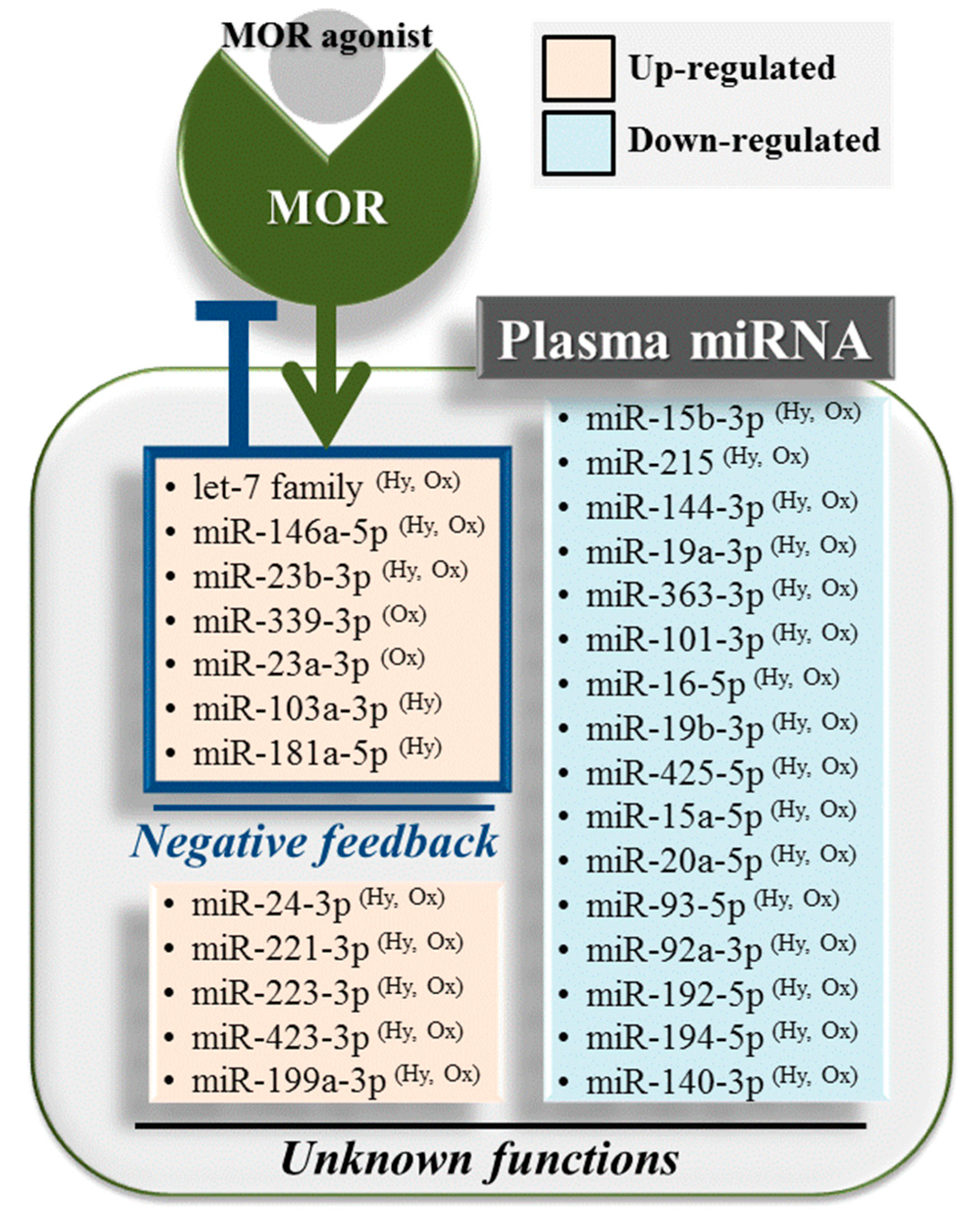
2.4. Hydromorphone- or Oxycodone-Induced miRNA Upregulation Is Associated with μ-Opioid Receptor (MOR) Signaling
3. Discussion
4. Materials and Methods
4.1. Summary of Clinical Studies
4.1.1. Hydromorphone Study (Japic-132167)
4.1.2. Oxycodone Study (Japic-163475)
4.2. miRNA Extraction from Plasma and qRT-PCR Analysis
4.3. Data Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| CNS | central nervous system |
| ER | extended release |
| miRNA | microRNA |
| MOR | μ-opioid receptor |
| PCR | polymerase chain reaction |
| qRT-PCR | quantitative real-time PCR |
| UTR | untranslated region |
References
- Al-Hasani, R.; Bruchas, M.R. Molecular mechanisms of opioid receptor-dependent signaling and behavior. Anesthesiology 2011, 115, 1363–1381. [Google Scholar] [CrossRef] [PubMed]
- Barbierato, M.; Zusso, M.; Skaper, S.D.; Giusti, P. MicroRNAs: Emerging role in the endogenous mu opioid system. CNS Neurol. Disord. Drug Targets 2015, 14, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Bruehl, S.; Apkarian, A.V.; Ballantyne, J.C.; Berger, A.; Borsook, D.; Chen, W.G.; Farrar, J.T.; Haythornthwaite, J.A.; Horn, S.D.; Iadarola, M.J.; et al. Personalized medicine and opioid analgesic prescribing for chronic pain: Opportunities and challenges. J. Pain 2013, 14, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, J.; Anderson, J.; Baumgartner, R.; Coimbra, A.; Schwarz, A.J.; Pendse, G.; Wallin, D.; Nutile, L.; Bishop, J.; George, E.; et al. Modulation of CNS pain circuitry by intravenous and sublingual doses of buprenorphine. NeuroImage 2012, 59, 3762–3773. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, J.; Anderson, J.; Schwarz, A.J.; Coimbra, A.; Baumgartner, R.; Pendse, G.; George, E.; Nutile, L.; Wallin, D.; Bishop, J.; et al. Imaging drugs with and without clinical analgesic efficacy. Neuropsychopharmacology 2011, 36, 2659–2673. [Google Scholar] [CrossRef] [PubMed]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef] [PubMed]
- Cortez, M.A.; Bueso-Ramos, C.; Ferdin, J.; Lopez-Berestein, G.; Sood, A.K.; Calin, G.A. MicroRNAs in body fluids—The mix of hormones and biomarkers. Nat. Rev. Clin. Oncol. 2011, 8, 467–477. [Google Scholar] [CrossRef] [PubMed]
- Vasudevan, S.; Tong, Y.; Steitz, J.A. Switching from repression to activation: MicroRNAs can up-regulate translation. Science 2007, 318, 1931–1934. [Google Scholar] [CrossRef] [PubMed]
