An Insight into T-DNA Integration Events in Medicago sativa
Abstract
:1. Introduction
- The ssT-DNA molecules can be directly integrated according to the model of Tinland [17]; however, such model does not explain the production of complex T-DNA insertions. Therefore, a model whereby the ssT-DNA is converted into double stranded T-DNA (dsT-DNA) prior to integration was proposed [5,18,19,20].
- The presence of the so-called filler DNA, that is, DNA sequences from unknown sources often found between tandemly repeated copies of T-DNA or between the borders and the gDNA, points to a role of the double strand break (DSB) repair machinery in T-DNA integration [5]. The dsT-DNA, that seems to be abundant in Agrobacterium infected plant cells [16], can be recruited by the plant cell’s own DSB repair machinery, thus leading to end joining between ds-molecules and/or integration; this pathway could represent the most likely route for T-DNA integration [21]. Notably, Singer et al. [16] observed the formation of complex T-DNA circular structures in infected cells resembling the observed complex patterns of integration. Recently, van Kregten et al. [22] demonstrated the involvement of polymerase theta (Pol θ), a DSB repair enzyme, in T-DNA integration in Arabidopsis: the primer–template switching ability of this polymerase can explain the presence of filler DNA.
- The availability of DSBs could be a limiting factor in the integration of T-DNA; however, a role of other types of lesions, such as single strand breaks (SSBs) cannot be excluded [4]. DSBs can be repaired either by the non-homologous end joining (NHEJ) pathway or by the homologous recombination (HR) pathway; the former seems to be the most frequent in plants, although the results are