- Im, H.I.; Kenny, P.J. MicroRNAs in neuronal function and dysfunction. Trends Neurosci. 2012, 35, 325–334. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chen, J.; Sen, S. MicroRNA as biomarkers and diagnostics. J. Cell. Physiol. 2016, 231, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Lopez, J.P.; Diallo, A.; Cruceanu, C.; Fiori, L.M.; Laboissiere, S.; Guillet, I.; Fontaine, J.; Ragoussis, J.; Benes, V.; Turecki, G.; et al. Biomarker discovery: Quantification of microRNAs and other small non-coding RNAs using next generation sequencing. BMC Med. Genom. 2015, 8, 35. [Google Scholar] [CrossRef] [PubMed]
- Brase, J.C.; Wuttig, D.; Kuner, R.; Sultmann, H. Serum microRNAs as non-invasive biomarkers for cancer. Mol. Cancer 2010, 9, 306. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Li, J.; Huang, B.; Liu, J.; Chen, X.; Chen, X.M.; Xu, Y.M.; Huang, L.F.; Wang, X.Z. Exosomes: Novel biomarkers for clinical diagnosis. Sci. World J. 2015, 2015, 657086. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.; Filant, J.; Moxley, K.M.; Sood, A.; McMeekin, S.; Ramesh, R. Exosomes: A role for naturally occurring nanovesicles in cancer growth, diagnosis and treatment. Curr. Gene Ther. 2015, 15, 182–192. [Google Scholar] [CrossRef] [PubMed]
- Turchinovich, A.; Weiz, L.; Burwinkel, B. Extracellular miRNAs: The mystery of their origin and function. Trends Biochem. Sci. 2012, 37, 460–465. [Google Scholar] [CrossRef] [PubMed]
- Higa, G.S.; de Sousa, E.; Walter, L.T.; Kinjo, E.R.; Resende, R.R.; Kihara, A.H. MicroRNAs in neuronal communication. Mol. Neurobiol. 2014, 49, 1309–1326. [Google Scholar] [CrossRef] [PubMed]
- Trescot, A.M.; Datta, S.; Lee, M.; Hansen, H. Opioid pharmacology. Pain Physician 2008, 11, S133–S153. [Google Scholar] [PubMed]
- Toyama, K.; Uchida, N.; Ishizuka, H.; Sambe, T.; Kobayashi, S. Single-dose evaluation of safety, tolerability and pharmacokinetics of newly formulated hydromorphone immediate-release and hydrophilic matrix extended-release tablets in healthy Japanese subjects without co-administration of an opioid antagonist. J. Clin. Pharmacol. 2015, 55, 975–984. [Google Scholar] [CrossRef] [PubMed]
- Toyama, K.; Furuie, H.; Ishizuka, H. Bioequivalence of an extended-release oxycodone hydrochloride tablet to a marketed reference product OXYCONTIN® in healthy Japanese subjects under fasting and fed conditions. Drugs R D 2017. [Google Scholar] [CrossRef]
- Fromm, B.; Billipp, T.; Peck, L.E.; Johansen, M.; Tarver, J.E.; King, B.L.; Newcomb, J.M.; Sempere, L.F.; Flatmark, K.; Hovig, E.; et al. A uniform system for the annotation of vertebrate microRNA genes and the evolution of the human microRNAome. Annu. Rev. Genet. 2015, 49, 213–242. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Yang, C.; Kirkmire, C.M.; Wang, Z.J. Regulation of opioid tolerance by let-7 family microRNA targeting the mu opioid receptor. J. Neurosci. 2010, 30, 10251–10258. [Google Scholar] [CrossRef] [PubMed]
- Dave, R.S.; Khalili, K. Morphine treatment of human monocyte-derived macrophages induces differential miRNA and protein expression: Impact on inflammation and oxidative stress in the central nervous system. J. Cell. Biochem. 2010, 110, 834–845. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Cao, D.L.; Jiang, B.C.; Yang, T.; Gao, Y.J. MicroRNA-146a-5p attenuates neuropathic pain via suppressing TRAF6 signaling in the spinal cord. Brain Behav. Immun. 2015, 49, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Zhang, L.; Law, P.Y.; Wei, L.N.; Loh, H.H. Long-term morphine treatment decreases the association of mu-opioid receptor (MOR1) mRNA with polysomes through miRNA23b. Mol. Pharmacol. 2009, 75, 744–750. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Zhang, Y.; Zheng, H.; Loh, H.H.; Law, P.Y. Morphine modulates mouse hippocampal progenitor cell lineages by upregulating miR-181a level. Stem Cells 2014, 32, 2961–2972. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekar, V.; Dreyer, J.L. MicroRNAs miR-124, let-7d and miR-181a regulate cocaine-induced plasticity. Mol. Cell. Neurosci. 2009, 42, 350–362. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Xu, J.; Xu, M.; Pasternak, G.W.; Pan, Y.X. Morphine regulates expression of mu-opioid receptor MOR-1A, an intron-retention carboxyl terminal splice variant of the mu-opioid receptor (OPRM1) gene via miR-103/miR-107. Mol. Pharmacol. 2014, 85, 368–380. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Hwang, C.K.; Zheng, H.; Wagley, Y.; Lin, H.Y.; Kim do, K.; Law, P.Y.; Loh, H.H.; Wei, L.N. MicroRNA 339 down-regulates mu-opioid receptor at the post-transcriptional level in response to opioid treatment. FASEB J. 2013, 27, 522–535. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014, 42, D68–D73. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Zeng, Y.; Zhang, X.; Chu, J.; Loh, H.H.; Law, P.Y. mu-Opioid receptor agonists differentially regulate the expression of miR-190 and NeuroD. Mol. Pharmacol. 2010, 77, 102–109. [Google Scholar] [CrossRef] [PubMed]