contradictory and a potential influence of the method of transformation used and of the cell type and/or developmental stage have been suggested. The evidences on the chromatin modifications that are essential for the DSB repair response and important for T-DNA integration have been reviewed, and models of integrations proposed [4].
2. Results
2.1. Isolation of Sequences Flanking T-DNA Insertions
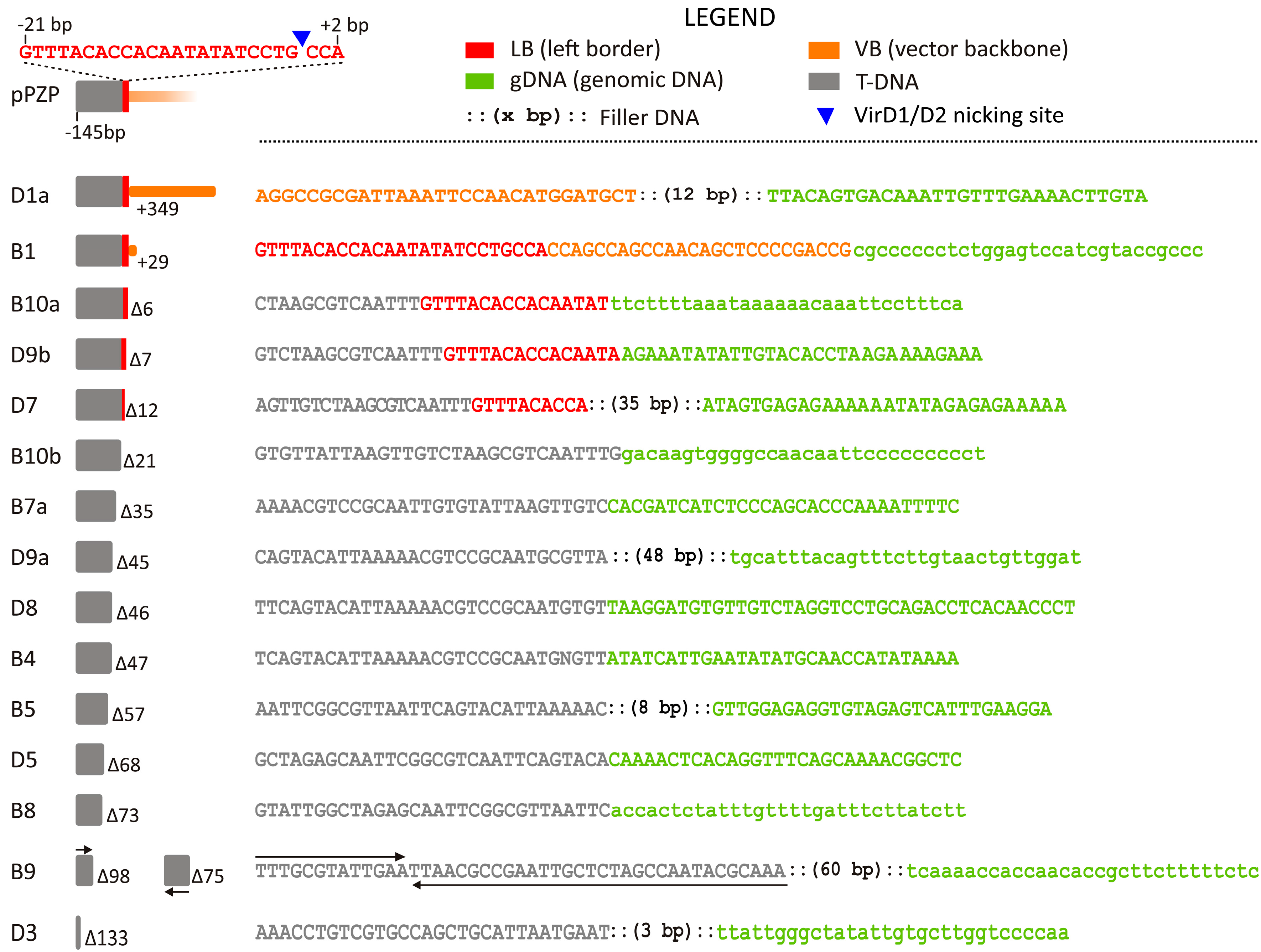
2.1.1. LB Junctions
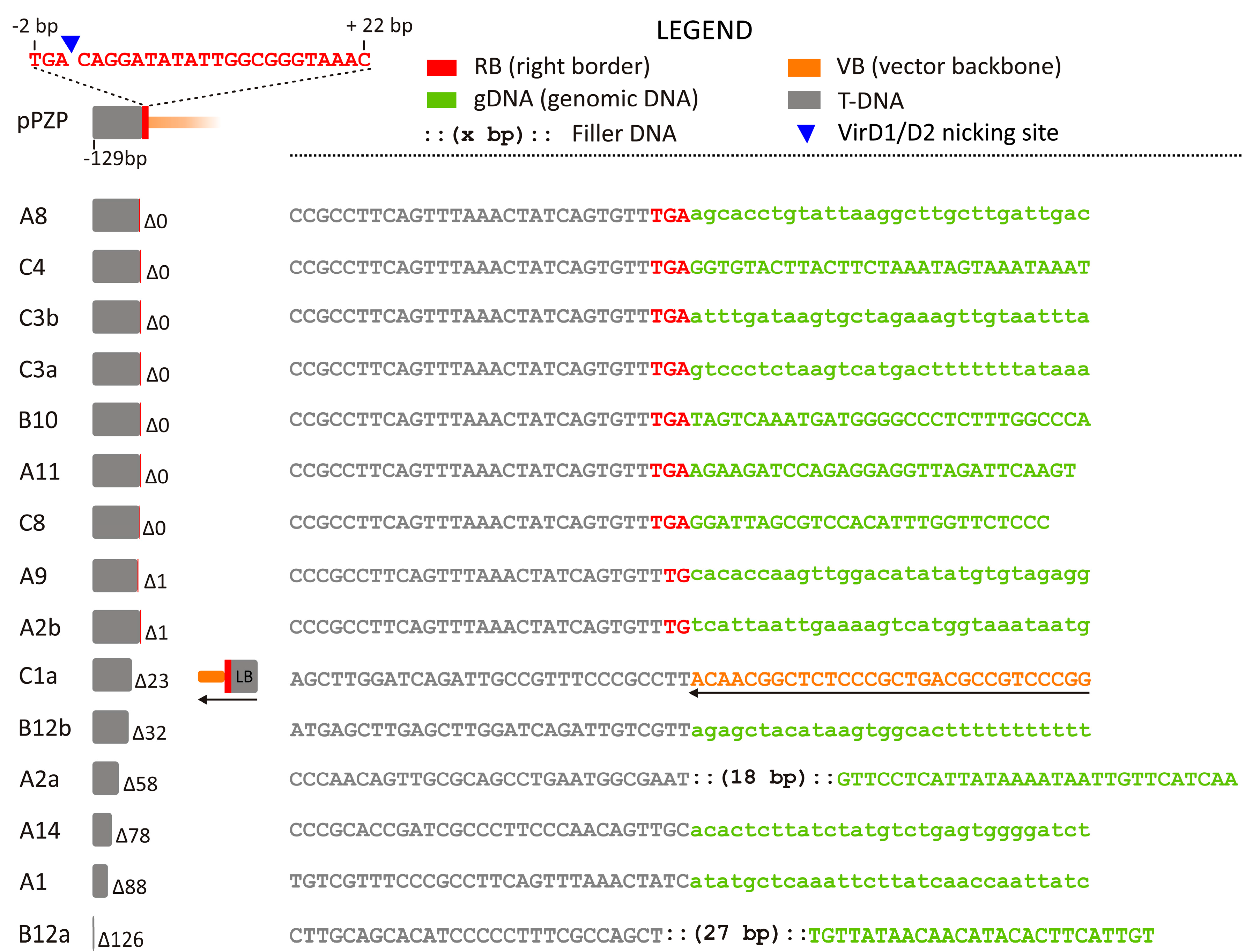
2.1.2. RB Junctions
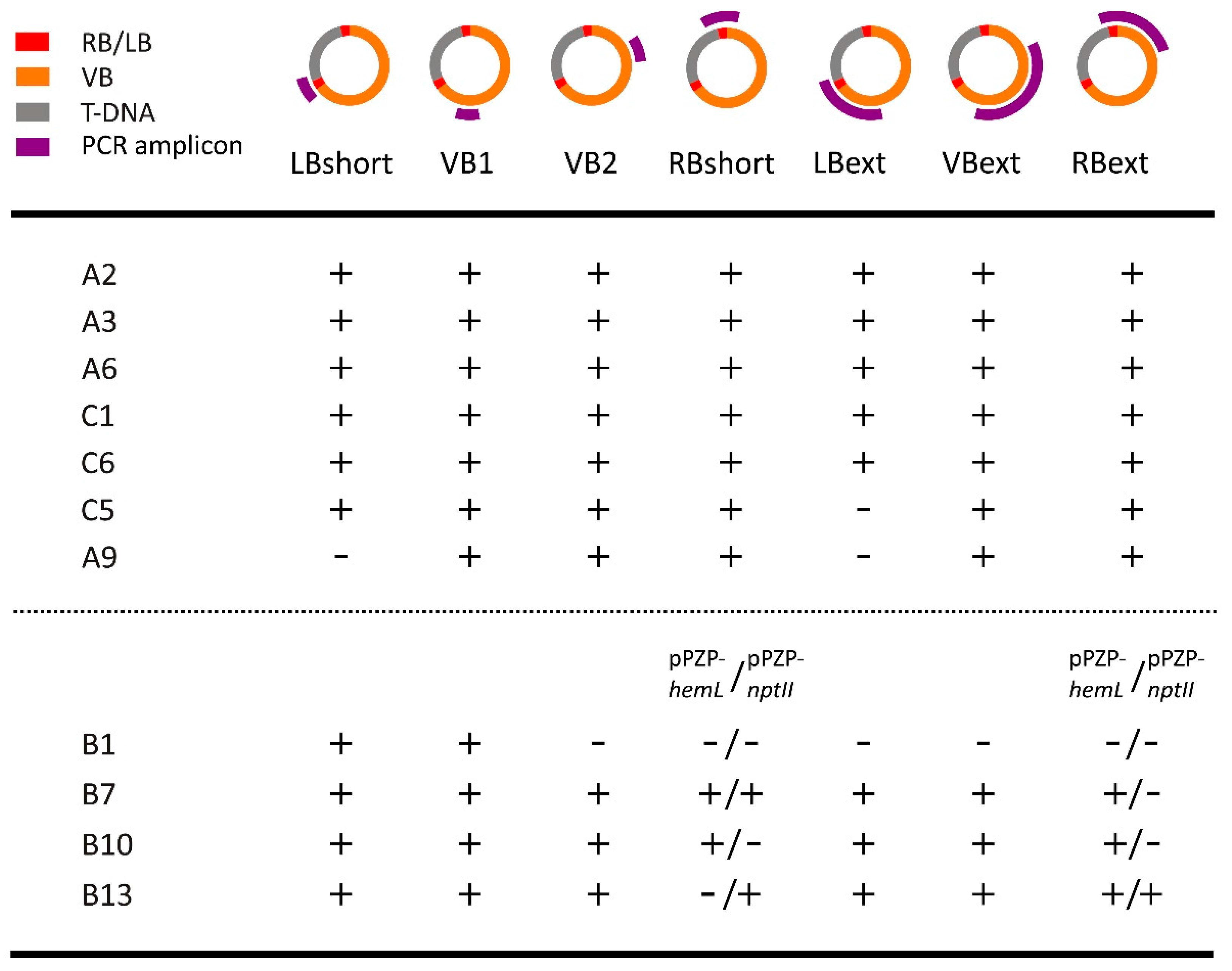
2.2. Polymerase Chain Reaction (PCR) Detection of Vector Backbone Sequences
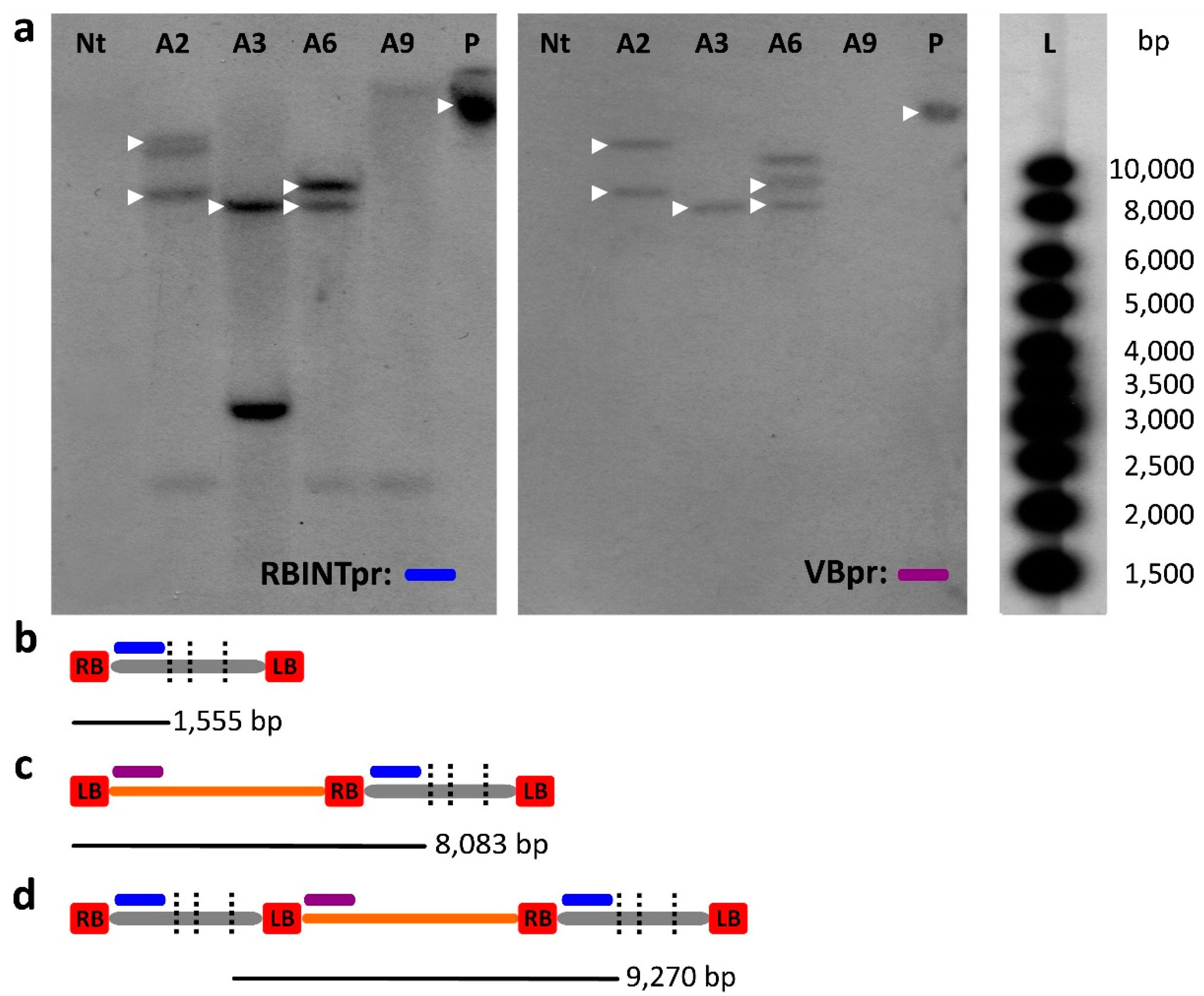
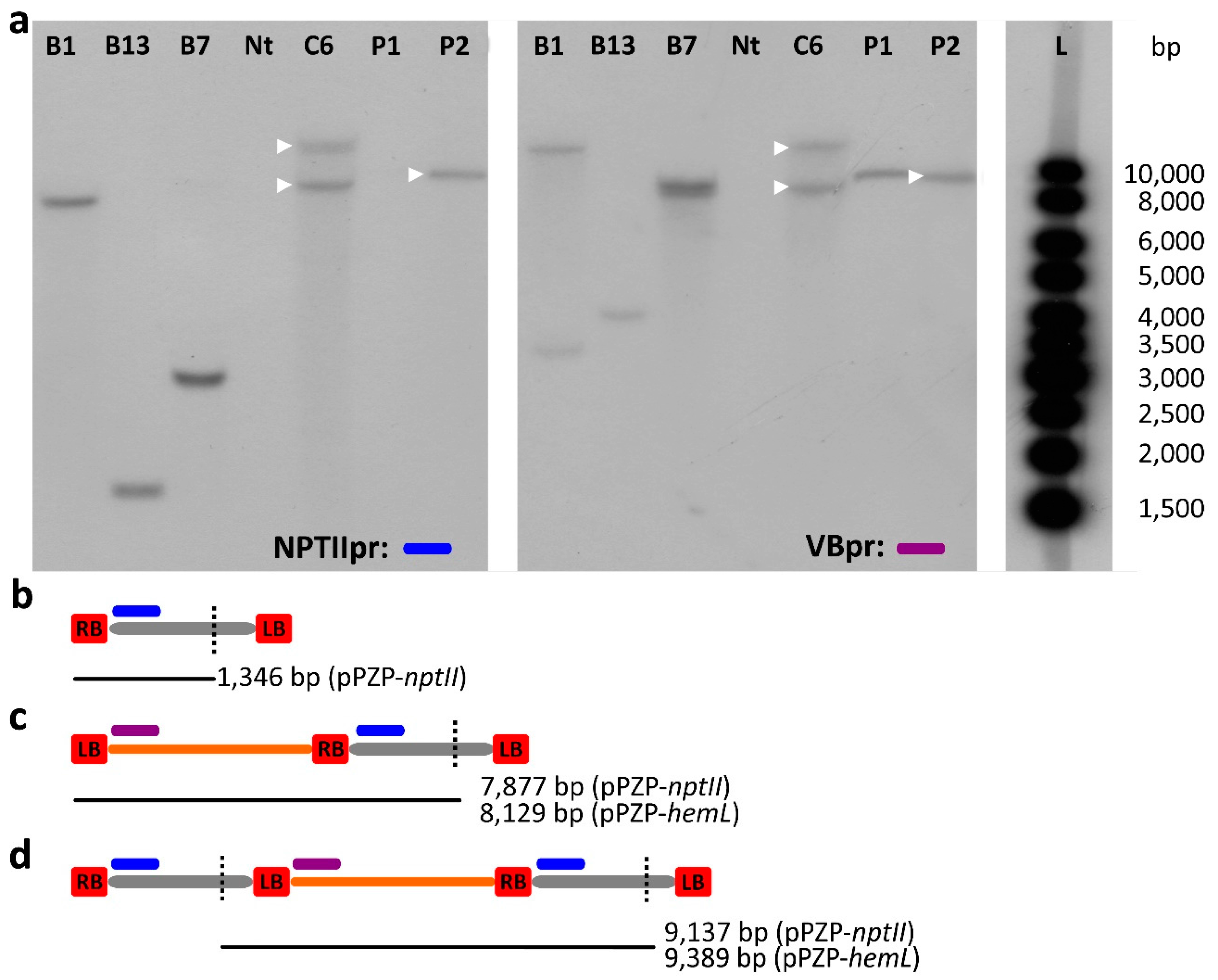
2.3. Southern Blot Analysis
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Isolation of Sequences Flanking T-DNA Insertions
4.3. PCR Detection of Vector Backbone Sequences
4.4. Southern Hybridization Analysis
5. Conclusions
- (a)
- Launching the T-DNA from the Agrobacterium chromosome may reduce the risk of transferring sequences belonging to the binary vector (including antibiotic resistance genes for bacterial selection); however, engineering the bacterial chromosome adds complication to the procedures, and a decrease in the transformation efficiency may result [27]; it should also be considered that rare cases of transfer of sequences belonging to Agrobacterium chromosome are documented [10,11,12];
- (b)
- (c)
- Using plant-derived sequences for vector construction (e.g., plant-derived SMGs, cisgenesis) [68,69,70] can relieve the perceived risk of genetically modified plants. The application of new breeding techniques such as genome editing is also offering new tools for precise modification of the alfafa genome.