- Boyerinas, B.; Park, S.M.; Hau, A.; Murmann, A.E.; Peter, M.E. The role of let-7 in cell differentiation and cancer. Endocr. Relat. Cancer 2010, 17, F19–F36. [Google Scholar] [CrossRef] [PubMed]
- Ohshima, K.; Inoue, K.; Fujiwara, A.; Hatakeyama, K.; Kanto, K.; Watanabe, Y.; Muramatsu, K.; Fukuda, Y.; Ogura, S.; Yamaguchi, K.; et al. Let-7 microRNA family is selectively secreted into the extracellular environment via exosomes in a metastatic gastric cancer cell line. PLoS ONE 2010, 5, e13247. [Google Scholar] [CrossRef] [PubMed]
- Kiyosawa, N.; Manabe, S.; Yamoto, T.; Sanbuissho, A. Practical application of toxicogenomics for profiling toxicant-induced biological perturbations. Int. J. Mol. Sci. 2010, 11, 3397–3412. [Google Scholar] [CrossRef] [PubMed]
- Martin, F.; Thomson, T.M.; Sewer, A.; Drubin, D.A.; Mathis, C.; Weisensee, D.; Pratt, D.; Hoeng, J.; Peitsch, M.C. Assessment of network perturbation amplitudes by applying high-throughput data to causal biological networks. BMC Syst. Biol. 2012, 6, 54. [Google Scholar] [CrossRef] [PubMed]
- Data Analysis Guide For the miRCURY LNA™ Universal RT MicroRNA Ready-to-Use PCR Panels Using Exiqon GenEx Software Version 3 (February 2014). Available online: https://www.exiqon.com/ls/Documents/Scientific/Exiqon-data-analysis-guide.pdf (accessed on 4 July 2017).
- Mestdagh, P.; van Vlierberghe, P.; de Weer, A.; Muth, D.; Westermann, F.; Speleman, F.; Vandesompele, J. A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. 2009, 10, R64. [Google Scholar] [CrossRef] [PubMed]


| Hydromorphone (N = 6) | Oxycodone (N = 23) | ||||
|---|---|---|---|---|---|
| miRNA | ΔCq(24 h–Pre) | q-Value | miRNA | ΔCq(24 h–Pre) | q-Value |
| hsa-miR-127-3p | −2.85 | 0.003 | hsa-miR-361-3p | −1.18 | 0.005 |
| hsa-miR-182-5p | −1.87 | 0.044 | hsa-miR-208a | −0.89 | 0.003 |
| hsa-miR-339-5p | −1.47 | 0.006 | hsa-miR-339-3p | −0.69 | 0.027 |
| hsa-miR-374b-5p | −1.39 | 0.003 | hsa-miR-10b-5p | −0.62 | 0.002 |
| hsa-miR-28-5p | −1.33 | 0.009 | hsa-miR-28-3p | −0.59 | 0.042 |
| hsa-miR-551b-3p | −1.33 | 0.008 | hsa-miR-125a-5p | −0.51 | 0.004 |
| hsa-let-7d-5p | −1.25 | 0.003 | hsa-miR-342-3p | −0.49 | 0 |
| hsa-miR-26a-5p | −1.15 | 0.009 | hsa-miR-150-5p | −0.48 | 0.001 |