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kyndt, T.; Quispe, D.; Zhai, H.; Jarret, R.; Ghislain, M.; Liu, Q.; Gheysen, G.; Kreuze, J.F. The genome of cultivated sweet potato contains Agrobacterium T-DNAs with expressed genes: An example of a naturally transgenic food crop. Proc. Natl. Acad. Sci. USA 2015, 112, 5844–5849. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, K.; Yamashita, I.; Tanaka, N. Tobacco plants were transformed by Agrobacterium rhizogenes infection during their evolution. Plant J. 2002, 32, 775–787. [Google Scholar] [CrossRef] [PubMed]
- Podevin, N.; de Buck, S.; de Wilde, C.; Depicker, A. Insights into recognition of the T-DNA border repeats as termination sites for T-strand synthesis by Agrobacterium tumefaciens. Transgenic Res. 2006, 15, 557–571. [Google Scholar] [CrossRef] [PubMed]
- Magori, S.; Citovsky, V. Epigenetic control of Agrobacterium T-DNA integration. Biochim. Biophys. Acta 2011, 1809, 388–394. [Google Scholar] [CrossRef] [PubMed]
- Tzfira, T.; Li, J.; Lacroix, B.; Citovsky, V. Agrobacterium T-DNA integration: Molecules and models. Trends Genet. 2004, 20, 375–383. [Google Scholar] [CrossRef] [PubMed]
- Pitzschke, A.; Hirt, H. New insights into an old story: Agrobacterium-induced tumour formation in plants by plant transformation. EMBO J. 2010, 29, 1021–1032. [Google Scholar] [CrossRef] [PubMed]
- Tzfira, T.; Citovsky, V. Agrobacterium-mediated genetic transformation of plants: Biology and biotechnology. Curr. Opin. Biotechnol. 2006, 17, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Kononov, M.E.; Bassuner, B.; Gelvin, S.B. Integration of T-DNA binary vector “backbone” sequences into the tobacco genome: Evidence for multiple complex patterns of integration. Plant J. 1997, 11, 945–957. [Google Scholar] [CrossRef] [PubMed]
- Wenck, A.; Czakó, M.; Kanevski, I.; Márton, L. Frequent collinear long transfer of DNA inclusive of the whole binary vector during Agrobacterium-mediated transformation. Plant Mol. Biol. 1997, 34, 913–922. [Google Scholar] [CrossRef] [PubMed]
- Ulker, B.; Li, Y.; Rosso, M.G.; Logemann, E.; Somssich, I.E.; Weisshaar, B. T-DNA-mediated transfer of Agrobacterium tumefaciens chromosomal DNA into plants. Nat. Biotechnol. 2008, 26, 1015–1017. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-R.; An, G. Bacterial transposons are co-transferred with T-DNA to rice chromosomes during Agrobacterium-mediated transformation. Mol. Cells 2012, 33, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Philips, J.G.; Naim, F.; Lorenc, M.T.; Dudley, K.J.; Hellens, R.P.; Waterhouse, P.M. The widely used Nicotiana benthamiana 16c line has an unusual T-DNA integration pattern including a transposon sequence. PLoS ONE 2017, 12, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Lacroix, B.; Guo, J.; Citovsky, V. The Agrobacterium VirE2 effector interacts with multiple members of the Arabidopsis VIP1 protein family. Mol. Plant Pathol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Pan, S.Q. Agrobacterium delivers VirE2 protein into host cells via clathrin-mediated endocytosis. Sci. Adv. 2017, 3, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Gheysen, G.; Villarroel, R.; van Montagu, M. Illegitimate recombination in plants: A model for T-DNA integration. Genes Dev. 1991, 5, 287–297. [Google Scholar] [CrossRef] [PubMed]
- Singer, K.; Shiboleth, Y.M.; Li, J.; Tzfira, T. Formation of complex extrachromosomal T-DNA structures in Agrobacterium tumefaciens-infected plants. Plant Physiol. 2012, 160, 511–522. [Google Scholar] [CrossRef] [PubMed]
- Tinland, B. The integration of T-DNA into plant genomes. Trends Plant Sci. 1996, 1, 178–184. [Google Scholar] [CrossRef]
- De Buck, S.; Jacobs, A.; van Montagu, M.; Depicker, A. The DNA sequences of T-DNA junctions suggest that complex T-DNA loci are formed by a recombination process resembling T-DNA integration. Plant J. 1999, 20, 295–304. [Google Scholar] [CrossRef] [PubMed]
- De Neve, M.; de Buck, S.; Jacobs, A.; van Montagu, M.; Depicker, A. T-DNA integration patterns in co-transformed plant cells suggest that T-DNA repeats originate from co-integration of separate T-DNAs. Plant J. 1997, 11, 15–29. [Google Scholar] [CrossRef] [PubMed]
- Mayerhofer, R.; Koncz-Kalman, Z.; Nawrath, C.; Bakkeren, G.; Crameri, A.; Angelis, K.; Redei, G.P.; Schell, J.; Hohn, B.; Koncz, C. T-DNA integration: A mode of illegitimate recombination in plants. EMBO J. 1991, 10, 697–704. [Google Scholar] [PubMed]
- Hu, Y.; Chen, Z.; Zhuang, C.; Huang, J. Cascade of chromosomal rearrangements caused by a heterogeneous T-DNA integration supports the double-stranded break repair model for T-DNA integration. Plant J. 2017, 90, 954–965. [Google Scholar] [CrossRef] [PubMed]