| hsa-miR-382-5p | −1.09 | 0.037 | hsa-miR-146b-5p | −0.46 | 0.005 |
| hsa-let-7f-5p | −1.05 | 0.018 | hsa-miR-197-3p | −0.40 | 0.01 |
| hsa-miR-154-5p | −1.01 | 0.039 | hsa-miR-146a-5p | −0.40 | 0.001 |
| hsa-let-7e-5p | −1.00 | 0.042 | hsa-miR-23b-3p | −0.38 | 0.001 |
| hsa-miR-191-5p | −0.98 | 0.013 | hsa-miR-221-3p | −0.36 | 0.027 |
| hsa-miR-103a-3p | −0.95 | 0.004 | hsa-miR-23a-3p | −0.33 | 0.001 |
| hsa-miR-423-3p | −0.91 | 0.004 | hsa-miR-199a-3p | −0.32 | 0.031 |
| hsa-miR-1 | −0.87 | 0.043 | hsa-let-7a-5p | −0.29 | 0.027 |
| hsa-miR-766-3p | −0.84 | 0.04 | hsa-miR-29a-3p | −0.29 | 0.027 |
| hsa-let-7a-5p | −0.80 | 0.009 | hsa-miR-223-3p | −0.27 | 0.011 |
| hsa-miR-181a-5p | −0.79 | 0.035 | hsa-miR-24-3p | −0.25 | 0.001 |
| hsa-miR-199a-3p | −0.77 | 0.003 | hsa-miR-142-3p | −0.24 | 0.011 |
| hsa-miR-146a-5p | −0.75 | 0.004 | hsa-miR-423-3p | −0.23 | 0.038 |
| hsa-miR-328 | −0.73 | 0.002 | hsa-miR-126-3p | −0.17 | 0.033 |
| hsa-let-7c | −0.73 | 0.019 | |||
| hsa-miR-221-3p | −0.67 | 0.012 | |||
| hsa-miR-151a-5p | −0.67 | 0.009 | |||
| hsa-miR-142-5p | −0.63 | 0.026 | |||
| hsa-miR-24-3p | −0.61 | 0.005 | |||
| hsa-miR-324-5p | −0.60 | 0.035 | |||
| hsa-miR-107 | −0.60 | 0.022 | |||
| hsa-miR-151a-3p | −0.60 | 0.011 | |||
| hsa-miR-23b-3p | −0.58 | 0.005 | |||
| hsa-miR-146b-5p | −0.56 | 0.035 | |||
| hsa-miR-223-3p | −0.52 | 0.02 | |||
| hsa-miR-652-3p | −0.49 | 0.006 | |||
| hsa-miR-140-5p | −0.41 | 0.043 | |||
| Hydromorphone (N = 6) | Oxycodone (N = 23) | ||||
|---|---|---|---|---|---|
| miRNA | ΔCq(24 h–Pre) | q-Value | miRNA | ΔCq(24 h–Pre) | q-Value |
| hsa-miR-885-5p | 2.47 | 0.011 | hsa-miR-182-5p | 1.34 | 0.004 |
| hsa-miR-122-5p | 1.71 | 0.001 | hsa-miR-15b-3p | 0.6 | 0 |
| hsa-miR-375 | 1.61 | 0.002 | hsa-miR-451a | 0.41 | 0.002 |
| hsa-miR-144-3p | 1.58 | 0.003 | hsa-miR-215 | 0.4 | 0.003 |
| hsa-miR-451a | 1.39 | 0.003 | hsa-miR-144-3p | 0.4 | 0.003 |
| hsa-let-7b-3p | 1.3 | 0.011 | hsa-miR-19a-3p | 0.39 | 0.001 |
| hsa-miR-192-5p | 1.26 | 0.001 | hsa-miR-363-3p | 0.39 | 0.003 |
| hsa-miR-215 | 1.25 | 0 | hsa-miR-192-5p | 0.35 | 0.003 |
| hsa-miR-363-3p | 1.17 | 0.003 | hsa-miR-101-3p | 0.34 | 0.003 |
| hsa-miR-194-5p | 1.14 | 0.004 | hsa-miR-16-5p | 0.33 | 0.003 |
| hsa-miR-660-5p | 1.14 | 0.003 | hsa-miR-106b-5p | 0.33 | 0.003 |
| hsa-miR-140-3p | 1.11 | 0.004 | hsa-miR-19b-3p | 0.32 | 0.002 |