- Van Kregten, M.; de Pater, S.; Romeijn, R.; van Schendel, R.; Hooykaas, P.J.; Tijsterman, M. T-DNA integration in plants results from Polymerase-θ-mediated DNA repair. Nat. Plants 2016, 2, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Kölliker, R.; Rosellini, D.; Wang, Z.Y. Development and application of biotechnological and molecular genetic tools. In Fodder Crops and Amenity Grasses; Springer: New York, NY, USA, 2010; pp. 89–113. [Google Scholar]
- Ferradini, N.; Nicolia, A.; Capomaccio, S.; Veronesi, F.; Rosellini, D. Assessment of simple marker-free genetic transformation techniques in alfalfa. Plant Cell Rep. 2011, 30, 1991–2000. [Google Scholar] [CrossRef] [PubMed]
- Ferradini, N.; Nicolia, A.; Capomaccio, S.; Veronesi, F.; Rosellini, D. A point mutation in the Medicago sativa GSA gene provides a novel, efficient, selectable marker for plant genetic engineering. J. Biotechnol. 2011, 156, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Rosellini, D.; Capomaccio, S.; Ferradini, N.; Savo Sardaro, M.L.; Nicolia, A.; Veronesi, F. Non-antibiotic, efficient selection for alfalfa genetic engineering. Plant Cell Rep. 2007, 26, 1035–1044. [Google Scholar] [CrossRef] [PubMed]
- Oltmanns, H.; Frame, B.; Lee, L.-Y.; Johnson, S.; Li, B.; Wang, K.; Gelvin, S.B. Generation of backbone-free, low transgene copy plants by launching T-DNA from the Agrobacterium chromosome. Plant Physiol. 2010, 152, 1158–1166. [Google Scholar] [CrossRef] [PubMed]
- Van der Graaff, E.; den Dulk-Ras, A.; Hooykaas, P.J. Deviating T-DNA transfer from Agrobacterium tumefaciens to plants. Plant Mol. Biol. 1996, 31, 677–681. [Google Scholar] [CrossRef] [PubMed]
- Lange, M.; Vincze, E.; Møller, M.G.; Holm, P.B. Molecular analysis of transgene and vector backbone integration into the barley genome following Agrobacterium-mediated transformation. Plant Cell Rep. 2006, 25, 815–820. [Google Scholar] [CrossRef] [PubMed]
- Confalonieri, M.; Borghetti, R.; Macovei, A.; Testoni, C.; Carbonera, D.; Fevereiro, M.P.; Rommens, C.; Swords, K.; Piano, E.; Balestrazzi, A. Backbone-free transformation of barrel medic (Medicago truncatula) with a Medicago-derived transfer DNA. Plant Cell Rep. 2010, 29, 1013–1021. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Williams, E.J.; Shen, J.; Johnson, S.; Lowe, B.; Radke, S.; Strickland, S.; Esser, J.A.; Petersen, M.W.; Gilbertson, L.A. Enhanced production of single copy backbone-free transgenic plants in multiple crop species using binary vectors with a pRi replication origin in Agrobacterium tumefaciens. Transgenic Res. 2011, 20, 773–786. [Google Scholar] [CrossRef] [PubMed]
- Fu, D.; St. Amand, P.C.; Xiao, Y.; Muthukrishnan, S.; Liang, G.H. Characterization of T-DNA integration in creeping bentgrass. Plant Sci. 2006, 170, 225–237. [Google Scholar] [CrossRef]
- Huang, S.; Gilbertson, L.A.; Adams, T.H.; Malloy, K.P.; Reisenbigler, E.K.; Birr, D.H.; Snyder, M.W.; Zhang, Q.; Luethy, M.H. Generation of marker-free transgenic maize by regular two-border Agrobacterium transformation vectors. Transgenic Res. 2004, 13, 451–461. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Cai, L.; Cheng, J.; Mao, H.; Fan, X.; Meng, Z.; Chan, K.M.; Zhang, H.; Qi, J.; Ji, L.; et al. Transgene integration and organization in cotton (Gossypium hirsutum L.) genome. Transgenic Res. 2008, 17, 293–306. [Google Scholar] [CrossRef] [PubMed]
- Gambino, G.; Chitarra, W.; Maghuly, F.; Laimer, M.; Boccacci, P.; Torello, M.D.; Gribaudo, I. Characterization of T-DNA insertions in transgenic grapevines obtained by Agrobacterium-mediated transformation. Mol. Breed. 2009, 24, 305–320. [Google Scholar] [CrossRef]
- Hanson, B.; Engler, D.; Moy, Y.; Newman, B.; Ralston, E.; Gutterson, N. A simple method to enrich an Agrobacterium-transformed population for plants containing only T-DNA sequences. Plant J. 1999, 19, 727–734. [Google Scholar] [CrossRef] [PubMed]
- Cluster, P.D.; O’Dell, M.; Metzlaff, M.; Flavell, R.B. Details of T-DNA structural organization from a transgenic Petunia population exhibiting co-suppression. Plant Mol. Biol. 1996, 32, 1197–1203. [Google Scholar] [CrossRef] [PubMed]
- Rommens, C.M.; Humara, J.M.; Ye, J.; Yan, H.; Richael, C.; Zhang, L.; Perry, R.; Swords, K. Crop improvement through modification of the plant’s own genome. Plant Physiol. 2004, 135, 421–431. [Google Scholar] [CrossRef] [PubMed]
- Yin, Z.; Wang, G.-L. Evidence of multiple complex patterns of T-DNA integration into the rice genome. TAG Theor. Appl. Genet. 2000, 100, 461–470. [Google Scholar] [CrossRef]
- Sallaud, C.; Meynard, D.; van Boxtel, J.; Gay, C.; Bès, M.; Brizard, J. P.; Larmande, P.; Ortega, D.; Raynal, M.; Portefaix, M.; et al. Highly efficient production and characterization of T-DNA plants for rice ( Oryza sativa L.) functional genomics. Theor. Appl. Genet. 2003, 106, 1396–1408. [Google Scholar] [CrossRef] [PubMed]
- Afolabi, A.; Worland, B.; Snape, J. W.; Vain, P. A large-scale study of rice plants transformed with different T-DNAs provides new insights into locus composition and T-DNA linkage configurations. Theor. Appl. Genet. 2004, 109, 815–826. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.-H.; Ramm, K.; Eamens, A.L.; Dennis, E.S.; Upadhyaya, N.M. Transgene structures suggest that multiple mechanisms are involved in T-DNA integration in plants. Plant Sci. 2006, 171, 308–322. [Google Scholar] [CrossRef] [PubMed]
- Wu, E.; Lenderts, B.; Glassman, K.; Berezowska, K.M.; Christensen, H.; Asmus, T.; Zhen, S.; Chu, U.; Cho, M.J.; Zhao, Z.Y. Optimized Agrobacterium-mediated sorghum transformation protocol and molecular data of transgenic sorghum plants. In Vitro Cell. Dev. Biol. Plant 2014, 50, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Abdal-Aziz, S.A.; Pliego, A.F.; Quesada, M.A.; Mercado, J.A. Evidence of frequent integration of non-T-DNA vector backbone sequences in transgenic strawberry plant. J. Biosci. Bioeng. 2006, 101, 508–510. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.P.; Yu, X.D.; Sun, Y.W.; Jones, H.D.; Xia, L.Q. Generation of marker- and/or backbone-free transgenic wheat plants via Agrobacterium-mediated transformation. Front. Plant Sci. 2016, 7, 1324. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Sparks, C.A.; Jones, H.D. Characterisation of T-DNA loci and vector backbone sequences in transgenic wheat produced by Agrobacterium-mediated transformation. Mol. Breed. 2006, 18, 195–208. [Google Scholar] [CrossRef]
- De Buck, S.; Podevin, N.; Nolf, J.; Jacobs, A.; Depicker, A. The T-DNA integration pattern in Arabidopsis transformants is highly determined by the transformed target cell. Plant J. 2009, 60, 134–145. [Google Scholar] [CrossRef] [PubMed]
- Petti, C.; Wendt, T.; Meade, C.; Mullins, E. Evidence of genotype dependency within Agrobacterium tumefaciens in relation to the integration of vector backbone sequence in transgenic Phytophthora infestans-tolerant potato. J. Biosci. Bioeng. 2009, 107, 301–306. [Google Scholar] [CrossRef] [PubMed]
- Hajdukiewicz, P.; Svab, Z.; Maliga, P. The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Mol. Biol. 1994, 25, 989–994. [Google Scholar] [CrossRef] [PubMed]
- Vain, P.; Harvey, A.; Worland, B.; Ross, S.; Snape, J.W.; Lonsdale, D. The effect of additional virulence genes on transformation efficiency, transgene integration and expression in rice plants using the pGreen/pSoup dual binary vector system. Transgenic Res. 2004, 13, 593–603. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.G.; Chen, Y. High-efficiency thermal asymmetric interlaced PCR for amplification of unknown flanking sequences. Biotechniques 2007, 43, 649–656. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-R.; Lee, J.; Jun, S.-H.; Park, S.; Kang, H.-G.; Kwon, S.; An, G. Transgene structures in T-DNA-inserted rice plants. Plant Mol. Biol. 2003, 52, 761–773. [Google Scholar] [CrossRef] [PubMed]
- De Buck, S.; De Wilde, C.; Van Montagu, M.; Depicker, A. T-DNA vector backbone sequences are frequently integrated into the genome of transgenic plants obtained by Agrobacterium-mediated transformation. Mol. Breed. 2000, 6, 459–468. [Google Scholar] [CrossRef]
- Kumar, S.; Fladung, M. Transgene integration in aspen: Structures of integration sites and mechanism of T-DNA integration. Plant J. 2002, 31, 543–551. [Google Scholar] [CrossRef] [PubMed]
- Bingham, G. Registration of alfalfa hybrid Regen-Sy germplasm for tissue culture and transformation research. Crop Sci. 1991, 31, 1098. [Google Scholar] [CrossRef]
- Rozen, S.; Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. In Bioinformatics Methods and Protocols; Humana Press: New York, NY, USA, 1999; pp. 365–386. [Google Scholar]
- Rong, L.J.; Karcher, S.J.; Gelvin, S.B. Genetic and molecular analyses of picA, a plant-inducible locus on the Agrobacterium tumefaciens chromosome. J. Bacteriol. 1991, 173, 5110–5120. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Russell, D.W. Molecular Cloning; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2012. [Google Scholar]
- Kirik, A.; Salomon, S.; Puchta, H. Species-specific double-strand break repair and genome evolution in plants. EMBO J. 2000, 19, 5562–5566. [Google Scholar] [CrossRef] [PubMed]
- Manova, V.; Gruszka, D. DNA damage and repair in plants-from models to crops. Front. Plant Sci. 2015, 6, 885. [Google Scholar] [CrossRef] [PubMed]
- Orel, N.; Puchta, H. Differences in the processing of DNA ends in Arabidopsis thaliana and tobacco: Possible implication for genome evolution. Plant Mol. Biol. 2003, 523–531. [Google Scholar] [CrossRef]