| hsa-miR-378a-3p | 1.1 | 0 | hsa-miR-185-5p | 0.32 | 0.005 |
| hsa-miR-486-5p | 1.08 | 0.006 | hsa-miR-17-5p | 0.29 | 0.038 |
| hsa-miR-424-5p | 1.02 | 0.003 | hsa-miR-324-3p | 0.29 | 0.006 |
| hsa-miR-144-5p | 1 | 0.03 | hsa-miR-425-5p | 0.29 | 0.004 |
| hsa-miR-16-5p | 1 | 0.003 | hsa-miR-18b-5p | 0.27 | 0.004 |
| hsa-miR-502-3p | 0.98 | 0.012 | hsa-miR-18a-5p | 0.27 | 0.003 |
| hsa-miR-16-2-3p | 0.97 | 0.011 | hsa-miR-15a-5p | 0.27 | 0.01 |
| hsa-miR-99a-5p | 0.95 | 0.004 | hsa-miR-106a-5p | 0.26 | 0.005 |
| hsa-miR-25-3p | 0.95 | 0.003 | hsa-miR-20a-5p | 0.26 | 0.01 |
| hsa-miR-365a-3p | 0.93 | 0.042 | hsa-miR-194-5p | 0.25 | 0.031 |
| hsa-miR-15a-5p | 0.91 | 0.003 | hsa-miR-30b-5p | 0.25 | 0.004 |
| hsa-miR-10b-5p | 0.88 | 0.042 | hsa-miR-140-3p | 0.24 | 0.031 |
| hsa-miR-92a-3p | 0.83 | 0.003 | hsa-miR-29b-3p | 0.24 | 0.028 |
| hsa-miR-101-3p | 0.78 | 0.011 | hsa-miR-93-5p | 0.23 | 0.032 |
| hsa-miR-20b-5p | 0.75 | 0.044 | hsa-miR-92a-3p | 0.23 | 0.028 |
| hsa-miR-210 | 0.74 | 0.012 | hsa-miR-484 | 0.21 | 0.014 |
| hsa-miR-497-5p | 0.73 | 0.042 | hsa-let-7g-5p | 0.21 | 0.014 |
| hsa-miR-19b-3p | 0.72 | 0.018 | hsa-miR-148b-3p | 0.2 | 0.017 |
| hsa-miR-30a-5p | 0.66 | 0.03 | hsa-miR-30e-5p | 0.2 | 0.01 |
| hsa-miR-19a-3p | 0.61 | 0.019 | |||
| hsa-miR-15b-3p | 0.59 | 0.036 | |||
| hsa-miR-425-5p | 0.42 | 0.031 | |||
| hsa-miR-93-5p | 0.37 | 0.042 | |||
| hsa-miR-20a-5p | 0.35 | 0.046 | |||
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Toyama, K.; Kiyosawa, N.; Watanabe, K.; Ishizuka, H. Identification of Circulating miRNAs Differentially Regulated by Opioid Treatment. Int. J. Mol. Sci. 2017, 18, 1991. https://doi.org/10.3390/ijms18091991
Toyama K, Kiyosawa N, Watanabe K, Ishizuka H. Identification of Circulating miRNAs Differentially Regulated by Opioid Treatment. International Journal of Molecular Sciences. 2017; 18(9):1991. https://doi.org/10.3390/ijms18091991
Chicago/Turabian StyleToyama, Kaoru, Naoki Kiyosawa, Kenji Watanabe, and Hitoshi Ishizuka. 2017. "Identification of Circulating miRNAs Differentially Regulated by Opioid Treatment" International Journal of Molecular Sciences 18, no. 9: 1991. https://doi.org/10.3390/ijms18091991
APA StyleToyama, K., Kiyosawa, N., Watanabe, K., & Ishizuka, H. (2017). Identification of Circulating miRNAs Differentially Regulated by Opioid Treatment. International Journal of Molecular Sciences, 18(9), 1991. https://doi.org/10.3390/ijms18091991