- Windels, P.; de Buck, S.; van Bockstaele, E.; de Loose, M.; Depicker, A. T-DNA integration in Arabidopsis chromosomes. Presence and origin of filler DNA sequences. Plant Physiol. 2003, 133, 2061–2068. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Krishnakumar, V.; Bidwell, S.; Rosen, B.; Chan, A.; Zhou, S.; Gentzbittel, L.; Childs, K. L.; Yandell, M.; Gundlach, H.; et al. An improved genome release (version Mt4.0) for the model legume Medicago truncatula. BMC Genom. 2014, 15, 312. [Google Scholar] [CrossRef] [PubMed]
- Porceddu, A.; Panara, F.; Calderini, O.; Molinari, L.; Taviani, P.; Lanfaloni, L.; Scotti, C.; Carelli, M.; Scaramelli, L.; Bruschi, G.; et al. An Italian functional genomic resource for Medicago truncatula. BMC Res. Notes 2008, 1, 129. [Google Scholar] [CrossRef] [PubMed]
- Scholte, M.; Erfurth, I.; Rippa, S.; Mondy, S.; Cosson, V.; Durand, P.; Breda, C.; Trinh, H.; Rodriguez, -L.I.; Kondorosi, E.; et al. T-DNA tagging in the model legume Medicago truncatula allows efficient gene discovery. Mol. Breed. 2002, 10, 203–215. [Google Scholar] [CrossRef]
- Nicolia, A.; Manzo, A.; Veronesi, F.; Rosellini, D. An overview of the last 10 years of genetically engineered crop safety research. Crit. Rev. Biotechnol. 2014, 34, 77–88. [Google Scholar] [CrossRef] [PubMed]
- Kondrák, M.; van der Meer, I.M.; Bánfalvi, Z. Generation of marker- and backbone-free transgenic potatoes by site-specific recombination and a bi-functional marker gene in a non-regular one-border agrobacterium transformation vector. Transgenic Res. 2006, 15, 729–737. [Google Scholar] [CrossRef] [PubMed]
- Rosellini, D. Selectable marker genes from plants: Reliability and potential. In Vitro Cell. Dev. Biol. Plant 2011, 47, 222–233. [Google Scholar] [CrossRef]
- Rosellini, D. Selectable markers and reporter genes: A well-furnished toolbox for plant science and genetic engineering. Crit. Rev. Plant Sci. 2012, 31, 401–453. [Google Scholar] [CrossRef]
- Holme, I.B.; Wendt, T.; Holm, P.B. Intragenesis and cisgenesis as alternatives to transgenic crop development. Plant Biotechnol. J. 2013, 11, 395–407. [Google Scholar] [CrossRef] [PubMed]
| Species | Agrobacterium Strain | Vector | VB % * | References |
|---|---|---|---|---|
| Arabidopsis | EHA101, GV3101, LBA4404 | pTF101.1, pTF::Bin19, pTF::UCD2, pTF:ri, pSDM1550, pITC15, pMAW2035HYG | 0–68 | [9,27,28] |
| Barley | AGL0 | pVec8-GFP | 48 | [29] |
| Barrel medic | EHA105 | pSIM843 | 56 | [30] |
| Canola | ABI | pMON67438 | 15 | [31] |
| Creeping bentgrass | EHA101 | pPMI-GFP, pUHVA1, pAHVA1 | 3 | [32] |
| Corn | ABI | pMON92726, pMON65153 | 30–33 | [31,33] |
| Cotton | AGL1 | pPZP-GFP | 31 | [34] |
| Grapevine | LBA4404 | pGA643, pBH710 | 29–50 | [35,36] |
| Maize | EHA101, GV3101, LBA4404 | pTF101.1, pTF::Bin19, pTF::UCD2, pTF:ri | 18–55 | [27] |
| Petunia | LBA4404 | pFLG5972 | 22 | [37] |
| Potato | LBA4404 | pSIM108 | 72 | [38] |
| Rice | LBA4404, AGL1, EHA105 | pCXa21K, pC30063, pGreen/pSOUP, pSK100/200, pEU334NA/NB, pNU393B2, pGA2144 | 4–60 | [39,40,41,42] |
| Sorghum | LBA4404, AGL1 | PHP32269 | 4–26 | [43] |
| Soybean | ABI | pMON83326 | 40 | [31] |
| Strawberry | LBA4404 | pBINPLUS, pGUSINT | 67–90 | [44] |
| Tobacco | LBA4404, GV3101, EHA105 | pBSG-1/BSG-2, pBH710 | 75–80 | [8,36] |
| Tomato | LBA4404 | pBH710 | 67 | [36] |
| Wheat | AGL1 | pCG181-1G+pCS167-1B, pCG185-1G+pCS167-1B, pCG185-2G+pAL154, pCG185-3G+pAL154, pCG185-4G+pAL154 | 8–62 | [45,46] |
| Plant Group | No. of Events | Agrobacterium Strain a | Binary Vector | FS b | VB |
|---|---|---|---|---|---|
| A | 15 | LBA4404 | pPZP-nptII-hemL | 6 (40.0%) | 4 (26.6%) |
| B | 13 | LBA4404 | pPZP-hemL + pZPZ-nptII | 8 (61.5%) | 4 (30.4%) |
| C | 9 | LBA4404 | pPZP-nptII | 4 (44.4%) | 3 (33.3%) |
| D | 9 | AGL1 | pPZP-MsGSAgr | 6 (66.6%) | nt |
| Total | 46 | 24/46 (52.2%) | 11/37 (29.7%) |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nicolia, A.; Ferradini, N.; Veronesi, F.; Rosellini, D. An Insight into T-DNA Integration Events in Medicago sativa. Int. J. Mol. Sci. 2017, 18, 1951. https://doi.org/10.3390/ijms18091951
Nicolia A, Ferradini N, Veronesi F, Rosellini D. An Insight into T-DNA Integration Events in Medicago sativa. International Journal of Molecular Sciences. 2017; 18(9):1951. https://doi.org/10.3390/ijms18091951
Chicago/Turabian StyleNicolia, Alessandro, Nicoletta Ferradini, Fabio Veronesi, and Daniele Rosellini. 2017. "An Insight into T-DNA Integration Events in Medicago sativa" International Journal of Molecular Sciences 18, no. 9: 1951. https://doi.org/10.3390/ijms18091951
APA StyleNicolia, A., Ferradini, N., Veronesi, F., & Rosellini, D. (2017). An Insight into T-DNA Integration Events in Medicago sativa. International Journal of Molecular Sciences, 18(9), 1951. https://doi.org/10.3390/ijms18091951





